Abstract
Fundus fluorescein angiography (FFA) is the gold standard for diagnosing chorioretinal diseases, but its interpretation requires significant expertise and time. Despite generative AI’s enormous potential in medical report generation, automatic FFA interpretation lacks robust models and sufficient evaluation metrics. This study introduces InterpreFFA, a diagnosis-supervised contrastive learning framework, to emulate ophthalmologists’ decision-making process in FFA report generation. Validated on multi-center datasets, InterpreFFA demonstrated superior performance and generalization compared to baseline models. In a simulated clinical setting, two residents used InterpreFFA to diagnose and report FFA cases, with six board-certified ophthalmologists rating the generated reports based on a five-point Likert scale. InterpreFFA significantly improved diagnostic accuracy (85.55 to 90.34%, p < 0.05) and shortened reporting time (153.93 to 108.08 s, p < 0.001). Although AI-generated reports scored slightly lower than manual reports (4.12 vs. 4.38, p < 0.01), InterpreFFA proves to be a promising and cost-effective ancillary tool for enhancing clinical efficiency.
Similar content being viewed by others
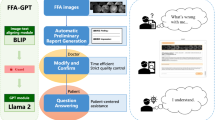
Introduction
Fundus fluorescein angiography (FFA), the widely accepted gold standard for visualizing retinal vasculature, has been crucial in diagnosing various chorioretinal diseases for over 30 years1. Due to the flexibility in the number, location and phase of FFA images when compared to other retinal examinations, FFA provides more dynamic and abundant fundus vascular information, yet is accompanied by a more complex and time-consuming interpretation, which requires significant ophthalmological expertise2. Given the increasing orders in FFA and scarcity of experienced ophthalmologists, particularly in remote or densely populated areas, a reliable and generalizable AI-assisted system for FFA image interpretation is urgently demanded3.
Generative artificial intelligence (AI), foundation models, and large language models (LLMs) are increasingly transforming medical image interpretation and clinical reporting. Generative AI techniques can generate new content including text, images and video, based on learned patterns from existing data4. Recently, advancements in generative AI have significantly expanded its application in medicine, yielding promising results in assisting physicians with interpreting different medical images, enhancing human-in-the-loop clinical decision-making and supporting disease diagnosis and prognosis prediction5,6,7. Studies have demonstrated that generative AI methods can produce chest radiograph reports of similar textual quality to radiologist reports, effectively shorten the reporting time and improve reader performance and efficiency8,9,10. These outcomes underscore generative AI’s potential to support clinicians in clinical image interpretation, which enables streamlining diagnostic workflows and delivering timely and accurate patient care.
Despite the abovementioned advantages, the development of AI-aided FFA interpretation systems faces several challenges. Firstly, the complexity of FFA images and the lack of open-source datasets with corresponding natural language reports cause limited research on automatic FFA interpretation11,12. Secondly, existing studies are also suffering from insufficient model performance and inadequate evaluation metrics10,13. Previous studies mostly focused on improving the literal similarity between generated reports and real-world reports, which are so-called natural language generation (NLG) metrics, instead of the clinical value14. Conventional NLG metrics do not reflect the improvements in clinical settings. To address these problems, Li et al. invited senior ophthalmologists to generally evaluate whether the generated reports can “accurately” describe the images based on several medical questions15,16. Chen et al. extracted keywords from the original FFA reports as classification labels and assessed the accuracy, which partially reflected the clinical efficacy of generated reports17. It is noteworthy that automatic report generation is currently accepted only for ancillary use, rather than for producing instant reports for patients. Despite the recent advances, there is still insufficient evidence on the comprehensive evaluation of the generated reports and the impact of these AI-aided systems in the clinical report-writing process, hindering the reliability and clinical applicability of these systems. Therefore, to further promote the adoption of AI-aided systems in FFA interpretation, it is crucial to validate the clinical efficacy (CE) of generated reports and the association between the assistance from the FFA captioning algorithm and improved efficiency in FFA report drafting8.
To address these pressing clinical needs, we established the InterpreFFA system, a diagnosis-supervised contrastive learning framework designed for FFA report generation. By comprehensively studying the impact of interpreFFA on the report drafting process using our proposed evaluation pipeline, we aim to validate InterpreFFA as a promising and cost-effective tool for enhancing the quality of ophthalmic reporting and supporting clinical decision-making, thereby facilitating the clinical adoption of generative AI models.
Results
We retrospectively collected and included 20052 FFA images and 1833 real-world reports under seven different eye conditions from three data sources: the Second Affiliated Hospital Zhejiang University School of Medicine (ZJU2, internal dataset), Taizhou First People’s Hospital and the Second Affiliated Hospital of Xi’an Jiaotong University (TZ and XJU2, external datasets). The detailed population characteristics and distribution of findings from the used datasets are shown in Table 1. The workflow of the entire study is illustrated in Fig. 1. Firstly, we did the automatic evaluation by comparing InterpreFFA against existing models to validate InterpreFFA’s satisfactory performance in generating FFA reports. Secondly, clinical effectiveness evaluation was conducted by inviting two residents to diagnose FFA cases in the test set and drafted reports with/without the assistance from InterpreFFA. We found that AI-aided FFA interpretation can effectively improve the junior ophthalmologists’ diagnostic accuracy and shorten the reporting time. Thirdly, the ophthalmologists’ ratings indicated an improvement in the quality of FFA reports with the assistance of InterpreFFA, validating the promising ancillary role of our model.
This study includes four major parts: I. Dataset establishment; II. Model development; III. Automatic evaluation; IV. Clinical evaluation. NLG natural language generation, CE clinical efficacy.
Automatic evaluation
By incorporating a novel diagnosis-supervised contrastive loss to generate more clinically accurate reports, InterpreFFA outperformed the other three baseline models and previous studies and achieved satisfactory performance in terms of both NLG metrics and CE metrics. The detailed performance was demonstrated in Table 2. The NLG metrics indicated a high literal similarity between AI-generated reports of InterpreFFA and real-world reports. The CE metrics demonstrated the InterpreFFA’s ability to interpret FFA images, correctly generating key ophthalmic findings in FFA reports with statuses of presence or absence. InterpreFFA excels in identifying key ophthalmic findings, especially in categories of “Normal”, “Aneurysm”, and “Occlusion” (AUC = 0.959, 0.890, and 0.886, respectively). The specific CE metrics of nine ophthalmic findings were demonstrated in Supplementary Table 1. To validate the generalization ability of InterpreFFA, we conducted the report generation test in two external datasets. According to NLG and CE metrics, InterpreFFA demonstrated comparable and satisfactory performance (see Supplementary Fig. 1).
Evaluation of clinical effectiveness
For the total 290 FFA cases in the test set of ZJU2, two residents wrote or modified FFA reports and diagnosed diseases using the report drafting and diagnosis tool (Supplementary Fig. 2). The difference between the normal template-aided mode and AI-aided is whether residents are assisted by the reports generated by InterpreFFA. The diagnostic accuracy for each type of disease is illustrated in Table 3. It is noteworthy that diagnostic accuracy of all types of conditions except Vogt–Koyamagi–Harada (VKH) achieved statistically significant improvements (P < 0.05) when readers wrote reports with the AI-aided mode. The proportion of VKH (35 cases in total) is the lowest across the whole dataset, which may lead to the underfitting problem. Consequently, the corresponding low-quality AI-generated reports may mislead the residents’ diagnosis, as the sensitivity dropped to 14.29% from 71.43% when readers used the AI-aided mode. For example, in one VKH case, the AI-generated report misclassified the presence of choroidal thickening as retinal edema, leading the readers to incorrectly diagnose central serous chorioretinopathy (CSC) instead of VKH. This highlights the potential risks of relying on AI-generated reports in cases of rare diseases.
The reporting time of each FFA case was recorded when the reader finished and pressed the “submit” button on the website. We calculated and compared the average reporting time of all cases when using the two modes. There was a significant reduction in the average reporting time in the AI-aided mode compared to the normal template-aided mode (mean ± SD seconds, 108.08 ± 19.29 vs 153.93 ± 14.67, p < 0.001). The abovementioned results indicated that AI-aided FFA captioning can effectively improve the junior ophthalmologists’ diagnostic accuracy and shorten the reporting time.
Evaluation by ophthalmologists
To further evaluate the quality and clinical accuracy of reports, we invited 6 certified ophthalmologists to rate three types of FFA reports (only AI, normal template-aided and AI-aided reports) based on the five-point Likert scale (see Supplementary Table 2). To examine the within-report-type rating consistency, every report was rated by two physicians and Kendall W for each report type was calculated. Kendall W values for only AI reports, normal template-aided reports and AI-aided reports were 0.628, 0.514, and 0.628, respectively, indicating the moderate consistency within each report type. The averages of each expert’s scores are listed in Table 4. In general, there is still a gap between the AI-generated FFA reports and real-world reports, as average scores of only AI reports (4.12 ± 1.29) are significantly lower than normal template-aided reports (4.38 ± 1.09, P < 0.01) and AI-aided reports (4.46 ± 0.99, P < 0.001). Every rater gave their highest scores to AI-aided reports, indicating that the prior information provided by AI-generated reports is consistent with the findings of residents and associated with improved efficiency in FFA report drafting (Fig. 2 and Table 4). However, statistically significant improvements in the AI-aided group compared to the normal templated-aided group are not seen (Fig. 3). The rating distribution by report types and scores showed that in the AI-aided group, a larger proportion of reports received ratings of 4 and 5 than the other two groups while the only AI group had the highest number of reports rated 3 or lower (Fig. 2 and Supplementary Fig. 3). Additionally, we found that 8.3% of AI-generated reports (rated as 1) need to be completely discarded. 13.1% of AI-generated reports (rated as 2 or 3) require major corrections and 20.9% (rated as 4) were accepted with minimal edits.
a Rating distribution proportion per score; b Rating distribution per report type.
Data presented as mean ± SD, ns means no significant difference, *p < 0.05, **p < 0.01, and ***p < 0.001.
For each report rated 3 or lower, reviewers needed to complete a multiple-choice form to describe the discrepant findings. Of the 281 (16.1%) records rated 3 or lower, 238 (84.7%) were errors in critical findings identification, 23 (8.2%) were errors in critical findings assessment, 80 (28.5%) were errors in non-critical findings identification, 20 (7.1%) were errors in non-critical findings assessment, 41 (14.6%) were errors in non-critical findings omission (Supplementary Fig. 3). All types of errors were least-occurred in AI-aided reports.
Discussion
In this study, we developed a diagnosis-supervised contrastive learning framework for FFA report generation, which outperformed three baseline models and previous studies in both NLG and CE metrics. The generalization ability of the InterpreFFA model was validated in multi-center datasets. Moreover, we invited two residents to write FFA reports with the assistance of AI-generated reports. To study the impact of AI on reader performance, we recorded key metrics such as diagnostic accuracy and the time required for report generation by residents when utilizing the AI-assisted mode. To further investigate the quality of generated reports, we collected and analyzed physician ratings of the reports based on a five-point Likert scale, focusing on accuracy, clarity and clinical relevance. Additionally, the errors in low-quality reports were analyzed. This study highlighted the potential of generative AI as a direct aid in the clinical report-writing process and filled the current gap of no valid clinical evaluation system for FFA report generation tasks.
Previous studies have reported the classification and segmentation approaches to diagnose fundus diseases and detect lesions18,19,20. Currently, the development of LLMs expanded the application of generative AI for medical report generation21,22,23. Compared to the traditional classification model, generative techniques provide more clinically relevant information, such as lesion location, quantity, shape, and severity. For example, vessel occlusion involving the macula area can significantly influence the clinical management and treatment plan for conditions such as BRVO. There have been several advances in the automatic generation of FFA reports. Li et al. and Lin et al. proposed their cross-modal clinical graph transformer models and contrastive pre-training methods for improving the performance in FFA report generation tasks16,24. Chen et al. introduced a pipeline that combines multi-modal transformers and an LLM-based question-answer module to enhance FFA and ICGA report interpretation17,25. These studies improved the quality of generated reports based on traditional NLG metrics and classification metrics and used LLMs to further explain reports to address patients’ questions. However, automatic report generation is currently recognized as a supplementary tool for physicians, as its clinical utility requires further validation. To facilitate clinical adoption, it is essential to assess the real-world impact of this tool on the report-writing process.
To our knowledge, this is the first study to comprehensively evaluate the ancillary ability of AI-generated FFA reports in a simulated clinical setting. By using the report drafting and diagnosis tool, a dramatic reduction in average reporting time and significant improvements in the diagnostic accuracy of most diseases can be seen at the same time when residents used the AI-aided mode with InterpreFFA (Table 3). Moreover, the improvements in sensitivity were not accompanied by large fluctuations in corresponding specificity, indicating that residents can selectively benefit from the true-positive findings present in the AI-generated reports based on their expertise. The moderate interrater agreement observed across the three report types suggests that while some uncertainties remain in the report presentation, the ratings were consistent enough to warrant further investigation. In fact, studies have reported similar or lower interrater agreement in other imaging methods10,26. There was still a quality gap between AI-generated FFA reports and manually generated/modified reports according to the ratings (Fig. 2). Despite their relatively lower quality, only AI reports were shown to modestly enhance resident reports, potentially increasing their sensitivity in detecting certain pathologies (Tables 3, 4). We selected three cases in Fig. 4. For example, in case 1, residents did not document macular involvement when using the normal template-aided mode, which may affect the management of BRVO. But after being alerted by InterpreFFA, residents added the description and generated a perfectly consistent report with the real-world report (Fig. 4a). In case 2, we speculated that residents misrecognized the leakage of microaneurysm in the late phase as neovascularization, which is the symbol of proliferative DR. InterpreFFA helped correct the description and prevented the potential misdiagnosis (Fig. 4b). In case 3, InterpreFFA provided a misleading report. However, we found that residents entirely discarded the AI-generated report and wrote an relatively accurate report instead, which indicates that obvious misleading results from InterpreFFA won’t seriously affect physician judgment due to the clinical expertise (Fig. 4c). The rating results were not significantly improved by comparing AI-aided group to normal template-aided group in this study, which is reasonable because residents have received clinical training and are expected to write relatively accurate reports for our included diseases which have specific pathologic manifestations on FFA examination (Fig. 3). Therefore, there is no much room for AI to improve when it comes to report quality. However, we are delighted to see the slight improvements in the average scores. Together with the improvements in reporting time and diagnostic accuracy, our proposed InterpreFFA model still offers high cost-effectiveness given the tremendous reporting volume and the shortage of ophthalmologists in real-world clinical settings27,28.
a one BRVO case, b one DR case, and c one AMD case. BRVO branch retinal vein occlusion, DR diabetic retinopathy.
Besides the aforementioned results, this study demonstrated several additional advantages. First, we emulate the decision-making process of ophthalmologists by presenting a diagnosis-supervised contrastive learning framework for generating FFA reports. Our InterpreFFA model could lead to better results on both NLG and CE metrics than previous methods. Second, FFA images and reports involved in this study were collected from three hospitals in different regions of China. These hospitals differ in the retina angiograph, imaging techniques, report-writing habits, data storage methods, and disease distributions. The multi-center validation results further mitigated concerns about overfitting, providing evidence of our model’s satisfactory generalization ability in generating FFA reports. Third, our evaluation system incorporated traditional NLG metrics, classification-based CE metrics and clinical quality evaluations. Although it only takes seconds for a pretrained model to generate an FFA report, the comprehensive evaluation of its performance and clinical applicability is the key step for translating this technology from research to practice.
There are several limitations in our study. First, only limited types of eye conditions were included in the datasets, and the diseases in the datasets were unbalanced. The reduction in diagnostic accuracy of VKH indicated that unbalanced data might affect the quality of generated reports, and sparse data from rare diseases can be a serious issue for clinical application of all image-to-text models. Second, although InterpreFFA outperforms other methods on both NLG and CE metrics, the quality of generated reports by our models needs improvement to match that of ophthalmologists. The results of the clinical evaluation indicated that traditional NLG metrics and CE metrics are insufficient to reflect the true quality of reports. Further studies should focus on enhancing report quality by incorporating clinical evaluation metrics that are more closely aligned with ophthalmologists’ standards. Third, although moderate interrater agreement was obtained in this study, there is still some discrepancy in ratings on the 5-point Likert scale. A more specific and standardized rubric for ratings is necessary to reduce discrepancies and improve the reliability of assessments. Third, although we invited human experts to rate AI-generated reports, we didn’t know exactly how residents deal with these reports (e.g., for AI-generated reports with slight errors, one resident may prefer to delete them and rewrite, while another resident may only modify the wrong part, and both practices are acceptable). In other words, the way AI assists physicians cannot be quantitatively compared in this study. Lastly, all real-world reports involved in this study were written in Chinese. In fact, smooth translation in input and output languages can be achieved with minor adjustments to our InterpreFFA model, which is a part of our ongoing research to validate its generalization ability across languages.
In conclusion, we presented a diagnosis-supervised contrastive learning framework designed to emulate ophthalmologists’ decision-making process in FFA report generation. The robust performance and generalization ability were validated in our multi-center datasets in terms of NLG and CE metrics. In a simulated clinical setting, these AI-generated reports improved residents’ diagnostic accuracy and quality of report while significantly shortening the average reporting time. However, hallucinated facts or logic are also observed in rare diseases, which may lead to incorrect diagnosis and need further evaluation. This study introduced a comprehensive evaluation pipeline to assess the effectiveness of AI models in the real-world FFA report-writing process and demonstrated that our model served as a promising and efficient ancillary tool for improving ophthalmologists’ clinical efficiency.
Methods
As illustrated in Fig. 1, this study mainly consists of four parts: (i) we collected and screened FFA cases for dataset establishment; (ii) our generative artificial intelligence model InterpreFFA was developed for FFA report generation; (iii) we evaluated InterpreFFA’s performance against baseline models and previous studies in terms of NLG and CE metrics; (iiii) two residents were invited to diagnose and interpret FFA cases with the assistance from InterpreFFA, their diagnostic accuracy and reporting time were calculated for assessing the clinical effectiveness and six ophthalmologists rated the generated reports for assessing the quality from the clinical perspective.
Dataset establishment
This study used data from three sources: ZJU2 (14684 images and 1453 reports), TZ (3078 images and 173 reports), and XJU2 (2290 images and 207 reports). This study was performed in accordance with the Declaration of Helsinki, and the protocol was obtained from the Ethics Committee of the Second Affiliated Hospital, Zhejiang University School of Medicine (No. Y2023-1073). All patient data were anonymized. The internal datasets consist of FFA cases at ZJU2 from August 2016 to December 2023 and were further divided into training, validation and test sets in a ratio of 3:1:1.
The inclusion criteria for FFA cases were adult patients regardless of gender, eye laterality, and the presence of 1 of the 7 eye conditions: normal, proliferative/non-proliferative diabetic retinopathy (DR), wet/dry age-related macular degeneration (AMD), branch retinal vein occlusion (BRVO), central retinal vein occlusion (CRVO), central serous chorioretinopathy (CSC), Vogt–Koyamagi–Harada (VKH). The FFA images and corresponding free-text reports (each report with 5 to 16 images) were retrieved from the picture archiving and communication system (PACS). For each FFA case, we extracted the temporal sequence of FFA images (each report with 5 to 16 images) from the corresponding PDF reports, where the images were arranged sequentially in a three-image-per-row format. Temporal information of the images was preserved through their file naming. The internal FFA images (JPEG format) were all performed using the tabletop systems HRA-II at 30° (Heidelberg, Germany) with a resolution of 768 × 768 pixels. In contrast, the external FFA images were performed using various systems, resulting in different formats and resolutions. Due to the variability in time points and retinal regions captured in FFA images, significant differences may be observed even between multiple examinations of the same patient. Therefore, in this study, all diagnosable FFA examinations were included. After excluding cases with FFA images of poor quality (images with significant blur issues, high noise quality caused by either pathological or technical issues) or those lacking corresponding free-text reports, two third-year residents (A.S. and W.S.) reviewed the included reports of cases to correct the misspelling and modify/exclude reports with incomplete information under the supervision of an experienced senior ophthalmologist (K.J.).
Model development
We implemented a diagnosis-supervised contrastive learning framework for FFA report generation, named InterpreFFA (Supplementary Fig. 4). The InterpreFFA leveraged a memory-driven Transformer, known as R2Gen, as the backbone, where the relational memory was used to capture information from previous generation processes and a novel layer normalization mechanism was designed to incorporate the memory into the transformer. Notably, R2Gen, one of the most widely used state-of-the-art medical report generation models, demonstrated an exceptional ability to generate long reports with essential medical terms and meaningful image-text attention mappings29.
To generate more clinically accurate text outputs, we propose a novel diagnosis-supervised contrastive loss to R2Gen for FFA report generation. For each FFA case, the extracted images were sequentially arranged by rows to form a large n × 3 input image, replicating the layout of the original PDF report while preserving their temporal order. If the number of images was insufficient, we supplemented it with one or two repeated images from the subsequent captures, where n represents the total number of merged FFA images. Therefore, the input image encompasses all diagnosable FFA images, capturing changes in various retinal regions over time, which enables the model to analyze these temporal changes in the retinal regions.
For the FFA images from each case, we employed the ResNet101 pretrained on ImageNet-1k at resolution 224 × 224 as the visual extractor to extract patch features with a dimension of 204830,31. These patch features were then input into the memory-driven transformer, which comprised an encoder and decoder, each consisting of three layers, eight attention heads, and 512 hidden units, along with relational memory module extensions in the decoder. During the training process, InterpreFFA was trained by minimizing the loss function consisting of two types of losses, which can be displayed as follows:
In Eq. 1, Loss denotes the overall loss of the InterpreFFA model, while \({L}_{\mathrm{CE}}\) and \({L}_{\mathrm{CL}}\) indicate the standard cross-entropy loss used for report generation and a diagnosis-supervised contrastive loss we proposed, respectively. α is a weight parameter that is employed to balance the two losses. The loss function \({L}_{\mathrm{CL}}\) enhances the model by maximizing the cosine similarity between pairs of source images and target sequences, while minimizing the cosine similarity between negative pairs. FFA cases with the same disease label were treated as semantically close, guiding the diagnosis-supervised contrastive learning process during training.
We also compared the InterpreFFA model with 3 baseline medical report generation models, including R2Gen, R2GenRL, and M2KT22,29,32. All the models were trained with one Nvidia GeForce RTX 3090 on the backend framework of PyTorch, using Adam optimizer with initial learning rates of 5e-5 for the visual extractor and 1e-4 for other parameters. The learning rate was decayed by a factor of 0.8 per epoch. The weight α for the diagnosis-supervised contrastive loss was set at 0.2.
Automatic evaluation of InterpreFFA
The performance of the four aforementioned models was automatically evaluated using NLG metrics and CE metrics on the ZJU2 test set and two external test datasets. The NLG metrics from the other three models (FFA-GPT, FFA-IR, and CGT) were directly cited from their published papers for comparison. The NLG metrics include Bilingual Evaluation Understudy (BLEU), Metric for Evaluation of Translation with Explicit Ordering (METEOR), and (Recall-Oriented Understudy for Gisting Evaluation-Longest common subsequence) ROUGE-L33,34,35. BLEU is widely used to evaluate machine translation quality by comparing generated texts with reference texts. It measures the overlap of n-grams (sequences of n words) between the two. In this study, we calculated BLEU1, BLEU2, BLEU3, and BLEU4 as evaluation metrics. METEOR enhances BLEU by considering synonyms and paraphrases, allowing for a more flexible evaluation of generated FFA reports. ROUGE-L emphasizes recall and assesses how well the generated text captures the main ideas of the reference, making it particularly useful for evaluating coherent descriptions of complex clinical information.
We innovatively constructed a keyword dictionary that contains key ophthalmic findings and terms mentioned in the free-text report, along with their synonyms based on ophthalmic knowledge. This dictionary enables the automatic identification and standardization of nine types of pathological terminology in the free-text reports. Notably, the presence of positive and negative expressions was also considered. The classification-based CE metrics were calculated by comparing the statuses of presence or absence between real-world and AI-generated reports. Therefore, unlike NLG metrics, which emphasize the literal similarity between AI-generated reports and real-world reports, potentially influenced by non-informative phrasing, CE metrics focus more on the proximity of clinical contents. The detailed information about the knowledge-based dictionary is in Supplementary Table 3. Sensitivity, specificity, AUC, and F1-score were used to assess model performance for these CE metrics.
Report drafting and diagnostic test
We built an online FFA report drafting and diagnosis tool. The tool mimics viewer functionalities of the real-world PACS, including image switching, zooming, labeling, measurement, and contrast adjustments without displaying other information about the patients. Besides, in the AI-aided mode, an AI-generated report appeared when viewers read the corresponding FFA images. To draft a report, viewers can copy, modify or give up the AI-generated report according to their own observations and knowledge. In the normal template-aided mode, template reports corresponding to the nine conditions are displayed for reference. Afterward, viewers can select one of the eye conditions as a diagnosis. In addition, the tool enables the recording of diagnosis and reporting time. Supplementary Fig. 1 illustrates the different display modes.
Clinical effectiveness evaluation
Two first-year residents from ZJU2, who were unaware of the study hypothesis and not involved in the data collection, participated in the study as independent and blinded readers. For each included FFA case, each participant only diagnosed and reported once, either with assistance from AI-generated reports or using normal templates (e.g., if one resident diagnosed a FFA case with the assistance of AI, the other resident diagnosed the same case using the corresponding normal template). To avoid complications associated with mixed captions of alternate FFA cases with and without AI-generated reports and assure that two participants receive the same amounts of AI-assisted cases, we did not randomly assign the 290 FFA cases in the internal test set to two participants but divided them into five sessions. Each session had a consecutive set of 29 cases without AI assistance and equal amounts of cases with AI assistance. Before each session, participants will review ten separate FFA cases, which were not part of the test set, to get familiar with the reporting and diagnosis tasks. Participants were asked to finish one session in a week. After finishing all the sessions, we evaluated the clinical effectiveness based on the summarized diagnostic accuracy and average time costs.
Clinical evaluation by ophthalmologists
Six board-certified ophthalmologists participated as the raters. For each of the 290 FFA cases, AI-generated reports (Only AI group), normal template-aided reports and AI-aided resident-generated reports were included for rating. A five-point Likert scale was used to demonstrate the degree of agreement or disagreement from raters, that is, the report quality and clinical efficacy. Raters decided the score based on the identification and assessment of the critical and non-critical findings that appeared in the reports. The detailed standard of the Likert scale in this study is shown in Supplementary Table 2. Each rater had access to the ground truth diagnosis of cases and corresponding FFA images but was blinded to the report type information. Each ophthalmologist rated all 290 cases (each case with a single report type) once in an individually randomized order so that each case received two ratings per report type. Besides, for cases rated 3 or lower, raters were instructed to complete a multiple-choice form to describe the reason. The choices include errors in the identification/assessment of critical/non-critical findings and the omission of non-critical findings (Supplementary Fig. 3).
Statistical analysis
The comparison of clinical efficacy between normal templated-aided reports and AI-aided reports was evaluated by diagnostic accuracy, sensitivity, specificity, F1-score, AUC, and report-writing time across all diseases and within disease subgroups. The Wilson score interval method was used to calculate the confidence intervals (CI) for these diagnostic metrics. McNemar’s test was used to evaluate per-disease diagnostic performance, while Bowker’s Test of Symmetry was conducted to assess the overall diagnostic performance. Additionally, two-sample t-tests were applied to compare report-writing times. For Likert score ratings, Kendall W was calculated for each report type, adjusting for tied rankings, to examine within-report-type rating concordance. Likert scores for three types of reports were described as means and standard deviations (SD). ANOVA was performed to test for significant differences in Likert scores between different report types, and post hoc analyses with Tukey’s honest significant difference test were conducted when ANOVA indicated significance, to identify specific differences between report types. All statistical tests were two-sided, with p values less than 0.05 considered statistically significant. All statistical analyses were conducted using R software (version 4.2.1).
Data availability
The datasets used and analysed during the current study are available from the corresponding author on reasonable request.
Code availability
The code is available at https://github.com/liuxc20/InterpreFFA.
References
Cole, E. D., Novais, E. A., Louzada, R. N. & Waheed, N. K. Contemporary retinal imaging techniques in diabetic retinopathy: a review. Clin. Exp. Ophthalmol. 44, 289–299 (2016).
Gao, Z. et al. Automatic interpretation and clinical evaluation for fundus fluorescein angiography images of diabetic retinopathy patients by deep learning. Br. J. Ophthalmol. 107, 1852–1858 (2022).
Resnikoff, S. et al. Estimated number of ophthalmologists worldwide (International Council of Ophthalmology update): will we meet the needs?. Br. J. Ophthalmol. 104, 588–592 (2020).
Howell, M. D., Corrado, G. S. & Desalvo, K. B. Three epochs of artificial intelligence in health care. JAMA 331, 242–244 (2024).
Christensen, M., Vukadinovic, M., Yuan, N. & Ouyang, D. Vision–language foundation model for echocardiogram interpretation. Nat. Med. 30, 1481–1488 (2024).
Wang, X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature 634, 970–978 (2024).
Lu, M. Y. et al. A multimodal generative AI copilot for human pathology. Nature 634, 466–473 (2024).
Ahn, J. S. et al. Association of artificial intelligence-aided chest radiograph interpretation with reader performance and efficiency. JAMA Netw. Open 5, E2229289 (2022).
Zhang, Y. et al. Comparison of chest radiograph captions based on natural language processing vs completed by radiologists. JAMA Netw. Open 6, E2255113 (2023).
Huang, J. et al. Generative artificial intelligence for chest radiograph interpretation in the emergency department. JAMA Netw. Open 6, E2336100 (2023).
Yu, T. et al. A systematic review of advances in AI-assisted analysis of fundus fluorescein angiography (FFA) images: from detection to report generation. Ophthalmol. Therapy 14, 599–619 (2025).
Koohi-Moghadam, M. & Bae, K. T. Generative AI in medical imaging: applications, challenges, and ethics. J. Med. Syst. 47, 94 (2023).
Kaur, N. & Mittal, A. RadioBERT: a deep learning-based system for medical report generation from chest X-ray images using contextual embeddings. J. Biomed. Inform. 135, 104220 (2022).
Ouis, M. Y. & A. Akhloufi, M. Deep learning for report generation on chest X-ray images. Comput. Med. Imaging Graph. 111, 102320 (2024).
Li, M. et al. FFA-IR: towards an explainable and reliable medical report generation benchmark. In Thirty-fifth Conference on Neural Information Processing Systems Datasets and Benchmarks Track (round 2) (2021).
Li, M. et al. Cross-modal clinical graph transformer for ophthalmic report generation. In IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 20624-20633 (IEEE, 2022).
Chen, X. et al. FFA-GPT: an automated pipeline for fundus fluorescein angiography interpretation and question-answer. NPJ Digit. Med. 7, 111 (2024).
Gao, Z. et al. End-to-end diabetic retinopathy grading based on fundus fluorescein angiography images using deep learning. Graefe’s Arch. Clin. Exp. Ophthalmol. 260, 1663–1673 (2022).
Jin, K. et al. Automatic detection of non-perfusion areas in diabetic macular edema from fundus fluorescein angiography for decision making using deep learning. Sci. Rep. 10, 15138 (2020).
Jin, K. & Ye, J. Artificial intelligence and deep learning in ophthalmology: current status and future perspectives. Adv. Ophthalmol. Pract. Res. 2, 100078 (2022).
Fink, M. A. et al. Potential of ChatGPT and GPT-4 for data mining of free-text CT reports on lung cancer. Radiology 308, E231362 (2023).
Yang, S. et al. Radiology report generation with a learned knowledge base and multi-modal alignment. Med. Image Anal. 86, 102798 (2023).
Liu, X. et al. Uncovering language disparity of ChatGPT on retinal vascular disease classification: cross-sectional study. J. Med. Internet Res. 26, E51926 (2024).
Lin, Z. et al. Contrastive pre-training and linear interaction attention-based transformer for universal medical reports generation. J. Biomed. Inform. 138, 104281 (2023).
Chen, X. et al. ICGA-GPT: Report generation and question answering for indocyanine green angiography images. Br. J. Ophthalmol. 108, 1450–1456 (2024).
Hlabangana, L. T. et al. Inter-rater reliability in quality assurance (QA) of pediatric chest X-rays. J. Med. Imaging Radiat. Sci. 52, 427–434 (2021).
Wang, Y. et al. Economic evaluation for medical artificial intelligence: accuracy vs. cost-effectiveness in a diabetic retinopathy screening case. NPJ Digit. Med. 7, 43 (2024).
Resnikoff, S., Felch, W., Gauthier, T. M. & Spivey, B. The number of ophthalmologists in practice and training worldwide: a growing gap despite more than 200 000 practitioners. Br. J. Ophthalmol. 96, 783–787 (2012).
Chen, Z., Song, Y., Chang, T.-H. & Wan, X. Generating radiology reports via memory-driven transformer. In Proc. 2020 Conference on Empirical Methods in Natural Language Processing 1439–1449 (Association for Computational Linguistics, 2020).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Deng, J. et al. ImageNet: a large-scale hierarchical image database. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 248–255 (IEEE, 2009).
Qin, H. & Song, Y. Reinforced cross-modal alignment for radiology report generation. In Findings of the Association for Computational Linguistics:ACL 2022 448–458 (Association for Computational Linguistics, 2022).
Papineni, K., Roukos, S., Ward, T. & Zhu, W.-J. Bleu: a method for automatic evaluation of machine translation. In Proc. 40th Annual Meeting of the Association for Computational Linguistics 311–318. (Association for Computational Linguistics, 2002).
Denkowski, M. & Lavie, A. Meteor 1.3: Automatic Metric for Reliable Optimization and Evaluation of Machine Translation Systems. Proceedings of the Sixth Workshop on Statistical Machine Translation 85–91 (Association for Computational Linguistics, 2011).
Lin, C.-Y. ROUGE: a package for automatic evaluation of summaries. In Text Summarization Branches Out 74–81 (Association for Computational Linguistics, 2004).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (82201195), the National Natural Science Foundation Regional Innovation and Development Joint Fund (U20A20386), Key Program of the National Natural Science Foundation of China (82330032), and Key Research and Development Program of Zhejiang Province (2024C03204). We thank the Second Affiliated Hospital of Xi'an Jiaotong University (Shanxi, China) and Taizhou First People’s Hospital (Zhejiang, China) for their contribution in the dataset construction.
Author information
Authors and Affiliations
Contributions
J.Y., K.J., A.S. and X.L. conceived the study. A.S. and W.S. collected the data. X.L. preprocessed the data and developed the NLG models. A.S. and X.L. designed the methods for model evaluation. Z.Y. designed the report drafting and diagnosis tool. Y.L. and H.W. performed the report drafting and diagnostic test. J.Y., K.J., Y.W., Y.X., T.Z. and X.P. rated the generated reports. X.L. performed the statistical analysis. A.S. and W.S. performed the results analysis. A.S. and X.L. wrote the manuscript. J.Y., K.J., J.W. and Y.C.T. did the critical review. All authors read and approved the final manuscript. A.S., X.L. and W.S. contributed equally to this study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shao, A., Liu, X., Shen, W. et al. Generative artificial intelligence for fundus fluorescein angiography interpretation and human expert evaluation. npj Digit. Med. 8, 396 (2025). https://doi.org/10.1038/s41746-025-01759-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01759-z
This article is cited by
-
Automated Report Generation in Ophthalmology: Integrating Artificial Intelligence, Multimodal Imaging, and Clinical Data
Ophthalmology and Therapy (2026)
-
Leveraging ChatGPT for Report Error Audit: An Accuracy-Driven and Cost-Efficient Solution for Ophthalmic Imaging Reports
Ophthalmology and Therapy (2025)






