Abstract
Electronic health records (EHRs) contain rich longitudinal information for clinical decision-making, yet LLMs struggle to reason across patient timelines. We introduce TIMER (Temporal Instruction Modeling and Evaluation for Longitudinal Clinical Records), a method to improve LLMs’ temporal reasoning over multi-visit EHRs through time-aware instruction tuning. TIMER grounds LLMs in patient-specific temporal contexts by linking each instruction-response pair to specific timestamps, ensuring temporal fidelity throughout the training process. Evaluations show that TIMER-tuned models outperform conventional medical instruction-tuned approaches by 6.6% in completeness on clinician-curated benchmarks, with distribution-matched training demonstrating advantages up to 6.5% in temporal reasoning. Qualitative analyses reveal that using TIMER enhances temporal boundary adherence, trend detection, and chronological precision, necessary for applications such as disease trajectory modeling and treatment response monitoring. Overall, TIMER provides a methodological basis for developing LLMs that can effectively engage with the inherently longitudinal nature of data for patient care. Code is available at TIMER.
Similar content being viewed by others
Introduction
Language models (LMs) have recently shown the ability to process exceptionally long contexts-spanning hundreds of thousands of tokens-yet their capacity for temporal reasoning over longitudinal documents remains limited1,2. In clinical settings, this limitation is particularly consequential: physicians must routinely synthesize complex longitudinal electronic health records (EHRs) across multiple visits to manage chronic conditions, make care decisions, and understand disease progression3,4. These tasks demand nuanced understanding of how temporally distributed information influences current and future care-capabilities that current large language models (LLMs) often lack.
Although biomedical LLMs have demonstrated promising performance on well-structured tasks such as medical knowledge retrieval and standardized exams5,6,7, recent studies have revealed deficiencies in their ability to perform longitudinal reasoning8,9. The inability of LLMs to integrate temporally dispersed evidence and maintain chronological consistency undermines their clinical use. While temporal reasoning has been examined in general domains10,11,12, how these abilities scale to complex, real-world medical contexts remains poorly understood.
Instruction tuning has emerged as a key strategy in aligning models with desired objectives by adapting LLMs to domain-specific tasks via curated instruction-response pairs13,14. However, existing instruction datasets for clinical applications suffer from fundamental limitations that hinder temporal reasoning development. Traditional clinical question-answer pairs often originate from medical examination scenarios or simplified case vignettes that present idealized, neatly organized clinical narratives, sharply contrasting with the fragmented and time-spanning nature of authentic patient records. When instructions do come from real clinical sources, they generally focus on single-visit events with a very limited time span. For instance, the MIMIC-Instr dataset15 draws exclusively from brief hospitalizations, averaging 7.2 days. Additionally, physician-curated instruction responses impose a significant cognitive burden when reviewing extensive patient timelines, resulting in a focus on isolated information retrieval rather than sophisticated temporal synthesis. For example, over 70% of questions in MedAlign16 focus on simple retrieval tasks rather than reasoning across multiple clinical encounters over time.
These limitations expose a fundamental gap in clinical LLM development and evaluation. Our analysis of existing evaluation datasets reveals a pronounced temporal skew-for example, over 55% of questions in MedAlign targeting only the final 25% of patient timelines-failing to evaluate models’ capabilities to synthesize data over longitudinal records.
To address the critical challenge of longitudinal reasoning in clinical LLMs, we introduce TIMER (Temporal Instruction Modeling and Evaluation for Longitudinal Clinical Records)-a method for improving temporal understanding in clinical language models. TIMER grounds LLMs in specific patient temporal contexts by instruction tuning with timestamp-linked instruction-response pairs generated from longitudinal EHR data. Doing so enables models to recognize temporal patterns, understand event sequences, and reason across extended patient histories-all of which are essential for meaningful clinical applications but largely absent in current systems.
Figure 1 illustrates this approach with an example: a TIMER-generated instruction asks “Explain how the patient’s response to lung cancer treatment changed after discovering the EGFR mutation...” with the corresponding response referencing specific timestamps: “Patient started chemotherapy for lung cancer in 01/2020 with good response. Genetic testing in 05/2020 revealed EGFR mutation...” This demonstrates how each instruction-response pair is grounded in specific temporal contexts within the patient’s longitudinal record. For evaluation, we employ both clinician-curated benchmarks and controlled temporal sampling strategies, revealing previously unrecognized biases in existing evaluation methods.
TIMER enhances model performance through instruction tuning with timestamp-linked instruction-response pairs generated across longitudinal EHR timelines. Our evaluation employs both clinician-curated benchmarks and a controlled sampling strategy to create instruction sets with varying temporal distributions, enabling assessment of how models reason across different time periods in patient histories.
Our findings demonstrate that TIMER-tuned models consistently outperform both general-purpose models and conventional medical instruction tuning approaches across multiple evaluation settings. Notably, while conventional medical instruction tuning (such as MedInstruct) sometimes degrades performance compared to the base model on clinician-curated benchmarks, TIMER consistently improves performance on both general clinical tasks and temporal reasoning tasks. Through detailed case studies and quantitative evaluation, we show that TIMER-tuned models have higher temporal boundary adherence (correctly limiting analyses to specified time periods), trend detection (identifying meaningful patterns in longitudinal data), and chronological precision (correctly ordering and contextualizing clinical events). Our exploration of temporal distribution effects reveals that aligning training and evaluation enhances performance, with distribution-matched training consistently outperforming misaligned approaches across all settings. Collectively, our findings demonstrate that instruction tuning with TIMER results in models that can effectively support longitudinal clinical care and complex medical decision-making in real-world scenarios.
Results
Study design and data source
Figure 1 summarizes the TIMER method for instruction-tuning and evaluating temporal reasoning capabilities of LLMs on longitudinal EHRs. Our study addresses two primary research questions (RQs):
-
RQ1: Can temporally-grounded instruction-response pairs from EHR data improve LLMs’ longitudinal reasoning capabilities compared to conventional medical question-answer pairs?
-
RQ2: How does the temporal distribution of instructions used in instruction-tuning affect model performance, specifically when we evaluate on varying temporal distributions?
We used de-identified longitudinal EHRs from the Stanford Medicine Research Data Repository (STARR)17. These records are accessible pre-IRB, cover Stanford Health Care (primarily adult care) and Lucile Packard Children’s Hospital, and are formatted in OMOP-CDM. Only data from patients who have previously consented to the research use of their de-identified data via the institutional privacy notice is included in STARR.
RQ1: Impact of temporal-aware instruction tuning
We compared models instruction-tuned with TIMER against both standard medical LLMs and models tuned with conventional medical QA datasets to quantify the specific benefits of temporal awareness in instruction tuning.
Overall performance
We evaluate multiple medical LLMs, including Meditron-7B, MedAlpaca, AlpaCare, MMed-Llama-3-8B, PMC-Llama-13B, MedLM-Medium, and MedInstruct (Conventional QA-tuned Llama-3.1-8 B-Instruct), using MedAlign and a model-generated evaluation set, called TIMER-Eval, that requires temporal reasoning. When models cannot process full patient timelines, inputs are truncated to recent tokens of their context limits. As shown in Table 1, even the strongest medical model baseline achieves just 30.85% correctness and 13.93% completeness in temporal reasoning evaluations. In contrast, models tuned with TIMER consistently outperform baselines across both evaluation sets. TIMER improves Llama-3.1-8B-Instruct’s performance from 30.69% to 34.32% correctness as measured using MedAlign and from 45.02% to 48.51% as measured via a temporal reasoning evaluation. Similar improvements are observed with Qwen-2.5-7B-Instruct, indicating that these gains are consistent across different base model architectures. TIMER’s performance gains on MedAlign are particularly significant given the dataset’s temporal characteristics. Despite MedAlign’s extended temporal coverage (median 3,895.1 days) and pronounced recency bias (55.3% of questions in the final 25% of timelines), TIMER-tuned models achieve consistent improvements in both correctness and completeness. This demonstrates TIMER’s ability to effectively utilize the full temporal scope of longitudinal records, even when evaluation questions exhibit temporal distribution misalignment.
To illustrate these improvements, Table 7 provides concrete examples of enhanced temporal reasoning. For instance, when asked to “Describe the trend in the patient’s weight over the past year” base models incorrectly assess trends from 2+ years prior, while TIMER-tuned models correctly limit analysis to the specified timeframe, demonstrating improved temporal boundary adherence.
Head-to-head comparison
To further identify performance improvement in addition to rubric-based scoring, we perform head-to-head analyses of outputs to given questions from different models. We identify in Table 2 that models instruction-tuned with temporal instruction data produce answers that are more generally preferred compared to existing medical finetuned models, with even the best medical model MedLM-Medium being preferred 20% less frequently on TIMER-Eval generated questions than models tuned with TIMER.
Additionally, we compare conventional QA-style tuning (MedInstruct) with our temporally grounded instruction-tuning (TIMER Tuning). While instruction-tuning with MedInstruct provided gains over baseline Llama-3.1-8B-Instruct performance, instruction tuning with TIMER provides additional gains of 6.3% on MedAlign and 8.45% on TIMER-Eval. This indicates the value of incorporating temporal structure into instruction tuning data.
RQ2: Effect of the temporal distribution of instructions on model performance
Temporal biases in existing clinical instruction sets
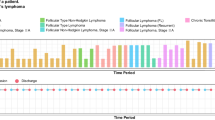
Existing clinical instruction data have pronounced temporal biases. Using our normalized temporal position metric, we found that MedAlign16, the first clinician-curated collection of clinical instructions, has a pronounced recency bias. Despite spanning an average of 3895 days (~10.7 years), 55.3% of its instructions reference only the final 25% of patient timelines, with 47.0% and 29.5% focused on just the last 15% and 5%, respectively (Fig. 2).
The majority of human-generated instructions focus on the most recent encounters.
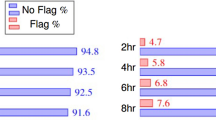
When examining model-generated instructions, we observed a “lost-in-the-middle" effect regarding the parts of the patient record in which the instructions were grounded (Fig. 3). These instructions cluster at the beginning (25.9%) and end (52.1%) of patient timelines while relatively underrepresenting middle periods (22.1%). These distribution biases in both human and model-generated instructions highlight the need for a more controlled approach to both instruction generation and evaluation.
Instructions cluster around early (0–25%) and late (75–100%) parts of patient timelines.
Temporal duration vs. utilization
An important distinction emerges between the temporal duration available in clinical records and the actual temporal scope utilized in instruction generation. MedAlign’s construction involved clinicians generating instructions independently without examining specific patient records, with these instructions subsequently matched to appropriate EHR data via retrieval. The pronounced recency bias (55.3% of instructions in the final 25% of timelines) reveals that MedAlign reflects the natural recency bias inherent in clinical question formulation when clinicians pose questions independent of specific patient cases. This observation motivates the need for systematic temporal control approaches that ensure evaluation coverage across extended patient timelines.
Development of controlled temporal distribution evaluation
Our analysis of existing temporal biases necessitated a new evaluation approach that could isolate and measure the specific effects of temporal distribution on model performance. Unlike existing approaches with inherent limitations (Table 3), we developed an evaluation method incorporating multi-visit records with explicit time evidence attribution. Table 3 highlights how TIMER-Eval overcomes limitations in prior approaches. MIMIC-Instr15 is restricted to single-visit episodes with limited temporal scope (median 7.2 days), while MedAlign’s human curation leads to recency bias. Our approach enables precise assessment of temporal reasoning while maintaining scalability through controlled sampling, allowing systematic manipulation of temporal distribution patterns in both instruction tuning and evaluation.
Clinician validation
To ensure the validity of the model-generated evaluation data, three clinicians assessed 100 randomly sampled instruction-response pairs generated with TIMER (Table 4). The pairs received high scores for clinical relevance (mean 95/100), temporal reasoning complexity (mean 80/100), and factual accuracy (mean 98/100), with strong inter-rater agreement. The results show high inter-rater agreement (86% clinical relevance, 93% accuracy) with low standard deviations (4.32, 1.89, respectively). Complexity scoring, being an inherently more qualitative metric, showed increased variability but remained significantly above chance (53% observed agreement vs. 12.5% random chance; std 14.87). Additional examples of where annotators agreed and disagreed on question complexity can be found in Supplementary Note 7. Disagreements primarily occurred on questions requiring temporal data retrieval with moderate synthesis, where annotators differed on whether the reasoning depth met clinical complexity standards. This variability demonstrates the rigor with which experienced clinicians evaluate temporal reasoning tasks, distinguishing between temporal mechanics and genuine clinical reasoning complexity. These validation results support the claim that our schema generates clinically meaningful evaluation scenarios.
Effect of instruction distributions on model performance
To understand how temporal distributions affect model performance, we created three distinct distribution patterns: Recency-Focused, Edge-Focused, and Uniformly-Distributed across timelines.
Table 5 demonstrates that across all evaluation patterns, models using distribution-matched training consistently outperform alternative training approaches. The advantage of matched training ranges from +1.20% to +6.50% in head-to-head comparisons. Notably, the largest performance difference appears in the Uniformly-Distributed evaluation setting. For instance, when evaluating on Uniformly-Distributed questions, Full-Timeline training shows a +6.50% advantage over Recent-Events training, highlighting how distribution alignment affects model performance on temporal reasoning tasks. These results indicate that while general alignment between train and test distribution is helpful, it provides the most significant gains in the harder evaluation schema of Uniformly-Distributed instructions.
LLM-judge and human correlation
To scale evaluation, we developed an LLM-based judge and validated it against clinician rankings for MedAlign responses. Table 6 shows a strong Spearman correlation between LLM scores and human ranks: ρ = −0.97 (average), −0.94 (correctness), and −0.89 (completeness). This inverse relationship (high LLM score = low human rank) supports that LLM-judge is a reliable proxy for human assessment in temporal reasoning evaluation. We additionally show the LLM Rank for a more direct comparison to human rank, and see general ranking trends being maintained across LLMs.
Case studies: temporal reasoning behavior
Table 7 presents qualitative examples comparing base and TIMER tuned models. TIMER-tuned models consistently show improved:
-
1.
Temporal boundary adherence (e.g., limiting responses to the past year),
-
2.
Trend detection (e.g., correctly summarizing longitudinal lab trends), and
-
3.
Temporal precision (e.g., associating measurements with exact dates).
In contrast, base models often conflate visits or provide temporally irrelevant information. Responses of models tuned with TIMER are more contextually grounded and clinically interpretable.
Discussion
Our results demonstrate that TIMER significantly enhances LLMs’ ability to reason over longitudinal clinical records through temporal-aware instruction tuning. By grounding instruction-response pairs at specific timestamps within patient histories, TIMER enables models to better integrate evidence across extended timeframes-a capability essential for clinical applications. Our evaluation reveals that models fine-tuned using the TIMER method consistently outperform those trained with standard medical question-answer pairs, particularly on tasks requiring synthesis across multiple timepoints.
We also identify shortcomings in current clinical LLM evaluation techniques regarding timeline length, question complexity, and temporal coverage. We find that existing evaluations often exhibit strong recency bias, with most questions focusing on recent visits rather than comprehensive patient histories. By developing evaluation approaches that explicitly sample across different points in a patient timeline, we demonstrate how the alignment between instruction distribution and evaluation context impacts model performance. This insight provides a methodological foundation for future development of LLMs capable of sophisticated temporal reasoning in clinical settings.
Our findings demonstrate the critical importance of temporal context when tuning LLMs for use with longitudinal clinical records, which is necessary for applications such as disease trajectory modeling, therapeutic response tracking, and longitudinal summarization. The performance improvements achieved through TIMER-a 39.50% win-rate over MedLM-Medium and 23.80% over the base model-reveal that current language models (both medical and general) lack the temporal reasoning capabilities necessary for clinical applications. The 6.3% advantage over traditional QA instruction tuning further emphasizes that temporal reasoning is a distinct capability that can be imbued with specialized training approaches. These results suggest that mere exposure to medical content is insufficient; what matters is how models learn to integrate information across time. By explicitly providing temporally-grounded instructions, TIMER instruction tunes models in a manner that mirrors the time-aware reasoning processes models have to support in healthcare workflows.
Our analysis reveals two key deficiencies in current medical evaluations: they primarily rely on single-timepoint data retrieval and exhibit recency bias, overemphasizing recent patient information while neglecting longer-term clinical patterns. TIMER mitigates this bias by sampling instructions generated from across the patient timeline. This approach increases instruction diversity and encourages models to learn more generalizable temporal patterns. Notably, ablations of TIMER’s sampling strategy reveal that while aligning training distribution to evaluation distribution of instructions consistently improves model performance, this alignment is most critical with Uniformly-Distributed evaluation questions, with Full-Timeline instruction-tuned models outperforming Recent-Events instruction-tuned models by 6.5%.
Human evaluation of a subset of model-generated instructions in TIMER-Eval rated the generated instructions as an average of 95/100 relevant, 80/100 complex, and 98/100 accurate. However, the use of model-generated data to investigate RQ2 remains a limitation, as these examples may not fully capture physician reasoning or align with clinical documentation patterns. Additionally, all model-generated instruction data cannot be easily screened by clinicians; therefore, this sampling assumes that other questions fall within the same distribution of quality.
This study has several limitations. The model-based generation process may encode training data biases, and while we include some manual review, large-scale validation is still needed. We do not yet assess fairness across demographic subgroups or calibration of the temporal reasoning outputs. Future work should include this fairness analysis to further ensure that generated training and evaluation data adequately represent patient subgroups. Additional work could explore expanding the modalities included in TIMER to further improve the richness of queries that can be posed to models (e.g., reasoning over labs, imaging, etc.). Integrating the generation of evaluation questions with real-world deployment tasks could offer more insights into clinical impact and safety. Developing methods for scalable verification of model-generated data would enable more refined evaluation that precisely measures model capabilities for specific clinical tasks. In addition, increasing the scope of models that are both fine-tuned and evaluated with TIMER would provide more insight into the capabilities of frontier models on medical temporal reasoning tasks. Extending the current fine-tuning framework to medical models such as Me-LLaMA or Meditron-70B would further illustrate the impact of fine-tuning for temporal reasoning tasks18,19.
In conclusion, our results establish TIMER as a practical method for improving temporal reasoning capabilities in clinical LLMs. TIMER improves generalization and performance across model architectures and evaluation datasets. Llama-3.1-8B-Instruct instruction-tuned with TIMER, as well as Qwen-2.5-8B-Instruct instruction-tuned with TIMER, both demonstrate improvements compared to the base model for both physician-generated questions (MedAlign) and synthetically generated questions (TIMER-Eval). Instruction tuning that reflects the temporal complexity of healthcare data yields substantial performance improvements and better aligns models with the needs of longitudinal clinical tasks. From a deployment standpoint, our findings highlight the potential of time-aware instruction tuning to enhance retrospective chart review, and forecasting tasks. Because we release the TIMER method open-source, it allows any group with access to EHR to generate instruction tuning data as well as the evaluation set to examine and improve temporal reasoning capabilities20.
Methods
Data source and preprocessing
We sourced the data for our study from the Stanford Medicine Research Data Repository (STARR)17, which contains EHR data from Stanford Health Care (primarily adult care) and Lucile Packard Children’s Hospital (primarily pediatric care). The source dataset is structured according to the Observational Medical Outcomes Partnership Common Data Model (OMOP-CDM) and contains records covering the time period from 1990 to February 8th, 2023. Patients had previously consented to the research use of their de-identified data via the institutional privacy notice. All data were de-identified, exempting this study from Institutional Review Board (IRB) review. Patient timelines were segmented into context-sized chunks (16,000 tokens) suitable for LLM processing.
TIMER methodology and technical design
TIMER uses timestamp-linked instruction-response pairs generated from longitudinal EHR data to fine-tune language models for improved temporal reasoning. The method leverages explicit time evidence integration to ground model responses in specific temporal contexts within patient histories. Figure 1 illustrates the overall approach, from EHR processing to instruction generation, instruction tuning, and evaluation.
Temporal instruction-tuning data generation
To generate temporally grounded instruction-response pairs for our instruction-tuning dataset, we used Gemini-1.5-Pro21 with carefully designed prompts that instructed the model to reference specific timestamps within patient records. The prompt template is described in Supplementary Note 2. These prompts specifically instruct models to “emphasize the temporal progression of clinical events” and require responses to include “specific timestamps (dates only, in the format MM/DD/YYYY) to justify responses and highlight temporal reasoning” (see Supplementary Note 2 for complete templates). We generated 5000 instruction-response pairs with explicit temporal grounding for each distribution pattern (Recent-Events, Timeline-Extremes, and Full-Timeline).
Model architecture and training details
We instruction-tuned Llama-3.1-8B-Instruct22,23 (8B parameters, 16K context) for 6 epochs over datasets of 5000 examples with an effective batch size of 16. We performed a hyperparameter search to identify optimal parameters. Training details are provided in Supplementary Note 3. We additionally instruction-tuned Qwen-2.5-7B-Instruct24 to demonstrate generalization across model architectures.
Evaluation metrics
We evaluated model responses using both automated metrics and LLM-based judges. For LLM-based evaluation, we used GPT-4o-mini to assess response correctness and completeness. The prompts used for LLM-Judge scoring are available in Supplementary Note 4. We validated this approach by comparing LLM-judge scores with clinician ratings, finding high correlation (correctness: ∣ρcorr∣ = 0.94; completeness: ∣ρcorr∣ = 0.89), as detailed in Section “LLM-Judge and Human Correlation”.
To provide additional evaluation insight, we also calculated standard automated metrics: BERTScore25 (DistilBERT-uncased), ROUGE-L26, CHRF27, and METEOR28. Details for these metrics are included in Supplementary Note 6.
Temporal reasoning instruction generation for evaluation
To evaluate longitudinal temporal reasoning, we created TIMER-Eval, an evaluation schema that generates clinical questions requiring evidence integration from multiple timepoints. We used Gemini-1.5-Pro21 to generate candidate questions from patient timelines disjoint from those used in instruction tuning. Each instance includes the question, answer, and associated timestamps \({{\bf{T}}}_{i}=\{{T}_{i,1},...,{T}_{i,{n}_{i}}\}\) corresponding to relevant timeline events.
Questions were filtered to retain only those requiring synthesis across at least two distinct, time-stamped evidence snippets, ensuring each requires genuine temporal reasoning rather than simple retrieval. For instance, questions like “What medication was prescribed on 03/15/2020?” are filtered out as simple retrieval, while questions such as “How did the patient’s treatment response change between the initial chemotherapy in 2020 and the targeted therapy switch in 2021?” could be retained as they require temporal synthesis across multiple timepoints. The prompt template used for question generation is provided in Supplementary Note 1. A validation protocol involving three clinicians was implemented to assess clinical relevance, temporal reasoning complexity, and factual accuracy (detailed in Section “RQ2: Effect of the Temporal Distribution of Instructions on Model Performance”).
Our quality control process included both automatic and manual review stages. Automatic filtering retained questions requiring synthesis across at least two distinct, timestamped evidence snippets from different time periods. Manual review subsequently eliminated low-quality questions that failed to meet temporal reasoning standards or contained factual inconsistencies. This two-stage process ensures that retained questions genuinely require longitudinal reasoning capabilities rather than simple information retrieval.
Normalized temporal position metric
To analyze temporal distribution patterns in clinical instructions, we defined a normalized position metric that enables consistent comparison across patient records of varying lengths. This normalization is applied to all datasets in this study. For each evidence timestamp Tj in an instruction-response pair (Qi, Ai), we compute the relative position:
This formulation standardizes temporal positions to a [0,1] range while preserving the temporal order, allowing cross-record comparison of evidence distribution patterns.
Temporal distribution sampling strategy
Based on our analysis of temporal distributions in existing datasets, we constructed three instruction-tuning datasets with matched size and source timelines, varying only in the distribution of instruction placement:
-
Recent-Events: Instructions placed in the final quartile of patient timelines, mirroring the recency bias found in many real-world and annotated datasets.
-
Timeline-Extremes: Instructions concentrated at the beginning and end of timelines, simulating LLM-generated data patterns.
-
Full-Timeline: Instructions uniformly distributed across the entire timeline, ensuring balanced temporal coverage.
This design isolates temporal placement as the sole variable, allowing evaluation of how different distributions affect models’ capacity for temporal reasoning across patient records.
Data availability
The datasets supporting the conclusions of this article are available with restrictions. The TIMER-Eval benchmark dataset generated for this study contains synthetic instruction-response pairs derived from clinical scenarios. The synthetic Electronic Health Record (EHR) data used for instruction-tuning was generated using Gemini-1.5-Pro based on prompts described in the Supplementary Information and is available under the same terms. A minimal synthetic dataset representative of the data structure and format will be released publicly alongside the code repository to facilitate understanding and reproduction of the methods. The original clinical data underlying the synthetic generation cannot be shared due to privacy and confidentiality requirements. The MedAlign16 benchmark used for comparative evaluation is publicly available on Stanford redivis: https://stanford.redivis.com/datasets/48nr-frxd97exb.
Code availability
The source code for TIMER is available at https://github.com/som-shahlab/TIMER. This includes scripts for data generation, instruction-tuning pipelines, evaluation benchmarks, automatic and LLM judge metrics, and all statistical analysis code used to support the findings of this study. The code is implemented in Python and can be reproduced on systems with NVIDIA H100 GPU or equivalent hardware. Documentation and installation instructions are provided in the repository.
References
Li, T., Zhang, G., Do, Q. D., Yue, X., & Chen, W. Long-context LLMs Struggle with Long In-context Learning. Trans. Mach. Learn Res. https://openreview.net/forum?id=Cw2xlg0e46 (2025).
Kuratov, Y. et al. BABILong: Testing the Limits of LLMs with Long Context Reasoning-in-a-Haystack. In A. Globerson, L. et al. (eds.) Advances in Neural Information Processing Systems (Vol. 37, pp 106519–106554) (2024).
Huguet, N. et al. Using electronic health records in longitudinal studies: estimating patient attrition. Med. Care 58, S46–S52 (2020).
Wornow, M. et al. Context Clues: Evaluating Long Context Models for Clinical Prediction Tasks on EHR Data. The Thirteenth International Conference on Learning Representations. https://openreview.net/forum?id=zg3ec1TdAP (2025).
Singhal, K. et al. Large language models encode clinical knowledge. Nature 620, 172–180 (2023).
Lu, Z. et al. Large language models in biomedicine and health: current research landscape and future directions. J. Am. Med. Inform. Assoc. 31, 1801–1811 (2024).
Lucas, M. M., Yang, J., Pomeroy, J. K. & Yang, C. C. Reasoning with large language models for medical question answering. J. Am. Med. Inform. Assoc. 31, 1964–1975 (2024).
Hager, P. et al. Evaluating and mitigating limitations of large language models in clinical decision making. Nat. Med. 30, 2613–2622 (2024).
Bedi, S. et al. Testing and Evaluation of Health Care Applications of Large Language Models: A Systematic Review. JAMA 333, 319–328, https://doi.org/10.1001/jama.2024.21700 (2025).
Wang, Y., & Zhao, Y. TRAM: Benchmarking Temporal Reasoning for Large Language Models. In L.-W. Ku, A. Martins, & V. Srikumar (eds.) Findings of the Association for Computational Linguistics, ACL 2024, Bangkok, Thailand and virtual meeting, 2024 (pp. 6389–6415). Association for Computational Linguistics. https://doi.org/10.18653/V1/2024.FINDINGS-ACL.382 (2024).
Fatemi, B. et al. Test of Time: A Benchmark for Evaluating LLMs on Temporal Reasoning. The Thirteenth International Conference on Learning Representations, ICLR 2025, Singapore, 2025. https://openreview.net/forum?id=44CoQe6VCq (2025).
Herel, D., Bartek, V. & Mikolov, T. Time awareness in large language models: benchmarking fact recall across time. Preprint at https://arxiv.org/abs/2409.13338 (2024).
Ouyang, L. et al. Training language models to follow instructions with human feedback. NeurIPS 35, 27730–27744 (2022).
Zhang, S. et al. Instruction tuning for large language models: a survey. Preprint at https://arxiv.org/abs/2308.10792 (2023).
Wu, Z., Dadu, A., Nalls, M., Faghri, F. & Sun, J. Instruction tuning large language models to understand electronic health records. In NeurIPS Datasets and Benchmarks Track https://openreview.net/forum?id=Dgy5WVgPd2 (2024).
Fleming, S. L. et al. MedAlign: A Clinician-Generated Dataset for Instruction Following with Electronic Medical Records. In M. J. Wooldridge, J. G. Dy, & S. Natarajan (eds.) Thirty-Eighth AAAI Conference on Artificial Intelligence, AAAI 2024, Thirty-Sixth Conference on Innovative Applications of Artificial Intelligence, IAAI 2024, Fourteenth Symposium on Educational Advances in Artificial Intelligence, EAAI 2014, February 20-27, 2024, Vancouver, Canada (pp. 22021–22030). https://doi.org/10.1609/AAAI.V38I20.30205 (AAAI Press, 2024).
Datta, S. et al. A new paradigm for accelerating clinical data science at Stanford medicine. Preprint at https://arxiv.org/abs/2003.10534 (2020).
Xie, Q. et al. Medical foundation large language models for comprehensive text analysis and beyond. npj Digit. Med. 8, 141 https://doi.org/10.1038/s41746-025-01533-1 (2025).
Chen, Z. et al. Meditron-70b: scaling medical pretraining for large language models. Preprint at https://arxiv.org/abs/2311.16079 (2023).
Zeng, D., Qin, Y., Sheng, B. & Wong, T. Y. Deepseek’s “low-cost" adoption across china’s hospital systems: Too fast, too soon? JAMA 333, 1866–1869 (2025).
Team, G. et al. Gemini 1.5: unlocking multimodal understanding across millions of tokens of context. Preprint at https://arxiv.org/abs/2403.05530 (2024).
Llama, M. Llama-3.1-8b-instruct. https://huggingface.co/meta-llama/Llama-3.1-8B-Instruct (2024).
Hu, E. J. et al. LoRA: Low-Rank Adaptation of Large Language Models. The Tenth International Conference on Learning Representations, ICLR 2022, Virtual Event, 2022. https://openreview.net/forum?id=nZeVKeeFYf9 (2022).
Qwen et al. Qwen2.5: A party of foundation models (2025).
Zhang, T., Kishore, V., Wu, F., Weinberger, K. Q., & Artzi, Y. BERTScore: Evaluating Text Generation with BERT. 8th International Conference on Learning Representations, ICLR 2020, Addis Ababa, Ethiopia, 2020. https://openreview.net/forum?id=SkeHuCVFDr (2020).
Chin-Yew Lin. ROUGE: A Package for Automatic Evaluation of Summaries. In Text Summarization Branches Out, pages 74–81, Barcelona, Spain. Association for Computational Linguistics. (2004).
Maja Popović. chrF: character n-gram F-score for automatic MT evaluation. In Proceedings of the Tenth Workshop on Statistical Machine Translation, pages 392–395, Lisbon, Portugal. Association for Computational Linguistics. (2015).
Satanjeev Banerjee and Alon Lavie. METEOR: An Automatic Metric for MT Evaluation with Improved Correlation with Human Judgments. In Proceedings of the ACL Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization, pages 65–72, Ann Arbor, Michigan. Association for Computational Linguistics. (2005).
Acknowledgements
We thank Kameron Black, Mehr Kashyap, and Akshay Swaminathan for their valuable clinical expertise in evaluating the quality, relevance, and accuracy of the instruction-response pairs used in this study. Their insights were instrumental in validating our approach and ensuring its clinical meaningfulness.
Author information
Authors and Affiliations
Contributions
H.C., A.U., and J.A.F. conceptualized the study and defined the research objectives. A.U. and H.C. led the development of the machine learning framework, implemented the training procedures, and evaluated models across datasets. H.C. developed the data preprocessing and generation pipelines, ensuring compatibility with the LLM workflow. B.C. contributed to the ablation studies evaluating model robustness and benchmarking against existing LLMs. H.C. and A.U. drafted the manuscript. E.A., S.K., and N.S. provided methodological contributions and guidance, with N.S. additionally providing computational resources for experimental execution. All authors reviewed the manuscript for critical content, contributed to revisions, and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cui, H., Unell, A., Chen, B. et al. TIMER: temporal instruction modeling and evaluation for longitudinal clinical records. npj Digit. Med. 8, 577 (2025). https://doi.org/10.1038/s41746-025-01965-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01965-9
This article is cited by
-
TempReasoner: neural temporal graph networks for event timeline construction
Scientific Reports (2026)