Abstract
AI-driven virtual cell models show the potential to transform the paradigm of life sciences research by integrating multimodal omics data (e.g., single-cell transcriptomics and proteomics) with advanced algorithms such as deep generative models and graph neural networks to enable high-precision predictions of drug responses, gene perturbations, and disease progression. These models enable high-precision predictions of drug responses, gene perturbations, and disease progression. This review outlines the technical pathways and validation mechanisms of virtual cells, emphasizing a closed-loop workflow from computational evaluation to experimental verification using CRISPR assays and organoid platforms. The applications of virtual cells in personalized drug screening and disease modeling are highlighted, showcasing their potential to reduce animal testing and optimize therapy. However, challenges in regulatory acceptance, data privacy, and model interpretability remain. Global policy and standardization trends are driving clinical translation, and future advancements will involve cross-disciplinary integration and greater standardization to enhance the impact of virtual cells in precision medicine and drug discovery.
Similar content being viewed by others
Introduction
AI-driven virtual cell models present a paradigm-shifting prospect for life-science research by supporting mechanistic, predictive simulation of functional responses1,2,3. A “Virtual Cell” is a computational model that simulates cellular functional states, signaling networks, and their dynamics under diverse perturbations1,4. Compared with conventional computational cell models, AI-driven virtual cells learn latent patterns from large-scale, multimodal biological data to construct predictive cellular state spaces5,6. Terminological fragmentation across the literature—spanning “virtual cell,” “digital cell,” and “digital twin”—drives our decision to foreground “virtual cell” as the field’s integrative label.
The emergence of this concept has been enabled by the rapid advancement of single-cell RNA sequencing (scRNA-seq), spatial transcriptomics (ST), proteomics, and other high-throughput omics, together with the introduction of novel Artificial Intelligence (AI) architectures such as deep generative models, graph neural networks, and physics-constrained neural networks7,8. By integrating multi-omics data from diverse sources, virtual cell models can reconstruct functional networks and signaling pathways across multiple scales, including the subcellular, single-cell, and cell-population levels1.
In preclinical drug research before Investigational New Drug (IND) submission, virtual cell models already demonstrate diverse applications: by simulating gene knockout, overexpression, or mutation, they predict functional roles in disease progression and, in combination with CRISPR validation, help screen potential drug targets9. In addition, these models can perform high-throughput in silico efficacy predictions across large candidate libraries to assess selectivity and potential adverse effects6,10. During lead optimization, simulations of absorption, distribution, and metabolism provide predictions that more closely approximate human physiological contexts11,12. Proof-of-concept, a multimodal single-cell virtual-cell model reconstructs mouse pancreatic ontogeny and validates NEUROD2-dependent ε-cell differentiation13; at scale, State—trained on >10^8 cells—yields superior predictions of drug-response perturbations across cell types14. Emerging practical utility of virtual cell models for mechanistic discovery and drug screening.
Virtual cell models also help elucidate tissue-specific differences in drug responses15. For example, integrating multi-omics data from hepatocytes and renal tubular epithelial cells enables prediction of tissue-specific toxicological responses under the same drug exposure6. This not only helps explain interindividual variability in adverse drug reactions but also offers a technical pathway to reduce animal experimentation and advance New Approach Methodologies (NAMs)6,16.
The advent of virtual cell models is shifting biomedicine from a wet-lab-dependent, “validation-driven” paradigm toward a data- and prediction-driven simulation–validation closed loop1,6. In sum, AI-driven virtual cell models are becoming a key bridge between molecular-mechanism research and preclinical drug evaluation, providing data-driven support throughout the pipeline from hypothesis generation to mechanistic verification1,5. Driven by an evidence-progression framework, the narrative advances from model construction to evaluation and validation, to virtual-cell use cases, translational barriers, and forward outlook. This translational driver integrates technical capability, testability, utility, and compliance pathways into a preclinical evidence chain (Fig. 1).
This figure systematically illustrates the entire lifecycle of AI-driven virtual cell models in preclinical research, encompassing technical pathways, validation mechanisms, application scenarios, as well as challenges and future directions in clinical translation. A Technical PathwaysMultimodal omics data (e.g., single-cell transcriptomics, spatial transcriptomics, proteomics) are integrated via advanced AI techniques—including deep generative models (e.g., UNAGI, scDiffusion), graph neural networks (e.g., DrugCell-GNN, GraphST), and physics-informed neural networks (e.g., BioPINN, VCBA)—to reconstruct cellular networks at subcellular, single-cell, and cell-population scales. Specialized platforms such as VCell (molecular/cellular level), PhysiCell (tissue level), and Chaste (organ level) enable multi-scale modeling, while foundation models like GeneFormer and State facilitate cross-level knowledge transfer from the gene/cell level to the organ and patient levels. B Validation MechanismsA two-tier validation framework is adopted: Computational Evaluation: Metrics such as distributional concordance, prediction accuracy, and uncertainty quantification are assessed using platforms like DepMap Portal and GEO.Experimental Validation: Predictions are verified via CRISPR assays, organoids (e.g., patient-derived intestinal organoids for toxicity testing), and organ-on-chip systems (e.g., hiPSC-CMs for cardiotoxicity prediction). For instance, a virtual cell model based on hiPSC-CMs accurately predicted doxorubicin-induced cardiotoxicity, consistent with clinical observations; VCBA-simulated hepatocyte mitochondrial toxicity matched in vitro experimental results. A closed-loop iteration of “computation→experiment→translation” ensures model robustness. C Application ScenariosVirtual cell models have diverse applications in preclinical research:Precision Drug Screening: Large-scale in silico screening of low-toxicity and high-efficacy compounds (e.g., million-compound virtual screening by the State model) and personalized drug response prediction (e.g., rare disease modeling integrated with digital twins).Mechanistic Inference: Elucidation of regulatory pathways (e.g., the role of NEUROD2 in pancreatic cell differentiation) and dynamic processes of drug-cell interactions.Reduction of Animal Experiments: Replacement of traditional animal models in toxicity and efficacy assessments (e.g., organoid-based antibody toxicity testing consistent with clinical reports). D Challenges and Future Directions in Clinical TranslationKey hurdles include regulatory ambiguity (e.g., FDA’s case-by-case IND review), data privacy (addressed by differential privacy and federated learning), intellectual property disputes, and algorithmic bias (mitigated by explainable AI tools like SHAP). Future developments will focus on standardization (e.g., SBML/CellML for model interoperability), interdisciplinary multi-scale integration (e.g., fusion of organoids, virtual cells, and digital twins), and international collaboration to advance clinical application. (Created in https://BioRender.com).
Technical Pathways
Precision virtual-cell construction sits at the forefront of the biology–AI nexus. Its core task is to integrate multimodal data, especially single-cell and spatial omics, thereby underpinning model development. Driven by complementary learners, deep generative models sculpt the cellular state space, graph neural networks capture intercellular interactions, and physics-informed neural networks embed biological laws, collectively shaping a virtual-cell model that delivers accurate predictions with defensible mechanistic interpretability.
Multimodal data integration
A primary challenge in building high-precision virtual cell models is integrating heterogeneous biological modalities, including scRNA-seq, ST, epigenomics, proteomics, and metabolomics1,2. Deep generative models and graph neural networks show strong potential for simulating single-cell states and predicting drug responses17, while physics-informed neural networks (PINNs) can enhance interpretability and biological plausibility16.
scRNA-seq resolves gene expression at single-cell resolution but loses spatial context18,19; ST preserves spatial structure yet faces limits in resolution, throughput, and gene coverage20,21. High-throughput imaging–driven advances in spatial transcriptomics now enable the simultaneous detection of ~20,000 genes, producing near-whole-transcriptome spatial maps and progressively mitigating the historical gene-coverage bottleneck of imaging-based ST22,23.
To bridge these gaps, AI-based cross-modal fusion methods have been proposed7,8,24,25,26,27. For instance, SpatialScope uses deep generative modeling to project high-dimensional scRNA-seq data into ST coordinates, enhancing resolution and imputing missing genes7. VAE (variational autoencoder)-based frameworks such as scVI and totalVI integrate multi-batch single-cell data and fuse transcriptome with proteome for joint denoising and analysis8,28. Domain adaptation reduces cross-platform bias: scAdapt employs adversarial training to align distributions between ST and scRNA-seq, improving spatial deconvolution accuracy7,24; conditional normalization and attention help detect rare cell types and low-abundance genes25,29. On high-dimensional integration, MultiVI maps transcriptomic and epigenomic modalities into a shared latent space to improve cell-type classification, trajectory inference, and lineage analysis26; GraphST couples graph structures with expression and spatial proximity to drive cell-type identification and spatial relationship reconstruction27.
Recent advances have led to a new wave of methods for integrating multi-omics data. For example, the GLUE (graph-linked unified embedding) model incorporates prior regulatory networks to project heterogeneous single-cell omics into a shared embedding space, achieving strong cross-species integration across millions of cells30. Driven by optimal-transport–based coupling across modalities, the moscot framework collectively shapes single-cell multi-omics into coherent spatiotemporal developmental trajectories in the mouse, with its model-predicted key regulators validated experimentally13. Moreover, a vision has been articulated for multimodal foundation models: that is, pretrained unified models that integrate genomics, transcriptomics, epigenomics, proteomics, metabolomics, and spatial omics to comprehensively characterize cellular states31. Such models would enable context-specific transfer learning for panoramic single-cell atlasing.
These strategies enable virtual cells to integrate multilayer omics more comprehensively, forming richer, more accurate inputs for downstream drug response prediction, perturbation simulation, and mechanism inference1,2.
Large public databases provide critical training data32,33,34: TCGA (The Cancer Genome Atlas) for tumor genomics/transcriptomics35,36; the Human Protein Atlas (HPA) for tissue/cell-level protein localization37,38,39; and GEO (Gene Expression Omnibus) for diverse omics datasets40,41,42. All harbor inherent biases43,44,45,46. Corrective techniques help: ComBat, Harmony, and related methods attenuate batch effects47,48,49; domain adaptation (e.g., scAdapt) aligns distributions within models50. Rigorous normalization and correction bolster generalization and reduce misleading predictions driven by data bias46,51,52,53,54 (see Table 1).
Omics foundation models and cross-hierarchical transfer
Foundation models for virtual cells trained on tens of millions of single cells deliver strong performance in cell-type annotation and gene-function inference (GeneFormer55; scFoundation56). Meanwhile, models that target complex genetic perturbations: GEARS57 and the Arc Institute’s State model58, exhibit measurable generalization across cell types. Driven by cross-scale knowledge transfer, the next critical task is to migrate capabilities learned at the gene/cell level to the patient level, where explicit modeling of tissue microenvironments, pharmacokinetics, and other systems constraints becomes indispensable. To this end, several studies use transfer learning to achieve cross-scale prediction: CODE-AE extends drug-response prediction from cell lines to patient tumor transcriptomes59, and scDEAL integrates population-level and single-cell data to improve individualized drug-sensitivity prediction60. CSG2A, by leveraging condition-specific gene and gene-to-gene attention, pretrains on LINCS L1000 and fine-tunes on GDSC to transfer perturbation knowledge from the gene level to drug-response prediction in cells and patients, enhancing the interpretability of virtual-cell models61. Together, these advances suggest that elevating gene-level pretraining to organ- and patient-level virtual models is feasible, provided that additional determinants—tissue context, ADME (absorption, distribution, metabolism, and excretion) processes, and related factors—are rigorously incorporated.
Deep generative models
Deep generative models are central to constructing predictive and generative cellular state spaces1,17. Beyond fitting high-dimensional data distributions, they can generate synthetic expression profiles in latent space for virtual experiments under drug perturbations or genetic edits5,17. Recent work has simulated disease trajectories and drug responses; for example, the graph-structured VAE-GAN model UNAGI captured single-cell dynamics in idiopathic pulmonary fibrosis and predicted potential anti-fibrotic activity of nifedipine, validated in patient lung tissues and proteomics62. Beyond classic frameworks such as VAEs, flow matching and diffusion models now define a new paradigm in generative modeling. Flow matching directly learns continuous-time transformations between data distributions to enable efficient generation of high-dimensional, structured data63; diffusion models formalize noise injection and denoising inversion with stochastic differential equations, progressively approaching the target distribution, and have achieved notable advances in image generation and single-cell data simulation64。)Driven by diffusion dynamics coupled with a pretrained foundation model, scDiffusion generates single-cell transcriptomes with high fidelity and strong diversity65,66. Pressure for cross–cell-type generalization drives model design: integrating perturbation data across lineages and enforcing priors from biological networks, mechanistically improves transfer to unseen perturbations. Against this backdrop, current deep learning approaches, in aggregate, do not surpass simple linear baselines by a large margin67, whereas a knowledge-graph-guided framework (GEARS) reports ~ 40% accuracy improvement for predicting transcriptional responses to multi-gene perturbations57. Mechanism-anchored validation remains sparse; only a few virtual cell predictions have been confirmed via targeted interventions that verify model-nominated key regulators13. Establishing standardized experimental validation pipelines should therefore be elevated as a field-level priority. Thus, generative models provide a powerful in silico experimental platform for hypothesis testing and therapeutic exploration1 (see Table 2).
Graph neural networks
Cells influence each other through signaling pathways, cell–cell communication networks, and spatial proximity2,29. Biologically faithful virtual cells must model such topology and information flow1. Graph neural networks (GNNs) naturally suit cell-relationship modeling in single-cell omics29. Treating each cell as a node, GNNs iteratively aggregate neighborhood information to capture contextual dependencies68,69,70. scGNN embeds scRNA-seq into a graph and applies multi-layer GCNs for feature aggregation, outperforming t-SNE (t-distributed Stochastic Neighbor Embedding)/UMAP (Uniform Manifold Approximation and Projection) for cell-type identification, imputation, and trajectory inference29.
For multimodal fusion, PINNACLE integrates scRNA-seq with protein–protein interaction networks via graph attention to better detect rare subpopulations and regulatory patterns71,72,73, aiding immune-cell subtyping and resistance mechanisms6. GNNs also extend to drug response prediction74,75,76. DrugCell-GNN unifies cellular transcriptomes, drug molecular graphs, and known target networks to predict sensitivity and synergy, improving accuracy in anticancer screening1,10. With the rise of ST, SpaGCN couples spatial adjacency with expression matrices to resolve cell types and tissue organization, excelling in tumor microenvironment heterogeneity analyses7.
Overall, GNNs endow virtual cells with multiscale modeling, from single-cell regulatory patterns to population-level coordination1,2,77. Integrating ST and live-cell imaging will further sharpen tissue-microenvironment realism and empower disease-mechanism and drug-development studies6,7.
Physics-informed neural networks
Although deep learning models demonstrate powerful data fitting and generation capabilities in virtual cell simulations, most are “black-box” models that lack explicit expression and constraints of known biological laws. The core idea of Physics-Informed Neural Networks is to incorporate known biophysical laws, dynamic equations, or constraints during the model training process to ensure that the predicted results are biologically plausible16,78,79,80,81,82. This approach combines theoretical models with data-driven deep networks, balancing flexibility and interpretability16.
In drug metabolism and toxicology simulations, the Virtual Cell-Based Assay (VCBA) platform combines organelle-level dynamic models (e.g., mitochondrial membrane potential changes, reactive oxygen species production) with dose-time integral models to simulate dynamic responses of hepatocytes and cardiomyocytes under drug exposure9,11. These models have demonstrated high consistency with some in vitro experimental data and clinical adverse event reports in predicting drug-induced liver injury (DILI) and cardiotoxicity9,12. Hybrid constraint methods are also a current trend83,84,85. For example, integrating the Flux Balance Analysis (FBA) model of metabolic pathways with neural networks can improve prediction accuracy for environmental disturbance responses while maintaining metabolic flux conservation84. Similarly, embedding the classic Hodgkin-Huxley equation of ion channel electrophysiology as a prior in cardiomyocyte models can more accurately predict drug-induced action potential changes85,86.
However, constructing PINNs still presents challenges1,6,16,87,88. Predictive improvement in PINNs hinges on the completion of biophysical knowledge and the identification of kinetic parameters, rather than hardware scaling; incomplete laws and parameter gaps impose a hard ceiling. Mathematical descriptions of biological systems are often incomplete, and some dynamic parameters are difficult to obtain16. The introduction of physical constraints increases the complexity of model training and sometimes conflicts with data fitting objectives, resulting in convergence difficulties1. Furthermore, in a multi-scale, multi-modal data environment, selecting and weighting constraints appropriately is a key challenge in model design6,16. With the improvement of computational hardware, PINNs are expected to become an important component of virtual cell models in the future1,6,16.
The various technical approaches mentioned above each have their own advantages, and in virtual cell modeling practices, they are not mutually exclusive but can work synergistically1. Currently, the integration of multimodal data ensures comprehensive and high-quality input for the models89,90. Next, deep generative models are used to establish baseline predictive capabilities for cell state spaces, such as generating cell transcriptomes under unseen perturbation conditions17,91. Furthermore, a graph neural network module is introduced, and previous research has explored combining generative models, graph networks, and physical constraints to build more powerful integrated models62,92,93. Despite the limitations of different methods, the comprehensive application of multiple techniques is expected to construct a more comprehensive and accurate virtual cell model1.
Platforms and toolboxes at a glance
A compact landscape of virtual-cell and cell-scale modeling platforms now anchors the fourth technical route, spanning molecular networks, multicellular systems, and organ-level physiology (see Table 1). These platforms can be broadly grouped by their core modeling targets and methodological foundations, each serving distinct research questions.
Driven by mechanistic representations of intracellular processes, several toolchains target gene regulation, signal transduction, and related pathways. VCell (Virtual Cell) provides a unified cell-biology environment that supports spatially resolved and well-mixed (nonspatial), deterministic and stochastic modeling, and it natively integrates image-analysis workflows94,95. For quantitative kinetics, COPASI (Complex Pathway Simulator) specializes in ODE/SDE (stochastic differential equation)-based biochemical network modeling and parameter estimation96,97,98,99. To address combinatorial explosion in rule-based signaling models, BioNetGen (BNG)100,101,102 and PySB103 enable compact specification of reaction rules and efficient state-space handling. Logic-network frameworks such as CellNOpt104,105,106 and the Cell Collective107 infer apoptosis signaling circuitry from perturbation datasets. Application-oriented platforms like VCBA focus on environmental toxicology and biokinetic simulation9,108,109.
At the multicellular and tissue scale, agent-based modeling (ABM) is widely used to capture population behaviors and tissue repair dynamics. PhysiCell emphasizes 3D, tissue-scale ABM and is particularly suited for simulating the tumor microenvironment110,111,112,113. Guided by cell–cell mechanics and morphodynamics, CompuCell3D114,115,116,117 and Morpheus118,119 implement Cellular Potts/GGH frameworks that couple cell mechanics with reaction–diffusion processes.
Specialized engines and emerging AI tools further extend virtual-cell modeling. At the subcellular scale, Smoldyn120,121, MCell102, and ReaDDy122,123 perform particle-based stochastic reaction–diffusion, enabling molecular-level transport and interaction studies. For organ-system modeling, Chaste124, OpenCOR/CellML (Cell Markup Language)125,126, and OpenSim/Physiome125,127 support multiscale cardiac electrophysiology and biomechanics. Recently, AI-driven platforms have gained traction: DeepCell128 and CellPose129,130,131 deliver high-accuracy cell image segmentation and analysis, while NVIDIA Modulus132provides a framework for physics-constrained neural networks—directly aligning with the three advanced AI routes discussed above (see Table 3) (Fig. 2).
A Multimodal data and representative resources include genomics, proteomics, and transcriptomics data types with mainstream databases and platforms, with a Multimodal Foundation Model serving as the hub that connects data to methods; B The technical routes comprise three paths:Deep generative models, Graph neural networks, and Physics-informed neural networks,where Deep generative models encompass VAE-GAN, Flow Matching, and Diffusion Models and connect to an in vivo/in vitro-referenced in silico experimental platform, Graph neural networks center on joint graph construction from single-cell and spatial transcriptomics and use SpaGCN as a representative implementation, and Physics-informed neural networks target Training Convergence and Accuracy of the Solution and present a PINN for the 1D Heat Equation to strengthen interpretability and physical consistency; C Application scenarios cover lineage analysis and cell-type annotation with constraint information to improve labeling robustness and include drug response prediction and trajectory inference with Targeting and model Refinement cues that form a feedback loop from applications to methods and data, while Alternative Platforms and Workbenches at the lower rim indicate where diverse tasks are executed and validated computationally. (Created in https://BioRender.com).
Validation Mechanisms
Collective shaping by a closed-loop validation architecture drives virtual cell models from methodological novelty to actionable biology. At its core is a systematized pipeline coupling computational assessment (fit, stability, uncertainty) with experimental corroboration, iterated in cycles. This mechanism foregrounds interpretability and accelerates translation while accumulating layered evidence for model robustness and portability across contexts.
Computational Evaluation
The performance validation of virtual cell models can be divided into two major aspects: computational evaluation and experimental validation9,133. Computational evaluation primarily focuses on the model’s fitting to existing data and its ability to predict unknown scenarios133. Common quantitative indicators are as follows: (1) Reconstruction error and distribution consistency: This includes measuring the distance between the generated data and the training data in high-dimensional space, statistical distribution differences, and comparing their clustering structures through dimensionality reduction methods such as t-SNE5. (2) Prediction accuracy and stability: For extrapolated scenarios, the model’s prediction is evaluated against real-world results134,135,136. (3) Model complexity and generalization ability: This involves statistical model parameters, computational time, and evaluating the model’s ability to adapt to new data through methods like cross-validation1. Uncertainty quantification enhances the interpretability and robustness of conclusions. For example, Monte Carlo dropout can approximate the posterior predictive distribution to compute confidence intervals, and model ensembling can evaluate cross-model output consistency, thereby quantifying the reliability of virtual-cell model predictions137,138. Driven by the destructive nature of single-cell assays—which precludes repeated measurements on the same cell139—evaluation should pivot to the population-distribution level by comparing the statistical distributions of perturbed simulated cell populations with those observed experimentally93. Accordingly, distributional distances—such as the Wasserstein distance, Kullback–Leibler divergence, and maximum mean discrepancy—serve as summary statistics to quantify agreement between model predictions and experimental data at the overall level. It is important to note that for high-dimensional biological data, “prediction accuracy” itself needs to be carefully defined and interpreted140,141,142. Therefore, a multi-layered evaluation index system should be designed based on specific tasks to comprehensively quantify the performance of virtual cell models143,144,145.
As virtual cell models become increasingly complex, the computational tools used to assess them are also evolving1,6,146. Through multi-angle, multi-index computational evaluation, models with superior performance and robust stability can be initially screened for further experimental validation146,147,148.
Experimental verification
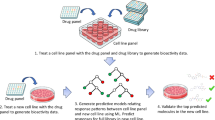
The models, after computational evaluation, need to be validated through biological experiments to confirm whether their predictions reflect real biological phenomena9,149,150,151,152,153. In terms of gene function prediction, CRISPR/Cas9 and other gene-editing technologies are used to verify the key targets predicted by the models through in vitro or in vivo validation154,155,156. For example, this process is commonly used in the discovery of new targets to help confirm whether the candidate targets selected by the model are worthy of further development9.
New in vitro models comprise organoids and organ-on-chip systems. Organoids are three-dimensional cell aggregates grown in vitro that partially recapitulate the structure and function of native tissues; organ-on-chip platforms culture cells within microfluidic devices using microengineering techniques to emulate organ-level microenvironments and functions. These innovative platforms offer physiologically relevant testbeds to validate predictions generated by virtual-cell models79,157. Furthermore, in drug response prediction, the virtual cell model’s predictions are validated through in vitro cell experiments or organoid models6,28,158. For instance, if the model suggests that a particular candidate compound may induce hepatocyte steatosis or other toxic reactions, its toxicological indicators can be verified in mouse models158,159,160. For the prediction of drug absorption, distribution, metabolism, and excretion properties, in vivo pharmacokinetic experiments can be conducted to validate the model’s inference about absorption pathways6,28. With the development of novel in vitro models, experimental systems that more closely mimic the in vivo environment are also being used to validate the predictions of virtual cells6,28,161. These 3D models highly replicate the microenvironment of native tissues at multiple levels, including cellular composition, tissue structure, and functional performance161. These models are expected to be deeply integrated with virtual cells to accelerate the transition of drugs from virtual screening to clinical trials6,161(Fig. 3).
A Computational evaluation: Quantitative indicators of distribution consistency and predictive ability. The figure shows commonly used quantitative indicators for computational evaluation and the computational tools to assess them. Common quantitative indicators include reconstruction error and distribution consistency, prediction accuracy and stability, model complexity and generalization ability. The computational tools to evaluate them include those based on cell phenotype space coverage, those evaluated using Spearman’s rank correlation coefficient, and those based on the “adversarial validation” strategy. Through multi-angle and multi-index computational evaluation, models with better performance and good robustness can be initially selected for the next stage of experimental verification. B Experimental verification: Process from CRISPR target editing to drug response testing. The figure shows three experimental verification methods, including biological experimental verification, drug response prediction, and in vivo environmental experimental systems with the development of new in vitro models. Biological experiments are used in gene function prediction by knocking out or overexpressing a certain gene for observation to discover new targets; drug verification validates the predictions of virtual cell models through in vitro cell experiments or organoid models; in vivo environmental experimental systems include organoids, organ-on-a-chip, etc. With the standardization and wide application of verification models, these models can be deeply integrated with virtual cells to accelerate the transformation process of drugs from virtual screening to clinical trials. (Created in https://biogdp.com).
Virtual-cell–experiment convergence is validating drug-effect predictions. Several concrete studies show that predictions from virtual cell models closely match results from wet-lab experiments. Driven by integrated modules for drug exposure, cardiomyocyte-drug interaction, and system-level response, Wang and colleagues built an in-vitro–in-vivo translation platform based on human iPSC (induced pluripotent stem cell)-derived cardiomyocytes (hiPSC-CMs). Its simulations predicted the incidence of cardiotoxicity for the anthracycline doxorubicin and the anti-HER2 antibody trastuzumab with high concordance to clinical observations162. Feasibility is demonstrated: hiPSC-CM–based virtual cells enable animal-free, quantitative prediction of anticancer-drug-induced cardiac dysfunction. Similarly, the Virtual Cell-Based Assay (VCBA) has been used to simulate intracellular and organellar pharmacokinetics. Driven by mechanistic transport and binding representations, Worth et al. used VCBA to model the distribution of selected compounds in HepaRG hepatocytes and cardiomyocytes and their effects on mitochondrial membrane potential. The simulations accurately recapitulated in-vitro changes in toxicity readouts for the uncoupler FCCP, caffeine, and amiodarone9. VCBA thus maps intracellular and mitochondrial concentrations to membrane potential shifts and cytotoxicity, reinforcing the agreement between virtual predictions and in vitro data.
“Next-generation in-vitro models” include organoids and organs-on-chips: organoids are three-dimensional cell aggregates cultured ex vivo that partially reproduce tissue architecture and function, whereas organs-on-chips culture cells in microfluidic devices to emulate organ microenvironments and functions79,157. These systems provide physiologically relevant platforms for validating virtual-cell predictions. Driven by patient-specific epithelium–immune crosstalk, patient-derived intestinal organoids co-cultured with immune cells have been used to assess on- and off-tumor toxicities of T-cell bispecific antibodies. Harter et al. showed that an EpCAM-targeted T-cell-engaging bispecific antibody induced apoptosis in healthy intestinal organoids, aligning with toxicities reported clinically163. Result: faithful reproduction of target-mediated toxicity in complex microenvironments, providing a reliable experimental anchor for validating virtual predictions. Driven by a policy shift toward animal reduction, the FDA (Food and Drug Administration) has recently signaled broader use of organs-on-chips, virtual models, and organoids in preclinical safety assessment164.
Collectively, these exemplars substantiate the effectiveness of virtual-cell models for predicting drug toxicity and ADME behavior. Driven by cross-validation against hiPSC-derived cardiomyocytes, organoids, and animal studies, virtual-cell platforms are accelerating drug development and enabling human-physiology-aligned alternatives to traditional animal testing. Regulatory agencies, including the FDA, are encouraging adoption of these integrated computational–biological approaches, steering screening and safety evaluation toward human relevance.
Closed-loop integration of computational evaluation and experimental validation
Closed-loop architecture—comprising a computational inner loop, an experimental middle loop, and a translational outer loop—links methodological assessment with application-level verification; entry, decision, and exit gates at key nodes operationalize bottom-up continuous improvement. In the computational inner loop, strict data partitioning, batch harmonization, and pre-analysis quality control are prerequisites. We quantify distributional concordance under perturbation scenarios and characterize predictive uncertainty, with primary criteria being a pre-registered concordance threshold and an upper bound on the confidence interval width. If criteria are not met, the workflow reverts to data cleaning, structural-prior specification, and regularization tuning165. Driven by near-physiological systems (e.g., organoids and organ-on-chip platforms), the experimental middle loop subjects model claims to targeted tests under pre-specified functional endpoints and statistical power. When effect direction or magnitude diverges from computational expectations—or power is insufficient to reject the null—the loop feeds back to joint revisions of modeling assumptions, feature selection, and experimental-protocol parameters166. In the translational outer loop, cross-platform replication and prospective datasets constitute the primary settings. Deployment readiness is adjudicated by integrated evidence on generalizability, stability, and interpretability, supplemented by safety review and failure-mode analysis; if out-of-domain performance departs from predefined bands, feedback triggers coordinated optimization of feature engineering and experimental design167. Tri-loop coupling secures distribution-level statistical reproducibility with bounded uncertainty while cross-validating functional evidence and translational feasibility, yielding a unified and transparent operating framework for comparable evaluation and reproducible validation across applications.
Application Scenarios
Post-validation priority: establishing the model’s real-world and translational value.Applications of virtual cells are steadily expanding to drug screening, mechanistic inference, digital twins, and multi-platform interoperability, opening new avenues for precision medicine.
Virtual-cell-enabled precise screening and mechanistic inference
The virtual cell model can assist in advancing and refining drug screening processes1,5,6,9,10. On one hand, during the target discovery phase, virtual cells can simulate gene knockout or overexpression, predicting downstream molecular networks and phenotypic changes to infer potential therapeutic targets and their mechanisms of action1,9. Researchers can prioritize the validation of targets predicted by the model to have significant effects, improving the hit rate of experimental screening168. On the other hand, in lead compound screening, virtual cell models can be used for in silico screening of large numbers of candidate compounds, predicting their ability to correct pathological cell states and assessing their risk of toxic side effects6,10. Regarding mechanism inference, virtual cell models can provide in-depth analysis of the dynamic processes of drug-cell interactions5,6,10. Moreover, virtual cells can also be used for hypothesis testing. Researchers can introduce a new regulatory factor or pathway hypothesis into the model to test whether it can reproduce the observed phenomena5. If the model validation supports the hypothesis, targeted experimental validation can then be designed at the experimental level169,170,171. This approach is expected to improve the efficiency of mechanistic research, quickly identifying key factors and pathways with the support of virtual cells169,170,171.
It is important to emphasize that the use of virtual cells should always be closely integrated with biological experiments172,173,174. In the short term, virtual results serve more as supporting evidence175,176, thus hypotheses generated through model predictions must be experimentally validated, with feedback used to refine the model and continually improve its consistency with reality1. This closed-loop process will drive the true precision of drug screening and mechanism analysis, enabling the efficient translation of vast amounts of data into knowledge discovery177,178. An integrative evaluation workflow, distilled from a systematic synthesis of prevailing standards and practices, enables comparable assessment and reproducible validation of virtual cell models across application contexts. We benchmark this workflow against existing methodological frameworks, specifying shared steps—data standardization, identification of bias sources, distribution-level concordance testing, and uncertainty quantification—and modules unique to virtual-cell settings—distributional distance evaluation for perturbation simulations, cross-modal alignment, and mapping of functional endpoints between organoids and organ-on-chip platforms. Driven by cell-state prediction and perturbation-response inference as core tasks, the workflow applies to studies with traceable data provenance and cross-batch calibration of experimental systems. Limitations: cross-species generalization and extrapolation to ultra-rare cell populations warrant particular caution.
Synergy between digital twins and virtual cells
The introduction of the concept of “Digital Twin” has provided new opportunities for the application of virtual cell models6. A digital twin typically refers to a high-fidelity virtual replica of a specific entity, which can receive real-time data from the physical entity and perform simulations179,180,181. By integrating virtual cell models into the digital twin framework, a multi-scale, comprehensive simulation from the molecular and cellular level to the whole organism can be achieved1,6,10,161,182,183. In rare disease research, by combining patient-specific stem cell-derived virtual models with patient phenotype data, the effects of personalized drug interventions can be simulated161,182. This strategy allows for large-scale virtual screening of efficient, low-toxicity candidate compounds before clinical trials10 and has shown potential to partially replace animal experiments in target discovery and other processes6,9.
The combination of digital twins and virtual cells also provides a new approach to disease prediction and monitoring6,181. By continuously inputting biomarker data from patients, virtual cell models can simulate the cellular evolution of disease progression, offering doctors predictions regarding the disease’s course and outcome6. For instance, in cancer treatment, if the virtual model predicts changes related to drug resistance signals, it can provide an early warning and suggest treatment adjustments184,185,186,187,188. These applications rely on high-quality individual data and the model’s ability to sensitively capture individual differences187,189,190,191. With advancements in data collection and modeling techniques, the integration of digital twins and virtual cells will become more seamless, playing a more active role in clinical decision support187,188,192.
Boundary setting and complementarity
Despite the powerful capabilities of virtual cell models, they cannot fully replace traditional methods in all scenarios. Therefore, it is essential to define their application boundaries and leverage their complementary role6. In the context of complex multicellular behaviors, virtual cells currently focus on molecular and intracellular processes. For phenomena involving tissue morphological changes, traditional multicellular modeling or experimental approaches remain indispensable1. Regarding model interpretability, physical models and rule-based models, which are directly grounded in known human laws, are generally more readily accepted by researchers. In contrast, although deep learning-driven virtual cells provide accurate predictions, they may struggle to offer deductive mechanistic explanations16. Thus, in studies requiring a clear mechanistic understanding, virtual cells serve as an exploratory tool, while the final mechanistic elucidation should be combined with traditional experiments and analytical models1. Additionally, virtual cells complement other emerging technologies5,6,193. For instance, when combined with single-cell lineage tracing technology, they can provide developmental pathways, which are difficult for models to simulate, serving as ground truth to calibrate model parameters5. Researchers should fully recognize the strengths and limitations of virtual cells and integrate them with other practice methods: capitalizing on their speed and scalability while using traditional methods to ensure reliable results and clear explanations194,195,196,197. Only in this way can virtual cell models find their optimal positioning and achieve maximum benefits in biomedical research and drug development198,199,200.
Virtual cells and cell-level modeling platforms: status and comparative analysis
Mechanism-driven alignment with the target scenario is the primary determinant of successful application. The selection of a modeling platform is driven by (i) the biological endpoints of the question and (ii) the complexity of implementing the pertinent molecular mechanisms. When the aim is Section 4.1’s precision screening and mechanistic deduction, intracellular network platforms are indicated—e.g., COPASI96,97 for simulating drug impacts on pathway behavior, or CellNOpt104,105 for data-driven reconstruction of signaling circuitry under perturbations. For Section 4.2’s digital-twin use cases involving multi-cellular coordination, tissue-level simulators—PhysiCell110,111 or CompuCell3D114,115—are preferable to capture tumor expansion and integrated tissue responses to drugs. Readers may consult Section 2.6’s computational-speed overview and Table 1’s comprehensive comparison (Fig. 4) to select the most suitable platform.
This figure elucidates the diverse applications of AI-driven virtual cell models in preclinical research, encompassing three dimensions: precision drug screening & mechanism deduction, synergistic integration with digital twins, and boundary definition, complementary roles, along with platform classification. A Precision Drug Screening & Mechanism DeductionIt demonstrates the core value of virtual cells in drug development, spanning drug target discovery, optimization of candidate compounds, to MOA deduction. The closed-loop integration of biological experiments and model predictions enables precise drug screening and mechanistic interpretation. B Synergistic Integration with Digital TwinsThis section illustrates their complementary value in multi-scale simulation: virtual cells focus on microscopic mechanisms at the cellular or cell population level (e.g., sensitive capture of individual differences in tumor drug resistance signals), while digital twins encompass the overall physiological-pathological state of an individual. Their synergy achieves multi-scale integration from molecule-cell-tissue to individual, exemplified by personalized drug efficacy simulation in rare disease research. C Boundary Definition, Complementary Roles, and Cell-Level Platform Classification.Boundary and Complementarity: Despite their capabilities, virtual cell models cannot fully replace traditional methods across all scenarios. They excel in molecular and intracellular processes but rely on traditional experiments or analytical models to explain complex tissue morphogenesis or provide deductive mechanistic. Meanwhile, they complement emerging technologies such as single-cell lineage tracing.Platform Classification: Cell modeling platforms are categorized by scale: the molecular and intracellular network level (e.g., COPASI for biochemical network dynamics), the multicellular-tissue level (e.g., PhysiCell for tissue-scale agent-based modeling, CompuCell3D for Cellular Potts-based simulation), and the organ-system level (e.g., OpenSim for biomechanical modeling). Future development trends lean toward cross-scale and multi-methodological integration. (Created in https://BioRender.com).
Clinical translation, ethics, and compliance
Driven by collective socio-technical shaping, the safe translation of virtual cell models from theoretical constructs to clinical use has become the decisive inflection point. The trajectory is conditioned not only by technical capability but by regulatory architectures, ethical safeguards, legal accountability, and trust-building mechanisms. We outline how compliance regimes, data privacy security, intellectual property ownership and stewardship, and fairness auditing collectively facilitate clinically acceptable deployment, paving the way for the field’s next phase.
Regulatory Trends
With the rapid development of non-animal testing methods, AI-driven virtual cell models have gradually entered the discussion within regulatory agencies6,16,201,202. In 2022, the U.S. Congress passed the FDA Modernization Act 2.0, which explicitly stated that animal testing data would no longer be a mandatory requirement for new drug IND submissions. It allowed for the use of humanized cell models, organoids, organ-on-a-chip systems, and artificial intelligence/machine learning (AI/ML) models as alternatives6,203,204. This legislation opened a potential path for the regulatory and compliant application of virtual cells, particularly in early-stage toxicology and pharmacology evaluations6,9,205. In 2025, the U.S. FDA released its first draft guidance on the use of AI models in drug development, titled “Considerations for the Use of Artificial Intelligence to Support Regulatory Decision-Making for Drugs and Biologics,” proposing a risk-based framework for model credibility assessment206. In Europe, the European Chemicals Agency (ECHA) and the EU REACH (Registration, Evaluation, Authorization and Restriction of Chemicals) regulations have also strengthened the adoption of non-animal testing methods in recent years, with some toxicology evaluation frameworks incorporating computational modeling and bioinformatics prediction tools207,208. Overall, global regulatory trends are cautiously supportive of AI-driven alternative methods: on one hand, they recognize their potential in accelerating drug development and reducing animal use, while on the other hand, they ensure their reliability and transparency through legislation and guidelines when used in decision-making6,16,203,204. For virtual cell model developers, understanding and adhering to these policy guidelines and considering regulatory requirements during the model design phase, will help increase the likelihood of success in subsequent clinical translation submissions209,210.
Regulatory and Implementation Challenges
Despite the gradual relaxation of the policy environment, virtual cell models still face significant challenges in fully replacing traditional in vivo and in vitro experiments under the current regulatory system6,12,182. The U.S. FDA still requires the submission of adequate in vitro and/or in vivo validation data during the IND review process, with AI models currently considered as supplementary evidence rather than the sole basis6. Differences in data sources, model architecture, parameter settings, and validation processes among research teams, along with the lack of a unified third-party benchmark testing platform, limit the comparability and reproducibility of model results16,211. At present, regulatory agencies have not established clear review guidelines for AI-based biological models, and projects are often subject to case-by-case review, making it difficult for applicants to predict the required level of validation16,211,212. For example, there is no unified standard regarding the level of predictive accuracy required to support an IND application, and regulatory demands may vary significantly across different cases6. Moreover, regulatory agencies’ personnel must also address challenges213,214,215. Reviewing a complex virtual cell model typically requires interdisciplinary experts in computational biology, machine learning, and disease biology. However, many regulatory agencies currently lack sufficient human resources in this area, potentially leading to delays or uncertainty in the review process16,211. As a result, agencies like the FDA are improving AI review capabilities through internal training and external consultants, but this cognitive gap remains a significant barrier to model translation in the short term216,217,218,219,220,221,222,223.
Another challenge arises from data sharing and privacy issues224,225,226,227,228,229,230,231. High-quality virtual cell models typically require the integration of patient-derived data for training, which often involves sensitive personal health information226,232,233,234. When submitting regulatory reviews, applicants need to provide sufficient information to demonstrate the reliability of the model while safeguarding patient privacy. This has led to the development of innovative solutions, such as secure multi-party computation and federated learning, which enable multiple institutions to collaborate on model validation without sharing raw data232,235,236. However, these technologies also add complexity and uncertainty to the review process237,238,239. Furthermore, legal issues such as model intellectual property ownership and compliance with training data regulations need to be clarified in submission materials, as failure to do so may trigger compliance investigations and delay the review process6. Finally, the application of AI models in regulatory decision-making lacks successful precedents240. To date, there are few cases where new drugs have been approved based solely on AI model results, with AI still predominantly used as an auxiliary tool241,242. At the regulatory implementation level, virtual cell models continue to face challenges related to model standards, review capabilities, data compliance, and successful demonstrations, requiring joint efforts from industry and regulators to resolve2,182,243.
Cross-scale coupling still constrains the transition from single-cell models to tissue-level applications.At the single-cell level, virtual models must be embedded within higher-order physiological contexts, encompassing cell–cell interactions, three-dimensional tissue architecture, and whole-body dynamics244. Driven by multiscale integration, existing frameworks (OpenSim, Physiome) can host cell-level models at organ and organism scales127,245, yet they prioritize macroscopic processes and remain limited in their explicit representation of signaling pathways and gene-regulatory mechanisms. Consequently, after in vitro verification, virtual cells require intermediate testing and parameter calibration in organoids and animal models to progressively approximate real-world patient contexts246,247. Clinical translation hinges on parallel progress in cross-scale technical concordance and regulatory–ethical alignment, both of which should be treated as coequal bottlenecks for virtual cells.
Data privacy and security compliance
Virtual cell models are constructed and trained on large-scale biomedical datasets that often contain sensitive patient information7,8. If mishandled, such data can lead to breaches of personal privacy and raise ethical concerns6,226,233,234,248. Many countries and regions have enacted laws governing the use of health data; for example, China’s Personal Information Protection Law (PIPL) requires health data to be stored locally and mandates obtaining the data subject’s explicit consent248, while the United States’ Health Insurance Portability and Accountability Act (HIPAA) requires encryption and de-identification of protected health information (PHI)6,248. European context: mandatory GDPR (generative adversarial network) -compliant protection of patient data privacy249. In parallel, stakeholders should closely follow the forthcoming European Health Data Space (EHDS) initiative, which aims to enable secure sharing and standardized reuse of research data within a defined regulatory framework250. These regulations mean that developers of virtual cell models must ensure legal compliance when collecting and processing data, and that cross-border data transfers may give rise to legal conflicts that affect the use of models in international multicenter studies248,251. Beyond regulatory compliance, technical measures are also needed to safeguard data security and privacy252,253,254,255,256,257,258,259. Evidence indicates that deep generative models can “memorize” unique features of training samples and, in extreme cases, reproduce portions of the original training data in their outputs16. To mitigate such risks, developers should deploy privacy-preserving mechanisms when releasing and sharing models260,261,262,263,264,265,266 For example, differential privacy techniques can inject calibrated noise into model parameters to obscure individual-level features and thereby reduce potential privacy leakage with minimal impact on model performance16,267,268,269. Likewise, federated learning frameworks allow models to be trained on local institutional data while only sharing model weight updates, thereby architecturally avoiding the risk of centralized data exposure232,270,271. A combined legal and technical approach is therefore required to realize the value of virtual cell models while ensuring data security272,273. Cybersecurity is another essential consideration. If virtual cell models are provided to healthcare institutions as cloud services, the databases and computing infrastructure on which they depend may become targets for attack274,275,276. Consequently, model deployment should be accompanied by strict access controls, vulnerability scanning, and incident-response mechanisms, and model outputs should be subject to anomaly monitoring so that any aberrant behavior triggers timely alerts and intervention277,278,279. These measures will help ensure that virtual cell models, as digital medical products, meet security standards at least as stringent as those applied to conventional clinical medical software6,280.
Intellectual property and liability
The ownership of intellectual property in outputs generated by AI models currently lacks a unified legal definition6,16. In the commercialization of virtual cell models, the boundaries of rights and interests among data providers, model developers, and end users remain unclear. Moreover, as a novel digital product, the regulatory and legal status of virtual cell models is still being explored6. When model predictions are used to support critical R&D or clinical decision-making, and an erroneous prediction leads to adverse outcomes, determining which party should bear responsibility—as well as disputes over the allocation of commercial benefits—becomes contentious16,281. For example, if a model is trained on large volumes of patient single-cell omics data provided by a hospital, should the intellectual property rights to a drug target predicted by the model belong to the model company or to the data-providing hospital282,283,284? Similarly, if the algorithm is developed by an AI company but a pharmaceutical firm uses the model to discover a new molecule, how should the related intellectual property be shared282,283,284? These questions currently have no precedent and must be progressively clarified in practice through contracts and legislation to address real-world problems6,16. Model developers, providers, and users may shift liability among themselves, which impedes compensation for harmed parties and undermines healthy industry development.
Therefore, regulatory authorities and the legal community should intervene early to establish liability-determination frameworks and insurance mechanisms tailored to AI model applications, clarify the rights and responsibilities of all parties, and reduce the legal risks associated with adopting new technologies285,286,287. Overall, clarifying intellectual property and liability allocation is a necessary condition for translating virtual cell technology from the laboratory into industry, requiring coordinated advancement through both legislation and industry self-regulation285,286,287.
Algorithmic fairness and explainability
The fairness and transparency of virtual cell models directly influence their adoption in clinical settings16,133. If the training data contain biases related to race or other factors, models can amplify these injustices and systematically disadvantage certain groups in prediction outcomes1,16. For example, if training data are drawn predominantly from Western populations, a model may perform with lower accuracy when predicting drug responses in East Asian populations, thereby reducing those groups’ opportunities to receive effective therapies288,289,290,291. Such AI bias should be mitigated by increasing data diversity and by incorporating fairness constraints during model training16,292. Moreover, the “black-box” nature of deep learning models undermines researchers’ and clinicians’ trust in their predictions16,133. Therefore, integrating explainable artificial intelligence (XAI) approaches into virtual cell models is essential292,293,294. Common XAI techniques—such as SHAP (SHapley Additive exPlanations) value analysis and integrated gradients—can quantify the dependence of model outputs on individual input features, thereby illuminating the rationale behind model decisions292,294. Through these methods, we can answer critical questions like “Why does the model consider a particular gene mutation pathogenic?” or “On what basis does the model predict that drug A is more effective than drug B?”292. Introducing fairness constraints during model development and augmenting interpretability analyses during deployment will markedly enhance the credibility of virtual cell models and increase their likelihood of broad adoption in both research and clinical practice16,292(Fig. 5).
Virtual Cell governance and compliance translational pathway. A The regulatory-policy core locates Virtual Cell within risk stratification, credibility assessment, and pertinent provisions, and specifies its interface with drug-regulatory decision-making; B The data-compliance core covers requirements for data acquisition and cross-domain transfer, together with technical pathways such as de-identification, federated learning, and trusted execution environments; C The algorithmic-fairness core emphasizes interpretability calibration and bias detection and constraint at both population and individual levels, culminating in a model credibility report; D The application-potential layer delineates the use boundaries and evidence needs of Virtual Cell in drug evaluation, toxicological inference, and scenario-based decision-making; E The responsibility-allocation layer depicts the duties and rights of data providers, model developers, and end users regarding IP attribution, responsibility determination, and feedback-loop optimization. (Created in https://BioRender.com).
Future development and outlook
With regulatory–ethical–compliance alignment achieved, virtual-cell development enters a new phase: foundation in terminology harmonization/standardization; delineation of technical hurdles; prospect of cross-disciplinary, multiscale integration; and establishment of open-science, international-collaboration platforms to expedite innovation.
Terminology harmonization and standardization
At present, terms such as “Virtual Cell”, “Digital Cell”, and “Digital Twin” are not used consistently across academia and industry1,182. This terminological overlap not only complicates cross-disciplinary communication but can also create ambiguities in regulation and review processes6. The International Organization for Standardization (ISO) has already begun promoting the harmonization of terms and definitions in the digital twin domain; however, in the life sciences, there is an urgent need for multinational academic organizations and regulatory authorities to take the lead in developing a standardized vocabulary and classification framework for virtual cells6. Standardization efforts should extend beyond nomenclature to include model description languages, data formats, and validation metrics1,295. For example, promoting the adoption of standard model description languages (e.g., SBML, CellML) to represent virtual cell models would facilitate model sharing and reuse across different platforms125,296; likewise, establishing public model and data repositories and encouraging researchers to submit validated models would enable model reuse and iterative improvement6. By unifying terminology and standards, the virtual cell research ecosystem will become more standardized, and collaboration among researchers will proceed more smoothly297.
Technical bottlenecks and challenges
The limited interpretability of virtual cell models represents a major bottleneck to their regulatory and clinical adoption16. As black-box models, they often fail to provide explicit biological causal explanations, undermining clinicians’ and regulatory experts’ trust in their predictions1. These issues are particularly pronounced in the preclinical stages of drug development, where the accuracy and reproducibility of model outputs directly affect the design and risk assessment of subsequent clinical trials251. Developing a graded framework for model credibility is therefore a promising avenue, stratifying predictive confidence according to factors such as the volume of training data and the rigor of validation, and aligning each credibility tier with the verification intensity required for different application scenarios1. Only by addressing these bottlenecks in a targeted manner can virtual cell models gain a foothold in more stringent regulatory and clinical environments6.
Cross-disciplinary, multiscale integration
Looking ahead, virtual cells will be deeply integrated with cutting-edge technologies such as organoids, organ-on-chip platforms, and digital twins, enabling multiscale, unified modeling that spans molecules, cells, tissues, organs, and whole organisms1,6,204. Such multiscale integration is expected to more systematically predict drug responses and disease progression, thereby enhancing the role of virtual cells in precision medicine1,298. At the same time, emerging generative artificial intelligence and large multimodal models will furnish virtual cells with stronger generalization and inferential capabilities299,300. However, realizing the interdisciplinary, multiscale integration described above requires overcoming the current fragmentation of research: the vast majority of studies remain confined to a single level or domain, and disciplinary barriers impede model interoperability1. Deep cross-domain integration and the consolidation of multiscale data modalities and model types are likely to become essential pathways for the evolution of the virtual cell field301.
International collaboration and open science
To accelerate the iterative development and clinical translation of virtual-cell technologies, international collaborative and open-science platforms should be established1,302. Such platforms could enable the sharing of large-scale multimodal omics datasets, the adoption of standardized model formats, and the dissemination of open-source tools, thereby promoting harmonized model performance evaluation criteria and clear pathways for regulatory acceptance125,207,251,296,303,304. Transnational cooperation would not only expand data diversity and reduce model bias, but also facilitate mutual recognition and coordination among regulatory agencies across countries207. Naturally, open collaboration must also address challenges related to intellectual property and data sovereignty. By exploring innovative cooperation mechanisms, it is possible—while safeguarding the privacy of all parties’ data—to realize the goal of “data sharing and collaborative modeling”232,270,271,281. Open science and benchmarking are advancing rapidly. The 2025 Virtual Cell Challenge tests perturbation-response prediction on unseen cell types via open competition, improving method comparability and iteration146. Therapeutic Data Commons (PyTDC) provides a unified evaluation framework by aggregating drug-discovery and single-cell benchmarks305. The “Open Problems in Single-Cell Analysis” initiative releases benchmark datasets and metrics to coordinate community solutions306. These efforts establish shared standards and reproducible resources, strengthen cross-dataset/platform external validity, and enable rapid, robust optimization of virtual cell models. These measures will cultivate a more open academic environment and accelerate the accumulation and iterative refinement of the underlying technologies2,307,308.
Outlook
Virtual cell models are poised to evolve from auxiliary tools in preclinical research into a core component of the drug-evaluation framework1,6,12. Their simulation scope will extend beyond current intracellular processes to higher organizational levels—including tissue morphogenesis and organ-level functional dynamics—thereby enabling a more complete depiction of complex physiological and pathophysiological processes309,310. In addition, establishing internationally recognized benchmark datasets and quantitative metrics will allow objective comparisons of different models’ predictive performance and credibility1,16,251,311,312. In parallel, explainable-AI approaches will be integrated into the development pipeline of virtual cell models to document and evaluate, end-to-end, the evidentiary basis of model decisions, improving the traceability of predictions and their clinical acceptability; as transparency increases and validation cases accumulate, clinical and regulatory confidence in these models is expected to strengthen progressively313,314,315,316. Looking ahead, virtual cell technologies are expected to play a central role in drug discovery, disease modeling, and personalized medicine, achieving a closed loop from theoretical innovation to clinical translation12. Driven by the explosive growth of biological big data, sustained advances in computing hardware, and broader societal acceptance of experimental alternatives, virtual cell models are likely to transition from frontier exploration to mainstream application in the near future, catalyzing a paradigm shift across the life sciences and the biopharmaceutical industry1,317(Fig. 6).
A Terminology unification and standardization: The figure shows that terms such as “Virtual Cell”, “Digital Cell”, and “Digital Twin” are not used consistently in academia and industry. Therefore, the International Organization for Standardization (ISO) has begun to promote the unification of terms and definitions in the field of digital twins, and promote the use of standard model description languages (such as SBML, CellML) to describe virtual cell models. B International cooperation and open science: The figure shows that internationally, to accelerate the iteration and clinical transformation of virtual cell technology, it is necessary to establish platforms for international cooperation and open science, and promote unified model performance evaluation standards and regulatory adoption paths. Governments and scientific research institutions around the world are paying increasing attention to open science, and the virtual cell field is expected to emerge a global cooperation network to promote the faster application of technology. C Technical bottlenecks and challenges: The figure shows black-box models, poor cross-platform compatibility, and lack of unified evaluation standards, and shows solutions. D Trend of interdisciplinary and multi-scale integration: The figure shows that virtual cells will be deeply integrated with cutting-edge technologies such as organoids, organ-on-a-chip, and digital twins to achieve multi-scale integrated modeling from molecule-cell-tissue-organ-individual. The challenge is to be limited to a single level or a single field, and it is necessary to explore new cooperation mechanisms and research paradigms to break down the barriers between biology, computer science, and engineering. In-depth cross-field integration and multi-scale data + model integration will become an inevitable path for the evolution of the virtual cell field. (Created in https://biogdp.com).
Data availability
No datasets were generated or analyzed during the current study.
References
Bunne, C. et al. How to build the virtual cell with artificial intelligence: Priorities and opportunities. Cell 187, 7045–7063 (2024).
Johnson, G. T. et al. Building the next generation of virtual cells to understand cellular biology. Biophys. J. 122, 3560–3569 (2023).
He, X. et al. Artificial intelligence-based multi-omics analysis fuels cancer precision medicine. Semin Cancer Biol. 88, 187–200 (2023).
Loew, L. M. & Schaff, J. C. The virtual cell: a software environment for computational cell biology. Trends Biotechnol. 19, 401–406 (2001).
Gupta, R. et al. Artificial intelligence to deep learning: machine intelligence approach for drug discovery. Mol. Divers 25, 1315–1360 (2021).
Gangwal, A. & Lavecchia, A. Artificial intelligence in preclinical research: enhancing digital twins and organ-on-chip to reduce animal testing. Drug Discov. Today 30, 104360 (2025).
Wan, X. et al. Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope. Nat. Commun. 14, 7848 (2023).
Gayoso, A. et al. Joint probabilistic modeling of single-cell multi-omic data with totalVI. Nat. Methods 18, 272–282 (2021).
Worth, A. P. et al. Virtual cell based assay simulations of intra-mitochondrial concentrations in hepatocytes and cardiomyocytes. Toxicol. Vitr. 45, 222–232 (2017).
Luo, Q. et al. Status and influential factors of water-borne diseases in bathing beaches in three cities of China from 2019 to 2020. Wei Sheng Yan Jiu 50, 472–475 (2021).
Paini, A. et al. Practical use of the virtual cell based assay: simulation of repeated exposure experiments in liver cell lines. Toxicol. Vitr. 45, 233–240 (2017).
Graepel, R. et al. The virtual cell based assay: current status and future perspectives. Toxicol. Vitr. 45, 258–267 (2017).
Klein, D. et al. Mapping cells through time and space with moscot. Nature 638, 1065–1075 (2025).
UNION E P C O T E. Regulation (EU) 2016/679 (General Data Protection Regulation) [Z]. Official Journal of the European Union. (EU, 2016)
Scheuher, B. et al. Towards a platform quantitative systems pharmacology (QSP) model for preclinical to clinical translation of antibody drug conjugates (ADCs). J. Pharmacokinet. Pharmacodyn. 51, 429–447 (2024).
Hartung, T., Maertens, A. & Luechtefeld, T. E-validation - Unleashing AI for validation. Altex 41, 567–587 (2024).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Sidaway, P. Neoadjuvant pembrolizumab shows promise. Nat. Rev. Clin. Oncol. 16, 7 (2019).
Schep, A. N. et al. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Asp, M. et al. A spatiotemporal organ-wide gene expression and cell atlas of the developing human heart. Cell 179, 1647–60.e19 (2019).
Tian, L. et al. Benchmarking single cell RNA-sequencing analysis pipelines using mixture control experiments. Nat. Methods 16, 479–487 (2019).
Eng, C. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Xia, C. et al. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl. Acad. Sci. USA 116, 19490–19499 (2019).
Soopairin, S., Patikorn, C. & Taychakhoonavudh, S. Antivenom preclinical efficacy testing against Asian snakes and their availability in Asia: a systematic review. PLoS One 18, e0288723 (2023).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Ashuach, T. et al. MultiVI: deep generative model for the integration of multimodal data. Nat. Methods 20, 1222–1231 (2023).
Li Y., Zhang S. Statistical batch-aware embedded integration, dimension reduction, and alignment for spatial transcriptomics. Bioinformatics 40, btae611 (2024).
Chen, P. C. et al. An augmented reality microscope with real-time artificial intelligence integration for cancer diagnosis. Nat. Med. 25, 1453–1457 (2019).
Wang, J. et al. scGNN is a novel graph neural network framework for single-cell RNA-Seq analyses. Nat. Commun. 12, 1882 (2021).
Cao, Z. J. & Gao, G. Multi-omics single-cell data integration and regulatory inference with graph-linked embedding [J]. Nat. Biotechnol. 40, 1458–1466 (2022).
Cui, H. et al. Towards multimodal foundation models in molecular cell biology. Nature 640, 623–633 (2025).
Xie, X. Q. Exploiting PubChem for virtual screening. Expert Opin. Drug Discov. 5, 1205–1220 (2010).
Antunes, D. A. et al. HLA-Arena: a customizable environment for the structural modeling and analysis of peptide-hla complexes for cancer immunotherapy. JCO Clin. Cancer Inform. 4, 623–636 (2020).
Campagne, F. et al. Quantitative information management for the biochemical computation of cellular networks. Sci. STKE 2004, pl11 (2004).
Tomczak, K., Czerwińska, P. & Wiznerowicz, M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp. Oncol. 19, A68–A77 (2015).
Weinstein, J. N. et al. The Cancer Genome Atlas pan-cancer analysis project. Nat. Genet 45, 1113–1120 (2013).
Thul, P. J. & Lindskog, C. The human protein atlas: a spatial map of the human proteome [J]. Protein Sci. 27, 233–244 (2018).
Digre, A. & Lindskog, C. The Human Protein Atlas-spatial localization of the human proteome in health and disease. Protein Sci. 30, 218–233 (2021).
Pontén, F., Jirström, K. & Uhlen, M. The Human Protein Atlas—a tool for pathology. J. Pathol. 216, 387–393 (2008).
Clough, E. & Barrett, T. The Gene Expression Omnibus Database. Methods Mol. Biol. 1418, 93–110 (2016).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995 (2013).
Edgar, R., Domrachev, M. & Lash, A. E. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Rasnic, R. et al. Substantial batch effects in TCGA exome sequences undermine pan-cancer analysis of germline variants. BMC Cancer 19, 783 (2019).
Ghaddar B. C., Blaser M. J., De S. Revisiting the cancer microbiome using PRISM [J]. bioRxiv, 2025.
Engin, B. & Güner, O. R. Negative evaluation of a pathergy test in hepatitis B surface antigen carriers. J. Dermatol 33, 547–549 (2006).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Wang, J. ComBat-met: adjusting batch effects in DNA methylation data. NAR Genom. Bioinform 7, lqaf062 (2025).
Leach, D. T. et al. malbacR: a package for standardized implementation of batch correction methods for omics data. Anal. Chem. 95, 12195–12199 (2023).
Antonsson, S. E. & Melsted, P. Batch correction methods used in single-cell RNA sequencing analyses are often poorly calibrated. Genome Res. 35, 1832–1841 (2025).
Kepp, O. et al. Consensus guidelines for the detection of immunogenic cell death. Oncoimmunology 3, e955691 (2014).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet 11, 733–739 (2010).
Chen, C. et al. Removing batch effects in analysis of expression microarray data: an evaluation of six batch adjustment methods. PLoS One 6, e17238 (2011).
Zhou et al. Outlier detection method based on high-density iteration. Inf. Sci. 662, 120268 (2024).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Hao, M. et al. Large-scale foundation model on single-cell transcriptomics. Nat. Methods 21, 1481–1491 (2024).
Roohani, Y., Huang, K. & Leskovec, J. Predicting transcriptional outcomes of novel multigene perturbations with GEARS. Nat. Biotechnol. 42, 927–935 (2024).
Adduri et al. Predicting cellular responses to perturbation across diverse contexts with State. bioRxiv, 2025.
He, D. et al. A context-aware deconfounding autoencoder for robust prediction of personalized clinical drug response from cell-line compound screening. Nat. Mach. Intell. 4, 879–892 (2022).
Chen, J. et al. Deep transfer learning of cancer drug responses by integrating bulk and single-cell RNA-seq data. Nat. Commun. 13, 6494 (2022).
Bang, D., Koo, B. & Kim, S. Transfer learning of condition-specific perturbation in gene interactions improves drug response prediction. Bioinformatics 40, i130–i139 (2024).
Zheng, Y. et al. A deep generative model for deciphering cellular dynamics and in silico drug discovery in complex diseases. Nat. Biomed. Eng. 10, 1038 (2025).
Morehead, A., Cheng, J. FlowDock: geometric flow matching for generative protein-ligand docking and affinity prediction. ArXiv, 2025.
Yu Z, Xia, Hao, Yu, Dahui, Cheng, Jiaoyang, Li, Jichun
Wang, P. et al. Score-based image-to-image brownian bridge. Proc. ACM Int Conf. Multimed. 2024, 10765–10773 (2024).
Luo, E. et al. scDiffusion: conditional generation of high-quality single-cell data using diffusion model. Bioinformatics 40, btae518 (2024).
Ahlmann-Eltze, C., Huber, W. & Anders, S. Deep-learning-based gene perturbation effect prediction does not yet outperform simple linear baselines. Nat. Methods 22, 1657–1661 (2025).
Feng, K. et al. Shared growth of graph neural networks via prompted free-direction knowledge distillation. IEEE Trans. Pattern Anal. Mach. Intell. 47, 4377–4394 (2025).
Sun J., et al. GNN codon adjacency tunes protein translation. Int. J. Mol. Sci. 25, 5914 (2024).
Mccardle, K. Shedding light on GNN affinity predictions. Nat. Comput Sci. 3, 1004 (2023).
Réau, M. et al. DeepRank-GNN: a graph neural network. Bioinformatics 39, 759 (2023).
Dong, Z., Feng, J. & Ji, Y. et al. SLI-GNN: a self-learning-input graph neural network for predicting crystal and molecular properties. J. Phys. Chem. A 127, 5921–5929 (2023).
Wang, R.H., Luo, T. & Zhang, H.L. et al. PLA-GNN: computational inference of protein subcellular location alterations under drug treatments with deep graph neural networks. Comput. Biol. Med. 157, 106775 (2023).
Chen, S. et al. MD-GNN: a mechanism-data-driven graph neural network for molecular properties prediction and new material discovery. J. Mol. Graph Model 123, 108506 (2023).
Wang, H. et al. CCF-GNN: a unified model aggregating appearance, microenvironment, and topology for pathology image classification. IEEE Trans. Med Imaging 42, 3179–3193 (2023).
Li, W. et al. Drug repurposing based on the DTD-GNN graph neural network: revealing the relationships among drugs, targets and diseases. BMC Genomics 25, 584 (2024).
Li S., Hua H., Chen S. Graph neural networks for single-cell omics data: a review of approaches and applications. Brief Bioinform. 26, 109 (2025).
Ning, X. et al. Physics-informed neural networks integrating compartmental model for analyzing COVID-19 transmission dynamics. Viruses 15, 1749 (2023).
Sharpee T. O., et al. 25th annual computational neuroscience meeting: CNS-2016. BMC Neurosci. 17, 54 (2016).
Song, Y. et al. SRS-Net: a universal framework for solving stimulated Raman scattering in nonlinear fiber-optic systems by physics-informed deep learning. Commun. Eng. 3, 109 (2024).
Lagergren, J. H. et al. Biologically-informed neural networks guide mechanistic modeling from sparse experimental data. PLoS Comput Biol. 16, e1008462 (2020).
Sattari, A. Machine learning in biofluid mechanics: a review of recent developments. Comput. Biol. Med. 193, 110410 (2025).
Colombo, M. et al. HER2 targeting as a two-sided strategy for breast cancer diagnosis and treatment: Outlook and recent implications in nanomedical approaches. Pharm. Res. 62, 150–165 (2010).
Yang S., et al. Up-regulation of CXCL8 expression is associated with a poor prognosis and enhances tumor cell malignant behaviors in liver cancer. Biosci. Rep. 40, BSR20201169 (2020).
Zhang, Z. & Qu, Z. Bistable nerve conduction. Biophys. J. 121, 3499–3507 (2022).
Griffith, B. E. & Peskin, C. S. Electrophysiology. Commun. Pure Appl Math. 66, 1837–1913 (2013).
Zhang, X. et al. Physics-informed neural networks (PINNs) for 4D hemodynamics prediction: an investigation of optimal framework based on vascular morphology. Comput Biol. Med. 164, 107287 (2023).
Malashin I., et al. Physics-informed neural networks in polymers: a review. Polymers 17, 1108 (2025).
Nam, Y. et al. Harnessing artificial intelligence in multimodal omics data integration: paving the path for the next frontier in precision medicine. Annu. Rev. Biomed. Data Sci. 7, 225–250 (2024).
Yang, X. et al. scCross: a deep generative model for unifying single-cell multi-omics with seamless integration, cross-modal generation, and in silico exploration. Genome Biol. 25, 198 (2024).
Wang G., et al. Modeling and predicting single-cell multi-gene perturbation responses with scLAMBDA [J]. bioRxiv, 2024.
Kana, O. et al. Generative modeling of single-cell gene expression for dose-dependent chemical perturbations. Patterns 4, 100817 (2023).
Bunne, C. et al. Learning single-cell perturbation responses using neural optimal transport. Nat. Methods 20, 1759–1768 (2023).
Denoeud, A. et al. Dynamic X-ray diffraction observation of shocked solid iron up to 170GPa. Proc. Natl. Acad. Sci. USA 113, 7745–7749 (2016).
Zhu L., et al. Microfluidic-based platforms for cell-to-cell communication studies. Biofabrication 16, 1116 (2023).
Hoops, S. et al. COPASI—a complex pathway simulator. Bioinformatics 22, 3067–3074 (2006).
Red. [Not Available]. MMW Fortschr. Med. 87, 158 (2016).
Bergmann, F. T. et al. COPASI and its applications in biotechnology. J. Biotechnol. 261, 215–220 (2017).
Sütterlin, T. et al. Bridging the scales: semantic integration of quantitative SBML in graphical multi-cellular models and simulations with EPISIM and COPASI. Bioinformatics 29, 223–229 (2013).
Cao, L. et al. Auditory perception modulated by word reading. Exp. Brain Res 234, 3049–3057 (2016).
Blinov, M. L. et al. BioNetGen: software for rule-based modeling of signal transduction based on the interactions of molecular domains. Bioinformatics 20, 3289–3291 (2004).
Husar, A. et al. MCell4 with BioNetGen: a Monte Carlo simulator of rule-based reaction-diffusion systems with Python interface. PLoS Comput. Biol. 20, e1011800 (2024).
Lopez, C. F. et al. Programming biological models in Python using PySB. Mol. Syst. Biol. 9, 646 (2013).
Vykoukal, J. V., Fahrmann, J. F. & Thompson, T. C. Caveolin and lipid domains-close companions in managing cellular pathways. Cancer Metastasis Rev. 39, 341–342 (2020).
Terfve, C. et al. CellNOptR: a flexible toolkit to train protein signaling networks to data using multiple logic formalisms. BMC Syst. Biol. 6, 133 (2012).
Gjerga, E. et al. Converting networks to predictive logic models from perturbation signalling data with CellNOpt. Bioinformatics 36, 4523–4524 (2020).
Helikar, T., Kowal, B. & Rogers, J. A. A cell simulator platform: the cell collective. Clin. Pharm. Ther. 93, 393–395 (2013).
Proença, S. et al. Insights into in vitro biokinetics using virtual cell based assay simulations. Altex 36, 447–461 (2019).
Comenges, J. M. Z. et al. Theoretical and mathematical foundation of the virtual cell based assay - a review. Toxicol. Vitr. 45, 209–221 (2017).
Ghaffarizadeh, A. et al. PhysiCell: an open source physics-based cell simulator for 3-D multicellular systems. PLoS Comput Biol. 14, e1005991 (2018).
Heiland R., et al. PhysiCell Studio: a graphical tool to make agent-based modeling more accessible. bioRxiv, 2023.
Heiland, R. et al. PhysiCell Studio: a graphical tool to make agent-based modeling more accessible. GigaByte 2024, gigabyte128 (2024).
Smeriglio, R. et al. Start & Stop: a physicell and physiBoSS 2.0 add-on for interactive simulation control. BMC Bioinforma. 26, 158 (2025).
Swat, M. H. et al. Multi-scale modeling of tissues using CompuCell3D. Methods Cell Biol. 110, 325–366 (2012).
Fortuna, I. et al. CompuCell3D simulations reproduce mesenchymal cell migration on flat substrates. Biophys. J. 118, 2801–2815 (2020).
Liu, R. et al. Development of a coupled simulation toolkit for computational radiation biology based on Geant4 and CompuCell3D. Phys. Med Biol. 66, 045026 (2021).
Palm, M. M. & Merks, R. M. Large-scale parameter studies of cell-based models of tissue morphogenesis using compucell3D or VirtualLeaf. Methods Mol. Biol. 1189, 301–322 (2015).
Starruß, J. et al. Morpheus: a user-friendly modeling environment for multiscale and multicellular systems biology. Bioinformatics 30, 1331–1332 (2014).
Ruffinatti, F. A. et al. MORPHEUS: An automated tool for unbiased and reproducible cell morphometry. J. Cell Physiol. 235, 10110–10115 (2020).
Andrews, S. S. et al. Detailed simulations of cell biology with Smoldyn 2.1. PLoS Comput. Biol. 6, e1000705 (2010).
Andrews, S. S. Smoldyn: particle-based simulation with rule-based modeling, improved molecular interaction and a library interface. Bioinformatics 33, 710–717 (2017).
Schöneberg, J. & Noé, F. ReaDDy—a software for particle-based reaction-diffusion dynamics in crowded cellular environments. PLoS One 8, e74261 (2013).
Hoffmann, M., Fröhner, C. & Noé, F. ReaDDy 2: fast and flexible software framework for interacting-particle reaction dynamics. PLoS Comput. Biol. 15, e1006830 (2019).
Mirams, G. R. et al. Chaste: an open source C++ library for computational physiology and biology. PLoS Comput. Biol. 9, e1002970 (2013).
Solmi, F. et al. Decomposing socio-economic inequality in colorectal cancer screening uptake in England. Soc. Sci. Med. 134, 76–86 (2015).
Garny, A. & Hunter, P. J. OpenCOR: a modular and interoperable approach to computational biology. Front Physiol. 6, 26 (2015).
Delp, S. L. et al. OpenSim: open-source software to create and analyze dynamic simulations of movement. IEEE Trans. Biomed. Eng. 54, 1940–1950 (2007).
Gautam, A. S. et al. Temporary reduction in air pollution due to anthropogenic activity switch-off during COVID-19 lockdown in Northern parts of India. Environ. Dev. Sustain. 23, 8774–8797 (2021).
Obermeier, M. M. et al. Plant resistome profiling in evolutionary old bog vegetation provides new clues to understand emergence of multi-resistance. ISME J. 15, 921–937 (2021).
Stringer, C. et al. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Riendeau, J. M. et al. Cellpose as a reliable method for single-cell segmentation of autofluorescence microscopy images. Sci. Rep. 15, 5548 (2025).
Etienam C. et al. A Novel A.I Enhanced reservoir characterization with a combined mixture of experts—NVIDIA modulus based physics informed neural operator forward model. arXiv 2404.14447 (2024).
Nazer, L. H. et al. Bias in artificial intelligence algorithms and recommendations for mitigation. PLOS Digit Health 2, e0000278 (2023).
Mansoor, S. et al. Zero-shot mutation effect prediction on protein stability and function using RoseTTAFold. Protein Sci. 32, e4780 (2023).
Ahmed, S. et al. Prediction of residue-specific contributions to binding and thermal stability using yeast surface display. Front. Mol. Biosci. 8, 800819 (2021).
Mao, S. et al. Development and validation of a novel preoperative clinical model for predicting lymph node metastasis in perihilar cholangiocarcinoma. BMC Cancer 24, 297 (2024).
Corrò, C., Novellasdemunt, L. & Li, V. S. W. A brief history of organoids. Am. J. Physiol. Cell Physiol. 319, C151–c65 (2020).
Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet 23, 467–491 (2022).
Forrow, A. & Schiebinger, G. LineageOT is a unified framework for lineage tracing and trajectory inference. Nat. Commun. 12, 4940 (2021).
Malepathirana, T. et al. Dimensionality reduction for visualizing high-dimensional biological data. Biosystems 220, 104749 (2022).
Kim, S. et al. Ensemble of sparse classifiers for high-dimensional biological data. Int J. Data Min. Bioinform. 12, 167–183 (2015).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Yang B., et al. Multi-view multi-level contrastive graph convolutional network for cancer subtyping on multi-omics data. Brief Bioinform. 26, 043 (2024).
Zhang, J. et al. Strategic multi-omics data integration via multi-level feature contrasting and matching. IEEE Trans. Nanobiosci. 23, 579–590 (2024).
Rabin, A. et al. SRCP: a comprehensive pipeline for accurate annotation and quantification of circRNAs. Genome Biol. 22, 277 (2021).
Roohani, Y. H. et al. Virtual cell challenge: toward a turing test for the virtual cell. Cell 188, 3370–3374 (2025).
Yin, W., Liu, Y. & Shen, C. Virtual normal: enforcing geometric constraints for accurate and robust depth prediction. IEEE Trans. Pattern Anal. Mach. Intell. 44, 7282–7295 (2022).
Xu et al. Virtual microfluidics for digital quantification and single-cell sequencing. Nat. Methods 13, 759–762 (2016).
Zhang, Y. K. et al. Identification, experimental validation, and computational evaluation of potential ALK inhibitors through hierarchical virtual screening. SAR QSAR Environ. Res 36, 271–285 (2025).
Carroll, G. T. et al. Experimental validation of convection-diffusion discretisation scheme employed for computational modelling of biological mass transport. Biomed. Eng. Online 9, 34 (2010).
Chen, S. et al. Machine learning-driven prediction of eye irritation toxicity: integration of in silico and in vitro study. Toxicol. Appl Pharm. 502, 117457 (2025).
Tanoli, Z., Schulman, A. & Aittokallio, T. Validation guidelines for drug-target prediction methods. Expert Opin. Drug Discov. 20, 31–45 (2025).
Reker, D. et al. Revealing the macromolecular targets of complex natural products. Nat. Chem. 6, 1072–1078 (2014).
Wei, J. & Li, Y. CRISPR-based gene editing technology and its application in microbial engineering. Eng. Microbiol 3, 100101 (2023).
Cao et al. Advances in precise regulation of CRISPR/Cas9 gene editing technology. Yi Chuan 42, 1168–1177 (2020).
Zhang, D. et al. CRISPR/Cas: a powerful tool for gene function study and crop improvement. J. Adv. Res 29, 207–221 (2021).
Akram, M. et al. Uncertainty-aware diabetic retinopathy detection using deep learning enhanced by Bayesian approaches. Sci. Rep. 15, 1342 (2025).
Mao, H., Martin, R. & Reich, B. J. Valid model-free spatial prediction. J. Am. Stat. Assoc. 119, 904–914 (2024).
Cabot, J. H. & Ross, E. G. Evaluating prediction model performance. Surgery 174, 723–726 (2023).
Huang, J. et al. Development of two-dimension epidemic prediction model. Infect. Dis. Model 10, 1190–1207 (2025).
Ooft S. N., et al. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Sci. Transl. Med. 11, 2574 (2019).
Sang, L. et al. An in silico platform to predict cardiotoxicity risk of anti-tumor drug combination with hiPSC-CMs based in vitro study. Pharm. Res. 41, 247–262 (2024).
Harter, M. F. et al. Analysis of off-tumour toxicities of T-cell-engaging bispecific antibodies via donor-matched intestinal organoids and tumouroids. Nat. Biomed. Eng. 8, 345–360 (2024).
FDA pushes to replace animal testing. Nat. Biotechnol. 43, 655 (2025).
Rosenblatt, M. et al. Data leakage inflates prediction performance in connectome-based machine learning models. Nat. Commun. 15, 1829 (2024).
Ma, C. et al. Organ-on-a-chip: a new paradigm for drug development. Trends Pharm. Sci. 42, 119–133 (2021).
Liu, X. et al. Reporting guidelines for clinical trial reports for interventions involving artificial intelligence: the CONSORT-AI extension. Nat. Med. 26, 1364–1374 (2020).
Debray, R. et al. Priority effects in microbiome assembly. Nat. Rev. Microbiol. 20, 109–121 (2022).
Chevalier, V. et al. First steps of experimental validation of a numerical model about mechanical behavior of NiTi endodontic instruments. Bull. Group Int Rech. Sci. Stomatol Odontol. 50, 46–47 (2011).
Albijanic B., et al. Induction time of wetting films between air bubbles and hydrophobic particles in the presence of dodecyl amine hydrochloride: first principles model analysis and experimental validation. Molecules 30, 695 2025.
Martin, K. et al. The Me first communication model. Nurs. Child Young People 31, 38–47 (2019).
Chen, L. et al. Discovery of anticancer activity of amentoflavone on esophageal squamous cell carcinoma: bioinformatics, structure-based virtual screening, and biological evaluation. J. Microbiol. Biotechnol. 32, 718–729 (2022).
Li, F. et al. 2-(2-Methylfuran-3-carboxamido)-3-phenylpropanoic acid, a potential CYP26A1 inhibitor to enhance all-trans retinoic acid-induced leukemia cell differentiation based on virtual screening and biological evaluation. Bioorg. Med Chem. 21, 3256–3261 (2013).
Sanachai, K. et al. Pharmacophore-based virtual screening and experimental validation of pyrazolone-derived inhibitors toward Janus kinases. ACS Omega 7, 33548–33559 (2022).
Naryzhny, S. N. et al. Combination of virtual and experimental 2DE together with ESI LC-MS/MS gives a clearer view about proteomes of human cells and plasma. Electrophoresis 37, 302–309 (2016).
Ukimura, O. et al. Real-time virtual ultrasonographic radiofrequency ablation of renal cell carcinoma. BJU Int. 101, 707–711 (2008).
Oikonomou, E. et al. BRAF(V600E) efficient transformation and induction of microsatellite instability versus KRAS(G12V) induction of senescence markers in human colon cancer cells. Neoplasia 11, 1116–1131 (2009).
Yu, G. et al. Facile dimension transformation strategy for fabrication of efficient and stable CsPbI(3) perovskite solar cells. ACS Appl Mater. Interfaces 15, 17825–17833 (2023).
Xu, B. et al. Concept and framework of digital twin human geographical environment. J. Environ. Manag. 373, 123866 (2025).
Asciak, L. et al. Digital twin assisted surgery, concept, opportunities, and challenges. npj Digit. Med. 8, 32 (2025).
Peshkova, M. et al. Digital twin concept: healthcare, education, research. J. Pathol. Inf. 14, 100313 (2023).
Bordukova, M. et al. Generative artificial intelligence empowers digital twins in drug discovery and clinical trials. Expert Opin. Drug Discov. 19, 33–42 (2024).
Guan A Z, Suyang G, Wei W, Zhi G, Mingyang L, Haiquan Z, Xiao-P. Dynamic simulation and parameter calibration-based experimental digital twin platform for heat-electric coupled system. IEEE Trans. Sustain. Energy 10, 3609042 (2025).
Camps, J. et al. Digital twinning of the human ventricular activation sequence to Clinical 12-lead ECGs and magnetic resonance imaging using realistic Purkinje networks for in silico clinical trials. Med. Image Anal. 94, 103108 (2024).
Li, H. et al. Digital quantitative detection for heterogeneous protein and MRNA expression patterns in circulating tumor cells. Adv. Sci. 12, e2410120 (2025).
Baandrup, L. et al. Development of a digital algorithm for assessing tumor-stroma ratio, tumor budding and tumor infiltrating lymphocytes in vulvar squamous cell carcinomas. Ann. Diagn. Pathol. 76, 152462 (2025).
Ștefănigă S. A., et al. Advancing precision oncology with digital and virtual twins: a scoping review. Cancers 16, 3387 (2024).
Wang, H. et al. From virtual patients to digital twins in immuno-oncology: lessons learned from mechanistic quantitative systems pharmacology modeling. npj Digit. Med. 7, 189 (2024).
Aghamiri S. S., et al. Digital twin technology in radiology. J. Imaging Inform. Med. 11, 6553 (2025).
Belik, M. & Rubanenko, O. Sensitivity analysis of digital twin model for energy community PV system. Sci. Rep. 15, 29097 (2025).
Shu, H. et al. Twin-S: a digital twin for skull base surgery. Int J. Comput. Assist. Radio. Surg. 18, 1077–1084 (2023).
Scott, A. K. & Oyen, M. L. Virtual pregnancies: predicting and preventing pregnancy complications with digital twins. Lancet Digit. Health 6, e436–e437 (2024).
Serrano D. R., et al. Artificial intelligence (AI) applications in drug discovery and drug delivery: revolutionizing personalized medicine. Pharmaceutics 16, 1328 (2024).
Brydon, N. Advantages and limitations of physical and virtual dose mapping. Biomed. Instrum. Technol. 58, 34–38 (2024).
Morel, M. et al. Advantages and limitations of virtual reality for balance assessment and rehabilitation. Neurophysiol. Clin. 45, 315–326 (2015).
Seemann, M. D., Schaefer, J. F. & Englmeier, K. H. Virtual positron emission tomography/computed tomography-bronchoscopy: possibilities, advantages and limitations of clinical application. Eur. Radio. 17, 709–715 (2007).
Grasso E., et al. Role of virtual iMRI in glioblastoma surgery: advantages, limitations, and correlation with iCT and brain shift. Brain Sci. 15, 35 (2024).
Kozłowska, E. et al. Mathematical modeling predicts response to chemotherapy and drug combinations in ovarian cancer. Cancer Res. 78, 4036–4044 (2018).
Del Rio, E. & Ferreira, L. F. An expression of uncertainty and its application to positioning: a quality-metric and optimal ranges for the identification of cells with RFID. Springerplus 4, 374 (2015).
Ramaswamy, R. K. et al. Virtual reality-guided left ventricular assist device implantation in pediatric patient: Valuable presurgical tool. Ann. Pediatr. Cardiol. 14, 388–392 (2021).
Weihe, W. H. Use and misuse of an imprecise concept: alternative methods in animal experiments. Lab Anim. 19, 19–26 (1985).
Kiani et al. Ethical considerations regarding animal experimentation. J. Prev. Med Hyg. 63, E255–e266 (2022).
Zhou, L. et al. Organoids and organs-on-chips: recent advances, applications in drug development, and regulatory challenges. Medicine 6, 100667 (2025).
Han, J. J. FDA modernization Act 2.0 allows for alternatives to animal testing. Artif. Organs 47, 449–450 (2023).
Chen, R. et al. Receptor conversion in metastatic breast cancer: analysis of 390 cases from a single institution. Mod. Pathol. 33, 2499–2506 (2020).
Deshmukh, A. D. & Wagner, J. K. FDA draft guidelines for AI and the need for ethical frameworks. JAMA Pediatr. 179, 937–938 (2025).
Herron, E. K. & Weeks, K. A. Development of a clinical competency pedagogy with senior nursing students as preparation for transition to practice. Nurse Educ. 46, E137–e138 (2021).
Yang, N., Dai, R. & Zhang, X. Global prevalence of human pegivirus-1 in healthy volunteer blood donors: a systematic review and meta-analysis. Vox Sang. 115, 107–119 (2020).
Wu, Y. et al. Beyond success: unveiling the hidden potential of radiotherapy and immunotherapy in solid tumors. Cancer Commun. 44, 739–760 (2024).
Zheng W., et al. Tumor-associated neutrophils in colorectal cancer development, progression and immunotherapy. Cancers 14, 4755 (2022).
Hartung, T. & Kleinstreuer, N. Challenges and opportunities for validation of AI-based new approach methods. Altex 42, 3–21 (2025).
Deng, Y. et al. Collective motility and mechanical waves in cell clusters. Eur. Phys. J. E Soft Matter 44, 137 (2021).
Schaufel, M. A. et al. Stretching oneself too thin and facing ethical challenges: healthcare professionals’ experiences during the COVID-19 pandemic. Nurs. Ethics 31, 1630–1645 (2024).
Edwards, C. et al. The role of patient outcomes in shaping moral responsibility in AI-supported decision making. Radiography 31, 102948 (2025).
Boyd, A. et al. How hospital survey teams function. J. Health Organ Manag 32, 206–223 (2018).
Zhang, K. et al. FDA review of radiologic AI algorithms: process and challenges. Radiology 310, e230242 (2024).
Harvey, H. B. & Gowda, V. How the FDA Regulates AI. Acad. Radio. 27, 58–61 (2020).
Ebrahimian, S. et al. FDA-regulated AI algorithms: trends, strengths, and gaps of validation studies. Acad. Radio. 29, 559–566 (2022).
Mcnamara, S. L., Yi, P. H. & Lotter, W. The clinician-AI interface: intended use and explainability in FDA-cleared AI devices for medical image interpretation. npj Digit. Med. 7, 80 (2024).
Windecker, D. et al. Generalizability of FDA-approved AI-enabled medical devices for clinical use. JAMA Netw. Open 8, e258052 (2025).
Muralidharan, V. et al. A scoping review of reporting gaps in FDA-approved AI medical devices. npj Digit. Med. 7, 273 (2024).
Benjamens, S., Dhunnoo, P. & Meskó, B. The state of artificial intelligence-based FDA-approved medical devices and algorithms: an online database. npj Digit. Med. 3, 118 (2020).
Lin, M. What’s needed to bridge the gap between US FDA clearance and real-world use of AI algorithms. Acad. Radio. 29, 567–568 (2022).
Halfpenny, W. & Baxter, S. L. Towards effective data sharing in ophthalmology: data standardization and data privacy. Curr. Opin. Ophthalmol. 33, 418–424 (2022).
Bonomi, L., Huang, Y. & Ohno-Machado, L. Privacy challenges and research opportunities for genomic data sharing. Nat. Genet. 52, 646–654 (2020).
Malin, B. & Goodman, K. Between access and privacy: challenges in sharing health data. Yearb. Med Inf. 27, 55–59 (2018).
Schreiber, R., Koppel, R. & Kaplan, B. What do we mean by sharing of patient data? Dash: a data sharing hierarchy of privacy and ethical challenges. Appl Clin. Inf. 15, 833–841 (2024).
White, T., Blok, E. & Calhoun, V. D. Data sharing and privacy issues in neuroimaging research: opportunities, obstacles, challenges, and monsters under the bed. Hum. Brain Mapp. 43, 278–291 (2022).
Chen, R. et al. Data sharing and privacy in pharmaceutical studies. Curr. Pharm. Des. 27, 911–918 (2021).
Alfawzan, N. et al. Privacy, data sharing, and data security policies of women’s mhealth apps: scoping review and content analysis. JMIR Mhealth Uhealth 10, e33735 (2022).
Conduah, A. K., Ofoe, S. & Siaw-Marfo, D. Data privacy in healthcare: global challenges and solutions. Digit. Health 11, 20552076251343959 (2025).
Drummond, D. & Gonsard, A. Definitions and characteristics of patient digital twins being developed for clinical use: scoping review. J. Med. Internet Res. 26, e58504 (2024).
Cramer, J. Privacy, data sharing, and other legal considerations. Surg. Clin. North Am. 103, 347–356 (2023).
Wirth, F. N. et al. Privacy-preserving data sharing infrastructures for medical research: systematization and comparison. BMC Med. Inf. Decis. Mak. 21, 242 (2021).
Huang, D., Ye, X. & Sakurai, T. Multi-party collaborative drug discovery via federated learning [J]. Comput. Biol. Med. 171, 108181 (2024).
Li Y., et al. LF3PFL: a practical privacy-preserving federated learning algorithm based on local federalization scheme. Entropy 26, 353 (2024).
Wang, X. Z., Wang, R. & Xu, C. Discovering the relationship between generalization and uncertainty by incorporating complexity of classification. IEEE Trans. Cyber 48, 703–715 (2018).
Gear, C., Koziol-Mclain, J. & Eppel, E. Engaging with uncertainty and complexity: a secondary analysis of primary care responses to intimate partner violence. Glob. Qual. Nurs. Res. 8, 2333393621995164 (2021).
Lawless W. F., Moskowitz I. S., Doctor K. Z. A Quantum-like model of interdependence for embodied human-machine teams: reviewing the path to autonomy facing complexity and uncertainty. Entropy 25, 1323 (2023).
Tajmir, S. H. et al. Artificial intelligence-assisted interpretation of bone age radiographs improves accuracy and decreases variability. Skelet. Radio. 48, 275–283 (2019).
Folgert, A. & Degroot, K. Using AI-generated podcasts as an adjunct to traditional teaching strategies. Nurse Educ. 50, 78 (2025).
Yang, Y. et al. Integration of AI-assisted in digital cervical cytology training: a comparative study. Cytopathology 36, 156–164 (2025).
Vinnakota, K. C. et al. Improving the physiological realism of experimental models. Interface Focus 6, 20150076 (2016).
Fletcher, A. G. & Osborne, J. M. Seven challenges in the multiscale modeling of multicellular tissues. WIREs Mech. Dis. 14, e1527 (2022).
Hunter, P. J. The IUPS physiome project: a framework for computational physiology [J]. Prog. Biophys. Mol. Biol. 85, 551–569 (2004).
Zhao Z., et al. Organoids. Nat. Rev. Methods Primers 2, e274 (2022).
Wang, H. et al. Human organoids-on-chips for biomedical research and applications. Theranostics 14, 788–818 (2024).
Huang, P. H., Kim, K. H. & Schermer, M. Ethical issues of digital twins for personalized health care service: preliminary mapping study. J. Med. Internet Res. 24, e33081 (2022).
UNION E P C O T E. Regulation (EU) 2016/679 (General Data Protection Regulation), (EU, 2016).
UNION E P C O T E. Regulation (EU) 2025/327 on the European Health Data Space and amending Directive 2011/24/EU, (EU, 2025).
Yu, L., Stokes, J. R. & Yakubov, G. E. Viscoelastic behaviour of rapid and slow self-healing hydrogels formed by densely branched arabinoxylans from Plantago ovata seed mucilage. Carbohydr. Polym. 269, 118318 (2021).
Mohammed Yakubu, A. & Chen, Y. P. Ensuring privacy and security of genomic data and functionalities. Brief. Bioinform 21, 511–526 (2020).
Thapa, C. & Camtepe, S. Precision health data: requirements, challenges and existing techniques for data security and privacy. Comput Biol. Med 129, 104130 (2021).
Oh S. R., et al. A comprehensive survey on security and privacy for electronic health data. Int. J. Environ. Res. Public Health 18, 9668 (2021).
Rai, H. M. et al. Enhancing data security and privacy in energy applications: Integrating IoT and blockchain technologies. Heliyon 10, e38917 (2024).
Kobayashi, S., Kane, T. B. & Paton, C. The privacy and security implications of open data in healthcare. Yearb. Med Inf. 27, 41–47 (2018).
Khalid M. I., Ahmed M., Kim J. Enhancing data protection in dynamic consent management systems: formalizing privacy and security definitions with differential privacy, decentralization, and zero-knowledge proofs. Sensors 23, 7604 (2023).
Yakubu, B. M., Sabi’u, J. & Bhattarakosol, P. Blockchain-based privacy and security model for transactional data in large private networks. Sci. Rep. 13, 17108 (2023).
Mackenzie, I. S. et al. Managing security and privacy concerns over data storage in healthcare research. Pharmacoepidemiol Drug Saf. 20, 885–893 (2011).
Zhang P., Ma J. Channel characteristic aware privacy protection mechanism in WBAN. Sensors 18, 2403 (2018).
Liu, L. et al. Dual blockchain-based data sharing mechanism with privacy protection for medical internet of things. Heliyon 10, e23575 (2024).
Liang, S., Lam, J. & Lin, H. Secure estimation with privacy protection. IEEE Trans. Cyber 53, 4947–4961 (2023).
Li, Z. et al. Local differential privacy protection for wearable device data. PLoS One 17, e0272766 (2022).
Lang, F. & Zhong, Y. Application of personal information privacy protection based on machine learning algorithm. Comput Intell. Neurosci. 2022, 6710631 (2022).
Altman, M. & Cohen, A. Natural differential privacy-a perspective on protection guarantees. PeerJ Comput Sci. 9, e1576 (2023).
Hu, Z. & Yang, J. Differential privacy protection method based on published trajectory cross-correlation constraint. PLoS One 15, e0237158 (2020).
Kaul, V. & Mukherjee, T. Equitable differential privacy. Front Big Data 7, 1420344 (2024).
Dyda, A. et al. Differential privacy for public health data: an innovative tool to optimize information sharing while protecting data confidentiality. Patterns 2, 100366 (2021).
Shen, Z. & Zhong, T. Analysis of application examples of differential privacy in deep learning. Comput. Intell. Neurosci. 2021, 4244040 (2021).
Nolte, D. et al. Federated learning framework integrating REFINED CNN and deep regression forests. Bioinform. Adv. 3, vbad036 (2023).
Zhang, F. et al. Secure and decentralized federated learning framework with non-IID data based on blockchain. Heliyon 10, e27176 (2024).
Biagioli, M. & Buning, M. Technologies of the law/ law as a technology”. Hist. Sci. 57, 3–17 (2019).
Qandeel, M. Facial recognition technology: regulations, rights and the rule of law. Front. Big Data 7, 1354659 (2024).
Bai Y., Guang X., Yeung R. W. Multiple linear-combination security network coding. Entropy 25, 1135 (2023).
Li, J., Ji, S. & Jiang, Y. Development of network security based on the neural network PSD algorithm. Comput. Intell. Neurosci. 2022, 9460985 (2022).
Wang Y., et al. Towards double defense network security based on multi-identifier network architecture. Sensors 22, 747 (2022).
Kou Z., et al. Identification of abnormal data for synchronous monitoring of transformer DC bias based on multiple criteria. Sensors 23, 4959 (2023).
Xue, H. et al. Abnormal data region discrimination and cross-monitoring points historical correlation repair of water intake data. Big Data 7, 99–113 (2019).
Spijkerboer, F. L., Overdyk, F. J. & Dahan, A. A machine learning algorithm for detecting abnormal patterns in continuous capnography and pulse oximetry monitoring. J. Clin. Monit. Comput 38, 915–925 (2024).
Gujral, H. et al. Design and implementation of a quantitative network health monitoring and recovery system. Wirel. Pers. Commun. 125, 367–397 (2022).
Sauer, K. et al. The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat. Rev. Microbiol. 20, 608–620 (2022).
Gilbert, P. et al. Intellectual property rights and vaccines. Methods Mol. Biol. 2412, 505–518 (2022).
Brown, W. M. Intellectual property. Methods Mol. Med. 40, 227–241 (2000).
Rake, B. Waiving intellectual property rights: boom or bust for medical innovation? Drug Discov. Today 27, 384–389 (2022).
Chisholm, O. & Critchley, H. Future directions in regulatory affairs. Front. Med. 9, 1082384 (2022).
Sneve M., Smith G. Reducing risks through regulatory cooperation: a review of bilateral regulatory cooperation between the Norwegian Radiation and Nuclear Safety Authority and corresponding authorities in countries of the former Soviet Union. J. Radiol. Prot. 45, 82 (2025).
Pejović, G. et al. Towards medicines regulatory authorities’ quality performance improvement: value for public health. Int J. Health Plan. Manag. 31, E22–E40 (2016).
Singhal, A. et al. Toward fairness, accountability, transparency, and ethics in ai for social media and health care: scoping review. JMIR Med Inf. 12, e50048 (2024).
Ueda, D. et al. Fairness of artificial intelligence in healthcare: review and recommendations. Jpn. J. Radio. 42, 3–15 (2024).
Inglada Galiana, L., Corral Gudino, L. & Miramontes González, P. Ethics and artificial intelligence. Rev. Clin. Esp. 224, 178–186 (2024).
Liefgreen, A. et al. Beyond ideals: why the (medical) AI industry needs to motivate behavioural change in line with fairness and transparency values, and how it can do it. AI Soc. 39, 2183–2199 (2024).
Van Der Velden, B. H. M. et al. Explainable artificial intelligence (XAI) in deep learning-based medical image analysis. Med. Image Anal. 79, 102470 (2022).
Toussaint, P. A. et al. Explainable artificial intelligence for omics data: a systematic mapping study. Brief Bioinform. 25, bbad453 (2023).
Ding, Q. et al. Explainable artificial intelligence in the field of drug research. Drug Des. Devel Ther. 19, 4501–4516 (2025).
Polimeni, M., Pasquier, C. & Lund, M. Virtual cell model for osmotic pressure calculation of charged biomolecules. J. Chem. Phys. 155, 194111 (2021).
Garny, A. et al. CellML and associated tools and techniques. Philos. Trans. A Math. Phys. Eng. Sci. 366, 3017–3043 (2008).
Douillet, A. & Ballet, P. A GPU algorithm for agent-based models to simulate the integration of cell membrane signals. Acta Biotheor. 68, 61–71 (2020).
Kobayashi, S. S. et al. Identification of myeloproliferative neoplasm drug agents via predictive simulation modeling: assessing responsiveness with micro-environment derived cytokines. Oncotarget 7, 35989–36001 (2016).
Waqas, A. et al. Revolutionizing digital pathology with the power of generative artificial intelligence and foundation models. Lab. Invest. 103, 100255 (2023).
Cascella, M. et al. The breakthrough of large language models release for medical applications: 1-year timeline and perspectives. J. Med. Syst. 48, 22 (2024).
Gu, Y. et al. A review of the development and challenges of cell mechanical models [J]. IEEE Trans. Nanobiosci. 22, 673–684 (2023).
Boycott K. M., et al. International collaborative actions and transparency to understand, diagnose, and develop therapies for rare diseases. EMBO Mol. Med. 11, e10486 (2019).
Keating et al. SBML Level 3: an extensible format for the exchange and reuse of biological models. Mol. Syst. Biol. 16, e9110 (2020).
Clerx M., et al. CellML 2.0. J. Integr. Bioinform. 17, 20200021 (2020).
Huang, K. et al. Artificial intelligence foundation for therapeutic science. Nat. Chem. Biol. 18, 1033–1036 (2022).
Luecken, M. D. et al. Defining and benchmarking open problems in single-cell analysis. Nat. Biotechnol. 43, 1035–1040 (2025).
Moher, D. et al. Increasing value and reducing waste in biomedical research: who’s listening? Lancet 387, 1573–1586 (2016).
Hildebrandt, M. G. et al. How to increase value and reduce waste in research: initial experiences of applying Lean thinking and visual management in research leadership. BMJ Open 12, e058179 (2022).
Acker, J. P. Editorial: advancing the cryopreservation of cells, tissues and organs using model biological systems. Cryobiology 117, 104975 (2024).
Holden, A. V. Development and application of human virtual excitable tissues and organs: from premature birth to sudden cardiac death. Alter. Lab Anim. 38, 87–99 (2010).
Qian, L., Dong, Z. & Guo, T. Grow AI virtual cells: three data pillars and closed-loop learning. Cell Res. 35, 319–321 (2025).
Runser, S., Vetter, R. & Iber, D. SimuCell3D: three-dimensional simulation of tissue mechanics with cell polarization. Nat. Comput Sci. 4, 299–309 (2024).
Kaczmarzyk et al. Explainable AI for computational pathology identifies model limitations and tissue biomarkers. ArXiv, 2024.
Sathyan A., Weinberg A. I., Cohen K. Interpretable AI for bio-medical applications. Complex Eng. Syst. 2, 18 (2022).
Auzine, M. M. et al. Development of an ensemble CNN model with explainable AI for the classification of gastrointestinal cancer. PLoS One 19, e0305628 (2024).
Walter, M., Webb, S. J. & Gillet, V. J. Interpreting neural network models for toxicity prediction by extracting learned chemical features. J. Chem. Inf. Model 64, 3670–3688 (2024).
Walker, D. C. & Southgate, J. The virtual cell—a candidate co-ordinator for ‘middle-out’ modelling of biological systems. Brief. Bioinform. 10, 450–461 (2009).
Storck, M. et al. [Carcinoid tumor of the stomach—aspects of surgical therapy]. Chirurg 62, 284–288 (1991).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No.82503242); the Natural Science Foundation of Hunan Province (2025JJ60596); and the China Postdoctoral Science Foundation (2025M772105).
Author information
Authors and Affiliations
Contributions
Author M.C.Y.: Conceptualization; Formal analysis; Visualization (overall design); Writing—original draft; Writing—review & editing (lead). Author Z.H.: Visualization (figures); Writing—review & editing. Author R.Y.W.: Visualization (figures); Writing—review & editing. Author J.X.Y.: Data curation; Writing—review & editing.Author L.B.H.: Writing—review & editing (language). Author S.Z.K.: Literature Collection and Management; Writing—review & editing. Author S.Z.Y.: Literature Collection and Management. Author G.Y.: Literature Collection and Management. Author C.Y.H.: Literature Collection and Management. Author L.X.Y.: Literature Collection and Management. Author L.Z.D.: Supervision; Writing—review & editing (critical revision and final approval).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All authors consent for publication.
Declaration of generative AI and AI-assisted technologies in the writing process
During the preparation of this work, the authors used ChatGPT-5Auto for language polishing. After using this tool, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, C., Zhang, H., Rao, Y. et al. AI-driven virtual cell models in preclinical research: technical pathways, validation mechanisms, and clinical translation potential. npj Digit. Med. 9, 25 (2026). https://doi.org/10.1038/s41746-025-02198-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02198-6