Abstract
Clinical insights from real-world data often require aggregating information from institutions to ensure sufficient sample sizes and generalizability. However, patient privacy concerns only limit the sharing of patient-level data, and traditional federated learning algorithms, relying on extensive back-and-forth communications, can be inefficient to implement. We introduce the Collaborative One-shot Lossless Algorithm for Generalized Linear Models (COLA-GLM), a novel federated learning algorithm that supports diverse outcome types via generalized linear models and achieves results identical to a pooled patient-level data analysis (lossless) with only a single round of aggregated data exchange (one-shot). To further protect aggregated institutional data, we developed a secure extension, secure-COLA-GLM, utilizing homomorphic encryption. We demonstrated the effectiveness and lossless property of COLA-GLM through applications to an international influenza cohort and a decentralized U.S. COVID-19 mortality study. COLA-GLM and secure-COLA-GLM offer a scalable, efficient solution for decentralized collaborative learning involving multiple data partners and diverse security requirements.
Similar content being viewed by others
Introduction
The integration of real-world data (RWD), such as electronic health records (EHRs), has significantly influenced healthcare research, as well as clinical and regulatory decision making. This transformation is primarily attributed to the widespread adoption of EHR systems across healthcare organizations worldwide1. To further promote the reuse of this data, distributed research networks, such as the Observational Health Data Sciences and Informatics (OHDSI)2 and the Patient-Centered Clinical Research Network (PCORnet)3, have established standardized data integration frameworks that enable multi-institutional studies, thus improving the generalizability and applicability of research findings. More recently, in response to the dynamic nature of the COVID-19 pandemic and the need to inform public health policies, collaborative networks and research initiatives such as Researching COVID to Enhance Recovery (RECOVER)4, the National COVID Cohort Collaborative (N3C)5, and the Consortium for Clinical Characterization of COVID-19 by EHR (4CE)6 were established as critical infrastructure for clinical evidence generation. In addition, the U.S. Food and Drug Administration’s (FDA’s) Sentinel Initiative and the Biologics Effectiveness and Safety (BEST) Initiative have collaborated with multiple data partners to conduct public health surveillance using common data models7,8. These, and related, initiatives facilitate collaboration among diverse organizations and stakeholders, fostering collective learning by leveraging insights from EHRs across multiple, disparate health systems to inform regulatory decision making.
The sharing of patient-level data across institutions and other entities is often limited by the need to protect patient privacy, even though such data is required for regulatory submissions9 and can offer substantial benefits in multi-institutional studies10,11. For example, PCORnets including STAR, REACHnet, PATH network, ADVANCE collaborative, GPC3, and 4CE6, are decentralized data models where data partners store their data at their institutions. To address the concerns in sharing patient-level data, federated learning algorithms, which partition statistical model estimation into discrete, local computations at each institution before aggregating them at a coordinating center, have been developed and implemented12. In federated learning systems, a coordinating center oversees the aggregation and communication of model updates while ensuring privacy and regulatory compliance across institutions.
Existing federated learning algorithms fall into two main types based on the infrastructure needed for communicating aggregated data. The first reflects algorithms that require iterative updates of aggregated data (e.g., updated estimates of model parameters at each iteration) from all data partners. Examples include the Grid Binary LOgistic REgression (GLORE)13 and WebDISCO14: a web service for distributed Cox model learning without patient-level data sharing, both of which were implemented in pSCANNER12 network, which connects data from over 20 million patients in California. These iterative algorithms can achieve lossless results, producing identical effect size estimates and standard errors compared to analyses based on the pooled patient-level data13,14. However, their implementation typically requires extensive infrastructure to facilitate frequent communication of aggregated data and requires specific institutional agreements among data partners, as in pSCANNER12.
The second type consists of “few-shot” algorithms, which require no more than a few rounds of communications among data partners. With minimal communication needed, these algorithms largely enhance the efficiency of multi-site studies and, at the same time, reduce the need to establish substantial infrastructure to automate the sharing of aggregated data, making them capable to support large-scale collaborations. Notably, the seminal work on surrogate likelihood15,16 for communication-efficient distributed inference has motivated a sequence of few-shot federated learning algorithms for integrating RWD with various types of outcomes, including binary17,18, count19, zero-inflated counts20, and time-to-event outcomes21,22. These algorithms have demonstrated consistent superiority over meta-analysis, particularly in cases when the outcome is relatively rare, or exposure is unbalanced. It makes these algorithms particularly well-suited for pharmacoepidemiologic and pharmacovigilance studies as well as public health surveillance, where rare adverse events are of primary interest. In addition, the robust performance of these algorithms in these challenging scenarios supports the evaluation of heterogeneity across underrepresented subpopulations—an essential goal in precision medicine research23.
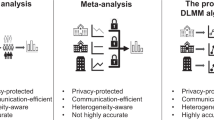
Still, each type of federated learning algorithms has its limitations. While iterative algorithms are capable of achieving lossless results, their implementation relies on (1) established multi-institutional data use agreements allowing automated sharing of aggregated data and (2) the pre-established secure computing infrastructure for data exchange across all participating partners. Both conditions create burdens for scaling these networks. In contrast, few-shot algorithms are easy to implement and scale, but their results are generally not lossless and can be sensitive to the initialization process, potentially yielding different findings if different initial values are chosen21. More importantly, both few-shot algorithms with two or more iterations and iterative algorithms can encounter synchronization issues, as updates to model parameters must wait until all data partners submit their results from the previous rounds24. A delayed update from one site holds up the entire process. These synchronization issues induce significant challenges to the scalability of the multi-shot frameworks (i.e., methods requiring multiple rounds of communications) when applied to a large number of data partners.
Ideally, federated algorithms should be devised to be lossless, one-shot, and not dependent on initializations, ensuring all these features are met. Yet, to date, only a select few federated learning algorithms achieve both lossless and one-shot properties, save for those focused on linear regression25 and linear mixed models26. The main challenge in designing algorithms that realize both properties lies in solving the non-linear estimating functions of regression models, which typically require multiple iterations.
To fill this gap, in this paper, we introduce a novel class of federated learning algorithms for fitting generalized linear models (GLMs) using multi-site data, named Collaborative One-shot Lossless Algorithms (COLA-GLM). These algorithms are provably lossless, one-shot, and do not require initializations. They eliminate the need for infrastructure to achieve automated updates of estimates and are scalable to a large number of data partners. COLA-GLM enables collaborative modeling of a broad range of clinical outcomes in the exponential family, including binary, categorical, and count outcomes. To further minimize the exposure of aggregated data from individual sites, we adopt fully homomorphic encryption to protect the aggregated data from participating sites, and developed the secure-COLA-GLM algorithm. Homomorphic encryption is a cryptographic technique that allows computations to be performed directly on encrypted data, producing results that, when decrypted, are identical to those obtained from operations on the original data. While encryption-based federated learning algorithms have been extensively studied in the broader literature, our work presents a novel contribution by focusing on its practical application in healthcare and integrating it within the COLA-GLM framework. We employ fully homomorphic encryption to protect aggregated data, offering greater computational and communication efficiency than methods that encrypt individual-level data. This design is particularly well-suited for real-world healthcare settings, where datasets have large sample sizes and low-to-moderate covariate dimensionality.
We empirically validate the properties of COLA-GLM by revisiting an international study launched by the OHDSI research network27, covering more than 3 million patients across six databases, which focused on collaboratively developing COVID-19 prediction models using influenza data during the early pandemic. Since patient-level data from all six databases were accessible for this evaluation, we could thoroughly evaluate the algorithm’s performance. Additionally, we demonstrate the practical applicability of COLA-GLM using four decentralized databases of clinical information in the U.S. to identify risk factors of COVID-19 mortality among hospitalized patients.
Results
The COLA-GLM and secure-COLA-GLM
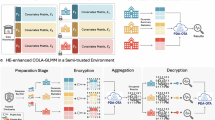
Figure 1 illustrates the workflows of the proposed COLA-GLM and secure-COLA-GLM. COLA-GLM enables collaborative modeling using generalized linear models (GLMs), requiring only aggregated data from each site, not individual patient-level data. It operates with a single round of communication of aggregated data, one-shot property, from the participating sites to the coordinating center, a third party operating the Privacy-Preserving Distributed Algorithms (PDA) framework28, referred to as PDA. Meanwhile, COLA-GLM achieves results identical to pooled analysis where patient-level data can be shared across sites, referred to as lossless property. Figure 2 highlights the aggregated data required for COLA-GLM and demonstrates the lossless property.
a The general workflow of the COLA-GLM, where each site generates raw aggregated data and shares it with the coordinating center, a third party running Privacy-Preserving Distributed Algorithms framework28, hereafter referred to as PDA. b Outlines the workflow of the secure-COLA-GLM, which involves four key steps: (1) a trusted third party or one of the participating sites generates a private key for each site and a public key; (2) each site computes their aggregated data, encrypts it using public key, and shares the encrypted aggregated data with PDA; (3) PDA reconstructs the multi-site data likelihood using the encrypted aggregated data, obtains the results, and broadcasts them to participating sites; and (4) each site decrypts the results using its private keys and sends the decrypted results back to PDA to consistency check.
a The workflow of the proposed COLA-GLM and the aggregated data (AD) that need to be shared in an illustrative example involving three hospitals. In this workflow, each hospital shares its aggregated data (\({S}_{j},{U}_{j}\)) with the coordinating center (e.g., 28PDA). The coordinating center then reconstructs the multi-site data likelihood function using these aggregated data based on Eq. (2) and obtains estimates of coefficients by optimizing the reconstructed multi-site data likelihood function. b The workflow of the pooled analysis using the same example, where patient-level data from all three hospitals are directly shared. Notably, COLA-GLM yields identical estimations of coefficients by only sharing summary-level information (i.e., aggregated data).
To further minimize the exposure of aggregated data from participating sites, secure-COLA-GLM employs fully homomorphic encryption, ensuring only encrypted aggregated data are shared with the coordinating center. Figure 1b outlines the four key steps in secure-COLA-GLM: (1) key generation and distribution, (2) encryption and communication of aggregated data, (3) homomorphic operations by the coordinating center, and (4) decryption and consistency checks.
A detailed explanation of the techniques embedded in both COLA-GLM and secure-COLA-GLM is provided in the Methods section.
Empirical Validation of COLA-GLM Using an International Cohort of Influenza
We demonstrate the utility and empirically validate properties of COLA-GLM using an international network of multiple databases on patients with influenza. During the COVID-19 pandemic, understanding the risk factors for severe COVID-19 is crucial for clinicians to identify high-risk patients who require prioritized treatment. At the onset of the pandemic, the COVID-19 data were limited, making historical data from similar respiratory diseases, such as influenza, a valuable proxy for early investigations27. Even today, influenza remains a significant global health burden, providing important insights that can inform future pandemic responses.
In this analysis, we focused on identifying risk factors for developing severe infections using historical data from patients with influenza. EHR and medical claims data were collected from six data sources (both within and outside of the United States, including IBM MarketScan Commercial Database (CCAE), IBM MarketScan® Medicare Supplemental Database (MDCR), IBM MarketScan® Multi‐State Medicaid Database (MDCD), Japan Medical Data Center (JMDC), Optum® de-identified Electronic Health Record Dataset (Optum® EHR), and Optum’s de-identified Clinformatics® Data Mart Database (Clinformatics®). All collected data were standardized to the Observational Medical Outcomes Partnership Common Data Model (OMOP-CDM)29. A detailed summary of these data sources is provided in Table 1 and we refer to the original paper27 for details of variable prevalence. Since patient-level data from all six databases were accessible, we conducted a pooled analysis, maximizing the global likelihood function across the six databases, as a gold standard baseline to evaluate the properties of COLA-GLM.
The study cohort consisted of patients aged 18 or older who presented with influenza or flu-like symptoms during a healthcare-provider interaction before early 2020, including fever, cough, shortness of breath, myalgia, malaise, and fatigue. Patients with fewer than 365 days of prior observation time and no symptoms in the preceding 60 days were excluded. The study cohort included 3,534,245 patients from all six international databases, including 1,000,000 from CCAE, 92,184 from MDCR, 378,900 MDCD, 311,870 from JMDC, 751,291 from Optum® EHR, and 1,000,000 from Clinformatics®. The index date was defined as the date of the initial healthcare provider interaction. The outcomes of interest are three binary variables of severe infections, including: (1) Death within 30 days after the index date; (2) Hospitalization with pneumonia within 30 days after the index date; (3) Hospitalization with pneumonia requiring critical care services (i.e., ICU) or death following hospitalization with pneumonia within 30 days after the index date. To analyze the risk of developing severe infections, we dichotomized age as less than 60 and 60 or older, and adjusted for sex and seven clinical factors: (1) history of cancer, (2) history of chronic obstructive pulmonary disease (COPD), (3) history of diabetes, (4) history of heart disease, (5) history of hypertension, (6) history of hyperlipidemia, and (7) history of kidney disease27.
Figure 3 presents the results of the pooled analysis and COLA-GLM in quantifying patients’ risk for severe infection outcomes using logistic regression models. Specifically, the estimated effect sizes (i.e., log odds ratio) and their standard errors align closely between the pooled analysis (x-axis) and COLA-GLM (y-axis) for all three outcomes and all covariates. For instance, among all the effect sizes and standard errors compared in Fig. 3, the largest difference between the pooled analysis and the COLA-GLM is 0.00013 for the history of COPD. The mean and median of the differences for all predictors are -3.09 \(\times {10}^{-6}\) and 1.20\(\times {10}^{-6}\). The small discrepancies in the results are an artifact of the limit of the machine’s precision in computing.
The x-axis shows the results from the pooled analysis (combining patient-level data from six data sources) and the y-axis shows the results for COLA-GLM. Each dot represents the estimated effect size or standard error for a specific covariate.
We investigate the sensitivity of COLA-GLM to threshold values recommended by OHDSI (a threshold of 5) and by Centers for Medicare & Medicaid Services (CMS) and PEDSnet (a value of 11)30,31. Cells with fewer than 5 or 11 observations were imputed with values of 3 or 6, respectively. A cell value of zero was reported as zero, as it does not violate these policies.
Figure 4 displays the results for COLA-GLM with a minimum cell size of 5 (Panel a) and 11 (Panel b) compared to the pooled analysis. The results from COLA-GLM align reasonably closely with the pooled analysis in these scenarios, with differences increasing slightly when a larger minimum cell size is used (e.g., 11). When using a minimum cell size of 5, the largest difference between the pooled analysis and COLA-GLM regarding the estimation of effect sizes is 0.011 for the intercept, with a mean difference of 5.21 \(\times {10}^{-5}\) and median difference of 0.0007 across all covariates. When using a minimum cell size of 11, the largest difference between the pooled analysis and COLA-GLM regarding the effect size estimates is 0.031 for the intercept, with a mean difference of 0.0005 and a median difference of 0.0009.
a Results using a minimum cell size of 5 (reported as 3). b Results using a minimum reporting cell size to of 11 (reported as 6).
Implementation of COLA-GLM on U.S. COVID-19 Mortality data
We further demonstrate the applicability of COLA-GLM in a truly decentralized network, where the patient-level data cannot be shared across sites, to identify the risk factors of COVID-19 mortality among hospitalized patients. Since the data are decentralized, a pooled analysis is not feasible in this study. Data partners for this study contributed EHR data from four databases within the OHDSI network: Optum® de-identified Electronic Health Record Dataset (Optum® EHR), IQVIA Hospital CDM Electronic Medical Record Database (IQVIA Hospital CDM), University of Florida Health Electronic Health Record Database (UF Health), and Columbia University Irving Medical Center (CUIMC) EHR Database, all standardized to the OMOP CDM. A detailed summary of these databases is provided in Table 2.
Our study focused on the period when the Omicron variant was predominant, from November 2021 to February 2022. The study cohort included patients aged 18 or older who had an inpatient visit with a diagnosis of COVID-19 or a positive COVID-19 test within 21 days prior to the visit. Patients with fewer than 365 days of prior observation were excluded. The study cohort consisted of 45,524 patients from all four databases. The outcome of interest was a binary mortality status, indicating whether the patient died during the inpatient visit or within 7 days afterward. The covariates included three age categories ( <65, 65-80, and ≥80), gender (male or female), and four clinical characteristics: (1) history of COPD, (2) history of diabetes, (3) history of hypertension, (4) history of kidney disease. We also included the Charlson Comorbidity Index (CCI) score as a measure of the patient’s overall health state, with higher scores indicating worse health conditions. Detailed characteristics of the study cohort are provided in the Supplementary Table 8.
We fit a logistic regression model using COLA-GLM. Following the policy of OHDSI, all cells with values smaller than 5 were reported as 3. Figure 5 displays the forest plot of the odds ratios (OR) estimated from COLA-GLM. The findings identify covariates significantly associated with higher risks of mortality, including age between 65 to 80 with an OR of 2.40 (95% CI 2.06-2.31), age above 80 with an OR of 2.72 (95% CI 2.46-2.88), being male with an OR of 1.33 (95% CI 1.24-1.42), a history of diabetes with an OR of 1.47 (95% CI 1.36-1.60), and a history of hypertension with an OR of 1.15 (95% CI 1.06-1.24). These findings are consistent with the existing literature, where age32, gender33, diabetes34 and hypertension35 were shown to be statistically significantly associated with mortality in COVID-19 patients. These insights enhance our understanding of the factors that elevate the severity of COVID-19 outcomes which helps inform clinical decision-making, guide public health policies, and improve targeted interventions to better manage and protect high-risk populations.
All cells with values smaller than 5 were imputed as 3 when sharing aggregated data from participating databases to PDA.
Discussion
This paper introduced and empirically evaluated COLA-GLM, a novel privacy-preserving distributed learning algorithm for analyzing diverse outcome types by fitting GLMs. COLA-GLM contributes to a communication-efficient, privacy-preserving framework for fitting GLMs using only a single round of communication of aggregated data, which achieves identical results to those from pooled data analysis. To address the potential risk of re-identification from sharing empirical distribution of categorical covariates, COLA-GLM incorporates a cell size suppression policy recommended by various US states and federal agencies. Specifically, any count below a predetermined threshold is substituted with a standardized value. The empirical evaluation of COLA-GLM in an international cohort of influenza data, where pooled data are accessible, demonstrated this lossless property. The computational and communication efficiency of COLA-GLM makes it highly scalable for distributed learning, especially with a large number of data contributors. The calculation of aggregated data at each individual site is implemented in the R package “pda”. We also developed an “over-the-air” online portal called PDA-OTA (http://pda-ota.pdamethods.org/) to facilitate secure and convenient collaboration on the basis of the “pda” package. Detailed instructions for using the PDA-OTA are provided in the Supplementary Materials.
COLA-GLM is proposed as a general federated learning framework capable of fitting GLMs with both canonical and non-canonical link functions, accommodating a wide range of outcome types. We consider our COLA-GLM algorithm privacy-preserving, as it requires only one-shot communication of aggregated data from participating sites, and these aggregates are shared exclusively among collaborators involved in the study. However, we acknowledge that our current data release mechanism has not been formally analyzed under rigorous privacy frameworks such as differential privacy36. Quantifying the potential risk of information leakage, such as establishing differential privacy guarantees, remains an important direction for future work.
COLA-GLM works with categorical covariates, which are commonly used in clinical research due to its practical advantages, such as alignment with administrative coding standards (e.g., ICD codes), ease of interpretation for clinical stakeholders, and privacy considerations (such as k-anonymity or minimum cell count suppression). Our empirical evaluation using influenza data indicates that imposing a minimum group size of 5 or 11 has a negligible impact on the algorithm’s performance when nine covariates are included in the model. Additional evaluations of accuracy across varying sample sizes and covariate dimensions are provided in Supplementary Results 3. In practice, categorizing continuous variables presents a trade-off between granularity, interpretability, and privacy protection. The number and boundaries of categories should be determined by balancing clinical interpretation, statistical power, and privacy constraints. As a general guideline, we recommend starting with quantile-based binning (such as tertiles or quartiles) or adopting clinically meaningful thresholds, with subsequent refinement based on empirical distributions and applicable data suppression thresholds.
In practice, we could encounter a semi-trusted environment where data contributors may have concerns about fully trusting the coordinating center. To address these concerns, we developed secure-COLA-GLM based on homomorphic encryption to minimize the exposure risk, where computations at the coordinating center are performed directly on encrypted data. Secure-COLA-GLM requires one additional round of communication compared to COLA-GLM, as well as extra computational time for data encryption and decryption. The implementation details of secure-COLA-GLM on the U.S. COVID-19 mortality data, along with the evaluation of its computational and communication overhead, are summarized in Supplementary Results 1. For a model with eight binary variables, encryption of the aggregated data takes only a few seconds on a system equipped with an AMD Ryzen 9 7900X (4.70 GHz, 12-core processor) and 64 GB of RAM. Despite these additional steps, secure-COLA-GLM ensures that sensitive information remains secure throughout the data-sharing and processing phases. Furthermore, it eliminates the need for minimum cell size policies for data sharing and allows for analyses with more variables and finer categorical distinctions. This capability is particularly important in pharmacoepidemiologic and pharmacovigilance studies of rare diseases, as well as in precision medicine research aimed at evaluating heterogeneity across subpopulations. Although model parameters estimated by secure-COLA-GLM show minor deviations from those produced by COLA-GLM in this real-data application, implementing homomorphic encryption in the COLA-GLM framework significantly enhances data privacy, allowing the utility of COLA-GLM in a wider range of scenarios that demand stronger privacy guarantees.
COLA-GLM is distinct from existing federated learning approaches for fitting GLMs. For instance, the GLORE framework by Wu et al.13, its online iteration WebGLORE by Jiang et al.37, and the techniques by Shu et al.38 necessitate iterative communication of aggregated information from participating sites. Additionally, the few-shot algorithms by Duan et al.18 and Edmondson et al.19 demand access to individual patient-level data at a central site. In contrast, COLA-GLM requires only aggregated data from all participating sites. It reconstructs a multi-site data likelihood that is identical to the pooled data analysis and naturally accommodates covariate distribution heterogeneity across sites. For addressing between-site heterogeneity in effect sizes, a generalized linear mixed model (GLMM) approach can be employed. Research on lossless, one-shot federated algorithms for GLMM estimation is ongoing and will be reported in the future.
This novel privacy-preserving algorithm COLA-GLM has the potential to make regulatory impact. The US FDA’s Sentinel initiative and BEST initiative both involve multiple data partners, and meta-analyses were typically used to draw a single conclusion to answer an important regulatory question. The algorithm COLA-GLM would allow the FDA and other agencies to analyze the aggregated data from data partners and obtain the same results as if analyzing pooled patient-level data from all data partners. In addition, access issue of patient-level data has been a concern in real-world evidence (RWE) regulatory submissions, especially when certain RWD are owned and controlled by entities other than the sponsors. This novel privacy-preserving algorithm COLA-GLM has the potential to enable the FDA and other regulatory agencies to perform independent review of the RWE generated from RWD with aggregated data.
In summary, COLA-GLM is lossless, requires only one round of communication of aggregated data for analyzing diverse outcome types using GLMs. It is highly suitable for collaborative analyses with a large number of data partners. It also addresses the patient-level data access issue for regulatory submissions and has the potential to make regulatory impact. The secure-COLA-GLM with homomorphic encryption provides additional privacy protection, extending the utility of COLA-GLM to a wider range of decentralized collaborative learning scenarios with varying security needs.
Methods
The proposed COLA-GLM
Generalized linear models (GLMs) refer to a broad class of models39 that have become a standard methodological approach in numerous statistical applications. GLMs extend the classic linear model by accommodating non-normal responses and allowing for a non-linear relationship between the response and covariates. For example, logistic regression models are frequently used in biomedical science to model binary outcomes, while Poisson regression models are commonly employed to analyze count data.
We consider a scenario to estimate a GLM using data from multiple sites. Assume there are \(J\) sites, with \({n}_{j}\) observations available at the \(j\)-th site, and a total sample size of \(N=\mathop{\sum }\nolimits_{j=1}^{J}{n}_{j}\). For the \(i\)-th subject at the \(j\)-th site, let \({y}_{{ij}}\) denote the response and \({{\boldsymbol{x}}}_{{ij}}\in {{\mathbb{R}}}^{d}\) be a vector of covariates. In GLMs, the response \({y}_{{ij}}\), conditional on covariates \({{\boldsymbol{x}}}_{{ij}}\), is assumed to follow an exponential family distribution independently. Specifically, a link function \({g}\left(\cdot \right)\) is pre-selected to describe the relationship between the conditional mean of response \({\mu }_{{ij}}{\mathbb{=}}{\mathbb{E}}[{y}_{{ij}}|{{\boldsymbol{x}}}_{{ij}}]\) and the linear predictor \({\eta }_{{ij}}={{\boldsymbol{x}}}_{{ij}}^{T}{\boldsymbol{\beta }}\), such that \(g({\mu }_{{ij}})={{\boldsymbol{x}}}_{{ij}}^{T}{\boldsymbol{\beta }}\). The canonical link function is the most commonly used in GLMs. It directly relates the linear predictor to the canonical parameter of the exponential family distribution, which simplifies the likelihood equations and estimation procedures. The canonical link function covers a wide range of statistical models with examples, as outlined in Supplementary Table 1.
With the canonical link function, the multi-site data log-likelihood of GLM can be expressed as
where \(b\left(\cdot \right)\) is the cumulant function depending on the exponential family distribution we specified. Standard estimation of parameters in GLMs can be based on maximizing the log-likelihood. However, unlike in the ordinary linear model25,26, the estimation of GLMs in a distributed setting is non-trivial due to the fact that the computation cannot be distributed to each site in a lossless fashion.
However, by rewriting the multi-site data likelihood as shown in Eq. (2), we observe that, in the scenario of categorical covariates, the multi-site data likelihood of a GLM can be exactly reconstructed by requiring each site to contribute some aggregated data only once.
In the first term of the multi-site data likelihood in Eq. (2), the data \(\{{\boldsymbol{x}}_{{ij}},{y}_{{ij}}\}\) are inherently separated from the model parameters \({\boldsymbol{\beta }}\). Therefore, reconstructing the first term of the multi-site data likelihood only requires each site to share the aggregated data \(\mathop{\sum }\nolimits_{i=1}^{{n}_{j}}{y}_{{ij}}{{\boldsymbol{x}}}_{{ij}}\). The second term in Eq. (2) contains \({{\boldsymbol{x}}}_{{ij}}^{T}{\boldsymbol{\beta }}\) inside the cumulant function \(b\left(\cdot \right)\), whose form depends on the exponential family specified in GLMs. This term generally does not allow us to separate the data \(\{{\boldsymbol{x}}_{{ij}}\}\) from model parameters \({\boldsymbol{\beta }}\). However, when covariates are categorical, we can reconstruct this term by sharing the joint empirical distribution of \({\boldsymbol{x}}\) at each site. The joint empirical distribution of \({\boldsymbol{x}}\) is defined by counting occurrences of all possible covariate values (\({\boldsymbol{x}}\)) at the \(j\)-th site. Note that communicating sufficient statistics \(\mathop{\sum }\nolimits_{i=1}^{{n}_{j}}{y}_{{ij}}{{\boldsymbol{x}}}_{{ij}}\) alone is not enough to reconstruct the data likelihood, which corresponds to information of model parameters under the scenario that the covariates are given. Thus, in practice, since covariates for each site are unknown to the coordinating center, communicating their empirical distribution is necessary to reconstruct the multi-site data likelihood.
This interesting observation motivated our development of the Collaborative One-shot Lossless Algorithm for GLMs (COLA-GLM). The COLA-GLM requires only a single round of communication of aggregated data from each site to recover the multi-site data likelihood, producing identical results to a pooled analysis that would require individual patient-level data from all sites. Figure 1a illustrates the general workflow of COLA-GLM. Figure 2a provides an example of how aggregated data is shared across sites. Figure 2b presents the workflow of pooled analysis using the same example, further demonstrating the lossless property of COLA-GLM. Let \({{\boldsymbol{x}}}^{1},\ldots ,{{\boldsymbol{x}}}^{m}\) denote all possible values of the covariate vector. To reconstruct the multi-site data likelihood, COLA-GLM requires each site \(j\) (\(j=1,\ldots ,J\)) to contribute the following aggregated data:
-
a \(d\)-dimensional vector \({{\boldsymbol{S}}}_{j}={{\boldsymbol{X}}}_{j}^{T}{{\boldsymbol{y}}}_{j}\)
-
and an \(m\times (d+1)\) matrix \({{\boldsymbol{U}}}_{j}\) counting the number of each possible value of the vector covariate \({{\boldsymbol{X}}}_{j}\):
where \({u}_{j}({{\boldsymbol{x}}}^{l})\) is the number of patients associated with covariate pattern\(\,{{\boldsymbol{x}}}^{l}\) at site \(j\).
The \(j\)-th site shares the summary-level data (\({S}_{j},{U}_{j}\)). The coordinating center, a third party operating the Privacy-Preserving Distributed Algorithms (PDA) framework28, referred to as PDA hereafter, reconstructs the multi-site data likelihood \({{\ell}}\left({\boldsymbol{\beta }}\right)\) using these summary-level data from participating sites. COLA-GLM then obtains the parameter estimate \(\hat{\beta }\) by maximizing the reconstructed log-likelihood.
Improving modeling flexibility by using non-canonical links
While the canonical link function is often the natural choice due to its mathematical properties, non-canonical link functions could be useful to offer more interpretable relationships between predictors and the response, and provide a better fit to the data, particularly in cases of non-normal response distributions, restricted ranges, or address issues such as overdispersion. Examples include Probit regression model40 for binary data and quasi-Poisson regression41 modeling count outcome with overdispersion. When using non-canonical link functions, the proposed COLA-GLM requires each site \(j\) (\(j=1,\ldots ,J\)) to communicate the following aggregated data:
-
an \(m\times (d+1)\) matrix \({{\boldsymbol{U}}}_{j}\) counting the number of each possible value of the vector covariate \({{\boldsymbol{X}}}_{j}\):
$${{\boldsymbol{U}}}_{j}=\left[\begin{array}{cc}{{\boldsymbol{x}}}^{1} & {u}_{j}\left({{\boldsymbol{x}}}^{1}\right)\\ \vdots & \vdots \\ {{\boldsymbol{x}}}^{m} & {u}_{j}\left({{\boldsymbol{x}}}^{m}\right)\end{array}\right]$$(4)where \({u}_{j}({{\boldsymbol{x}}}^{l})\) is the number of patients associated with covariate pattern \({{\boldsymbol{x}}}^{l}\) in site \(j\),
-
and an \(m\times (d+1)\) matrix \({{\boldsymbol{W}}}_{j}\) counting the occurrence of outcomes for each possible value of the vector covariate \({{\boldsymbol{X}}}_{j}\):
where \({w}_{j}({{\boldsymbol{x}}}^{l})\) is the number of outcomes associated with covariate pattern \({{\boldsymbol{x}}}^{l}\) at site \(j\).
Maintaining cell suppression policy
COLA-GLM collects the aggregated data from each local site once, including counts of each possible value of the covariate vector, which may be subject to a risk of patient re-identification when the cell counts are small42. To mitigate this risk, cell suppression policies are recommended by various US states and federal agencies30,31,43,44 when sharing data. These policies require that cells containing non-zero counts below a certain threshold be suppressed. For example30, a cell size suppression policy with a threshold 11 specifies that any cell with a count between 1 and 10 cannot be reported directly. Specifically, OHDSI recommends a minimum reporting cell size of 5, while the Centers for Medicare & Medicaid Services (CMS) and PEDSnet require that no cell containing values fewer than 11 be reported30,31. The proposed COLA-GLM comply with such cell suppression policy by replacing counts below the threshold with a standard value. The sensitivity of COLA-GLM in terms of threshold values are evaluated in the Results section.
Secure-COLA-GLM with homomorphic encryption
COLA-GLM operates in an environment where the coordinating center (e.g., a third party running PDA) being responsible for implementing COLA-GLM, is entrusted with collecting aggregated data from local sites, as illustrated in Fig. 1a. Such an assumption is reasonable because, in practice, the activities of all participating parties would typically be governed by contractual agreements, such as data use agreements, that forbid unauthorized data sharing and tampering. Established collaborative networks like OHDSI2, PCORnet3, and Sentinel42 already adhere to similar frameworks, demonstrating the viability of such a cooperative environment.
However, in practice, the data contributors may not fully trust the coordinating center, creating what is referred to as a semi-trusted environment. While all parties are expected to adhere to the protocol and avoid malicious actions, contributors may remain concerned that others may try to infer information from data they observe passively45,46,47. In COLA-GLM, for example, the patient-level covariate information at site \(j\) could potentially be inferred from the shared empirical distribution of covariates, represented by the matrix \({{\boldsymbol{U}}}_{j}\). To ensure that the coordinating center cannot infer patient-level information in a semi-trusted environment, we employ a fully homomorphic encryption scheme and propose a secure extension of COLA-GLM, named secure-COLA-GLM. Fully homomorphic encryption has been extensively adopted in privacy-preserving federated learning due to its ability to perform computations directly on encrypted data46,48,49,50,51,52. For the development of secure-COLA-GLM, we specifically utilize the CKKS53 which is well-suited for approximate arithmetic operations in encrypted domains. In this algorithm, only encrypted aggregated data are communicated to the coordinating center, PDA, to achieve identical results without accessing the raw aggregated data from local sites. The workflow of secure-COLA-GLM is illustrated in Fig. 1b. We refer to Supplementary Table 2 for a detailed walkthrough of CKKS.
Secure-COLA-GLM requires a trusted third party to generate encryption key pairs. This third party must operate independently of the coordinating center and could potentially be a selected leader among the participating data partners or an authenticated organization. For example, one of the participating hospitals can be selected to serve as the trusted third party to generate and distribute key pairs to the other participating hospitals. During this process, the trusted third party will generate a public key, which will be distributed to by all participating sites and the PDA, and a private key for each site, which is accessible only by the respective site and the trusted third party. The primary steps of secure-COLA-GLM are as follows:
-
Step 1 (Key generation and distribution): The trusted third party, e.g., one of the participating sites, generates a private key for each site and a public key and distributes them to all participating sites via a secure information channel. The public key is also shared with the coordinating center PDA.
-
Step 2 (Encryption and communication of aggregated data): After receiving the key pairs, each participating site generates the raw aggregated data using local patient-level data, encrypts the aggregated data using the public key, and communicates the encrypted aggregated data to the coordinating center, PDA.
-
Step 3 (Homomorphic operations by PDA): PDA, who also has access to the public key, employs homomorphic operations on the encrypted aggregated data to reconstruct the multi-site data likelihood and produce encrypted model estimations. By using a common public key across the participating sites and the PDA, the homomorphic properties of the encryption function ensure that the operations performed on the encrypted aggregated data yield identical results to those on the original aggregated data once decrypted.
-
Step 4 (Decryption and consistency check): The encrypted estimation results from PDA are shared with participating sites. Each site decrypts the results using its private key to obtain the original, unencrypted results. These decrypted results, which should be consistent across all sites, are then sent back to the coordinating center for a final consistency check.
The design of independence between the trusted third party and the coordinating center, PDA, in secure-COLA-GLM ensures robust privacy protection. While the PDA has access to the public key for performing homomorphic operations, it does not have any access to any private keys for decryptions. With this infrastructure, the secure-COLA-GLM requires two rounds of communication. One round of communication is defined as one instance of transferring information to PDA. Specifically, in a semi-trusted environment, the PDA is involved twice: (1) receiving encrypted aggregated data and producing encrypted results, and (2) receiving decrypted results from participating sites and checking their consistency. By contrast, COLA-GLM involves the PDA only once, for collecting the raw aggregated data and producing the final results.
Though secure-COLA-GLM requires one additional round of communication compared to COLA-GLM, the single additional round of communication may be seen as burdensome if it can eliminate concerns about the trustworthiness of the coordinating center. Additionally, because of the one-shot design of COLA-GLM, the additional communication overhead for the secure version is minimal, making the algorithm highly scalable to a large number of data partners. Moreover, the consistency check in Step 4 is essential to verify that the decryption process, performed independently at each site, was carried out correctly and without error. It also serves to detect any discrepancies that may arise from implementation issues or misalignment in key management, thereby ensuring the correctness of the overall process.
Data availability
The data are not publicly available due to privacy concerns. The individual de-identified participant data will not be shared. The data that support the findings of this study may be available through request and DUA process to the corresponding authors.
Code availability
The code used to implement the methods in this study is publicly available on GitHub at https://github.com/Penncil/pda.
References
Friedman, C. P., Wong, A. K. & Blumenthal, D. Achieving a nationwide learning health system. Sci. Transl. Med. 2, 57cm29 (2010).
Hripcsak, G. et al. Observational health data sciences and informatics (OHDSI): opportunities for observational researchers. Stud. Health Technol. Inf. 216, 574–578 (2015).
Fleurence, R. L. et al. Launching PCORnet, a national patient-centered clinical research network. J. Am. Med Inf. Assoc. 21, 578–582 (2014).
Horwitz, L. I. et al. Researching COVID to Enhance Recovery (RECOVER) adult study protocol: Rationale, objectives, and design. PLoS One 18, e0286297 (2023).
Haendel, M. A. et al. The National COVID Cohort Collaborative (N3C): Rationale, design, infrastructure, and deployment. J. Am. Med. Inform. Assoc. 28, 427–443 (2021).
Brat, G. A. et al. International electronic health record-derived COVID-19 clinical course profiles: the 4CE consortium. NPJ Digit Med 3, 109 (2020).
The Biologics Effectiveness and Safety (BEST) Initiative. https://bestinitiative.org/.
U.S. Food & Drug Administration. FDA’s Sentinel Initiative. https://www.fda.gov/safety/fdas-sentinel-initiative.
U.S. Food & Drug Administration. Considerations for the Use of Real-World Data and Real-World Evidence to Support Regulatory Decision-Making for Drug and Biological Products. https://www.fda.gov/media/171667/download.
Zuo, X., Chen, Y., Ohno-Machado, L. & Xu, H. How do we share data in COVID-19 research? A systematic review of COVID-19 datasets in PubMed Central Articles. Brief. Bioinform. 22, 800–811 (2021).
Ohno-Machado, L. To Share or Not To Share: That Is Not the Question. Sci Transl Med 4, (2012).
Ohno-Machado, L. et al. pSCANNER: patient-centered Scalable National Network for Effectiveness Research. J. Am. Med. Inform. Assoc. 21, 621–626 (2014).
Wu, Y., Jiang, X., Kim, J. & Ohno-Machado, L. Grid Binary LOgistic REgression (GLORE): building shared models without sharing data. J. Am. Med. Inform. Assoc. 19, 758–764 (2012).
Lu, C.-L. et al. WebDISCO: a web service for distributed cox model learning without patient-level data sharing. J. Am. Med. Inform. Assoc. 22, 1212–1219 (2015).
Wang, J., Kolar, M., Srebro, N. & Zhang, T. Efficient distributed learning with sparsity. In International Conference on Machine Learning 3636–3645 (PMLR, 2017).
Jordan, M. I., Lee, J. D. & Yang, Y. Communication-efficient distributed statistical inference. J. Am. Stat. Assoc. 114, 668–681 (2019).
Duan, R. et al. Learning from electronic health records across multiple sites: A communication-efficient and privacy-preserving distributed algorithm. J. Am. Med. Inform. Assoc. 27, 376–385 (2020).
Duan, R., Boland, M. R., Moore, J. H. & Chen, Y. ODAL: A one-shot distributed algorithm to perform logistic regressions on electronic health records data from multiple clinical sites. Pac. Symp. Biocomput 24, 30–41 (2019).
Edmondson, M. J. et al. Distributed Quasi-Poisson regression algorithm for modeling multi-site count outcomes in distributed data networks. J. Biomed. Inf. 131, 104097 (2022).
Edmondson, M. J. et al. An efficient and accurate distributed learning algorithm for modeling multi-site zero-inflated count outcomes. Sci. Rep. 11, 19647 (2021).
Duan, R. et al. Learning from local to global: An efficient distributed algorithm for modeling time-to-event data. J. Am. Med Inf. Assoc. 27, 1028–1036 (2020).
Luo, C. et al. ODACH: a one-shot distributed algorithm for Cox model with heterogeneous multi-center data. Sci. Rep. 12, 6627 (2022).
Li, R., Romano, J. D., Chen, Y. & Moore, J. H. Centralized and federated models for the analysis of clinical data. Annu Rev. Biomed. Data Sci. 7, 179–199 (2024).
dGEM: Decentralized algorithm for Generalized mixed Effect Models with the Application in Hospital Profiling. https://www.ohdsi.org/2022showcase-72/.
Chen, Y. et al. Regression cubes with lossless compression and aggregation. IEEE Trans. Knowl. Data Eng. 18, 1585–1599 (2006).
Luo, C. et al. DLMM as a lossless one-shot algorithm for collaborative multi-site distributed linear mixed models. Nat. Commun. 13, 1678 (2022).
Williams, R. D. et al. Seek COVER: using a disease proxy to rapidly develop and validate a personalized risk calculator for COVID-19 outcomes in an international network. BMC Med Res Methodol. 22, 35 (2022).
Penn Computing Inference Learning (PennCIL) lab. Privacy-Preserving Distributed Algorithms. https://github.com/Penncil/pda.
OHDSI Community. OMOP Common Data Model. https://ohdsi.github.io/CommonDataModel/.
CMS Cell Size Suppression Policy. https://resdac.org/articles/cms-cell-size-suppression-policy.
California Department of Health Care Services. Data De-identification Guidelines (DDG). https://www.dhcs.ca.gov/dataandstats/Documents/DHCS-DDG-V2.0-120116.pdf.
Bonanad, C. et al. The Effect of Age on Mortality in Patients With COVID-19: A Meta-Analysis With 611,583 Subjects. J. Am. Med Dir. Assoc. 21, 915–918 (2020).
Nguyen, N. T. et al. Male gender is a predictor of higher mortality in hospitalized adults with COVID-19. PLoS One 16, e0254066 (2021).
Corona, G. et al. Diabetes is most important cause for mortality in COVID-19 hospitalized patients: Systematic review and meta-analysis. Rev. Endocr. Metab. Disord. 22, 275–296 (2021).
Du, Y., Zhou, N., Zha, W. & Lv, Y. Hypertension is a clinically important risk factor for critical illness and mortality in COVID-19: A meta-analysis. Nutr., Metab. Cardiovascular Dis. 31, 745–755 (2021).
Wasserman, L. & Zhou, S. A Statistical Framework for Differential Privacy. J. Am. Stat. Assoc. 105, 375–389 (2010).
Jiang, W. et al. WebGLORE: a Web service for Grid LOgistic REgression. Bioinformatics 29, 3238–3240 (2013).
Shu, D., Yoshida, K., Fireman, BH. & Toh, S. Inverse probability weighted Cox model in multi-site studies without sharing individual-level data. Stat. Methods Med. Res. 29, 1668–1681 (2019).
McCullagh, P. & Nelder, J. A. Generalized Linear Models. (Routledge, 2019). https://doi.org/10.1201/9780203753736.
Hilbe, J. M. Negative Binomial Regression. (Cambridge University Press, 2011). https://doi.org/10.1017/CBO9780511973420.
Ver Hoef, J. M. & Boveng, P. L. Quasi-Poisson vs. negative binomial regression: how should we model overdispersed count data? Ecology 88, 2766–2772 (2007).
Xia, W. et al. Managing re-identification risks while providing access to the All of Us research program. J. Am. Med Inf. Assoc. 30, 907–914 (2023).
Department of Health Agency Standards for Reporting Data with Small Numbers. https://doh.wa.gov/sites/default/files/legacy/Documents/1500//SmallNumbers.pdf (2018).
Utah Department of Health Data Suppression/Data Aggregation Guidelines Summary. https://ibis.health.utah.gov/ibisph-view/pdf/resource/DataSuppressionSummary.pdf (2022).
Shokri, R. & Shmatikov, V. Privacy-preserving deep learning. In 2015 53rd Annual Allerton Conference on Communication, Control, and Computing (Allerton) 909–910 (IEEE, 2015). https://doi.org/10.1109/ALLERTON.2015.7447103.
Phong, L. T., Aono, Y., Hayashi, T., Wang, L. & Moriai, S. Privacy-preserving deep learning via additively homomorphic encryption. IEEE Trans. Inf. Forensics Security 13, 1333–1345 (2018).
Zhang, C. et al. BatchCrypt: efficient homomorphic encryption for cross-silo federated learning. In Proc. 2020 USENIX Conference on Usenix Annual Technical Conference (USENIX Association, USA, 2020).
Fang, H. & Qian, Q. Privacy preserving machine learning with homomorphic encryption and federated learning. Future Internet 13, 94 (2021).
Xie, Q. et al. Efficiency optimization techniques in privacy-preserving federated learning with homomorphic encryption: a brief survey. IEEE Int. Things J. 11, 24569–24580 (2024).
Aono, Y., Hayashi, T., Phong, L. T. & Wang, L. Privacy-preserving logistic regression with distributed data sources via homomorphic encryption. IEICE Trans. Inf. Syst. E99.D, 2079–2089 (2016).
Park, J. & Lim, H. Privacy-preserving federated learning using homomorphic encryption. Appl. Sci. 12, 734 (2022).
Han, K., Hong, S., Cheon, J. H. & Park, D. Logistic regression on homomorphic encrypted data at scale. Proc. AAAI Conf. Artif. Intell. 33, 9466–9471 (2019).
Cheon, J. H., Kim, A., Kim, M. & Song, Y. Homomorphic encryption for arithmetic of approximate numbers. In Advances in Cryptology–ASIACRYPT 2017: 23rd International Conference on the Theory and Applications of Cryptology and Information Security, Proceedings, part I 23, 409–437 (Springer International Publishing, Hong kong, China, 2017).
Acknowledgements
This work was supported in part by the National Institutes of Health (1R01LM012607, 1R01AI130460, 1R01AG073435, 1R56AG074604, 1R01LM013519, 1R56AG069880, 1R01AG077820, 1U01TR003709). This work was supported partially through the Patient-Centered Outcomes Research Institute (PCORI) Project Program Awards (ME-2019C3-18315 and ME-2018C3-14899). All statements in this report, including its findings and conclusions, are solely those of the authors and do not necessarily represent the views of the Patient-Centered Outcomes Research Institute (PCORI), its Board of Governors, or the Methodology Committee. The article reflects the views of the author and should not be construed to represent FDA’s views or policies.
Author information
Authors and Affiliations
Contributions
Q.W. and Y.C. conceived and designed the study. Q.W., J.R., L.L., B.Z., Y.L., J.T., D.Z. and G.T. contributed to the analysis and interpretation of the data. Data collection and assembly were carried out by J.R., M.B., M.V., T.F., G.H. and M.S. The manuscript was drafted by Q.W. and Y.C., with critical revisions for important intellectual content provided by Q.W., C.Y., B.M., Y.L., S.D., J.Z. and Y.C. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Q., Reps, J.M., Li, L. et al. COLA-GLM: collaborative one-shot and lossless algorithms of generalized linear models for decentralized observational healthcare data. npj Digit. Med. 8, 442 (2025). https://doi.org/10.1038/s41746-025-01781-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01781-1