Abstract
Cardiovascular diseases remain the leading cause of death in developed countries. This study introduces deep learning ensembles for arrhythmia detection and atrial fibrillation (AF) recurrence prediction from electrocardiogram scans, supported by explainable artificial intelligence (XAI) methods. Validation used two datasets: Guangdong Provincial People’s Hospital, China (Dataset G, 1172 patients, 71.4 ± 6.3 years, 66% women, 20.5% with arrhythmia) and Liverpool Heart and Chest Hospital, UK (L, 909 patients, 60.5 ± 10.71 years, 33% women, 29.7% with arrhythmia). Our ensembles outperformed individual and voting models with the area under the receiver operating characteristic curve (ROC-AUC): 0.980 (95%CI: 0.956–0.998, p = 0.03) for Dataset G, 0.799 (95%CI: 0.737–0.856, p = 0.07) for Dataset L. The models trained on combined training sets achieved ROC-AUC: 0.980 (95%CI: 0.952–1.0) and 0.800 (95% CI: 0.739–0.861) for the G and L test sets. Precision-recall AUC for AF recurrence was 0.765 (95%CI: 0.669–0.849) for ensembles vs. 0.737 (95%CI: 0.648–0.821) for individual models. XAI enhanced interpretability for clinical applications.
Similar content being viewed by others
Introduction
Cardiovascular diseases (CVDs) are among the deadliest diseases in developing countries1. They claim more than 3.1 million lives annually in Europe2, with some of the countries reporting as much as 67% of deaths from purely cardiovascular issues. About half of all cardiovascular deaths are sudden, and about 80% of those are caused by cardiac arrhythmia3. The severity of the impact of CVD on society depends on the quality of medical care and its early detection, which is often less accessible in developing countries4.
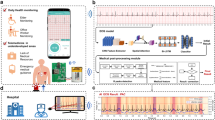
Among the most common, noninvasive, and cost-effective tools to diagnose early signs of arrhythmia is electrocardiogram (ECG) analysis. Despite its widespread adoption, challenges remain in interpreting ECG data and ensuring reproducibility and standardization of the analysis, particularly in resource-constrained clinical settings. To tackle these challenges, it is necessary to implement an algorithm that learns to mimic the decisions of experienced cardiologists. This may allow for delivering immediate, objective, and continuous observation of a patient. Timely medical intervention and prevention can result in lower hospitalization costs across a given population, along with a decreased risk of long-term health complications. It is important as CVDs are prevalent in growing economies, which may not have funding to provide healthcare institutions with enough funding.
Due to the amount of available ECG scans, developing large-capacity deep learning models has become increasingly popular. Such methods are characterized by the ability to self-derive complex characteristics and intrinsic features of an analyzed dataset through automated representation learning5. This property made deep learning solutions widely spread in a range of research areas within the medical data analysis field, such as drug discovery6, patient diagnostics7, or medical imaging8, among others. Due to their complexity, the reasoning process of deep learning models is rarely explained. The inability to provide coherent reasoning and understanding behind the automatically elaborated diagnosis is a direct reason for a relatively low trust put into machine learning solutions in medicine, both from the perspective of doctors and patients9. To increase the perceived usefulness and trust in deep learning solutions, various techniques for explaining their predictions were developed10. The local explainable artificial intelligence (XAI) methods highlight the input data features, e.g., parts of the medical image fed into a deep learning model, which strongly influences the prediction outcome, letting a practitioner understand which image parts were important during inference11. Such techniques are particularly useful in clinical settings, as they elaborate explanations for each input data sample (e.g., an ECG scan)12. There exist both classic13 and deep machine learning (ML)14 methods for analyzing ECG scans, with the former requiring the design of manual feature extractors. Such techniques are often validated using local datasets and may not generalize well over different cohorts, and they lack interpretability, limiting clinical utility.
To enhance the generalizability of ML systems, ensemble learning has been widely exploited in various pattern recognition and data analysis tasks15. However, building such cooperative committees of ML models is a multi-objective optimization problem: the classification performance should be maximized, the heterogeneity of an ensemble should usually be maximized too (to benefit from different model architectures), and their size is often minimized to avoid overfitting and accelerate their operations. Although there are a variety of automated ML (AutoML) approaches toward automatically designing ML pipelines16, this research area is under research in the context of arrhythmia identification and ECG analysis. We address this research gap and tackle the problem of automated detection of signs of arrhythmia from ECG scans using automatically evolved, interpretable, and rigorously validated—in both intra- and inter-dataset settings—deep learning ensembles, which are built using a genetic programming approach in a fully data-driven manner.
Results
The experimental results (Tables 2 and 3) showed that—for Dataset G—the best GIRAFFE ensemble achieved ROC-AUC of 0.980 (95% CI: 0.956–0.998), which was a significant (p = 0.03) improvement over the baseline with ROC-AUC of 0.961 (95% CI: 0.931–0.986). For Dataset L, GIRAFFE improved the arrhythmia detection performance metrics, achieving ROC-AUC of 0.799 (95% CI: 0.737–0.856) compared to 0.773 (95% CI: 0.708–0.837) of the baseline model (p = 0.07). The comparison of Tables 2 and 3 reveals challenges in ensuring inter-dataset generalizability of ML models trained using training data from one dataset. The ML model achieved ROC-AUC of 0.980 (95% CI: 0.956–0.998) when both trained and tested on Dataset G, using respective training and test subsets of this dataset (Table 1). When the ML model was trained on the training set T from Dataset L and then tested on test set Ψ from Dataset G, its classification performance dropped to ROC-AUC of 0.494 (95% CI: 0.390–0.603). Similarly, for Dataset L, ROC-AUC decreased from 0.799 (95% CI: 0.737–0.856) (when the model was trained and tested on the training and test subsets of Dataset L) to 0.556 (95% CI: 0.521–0.595), when it was trained on the training set T from Dataset G and tested on the test set Ψ from Dataset L.
The performance of the models trained on the combined datasets was also evaluated (Table 2). When trained using both Dataset G and Dataset L training data simultaneously, the model maintained its high-performance levels on the respective test subsets. For Dataset G, this combined training approach achieved ROC-AUC of 0.980 (95% CI: 0.952–1.000), matching the performance of the dataset-specific model with ROC-AUC of 0.980 (95% CI: 0.956–0.998). Similarly, for Dataset L, the combined training yielded ROC-AUC of 0.800 (95% CI: 0.739–0.861), comparable to the dataset-specific model’s ROC-AUC of 0.799 (95% CI: 0.737–0.856).
Figure 1 displays the ROC and PR curves for the intra-dataset models, along with their corresponding probability distributions. These distributions reveal that the GIRAFFE ensembles produce sharper probability distributions, with more concentrated probability mass in high-confidence regions and lower probabilities assigned to peripheral areas. This means that GIRAFFE ensembles are more confident in their predictions compared to baseline models.
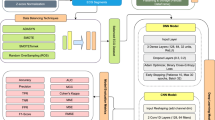
a Dataset split and training strategy: models are trained on different subsets with early stopping. b Ensemble creation using GIRAFFE: model predictions are fused using evolved strategies (mean, thresholding, weighted sum). c Generalization and performance metrics on the G and L datasets: Precision-Recall/ROC curves and probability distributions highlight GIRAFFE’s improvements. d Explainable artificial intelligence (XAI) interpretability: Local Interpretable Model-Agnostic Explanations (LIME) and Integrated Gradients (IG) highlight image regions; results are ensembled to showcase model reasoning.
The ML models underwent the explanation process (Fig. 1)—the example results of this process are rendered in Fig. 2, highlighting the parts of the image affecting the prediction process of the corresponding models (in the visual examples, the green color is used to highlight areas which were crucial for making the model prediction, whereas the red color corresponds to counter examples highlighting areas making model consider the opposite class). The practical scenarios indicating the correct and incorrect operation of the ML model being explained (i.e., correctly identifying healthy and unhealthy patients, and classifying a healthy patient as one with arrhythmia, or vice versa) are presented in Fig. 2.
The meaning of the colors is as follows: the green color indicates regions which are relevant for the current prediction, while the red color is used for highlighting regions that may support opposite prediction.
AF detection
While our primary analysis examined arrhythmia detection broadly, we conducted additional sensitivity analyses to better understand the model’s performance for specific clinical contexts, particularly for AF. For Dataset L (LHCH), which specifically focused on AF recurrence following Catheter Ablation, we performed a dedicated analysis to evaluate our model’s performance in assessing the utility of our approach for predicting AF recurrence from pre-procedure ECGs, which has significant implications for post-ablation management and outcomes.
Among the 909 patients in Dataset L, 270 (29.7%) experienced AF recurrence following catheter ablation (Table 1). GIRAFFE ensemble achieved ROC-AUC of 0.799 (95% CI: 0.737–0.856), outperforming the individual model (ROC-AUC: 0.773, 95% CI: 0.708–0.837) with borderline statistical significance (p = 0.07). The PR-AUC for AF recurrence detection was 0.765 (95% CI: 0.669–0.849) for the GIRAFFE ensemble compared to 0.737 (95% CI: 0.648–0.821) for the individual model (Table 2).
When examining the pre-procedure ECGs of patients who later experienced AF recurrence, we observed enhanced prediction stability with the ensemble approach. For patients with multiple pre-procedure ECG scans, the GIRAFFE ensemble produced consistent predictions (all scans classified identically) in 81.5% of cases, compared to 75.0% with the individual model (Table 4). When examining the XAI explanation maps specifically for AF recurrence cases in Dataset L, we observed that the ensemble model consistently highlighted regions corresponding to known electrophysiological markers associated with AF, particularly focusing on P-wave morphology and rhythm irregularities.
Discussion
It is important to contextualize our findings with respect to the specific data modality used. While large public datasets of raw ECG signals exist, such as PTB-XL with over 21,000 recordings17, our study addresses the distinct challenge of analyzing scanned images of legacy paper ECGs. The clinical relevance and technical challenge of this data type have been recently highlighted by the 2024 George B. Moody PhysioNet Challenge, which focused exclusively on this problem (analysis of ECG printouts)18. This modality, common in many clinical settings, introduces variability from scanner hardware and print quality, making it a challenging real-world clinical use case. The primary aim of this work was therefore to develop and validate a methodology robust to these challenges, rather than to train a state-of-the-art model on a large-scale signal dataset.
One of the important strengths of the reported study lies in its rigorous multi-center, cross-dataset validation approach. By evaluating ML models across distinct datasets (Dataset G and Dataset L) from different clinical centers, it is possible to assess not only the ML models’ classification performance but also their real-world generalizability. This comprehensive validation strategy revealed important insights concerning the challenges and limitations of ECG classification systems when deployed across different clinical settings. Several important insights may be learned from our study. The analysis of the results indicates that the GIRAFFE ensembles demonstrate substantial performance gains compared to the baseline models, which lead to consistently statistically significant improvements over the baseline models (Table 3). Notably, the naïve ensembles (averaging the predictions of base ML models) also showed performance gains, but these were never statistically significant. Importantly, across all experiments and ensemble models, the performance never degraded significantly below that of the respective baselines (Table 3).
An important finding of our study is the limited cross-dataset generalizability observed when models trained on one cohort were tested on the other (Table 4). This performance degradation is strong evidence of a significant domain shift between the two datasets, which can be attributed to at least three distinct sources of heterogeneity.
First, the clinical tasks tackled within the considered datasets are fundamentally different. Dataset G addresses a diagnostic problem (detecting current arrhythmia), whereas Dataset L tackles a more challenging prognostic problem (predicting future AF recurrence post-ablation). The underlying electrophysiological features for these two tasks are not necessarily the same, creating a significant task-level domain gap.
Second, the patient demographics differ substantially, as shown in Table 1. The cohort from Dataset G is, on average, a decade older than that from Dataset L (71.4 ± 6.3 vs. 60.5 ± 10.7 years, respectively) and has a much higher percentage of female patients (66% vs. 33%). These differences represent distinct population distributions that challenge model generalization.
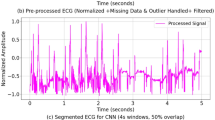
Finally, there are variations in data acquisition and format. Although both datasets consist of scanned 12-lead ECGs, they originate from different clinical centers with different equipment and protocols, leading to variations in printout quality, scanner artifacts, and the number of available long leads. This source of heterogeneity is further explored in Table 1, showcasing the difference between the mean and standard deviation of image channels, which may contribute to the impeded knowledge transfer between datasets. The comparison of example ECG scans is presented in Fig. 3, showing major differences between the collected ECGs. Collectively, these factors contribute to the observed performance drop and highlight a critical challenge for the deployment of AI models in real-world clinical settings: the models must be robust not only to population differences but also to variations in clinical tasks and data sources.
Examples of ECG scans from each dataset showing their heterogeneity.
Combining the datasets to create a larger, more diverse training set did not yield substantial performance improvements when compared to the ML models specialized in either Dataset G or Dataset L. Despite the theoretical advantage of a larger training set for deep neural networks, the experiments showed no notable gains when evaluating the combined model against single-dataset models. Specifically, when tested on Dataset G’s test set Ψ, both Dataset G-specific ML model and the combined-data model achieved identical ROC-AUC scores of 0.980 (Tables 2 and 3, respectively). Similarly, when evaluated on Dataset L’s test set—, the Dataset L-specific model achieved ROC-AUC of 0.799, while the combined-data model scored nearly identically at 0.800 (Tables 2 and 3, respectively). Hence, rather than learning generalizable features from the combined data, the model developed dataset-specific and task-specific classification patterns for each dataset.
Patients in Dataset L had varying numbers of ECG scans per individual (on the other hand, each patient in Dataset G had exactly one ECG scan, hence the number of scans equals the number of patients in Dataset G). All scans for a given patient shared the same label, as they were acquired prior to cardiac ablation surgery, after which AF recurrence might occur. The model’s goal was to predict this likelihood of recurrence. The consistency of model predictions at the patient level was analyzed using a 50% probability threshold, examining the number and percentage of patients for whom all scans received consistent predictions—either all normal or all abnormal (Table 4).
The consistency values range from 75 to 85%. The ensemble models generally demonstrated higher consistency than baseline (individual) models, aligning with the probability distributions shown in Fig. 1. However, this higher consistency does not appear to correlate with overall model performance. This suggests that for patients who experienced AF recurrence, only some of their ECG scans may display abnormal characteristics. This heterogeneity in ECG presentations makes both training and evaluation more challenging and may partially explain Dataset L’s lower performance metrics compared to Dataset G.
With the upcoming changes in the legal landscape, complex artificial AI solutions in high-impact domains such as medicine need to incorporate a level of transparency19. It might be accomplished by providing probabilities associated with the model’s prediction, but this approach does not provide insights concerning the decision-making process of the analyzed model. It, in turn, leads to lower trust in black-boxed solutions, especially when comparing them with those that are easier to interpret20. The model-agnostic explainability allows for using a diverse set of models, independently of their ML architecture, by highlighting parts of the input data that were crucial during the inference process.
The reported explanation examples (Fig. 2) show that by identifying specific patterns in ECG scans, it is possible to select scans that are more likely to be misclassified due to the uncertainty of the model’s prediction could be analyzed further by a human reader. When considering the correct classification of a healthy patient, the baseline model does not focus on any region in particular. On the other hand, the GIRAFFE ensemble focuses on peaks of the leads to verify that they could be associated with a healthy patient. The example of a correctly classified patient with arrhythmia showcases different behaviors by both baseline and GIRAFFE algorithms. GIRAFFE focuses its attention on a singular lead. That implies that the lead is most likely to contain signs of abnormal heart activity according to most of the individual models making up the ensemble. This, however, is not the case for the baseline model, which does not seem to favor a single area, highlighting all possible areas relevant to the prediction. The next examples bring an incorrect prediction by both models, starting with a healthy patient incorrectly classified as having an abnormal heart rhythm. The importance map in the case of both models is highly focused on a single lead, with regions of interest taking a sparser form, with both models focusing on the lower right corner of the ECG. The sparsity can be a result of low confidence in prediction and the presence of noisy regions, which can alter the prediction. The trend of lower sparsity compared to correct classification continues during the example of the incorrectly classified unhealthy patient, along with more frequent occurrences of red regions representing regions that could be attributed to the opposite prediction—the patient being unhealthy.
The last four examples showcase explanations when only one of the models delivered the correct prediction. When the GIRAFFE ensemble provides a correct prediction, it can be observed that the model looks at similar parts of an image. It indicates that the GIRAFFE ensemble model can interpret the same signal more thoroughly. In the opposite example (the baseline model elaborates the correct prediction), it can be observed that the baseline model focused on the regions ignored by GIRAFFE that have their importance diminished if they are not utilized consistently across models incorporated into the ensemble classifier.
Limitations
There are some limitations of this study. The datasets were relatively small for deep learning image classification, containing only 1000–2000 examples each, especially compared to large-scale public corpora of raw digital ECG signals17. However, they represent a substantial collection for the specific modality of scanned paper ECGs, for which large, publicly available datasets only begin to emerge, yet remain not publicly available21. Although the models were pre-trained on ImageNet, no task-specific pre-trained models for ECG scan classification were available–using such models might have enhanced the performance of the ensemble classifiers. Second, despite both datasets addressing the arrhythmia ECG classification from paper ECG scans, their heterogeneity limited the cross-dataset (and cross-task) generalizability of the ML models. Specifically, Dataset G from Guangzhou represents a population study focused on detecting the presence or absence of AF, while Dataset L from LHCH aimed to predict AF recurrence using pre-operative ECGs. Dataset L presents an additional challenge for model training and validation, as each patient had multiple ECG scans sharing a single label, though not all ECGs may have exhibited characteristics indicative of the same prognosis. Furthermore, this study did not include a comparison to classical machine learning models that rely on handcrafted features. Applying such models directly to the ECG printouts is not necessarily feasible, as global image features are dominated by non-signal background pixels. A fair comparison would necessitate a complex multi-step pipeline involving ECG signal segmentation and digitization to extract standard clinical (signal) features. The development of such a pipeline is a significant research challenge, as highlighted by its inclusion as a dedicated task in the recent PhysioNet Challenge18 and would require segmentation ground truth, which was not available for our datasets. Therefore, we focused on end-to-end deep learning methods which learn representations directly from the image. Finally, although GIRAFFE ensembling achieved superior results compared to traditional approaches, the challenge of cross-regional generalization persists and could be tackled using explainable AI techniques. They provided insights into model decision-making understanding this process for ECG scans captured in different sites might help investigate the causes why the ML models failed (different examination procedures, devices, other clinical/technical reasons), potentially suggesting a path toward making them more robust for target data distributions, through e.g., appropriate pre-processing of incoming data or fine-tuning the models. Our current research efforts are focused on designing a Mean Opinion Score experiment22,23 for a follow-up large-scale study, in which the explanations will be shown to cardiologists (with different years of experience)—the practitioners will be asked to interpret the explanations, as well as the way they are reported, to validate their potential clinical utility. Incorporating a human in the explainable AI loop is key in accelerating such approaches in various fields24, with medicine being a prominent example, where transparent and standardized reporting of XAI outcomes became crucial25.
In conclusion, deep learning ensembles are built using a genetic programming approach to obtain accurate identification of patients with arrhythmia from ECG scans, and their predictions can be interpreted using local explainable artificial intelligence techniques for improving their potential clinical utility.
Methods
Datasets
In this study, two clinical datasets were used: the Guangzhou Heart Study Dataset (GHS)26 (Dataset G), and the Liverpool Heart and Chest Hospital Study Dataset (LHCH)27 (Dataset L).
Dataset G comprised N = 1172 patients, and each patient was associated with a 12-lead ECG scan and a label reflecting the patient status, either normal (sinus rhythm) or abnormal (arrhythmia), acquired between May 1, 2019, and August 31, 2019 (Table 1). The arrhythmia category included the following conditions: (i) atrial fibrillation (AF) and flutter (AFL), (ii) premature ventricular contractions, (iii) premature atrial contractions, (iv) ventricular tachycardia, (v) Wolff-Parkinson-White (WPW) syndrome, (vi) pacing rhythm, and (vii) borderline rhythm. Dataset L was prospectively acquired in a study focused on the AF recurrence following catheter ablation, which has reported rates of 20–45%28, and it included N = 909 patients (33% women, 60.5 ± 10.7 years; 1821 ECG scans in total) acquired between June 2013 and December 2019. ECGs were included if the full clinical record was available for a patient and if ECG scans were obtained within 6 months prior to ablation, with patients having between one and 10 ECGs (with mean of 2 ECGs per patient). The first dataset (Dataset G) is from the Guangzhou Heart Study26. This study was approved by the Guangzhou Medical Ethics Committee of the Chinese Medical Association. The second dataset (Dataset L) is approved from the Liverpool Heart and Chest Hospital NHS Foundation Trust as a service evaluation focused on the recurrence of AF after catheter ablation.
Intra- and inter-dataset validation strategy
The datasets were split into the training and test sets as suggested in refs. 27,29—these splits are used in the inter-dataset and intra-dataset validation. Furthermore, each of the training sets was split into six partly overlapping, stratified folds, ensuring similar proportions of arrhythmia cases in each fold. For Dataset L, the data were split at the patient level, as patients had varying numbers of acquired scans, thus ensuring no information leakage across folds. Therefore, each dataset (Dataset G and Dataset L) was divided into the following subparts (Table 1):
-
Training folds (collectively referred to as T)—Subsets T1–T5—these sets are used for training ML models,
-
Validation set–Subset V (6th fold)—it is used for ranking separate ML models and for training the ensemble models.
-
Test set—Subset Ѱ—it is used for verifying the generalization of the models.
To verify the models’ generalization across datasets and across different tasks (diagnosis of AF using ECGs for Dataset G, and prediction of post-ablation AF recurrence using pre-operative ECGs for Dataset L), we followed the inter-dataset validation strategy, where (i) the models were trained using T from Dataset G and they were tested over Ψ from Dataset L, (ii) the models were trained using T from Dataset L and tested over Ψ from Dataset G, and (iii) the models were trained using the combined T (from Dataset G and Dataset L), and tested over the combined Ψ from Dataset G and Dataset L (Fig. 1).
Deep learning algorithms for arrhythmia detection in ECG scans
This study focuses on end-to-end deep learning approaches that perform automated representation learning directly from the scanned ECG images. This strategy was chosen because the clinically relevant signal constitutes only a small fraction of each image, making classical approaches based on handcrafted features challenging to implement without a complex pre-processing pipeline for signal extraction and digitization. Our deep learning ensemble approach builds upon two distinct deep network architectures: InceptionNetV330 and EfficientNet31, pre-trained on the large-scale ImageNet dataset32. Each architecture was implemented with either a standard Sigmoid activation function or a Generalized Extreme Value (GEV) activation function33, the latter being particularly effective for handling class imbalance scenarios. For each architectural configuration, we developed multiple models using two training strategies:
-
A five-fold cross-validation approach, where five separate models were trained using four folds from T, with the remaining fifth fold serving as an internal validation set.
-
A full-dataset training approach, where a single model was trained on the complete training dataset T from either Dataset G or Dataset L.
This approach resulted in six trained deep learning models per architecture-activation combination (5 fold-specific models together with 1 full-dataset model). During the training process, an early stopping mechanism was used to monitor the validation loss, terminating the training if no improvement was observed over ten consecutive epochs. The training sets were augmented using random rotations of the T images, to up to 5°. Overall, the combination of two architectures (InceptionNetV3, EfficientNet) and two activation functions (Sigmoid, GEV) resulted in four distinct base model configurations., Six deep learning models were trained per architecture-activation combination (5 fold-specific models together with 1 full-dataset model), yielding a comprehensive set of 24 individual (base) models for subsequent ensemble learning. To evolve ensemble ML models, the GIRAFFE algorithm34 was utilized—it leverages genetic programming to select relevant base models and evolve their fusion schema. The predictions of base models are aggregated in a way delineated by GIRAFFE to produce the final arrhythmia probability estimation (Fig. 1). The ensembles created by GIRAFFE are inherently interpretable and are differentiable, which is a requirement for many XAI techniques.
Understanding predictions of deep learning algorithms using explainable AI methods
We exploit local XAI methods, which provide explanations for each classified ECG scan—they highlight parts of the scan that the model found important while figuring out whether the patient has arrhythmia or not. This approach is built from two XAI methods: Local Interpretable Model-Agnostic Explanations (LIME)35 and Integrated Gradients (IG)36. In LIME, an input ECG scan is automatically segmented into super-pixels (locally coherent image areas sharing similar pixel characteristics), which are later sequentially masked during the analysis process. The model’s predictions obtained for such perturbed input images (with masked super-pixels) are used to quantify the importance of each super-pixel, which contributes to the decision process. In IG, the gradients of the ML model’s outputs are calculated along a path from the baseline (empty) image, which is progressively changed to the actual input ECG scan, and the contribution of each pixel to the prediction is quantified.
Once the individual explanations (LIME and IG) were obtained, they were combined using the Second Moment Scaling method37 and then averaged into a single attribution map (Fig. 1). By combining two types of XAI techniques—input data manipulation (LIME) and gradient-based analysis (IG)—into a unified scoring system, the framework provides a holistic understanding of the model’s decision-making process, thereby enhancing its clinical utility.
Quality metrics and statistical analysis
To quantify the classification performance of ML models, we investigate the Receiver Operating Characteristic (ROC) and Precision-Recall (PR) curves and calculate the areas under these curves (ROC-AUC and PR-AUC, respectively). For each experimental setup, defined by a combination of the training and test sets, we report the results for four approaches: (i) the best individual ML model, selected based on its ROC-AUC score on the validation set V, (ii) a naïve ensemble ML model, calculated as the average prediction across all base models, (iii) an upper-bound GIRAFFE model, representing the best ROC-AUC achievable on Ψ; and (iv) the GIRAFFE model selected by maximizing the average ROC-AUC on \({\bf{T}}{\boldsymbol{\cup }}{\bf{V}}\).
The bootstrap method with 1000 sampling repetitions (with replacement) was used to estimate confidence intervals and p-values. Each sampled set was used to calculate the performance metrics (ROC-AUC and PR-AUC). Confidence intervals were determined by the 5th and 95th percentiles of the resulting metric distributions for each model. To calculate p-values, each model is compared to its respective baseline by computing the difference in their metrics for each resampled set. These differences are then centered around zero to simulate the null hypothesis (no difference between the models). Finally, the p-value is determined as the proportion of these centered differences that were more extreme (in a single direction for a one-sided test) than the originally observed difference between the model and its baseline.
Python (version 3.12) was an implementation framework for all ML models and statistical analysis. The most important packages included: Pytorch (v. 2.6.0) for implementing deep learning algorithms, Pytorch-lightning (v. 2.5.0) for training process, Scikit-learn (v. 1.6.1) for data stratification and handling, Numpy (v. 2.2) for tensor operations, and Pandas (v. 2.2.3) and Matplotlib (3.10) for statistics and data visualizations. All experiments were executed and tracked in Weights and Biases to ensure full reproducibility of this study. The figures were prepared using the Microsoft PowerPoint 16.99 software.
Data availability
The datasets generated during and analyzed in the current study are available from the corresponding author upon reasonable request.
Code availability
The code associated with this submission is publicly available through the GitHub repository (https://github.com/smile-research).
Abbreviations
- AF :
-
atrial fibrillation
- AFL :
-
atrial flutter
- AI :
-
artificial intelligence
- AUC :
-
area under the curve
- CC :
-
correctly classified
- CI :
-
confidence interval
- CVD :
-
cardiovascular disease
- CV :
-
cardiovascular
- GEV :
-
generalized extreme value (distribution)
- GIRAFFE :
-
genetic programming algorithm to build deep learning ensembles for ECG arrhythmia classification
- ECG :
-
electrocardiogram
- IG :
-
integrated gradients
- LIME :
-
local interpretable model-agnostic explanations
- ML :
-
machine learning
- PR :
-
precision-recall (curve)
- ROC :
-
receiver operating characteristic (curve)
- WPW :
-
Wolf-Parkinson-White syndrome
- XAI :
-
explainable artificial intelligence
References
Dalakoti, M. et al. Primary prevention of cardiovascular disease in Asia: challenges: a narrative review. JACC Adv. 4, 101670 (2025).
Timmis, A. et al. European Society of Cardiology: the 2023 Atlas of Cardiovascular Disease Statistics. Eur. Heart J. 45, 4019–4062 (2024).
Mehra, R. Global public health problem of sudden cardiac death. J. Electrocardiol. 40, S118–S122 (2007).
Cardiovascular diseases (CVDs). https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds) (2021).
Sarker, I. H. Deep learning: a comprehensive overview on techniques, taxonomy, applications and research directions. SN Comput. Sci. 2, 420 (2021).
Gupta, R. et al. Artificial intelligence to deep learning: machine intelligence approach for drug discovery. Mol. Divers. 25, 1315–1360 (2021).
Hama, T. et al. Enhancing patient outcome prediction through deep learning with sequential diagnosis codes from structured electronic health record data: systematic review. J. Med. Internet Res. 27, e57358 (2025).
Thakur, G. K., Thakur, A., Kulkarni, S., Khan, N. & Khan, S. Deep learning approaches for medical image analysis and diagnosis. Cureus 16, e59507 (2024).
Teng, Q., Liu, Z., Song, Y., Han, K. & Lu, Y. A survey on the interpretability of deep learning in medical diagnosis. Multimed. Syst. 28, 2335–2355 (2022).
van der Velden, B. H. M., Kuijf, H. J., Gilhuijs, K. G. A. & Viergever, M. A. Explainable artificial intelligence (XAI) in deep learning-based medical image analysis. Med. Image Anal. 79, 102470 (2022).
Lesley, U. & Kuratomi Hernández, A. Improving XAI explanations for clinical decision-making—physicians’ perspective on local explanations in healthcare. In Proc. 22nd International Conference on Artificial Intelligence in Medicine: AIME 2024 (eds. Finkelstein, J., Moskovitch, R. & Parimbelli, E.) 296–312 (Springer Nature Switzerland, 2024).
Ganeshkumar, M. R. & Ravi, V. Explainable deep learning-based approach for multilabel classification of electrocardiogram. IEEE Trans. Eng. Manag. 70, 2787–2799 (2023). V, S., E. A., G. & K. P., S.
Rajpal, H. et al. Interpretable XGBoost-based classification of 12-lead ECGs applying information theory measures from neuroscience. Comput. Cardiol. 47, 1 (2020).
Daydulo, Y. D., Thamineni, B. L. & Dawud, A. A. Cardiac arrhythmia detection using deep learning approach and time frequency representation of ECG signals. BMC Med. Inform. Decis. Mak. 23, 232 (2023).
Dang, T., Nguyen, T. T., McCall, J., Elyan, E. & Moreno-García, C. F. Two-layer ensemble of deep learning models for medical image segmentation. Cogn. Comput. 16, 1141–1160 (2024).
Shin, S., Park, D., Ji, S., Joo, G. & Im, H. Medical data analysis using AutoML frameworks. J. Electr. Eng. Technol. 19, 4515–4522 (2024).
Wagner, P. et al. PTB-XL, a large publicly available electrocardiography dataset. Sci. Data 7, 154 (2020).
Reyna, M. A. et al. Digitization and classification of ECG images: the George B. Moody PhysioNet Challenge 2024. Comput. Cardiol. 51, 1–4 (2024).
Aboy, M., Minssen, T. & Vayena, E. Navigating the EU AI Act: implications for regulated digital medical products. npj Digit. Med. 7, 229 (2024).
Rosenbacke, R., Melhus, Å, McKee, M. & Stuckler, D. How explainable artificial intelligence can increase or decrease clinicians’ trust in AI applications in health care: systematic review. JMIR AI 3, e53207 (2024).
Reyna, M. A. et al. ECG-Image-Database: a dataset of ECG images with real-world imaging and scanning artifacts; a foundation for computerized ECG image digitization and analysis. Preprint at https://doi.org/10.48550/arXiv.2409.16612 (2024).
Nalepa, J. et al. Fully-automated deep learning-powered system for DCE-MRI analysis of brain tumors. Artif. Intell. Med. 102, 101769 (2020).
Grabowski, B. et al. Squeezing adaptive deep learning methods with knowledge distillation for on-board cloud detection. Eng. Appl. Artif. Intell. 132, 107835 (2024).
Hryniewska, W., Grudzień, A. & Biecek, P. LIMEcraft: handcrafted superpixel selection and inspection for visual explanations. Mach. Learn. 113, 3143–3160 (2024).
Brankovic, A. et al. Clinician-informed XAI evaluation checklist with metrics (CLIX-M) for AI-powered clinical decision support systems. npj Digit. Med. 8, 364 (2025).
Deng, H. et al. Epidemiological characteristics of atrial fibrillation in Southern China: results from the Guangzhou Heart Study. Sci. Rep. 8, 17829 (2018).
Aldosari, H., et al. Electrocardiogram two-dimensional motifs: a study directed at cardiovascular disease classification. in Knowledge Discovery, Knowledge Engineering and Knowledge Management (eds. Coenen, F. et al.) 3–27 (Springer Nature Switzerland, 2023).
Dretzke, J. et al. Predicting recurrent atrial fibrillation after catheter ablation: a systematic review of prognostic models. Europace 22, 748–760 (2020).
Bridge, J. et al. Artificial intelligence to detect abnormal heart rhythm from scanned electrocardiogram tracings. J. Arrhythmia 38, 425–431 (2022).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2818–2826; (IEEE, 2016).
Tan, M. & Le, Q. V. EfficientNet: rethinking model scaling for convolutional neural networks. In Proc. 36th International Conference on Machine Learning (PMLR) Vol. 97 (eds. Chaudhuri, k. & Salakhutdinov, R. et al.) 6105–6114 https://doi.org/10.48550/arXiv.1905.11946 (2019).
Deng, J. et al. ImageNet: a large-scale hierarchical image database. in Proc. IEEE Conference on Computer Vision and Pattern Recognition 248–255; https://doi.org/10.1109/CVPR.2009.5206848 (IEEE, 2009).
Bridge, J. et al. Introducing the GEV activation function for highly unbalanced data to develop COVID-19 diagnostic models. IEEE J. Biomed. Health Inform. 24, 2776–2786 (2020).
Kucharski, D. et al. Giraffe: a genetic programming algorithm to build deep learning ensembles for ECG arrhythmia classification. in Proc.IEEE International Conference on Image Processing (ICIP) 3070–3076; https://doi.org/10.1109/ICIP51287.2024.10647780 (2024).
Ribeiro, M. T., Singh, S. & Guestrin, C. “Why should I trust you?”: explaining the predictions of any classifier. in Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 1135–1144; https://doi.org/10.1145/2939672.2939778 (2016).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. in Proc. 34th International Conference on Machine Learning 3319–3328 (2017).
Miró-Nicolau, M., Jaume-i-Capó, A. & Moyà-Alcover, G. Meta-evaluating stability measures: MAX-sensitivity and AVG-sensitivity. in Explainable Artificial Intelligence (eds. Longo, L., Lapuschkin, S. & Seifert, C.) 356–369 https://doi.org/10.48550/arXiv.2412.10942 (Springer Nature Switzerland, 2024).
Acknowledgements
A.M.W. and J.N. were supported by the Silesian University of Technology funds through the Excellence Initiative—Research University program (grant 02/080/SDU/10-21-01). This study was supported by the Silesian University of Technology funds through the grant for maintaining and developing research potential. The Article Processing Charge was financed under the European Funds for Silesia 2021–2027 Program, co-financed by the Just Transition Fund—project entitled “Development of the Silesian biomedical engineering potential in the face of the challenges of the digital and green economy (BioMeDiG)”. The project number: FESL.10.25-IZ.01-07G5/23. We gratefully acknowledge Polish high-performance computing infrastructure PLGrid (HPC Center: ACK Cyfronet AGH) for providing computer facilities and support within the computational grant no. PLG/2024/017393.
Author information
Authors and Affiliations
Contributions
Arkadiusz Czerwiński: conceptualization, methodology, design and implementation of the explanation and training frameworks, visualization, data curation, statistical analysis, computational experiments, writing—original draft, writing—revised draft. Damian Kucharski: conceptualization, methodology, design and development of the GIRAFFE algorithm and training frameworks, data curation, computational experiments, statistical analysis, visualization, writing–original draft, writing—revised draft. Agata M. Wijata: conceptualization, methodology, investigation, validation, visualization, formal analysis, supervision, resources, funding acquisition, writing—original draft, writing—revised draft. Dhiraj Gupta: data acquisition, data preparation.Lu Fu: data acquisition, data preparation. Weidong Lin: data acquisition, data preparation. Yumei Xue: data acquisition, data preparation. Jacek Kawa: resources, manuscript revision. Hanadi Aldosari: data acquisition, data preparation. Frans Coenen: data acquisition, data preparation. Yalin Zheng: conceptualization, supervision, resources, manuscript revision. Gregory Y. H. Lip: conceptualization, supervision, resources, manuscript revision, writing—revised draft. Jakub Nalepa: conceptualization, methodology, investigation, validation, formal analysis, supervision, resources, writing—original draft, writing—revised draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Czerwinski, A., Kucharski, D., Wijata, A.M. et al. Interpretable arrhythmia detection in ECG scans using deep learning ensembles: a genetic programming approach. npj Digit. Med. 8, 642 (2025). https://doi.org/10.1038/s41746-025-01932-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01932-4