Abstract
Radiologists can miss subtle secondary findings relevant to tumor staging and management. Large reasoning models (LRMs) may mitigate this by improving interpretive completeness and transparency, yet systematic evaluation remains lacking. We studied 900 multicenter oncologic cases from three Chinese hospitals to compare an LRM’s reasoning processes with its conclusion-only format and two non-reasoning models. Three senior radiologists assessed diagnostic errors and qualitative attributes, and a human-in-the-loop study with six radiologists evaluated workflow-related effects. Cross-language generalization was tested using an English MIMIC-Cancer-90 cohort. Reasoning processes showed the fewest missed or misclassified errors and the highest ratings for comprehensiveness, explainability, and unbiasedness, though with reduced conciseness. Performance dropped when only conclusions were used, and non-reasoning models underperformed across metrics. Improvements were consistent across cancer types, modalities, institutions, and languages. Reader studies confirmed greater perceived completeness and reasoning clarity, especially among juniors, while revealing workflow costs requiring optimization for clinical use.
Similar content being viewed by others
Introduction
Formulating clinically sound impressions from imaging findings is a central yet demanding component of radiologic practice1. Whereas findings describe the radiologist’s direct observations and convey elements of interpretive reasoning, the impression requires a deliberate synthesis that prioritizes, contextualizes, and integrates these details2. Currently, impressions are drafted manually by radiologists, which is inherently subjective and vulnerable to omissions and misdiagnoses3,4,5. These errors are far from rare, with studies estimating an everyday error or discrepancy rate of 3–5% in radiology practice6. Particularly, missing subtle secondary findings (e.g., small metastases) is a documented challenge7, and such omissions in oncologic imaging could lead to under-staging or suboptimal treatment planning. At the same time, impression generation represents the remaining bottleneck in the pipeline of AI report generation, as current AI techniques for identifying imaging findings have already reached a relatively mature stage8,9. These challenges underscore a need not only for automation but for approaches that preserve interpretive transparency and ensure information completeness for impression generation to support diagnostic reliability and trust in AI-assisted reporting.
Recent advances in large language models (LLMs) have enabled partial automation of radiology reporting10,11,12. Researchers have evaluated the performance of LLM on various radiology-related tasks, including report simplification13, error correction14,15, and impression generation1,16,17,18. Sun et al.1 assessed the performance of GPT-4 in generating diagnostic impressions from chest radiograph findings. In a study involving 50 radiology reports, the impressions generated by GPT-4 were found to be inferior to those written by human radiologists due to including statements not mentioned in the original findings. Zhang et al.17 developed a specialized LLM for radiology impression generation. On an external test set (n = 3988) encompassing various imaging modalities and anatomical sites, the LLM was generally able to generate both linguistically and professionally appropriate impressions. However, its performance in generating specific diagnoses remained suboptimal. While these studies highlight the considerable advances and clinical potential of LLMs in radiology, most LLMs generate conclusions without explaining their underlying reasoning. The absence of transparency limits clinical trust19, as radiologists cannot verify how an impression was derived. Moreover, purely conclusion-based outputs may omit subtle but clinically significant findings, reducing the reliability of the generated impressions.
The emergence of OpenAI o1-like20 large reasoning models (LRMs)21 has marked a paradigm shift in LLM development, i.e., from the train-time compute to the test-time compute that enhances performance by enabling the LLM to engage in more “thinking” during inference22. Taking the pioneering open-source LRM DeepSeek-R123 as an example, thanks to the chain-of-thought (CoT)24 and reinforcement learning (RL)25 techniques, it demonstrates exceptional reasoning capability, featuring outputting native and explicit reasoning processes. More importantly, the model enables self-reflection in the reasoning processes that can greatly improve the generative performance, particularly in solving mathematical and programming tasks26. The advancement in reasoning positions it to potentially overcome limitations observed in non-reasoning LLMs like GPT-4, enabling more accurate medical diagnoses through deeper and more deliberate reasoning. Tordjman et al.27 have conducted a comprehensive evaluation of DeepSeek-R1 with OpenAI-o1 and Llama3.1-405B28 on radiological impression generation with 200 cases. DeepSeek-R1 achieved competitive impression quality compared to OpenAI-o1 (five-point Likert score: 4.5 vs 4.8). Sandmann et al.29 compared the clinical decision-making capability of DeepSeek-R1 against proprietary LLMs (GPT-4o and Gem2FTE) on 125 cases. DeepSeek-R1 performed equally well and better than proprietary LLMs in some cases. Collectively, these studies verified the advancement and potential of LRM in the medical field. Nevertheless, they have primarily evaluated the quality of the final reasoning-derived conclusions, such as diagnostic accuracy or linguistic quality, rather than systematically assessing the effect of the reasoning processes themselves. Understanding how these reasoning processes influence diagnostic completeness, interpretability, and clinical usability is essential to determine whether such models can truly enhance radiological impression generation and align with clinical practice needs.
Building upon these advances, this study aimed to systematically evaluate the effect of the reasoning processes generated by an LRM on radiological impression generation for oncologic imaging. Using the open-source model DeepSeek-R1 as a representative LRM, we compared the diagnostic, qualitative, and workflow-related performance of reasoning-based outputs with both the model’s own conclusions and the outputs of two non-reasoning LLMs across diverse cancer types, imaging modalities, institutions, and languages. By focusing on the reasoning processes themselves rather than solely the final reasoning-derived conclusions, this work sought to determine whether explicit reasoning could enhance diagnostic completeness, interpretability, and clinical reliability.
Results
Study overview
A total of 990 oncologic cases were analyzed across breast, lung, and colorectal cancers, covering three imaging modalities of CT, MRI, and mammography (MG). We compared the two components of DeepSeek-R1 outputs: (i) DeepSeek-R1 (Rea.): the stepwise reasoning processes, and (ii) DeepSeek-R1 (Con.): the subsequent conclusion derived from the reasoning processes. In addition, DeepSeek-R1 (Rea.) was also compared against two non-reasoning LLMs (DeepSeek-V3_0324 and GPT-4.5). Model outputs were evaluated across eight diagnostic and qualitative metrics. The diagnostic metrics were designed to target four kinds of common errors in oncologic radiology, including missed primary diagnoses (MPD), missed secondary diagnoses (MSD), primary misdiagnoses (PMisD), and secondary misdiagnoses (SMisD). The qualitative metrics are comprehensiveness, explainability, conciseness, and unbiasedness. Additionally, a reader study on 54 Chinese cases involving three junior (<10 years) and three senior (≥10 years) radiologists assessed the information completeness, reasoning helpfulness, and short-term editability of different model outputs. The evaluation time on each case in the reader study was also analyzed.
Dataset
Oncologic cases (n = 900) from three Chinese medical institutions were included for model evaluation. Table 1 summarizes the demographic and clinical characteristics of the 900 Chinese cases, with each institution contributing 300 cases (150 breast, 50 lung, and 100 colorectal cancer cases). The distribution of age, sex, histological subtypes, and imaging modalities was largely similar across the three institutions. This stratified multicenter design ensured balanced case numbers across cancer types and modalities, enabling consistent subgroup analyses. To assess cross-language generalization, an additional English-language cohort consisting of 90 cases (MIMIC-Cancer-90) was constructed and evaluated. Details of the MIMIC-Cancer-90 dataset are in the “Data” section of the “Methods”.
Reasoning processes versus the conclusion only
To assess the contribution of the reasoning processes themselves, we first compared DeepSeek-R1 (Rea.) with DeepSeek-R1 (Con.). Overall performance on the Chinese reports (n = 900) of DeepSeek-R1 (Rea.) and that of DeepSeek-R1 (Con.) is illustrated in Fig. 1. As shown in Fig. 1a, in terms of the four diagnostic metrics, DeepSeek-R1 (Rea.) achieved significant superiority on DeepSeek-R1 (Con.): 0.67% (6 cases; 95% CI: 0.31%–1.45%) versus 5.56% (50 cases, 95% CI: 4.24%–7.25%) on MPD, 3.22% (29 cases; 95% CI: 2.25%–4.59%) versus 23.11% (208 cases, 95% CI: 20.47%–25.98%) on MSD, 0.56% (5 cases; 95% CI: 0.24%–1.29%) versus 4.78% (43 cases, 95% CI: 3.57%–6.37%) on PMisD, and 6.11% (55 cases; 95% CI: 4.72%–7.87%) versus 11.33% (102 cases, 95% CI: 9.42%–13.57%) on SMisD (all p < 0.001).
a Bar charts of diagnostic metric comparison. Blue bars represent DeepSeek-R1 (Rea.) and orange bars represent DeepSeek-R1 (Con.). b Box plots of qualitative metric comparison. Blue boxplots represent DeepSeek-R1 (Rea.) and yellow boxplots represent DeepSeek-R1 (Con.). ×: mean values. Lines in the boxes: median values. Whiskers: 1.5×IQR. Circles outside the boxes: outliers.
As shown in Fig. 1b, DeepSeek-R1 (Rea.) demonstrated consistent and statistically significant advantages over DeepSeek-R1 (Con.) in three of the four qualitative evaluation metrics. For comprehensiveness, DeepSeek-R1 (Rea.) achieved higher scores (Median = 4.333, IQR = [4.333–4.667]) than DeepSeek-R1 (Con.) (Median = 4.000, IQR = [3.667–4.333]) (p < 0.001, Cohen’s d = 0.784), indicating more complete and detailed coverage of key diagnostic elements. In explainability, DeepSeek-R1 (Rea.) also outperformed DeepSeek-R1 (Con.) (Median = 4.667, IQR = [4.333–5.000] vs. 4.333, IQR = [4.000–4.667]; p < 0.001, Cohen’s d = 0.836), reflecting greater interpretive transparency and logical clarity in the reasoning processes. For unbiasedness, both models maintained high performance, with DeepSeek-R1 (Rea.) showing a modest but statistically significant advantage (Median = 4.333, IQR = [4.333–5.000] vs. 4.333, IQR = [3.833–4.667]; p < 0.001, Cohen’s d = 0.670), suggesting that explicit reasoning did not introduce systematic bias and may even improve reporting consistency. In contrast, conciseness exhibited an opposite trend. DeepSeek-R1 (Rea.) generated considerably longer and more detailed outputs (Median = 2.667, IQR = [2.333–3.000]) compared with DeepSeek-R1 (Con.) (Median = 5.000, IQR = [4.333–5.000]) (p < 0.001, Cohen’s d = 4.673), representing an extremely large effect size. Gwet’s AC1 was used to assess inter-rater consistency across the four qualitative scores from three raters. High agreement was observed for most metrics (all > 0.69), whereas conciseness showed only moderate agreement (0.468) (Table S1).
Overall, these findings indicate that explicit reasoning improves the comprehensiveness and interpretability of oncologic diagnoses, particularly in capturing secondary findings relevant to cancer staging and potentially informing treatment decisions, albeit with a corresponding loss of conciseness. In Tables S2–S5, we illustrate two representative cases, where DeepSeek-R1 (Con.) failed to point out the tumor-related secondary lesions (MSD). In contrast, DeepSeek-R1 (Rea.) mentioned all of these.
Subgroup analyses further confirmed that the benefits of the reasoning processes were consistent across cancer types, imaging modalities, and institutions (Fig. 2). On all subsets, DeepSeek-R1 (Rea.) achieved higher scores than DeepSeek-R1 (Con.) on all four diagnostic metrics and three qualitative metrics except conciseness. For diagnostic metrics, DeepSeek-R1 (Rea.) achieved notably lower MPD on subsets of breast cancer, MRI, and Institution 2, MSD on all subsets, PMisD on subsets of breast cancer, MRI, and Institution 1, and SMisD on subsets of all three cancers, imaging modalities of CT, MRI, Institution 1, and Institution 2. In terms of generation quality, the improvements were particularly pronounced in three dimensions except conciseness, with DeepSeek-R1 (Rea.) outperforming DeepSeek-R1 (Con.) in nearly all subgroups. This consistency across diverse clinical contexts suggests that the advantage of explicit reasoning is robust to variations in disease type and imaging modality.
a Breast cancer (n = 450). b Lung cancer (n = 150). c Colorectal cancer (n = 300). d CT (n = 450). e MG (n = 150). f MRI (n = 300). g Institution 1 (n = 300). h Institution 2 (n = 300). i Institution 3 (n = 300). Blue lines represent DeepSeek-R1 (Rea.) condition and yellow lines represent DeepSeek-R1 (Con.). For the diagnostic metrics, the illustrated score was computed with 1-error rate. For the qualitative metrics, the score was calculated by dividing the mean score by 5. The asterisk in parentheses denotes the significance level.
Reasoning processes versus non-reasoning models
To contextualize the effect of reasoning within the broader landscape of LLM performance, DeepSeek-R1 (Rea.) was further compared with two non-reasoning models (DeepSeek-V3_0324 and GPT-4.5). As illustrated in Fig. 3a, significant diagnostic gains were achieved by DeepSeek-R1 (Rea.) compared with the two LLMs on the whole 900 cases in terms of MPD, MSD, and SMisD (all p < 0.01). Particularly, for MSD and SMisD, the error rates of DeepSeek-V3_0324 and GPT-4.5 are much higher. DeepSeek-V3_0324 achieves 17.78% (160 cases, 95% CI: 15.42%–20.41%), and GPT-4.5 achieves 7.89% (71 cases, 95% CI: 6.30%–9.83%) regarding MSD. For SMisD, DeepSeek-V3_0324 achieves 14.11% (127 cases, 95% CI: 11.99%–16.54%) and GPT-4.5 achieves 18.33% (165 cases, 95% CI: 15.94%–20.99%).
a Bar charts of diagnostic metric comparison. Blue bars represent DeepSeek-R1 (Rea.), light green bars represent DeepSeek-V3_0324, and pink bars represent GPT-4.5. b Box plots of qualitative metric comparison. Dark blue boxplots represent DeepSeek-R1 (Rea.), light green boxplots represent DeepSeek-V3_0324, and pink boxplots represent GPT-4.5. ×: mean values. Lines in the boxes: median values. Whiskers: 1.5×IQR. Circles outside the boxes: outliers.
As shown in Fig. 3b, DeepSeek-R1 (Rea.) also achieved significant superiority in most comparisons in terms of generation quality. For explainability, DeepSeek-R1 (Rea.) achieved the highest ratings, showing statistically significant improvements compared with both DeepSeek-V3_0324 and GPT-4.5 (p < 0.001, Cohen’s d = 0.51–0.63). The magnitude of these effects indicates that explicit reasoning contributes substantially to enhancing the interpretability and clarity of the model output. Comprehensiveness was similarly robust, significantly higher than DeepSeek-V3_0324 (p < 0.001, Cohen’s d = 0.33) and comparable to GPT-4.5. DeepSeek-R1 (Rea.) still scored lowest on conciseness, producing substantially longer outputs than DeepSeek-V3_0324 (5.000, IQR = [4.333–5.000]) and GPT-4.5 (4.333, IQR = [4.333–5.000]) (p < 0.001, Cohen’s d = 3.65–4.24). Unbiasedness remained consistently high across models, DeepSeek-R1 (Rea.) showing moderate but statistically significant advantages over DeepSeek-V3_0324 and GPT-4.5 (p < 0.001, Cohen’s d = 0.47–0.48). Inter-rater consistency for all metrics remained high for both DeepSeek-V3_0324 and GPT-4.5 (all AC1 > 0.797) (Table S1).
These findings indicate that explicit reasoning not only improves internal consistency relative to the model’s own conclusions but also yields diagnostic advantages over conventional non-reasoning architectures, particularly in identifying secondary lesions.
As illustrated in Fig. 4, model comparison on the nine subsets between DeepSeek-R1 (Rea.) and two non-reasoning LLMs was also conducted. In general, DeepSeek-R1 (Rea.) outperforms the DeepSeek-V3_0324 and GPT-4.5 across most axes in the radar chart, independent of cancer type, imaging modality, and institution, underscoring a consistent and extensive advantage of leveraging the reasoning processes of LRM across diverse medical settings.
a Breast cancer (n = 450). b Lung cancer (n = 150). c Colorectal cancer (n = 300). d CT (n = 450). e MG (n = 150). f MRI (n = 300). g Institution 1 (n = 300). h Institution 2 (n = 300). i Institution 3 (n = 300). Blue lines represent DeepSeek-R1 (Rea.), green lines represent DeepSeek-V3_0324, and red lines represent GPT-4.5. For the diagnostics metrics, the illustrated score is computed as (1 - error rate). For the qualitative metrics, the score is calculated by dividing the mean score by 5. Green and red asterisks in parentheses denote the significance level between DeepSeek-R1 and DeepSeek-V3_0324 and DeepSeek-R1 and GPT-4.5, respectively.
Non-Chinese validation on MIMIC-Cancer-90
As shown in Fig. 5, DeepSeek-R1 (Rea.) consistently outperforms DeepSeek-R1 (Con.) on the English reports from MIMIC-Cancer-90 across both diagnostic and qualitative metrics. Notably, it achieves a significantly lower MSD error rate (p < 0.001) and substantially higher scores in comprehensiveness (p < 0.001, Cohen’s d = 1.398) and explainability (p < 0.001, Cohen’s d = 1.035). The large effect sizes indicated by Cohen’s d highlight that these improvements are not only statistically significant but also of considerable practical magnitude. These patterns closely mirror the results obtained from the 900 Chinese reports, reinforcing that DeepSeek-R1’s explicit reasoning processes provide more complete and interpretable outputs across languages.
a Bar charts of diagnostic metric comparison. Blue bars represent DeepSeek-R1 (Rea.) and orange bars represent DeepSeek-R1 (Con.). b Box plots of qualitative metric comparison. Dark blue boxplots represent DeepSeek-R1 (Rea.) and yellow boxplots represent DeepSeek-R1 (Con.). ×: mean values. Lines in the boxes: median values. Whiskers: 1.5×IQR. Circles outside the boxes: outliers.
Residual errors
Despite the overall improvements, several diagnostic errors persisted in the reasoning-based impressions. As illustrated in Fig. 6, the top eight reasons for the failure of DeepSeek-R1 (Rea.) are listed. Misclassifications were dominated by over-calling benign entities as metastatic disease: most commonly hepatic hemangioma labelled as liver metastasis (n = 6), followed by bilateral scattered pulmonary nodules (n = 4) and multiple hepatic cysts (n = 4) read as metastases. A second cluster reflected under-recognition of tumor-associated findings, including obstructive pneumonia (n = 3), pectoral muscle invasion in breast cancer (n = 3), and nodal metastasis (n = 3). A smaller fraction involved within-organ misclassification, such as peripheral lung cancer called central type (n = 2) and a breast cancer mass reported as benign (n = 3). These findings underscore that reasoning transparency does not fully prevent diagnostic errors and highlight the need to ensure alignment between reasoning quality and final conclusions.
→ : The disease before the arrow was misdiagnosed as the one after the arrow.
Concluding failure
The observed performance gap between DeepSeek-R1’s reasoning and conclusion outputs reveals an important reliability issue, termed here as concluding failure. This phenomenon occurs when the model’s reasoning is diagnostically correct, but the final conclusion contradicts it. Concluding failures may cause serious clinical harm. Such plausible but defective outputs may directly misguide image interpretation and clinical decision-making, while concealing critical errors that are difficult to detect under workload pressure, potentially resulting in delayed diagnoses or inappropriate treatments. They can also foster automation bias, reducing radiologists’ willingness to independently analyze and question results, and, over time, introduce systematic biases that compromise diagnostic quality and consistency. Due to their high surface credibility, these errors are less likely to be identified during routine review. To analyze the level of concluding failure, we calculated the concluding failure rates of DeepSeek-R1 on four diagnostic metrics, defined as the number of concluding failures divided by the corresponding number of cases. As shown in Fig. 7, DeepSeek-R1 achieved concluding failure rates of 4.67%, 19.33%, 3.44%, and 7.33% in terms of MPD, MSD, PMisD, and SMisD, respectively. Moreover, the concluding failures of DeepSeek-R1 are prevalent on all subsets, and particularly on secondary diagnoses (MSD and SMisD). Quantifying this discrepancy provides an objective measure of the reliability gap between reasoning and conclusion generation, offering an important diagnostic signal for evaluating future reasoning-enabled models. Two examples of the concluding failure can be found in Tables S2–S5.
Failure rates for MPD, MSD, PMisD, and SMisD are compared across cancer types, imaging modalities, and institutions. Dark blue bars represent MPD, teal bars represent MSD, light blue bars represent PMisD, and orange-red bars represent SMisD.
Human-in-the-loop reader study
Across all six radiologists, DeepSeek-R1 (Rea.) consistently improved information completeness (Fig. 8a) and reasoning helpfulness compared with DeepSeek-R1 (Con.) (Fig. 8b). For information completeness, the proportion of top-tier scores (level 3: “sufficient and clinically sound”) was significantly higher for the reasoning condition in five of six readers (all p < 0.05). Similarly, reasoning helpfulness was rated significantly higher by four readers (all p < 0.05), and the higher mean scores of DeepSeek-R1 (Rea.) were achieved for all readers. In contrast, the short-term editability scores favored the conclusion-only outputs in four readers (all p < 0.05), suggesting that the more concise DeepSeek-R1 (Con.) required less effort to refine into deliverable clinical impressions (Fig. 8c). For the evaluation time, DeepSeek-R1 (Rea.) required significantly longer per-case reading time in four readers (all p < 0.05), while two readers showed comparable efficiency between the two versions (Fig. 8d).
a Stacked column chart of the information completeness. For each radiologist, the left bar represents the DeepSeek-R1 (Rea.) and the right bar represents the DeepSeek-R1 (Con.). Within each bar, dark blue segments correspond to rating level 3, medium blue segments to rating level 2, and light blue segments to rating level 1. Numbers in the column are the cases with the highest score. b Bar charts of the reasoning helpfulness. Blue boxplots represent DeepSeek-R1 (Rea.) and orange boxplots represent DeepSeek-R1 (Con.). c Stacked column chart of the short-term editability. The left bar represents the DeepSeek-R1 (Rea.) and the right bar represents the DeepSeek-R1 (Con.). Within each bar, dark lavender segments correspond to rating level 3, medium lavender segments to rating level 2, and light lavender segments to rating level 1. d Box plots of the evaluation time. Blue boxplots represent DeepSeek-R1 (Rea.) and orange boxplots represent DeepSeek-R1 (Con.). For the evaluation time, data were preprocessed by removing outliers on a paired basis. Outliers were identified using the IQR rule applied to the paired differences; any pair with a difference outside the range [Q1 − 1.5×IQR, Q3 + 1.5×IQR] was excluded. ×: mean values. Lines in the boxes: median values.
When stratified by experience, radiologists 1–3 (junior, <10 years) exhibited larger performance gaps between DeepSeek-R1 (Rea.) and DeepSeek-R1 (Con.), with greater gains in completeness and reasoning helpfulness but also greater time costs. By contrast, radiologists 4–6 (senior, ≥10 years) demonstrated more stable performance across the two conditions, maintaining comparable ratings across all dimensions in general. These results indicate that explicit reasoning generally enhances diagnostic completeness and interpretability but imposes a measurable efficiency cost, particularly for less-experienced readers.
Discussion
This study systematically investigated the effect of the reasoning processes generated by an LRM on radiological impression generation. Across cancer types, imaging modalities, institutions, and report languages, the reasoning-based outputs of DeepSeek-R1 consistently improved diagnostic completeness and interpretive quality compared with both its own conclusion-only outputs and two non-reasoning LLMs. These findings highlight that the diagnostic benefit of LRMs arises not merely from more powerful language generation, but also from the reasoning processes themselves.
In the human-in-the-loop reader study, reasoning-based outputs consistently achieved higher scores in information completeness and reasoning helpfulness, further confirming that explicit reasoning enhanced diagnostic transparency and reduced omission of key findings. These benefits, however, came at a workflow cost, as longer reasoning outputs required more time for review and editing, particularly among junior readers. Conclusion-only texts were rated as more readily editable within a short time frame and required shorter reading durations. In contrast, senior readers maintained comparable efficiency between reasoning and conclusion outputs, suggesting that diagnostic experience mitigates the cognitive load of interpreting explicit reasoning. These results highlight a quantifiable trade-off between interpretability and workflow efficiency and indicate that reasoning transparency may be particularly valuable for junior radiologists as an assistive or educational tool, whereas experienced readers may prefer condensed reasoning summaries for routine clinical deployment.
The clinical significance of this improvement is multifold. First, the substantial reductions in secondary errors (particularly in MSD) mean that explicit reasoning could help capture subtle but clinically decisive information that is relevant to staging, treatment planning, and prognostic evaluation. Second, the reasoning transparency has the potential to provide radiologists with a verifiable decision pathway, enabling them to audit model logic and reconcile automated impressions with human judgment. This interpretive traceability may add additional value for multidisciplinary tumor boards and institutional quality assurance, where the rationale behind diagnostic statements must be transparent and defensible. Collectively, these results demonstrate that reasoning transparency makes a functional advantage that may strengthen diagnostic reliability and trust in AI-assisted reporting.
Despite the advantages, using the reasoning processes alone introduces several practical and methodological challenges. The first challenge concerns workflow efficiency and conciseness. The human-in-the-loop study revealed that explicit reasoning, while improving diagnostic completeness and transparency, increased reading and editing time for most radiologists, especially for those with less experience. DeepSeek-R1 produced substantially longer reasoning chains than conventional impressions, reflecting a tendency toward overthinking30. Although detailed reasoning enhances analytical coverage, excessive verbosity may impose cognitive burden and limit real-world usability. Methods for reasoning-length control31 and adaptive thinking32 should be explored to achieve a balance between completeness and efficiency.
The second challenge involves residual diagnostic errors. While reasoning reduced major failure modes, it did not fully eliminate confusion between benign and metastatic findings or under-recognition of tumor-associated features. These errors underscore that explicit reasoning alone cannot compensate for missing domain priors or insufficient spatial understanding. Integrating radiology-specific knowledge33 and structured spatial reasoning mechanisms34 could further improve accuracy in complex oncologic scenarios.
The third challenge relates to reasoning–conclusion misalignment. The phenomenon of concluding failure, where a model reasons correctly but produces an incorrect final statement, highlights a gap between analytical reasoning and generative summarization. The underlying reasons could lie in the RL reward misalignment and the positional bias. First, models such as DeepSeek-R1 were optimized using Reinforcement Learning with Verifiable Rewards (RLVR) with rule-based rewards primarily designed for mathematical and programming tasks23, which do not align with the demands of oncologic radiology that requires the integration of imaging findings, the identification of key abnormalities, and logical diagnostic synthesis and inference2,35. With respect to this, the reasoning corpus used for RLVR largely consists of non-radiology texts, further resulting in substantial domain-knowledge gaps that may impair diagnostic reasoning. Second, LLMs exhibit positional bias36, i.e., they tend to under-attend to information appearing in the middle of long inputs due to the inherent limitation of the attention mechanism37, leading to the omission of clinically relevant details (e.g., the obstructive inflammation in the given lung example [Table S2]) when generating conclusions.
The above analysis could also be implied by the superiority of the two non-reasoning models (DeepSeek-V3_0324 and GPT-4.5) over DeepSeek-R1 (Con.) on MSD and comprehensiveness. Although being all conclusion-only, the non-reasoning models generate outputs directly, suffering no effect from the RL reward misalignment and the positional bias resulting from reasoning training. However, as DeepSeek-R1 (Rea.) is more comprehensive due to its better exploratory ability arising from RL training38, it outperforms the non-reasoning models.
In clinical use, radiologists would either need to manually synthesize impressions from the reasoning text, risking inconsistency and time inefficiency, or rely on the model’s conclusion, risking the concluding failures of LRMs and the suboptimal performance of non-reasoning LLMs. Future LRMs can explore radiology-aligned optimization39, focused prompting36, and self-refinement mechanisms40 to ensure that the final outputs remain faithful to their underlying reasoning.
From a methodological perspective, this study demonstrates a new paradigm for evaluating reasoning-enabled models: not only measuring performance by final outcomes, but also analyzing the process-level behavior that leads to those outcomes. This approach aligns with the growing emphasis on explainability and safety in medical AI evaluation frameworks8. By quantifying the diagnostic effect of reasoning itself, this study provides an analytic foundation for auditing reasoning-based systems before clinical deployment. From a translational standpoint, explicit reasoning can serve as a bridge between human and artificial decision-making. Reasoning traces could be selectively surfaced in reporting systems as “auditable evidence,” allowing radiologists to inspect how the model integrated findings, cross-check against raw imaging data, and flag potential inconsistencies. Such integration could improve reporting transparency, training of junior readers, and consistency in multidisciplinary discussions. Ultimately, reasoning-based systems may evolve from report generators to interactive diagnostic assistants that augment, rather than replace, radiologists’ analytical workflows.
This study has several limitations. First, all evaluations were retrospective. Although we conducted a human-in-the-loop reader study to assess interpretability and usability, the work does not constitute a real-time or prospective deployment. Future studies should incorporate prospective clinical testing, workflow integration, and real-world assessments of reporting efficiency and user satisfaction. Moreover, the observed increase in reading and editing time for reasoning-based outputs indicates a non-negligible workflow cost that needs to be addressed before large-scale clinical adoption. Second, as DeepSeek-R1 is a representative reasoning-capable model, and given that the latest LRMs such as Qwen341 and Kimi K2 Thinking42, which also rely on RL and the attention mechanism, may be similarly influenced by RL reward misalignment and positional bias, the findings of this study are more likely to reflect a reasoning-paradigm effect rather than a model-specific phenomenon. Nevertheless, future work will extend the analysis to additional LRMs to delineate the contributions of different reasoning processes. In addition, the present study focused exclusively on the findings-to-impression step and did not evaluate end-to-end image interpretation or multimodal vision–language systems, so the results should be interpreted as evidence about impression generation from textual findings rather than full image-level diagnostic performance. Finally, future research should also focus on reasoning conciseness control, radiology-specific alignment, and verification mechanisms to ensure that reasoning-enabled systems can deliver concise, accurate, and trustworthy support for radiologic decision-making.
In conclusion, this study shows that explicit reasoning in an LRM improves the completeness and interpretability of radiological impressions compared with conclusion-only generation. These benefits were consistent across Chinese-language and English-language cohorts and were further supported by a human-in-the-loop reader study demonstrating clearer diagnostic logic but also a measurable increase in reading and editing time. The findings highlight both the potential value and the practical constraints of integrating reasoning-based systems into clinical workflows, including challenges related to reasoning–conclusion alignment and residual secondary errors. Future work should focus on radiology-specific alignment, verification mechanisms, and workflow-aware optimization to enable reliable and efficient deployment of reasoning-enabled models in clinical practice.
Methods
The Ethics Committee of The Second Xiangya Hospital approved this retrospective study (protocol code 2022JJ20089) and waived informed consent, as the information prior to the routine reporting process does not pose any risk to patients.
Study design
As illustrated in Fig. 9, this study was designed to investigate the diagnostic, interpretive, and workflow-related effects of the reasoning processes generated by DeepSeek-R1 (version: 2025-01-20) during radiological impression generation. DeepSeek-R1 was selected as the representative LRM in this study because it provides open access to both model weights and reasoning traces, and adopts an RL post-training paradigm comparable to other frontier reasoning systems (e.g., OpenAI-o1 and Qwen341). This selection enables a transparent evaluation of the explicit reasoning processes of LRMs and broader clinical application. Imaging findings extracted from radiology reports of cancer cases were input into DeepSeek-R1 to generate outputs. The original radiologist-written impressions that have undergone two rounds of quality control at their respective institutions served as the reference standard for evaluation. To study the effect of the reasoning processes, the same prompt was also input into two state-of-the-art non-reasoning LLMs to generate impressions: the proprietary GPT-4.5 (version: 2025-02-07) and the open-source DeepSeek-V3_0324 (version: 2025-03-24).
A total of 990 radiology reports of cancer cases were collected from three Chinese medical institutions and one public English corpus, with the imaging findings extracted and input into the open-source LRM DeepSeek-R1 to generate the reasoning processes (DeepSeek-R1 (Rea.)) and final conclusion (DeepSeek-R1 (Con.)). The performance of DeepSeek-R1 (Rea.) was then compared with DeepSeek-R1 (Con.) and two state-of-the-art non-reasoning LLMs (DeepSeek-V3_0324 and GPT-4.5) by three senior radiologists on diagnostic, qualitative, and workflow-related metrics. Human-in-the-loop evaluations were also conducted beyond model-level comparison.
DeepSeek-R1 outputs consist of two structured segments: (i) a reasoning segment enclosed by <think> and </think> tags, representing the model’s explicit reasoning processes (DeepSeek-R1 (Rea.)); and (ii) a conclusion segment that follows, representing the model’s final impression (DeepSeek-R1 (Con.)). To specifically examine the independent effect of the reasoning processes, these two segments were separately extracted and analyzed. For DeepSeek-V3_0324 and GPT-4.5, the whole generated outputs were used as the impression. The three LLMs were accessed through their official application programming interface (API). Senior radiologists independently reviewed the LLM-generated outputs and evaluated them with four diagnostic metrics and four qualitative metrics. Specific definitions of these metrics are illustrated in Table 2.
Specifically, MPD refers to cases in which the primary malignant tumor is present but is entirely unrecognized or omitted from the model outputs. MSD denotes cases in which tumor-related secondary lesions, such as lymph node metastases, adjacent tissue invasion, or distant organ metastases, are present but entirely unrecognized or omitted. PMisD means that the lesions within the affected organ are recognized but misclassified with respect to malignancy status (benign versus malignant), malignant subtype, or location. This category includes (i) coexisting benign lesions misclassified as malignant (e.g., pulmonary hamartoma diagnosed as lung cancer), (ii) the primary malignant tumor misclassified as benign (e.g., breast cancer mass reported as benign), and (iii) the primary malignant tumor misclassified as another malignant subtype or location (e.g., peripheral lung cancer diagnosed as central type). It excludes non-recognition (which belongs to MPD/MSD). SMisD refer to cases in which benign lesions in other organs or anatomic sites outside the primary organ are recognized but misclassified as malignant tumors or metastases. (e.g., hepatic cysts or hemangiomas misdiagnosed as liver metastases). Errors not falling into these four predefined categories were beyond the scope of this study. Notably, for DeepSeek-R1 (Rea.), a diagnosis is credited only if it is (i) identified consistently at key reasoning stages, (ii) explicitly justified via a traceable chain of reasoning (linking imaging findings to hypotheses to diagnosis), and (iii) coherently integrated across the full reasoning trajectory. On the other hand, if the reasoning merely mentions a diagnosis intermittently and subsequently builds integration on incorrect features, it is classified as a misdiagnosis; if the reasoning fails to provide a continuous, traceable description of that diagnosis (even if mentioned), it is classified as a missed diagnosis. This approach is supported by empirical evidence showing that diagnostic justification, which requires a coherent and traceable reasoning path linking data through hypotheses to a final diagnosis, is associated with improved diagnostic accuracy and fewer reasoning errors in clinical reasoning assessments43.
A human-in-the-loop reader study was also conducted to assess the clinical interpretability and workflow usability of the reasoning processes. Six radiologists participated, including three senior radiologists (≥10 years of independent reporting experience) and three junior radiologists (<10 years of experience). A total of 54 cases were independently evaluated by each reader, comprising 18 cases per institution. For each institution, three cancer types and three imaging modalities were included, with three representative cases randomly selected for each cancer or modality. Readers rated the outputs of DeepSeek-R1 (Rea.) and DeepSeek-R1 (Con.) according to three criteria: (A) information completeness, (B) reasoning helpfulness, and (C) short-term editability. Definitions of the three criteria are illustrated in Table 3. Reader evaluation time was also recorded for efficiency analysis.
Evaluation procedure
The evaluation was designed to assess whether the reasoning processes themselves contribute to improved diagnostic completeness, interpretability, and workflow efficiency beyond the final conclusion. The diagnostic metrics quantified the impact of reasoning on error reduction across four common diagnostic failure modes in oncologic radiology. Specifically, two senior radiologists (W.Z., 10 years of experience; C.Z., 13 years of experience) independently evaluated the DeepSeek-R1 (Rea.) and DeepSeek-R1 (Con.), along with the generated impressions from DeepSeek-V3_0324 and GPT-4.5. The numbers of missed diagnoses and misdiagnoses were recorded at the case level, i.e., a binary classification (1 = exist, 0 = not exist) for each case was conducted on four error types. In cases where the two radiologists disagreed in their assessments, a third senior radiologist (J.L., 25 years of experience) reviewed the case, and the result was taken as the final decision.
The qualitative metrics were used to assess whether explicit reasoning improved the interpretive quality of generated impressions in terms of comprehensiveness, explainability, conciseness, and unbiasedness. Specifically, the qualitative metrics were assessed by three senior radiologists (W.Z., C.Z., and J.L.) using a five-point Likert scale (1 = strongly disagree, 5 = strongly agree) (Fig. 10). To ensure unbiased assessment, all evaluators were blinded to the identity of the models, and model outputs were anonymized and randomly ordered before evaluation. For each case, the final score for each dimension was the average of all evaluators’ scores. The error numbers made by each LLM and the scores of the Likert scale were then used for statistical analysis to evaluate the effect of the reasoning processes of the LRM.
The original scale was developed in Chinese; the English translation is presented here for illustrative purposes.
For the three human-in-the-loop evaluation dimensions (A–C) described above, six radiologists of various experience independently rated 54 cases using predefined ordinal or categorical scales. Information completeness (A) and short-term editability (C) were graded on a three-level ordinal scale (1–3). Reasoning helpfulness (B) was rated on a five-point Likert scale (1–5). All evaluations were performed through a custom graphical interface developed for this evaluation (Fig. S1). The interface also recorded time stamps at the start and end of each case evaluation, allowing derivation of per-case reading times as an objective indicator of efficiency. The specific scoring criteria are shown in Table 4. For each case, the corresponding imaging findings and one randomly selected model-generated output, i.e., the DeepSeek-R1 (Rea.) or the DeepSeek-R1 (Con.), were presented in a blinded and randomized order. All ratings and timing metadata were exported in JavaScript Object Notation (JSON) format for subsequent statistical analysis. This experimental design enabled a quantitative assessment of inter-observer agreement and experience-related differences in perceived utility between senior and junior radiologists.
Data
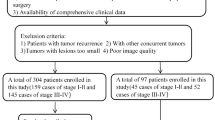
We retrospectively collected the 900 radiology reports (corresponding to 900 cancer cases) from three Chinese medical institutions: The Second Xiangya Hospital, Changsha, China (Institution 1), The Affiliated Cancer Hospital of Xiangya School of Medicine, Central South University, Changsha, China (Institution 2), and The First People’s Hospital of Changde City, Changde, China (Institution 3) from January 2024 to January 2025. Among these cases, those with breast cancer received bilateral MG (craniocaudal and mediolateral oblique views), non-contrast and contrast-enhanced chest CT, and contrast-enhanced breast MRI with DWI; lung cancer patients underwent non-contrast and contrast-enhanced chest CT; colorectal cancer patients underwent contrast-enhanced 3D whole-abdomen CT, pelvic MRI, and contrast-enhanced liver MRI. All radiology reports met local medical quality control standards and were confirmed by three senior radiologists (W.Z., C.Z., and J.L.) after being collected together. The report collection process is illustrated in Fig. 11. Only reports in which the impression section explicitly mentioned malignancy and were pathologically confirmed within several days of the report review were included. Cases that only mentioned an indeterminate nature, lacked pathological results, or had pathology results that could not definitively confirm cancer were excluded.
n1, n2, and n3 denote Institution 1, 2, and 3, respectively.
Ultimately, the following cases were randomly selected from Institution 1: 50 MG reports of breast cancer, 50 CT reports of breast cancer, 50 MRI reports of breast cancer, 50 CT reports of lung cancer, 50 CT reports of colorectal cancer, and 50 MRI reports of colorectal cancer. Similarly, from Institution 2 and Institution 3, 50 cases were randomly selected for each of the same six cancer-modality categories. This stratified sampling design mitigates the influence of case number disparities on model evaluation, permits consistent subgroup analyses, and supports the generalizability of our findings across various clinical contexts. All these reports were written in Chinese.
To further validate the reasoning effect of DeepSeek-R1, an English-language cohort was also utilized for diagnostic and qualitative evaluation. Specifically, we constructed a dataset named MIMIC-Cancer-90 from MIMIC-IV-Note v2.244. MIMIC-Cancer-90 consists of 90 cancer cases, with 30 cases for each of the three cancer types. The detailed construction procedures of MIMIC-Cancer-90 can be found in Fig. S2.
For both the Chinese and English reports, prompts were constructed in their raw languages and then input into the models. The outputs were thus generated in the corresponding language of the prompts. The English outputs of MIMIC-Cancer-90 were translated into Chinese using GPT-4.1 before being evaluated. All evaluations were performed in Chinese.
The detailed prompt texts and API call parameters are presented in Tables S9, S10.
Statistical analysis
Statistical analyses were performed to test whether differences between DeepSeek-R1 (Rea.) and comparator outputs were statistically significant. Since all models were applied to the same dataset, a repeated-measures design was adopted. Within each clinical subgroup (defined by cancer type, imaging modality, and institution) and in the overall dataset, pairwise comparisons between LLMs were conducted using McNemar tests on the four diagnostic metrics and paired t-tests on the 5-point scores of the four qualitative metrics and the reasoning helpness. For the error rate, the 95% confidence intervals were calculated using the Wilson score interval. For 3-level ordinal scores, Mann–Whitney tests were used to compare the distribution difference. For the evaluation time, paired t-tests were used to compare the mean evaluation time. All tests were two-tailed, and a p-value < 0.05 was considered statistically significant. All statistical analyses were performed using a combination of OriginPro 2025 (SR1, 10.2.0.196) and Python 3.10 (SciPy 1.5.4, Statsmodels 0.13.5).
Data availability
The private datasets generated and analyzed during the current study are available from the corresponding author on reasonable request. MIMIC-Cancer-90 can be accessed from: https://docs.google.com/spreadsheets/d/1TOn24shOVokCaWutVBDoFgkESwR4ln7X/edit?usp=drive_link&ouid=109800280201095538009&rtpof=true&sd=true.
Code availability
The underlying code for this study is available to qualified researchers on reasonable request from the corresponding author.
References
Sun, Z. et al. Evaluating GPT-4 on impressions generation in radiology reports. Radiology 307, e231259 (2023).
Hartung, M. P., Bickle, I. C., Gaillard, F. & Kanne, J. P. How to create a great radiology report. Radiographics 40, 1658–1670 (2020).
Onder, O. et al. Errors, discrepancies and underlying bias in radiology with case examples: a pictorial review. Insights Imaging 12, 51 (2021).
Zhang, L. et al. Diagnostic error and bias in the department of radiology: a pictorial essay. Insights Imaging 14, 163 (2023).
Wildenberg, J. C., Chen, P. H., Scanlon, M. H. & Cook, T. S. Attending radiologist variability and its effect on radiology resident discrepancy rates. Acad. Radiol. 24, 694–699 (2017).
Brady, A. P. Error and discrepancy in radiology: inevitable or avoidable? Insights Imaging 8, 171–182 (2017).
Gefter, W. B., Post, B. A. & Hatabu, H. Commonly missed findings on chest radiographs: causes and consequences. Chest 163, 650–661 (2023).
Seah, J. C., Tang, J. S. & Tran, A. Drafting the future: the dawn of AI report generation in radiology. Radiology 316, e243378 (2025).
Sloan, P., Clatworthy, P., Simpson, E. & Mirmehdi, M. Automated radiology report generation: a review of recent advances. IEEE Rev. Biomed. Eng. 18, 368–387 (2024).
Singhal, K. et al. Toward expert-level medical question answering with large language models. Nat. Med. 31, 1–8 (2025).
Liu, X. et al. A generalist medical language model for disease diagnosis assistance. Nat. Med. 31, 1–11 (2025).
Qiu, P. et al. Towards building multilingual language model for medicine. Nat. Commun. 15, 8384 (2024).
Amin, K. S. et al. Accuracy of ChatGPT, Google Bard, and Microsoft Bing for simplifying radiology reports. Radiology 309, e232561 (2023).
Gertz, R. J. et al. Potential of GPT-4 for detecting errors in radiology reports: implications for reporting accuracy. Radiology 311, e232714 (2024).
Sun, C. et al. Generative large language models trained for detecting errors in radiology reports. Radiology 315, e242575 (2025).
Tie, X. et al. Personalized impression generation for PET reports using large language models. J. Imaging Inform. Med. 37, 471–488 (2024).
Zhang, L. et al. Constructing a large language model to generate impressions from findings in radiology reports. Radiology 312, e240885 (2024).
Mitsuyama, Y. et al. Comparative analysis of GPT-4-based ChatGPT’s diagnostic performance with radiologists using real-world radiology reports of brain tumors. Eur. Radiol. 35, 1938–1947 (2024).
Kim, C., Gadgil, S. U. & Lee, S. I. Transparency of medical artificial intelligence systems. Nat. Rev. Bioeng. https://doi.org/10.1038/s44222-025-00363-w (2025).
Jaech, A. et al. OpenAI o1 system card. Preprint at arXiv https://doi.org/10.48550/arXiv.2412.16720 (2024).
Mitchell, M. Artificial intelligence learns to reason. Science 387, eadw5211 (2025).
Ji, Y. et al. Test-time computing: from System-1 thinking to System-2 thinking. Preprint at arXiv:2501.02497 (2025).
Guo, D. et al. Deepseek-r1 incentivizes reasoning in LLMs through reinforcement learning. Nature 645, 633–638 (2025).
Wei, J. et al. Chain-of-thought prompting elicits reasoning in large language models. Adv. Neural Inf. Process. Syst. 35, 24824–24837 (2022).
Tie, G. et al. A survey on post-training of large language models. Preprint at https://doi.org/10.48550/arXiv.2503.06072 (2025).
Deng, Z. et al. Exploring DeepSeek: a survey on advances, applications, challenges and future directions. IEEE/CAA J. Autom. Sin. 12, 872–893 (2025).
Tordjman, M. et al. Comparative benchmarking of the DeepSeek large language model on medical tasks and clinical reasoning. Nat. Med. 31, 2550–2555 (2025).
Grattafiori, A. et al. The LLaMA 3 herd of models. Preprint at arXiv: https://doi.org/10.48550/arXiv.2407.21783 (2024).
Sandmann, S. et al. Benchmark evaluation of DeepSeek large language models in clinical decision-making. Nat. Med. 31, 2546–2549 (2025).
Wu, Y. et al. When more is less: understanding chain-of-thought length in llms. Preprint at arXiv https://doi.org/10.48550/arXiv.2502.07266 (2025).
Aggarwal, P. & Welleck, S. L1: Controlling how long a reasoning model thinks with reinforcement learning. Preprint at arXiv https://doi.org/10.48550/arXiv.2503.04697 (2025).
Aytes, S. A., Baek, J. & Hwang, S. J. Sketch-of-thought: efficient llm reasoning with adaptive cognitive-inspired sketching. Preprint at arXiv https://doi.org/10.48550/arXiv.2503.05179 (2025).
Pal, S., Bhattacharya, M., Lee, S. S. & Chakraborty, C. A domain-specific next-generation large language model (LLM) or ChatGPT is required for biomedical engineering and research. Ann. Biomed. Eng. 52, 451–454 (2024).
Li, F., Hogg, D. C. & Cohn, A. G. Advancing spatial reasoning in large language models: an in-depth evaluation and enhancement using the stepgame benchmark. Proc. AAAI Conf. Artif. Intell. 38, 18500–18507 (2024).
Spînu-Popa, E. V., Cioni, D. & Neri, E. Radiology reporting in oncology—oncologists’ perspective. Cancer Imaging 21, 63 (2021).
Wan, D., Vig, J., Bansal, M. & Joty, S. On positional bias of faithfulness for long-form summarization. In Proc. Conference of the Nations of the Americas Chapter of the Association for Computational Linguistics: Human Language Technologies 1, 8791–8810 (ACL Anthology, 2025).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 5998–6008 (2017).
Wang, S. et al. Benefits and pitfalls of reinforcement learning for language model planning: a theoretical perspective. Preprint at arXiv https://doi.org/10.48550/arXiv.2509.22613 (2025).
Fan, Z. et al. ChestX-Reasoner: advancing radiology foundation models with reasoning through step-by-step verification. Preprint at arXiv https://doi.org/10.48550/arXiv.2504.20930 (2025).
Madaan, A. et al. Self-refine: iterative refinement with self-feedback. Adv. Neural Inf. Process. Syst. 36, 46534–46594 (2023).
Yang, A. et al. Qwen3 technical report. arXiv https://doi.org/10.48550/arXiv.2505.09388 (2025).
Team, K. et al. Kimi k2: open agentic intelligence. Preprint at arXiv https://doi.org/10.48550/arXiv.2507.20534 (2025).
Staal, J., Waechter, J., Allen, J., Lee, C. H. & Zwaan, L. Deliberate practice of diagnostic clinical reasoning reveals low performance and improvement of diagnostic justification in pre-clerkship students. BMC Med. Educ. 23, 684 (2023).
Johnson, A., Pollard, T., Horng, S., Celi, L. A. & Mark, R. MIMIC-IV-note: deidentified free-text clinical notes (version 2.2). PhysioNet. RRID:SCR_007345 https://doi.org/10.13026/1n74-ne17 (2023).
Acknowledgements
We thank Li Fan, Shiwei Luo, Cong Li, Minglei Xiao, Yule Zeng, and Ying Deng of the Department of Radiology, The Second Xiangya Hospital, and Jiacheng Pang of the Department of Radiology, The First People’s Hospital of Changde City, for their professional expertise with the model evaluation and raw data collection and organization in this study. This work was supported by NSFC-FDCT Grants 62361166662; National Key R&D Program of China 2023YFC3503400, 2022YFC3400400; National Natural Science Foundation of China U23A20479, 61971451, U22A20303, 62476291; The Innovative Research Group Project of Hunan Province 2024JJ1002; Key R&D Program of Hunan Province 2023GK2004, 2023SK2059, 2023SK2060; Top 10 Technical Key Project in Hunan Province 2023GK1010; Key Technologies R&D Program of Guangdong Province (2023B1111030004 to FFH); Hunan Provincial Natural Science Foundation for Distinguished Young Scholars (2025JJ20097); the Research Foundation of Education Bureau of Hunan Province (24B0003).
Author information
Authors and Affiliations
Contributions
R.W., J.L., and J.W. conceived and designed the study. R.W. and J.W. drafted the original manuscript. J.W., C.Z., S.H., and X.L. collected the data. W.Z., C.Z., S.H., Y.W., X.L., and Z.W. contributed to data interpretation. R.W., J.W., Y.W., G.T., and S.P. contributed to data analysis. W.Z., C.Z., and J.L. contributed to model evaluation. W.Z., Z.W., S.P., and J.L. provided supervision and contributed to writing review. All authors critically reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, R., Wang, J., Wang, Y. et al. Diagnostic and interpretive gains from reasoning over conclusions with a large reasoning model in radiology. npj Digit. Med. 9, 100 (2026). https://doi.org/10.1038/s41746-025-02285-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02285-8