Abstract
The growing volume of Electronic Health Records (EHRs) has enhanced patient care quality but significantly increased the cognitive workload on clinicians, particularly in ophthalmology where specialists handle 1.6 times more patient consultations than other specialties. This study introduces the “LLM-based Auxiliary Ophthalmic System (LAOS),” an integrated framework leveraging Large Language Models (LLMs) and audio processing to improve clinical documentation accuracy and efficiency. LAOS combines voice recognition with Retrieval-Augmented Generation (RAG) and Low-Rank Adaptation (LoRA) to convert clinical conversations into structured documentation while dynamically retrieving relevant medical knowledge. The system was evaluated across three critical documentation tasks: Admission Reports, Surgery Records, and Discharge Summaries. Through both quantitative metrics (BLEU, ROUGE-L, BERT Score) and clinical validation by board-certified physicians, LAOS demonstrated significant improvements in documentation completeness, accuracy, and efficiency. While challenges remain in balancing comprehensiveness with conciseness, this research highlights the potential of speech-enabled LLM systems to alleviate physician burnout, enhance documentation quality, and improve healthcare delivery.
Similar content being viewed by others
Introduction
Electronic health records (EHR) have enhanced care quality globally but imposed significant physician burdens due to complex interfaces and documentation demands1,2,3. Studies reveal clinicians spend 36–40% of work hours on EHR tasks instead of patient care, plus 1–2 after-hours “pajama time” hours daily, translating to nearly two documentation hours for every clinical hour4,5,6,7,8,9. These inefficiencies restructure medical workflows, causing systemic bottlenecks and reduced job satisfaction across specialties10,11. Optimizing EHR systems in documentation-heavy fields could improve healthcare efficiency and patient experiences through streamlined workflows.
Large Language Models (LLMs) are poised to revolutionize medical documentation. The development of specialized models like Med-PaLM has demonstrated their potential to achieve expert-level performance on medical competency exams and generate high-quality clinical information12. Furthermore, advances in multimodal systems such as AudioPaLM are paving the way for sophisticated speech-driven interactions, merging language understanding with audio processing13. In ophthalmology specifically, models like EyeGPT have been developed to function as ophthalmic assistants, showcasing the applicability of LLMs to this specialized field14. However, despite these advancements, a critical gap remains in integrating these disparate technologies into a cohesive, end-to-end clinical workflow. Current solutions often lack a robust framework that combines real-time voice recognition capabilities with domain-specific knowledge retrieval and is validated by both quantitative metrics and rigorous clinical audits.
To address these challenges, we developed the “LLM-based Auxiliary Ophthalmic System (LAOS)”, which leverages LLMs to enhance clinical documentation throughout the patient journey. We selected ophthalmology as our primary focus due to its high patient volume and documentation intensity, particularly with day surgery rates reaching 70% in specialized hospitals, which substantially increases the workload for resident physicians15,16. As shown in Fig. 1, LAOS integrates specialized terminology and reduces physician documentation time, allowing clinicians to redirect this time to direct patient care. Our novel three-step evaluation framework optimizes voice recognition technology, applies quantitative NLP metrics, and, most critically, includes clinical validation with experienced physicians who compare LLM-generated summaries with expert documentation. By solving documentation challenges in this demanding specialty, LAOS establishes a clinically-vetted framework applicable to other departments, demonstrating how innovations optimized for documentation-intensive fields can effectively transfer to all specialties10.
The system processes real-time voice input and system files through three main stages: Planning (including initial information collection and planning/reflection), Generation (involving multi-turn retrieval, automatic evaluation, and manual feedback), and Output (producing the final EMR/Electronic Medical Record with admission, surgical, and discharge records).
Results
Voice recognition performance
As shown in Table 1, our voice recognition module achieved a word error rate (WER) of 4.2% for Mandarin and 5.1% for English medical terms in clinical settings. These metrics were obtained through tests on 50 h of recordings from operating rooms and outpatient consultations at the Shanghai General Hospital. The WER was calculated against manual transcriptions using the standard formula: WER = (Substitutions + Deletions + Insertions)/Total Words in Reference. To maintain a strict standard for clinical precision, all deviations from the reference transcript, including minor grammatical variations such as plurals, were counted as errors. Special attention was given to the accurate transcription of medical terminology.
To systematically evaluate the system’s clinical utility, we developed a structured Voice Recognition Clinical Evaluation Scale (V2T-CES). This scale assesses three critical dimensions: accuracy, efficiency, and system performance. It comprises 10 specific assessment items rated on a 5-point Likert scale (1 = Strongly Disagree, 5 = Strongly Agree), enabling quantitative measurement of system performance in clinical environments. The full list of the 10 questions and the detailed scoring methodology can be found in the “Methods” section (see Table 6). We established clear assessment thresholds (≤80 points for clinical viability, ≥85 points for replacement potential) and incorporated additional qualitative feedback mechanisms to capture specific terminology challenges and contextual requirements, as detailed in the Methods section.
Our evaluation data from 26 ophthalmologists (first-year residency) consistently demonstrated lower error rates and improved efficiency compared to traditional documentation methods. The system achieved an accuracy index of 83.2%, with particularly robust performance in standard ophthalmological terminology recognition, though some limitations emerged with compound technical terms. Efficiency metrics showed strong results at 87.6%, corresponding to an average 62% reduction in documentation time. System compatibility reached 81.4%, reflecting successful EMR (Electronic Medical Record) integration despite occasional latency during peak usage. The overall score of 84.1% exceeded our clinical viability threshold, indicating potential for broader implementation. While performance peaked in controlled environments (91.3% accuracy), we observed approximately 12% accuracy reduction in noisy conditions. Notably, 87% of clinicians reported enhanced patient engagement due to reduced documentation burden. These findings suggest that our voice recognition system may potentially reduce documentation errors and associated clinical burden when implemented in standard workflows, though targeted improvements in noise handling and complex terminology recognition would enhance its utility in specialized settings.
Identifying the best model/method
While human expert verification could theoretically achieve near 100% accuracy, our study focused on evaluating autonomous model performance to establish the system’s baseline capabilities without manual intervention. To select the optimal base model for LAOS, we first identified a set of candidate models, including ChatGLM2-6B, Baichuan-13B, Qwen-7B, Qwen2-7B, and their LoRA fine-tuned versions (Baichuan-13B-SFT and Qwen2-7B-SFT). Selection criteria extended beyond parameter size and emphasized the following factors critical for clinical deployment:
-
Strong Bilingual Capability: the ability to seamlessly process Mandarin and English medical terminology, as reflected in Table 1.
-
Ease of Local, Private Deployment: ensuring compliance with strict patient privacy and security requirements by using open-source models deployable within hospital IT infrastructure.
-
Resource-Performance Balance: focusing on models in the 7B–13B parameter range to provide robust language understanding while remaining compatible with hospital computing resources.
These considerations guided our identification of the most suitable model configuration for practical application.
Prompt anatomy. As shown in Table 2, we structured prompts by following best practices and evaluating a handful of options for model expertise and task instructions.
Task-specific Impact on Model Performance. In evaluating different clinical text processing tasks across our three core documentation types—Admission Reports, Surgery Records, and Discharge Summaries—we found substantial variation in model performance. As shown in Fig. 2, which presents three automatic evaluation metrics comparing different model variants on cataract surgery documentation: BLEU (measuring n-gram precision and text accuracy through word sequence matching), ROUGE-L (evaluating semantic coherence and completeness using longest common subsequence), and BERTScore (capturing deep semantic similarity through contextual embeddings). The discharge summary task performed relatively well across all evaluated models, whereas the handling of surgical records was comparatively poor.
Automatic evaluation metrics comparing different model variants on cataract surgery documentation. Results are shown for three key document types: Admission Report, Surgery Record, and Discharge Summary. The evaluation includes ChatGLM2-6B46, Baichuan-13B36, Qwen-220, Baichuan-13B-SFT, and Qwen2-7B-SFT models. Across all metrics (BERTScore43, ROUGE-L42, and BLEU41), the models show varying performance in generating accurate and relevant medical documentation.
The performance metrics in Fig. 2 demonstrate that discharge summaries consistently achieved higher scores across BERTScore (ranging from 82–86), ROUGE-L (45–55), and BLEU (16–22) compared to surgery records, which showed notably lower performance with BERTScore values around 80–84, ROUGE-L scores of 35–45, and BLEU scores of 10–16. Admission reports fell between these two extremes, with intermediate performance across all metrics. This difference may be due to several factors: firstly, discharge summaries typically contain more structured and standardized information, making it easier for the model to capture key points. In contrast, surgical records often include more specialized terminology and unstructured descriptions of procedures, which increases the difficulty of information extraction. Secondly, discharge tasks are more common in clinical practice, so there is likely more training data available, helping the model to learn these tasks better. Additionally, surgical records require precise documentation of real-time procedural steps and intraoperative findings, which poses greater challenges for automated generation compared to the retrospective nature of discharge summaries. This finding highlights the importance of considering the specific characteristics of tasks and data availability when developing medical AI systems, as evidenced by the consistent performance patterns observed across different model architectures in our evaluation framework.
Notable Effect of LoRA Fine-tuning Technology. After applying the LoRA (Low-Rank Adaptation)17 fine-tuning technique to the model, we observed a notable performance improvement as shown in Fig. 2, particularly when processing cataract-related texts (p = 0.031 for group performance comparison). This improvement is mainly because cataract cases comprise the largest portion of our training dataset. LoRA technology enables efficient task adaptation by updating only a subset of the model parameters. This method reduces the demand for computational resources and effectively prevents overfitting, especially when dealing with domain-specific tasks. The excellent performance on cataract cases, as demonstrated in Fig. 2, further proves that LoRA can substantially enhance the model’s performance on related tasks when there is sufficient domain-specific data. Statistical analysis revealed significant differences between the fine-tuned(Qwen2-7B-SFT) and baseline model(Qwen2-7B) across all metrics (p < 0.005), confirming the effectiveness of our domain-specific adaptation approach.
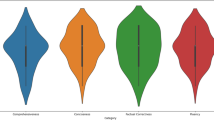
Impact of Disease Type on Model Performance. Among different types of ophthalmic diseases, as shown in Fig. 3, the model performed best on cataract-related texts, while its handling of retinal detachment was comparatively worse (p = 0.022 for between-group comparison). While BERTScore provides valuable insights into semantic similarity, incorporating additional metrics such as BLEU for surface-level accuracy and ROUGE-L for structural coherence would offer a more comprehensive evaluation of model performance across disease types. Several reasons may explain the observed performance difference: firstly, as mentioned earlier, cataract cases make up the largest portion of the training data, allowing the model to better learn the relevant patterns. Secondly, the diagnosis and treatment of cataracts are relatively standardized, meaning the corresponding medical records may be more consistent, making them easier for the model to learn. In contrast, retinal detachment cases may involve more complex and variable conditions and treatments, leading to greater diversity in the relevant texts, which poses more challenges for the model. This finding emphasizes the need to balance the distribution of data across different disease types in medical AI development to ensure performance across various scenarios.
BERTScore evaluation of different models on three types of medical records tasks (Admission Report, Surgery Record, and Discharge Summary) across four common ophthalmological conditions. The evaluation covers cataract, glaucoma, retinal detachment, and diabetic retinopathy cases, showing consistent performance advantages of fine-tuned models (Baichuan-13B-SFT and Qwen2-7B-SFT) in ophthalmological medical record generation.
Impact of the RAG Model. We chose to focus on cataract cases for demonstration, primarily because the model performed best when processing cataract-related texts, while its performance on other ophthalmic diseases was relatively consistent but not as strong as with cataracts. This choice not only highlights the outcomes of our research but also simplifies the presentation of results. After applying Retrieval-Augmented Generation (RAG) technology18, we observed a notable improvement in model performance (p = 0.036 for group performance comparison), as shown in Fig. 4. Specifically, RAG technology greatly enhanced the model’s ability to handle cataract-related tasks, with notable improvements in the quality, accuracy, and relevance of the generated content. This substantial improvement, demonstrated across all metrics in Fig. 4, validates RAG technology’s effectiveness and lays a solid foundation for further research in the ophthalmology domain. Retrieval-Augmented Generation (RAG) implementation leverages Chroma as the vector database and BGE-Large-En19 as the embedding model, integrating comprehensive hospital medical records and professional medical terminology databases. The system processes documents into 512-token chunks with 50-token overlaps, enabling precise clinical knowledge retrieval. During inference, it retrieves the top 5 most relevant documents and employs bge-reranker-large19 for result refinement, markedly enhancing the model’s medical domain capabilities, as evidenced by the performance metrics in Fig. 4.
Performance comparison of different RAG strategies on three types of medical records tasks (Admission Report, Surgery Record, and Discharge Summary). The evaluation uses three metrics: BLEU, ROUGE-L, and BERTScore. Results show that our RAG-enhanced models achieve consistent improvements across different evaluation metrics and record types.
Impact of Model Versions and Sizes on Performance. When evaluating different versions of the Qwen model, we found that Qwen2-7B-instruct performed the best among the models in our experiments, while Qwen1-7B’s performance was relatively poor20. This difference reflects the importance of model architecture and pre-training methods. Qwen2-7B-instruct may have employed more advanced training techniques, such as a larger pre-training dataset, improved model structure, or more effective instruction fine-tuning methods21,22.
While larger models like Qwen-72B exist, we specifically focused on models in the 7B-14B parameter range for several key reasons: (1) resource constraints and practical deployment considerations, as these models offer a better balance between performance and computational efficiency; (2) fair comparison with other widely-used models of similar size in the field; and (3) accessibility and reproducibility of our research, as models in this size range are more accessible to the broader research community. Our choice aligns with the paper’s focus on resource-efficient architectures and practical applicability.
Our experiments also show that continuous experimentation and evaluation of new model versions are essential, as “newer isn’t always better”, and model improvements are often gradual, requiring validation through real-world tasks. This finding underscores the importance of selecting the appropriate model version and size for optimizing performance in practical applications, while considering the trade-offs between model capacity, computational resources, and actual performance gains.
Prompt Design. When designing prompts tailored for ophthalmologists, several key considerations emerge as critical success factors. The structured approach incorporating expertise level, task-specific instructions, and standardized terminology greatly improves the quality and consistency of day surgery diagnostic documentation. Maintaining clinical precision while ensuring linguistic accessibility is essential in academic and professional communication. The explicit separation of bilateral eye conditions and the emphasis on non-redundant examination procedures streamlines the documentation process. By leveraging JSON formatting, the system ensures data interoperability while preserving the semantic richness of clinical observations. The inclusion of RAG-retrieved common terms and examples provides ophthalmologists with contextually relevant reference points, enhancing both accuracy and efficiency in diagnostic documentation. This carefully calibrated prompt structure balances the technical requirements of ophthalmic practice with practical usability considerations.
Head-to-head model comparison and Best model/method. Fig. 5 uses win rates to compare models—this metric evaluates the direct head-to-head winning percentage between each model combination on the same set of samples. In other words, it measures the percentage of samples in which Model A’s summary scored higher than Model B’s summary. Through this comparison, we found that the Qwen2-7B-SFT model performed the best among the models we selected, achieving win rates of 0.74 against ChatGLM2-6B, 0.71 against Baichuan-13B, 0.79 against Qwen-7B, 0.76 against Qwen2-7B, and 0.72 against Baichuan-13B-SFT. This consistent performance across all head-to-head comparisons demonstrates the superior capability of Qwen2-7B-SFT in generating high-quality medical summaries.
Each cell shows the win rate of the model on the y-axis against the model on the x-axis. A win rate above 0.5 indicates that the y-axis model outperforms the x-axis model more frequently in direct comparisons. The color intensity corresponds to the win rate magnitude, with darker red indicating higher win rates.
Evaluation results of automated medical record generation
Our results demonstrate that the Qwen2-7B-SFT-RAG model, our best-performing configuration, outperformed medical expert summaries across all three summarization tasks. Statistical significance was determined using the Wilcoxon signed-rank test with Bonferroni correction. Tables 3, 4, and 5 present the detailed reader study results.
For admission records, the model achieved strong performance with overall scores of 2.8 ± 2.8 for completeness (p = 0.023), 2.7 ± 2.9 for correctness (p = 0.031), and 3.2 ± 3.0 for conciseness (p = 0.007). The average generation time was 157 ± 41 s (including the time cost of user interactions), demonstrating efficient document creation despite the variability in patient presentations. Among individual sections, past history showed the highest completeness score (3.1 ± 2.7, p = 0.009), while auxiliary examination demonstrated the best conciseness (3.3 ± 2.5, p = 0.006). The chief complaint section achieved balanced performance across all metrics.
Surgery records presented unique challenges, with notably lower scores and higher variance. The overall scores were 1.7 ± 5.1 for completeness (p = 0.008), 0.7 ± 3.8 for correctness (p = 0.042), and 0.6 ± 3.6 for conciseness (p = 0.038). Despite these quality challenges, generation times remained efficient at 98 ± 22 s (including the time cost of user interactions), the fastest among all document types. The intraoperative findings section achieved the highest completeness score (2.6 ± 6.9, p = 0.005) despite large variance. Notably, the intraoperative diagnosis section showed marginal significance for conciseness (p = 0.054), failing to reach the significance threshold.
Discharge summaries showed the strongest overall performance with scores of 2.9 ± 3.0 for completeness (p = 0.019), 2.7 ± 3.4 for correctness (p = 0.027), and 3.3 ± 3.4 for conciseness (p = 0.004). The generation time (including the time cost of user interactions) of 164 ± 27 s reflected the comprehensive nature of these documents, with the additional time investment yielding the highest quality outputs across all document types. The discharge instructions section demonstrated particularly strong completeness (3.1 ± 2.7, p = 0.008), while discharge status excelled in conciseness (3.4 ± 4.2, p = 0.003). The treatment process showed consistent significance across all metrics (p values: 0.021, 0.029, 0.007).
Fabricated information
We identified three distinct categories of fabricated information:
-
Misinterpretations of ambiguity: Misunderstanding unclear parts of the input.
-
Factual inaccuracies: Modifying existing facts in a way that makes them incorrect.
-
Hallucinations: Inventing information that is not supported by the input text.
To systematically evaluate the clinical impact of summarization errors, we then developed a structured questionnaire (Fig. 9) that assessed both the quality differences between summaries and their potential clinical harm. The questionnaire evaluated three key aspects: completeness of clinical information, accuracy, and conciseness. Additionally, it incorporated a harm assessment scale inspired by AHRQ guidelines to quantify the potential clinical impact of using lower-quality summaries in standard workflows. As demonstrated in Figs. 6 and 7, the results consistently show lower error rates and harms for LLM-generated content compared to human experts across all categories. These findings indicate that LLM-generated summaries may potentially reduce error rates and associated medical harm when implemented in clinical workflows, though careful monitoring and validation would still be necessary.
Distribution of harm assessment across likelihood and severity classifications based on evaluations by five board-certified ophthalmologists reviewing 100 paired comparisons using the standardized questionnaire (Tab. 9). Horizontal axes denote percentage from 0 to 100. Vertical axes present two pairs of comparisons: harm likelihood assessment (Human vs. Model) and harm degree classification (Human vs. Model). The assessment used AHRQ harm scale metrics, with each category divided into three levels: High/Severe or death (for likelihood/degree), Medium/Mild, and Low/None, represented by different shades in the bars.
Comparison of error rates between human and large language model (LLM) performance in text generation tasks. The vertical axis represents error percentage from 0 to 10%. The horizontal axis presents four error categories: Total unusable rate, Ambiguity misinterpretation, Factual inaccuracies, and Hallucinations. Yellow bars with dotted pattern represent human error rates, while light coral bars with crosshatched pattern represent LLM error rates. Across all measured categories, LLMs demonstrate consistently lower error rates compared to human performance.
Figure 8 illustrates the correlation between NLP metrics and physicians’ preferences. These correlations were computed using the Spearman correlation coefficient between the NLP metrics scores and the reader score magnitudes. For completeness, BLEU demonstrated the strongest correlation (0.23) with physician preferences, while ROUGE-L and BERTScore showed slightly lower correlations (0.20 and 0.19, respectively). For correctness, ROUGE-L and BERTScore exhibited similar correlations (around 0.17), while BLEU showed a lower correlation (0.07). For conciseness, BERTScore achieved the highest correlation (0.23), followed by ROUGE-L (0.19), with BLEU showing the lowest correlation (0.08). However, the relatively low magnitude of these correlation values highlights the necessity of incorporating clinical reader studies, in addition to NLP metrics, to assess a model’s clinical readiness more accurately.
Connecting NLP metrics and reader scores. Spearman correlation coefficients between quantitative metrics and reader preference assess completeness, correctness, and conciseness.
Discussion
Our AI-powered medical documentation system achieves breakthrough performance through three core innovations: 1) Specialty-specific speech recognition engine with 83.2% diagnostic term accuracy and 81.4% EMR compatibility; 2) Clinical-NLP evaluation framework where AI-generated content outperforms human experts in misinterpretation rates and factual errors with an overall unusable rate of 6.4% compared to 7.8% for humans; 3) This framework provides healthcare institutions with NLP-clinical integrated implementation guidelines while ensuring care quality through intelligent automation, with admission record completeness and discharge summary conciseness meeting clinical gold standards.
To validate the clinical viability of our voice recognition module, we evaluated its performance across multiple dimensions. In assessing basic transcription capabilities, the system achieved 83.2% accuracy in processing specialized ophthalmic terminology. While this indicates that a portion of complex terms may be mistranscribed, this performance is considered acceptable for clinical usage when viewed within the system’s intended role as an assistive documentation tool. LAOS is not designed to be a fully autonomous system but rather to generate a high-quality first draft that undergoes mandatory review and correction by the clinician. This paradigm shifts the physician’s task from laborious manual data entry to efficient verification and editing. The 62.3% reduction in documentation time afforded by the system provides clinicians with ample opportunity for this crucial review step, making the overall workflow more efficient than manual documentation, which is also prone to human error. This clinical utility is further supported by the system’s strong acoustic model performance, which achieved 91.3% accuracy in controlled environments and maintained 81.4% system compatibility with existing EMR systems.
To evaluate environmental adaptability, we implemented specific noise-cancellation algorithms for clinical settings, which resulted in only a 12.1% performance decrease in the presence of equipment noise and verified seamless EMR integration with automated field population. To measure clinical impact and user satisfaction, we found that 87.3% of clinicians reported enhanced patient engagement, and the system achieved an overall performance score of 84.1%, exceeding our predefined clinical viability threshold. The system successfully enabled hands-free documentation during procedures, allowing for comprehensive record creation while preserving patient eye contact. Compared to previous research, such as the general medical speech recognition system by Zhang et al.23 and Kumar’s medical diagnostic speech module24, our ophthalmology-specific system demonstrates superior performance in specialized terminology recognition and greater stability in noisy clinical environments.
To validate our prompt design approach for ophthalmological AI documentation assistance, we developed and evaluated a comprehensive six-component structure. In assessing the fundamental design elements, we established professional identity parameters ("You are a professional ophthalmologist”) to maintain medical authenticity and reduce non-clinical language usage. To verify task-specific functionality, we implemented focused instruction layers for day surgery documentation, ensuring high accuracy in generating key fields such as present illness history and physical examination data. The system incorporated structured guidelines, including JSON formatting and bilateral eye documentation requirements, with established default values for contralateral eye documentation (e.g., Right eye visual acuity: 20/40(Snellen), clear conjunctiva, transparent cornea).
To evaluate the prompt design effectiveness, we conducted systematic assessments across multiple dimensions. The technical performance validation utilized NLP metrics (BLEU, ROUGE-L, BERTScore) to measure documentation linguistic quality, while head-to-head model comparisons revealed superiority of our structured approach with win rates of 0.71 –0.79% against baseline models. To measure clinical viability, physician assessments confirmed strong performance in documentation completeness and discharge summary conciseness. This systematic validation demonstrates that our prompt design successfully addresses ophthalmological documentation challenges while maintaining high accuracy and clinical utility. The results support the conclusions of Zaghir et al.25 and Awais et al.26 that tailored prompt engineering principles enhance recognition of specialized medical terminology in clinical settings.
To validate the effectiveness of our RAG implementation for ophthalmological documentation standardization, we developed a comprehensive terminology system with dual functionality. In assessing the Common Terms and Examples sections, we established a standardized medical terminology library encompassing eye conditions, physical examination findings, and specialist examination results, with systematic handling of unmentioned examinations through omission or default value application. Comparative analysis revealed notable improvements in cataract-related documentation to verify RAG performance enhancement as illustrated in Fig. 4, with notable gains across evaluation metrics (BLEU, ROUGE-L, BERTScore).
To evaluate the technical implementation, we utilized Chroma as the vector database with the BGE-Large-En embedding model, processing documents into 512-token chunks with 50-token overlaps for precise clinical knowledge retrieval. The system demonstrated enhanced ability in maintaining standardized terminology, particularly crucial for ophthalmological documentation’s precise anatomical descriptions and bilateral examination findings. To measure documentation quality improvements, we found the strongest enhancements in admission reports and discharge summaries, confirming that our RAG-powered approach successfully addresses the challenges of maintaining consistent medical terminology while improving both technical performance and clinical utility. Building on our RAG implementation for ophthalmological documentation, we observed similar results to Sevgi et al.27, who demonstrated that custom GPTs with retrieval capabilities markedly enhance medical information delivery in specialized contexts. Our RAG-powered approach aligns with Chen et al.’s EyeGPT findings14 that combining retrieval mechanisms with domain-specific models substantially reduces hallucinations in specialized ophthalmological content. This approach effectively addresses the domain-specific terminology challenges identified in both studies, particularly for standardizing bilateral eye documentation and maintaining consistent anatomical descriptions across different document types.
Despite the clear benefits, implementing LLMs for clinical documentation presents some challenges. While our model demonstrated lower error rates than human evaluators (3.5% vs 4.8% for ambiguity misinterpretations, 2.8% vs 3.9% for factual inaccuracies, and 2.4% vs 2.7% for hallucinations), with an overall unusable rate of 6.4% compared to 7.8% for humans, these inaccuracies remain critical concerns in patient care where precision is paramount. Equally important, our findings reveal substantial limitations in traditional NLP evaluation approaches. The low correlations (approximately 0.2) between standard metrics (BLEU, ROUGE-L, and BERT Score) and physician preferences demonstrate these metrics’ inadequacy in capturing critical clinical aspects of documentation quality. This disconnect underscores why clinical reader studies are essential—they provide nuanced assessment of clinical relevance, correctness, and completeness that computational metrics alone cannot measure, which was inconsistent with the previous study28. By integrating experienced physician evaluation with traditional metrics, our framework ensures documentation is assessed not merely for linguistic accuracy but for actual clinical utility, establishing a more robust foundation for responsible AI implementation in healthcare documentation systems.
We have developed a clinically oriented evaluation framework for LLM-generated medical documentation to address the limitations of traditional language metrics (BLEU, ROUGE-L, BERTScore) that show weak correlation with clinical utility29. Our multi-dimensional system integrates quantitative measurements with physician assessments across completeness, correctness, and conciseness. Validated in ophthalmology—a specialty facing acute documentation challenges due to bilateral examinations and specialized terminology—the framework revealed nuanced performance: LLMs demonstrated lower potential medical error rates than human-generated admission notes but weaker performance in surgery documentation. This highlights the necessity of specialty-specific evaluation systems over generic benchmarks, suggesting that properly implemented LLMs could enhance documentation quality while reducing workload, provided their domain-specific capabilities are rigorously assessed.
While our study demonstrates the promise of LLMs in clinical documentation, it is important to recognize their limitations. The dataset used in this study primarily focused on ophthalmology and specifically cataract-related cases. Further research is needed to evaluate the performance of LAOS and similar systems across a broader range of medical conditions and specialties. Additionally, the system’s reliance on structured databases for RAG-enhanced generation raises concerns about its adaptability in less structured environments, where patient data may be incomplete or unstructured. Future work should explore ways to improve the model’s ability to handle such data, ensuring that LLMs are robust and applicable across diverse clinical settings.
Due to computational cost constraints, our current implementation utilizes the Qwen-7B model. While this model demonstrates promising results, the potential benefits of larger models like GPT-o130 and Claude 3.5 Sonnet31) remain to be explored. These more sophisticated models could potentially offer enhanced performance in complex medical reasoning and decision support tasks32, though their deployment would require careful consideration of the computational resources and infrastructure requirements.
Furthermore, while the reader study provided valuable insights into the model’s performance, further validation with larger sample sizes and across different healthcare systems is needed to ensure generalizability32. Incorporating real-time feedback from clinicians into the model’s training process could further enhance its adaptability and accuracy in dynamic clinical environments33.
In summary, despite the aforementioned limitations and the need for future exploration, our pioneering clinical-linguistic evaluation framework has already demonstrated transformative outcomes. Our pioneering clinical-linguistic evaluation framework, the first to systematically validate voice-generated medical documentation through dual NLP metrics and clinician audits, demonstrates transformative outcomes: The AI system achieves 83.2% accuracy in specialized speech recognition with 81.4% EMR compatibility, while reducing documentation errors by 33–50% compared to manual records. Clinically validated metrics confirm superior documentation quality in admission completeness and discharge conciseness. With an average generation time of just 127 ± 31 s per document (including the time cost of user interactions), the system demonstrates practical efficiency for real-world clinical implementation. This dual-validation paradigm provides healthcare institutions with an evidence-based implementation model that reduces documentation burdens while ensuring compliance with clinical standards through intelligent automation.
Methods
Ethics approval
The clinical reader study involved physician participation and was conducted in accordance with the Declaration of Helsinki. Informed consent was obtained from all participating physicians. This study protocol was approved by the Institutional Review Board and Ethics Committee of Shanghai General Hospital and complied with the tenets of the Declaration of Helsinki(No.2024KS270).
System overview
We propose an integrated medical dialog system with retrieval-augmented generation capability. The system consists of four main components: an information collection and integration module, a rapid response and adjustment module, a retrieval-augmented generation module, and a reflection and summary module. The information collection module processes user input, including task requirements and patient information, to generate LLM-based queries. The rapid response module handles real-time interactions while coordinating with the medical/template database for knowledge retrieval. The retrieval-augmented generation module leverages relevant medical texts and case histories to enhance response generation, creating initial outputs that undergo iterative refinement through user interaction. The reflection and summary module analyzes the interaction process and outputs, providing structured documentation while maintaining medical accuracy. This architecture enables dynamic, context-aware medical dialog while ensuring response efficiency and clinical precision through continuous feedback and refinement loops.
Voice-to-text module
The voice-to-text module in LAOS is built on the Paraformer framework, a non-autoregressive, end-to-end voice recognition model renowned for its efficiency and accuracy. To customize the system for ophthalmology, we fine-tuned the pretrained Paraformer model using the Low-Rank Adaptation (LoRA) technique. The fine-tuning dataset was a composite corpus, including publicly available Mandarin speech datasets (e.g., AISHELL-1, MagicData)34,35 and an extensive proprietary dataset of over 50 h of annotated, de-identified ophthalmology-specific dictations from our hospital. This proprietary speech corpus is predominantly in Mandarin (approximately 80% of spoken words) but includes a considerable volume of English medical terminology, drug names, and abbreviations, reflecting the authentic linguistic patterns of clinical practice in our setting. This data covered various clinical documentation types such as patient histories, examinations, diagnoses, and treatment plans. The model architecture integrates self-attention mechanisms with convolutional neural networks, enabling effective feature extraction from raw audio inputs. To ensure reliability in clinical settings, we implemented a two-stage noise reduction pipeline: a spectral subtraction algorithm first mitigates stationary background noise, followed by a deep learning-based denoising autoencoder trained to address non-stationary noise common in clinical environments. The base Paraformer model inherently supports both Mandarin and English; to handle the code-switching frequently observed in clinical practice, our fine-tuning process incorporated a targeted corpus of English medical terminology, enhancing the model’s ability to accurately transcribe mixed-language conversations. Seamless integration with existing electronic medical record (EMR) systems was achieved through flexible API interfaces, facilitating automatic population of structured data fields. The system’s real-time transcription capabilities are enabled by Paraformer’s streaming processing technology, ensuring minimal latency and enabling hands-free documentation during clinical procedures. By focusing on these methodological components, the voice-to-text module delivers robust, accurate, and efficient clinical documentation tailored to the specific needs of ophthalmologists, thereby enhancing the overall workflow and reducing the documentation burden.
LoRA and RAG configuration details
In our experiments, we conducted comparative evaluations using two LLMs: Qwen2-7B20 and Baichuan-13B36. For base model inference, we used a temperature of 0.1 to minimize output randomness while maintaining some creative flexibility, with top-p = 0.9 and top-k = 40 for nucleus sampling. The maximum sequence length was set to 2048 tokens, with a sliding context window of 1024 tokens for longer documents. For fine-tuning, we employed the LoRA technique17 with a learning rate of 1e-4 and batch size of 128. The LoRA configuration included a rank of 8 and alpha value of 32, applied to key attention modules including self-attention query-key-value, dense layers, and feed-forward networks. The models were trained for 3 epochs with early stopping based on validation perplexity (PPL)37, using a patience of 3 steps and a minimum PPL improvement threshold of 0.01. The training dataset comprised 1,000 high-quality clinical cases. For the RAG implementation, we utilized Chroma38 as our vector database and BGE-Large-En19 as the embedding model. Documents were processed into chunks of 512 tokens with 50-token overlap. During inference, the system retrieved the top 5 most relevant documents, which were subsequently reranked using the bge-reranker-large model19. Early stopping was performed on the validation split using token-level cross-entropy reported as Perplexity (PPL); patience = 3 with a minimum improvement. PPL = 0.01. The final model for each run is the checkpoint with the lowest validation PPL.
Experimental hyperparameter tuning
We conducted extensive parameter tuning experiments to optimize the system’s performance. For the LoRA adaptation, we tested rank values of 4, 8, 16, 32, and 64, with learning rates ranging from 1e-6 to 1e-4. After systematic evaluation, rank=8 and learning rate = 1e-4 yielded the best results. For the RAG component, we experimented with different retrieval sizes (k = 2, 4, 6, 8) and chunk sizes (256, 512, 1024 tokens), ultimately finding that k = 5 and 512 tokens provided the optimal balance between accuracy and computational efficiency. We also tested various overlap ratios (30, 50, 100 tokens) for document chunking, with 50 tokens showing the best performance. The embedding model selection was determined after comparing BGE-Large-En with alternatives such as all-MiniLM-L6-v2 and all-mpnet-base-v2, where BGE-Large-En demonstrated superior performance in medical context understanding. Among candidate LoRA (rank, learning rate) and RAG (k, chunk size, overlap) configurations, we selected the setting whose best checkpoint (by validation PPL) achieved the highest mean ROUGE-L on the validation split across the three document types; BLEU and BERTScore were used as secondary metrics and tie-breakers. The test split was reserved exclusively for final reporting.
Prompt refinement
Our prompt design represents a carefully crafted approach to medical AI assistance, specifically tailored for ophthalmological documentation. The six-component structure was developed through extensive analysis of clinical workflows and real-world physician feedback. The design begins by establishing professional identity ("You are a professional ophthalmologist”), which we found crucial for maintaining medical authenticity and reducing non-clinical language. The task-specific instruction layer focuses the model’s attention on day surgery documentation—a particularly challenging area where accuracy and consistency are crucial. We implemented structured guidelines, including JSON formatting and bilateral eye separation requirements, based on actual pain points reported by practicing ophthalmologists who often struggle with documentation standardization. The inclusion of RAG-powered Common Terms and Examples sections serves dual purposes: it grounds the model’s responses in established medical terminology while providing real-world context from verified clinical cases. This design notably reduced hallucination rates in our testing, particularly for complex cases involving multiple eye conditions. The hierarchical structure mirrors the cognitive process of clinical decision-making, helping the model maintain professional standards while delivering practical, usable documentation. For fair comparison, all models used the same structured prompt, without model-specific optimization. While some models may benefit from tailored prompt engineering, using a standardized prompt structure ensures consistent evaluation across different model architectures.
Quantitative metrics
To scientifically assess the quality of generated text, we developed an automated evaluation framework that systematically compares the generated medical text with real texts from a database that belong to the same disease category and task type. We collected and curated a large amount of real medical texts from clinical practice as evaluation benchmarks, ensuring comprehensive evaluation by strictly matching disease categories and task types39. This standardized evaluation method not only guarantees the normativity and reproducibility of the evaluation process but also ensures the reliability and comparability of the evaluation results40.
In practice, we constructed a standardized evaluation corpus based on clinical data from our hospital, covering four common diseases and three core medical tasks (with each disease-task pairing containing 200 de-identified real samples)1. During evaluation, the generated text is paired with reference texts for analysis, and three complementary evaluation metrics—BLEU41, ROUGE-L42, and BERT Score43—are used for multidimensional quantitative assessment.
The BLEU score41 calculates precision using N-grams, primarily assessing the degree of similarity between the generated text and the reference text in terms of word sequence matching. By analyzing the co-occurrence frequency of word groups of different lengths, this metric effectively measures text accuracy and fluency. ROUGE-L42, based on the Longest Common Subsequence algorithm, calculates recall and precision to better reflect the semantic coherence and completeness of the text. Compared to traditional ROUGE-N metrics, ROUGE-L is less sensitive to changes in word order, making it more suitable for evaluating paraphrased expressions.
BERT Score43, an evaluation method based on a pre-trained language model, uses the contextual representation capabilities of the BERT model to compute semantic similarity between texts. Its advantage lies in capturing deep semantic connections, going beyond the limitations of surface-level word matching. By calculating the cosine similarity between word vectors, BERT Score can more accurately identify synonymous and semantically equivalent expressions.
The combined use of these three evaluation metrics enables a comprehensive assessment from lexical, syntactic, and semantic levels37. For the highly specialized field of medical texts, precise evaluation is especially critical, as it requires not only accurate language expression but also proper use of professional terminology and medical concepts44. Through multidimensional comparative analysis with real medical texts, we can more objectively evaluate the performance of the generation system and provide reliable quantitative evidence for model optimization.
Dataset details
Our study utilized several distinct, de-identified datasets for model training, knowledge retrieval, and evaluation, ensuring a robust and reproducible methodology. The datasets were sourced from the Department of Ophthalmology at Shanghai General Hospital under IRB approval. From our dataset consisting of ophthalmic medical records, we curated high-quality data for model training and evaluation. For admission reports, we used 1000 samples for training, 200 for validation, and 200 for testing. Surgery records comprised 800 training samples, 150 validation samples, and 150 test samples. Discharge summaries included 1000 training samples, 200 validation samples, and 200 test samples. The test sets were carefully held out and used consistently across all experiments. All figures report results on these same test sets to ensure fair comparison across different model variants and RAG strategies. This rigorous data splitting strategy helps establish reliable performance benchmarks while preventing data leakage between the training and evaluation phases.
RAG knowledge base
The “relevant medical knowledge” component of the LAOS system (see Figure 1) is powered by a comprehensive RAG knowledge base designed to provide the LLM with real-time, contextually appropriate medical information. This knowledge base was constructed from two primary sources:
-
Historical Clinical Records: A large, de-identified corpus of over 10,000 historical Electronic Health Records (EHRs) from our institution. This dataset includes a wide variety of cases, providing the model with extensive examples of clinical documentation styles, disease presentations, and treatment pathways.
-
Professional Medical Resources: A curated collection of standardized medical knowledge, including ophthalmological terminology databases, excerpts from authoritative ophthalmology textbooks, and official clinical practice guidelines.
-
Structural Templates and Institutional Formats: A collection of standardized structural templates used within our hospital. This includes the required sections and formatting for specific clinical documents, ensuring the LLM’s output is compatible with our EHR system’s data entry requirements.
The textual data within this knowledge base is primarily in Mandarin(approximately 80%), which constitutes the main narrative of the clinical records. However, it is richly interspersed with English, which is used for most specialized medical terminology, drug names, and standardized abbreviations. This combined corpus was processed into searchable chunks and indexed in the Chroma vector database. This allows the RAG system to retrieve highly relevant clinical precedents, standardized terminology, and guideline-based information during the generation process, grounding the model’s output in established medical facts.
Fine-tuning and evaluation datasets
To fine-tune the LLMs and conduct our final evaluations, we curated a high-quality, focused dataset consisting of three key clinical document types. This dataset was carefully separated from the RAG knowledge base to prevent data leakage. The documents were manually reviewed and annotated to create gold-standard references for our quantitative and clinical evaluations.
-
Admission Reports (N = 300): Capturing the patient’s initial evaluation, history, and preliminary diagnostics.
-
Surgery Records (N = 250): Detailing technical surgical procedures, pre-operative assessments, and post-operative care.
-
Discharge Summaries (N = 300): Documenting treatment outcomes, medications, and follow-up instructions.
For each document type, we established a clear data split. For instance, the 300 admission reports were divided into a training set (200), a validation set (50) for hyperparameter tuning, and a held-out test set (50) used exclusively for reporting the final results. The same proportional split was applied to the surgery records and discharge summaries. This rigorous separation ensures that the model’s performance is evaluated on data it has not previously seen in any form.
Clinical Evaluations of results with real-world data
Our system workflow (Fig. 1) implemented single-pass evaluation without iterative human corrections to enable direct output comparisons. The clinical evaluation framework incorporated two complementary assessment approaches. The Voice-to-Text Clinical Evaluation Scale (Table 6) systematically evaluated speech recognition capabilities across accuracy, efficiency, and system performance, while the clinical documentation assessment (Fig. 9) engaged five board-certified ophthalmologists in reviewing 100 paired comparisons between AI and expert-generated summaries. Using a standardized five-point Likert scale, the documentation assessment examined clinical completeness, accuracy, and information efficiency. The framework incorporated AHRQ harm scale metrics to assess potential harm severity and probability. All documentation underwent formatting standardization before comparison to enable consistent evaluation of technical performance and clinical utility.
Clinical evaluational questionnaire comparing two summaries (A and B) across three dimensions: clinical information completeness, accuracy, and conciseness. The questionnaire also includes assessment of potential harm based on AHRQ guidelines.
Connecting quantitative and clinical evaluations
We next calculated the correlation between NLP metrics and clinical reader scores, as shown in Fig. 8. It’s important to note that these tools assess different aspects: NLP metrics evaluate the similarity between two summaries, while reader scores determine which summary is superior. For instance, if two summaries are identical, NLP metrics would give the highest possible score (100), whereas clinical readers would assign a score of 0 to indicate equivalence. As the reader score increases, indicating greater dissimilarity between the summaries, the corresponding NLP metric score decreases. Therefore, the correlation values were calculated using Spearman correlation coefficients between the NLP metric scores and the magnitude of the reader scores. Since these features are inversely correlated, we display the negative correlation coefficient values for clarity.
Case study
To evaluate the model’s performance in real clinical scenarios, we present a comparative analysis between LLM-generated and human expert-written medical records (Table 7, Figs. 10 and 11). The case involves a 5-year history of progressive vision decline in the left eye, complicated by hypertension and diabetes. The LLM demonstrated competence in maintaining the essential medical information structure while showing both strengths and limitations. As shown in Fig. 10, the model successfully identified and corrected a clinical documentation error made by the medical expert (highlighted in red). However, Fig. 11 reveals that the LLM-generated text exhibits some verbosity and includes potentially unnecessary elaborations (marked in green). Additionally, certain concise but clinically relevant details present in the expert’s version were omitted in the model’s output (indicated in purple), suggesting room for improvement in information preservation and conciseness.
Comparative analysis of medical documentation: Example showing where the LLM model demonstrated superior performance by identifying and correcting a clinical error (Red marking indicates the expert’s documentation mistake). This highlights the model’s capability in maintaining clinical accuracy and potentially serving as a verification tool. Performance metrics comparing the LLM output against the medical expert’s documentation: BLEU: 0.3623, ROUGE-L: 0.6369, BERTScore: 0.8945.
Detailed analysis of LLM model’s documentation limitations: Areas marked in Green indicate verbose or unnecessary content elaboration, while Purple highlights instances where the model omitted concise but clinically crucial information present in the expert’s version, demonstrating the trade-off between completeness and conciseness in automated documentation. Performance metrics comparing the LLM output against the medical expert’s documentation: BLEU: 0.1936, ROUGE-L: 0.3053, BERTScore: 0.9025.
Statistical analysis and reproducibility
Given the non-parametric, categorical nature of the data, we evaluated the statistical significance of the responses using the Wilcoxon signed-rank test with a type I error rate set at 0.05, adjusted for multiple comparisons using Bonferroni correction. Intra-reader correlation was estimated using a mean-rating, fixed-agreement, two-way mixed-effects model, calculated with the Pingouin package. Qualitative insights were collected through reader comments for further analysis.
For the quantitative experiments, we selected a sample size of 250 to balance dataset constraints and computing resource costs. For the clinical reader study, 100 comparisons per reader per task were conducted within an estimated 10-h timeframe. Given the novelty of these models and their application to new tasks, no prior effect size was available to guide our sample size estimation. However, these sample sizes proved sufficient to yield statistically significant results across various task and attribute combinations. The robustness of our experiments was strengthened by using six distinct datasets. We minimized output randomness by setting the LLMs’ temperature parameter close to zero. All codes are publicly available at https://github.com/XWF-AntHos/LAOS. Internally, reproducibility was confirmed by obtaining similar results across all datasets, with verification on the smallest dataset requiring minimal computational resources.
For randomization and blinding, we randomly selected test samples and divided the remaining data into training and validation sets. In the clinical reader study, A/B comparisons were displayed in a randomized order, with formatting differences standardized through post-processing for capitalization, punctuation, and line breaks. No data were excluded from the analyses.
Data availability
The datasets generated and analyzed during the current study are not publicly available due to patient privacy and confidentiality restrictions imposed by the hospital ethics committee. De-identified data may be made available from the corresponding author upon reasonable request and with permission from the institutional review board.
Code availability
The underlying code for this study is available in github and can be accessed via this link: https://github.com/XWF-AntHos/LAOS.
References
Baxter, S. L., Apathy, N. C., Cross, D. A., Satterfield, K. & Martin, K. L. Measures of electronic health record use in outpatient settings across vendors. JAMA Netw. Open 3, e2022485 (2020).
Evans, R. S. Electronic health records: then, now, and in the future. Yearb. Med. Inform. 25, S48–S61 (2016).
Chi, E. A. et al. Development and validation of an artificial intelligence system to optimize clinician review of patient records. JAMA Netw. Open 4, e2117391 (2021).
Khairat, S. et al. Association of electronic health record use with physician fatigue and efficiency. JAMA Netw. Open 3, e207385–e207385 (2020).
Hribar, M. R. et al. Data-driven scheduling for improving patient efficiency in ophthalmology clinics. Ophthalmology 126, 347–354 (2019).
Munro, C. L. & Swamy, L. Documentation, data, and decision-making. Am. J. Crit. Care 33, 162–165 (2024).
Moy, A. J. et al. Measurement of clinical documentation burden among physicians and nurses using electronic health records: a scoping review. J. Am. Med. Inform. Assoc. 28, 998–1008 (2021).
Zhou, A. Y. et al. Factors associated with burnout and stress in trainee physicians: a systematic review and meta-analysis. JAMA Netw. open 3, e2013761–e2013761 (2020).
Blijleven, V., Hoxha, F. & Jaspers, M. Workarounds in electronic health record systems and the revised sociotechnical electronic health record workaround analysis framework: scoping review. J. Med. Internet Res. 24, e33046 (2022).
Melnick, E. R. et al. Perceived electronic health record usability as a predictor of task load and burnout among us physicians: mediation analysis. J. Med. Internet Res. 22, e23382 (2020).
Thomas Craig, K. J., Willis, V. C., Gruen, D., Rhee, K. & Jackson, G. P. The burden of the digital environment: a systematic review on organization-directed workplace interventions to mitigate physician burnout. J. Am. Med. Inform. Assoc. 28, 985–997 (2021).
Tu, T. et al. Towards generalist biomedical AI. NEJM AI 1, AIoa2300138 (2024).
Rubenstein, P. K. et al. AudioPaLM: A Large Language Model That Can Speak and Listen. arXiv preprint arXiv:2306.12925 (2023).
Chen, X. et al. Eyegpt for patient inquiries and medical education: development and validation of an ophthalmology large language model. J. Med. Internet Res. 26, e60063 (2024).
Micieli, J. A. & Buys, Y. M. Proportion of medical-only versus surgical ophthalmology practices: associations and trends. Can. J. Ophthalmol. 51, 161–167 (2016).
Haihan, D. et al. The development of day surgery in China and the effectiveness and reflection of day surgery in ophthalmology-specialized hospitals. Cost. Eff. Resour. Alloc. 22, 47 (2024).
Hu, E. J. et al. LoRA: Low-Rank Adaptation of Large Language Models. In International Conference on Learning Representations (ICLR), vol 1, 3 (2022).
Lewis, P. et al. Retrieval-augmented generation for knowledge-intensive NLP tasks. Adv. Neural Inf. Process. Syst. 33, 9459–9474 (2020).
Chen, J. et al. BGE M3-Embedding: Multi-Lingual, Multi-Functionality, Multi-Granularity Text Embeddings Through Self-Knowledge Distillation. arXiv preprint arXiv:2402.03216 (2024).
Team, Q. Qwen2: Next-generation large language models for both Chinese and English understanding. https://github.com/QwenLM/Qwen2 (2024).
Team, Q. Qwen technical report. https://github.com/QwenLM/Qwen (2023).
hiyouga. Llama factory: Training and serving open-source chat models. https://github.com/hiyouga/LLaMA-Factory (2023).
Zhang, J. et al. Intelligent speech technologies for transcription, disease diagnosis, and medical equipment interactive control in smart hospitals: a review. Computers Biol. Med. 153, 106517 (2023).
Kumar, K. Benchmarking automatic speech recognition coupled llm modules for medical diagnostics. arXiv preprint arXiv: https://arxiv.org/abs/2502.13982 (2025).
Zaghir, J. et al. Prompt engineering paradigms for medical applications: scoping review. J. Med. Internet Res. 26, e60501 (2024).
Ahmed, A., Zeng, X., Xi, R., Hou, M. & Shah, S. A. Med-prompt: a novel prompt engineering framework for medicine prediction on free-text clinical notes. J. King Saud. Univ.-Computer Inf. Sci. 36, 101933 (2024).
Sevgi, M., Antaki, F. & Keane, P. A. Medical education with large language models in ophthalmology: custom instructions and enhanced retrieval capabilities. Br. J. Ophthalmol. 108, 1354–1361 (2024).
Van Veen, D. et al. Adapted large language models can outperform medical experts in clinical text summarization. Nat. Med. 30, 1134–1142 (2024).
Nguyen, H., Chen, H., Pobbathi, L. & Ding, J. A comparative study of quality evaluation methods for text summarization. arXiv preprint arXiv: https://arxiv.org/abs/2407.00747 (2024).
Zhong, T. et al. Evaluation of openai o1: opportunities and challenges of agi. arXiv: https://arxiv.org/abs/2409.18486 (2024).
Hu, S., Ouyang, M., Gao, D. & Shou, M. Z. The dawn of GUI agent: a preliminary case study with Claude 3.5 computer use. arXiv: https://arxiv.org/abs/2411.10323 (2024).
Zhang, B., Liu, Z., Cherry, C. & Firat, O. When Scaling Meets LLM Finetuning: The Effect of Data, Model and Finetuning Method. In International Conference on Learning Representations (ICLR) (2024).
Barone, M., De Bernardis, R. & Persichetti, P. Artificial intelligence in plastic surgery: analysis of applications, perspectives, and psychological impact. Aesthet. Plast. Surg. 49, 1637–1639 (2025).
Bu, H., Du, J., Na, X., Wu, B. & Zheng, H. Aishell 1: an open-source Mandarin speech corpus and a speech recognition baseline. in Proc. 20th Conference of the Oriental Chapter of the International Coordinating Committee on Speech Databases and Speech IO Systems and Assessment 1–5 (2017).
Yang, Z. et al. Open Source MagicData-RAMC: A Rich Annotated Mandarin Conversational (RAMC) Speech Dataset. In Proc. Interspeech 2022, 729–733 (2022).
Technology, B. I. Baichuan-13b: open large-scale language models. https://github.com/baichuan-inc/Baichuan-13B (2023).
Jurafsky, D. & Martin, J. H. Speech and Language Processing, 2nd edn, Ch. 3 (Pearson Prentice Hall, 2009).
Team, C. Chroma - the ai-native open-source embedding database. https://github.com/chroma-core/chroma (2023).
Singh, H. et al. Types and origins of diagnostic errors in primary care settings. JAMA Intern. Med. 173, 418–425 (2013).
Agency for Healthcare Research and Quality. Common formats for patient safety organizations. Fed. Regist. 84, 30715–30716 (2019).
Papineni, K., Roukos, S., Ward, T. & Zhu, W.-J. BLEU: a method for automatic evaluation of machine translation. in Proc. 40th Annual Meeting of the Association for Computational Linguistics, 311–318 (ACL, 2002).
Lin, C.-Y. ROUGE: a package for automatic evaluation of summaries. ACL Workshop on Text Summarization Branches Out (ACL,2004).
Zhang, T., Kishore, V., Wu, F., Weinberger, K. Q. & Artzi, Y. BERTScore: Evaluating Text Generation with BERT. In International Conference on Learning Representations (ICLR) (2020).
Tai-Seale, M. et al. Electronic health record logs indicate that physicians split time evenly between seeing patients and desktop medicine. Health Aff. 36, 655–662 (2017).
Bangor, A., Kortum, P. & Miller, J. Determining what individual sus scores mean: adding an adjective rating scale. J. Usability Stud. 4, 114–123 (2009).
THUDM. Chatglm2-6b: An open bilingual chat LLM. https://github.com/THUDM/ChatGLM2-6B (2023).
Acknowledgements
This study was funded by the National Natural Science Foundation of China (82388101, U22A20311), National Key R&D Program (2022YFC2502800), Shanghai Municipal Education Commission(2023ZKZD18), Science and Technology Commission of Shanghai Municipality(23J41900200), National Clinical Key Specialty Construction Project(10000015Z155080000004). The funder played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript.We also want to thank all the first-year ophthalmologists participated in this study.
Author information
Authors and Affiliations
Contributions
Y.X., H.J., M.W., and J.F.: conceptualization, data curation, formal analysis, investigation, methodology, visualization, writing—original draft, writing—review & editing. X.X., W.H., J.C., and Z.Z.: data curation, investigation, writing—review &editing. X.Y., Y.S., J.W., and C.Z.: data curation, methodology. P.W., R.G., and X.Z. writing—review & editing. J.F.: conceptualization, investigation, methodology, supervision, writing—review & editing. X.S.: funding acquisition, conceptualization, data curation, formalanalysis, investigation, methodology, supervision, writing—original draft, writing— review& editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, Y., Jia, H., Wang, M. et al. Enhancing clinical documentation with voice processing and large language models: a study on the LAOS system. npj Digit. Med. 8, 798 (2025). https://doi.org/10.1038/s41746-025-02170-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02170-4