Abstract
Continuous prediction of glucose levels and hypoglycemia events is critical for managing type 1 diabetes mellitus (T1DM) under intensive insulin therapy. Existing models focus on a single task, limiting their practicality and adaptability in automated insulin delivery (AID) systems. To address this, a domain-agnostic continual multi-task learning (DA-CMTL) framework that simultaneously performs glucose level forecasting and hypoglycemia event classification within a unified framework is proposed. Trained on simulated datasets via Sim2Real transfer and adapted using elastic weight consolidation, DA-CMTL supports cross-domain generalization. Evaluation on public datasets (DiaTrend, OhioT1DM, and ShanghaiT1DM) yielded a root mean squared error of 14.01 mg/dL, mean absolute error of 10.03 mg/dL, and sensitivity/specificity of 92.13%/94.28% on 30 min prediction. Real-world validation using diabetes-induced rats demonstrated a reduction in time below range from 3.01% to 2.58%, supporting reliable integration as a safety layer in AID systems. These results highlight DA-CMTL’s robustness, scalability, and potential to improve safety in AID.
Similar content being viewed by others
Introduction
Intensive insulin therapy is essential for blood glucose management in individuals with T1DM. In this context, the automated insulin delivery (AID) system, also known as the artificial pancreas system, has led to remarkable advancements in T1DM management by enhancing the accuracy and convenience of insulin administration1,2,3. This system comprises continuous glucose monitoring (CGM) sensors, insulin pumps, and control algorithms that process CGM glucose readings and user inputs to dynamically adjust insulin delivery in real time. This approach helps maintain optimal time in range (TIR)—the proportion of time glucose readings remain between 70 and 180 mg/dL4,5. Numerous clinical studies have validated the effectiveness of AID systems, revealing substantial improvements in glycemic outcomes such as increased TIR and reduced HbA1c levels6,7,8,9. For instance, a UK study demonstrated a reduction in HbA1c from 9.4% to 7.8% and a notable increase in TIR from 34.2% to 61.7% among adults using AID7. Additionally, Brown et al. observed TIR improvements of 15.6% in children and 9.3% in adults, further confirming the benefits of AID systems across different age groups8.
As the adoption of AID systems continues to expand, the development of accurate glucose level prediction and robust hypoglycemia event classification algorithms is essential to ensure the safety and efficacy of insulin therapy. This need is clarified by two key considerations. First, correction boluses are frequently required, even after the administration of a meal bolus4. For example, an analysis of 996 insulin pump users in the United States found that correction boluses accounted for ~12% of the total daily insulin dose, reflecting a significant burden of postprandial hyperglycemia10. This requirement is largely attributed to delayed insulin dosing, inaccuracies in carbohydrate estimation, and the conservative dosing strategies utilized by many current AID algorithms. Accurate glucose forecasting is therefore critical to enable timely and autonomous correction dosing, thereby supporting optimal TIR. Second, AID systems represent a form of intensive insulin therapy, inherently increasing the risk of hypoglycemia11. Given the potential severity of hypoglycemic events (e.g., seizures, coma, and vision impairment), early and reliable detection is essential to enable prompt suspension of insulin delivery and mitigate adverse outcomes. Such safety mechanisms are fundamental to the successful and responsible deployment of AID technologies.
Recent advancements in prediction algorithms for AID systems have increased interest in deep learning (DL)-based glucose prediction and hypoglycemia event classification12. DL models offer the capacity to learn complex nonlinear relationships between glucose dynamics and various physiological and behavioral factors (e.g., insulin, meals, patient characteristics), thereby improving glycemic prediction performance. In glucose level prediction, Pérez-Gandía et al. first applied artificial neural networks for glucose prediction, laying the foundation for subsequent research13. Recurrent neural networks (RNNs), such as long short-term memory (LSTM) and gated recurrent units (GRU), have since gained prominence due to their effectiveness in modeling temporal dependencies14,15,16. For instance, Martinsson et al. reported a root mean squared error (RMSE) of 18.87 mg/dL on the OhioT1DM dataset using an LSTM-based model with a 30 min prediction horizon (PH)17, while Alshehri et al. demonstrated the efficiency of GRU-based models, citing comparable accuracy with reduced computational complexity18. More recently, advanced models have emerged to further enhance predictive performance. Zhu et al. employed a temporal fusion transformer (TFT), achieving RMSEs of 19.10 and 12.70 mg/dL on the OhioT1DM and ShanghaiT1DM, respectively, under a 30-minute PH19. Piao et al. introduced a graph attentive RNN (GARNN) that yielded RMSEs of 18.97 and 13.62 mg/dL on the same datasets20. In parallel, Montaser et al. proposed seasonal local modeling frameworks for glucose prediction using variable-length, time-stamped events, demonstrating strong adaptability and performance across patient-specific trajectories21. These efforts collectively reflect a growing emphasis on capturing individual variability, temporal complexity, and multimodal dependencies. However, many models remain vulnerable to overfitting on dataset-specific patterns, limiting their applicability to diverse populations and undermining generalizability across domains16,22,23,24. While glucose forecasting has progressed as a distinct research pillar, hypoglycemia event prediction has evolved largely in parallel, using both feature-based and time series–based methodologies25. Earlier approaches relied on handcrafted features such as CGM trends, insulin-on-board (IOB), and carbohydrates-on-board (COB), processed through traditional machine learning (ML) models such as random forests (30 min PH; sensitivity/specificity: 89.6/91.3%), support vector regression (30 min PH; 96.0/97.0%), support vector machine (100% sensitivity in 17 cases). Although these models offered computational simplicity, they were often inadequate for modeling dynamic glucose fluctuations, especially under postprandial, nocturnal, or exercise-induced variability26,27,28. Subsequent approaches leveraged DL architectures such as deep belief networks and fully connected neural networks (FCNNs) to capture richer feature representations and improve sensitivity29. However, their reliance on manually engineered inputs limited scalability and hindered integration with glucose forecasting frameworks. More recent models have shifted toward time series-based classifiers that directly learn from CGM glucose readings. For example, LSTM-based30 and transformer-based31 architectures have demonstrated improved performance and generalization across populations. Notably, the LSTM model proposed by Shao et al., trained on Chinese patient data, outperformed traditional ML baselines when tested on European-American cohorts, highlighting cross-domain potential. Despite these advances, most hypoglycemia prediction models remain decoupled from glucose forecasting pipelines or are developed in isolation for specific datasets. Treating prediction and classification as independent tasks impairs the coordinated operation required for real-time insulin delivery systems. In particular, this separation leads to a fragmented model architecture, necessitating distinct inference pathways for each task and resulting in asynchronous outputs. Such a design increases computational burden and latency, which directly undermines the feasibility of deploying responsive and unified feedback mechanisms essential for closed-loop control in AID systems. Beyond algorithmic development, practical and ethical barriers further hinder the deployment of adaptive AID systems. Collecting real-world data from a single patient over one year is estimated to cost at least $2940, excluding the time-consuming processes of recruitment, experimentation, and ethical approval, which can take a minimum of one year32. Additionally, privacy concerns remain a major barrier to data sharing, with over 20 studies citing them as a leading issue in healthcare AI deployment33. Thus, integration, generalization, and large-scale real-world data collection remain critical challenges to be addressed for reliable and responsible deployment.
In summary, despite promising advancements, current DL-based research is limited by three key factors: system complexity resulting from task separation, poor generalization across populations, and high dependence on real-world data34. These limitations hinder the deployment of scalable and adaptive insulin delivery solutions. To address these challenges, we propose a domain-agnostic continual multi-task learning (DA-CMTL) framework that leverages simulation-to-real (Sim2Real) transfer. DA-CMTL adopts a multi-head architecture to jointly perform glucose prediction and hypoglycemia event classification, allowing task-specific modeling while leveraging shared temporal features. This unified design enhances task synergy and supports efficient inference within real-time AID applications. Moreover, Sim2Real transfer—where a model trained in simulated environments is deployed in real-world applications—is employed to enable generalization while reducing the cost associated with real-world data collection. In this context, we conceptualize Sim2Real not merely as a data domain transfer but as a generalization strategy that leverages physiologically diverse, simulated scenarios to build robust representations. Crucially, model performance was ultimately evaluated on real-world datasets collected under free-living conditions, confirming its applicability beyond controlled simulations. Unlike traditional methods, such as data augmentation or SMOTE, which improve data diversity by modifying or oversampling existing samples, Sim2Real leverages physiologically validated simulators to generate synthetic patient profiles with systematic variability. This includes adverse and infrequent conditions, such as prolonged hypoglycemia or atypical glucose–insulin responses, which are often underrepresented in real-world datasets. Incorporating such difficult-to-capture scenarios into training enables the model to generalize across a broader range of clinically meaningful conditions while reducing reliance on large-scale data collection from real patients. Similar approaches have been applied in various healthcare AI, including diabetes management35, tumor segmentation36, and autonomous surgical navigation37, demonstrating that simulation-based training can achieve high performance and safety in clinical environments. While simulated data enables scalable training, the transition to real-world application introduces a critical barrier: domain shift, which arises from discrepancies in data characteristics between simulated and real-world environments. These distributional differences can degrade model performance when applied across domains. To address this issue, elastic weight consolidation (EWC), a continual learning (CL) method, is incorporated. EWC facilitates sequential learning from diverse simulated datasets while preventing catastrophic forgetting by introducing a regularization term in the loss function38. This allows the model to retain knowledge from previously learned domains, thereby improving robustness and ensuring reliable deployment in practical settings. We refer to this property as domain-agnostic, indicating the model’s ability to generalize without relying on domain-specific adaptation. The proposed DA-CMTL framework advances algorithm development for insulin delivery systems through the integration of multi-task learning (MTL) while mitigating both data acquisition costs and domain-specific performance issues via Sim2Real transfer and CL techniques.
The primary contributions of this research are threefold:
-
We propose a unified multi-task framework that jointly models glucose prediction and hypoglycemia event classification, addressing the issue of task separation and structural inefficiency in current AID system pipelines.
-
We introduce a novel Sim2Real transfer strategy, enabling scalable model training while minimizing reliance on labeled real-world data.
-
We implement continual learning with elastic weight consolidation to support domain-agnostic generalization across multiple datasets without data retention.
The remainder of this paper is organized as follows: section “Results” presents our experimental results, section “Discussion” discusses key findings and clinical implications, and section “Methods” details the model architecture and training methodology.
Results
Study design and overview
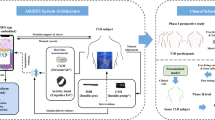
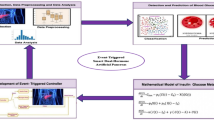
This study introduces a generalized DA-CMTL framework developed for reliable glucose level prediction and hypoglycemia event classification across diverse real-world environments. The model integrates three core components: MTL, CL, and Sim2Real transfer, as illustrated in Fig. 1. To achieve this integration, a multi-head architecture was utilized to enable simultaneous learning of both glucose prediction and hypoglycemia event detection tasks, thereby optimizing an integrated loss function (Eq. (11)) to improve performance.
The DA-CMTL framework integrates multi-task learning (MTL), Sim2Real transfer, and continual learning (CL) to enhance glucose prediction and hypoglycemia event detection. a MTL uses a multi-head architecture to simultaneously predict glucose levels and detect hypoglycemia events via CGM glucose readings and IOB features. b Sim2Real transfer improves real-world adaptability by pre-training the model on virtual patient data generated by the UVA/Padova simulator before clinical deployment. c CL mitigates domain shift by sequentially training the model on simulated datasets that represent diverse patient characteristics.
To enhance the model’s generalization capability, synthetic datasets representing a wide range of patient distributions—including variations in glucose dynamics, demographics, and physiological traits—were generated using the FDA-approved UVA/Padova simulator, which emulates virtual patients based on validated physiological glucose-insulin interaction models and simulates realistic daily scenarios such as meal intake, insulin dosing. To effectively learn from these heterogeneous datasets, the CL framework was adapted. This framework enables the model to train progressively to progressively acquire knowledge from diverse simulated domains while preserving previously learned information. By preventing catastrophic forgetting and promoting balanced learning across sequential datasets, the CL strategy (Eq. (10)) mitigates performance degradation caused by domain shift and supports the development of a more robust and adaptable model. In our implementation, both MTL and CL components are jointly optimized through a unified training objective (Eq. (12)), allowing seamless integration of task- and domain-level learning dynamics. The trained model was subsequently validated using real-world datasets to assess its adaptability and robustness in practical clinical scenarios. Performance comparisons against state-of-the-art (SOTA) models were conducted to further evaluate the model’s clinical efficacy.
Effects of different scenarios in CL
CL is essential for addressing domain shift—a phenomenon in which a model trained on one data distribution fails to effectively generalize to a different distribution. Especially, in the hypoglycemia event detection task, domain shift arises due to variations in time below range (TBR), defined as the proportion of time spent in the hypoglycemic range (<70 mg/dL). The low-hypoglycemia-risk (LHR) group has low TBR and insulin sensitivity, with infrequent hypoglycemia; the high-hypoglycemia-risk (HHR) group shows high insulin sensitivity and frequent, rapid glucose drops, associated with greater glycemic variability (ρ = 0.75; Supplementary Fig. 1). To examine the effect of training order, we tested two CL scenarios: scenario 1 (LHR → HHR), representing a progression from low to high risk, and scenario 2 (HHR → LHR), the reverse. Since HHR cases are more difficult due to sharp fluctuations, learning them later aligns with a difficulty-increasing curriculum that may improve CL stability and performance39,40,41. These scenarios were designed to reflect increasing versus decreasing clinical risk conditions, respectively. This comparison aims to assess how domain scheduling influences the model’s generalizability to high-risk cases, which are critical for ensuring safe and effective hypoglycemia detection.
Table 1 presents the zero-shot performance of the generalized DA-CMTL model for 30-min glucose level prediction and hypoglycemia event detection. For each scenario, the decision threshold for classification was determined based on the maximum Youden’s Index42. To assess generalization, the model was evaluated on three public datasets (ShanghaiT1DM, OhioT1DM, and DiaTrend), capturing diversity in glucose profiles, sensors, and demographics. Results show that training order significantly affects model performance, especially in hypoglycemia event detection sensitivity (Supplementary Table 1). In scenario 1 (LHR → HHR), sensitivity improved by 18.88% (79.04–97.92%) in ShanghaiT1DM, 16.37% (82.21–98.58%, p = 0.0178) in OhioT1DM, and 19.46% (67.96–87.42%, p = 0.0007) in DiaTrend, compared with scenario 2 (HHR → LHR). These improvements clearly suggest that starting from stable glycemic conditions (LHR) promotes better adaptation to high-variability domains (HHR). This result aligns with the concept of curriculum learning, where training starts from simpler patterns and gradually progresses to more complex ones. In our case, early exposure to low-variability patterns helps the model build reliable temporal representations, improving generalization and robustness in safety-critical tasks like hypoglycemia event detection. Given these results, scenario 1 was selected for further evaluations due to its superior sensitivity, predictive stability, and adaptability. These outcomes underscore the importance of structured CL, where progressive learning from stable to variable glucose conditions enhances model generalization and clinical relevance.
Performance evaluation
The predictive accuracy of the generalized DA-CMTL model was evaluated by Clarke error grid analysis (CEGA) for a 30-min PH across three public datasets (Fig. 2). CEGA categorizes predictions into five clinical risk zones A–E, with zone A indicating the highest accuracy and lowest clinical risk. In our evaluation, the model achieved high A-zone proportions: 98.74% (ShanghaiT1DM), 92.93% (OhioT1DM), and 92.57% (DiaTrend). The combined A + B zone proportions exceeded 98% (Supplementary Table 2), indicating clinically acceptable accuracy. Notably, DA-CMTL outperformed prior models on OhioT1DM, surpassing results from McShinsky et al. (97.50%)43 and Zhu et al. (98.82%)44, achieving 99.40%. A supplementary breakdown of the OhioT1DM dataset further showed that the majority of zone D predictions (0.48% of total) were associated with hypoglycemia, representing a small portion of hypoglycemic events and thus indicating minimal clinical risk. In addition, predictions in zones C–E, representing clinically unsafe errors, were negligible across datasets, further supporting model reliability.
This figure presents the CEGA for a 30-min PH, displayed from left to right for the ShanghaiT1DM, OhioT1DM, and DiaTrend datasets. Predictions for all subjects across each dataset are included in the figure.
Dataset-specific characteristics may have contributed to the observed variations in accuracy. For instance, ShanghaiT1DM showed the highest proportion of zone A predictions, which may be attributed to its higher time-below-range (TBR) and smoother CGM glucose readings (Table 4), both of which align well with the model’s recent high hypoglycemia risk (HHR) training phase. In contrast, the OhioT1DM and DiaTrend datasets exhibited greater glycemic variability and lower TBR, posing more challenging prediction conditions.
Effectiveness analysis of MTL and CL
Artificial intelligence-driven glucose prediction models must balance accuracy and complexity, requiring key components that enhance performance while avoiding overfitting and excessive computation. To assess the contributions of multi-task learning (MTL) and continual learning (CL), we conducted an ablation study on the generalized DA-CMTL model (Supplementary Table 3). MTL was evaluated by removing either the classification or prediction task. When classification was removed, hypoglycemia event was determined from predicted glucose levels, defined as three consecutive readings below 70 mg/dL. Removing classification (without classification) yielded the lowest RMSE: 10.24 mg/dL (ShanghaiT1DM), 13.21 mg/dL (OhioT1DM), and 15.82 mg/dL (DiaTrend), indicating improved glucose levels prediction accuracy. However, sensitivity dropped by 38.97%, 22.41%, and 33.86%, respectively, compared to the full model, highlighting a trade-off, where slight RMSE gains at the cost of markedly reduced hypoglycemia event detection. This trade-off underscores the need for explicitly modeling hypoglycemia event detection as a classification task, rather than relying solely on thresholding predicted glucose. This design choice improves robustness near the clinical threshold, such as the 70 mg/dL boundary, especially under high-risk conditions (Supplementary Fig. 2, Supplementary Table 4). Furthermore, Welch’s t-test confirmed that RMSE differences between the models without classification and the full DA-CMTL model (ours) were not statistically significant (p = 0.85, 0.78, 0.77), indicating that adding the classification task enables simultaneous glucose prediction and hypoglycemia event detection without compromising glucose prediction accuracy. Conversely, removing prediction maintained high sensitivity (97.92% in ShanghaiT1DM) but eliminated glucose prediction, limiting clinical applicability. Together, these results highlight that MTL is essential for balancing prediction accuracy and safety.
Next, the role of CL was analyzed by excluding multiple domain learning (MDL) and EWC on model stability and generalization. Without MDL, training on HHR improved sensitivity (99.68% in OhioT1DM)—but increased RMSE values (13.88 mg/dL in OhioT1DM), indicating reduced stability. Training solely on LHR reduced sensitivity (74.70% in OhioT1DM), suggesting the importance of exposure to diverse glucose. Removing EWC preserved high sensitivity (98.58% in OhioT1DM), but worsened RMSE (13.80 mg/dL in OhioT1DM), indicating overfitting to the most recently trained HHR domain. These outcomes confirm that MDL ensures generalization across glucose distributions, whereas EWC stabilizes sequential learning and prevents catastrophic forgetting. In summary, DA-CMTL consistently outperformed all ablation variants across multiple performance metrics, including RMSE, MAE, TG, sensitivity, and specificity (Fig. 3). It achieved the largest enclosed performance area per dataset—ShanghaiT1DM (1.25), OhioT1DM (1.24), and DiaTrend (1.61)—demonstrating superior overall performance. These results affirm that removing core components (MDL, EWC, classification) undermines generalization and robustness, supporting the necessity of the full DA-CMTL design for real-world AID applications.
This figure visualizes the ablation study results across three datasets, where each method is represented by a distinct color. The axes display normalized metrics, including -RMSE, -MAE, TG, sensitivity, and specificity. The area, calculated from Cartesian coordinates transformed from polar coordinates, represents overall performance; a larger area indicates stronger performance across the evaluated metrics.
Comparison with recent SOTA models for glucose level prediction
To benchmark DA-CMTL, we compared its performance with recent SOTA glucose prediction models, covering both generalized and personalized approaches. These models span a variety of architectures and learning strategies, including transformers, GANs, and recurrent networks (Supplementary Note: Description of baseline models used for benchmarking).
As shown in Table 2, a comparison between the generalized DA-CMTL-based predictor and recent SOTA models, including TFT19 and GARNN20. Because results for the DiaTrend dataset were unavailable for competing models, the comparison primarily focuses on the ShanghaiT1DM and OhioT1DM datasets. At a 30-min PH, DA-CMTL achieved the lowest RMSE across both datasets—10.58 mg/dL for ShanghaiT1DM and 13.38 mg/dL for OhioT1DM—outperforming TFT, GARNN, and Meta-GPformer. As a domain-agnostic model, DA-CMTL demonstrated superior adaptability across patient populations exhibiting diverse glucose dynamics. However, at a 60-min PH, performance degradation was observed. This reduction in accuracy is likely due to the advantage that competing models gain from utilizing longer historical input sequences and additional physiological features. Importantly, this performance drop does not critically impair the model’s applicability in real-world automated insulin delivery (AID) systems. In clinical practice, 30-min prediction results are most commonly used to inform preventative interventions, such as carbohydrate intake, to mitigate impending hypoglycemic events45,46,47. Given that DA-CMTL maintains strong accuracy at this horizon, it remains highly effective for proactive glucose management within AID frameworks.
To complement the generalization analysis, the personalized performance of DA-CMTL was evaluated to assess its capacity for individual-level adaptation under limited data conditions. Table 3 summarizes the comparison between personalized DA-CMTL and recent personalized SOTA models, primarily on the OhioT1DM dataset, a widely adopted benchmark in previous studies. In clinical applications, accurate prediction with minimal patient-specific data is particularly critical during early system deployment or initial monitoring phases. To reflect this constraint, the evaluation focused on models’ ability to achieve effective personalization from limited data. At a 30-min PH, DA-CMTL achieved an RMSE of 13.38 mg/dL and an MAE of 9.42 mg/dL, comparable to competing models but requiring only 2 days of fine-tuning—a key advantage, as others often rely on larger labeled datasets. Although certain models showed marginal improvements at a 60-min horizon, a trend consistent with our generalized model comparison, the DA-CMTL-based predictor retained a competitive edge by enabling efficient and scalable personalization from minimal data. The inclusion of MTL-based model48 trained on real clinical datasets further underscores DA-CMTL’s strengths: despite limited data, our model outperformed in terms of both scalability and efficiency. These findings validate our goal of building a model capable of rapid adaptation with low data burden, supporting its practical use in real-world AID systems.
In summary, DA-CMTL demonstrated superior accuracy in short-term glucose prediction at a 30-min PH, outperforming domain-specific SOTA models and confirming its adaptability as a fully generalized model. With only 2 days of fine-tuning, it also surpassed existing personalized models, demonstrating efficient personalization from a robust generalized framework. At 60-min PH, performance declined due to challenges in long-term foreca(Asting, including the increasing influence of external factors such as delayed meal and insulin effects, and the GRU’s limited capacity for long-range dependencies (Fig. 4). Nevertheless, the model remained stable at the clinically relevant 30-min horizon, supporting timely decision-making in AID systems.
Predicted glucose at 30- and 60-min horizons. The shaded area shows a 99% confidence interval from the Monte Carlo Dropout across sampled predictions (Supplementary Note: Uncertainty prediction). Purple indicates 30-min predictions, and blue represents 60-min predictions. Circles mark timepoints classified as hypoglycemia events.
Impact of glycemic traits on model generalization
To examine the dynamics of continual learning, step-wise performance changes were evaluated by progressively incorporating domain-specific data (Supplementary Fig. 3a). While RMSE generally decreased and sensitivity improved from Step 1 (LHR-only training) to Step 2 (HHR added), a performance drop, particularly an RMSE increase, was observed in some cases during Step 3 (domain-specific adaptation). To determine whether this was attributable to patient characteristics rather than model limitations, linear regression was performed on RMSE changes (Step 2 → 3) using glycemic features as predictors (Supplementary Fig. 3b). Higher HbA1c and CV were significantly associated with increased RMSE, suggesting that performance degradation occurred primarily in clinically complex cases. This pattern is consistent with Fig. 5a, where higher CV was positively correlated with RMSE, highlighting the need for personalized strategies in patients with high glycemic variability.
a Heatmap of standardized OLS regression coefficients between glycemic traits and performance metrics. b Scatter plots showing the relationship between CV and RMSE (left), and between HbA1c and hypoglycemia event sensitivity (right), with 95% confidence intervals. c Violin plots illustrating the distribution of glycemic traits and model performance across five clusters. d Normalized Radar chart summarizing average cluster-wise glycemic features and DA-CMTL performance.
We next examined dataset-level generalizability by assessing the distributional shifts between training and external datasets. Kullback–Leibler (KL) divergence was computed for CGM distributions, identified as most important through SHAP analysis (Supplementary Fig. 4), to quantify the mismatch between each target dataset and the training domains (HHR, LHR, and HHR + LHR). As summarized in Supplementary Table 5, DiaTrend exhibited the highest divergence (0.24 with HHR + LHR; 0.39 with HHR), suggesting a greater distributional mismatch that may partially account for its relatively lower predictive performance.
Building upon these findings, a patient-level analysis was conducted to further examine heterogeneity in model performance (Fig. 5). Five common clinical features (HbA1c, CV, TBR, TAR, and gender) were used as predictors in a standardized ordinary least squares (OLS) regression (Fig. 5a). The results indicated a significant negative association between HbA1c and Sensitivity, and a positive association between CV and RMSE (p < 0.01) (Fig. 5b). Additionally, k-means clustering based on these features identified five distinct patient subgroups (Fig. 5c). Notably, Cluster 4, defined by high HbA1c and low CV, exhibited low RMSE and MAE but also the lowest sensitivity (Fig. 5d). This result likely reflects a low incidence of hypoglycemic events in this group, which limits the model’s opportunity to detect such events and consequently leads to reduced sensitivity.
Preliminary evaluation of the AID system
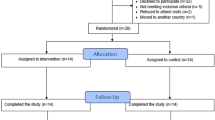
To assess the feasibility of real-time integration, the DA-CMTL model was implemented as a predictive safety layer (SL) within a closed-loop AID system (Supplementary Fig. 5c, Supplementary Note: Application in AID systems). This included an in vivo evaluation using a T2DM-induced rat model to assess cross-pathophysiology generalization, based on the model’s domain-agnostic design using only CGM and insulin data, and to complement prior validations on multiple external T1D human datasets. In a 31-day in silico simulation involving virtual patients, the generalized DA-CMTL–based SL (GSL) significantly reduced TBR from 7.15% to 0.50% (p = 0.0013) and improved time in range (TIR) from 55.39% to 59.89% (p = 0.0025), outperforming rule-based baselines (Supplementary Fig. 5d, Supplementary Table 6). In the subsequent in vivo evaluation, the system was configured with a Dana-i insulin pump (Sooil Development), a Dexcom G7 sensor, and a control algorithm deployed via a dedicated mobile application on an Android smartphone, emulating a clinically relevant hybrid AID environment. The in vivo experiment, conducted in T2DM-induced rats49, showed an unexpected increase in TBR under GSL, which may reflect interspecies physiological differences50 and the inability to administer carbohydrate corrections (Supplementary Fig. 5a). Although the personalized SL (PSL) achieved superior predictive performance, its control efficacy did not show statistically significant improvement over GSL, likely due to the limited sample size and single-day evaluation per condition (Supplementary Fig. 5e). These findings suggest that the proposed model holds promise for real-time AID applications, warranting further validation in larger-scale clinical studies.
Discussion
In this study, we proposed the DA-CMTL model, a unified modeling framework that simultaneously performs glucose level prediction and hypoglycemia event detection by integrating three key components: multi-task learning (MTL), continual learning (CL), and simulation-to-real (Sim2Real) transfer. Rather than operating independently, these components are jointly optimized to form a cohesive system in which each part reinforces the others. Specifically, MTL enables the model to learn task-specific features while leveraging shared representations; CL mitigates domain shift between simulated and real-world distributions while avoiding catastrophic forgetting in sequential learning, and Sim2Real transfer allows the model to benefit from large-scale, diverse virtual patient data without requiring extensive labeled clinical data. This synergistic design contributes to improved generalization, robustness, and practical adaptability of the DA-CMTL glucose prediction model for real-world applications.
To assess the generalizability of the proposed DA-CMTL glucose prediction model, we compared its performance with recent SOTA methods across three public datasets under a 30-min PH. Without requiring any domain-specific tuning, DA-CMTL consistently achieved high predictive accuracy, with RMSE values of 10.58 mg/dL on ShanghaiT1DM, 13.38 mg/dL on OhioT1DM, and 15.74 mg/dL on DiaTrend. These results demonstrate the model’s robustness in handling heterogeneous glucose dynamics, sensor types, and population characteristics. While performance naturally declined at a 60-min PH due to the inherent challenge of long-term forecasting, the model maintained clinically acceptable accuracy, supporting its applicability for anticipatory interventions in AID systems. In addition, we evaluated the fine-tuned DA-CMTL for personalization using only 2 days of data. Despite the limited tuning duration, the model confirmed its capacity for rapid adaptation with minimal data. These findings indicate that DA-CMTL offers an efficient and scalable solution for both generalized and personalized glucose forecasting in real-world scenarios.
Next, an ablation analysis was conducted to assess the contribution of key architectural components. Removing the classification head resulted in slightly lower RMSE (13.21 mg/dL in OhioT1DM) but led to a notable drop in hypoglycemia event sensitivity across datasets. This highlights the importance of explicitly modeling event detection as a classification task, as opposed to relying exclusively on threshold-based interpretation of predicted glucose values. Conversely, exclusion of continual learning elements, such as multi-domain learning or elastic weight consolidation (EWC), degraded performance stability and generalization, indicating their critical role in maintaining knowledge across shifting domains.
The DA-CMTL model offers practical advantages in real-world settings by balancing accuracy and efficiency. Despite using a lightweight GRU-based architecture, the model achieved reliable 30-min prediction accuracy across datasets while maintaining high sensitivity in hypoglycemia event detection. These outcomes, along with fast inference speed and minimal reliance on extensive input features, make the model suitable for integration into resource-constrained platforms such as mobile-based AID systems. To further enhance deployment efficiency, future implementations could consider conditionally activating the classification module only when predicted glucose levels approach hypoglycemic thresholds. Such adaptive scheduling could reduce computational overhead while preserving critical event detection functionality in high-risk scenarios. Additionally, the modular structure of DA-CMTL enables flexibility for future integration of additional physiological signals when available.
While DA-CMTL shows promising results, it still has several limitations. First, prediction accuracy declines at the 60-min horizon, likely due to the increasing influence of external variables and the limited temporal capacity of GRUs. To address this, future work may explore replacing or augmenting the GRU backbone with Transformer-based architectures to better capture long-range temporal dependencies and improve long-term forecasting performance. Second, to ensure high usability and facilitate generalization across diverse real-world AID settings, the model was intentionally developed using only CGM and insulin data. While this input restriction supports practical deployment, it precludes the integration of contextual factors such as meals or physical activity, which are known to affect glycemic dynamics. In response, incorporating physiological variables such as meals or activity through attention-based modules may enhance accuracy without compromising real-time usability. Third, the in vivo validation was limited to a T2D-induced rat model, which differs in pathophysiology from T1D, the primary target population. Future studies should include T1D animal models and diverse human cohorts to fully assess clinical adaptability. Lastly, although uncertainty estimates were not directly incorporated into the control strategies in this study, providing uncertainty information to user-facing AID systems may enhance reliable decision-making by allowing users to assess the confidence of predictions in real time. In addition, while the current multi-task design was adopted to explicitly optimize both glucose prediction and hypoglycemia event detection, future work may consider a single-task design, provided that predictive accuracy is sufficient to reliably fulfill both functions.
In summary, the findings in this study demonstrate that the DA-CMTL framework enables robust and scalable performance across heterogeneous datasets by jointly modeling glucose level forecasting and hypoglycemia event detection within a unified framework. By leveraging simulation-based training and continual learning, the model effectively reduces reliance on large-scale labeled clinical data, supporting efficient adaptation across domains toward full generalization. Despite its lightweight design, the DA-CMTL-based predictor consistently achieved clinically acceptable accuracy at the 30-min horizon, making it suitable for real-time use in adaptive insulin delivery systems. These attributes suggest that the DA-CMTL framework offers a practical backbone for future AI-driven glycemic control technologies, particularly in mobile or resource-constrained environments.
Methods
This section introduces CL and MTL, two key concepts for implementing the generalized DA-CMTL model. It presents the learning strategy using a loss function and details the model’s application through transfer learning, where the model is trained in a simulated environment and subsequently applied to real-world data.
Training datasets from simulator
For this study, training datasets were developed using a modified version of the Simglucose simulator51, incorporating the S2013 version of the FDA-approved UVA/Padova virtual patient (VP) population consisting of 10 adult T1DM subjects52. The simulator was internally adapted by our research group to integrate the S2013 profiles and implement parameter perturbation for insulin-related physiology, as previously described35,53. Two simulation scenarios were created, each spanning 61 days, with parameters such as total daily insulin (TDI), CR, and CF randomly varied. To generate the HHR condition, TDI, CR, and CF were randomized within ±20% of their optimal values under basal-bolus insulin settings. Conversely, for the LHR condition, the optimal values were retained. Each simulation introduced three meals per day with basal-bolus insulin dosing. Carbohydrate intake was set to 70, 110, and 90 g for breakfast, lunch, and dinner, respectively, with variability modeled as a uniform distribution of ±30 g. Mealtimes were set at 7:00 AM, 1:00 PM, and 9:00 PM, with a random variation of ±30 min. After each meal, there was a 50% chance of snack consumption. If a snack occurred, its timing was set between 60 and 90 min post-meal, and its carbohydrate content was uniformly distributed between 5.0 and 25.0 g.
Real-world validation datasets
Three widely recognized, publicly available datasets were adopted for evaluation: DiaTrend54, OhioT1DM55, and ShanghaiT1DM56. To ensure consistency in model inputs and relevance to AID applications, we selected datasets that included CGM glucose readings alongside both basal and bolus insulin records. All datasets were derived from individuals with T1DM, aligning with the clinical scope of this study. Key dataset characteristics are summarized below:
-
(1)
DiaTrend: This recently released dataset includes CGM data from 54 patients aged 19–74 years (17 men, 37 women), with an average of 510 days (range: 31–780 days) of data per subject. For this study, we selected 17 subjects whose records included both basal and bolus insulin data, which are critical inputs for modeling insulin dynamics in AID systems. The selected files were S29–S31, S36–S39, S42, S45–S47, and S49–S54. CGM data were recorded at 5-min intervals using devices from Dexcom, Abbott, and Medtronic. The dataset is accessible via Synapse with required registration.
-
(2)
OhioT1DM: This dataset was introduced as part of the Blood Glucose Level Prediction (BGLP) Challenge in 2018 and 2020. It includes 8 weeks of data from 12 T1DM patients (seven men, five women) aged 20–80 years. CGM readings were recorded every 5 minutes, along with bolus and basal insulin doses from Medtronic 530 or 630 G insulin pumps paired with Medtronic Enlite CGM sensors. Access requires academic credentials.
-
(3)
ShanghaiT1DM: This dataset was collected from 12 Chinese T1DM patients (seven men, five women) aged 37–73 years. CGM data were collected over 3–14 days under real-life conditions using the FreeStyle Libre H device. Readings were recorded every 15 min and are publicly available for research use.
Data preprocessing
A consistent data preprocessing pipeline was applied to both simulated and real-world datasets, in accordance with the methodology outlined in a previous study57. For CGM data, cubic interpolation was used to fill missing values if gaps were shorter than 20 min, while days with longer gaps were excluded to prevent distorting rapidly changing CGM signals. Additionally, for the ShanghaiT1DM dataset, which originally had 15-min sampling intervals, cubic interpolation was applied to resample the data at 5-minute intervals to ensure consistency across datasets during model training. Interpolated values were excluded from the evaluation phase to ensure fair performance assessment. Missing basal and bolus insulin values were imputed with zeros. CGM values were clipped within the range of 40–400 mg/dL to handle outliers. After cleaning, the CGM data were merged with basal and bolus insulin data for overlapping periods. To estimate IOB, which represents the amount of active insulin remaining in the body, a two-compartment pharmacokinetic model, originally introduced by Wilinska et al.58 and adopted in previous studies59,60,61, was utilized. In this model, \({C}_{1}\) and \({C}_{2}\) represent the two compartments, \({\rm{u}}\left(t\right)\) denotes the insulin injection, and \({K}_{{\rm{D}}{\rm{I}}{\rm{A}}},\) set to 0.025, indicates the constant related to the duration of insulin action. The related formulas are detailed in Eqs. (1)–(3). Using the processed variables as input data, we specified input variables with p CGM and IOB sequences to derive \({\hat{{\rm{y}}}}_{{\rm{r}}{\rm{e}}{\rm{g}}}\) and \({\hat{{\rm{y}}}}_{{\rm{c}}{\rm{l}}{\rm{f}}}\) as shown in Fig. 6, with \({X}_{t}=({\rm{C}}{\rm{G}}{\rm{M}}(t),{\rm{I}}{\rm{O}}{\rm{B}}(t))\). These input sequences were constructed using a sliding window approach with a 60-min window length and a 5-min step size, resulting in overlapping time-series segments that reflect realistic clinical prediction settings.
DA-CMTL consists of two main modules: shared layers and task-specific layers. It uses a multi-head structure to implement task-specific layers.
A validation strategy was employed within each dataset. Subjects were randomly partitioned into training and validation subsets at an 8:2 ratio, ensuring that all data from a given individual were included in only one subset. This subject-level split was adopted to prevent data leakage and to enable a reliable assessment of the model’s performance. To ensure consistency, min–max normalization was applied to each feature based on the training data statistics. A detailed overview of the processed datasets is provided in Table 4, and their distributions are illustrated in Supplementary Fig. 1.
Model development
Blood glucose prediction and hypoglycemia event classification were conducted using a stacked architecture consisting of GRU layers followed by MC-dropout layers. The model was implemented using Python 3.10.9, PyTorch 1.13.1, CUDA 12.2, and cuDNN 8.9.2, and training was accelerated using an NVIDIA GeForce RTX 3090 GPU. The Adaptive Moment Estimation (Adam) optimizer was used to minimize the loss function, and a Cosine Annealing learning rate scheduler62 was applied. Early stopping was utilized to prevent overfitting. During inference, 20 MC-dropout samples were used, based on the elbow point identified during experimentation. Specific hyperparameters used for general training, optimized via Bayesian optimization on the simulator dataset (Supplementary Table 7). For CL, the model was sequentially trained on two simulated datasets. To minimize forgetting of hypoglycemia-related features, Simulated-1 (the dataset with more frequent hypoglycemia events) was placed later in the training sequence. During personalization, the learning rate and batch size were reduced by a factor of 0.1 to enhance training stability.
Evaluation metrics
The model’s performance was evaluated using two categories of metrics. First, glucose prediction accuracy was assessed using RMSE and MAE, which are common metrics in glucose forecasting63. RMSE reflects the average magnitude of prediction errors, emphasizing larger errors, whereas MAE provides a more robust measure, less sensitive to outliers. Second, hypoglycemia event classification was evaluated using sensitivity, specificity, and TG, which are standard metrics for hypoglycemia event detection64. Sensitivity measures the model’s ability to correctly identify hypoglycemic events, and Specificity quantifies its accuracy in detecting non-hypoglycemic cases. TG represents the model’s ability to detect glucose changes ahead of time, calculated by determining the delay through cross-correlation between predicted and actual time series, and subtracting it from the PH. The evaluation metrics are formally defined as follows:
To further evaluate clinical relevance, CEGA was performed65. CEGA categorizes glucose predictions into five zones (A–E) based on their potential clinical impact. Predictions in zone A are considered clinically accurate and safe, whereas those in zone B are acceptable but less precise. Zone C may lead to unnecessary corrective actions; zones D and E indicate dangerous or erroneous predictions. A high proportion of data points in zones A and B indicates better clinical reliability.
CL on the DA-CMTL
In this study, we implemented a generalized, domain-agnostic model that learns sequentially from multiple datasets using a continual learning strategy66. While conventional deep learning (DL) assumes a stationary data distribution, real-world applications frequently involve domain shifts, leading to a phenomenon known as catastrophic forgetting, where newly acquired knowledge interferes with previously learned information, ultimately degrading model performance (Supplementary Fig. 6a).
To address this challenge, we considered several CL paradigms. Rehearsal-based method67 retains previous data to mitigate forgetting, but poses privacy and storage concerns. Architectural approaches68 introduce additional parameters or modules to preserve prior knowledge, but increase computational overhead. As a lightweight and data-independent alternative, we adopted EWC—a regularization-based method that penalizes updates to parameters critical for prior tasks. Specifically, EWC estimates parameter importance using the Fisher Information Matrix and incorporates a corresponding penalty term into the loss function, thereby constraining significant deviations during training38. As illustrated in Supplementary Fig. 6b and Eq. (10), this mechanism enables the model to maintain previously acquired knowledge while adapting to new data distributions, supporting both stability and plasticity throughout the learning process.
MTL on the DA-CMTL
The DA-CMTL model employs MTL to simultaneously predict glucose levels and detect hypoglycemic events, aiming to support efficient and adaptive closed-loop insulin delivery. MTL is a learning paradigm that jointly optimizes multiple related tasks, enabling the model to leverage shared information and enhance generalization performance across tasks69. A primary benefit of MTL is its ability to improve the performance of individual tasks through shared learning; it has been widely utilized in areas such as natural language processing and computer vision70,71. In this study, we adopted the hard parameter sharing strategy, wherein hidden layers are shared across tasks while output layers remain task-specific.
Module 1 consists of a stacked gated recurrent unit (GRU) architecture that encodes temporal dependencies in CGM and insulin-on-board (IOB) data. GRUs are lightweight recurrent networks designed to capture long-term patterns with reduced computational overhead. The GRU structure used is shown in Supplementary Fig. 6c72. Extracted features are subsequently passed to Module 2 for task-specific processing. Given the distinct objectives of continuous glucose forecasting and binary event detection, each output branch includes a dedicated fully connected (FC) layer. The regression branch outputs a continuous glucose estimate \({\hat{y}}_{{\rm{r}}{\rm{e}}{\rm{g}}}\), while the classification branch applies a sigmoid activation func\({\rm{tion}}\) to produce a binary hypoglycemia classification \({\hat{y}}_{{\rm{c}}{\rm{l}}{\rm{f}}}\). This architecture enables efficient reuse of temporal features while tailoring outputs to the unique requirements of each task.
Loss function
To integrate CL and MTL into model training, a composite loss function was formulated, combining task-specific objectives with a regularization term from elastic weight consolidation (EWC). The loss for CL (Eq. (10)) introduces a penalty on deviations from previously learned parameters, where F denotes the Fisher Information Matrix and λ controls the regularization strength. Each model parameter \({\theta }_{i}\) is penalized based on its divergence from the previously learned value \({\theta }_{p,i}^{\ast }\), scaled by its estimated importance \({F}_{i}.\) This mechanism mitigates catastrophic forgetting by preserving essential parameters across sequential tasks. The MTL objective (Eq. (11)) jointly minimizes the regression loss \({L}_{{\rm{R}}{\rm{E}}{\rm{G}}}(\theta )\), defined as mean square error, and classification loss \({L}_{{\rm{C}}{\rm{L}}{\rm{F}}}(\theta )\), defined as binary cross-entropy. A task-scaling factor \({\lambda }_{{\rm{C}}{\rm{L}}{\rm{F}}}\) was heuristically selected to balance the contributions of both tasks. Notably, the effective influence of the classification term varies with the prevalence of TBR in the training data, enabling implicit task reweighting without manual tuning. Unlike prior study48 that assigned large fixed weights to the classification task \(({\lambda }_{{\rm{C}}{\rm{L}}{\rm{F}}}\) = 100), our approach adopts a moderate and adaptive task balancing strategy (\({\lambda }_{{\rm{C}}{\rm{L}}{\rm{F}}}\,\)≤ 10), which prevents overfitting to rare events while preserving prediction accuracy. The final loss function (Eq. (12)) combines both the MTL objective and CL regularization, facilitating stable knowledge retention while supporting adaptive learning across heterogeneous data distributions.
Application: Sim2Real transfer
This section describes how the generalized model, developed using CL and MTL, was applied in real-world settings. Transfer learning is an efficient strategy for reusing pre-trained models on new tasks or datasets73. As illustrated in Supplementary Fig. 6d, this study categorized transfer learning into three approaches: Frozen, Head-Tuned, and Fully-Tuned. In the Frozen method, the pre-trained generalized model is used without modification. All parameter values of the original layers are retained and applied directly to the target dataset. In the Head-Tuned method, only the weights of FC layers in the task-specific heads—those responsible for multi-task outputs—are retrained based on individual real-world subjects. In contrast, the Fully-Tuned method retrains all layers of the model from the generalized initialization. Previous research has shown that selectively retraining parts of a model is often more efficient, more robust, and yields better performance compared with full model retraining22,74,75. For this reason, the present study applied the Head-Tuned approach to personalize the generalized model for specific individuals. Inspired by continual learning principles, personalization was performed using a subset of individual data to efficiently adapt the model while minimizing overfitting.
Data availability
Datasets used in this study included Simglucose, DiaTrend (Prioleau, T., Bartolome, A., Comi, R. et al. DiaTrend: a dataset from advanced diabetes technology to enable the development of novel analytic solutions. Sci. Data 10, 556 (2023). https:/doi.org/10.1038/s41597-023-02469-5), OhioT1DM (Marling, C., & Razvan, B. The OhioT1DM dataset for blood glucose level prediction: update 2020. CEUR Workshop Proceedings, Vol. 2675 (2020)), and ShanghaiT1DM (Jee, J., Fong, C., Pichotta, K. et al. Automated real-world data integration improves cancer outcome prediction. Nature 636, 728–736 (2024). https:/doi.org/10.1038/s41586-024-08167-5), all of which are publicly accessible. Access to these datasets may require permission from the respective data holders. Additional datasets utilized in this study are not provided within this publication. Access to such data may be granted upon reasonable request to the corresponding author.
Code availability
The source code used for data preprocessing, analysis, and model development is publicly available via GitHub: https://github.com/Hwang-Minjoo/DA-CMTL/. Implementation for model training and evaluation was conducted using Python 3.10.9, whereas model deployment within the real-time control application utilized Python 3.9.19.
References
Limbert, C., Kowalski, A. J. & Danne, T. P. Automated insulin delivery: a milestone on the road to insulin independence in type 1 diabetes. Diabetes Care 47, 918–920 (2024).
Perkins, B. A., Sherr, J. L. & Mathieu, C. Type 1 diabetes glycemic management: insulin therapy, glucose monitoring, and automation. Science 373, 522–527 (2021).
Latres, E., Finan, D. A., Greenstein, J. L., Kowalski, A. & Kieffer, T. J. Navigating two roads to glucose normalization in diabetes: automated insulin delivery devices and cell therapy. Cell Metab 29, 545–563 (2019).
Phillip, M. et al. Consensus recommendations for the use of automated insulin delivery technologies in clinical practice. Endocr. Rev. 44, 254–280 (2023).
Sherr, J. L. et al. Automated insulin delivery: benefits, challenges, and recommendations. A Consensus Report of the Joint Diabetes Technology Working Group of the European Association for the Study of Diabetes and the American Diabetes Association. Diabetes Care 45, 3058–3074 (2022).
Lee, T. T. et al. Automated insulin delivery in women with pregnancy complicated by type 1 diabetes. N. Engl. J. Med. 389, 1566–1578 (2023).
Crabtree, T. S. et al. Hybrid closed-loop therapy in adults with type 1 diabetes and above-target HbA1c: a real-world observational study. Diabetes Care 46, 1831–1838 (2023).
Brown, S. A. et al. Multicenter trial of a tubeless, on-body automated insulin delivery system with customizable glycemic targets in pediatric and adult participants with type 1 diabetes. Diabetes Care 44, 1630–1640 (2021).
Mosquera-Lopez, C. et al. Enabling fully automated insulin delivery through meal detection and size estimation using artificial intelligence. NPJ Digit. Med. 6, 39 (2023).
Walsh, J., Roberts, R., Bailey, T. S. & Heinemann, L. Insulin titration guidelines for patients with type 1 diabetes: it is about time! J. Diabetes Sci. Technol. 17, 1066–1076 (2023).
Shalitin, S. & Phillip, M. Hypoglycemia in type 1 diabetes: a still unresolved problem in the era of insulin analogs and pump therapy. Diabetes Care 31, S121–S124 (2008).
Zhu, T., Li, K., Herrero, P. & Georgiou, P. Deep learning for diabetes: a systematic review. IEEE J. Biomed. Health Inform. 25, 2744–2757 (2020).
Pérez-Gandía, C. et al. Artificial neural network algorithm for online glucose prediction from continuous glucose monitoring. Diabetes Technol. Ther. 12, 81–88 (2010).
Kim, D. Y. et al. Developing an individual glucose prediction model using recurrent neural network. Sensors 20, 6460 (2020).
Li, K., Daniels, J., Liu, C., Herrero, P. & Georgiou, P. Convolutional recurrent neural networks for glucose prediction. IEEE J. Biomed. Health Inform. 24, 603–613 (2019).
Zhu, T., Li, K., Chen, J., Herrero, P. & Georgiou, P. Dilated recurrent neural networks for glucose forecasting in type 1 diabetes. J. Healthc. Inform. Res. 4, 308–324 (2020).
Martinsson, J., Schliep, A., Eliasson, B. & Mogren, O. Blood glucose prediction with variance estimation using recurrent neural networks. J. Healthc. Inform. Res. 4, 1–18 (2020).
Alshehri, O. S., Alshehri, O. M. & Samma, H. Blood glucose prediction using RNN, LSTM, and GRU: a comparative study. In Proc. IEEE Int. Conf. Advanced Systems and Emergent Technologies (eds. Amor, A. B. et al.) 1–5 (2024).
Zhu, T. et al. Population-specific glucose prediction in diabetes care with transformer-based deep learning on the edge. IEEE Trans. Biomed. Circuits Syst. 18, 236–246 (2024).
Piao, C. et al. GARNN: an interpretable graph attentive recurrent neural network for predicting blood glucose levels via multivariate time series. Neural Networks 185, 107229 (2025).
Montaser, E. et al. Seasonal local models for glucose prediction in type 1 diabetes. IEEE J. Biomed. Health Inform. 24, 2064–2072 (2019).
Seo, W., Park, S.-W., Kim, N., Jin, S.-M. & Park, S.-M. A personalized blood glucose level prediction model with a fine-tuning strategy: a proof-of-concept study. Comput. Methods Prog. Biomed. 211, 106424 (2021).
Shuvo, M. M. H. & Islam, S. K. Deep multitask learning by stacked long short-term memory for predicting personalized blood glucose concentration. IEEE J. Biomed. Health Inform. 27, 1612–1623 (2023).
Deng, Y. et al. Deep transfer learning and data augmentation improve glucose levels prediction in type 2 diabetes patients. NPJ Digit. Med. 4, 109 (2021).
Tsichlaki, S., Koumakis, L. & Tsiknakis, M. Type 1 diabetes hypoglycemia prediction algorithms: systematic review. JMIR Diabetes 7, e34699 (2022).
Seo, W., Lee, Y.-B., Lee, S., Jin, S.-M. & Park, S.-M. A machine-learning approach to predict postprandial hypoglycemia. BMC Med. Inform. Decis. Mak. 19, 1–13 (2019).
Georga, E. I., Protopappas, V. C., Ardigo, D., Polyzos, D. & Fotiadis, D. I. A glucose model based on support vector regression for the prediction of hypoglycemic events under free-living conditions. Diabetes Technol. Ther. 15, 634–643 (2013).
Jensen, M. H. et al. Real-time hypoglycemia detection from continuous glucose monitoring data of subjects with type 1 diabetes. Diabetes Technol. Ther. 15, 538–543 (2013).
Zhu, T., Li, K., Herrero, P. & Georgiou, P. Personalized blood glucose prediction for type 1 diabetes using evidential deep learning and meta-learning. IEEE Trans. Biomed. Eng. 70, 193–204 (2022).
Shao, J. et al. Generalization of a deep learning model for continuous glucose monitoring–based hypoglycemia prediction: algorithm development and validation study. JMIR Med. Inform. 12, e56909 (2024).
Lee, S.-M., Kim, D.-Y. & Woo, J. Glucose transformer: forecasting glucose level and events of hyperglycemia and hypoglycemia. IEEE J. Biomed. Health Inform. 27, 1600–1611 (2023).
Lee, K. W. Costs of diabetes mellitus in Korea. Diabetes Metab. J. 35, 567 (2011).
Ahmed, M. I. et al. A systematic review of the barriers to the implementation of artificial intelligence in healthcare. Cureus 15, e46454 (2023).
Murdoch, B. Privacy and artificial intelligence: challenges for protecting health information in a new era. BMC Med. Ethics 22, 1–5 (2021).
Rachim, V. P., Yoo, J., Lee, J., Lee, Y. & Park, S.-M. Generalized reinforcement learning control algorithm for fully automated insulin delivery system. Expert Syst. Appl. 274, 126909 (2025).
Zhang, X. et al. Self-supervised tumor segmentation with sim2real adaptation. IEEE J. Biomed. Health Inform. 27, 4373–4384 (2023).
Shademan, A. et al. Supervised autonomous robotic soft tissue surgery. Sci. Transl. Med. 8, 337ra364 (2016).
Kirkpatrick, J. et al. Overcoming catastrophic forgetting in neural networks. Proc. Natl Acad. Sci. USA 114, 3521–3526 (2017).
Yifan, C., Yulu, C., Yadan, Z. & Wenbo, L. Continual learning in an easy-to-hard manner. Appl. Intell. 53, 20626–20646 (2023).
Bengio, Y., Louradour, J., Collobert, R. & Weston, J. Curriculum learning. in Proc. of the 26th Annual International Conference on Machine Learning. 41–48 (ICML, 2009).
Wang, Y., Gao, J., Wang, W., Yang, X. & Du, J. Curriculum learning-based domain generalization for cross-domain fault diagnosis with category shift. Mech. Syst. Signal Process. 212, 111295 (2024).
Smits, N. A note on Youden’s J and its cost ratio. BMC Med. Res. Methodol. 10, 89 (2010).
McShinsky, R. & Marshall, B. Comparison of Forecasting Algorithms for Type 1 Diabetic Glucose Prediction on 30 and 60-Minute Prediction Horizons. in Knowledge Discovery in Healthcare Data@ European Conference on Artificial Intelligence. 12–18 (CEUR-WS, 2020).
Zhu, T., Yao, X., Li, K., Herrero, P. & Georgiou, P. Blood Glucose Prediction for Type 1 Diabetes Using Generative Adversarial Networks. In CEUR Workshop Proc. 2675, 90–94 (CEUR-WS, 2020).
Shroff, P., Arefeen, A. & Ghasemzadeh, H. GlucoseAssist: Personalized blood glucose level predictions and early dysglycemia detection. In Proc. 2023 IEEE 19th International Conference on Body Sensor Networks (BSN). 1–4 (IEEE, 2023).
Pappada, S. M. et al. Neural network-based real-time prediction of glucose in patients with insulin-dependent diabetes. Diabetes Technol. Ther. 13, 135–141 (2011).
Buckingham, B. et al. Preventing hypoglycemia using predictive alarm algorithms and insulin pump suspension. Diabetes Technol. Ther. 11, 93–97 (2009).
Yang, M., Dave, D., Erraguntla, M., Cote, G. L. & Gutierrez-Osuna, R. Joint Hypoglycemia Prediction and Glucose Forecasting via Deep Multi-Task Learning. IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP). 1136–1140 (IEEE, 2022).
Srinivasan, K., Viswanad, B., Asrat, L., Kaul, C. & Ramarao, P. Combination of high-fat diet-fed and low-dose streptozotocin-treated rat: a model for type 2 diabetes and pharmacological screening. Pharmacol. Res. 52, 313–320 (2005).
De Vos, A. et al. Human and rat beta cells differ in glucose transporter but not in glucokinase gene expression. J. Clin. Investig. 96, 2489–2495 (1995).
J Xie. Simglucose v0.2.1 https://github.com/jxx123/simglucose (2018).
Man, C. D. et al. The UVA/PADOVA type 1 diabetes simulator: new features. J. Diabetes Sci. Technol. 8, 26–34 (2014).
Lee, S., Kim, J., Park, S. W., Jin, S.-M. & Park, S.-M. Toward a fully automated artificial pancreas system using a bioinspired reinforcement learning design: in silico validation. IEEE J. Biomed. Health Inform. 25, 536–546 (2020).
Prioleau, T., Bartolome, A., Comi, R. & Stanger, C. DiaTrend: a dataset from advanced diabetes technology to enable development of novel analytic solutions. Sci. Data 10, 556 (2023).
Marling, C. & Bunescu, R. The OhioT1DM Dataset for Blood Glucose Level Prediction: Update 2020. CEUR Workshop Proc 2675, 71 (2020).
Zhao, Q. et al. Chinese diabetes datasets for data-driven machine learning. Sci. Data 10, 35 (2023).
Jacobs, P. G. et al. Artificial intelligence and machine learning for improving glycemic control in diabetes: best practices, pitfalls and opportunities. IEEE Rev. Biomed. Eng. 17, 19–41 (2023).
Wilinska, M. E. et al. Insulin kinetics in type-1 diabetes: continuous and bolus delivery of rapid acting insulin. IEEE Trans. Biomed. Eng. 52, 3–12 (2004).
Ma, N., Yu, X., Yang, T., Zhao, Y. & Li, H. A hypoglycemia early alarm method for patients with type 1 diabetes based on multi-dimensional sequential pattern mining. Heliyon 8, e11372 (2022).
Contreras et al. KDH@ IJCAI 91–96.
Fushimi, E., Rosales, N., De Battista, H. & Garelli, F. Artificial pancreas clinical trials: Moving towards closed-loop control using insulin-on-board constraints. Biomed. Signal Process. Control 45, 1–9 (2018).
Loshchilov, I. & Hutter, F. Sgdr: Stochastic gradient descent with warm restarts. arXiv preprint arXiv:1608.03983 (2016).
Liu, K. et al. Machine learning models for blood glucose level prediction in patients with diabetes mellitus: systematic review and network meta-analysis. JMIR Med. Inform. 11, e47833 (2023).
Zhang, L., Yang, L. & Zhou, Z. Data-based modeling for hypoglycemia prediction: Importance, trends, and implications for clinical practice. Front. Public Health 11, 1044059 (2023).
Clarke, W. L., Cox, D., Gonder-Frederick, L. A., Carter, W. & Pohl, S. L. Evaluating clinical accuracy of systems for self-monitoring of blood glucose. Diabetes care 10, 622–628 (1987).
Wang, L., Zhang, X., Su, H. & Zhu, J. A comprehensive survey of continual learning: theory, method and application. IEEE Trans. Pattern Anal. Mach. Intell. 46, 5362–5383 (2024).
Rebuffi, S. A., Kolesnikov, A., Sperl, G. & Lampert, C. H. In Proc. of the IEEE conference on Computer Vision and Pattern Recognition. 2001–2010 (IEEE, 2017).
Madotto, A. et al. Continual learning in task-oriented dialogue systems. arXiv preprint arXiv:2012.15504 (2020).
Ruder, S. An overview of multi-task learning in deep neural networks. arXiv preprint arXiv:1706.05098 (2017).
Wang, Y., Zhai, C. & Awadalla, H. H. Multi-task learning for multilingual neural machine translation. arXiv preprint arXiv:2010.02523 (2020).
Graham, S. et al. One model is all you need: multi-task learning enables simultaneous histology image segmentation and classification. Med. Image Anal. 83, 102685 (2023).
Cho, K. et al. Learning phrase representations using RNN encoder-decoder for statistical machine translation. arXiv preprint arXiv:1406.1078 (2014).
Pan, S. J. & Yang, Q. A survey on transfer learning. IEEE Trans. Knowl. Data Eng. 22, 1345–1359 (2009).
Tajbakhsh, N. et al. Convolutional neural networks for medical image analysis: Full training or fine tuning? IEEE Trans. Med. imaging 35, 1299–1312 (2016).
Han, Z., Gao, C., Liu, J. & Zhang, S. Q. Parameter-efficient fine-tuning for large models: a comprehensive survey. arXiv preprint arXiv 2403, 14608 (2024).
Zhu, T. et al. Multi-horizon glucose prediction across populations with deep domain generalization. IEEE J. Biomed. Health Inform. 29, 5424–5437 (2024).
Chu, S. J., Amarasiri, N., Giri, S. & Kafle, P. Blood glucose level prediction in type 1 diabetes using machine learning. arXiv preprint arXiv:2502.00065 (2025).
Zhu, T., Li, K., Herrero, P. & Georgiou, P. Glugan: generating personalized glucose time series using generative adversarial networks. IEEE J. Biomed. Health Inform. 27, 5122–5133 (2023).
Domanski, P. et al. Advancing blood glucose prediction with neural architecture search and deep reinforcement learning for type 1 diabetics. Biocybern. Biomed. Eng. 44, 481–500 (2024).
Acknowledgements
This work was supported by the Institute of Information & Communications Technology Planning & Evaluation (IITP) grant funded by the Korea government (MSIT) (No. RS-2019-II191906, Artificial Intelligence Graduate School Program, POSTECH); the Pioneer Research Center Program through the NRF funded by the Ministry of Science and ICT (No. 2022M3C1A3081294); the University Technology Commercialization Promotion Program through the Commercializations Promotion Agency for R&D Outcomes (COMPA) funded by the NRF (No. RS-2024-00426901); and the Basic Science Research Program through the NRF funded by the Ministry of Education (No. RS-2025-00517742).
Author information
Authors and Affiliations
Contributions
M.H. conceptualized the study, developed the methodology, performed the data analysis, interpreted the results, and wrote the first draft of the manuscript. V.P.R. contributed to data extraction and supported the development of the methodology. J.Y. and Y.L. contributed to data collection. S.M.P. supervised the study as the corresponding author, provided critical revisions, and oversaw the overall research direction. All authors contributed to the final manuscript and approved its final version.
Corresponding author
Ethics declarations
Competing interests
V.P.R. and S.M.P. are Curestream employees and shareholders. M.H., J.Y. and Y.L. have no competing interests to disclose for the publication of this paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hwang, M., Rachim, V.P., Yoo, J. et al. Generalized multi task learning framework for glucose forecasting and hypoglycemia detection using simulation to reality. npj Digit. Med. 8, 612 (2025). https://doi.org/10.1038/s41746-025-01994-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01994-4