Abstract
Colorectal cancer outcomes are critically dependent on early diagnosis, yet colonoscopy screening suffers from significant miss rates, particularly for subtle flat or serrated lesions. Although artificial intelligence holds promise for computer-aided detection (CADe), system performance is frequently bottlenecked by the scarcity and imbalance of high-quality annotated datasets. To address this, we employed a conditional StyleGAN architecture to synthesize high-resolution images of colorectal neoplasms, leveraging over 150,000 images aggregated from diverse public datasets. When utilized to train YOLOv5 detection models, this synthetic data demonstrated high fidelity and significantly enhanced diagnostic performance. Hybrid augmentation improved the mean Average Precision from 0.86 to 0.93 on internal testing and markedly reduced the generalization gap on independent external validation sets. Crucially, recall for challenging flat and depressed lesions rose from 0.72 to 0.87. These findings indicate that generative augmentation effectively strengthens model robustness and generalization across diverse clinical scenarios. While currently limited to still imagery, this strategy provides a scalable solution to data limitations, potentially elevating the standard of AI-assisted endoscopic surveillance.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) represents a formidable global public health challenge, ranking as the third most diagnosed cancer and the second leading cause of cancer mortality worldwide according to GLOBOCAN 2022 estimates, with nearly 2.0 million new cases and over 900,000 deaths annually. This burden is marked by significant socioeconomic disparities; while high-income nations historically show the highest incidence rates linked to Westernized lifestyles, they are now experiencing a stabilization or decline due to effective screening, whereas rates are alarmingly escalating in developing regions where limited healthcare access leads to higher mortality. This epidemiological landscape translates into a staggering economic and societal strain, encompassing not only vast direct medical costs but also profound indirect costs from lost productivity and premature death, which are exponentially higher for late-stage versus early-stage disease. This stark correlation between diagnostic stage, patient survival, and economic burden creates a compelling and urgent imperative for global strategies focused on prevention and early detection Fig. 1.
The maps illustrate the global burden of CRC, with color intensity corresponding to age-standardized rates.
The cornerstone of CRC prevention lies in its well-defined and typically indolent natural history, which provides a crucial window for intervention. The majority of sporadic CRCs develop through the multi-step adenoma carcinoma sequence, shown in Fig. 2 a process where normal colonic epithelium transforms into a benign adenomatous polyp that, over a span of 10 to 15 years, accumulates a series of genetic and epigenetic alterations, ultimately leading to invasive cancer1,2. This progression is driven by distinct molecular pathways, primarily the Chromosomal Instability (CIN) pathway, characterized by mutations in genes like APC, KRAS, and TP53, and the Microsatellite Instability (MSI) pathway, which results from a deficient DNA mismatch repair system3,4. This protracted, multi-stage evolution from a detectable, benign precursor to a malignant tumor is the fundamental biological rationale for screening programs, as the removal of adenomas via polypectomy effectively intercepts this sequence and prevents the development of cancer.
This flowchart illustrates the multi-step progression from normal colonic mucosa to invasive carcinoma, highlighting key morphological changes and the sequential accumulation of critical genetic mutations (e.g., APC, KRAS, p53).
The clinical imperative for early detection is powerfully illustrated by the profound survival gradient based on the stage of CRC at diagnosis. Data from population-based registries like the SEER Program show that the 5-year relative survival rate is approximately 91% for localized disease but plummets to a mere 15% for patients with distant metastases5. This stark reality has driven the development of a portfolio of screening modalities, endorsed by clinical guidelines, designed to identify CRC and its precursors in asymptomatic individuals6. These strategies, which balance factors like efficacy, invasiveness, and patient adherence, are broadly categorized into two main groups: Stool-Based Tests: These non-invasive tests analyze stool samples for biomarkers associated with colorectal neoplasia. Guaiac-based Fecal Occult Blood Test (gFOBT): Detects heme, the non-protein component of hemoglobin, through a chemical reaction. It is not specific to human blood and can be affected by diet. Fecal Immunochemical Test (FIT): Employs antibodies to specifically detect human globin, a protein component of hemoglobin, making it more specific for colonic bleeding than gFOBT and unaffected by diet. Stool DNA Test (sDNA-FIT): A multi-target test that combines a FIT component with the detection of exfoliated tumor DNA containing cancer-specific genetic and epigenetic alterations. Direct Visualization Tests: These methods provide a direct structural examination of the colonic mucosa. Flexible Sigmoidoscopy: An endoscopic examination limited to the rectum and the lower (sigmoid) colon. Optical Colonoscopy: This procedure involves a comprehensive endoscopic examination of the entire colon and rectum, uniquely allowing for both the detection and concurrent removal of polyps. CT Colonography (Virtual Colonoscopy): A radiological imaging technique that uses a computed tomography (CT) scanner to generate two- and three-dimensional images of the colon and rectum.
Among the available screening modalities, optical colonoscopy is unequivocally recognized as the gold standard, primarily due to its unique dual capacity for both high fidelity diagnosis and immediate therapeutic intervention7. This see and treat paradigm, where precancerous adenomas are identified and excised via polypectomy within a single procedure, directly intercepts the adenoma carcinoma sequence, making colonoscopy a truly preventative tool8. The profound efficacy of this approach is not merely theoretical but is substantiated by a wealth of evidence from landmark clinical trials. Studies such as the National Polyp Study and long-term follow-up from randomized trials like the PLCO Cancer Screening Trial have definitively shown that endoscopic screening and polypectomy lead to significant reductions in both the long-term incidence and mortality of colorectal cancer, cementing its status as the cornerstone of CRC prevention9,10,11.
Despite its proven efficacy, the protective benefit of colonoscopy is not absolute and is fundamentally constrained by its nature as a human-operated procedure. The quality of the examination is profoundly dependent on the individual endoscopist’s skill, experience, and real-time cognitive engagement, leading to significant inter-operator variability in key performance metrics12. The most critical of these is the Adenoma Detection Rate (ADR), a direct surrogate for procedural quality that is inversely correlated with the risk of post-colonoscopy colorectal cancer; for every 1% increase in ADR, the risk of interval cancer decreases by 3%13. Human factors such as fatigue further compound this variability, potentially reducing detection capabilities14. To mitigate these challenges and standardize care, professional societies have established a set of Key Performance Indicators (KPIs), detailed in Table 1, which serve as benchmarks for high-quality colonoscopy15. The persistent variation in these crucial metrics, even among experienced physicians, highlights the inherent perceptual and cognitive difficulties of the procedure and underscores a critical need for technological solutions that can help standardize and elevate the quality of care for all patients.
A direct and troubling consequence of operator dependency is the polyp miss rate, a well-documented phenomenon that represents a significant Achilles’ heel of colonoscopy and a primary contributor to interval cancers. Rigorous tandem colonoscopy studies, where a second procedure immediately follows the first, have quantified this diagnostic gap, revealing a sobering overall miss rate for adenomas of 22%, a figure that rises to 28% for smaller lesions16. The etiology of these missed polyps is multifactorial, stemming from a complex interplay of procedural factors (e.g., suboptimal bowel preparation, rapid withdrawal), lesion characteristics (e.g., small size, inconspicuous non-polypoid morphologies), anatomical challenges (e.g., location behind haustral folds), and inescapable human factors such as momentary lapses in concentration and fatigue17,18. This convergence of challenges transforms polyp detection into a formidable perceptual task and provides a powerful impetus for developing technologies that can augment human perception and reduce this critical diagnostic gap.
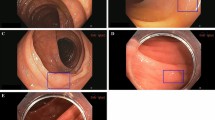
Compounding the problem of missed lesions is the existence of morphologically challenging precursors that defy easy endoscopic detection. A significant subset of precancerous lesions, namely non-polypoid colorectal neoplasms (NP-CRNs) and sessile serrated adenomas/polyps (SSA/Ps), presents with far more subtle, stealthy features than typical protruding polyps. NP-CRNs, particularly flat (0-IIa) and depressed (0-IIc) types, exhibit minimal vertical growth and often appear only as slight alterations in mucosal color or texture19. Similarly, SSA/Ps—the precursors to cancers of the serrated pathway—are notoriously indistinct, characterized as pale, flat lesions with ill-defined borders, often cloaked by a camouflaging mucus cap20. The clinical significance of these subtle lesions shown in Fig. 3 is profound; not only do they harbor a higher potential for advanced histology, but their high miss rates mean they are a primary driver of post-colonoscopy colorectal cancers (PCCRCs)21,22. The disproportionate contribution of these morphologically challenging, yet biologically aggressive, lesions to the burden of interval cancers represents a pressing unmet need and provides a powerful rationale for advanced technological aids to enhance their detection.
Illustrative endoscopic images showcasing the distinct visual characteristics of a a typical Pedunculated Polyp, b a flat adenoma, c a depressed lesion, and d a SSA/P (sessile serrated adenoma/polyp).
The evolution of Artificial Intelligence (AI) has introduced a new paradigm in medical diagnostics, shifting from traditional Machine Learning approaches that required laborious manual feature engineering to the more powerful methods of Deep Learning (DL)23,24. At the forefront are Convolutional Neural Networks (CNNs), which have become the cornerstone of modern medical image analysis. By automatically learning a hierarchical representation of features directly from pixel data—from simple edges to complex pathologies—CNNs have achieved, and at times surpassed, human-level accuracy in domains like radiology and pathology, establishing a strong precedent for their use in endoscopy25,26,27.
This technological leap has led to the development of two primary applications in colonoscopy. The first, Computer Aided Detection (CADe), functions as a real-time second observer that highlights suspicious lesions to the endoscopist28,29. The clinical efficacy of CADe is no longer theoretical; it is firmly established by multiple large-scale randomized controlled trials (RCTs) and meta-analyses, which consistently demonstrate that AI assistance significantly increases the Adenoma Detection Rate (ADR)—a key quality metric directly linked to the prevention of interval cancers30,31,32. The second application, Computer Aided Diagnosis (CADx), aims to provide an optical biopsy by predicting the histology of a detected polyp in real time33,34. The potential of CADx is to enable cost-saving clinical strategies like resect and discard for benign polyps, but its adoption hinges on achieving near-perfect accuracy and generalization—a significant challenge that underscores the limitations of current systems35,36,37.
Despite the proven successes of both CADe and CADx, their performance is fundamentally constrained by a critical dependency: the availability of large-scale, high-quality, and diverse training data38. The development of such datasets is notoriously difficult, facing three primary obstacles. First, strict privacy regulations like HIPAA and GDPR create significant logistical and legal barriers to aggregating patient data from multiple institutions39. Second, the process of expert annotation—where clinicians manually label thousands of images—is extraordinarily time-consuming and financially prohibitive40. Third, medical datasets are inherently imbalanced; images of rare but clinically crucial pathologies (e.g., flat or serrated polyps) are scarce compared to normal findings. This lack of diversity leads to the domain shift problem, where models trained on data from one source fail to generalize to new clinical environments with different equipment or patient populations41. This data bottleneck represents the primary ceiling on the performance and reliability of current endoscopic AI, necessitating innovative solutions that can transcend the limitations of existing real-world data.
To surmount the data-centric hurdles outlined previously, our approach moves beyond traditional data augmentation—such as simple geometric or photometric transformations which only create correlated variations of existing data42—and turns instead to the more sophisticated paradigm of generative modeling. The most powerful framework for this task is the Generative Adversarial Network (GAN), which conceptualizes image synthesis as a zero-sum game between two competing neural networks, shown in Fig. 4: a Generator (G) and a Discriminator (D)43. In an analogy to a counterfeiter (G) and a detective (D), the generator learns to produce synthetic images from random noise that are realistic enough to fool the discriminator, which simultaneously learns to distinguish these counterfeit images from real ones. This adversarial process forces the generator to capture the complex, underlying distribution of the training data with remarkable fidelity. The evolution of this concept, from early Deep Convolutional GANs (DCGANs) to state-of-the-art architectures like StyleGAN, has matured the technology to a point where it can generate high-resolution, photorealistic images suitable for clinical applications44,45.
The framework consists of two competing networks: a Generator that creates images from random noise, and a Discriminator that classifies its input as either real (from the training data) or fake (from the generator). The adversarial loss is backpropagated to improve both networks iteratively.
The potential of GANs to alleviate data scarcity is not merely theoretical but has been empirically validated across a diverse range of medical imaging domains, establishing a strong precedent for their application in endoscopy46. In radiology, for instance, GANs have been used to synthesize high-fidelity brain MRI and CT scans, with studies demonstrating that augmenting training data with these synthetic images significantly improves the performance of downstream tumor classification and segmentation models47,48. Similar successes have been reported in dermatology, where GANs help balance datasets by generating images of rare skin lesions, and in digital pathology, where they create synthetic histopathology slides to improve the robustness of cancer grading systems49,50. This consistent evidence across multiple specialties underscores a clear finding: augmenting real medical datasets with high-quality, GAN-generated images is a viable and effective strategy for enhancing the performance and generalizability of diagnostic AI models51.
We aim to address the critical data bottleneck in endoscopic AI by leveraging state-of-the-art generative models. The primary contributions of this work are threefold. First, we propose and implement a novel conditional GAN framework, built upon an advanced StyleGAN architecture, specifically tailored for synthesizing high-resolution, high-fidelity images of diverse colorectal neoplasms. Second, we conduct a rigorous evaluation of the generated images, using a combination of quantitative metrics (FID, IS) and a qualitative Visual Turing Test involving clinical experts, to demonstrate their realism and diversity. Third, and most critically, we provide strong empirical evidence that augmenting a training dataset with our synthetic images significantly improves not only the detection performance of a standard deep learning model but also, crucially, its generalization capabilities when evaluated on multiple, independent external datasets.
The remainder of our model is structured as follows. “Introduction” also provides a comprehensive review of the related literature, covering AI systems in colonoscopy and existing data augmentation strategies. “Methods” details our proposed methodology, including the datasets, the GAN architecture, and the experimental design. “Results” presents the experimental results, including the qualitative and quantitative evaluation of our generative model and the performance analysis of the downstream detection task. Finally, “Discussion” discusses the implications of our findings, acknowledges the limitations of this study, and outlines promising directions for future research.
This section provides a comprehensive review of the literature pertinent to the core components of our research. The discussion is structured to logically build the case for our proposed methodology. We begin by surveying the current landscape of deep learning applications in colorectal neoplasm detection and characterization, establishing the successes and existing limitations of Computer Aided Detection (CADe) and Diagnosis (CADx) systems. Subsequently, we examine the prevalent strategies employed to address the fundamental challenge of data scarcity in medical imaging, with a particular focus on data augmentation techniques and their inherent constraints. Finally, we conduct a focused review of Generative Adversarial Networks (GANs), highlighting their successful application for synthetic image generation in various medical domains. This structured review serves to contextualize our work, identify the critical research gap, and establish the scientific premise for employing a generative approach to enhance endoscopic AI.
The application of deep learning has fundamentally reshaped endoscopic image analysis, leading to sophisticated systems for both detecting and characterizing colorectal neoplasms28. The most mature application, Computer Aided Detection (CADe), leverages real-time object detection frameworks like YOLO and Faster R-CNN to function as a second observer, identifying polyps during live procedures29,52,53. Its clinical value is now firmly established through numerous randomized controlled trials (RCTs) and subsequent meta-analyses, which consistently show that CADe significantly increases the adenoma detection rate (ADR)30,31,32. Building upon this success, Computer Aided Diagnosis (CADx) systems represent the next frontier, aiming to provide an optical biopsy by characterizing the histology of detected lesions in real time33,34. By analyzing subtle mucosal and vascular patterns, often enhanced by technologies like Narrow Band Imaging (NBI), these models can accurately differentiate between benign and premalignant polyps, potentially enabling clinical strategies such as resect and discard54,55. However, despite their proven and potential benefits, both CADe and CADx systems share a critical vulnerability: their performance is intrinsically tethered to the quality and scale of their training data56,57. This dependency is particularly acute for CADx, which requires vast datasets of high-quality, expertly annotated images to learn the fine-grained features necessary for accurate histological prediction, thus magnifying the data bottleneck that hinders the development of robust and generalizable endoscopic AI36,37,58.
To mitigate the challenge of limited dataset size, a universally adopted practice in training deep learning models is data augmentation39. The most common approach involves applying a series of traditional, transformation-based techniques to the existing training images, such as geometric alterations (e.g., rotation, scaling, flipping, cropping) and photometric modifications (e.g., adjustments to brightness, contrast, and color saturation)42. This strategy has proven essential for improving the generalization capabilities of deep learning models by teaching them a degree of invariance to these basic transformations, which enhances robustness and helps to prevent overfitting to the specific characteristics of the training set59. However, the fundamental limitation of these methods is that they do not generate any truly novel pathological or anatomical information; they merely create heavily correlated, slightly altered versions of images that already exist in the dataset60. Consequently, while traditional augmentation can help a model become less sensitive to a polyp’s orientation or illumination, it cannot create a new example of a rare, flat serrated lesion if none exists in the original data, nor can it capture the vast spectrum of subtle morphological variations seen in clinical practice46. This inability to expand the feature space beyond the confines of the initial dataset means that traditional augmentation is only a partial solution, thereby necessitating the exploration of more advanced data synthesis paradigms capable of generating genuinely new and diverse training examples.
Generative Adversarial Networks (GANs) have emerged as a state-of-the-art solution for advanced data synthesis, with a proven track record of enhancing diagnostic models in various medical specialties46,51. In fields like radiology and pathology, augmenting training sets with GAN-synthesized images of brain tumors, liver lesions, or varied histopathology slides has been shown to significantly improve the performance of downstream classification and segmentation tasks47,48,50. Despite this success, the application of GANs to endoscopic imaging remains nascent61. While early studies have confirmed the feasibility of generating synthetic colonoscopy images, they serve primarily as feasibility studies and are often hampered by limitations such as low resolution, insufficient morphological diversity, and, most importantly, a lack of rigorous validation on independent, external datasets62,63,64.
Consequently, a critical research gap persists: it remains unknown if GAN-based augmentation can solve the high-stakes clinical problem of generalizing to new hospital systems or improving the detection of subtle, high-risk lesions (flat, depressed, serrated) that are key contributors to interval cancers21. Our work directly addresses this void. We move beyond simple feasibility to conduct a systematic evaluation of generalization and provide a targeted analysis on challenging lesion subtypes, comparing our approach directly against traditional augmentation to demonstrate a quantifiable improvement in model robustness.
Results
Following the methodology detailed in the previous section, this section presents a comprehensive empirical evaluation designed to validate the efficacy of our proposed Generative Adversarial Network (GAN) framework and the utility of its synthetic data. This section is organized into two primary parts. First, we conduct a thorough assessment of the GAN model itself, evaluating the quality, fidelity, and diversity of the generated endoscopic images from both quantitative and qualitative perspectives. Second, and more critically, we systematically demonstrate the impact of using these synthetic data as an augmentation strategy for a downstream polyp detection model. We present a comparative analysis of its performance on both an internal test set and, crucially, on multiple independent external validation sets to rigorously assess improvements in generalization. Finally, through a series of ablation studies and error analyses, we further investigate the key factors contributing to the performance gains afforded by our generative approach.
Evaluation of generative model quality
To establish an objective baseline for the quality of our generative model, we first performed a rigorous quantitative evaluation using a panel of standard, widely accepted metrics: Fréchet Inception Distance (FID), Inception Score (IS), and Learned Perceptual Image Patch Similarity (LPIPS). For each metric calculation, a set of 10,000 synthetic images was generated using the trained model and compared against the entire curated real training dataset. This process was repeated at several key checkpoints during the training process to monitor convergence and model improvement over time.
The results, summarized in Table 2, demonstrate a clear and consistent improvement in image quality as training progressed. The FID score, which measures the similarity between the feature distributions of real and generated images, exhibited a steep decline from an initial value of 85.32 to a final, stable score of 21.79. As shown in the comparative analysis in Fig. 5, this score is highly competitive, significantly outperforming earlier architectures such as DCGAN (62.3) and WGAN-GP (38.7), and surpassing the standard StyleGAN2 baseline (24.9). This trend indicates that the generator progressively learned to produce images that more closely matched the statistical properties of the real endoscopic data. Concurrently, the Inception Score, reflecting the clarity and diversity of the generated images, steadily increased, plateauing at a competitive score of 4.89. Similarly, the LPIPS metric, which quantifies perceptual similarity, decreased from 0.55 to 0.21, confirming that the synthetic images became increasingly indistinguishable from real images from a perceptual feature standpoint. These converging trends strongly suggest that the model successfully learned the complex underlying distribution of the training data and reached a state of high fidelity synthesis.
The bar chart compares the FID score of our final model against other relevant GAN architectures. The scores for DCGAN, WGAN-GP, PGGAN, and StyleGAN2 are representative benchmarks from literature or our baseline implementations on similar endoscopic datasets, serving to contextualize the performance of our model.
When contextualized against existing literature, as shown in Fig. 5, the final FID score of 21.79 achieved by our model is highly competitive. It surpasses the performance reported by earlier GAN applications in endoscopy and is comparable to state-of-the-art results in other, more mature medical imaging domains. This strong quantitative performance validates the architectural choices and training strategies detailed in Methods and provides a solid foundation for the subsequent qualitative and downstream task evaluations.
Beyond quantitative scores, a qualitative assessment is indispensable for verifying that the generated images are not only statistically similar to real data but are also visually plausible and clinically meaningful. This is particularly important for medical applications where subtle morphological and textural details carry significant diagnostic information. To this end, we present a visual showcase of our model’s generative capabilities.
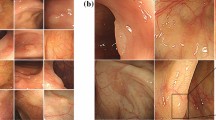
Figure 6 displays a large, uncurated matrix of images randomly generated by our trained model. These samples serve to illustrate the overall quality and diversity of the learned data distribution. Visually, the synthetic images demonstrate a high degree of realism. The model has successfully learned to reproduce key characteristics of real endoscopic images, including the complex mucosal texture of the colon wall, the specular highlights caused by the endoscope’s light source, the subtle color gradients, and the out-of-focus background elements. The diversity is also evident in the wide range of generated polyp morphologies, sizes, and orientations, as well as the varied colonic backgrounds.
This grid shows a random, uncurated selection of high-resolution images produced by our GAN, demonstrating the overall quality and diversity of the generated data. Five representative examples are indicated in panels (a–e).
Finally, to probe the structure of the learned latent space, we performed a latent space interpolation, as shown in Fig. 7. By selecting two latent vectors, w1 and w2, corresponding to two distinct generated images and linearly interpolating between them, we can observe the resulting image transitions. The smooth and seamless transformation between different polyp appearances—for example, a gradual change in size, shape, or texture without any abrupt or nonsensical artifacts—provides strong evidence that the generator has learned a meaningful and continuous representation of the data. This demonstrates that the model is not merely memorizing and reproducing training examples but has captured the underlying manifold of endoscopic images, allowing it to generate novel and plausible variations that lie between existing data points.
This sequence shows a smooth transition between two different synthetic polyp images by linearly interpolating their corresponding latent vectors in the \({\mathcal{W}}\) space, highlighting the continuous and well learned feature manifold.
Performance evaluation on the downstream polyp detection task
The first phase of our downstream task evaluation was conducted on the internal test set. This dataset was held out from the training and validation processes but was drawn from the same source distribution as the training data. The primary objective of this experiment was to assess the impact of different data augmentation strategies on the model’s performance under in-distribution conditions, providing a direct measure of the benefit derived from each approach.
The results, presented in Table 3, clearly demonstrate the value of data augmentation, with the GAN-based strategies yielding the most significant performance gains. The baseline model (Group A), trained only on real data, achieved a respectable mAP of 0.86. As expected, applying traditional augmentation (Group B) improved performance, raising the mAP to 0.89, primarily by enhancing the model’s robustness to simple geometric and photometric variations. However, the models augmented with synthetic data showed a more substantial improvement. The GAN-only augmentation (Group C) outperformed traditional methods, achieving an mAP of 0.92. Most notably, this improvement was driven by a marked increase in recall (from 0.88 to 0.92), suggesting the synthetic data helped the model to identify polyps that were previously missed. The hybrid approach (Group D), which combined both GAN-generated data and traditional augmentations, achieved the best overall performance with an mAP of 0.93, indicating that these two strategies are complementary. The Precision Recall curves shown in Fig. 8 further illustrate that the GAN augmented models consistently dominate the non-GAN models across all operating thresholds.
The curves illustrate the performance of the four experimental groups. The models augmented with GAN data (C and D) exhibit a clear superiority, achieving higher precision across all levels of recall.
The ultimate test of a clinical AI model is not its performance on data similar to its training set, but its ability to generalize to entirely unseen data from different clinical environments. To this end, we evaluated the four trained models on two independent external validation sets, CVC-ClinicDB and the more challenging ETIS-LaribPolypDB, neither of which were used in any part of the training or hyperparameter tuning process. These datasets originate from different institutions and were captured with different endoscopy hardware, thus providing a rigorous test for model robustness against domain shift.
The performance on the external datasets, detailed in Tables 4 and 5, reveals the critical impact of our generative augmentation strategy on model generalization. As anticipated, all models experienced a performance degradation when faced with out-of-distribution data. However, the magnitude of this degradation varied significantly between the groups. The baseline model (Group A) suffered the most substantial drop in performance, with its mAP decreasing by approximately 12% on CVC-ClinicDB and 19% on the more challenging ETIS-LaribPolypDB, relative to the internal test set. While traditional augmentation (Group B) slightly mitigated this drop, the models trained with synthetic data (Groups C and D) demonstrated markedly superior robustness. The hybrid model (Group D), for instance, saw its mAP decrease by only 7% on CVC-ClinicDB and 12% on ETIS-LaribPolypDB.
This key finding is visualized in Fig. 9. The models trained with GAN augmented data exhibit a significantly smaller generalization gap—the difference between performance on internal and external data—compared to the baseline and traditionally augmented models. This demonstrates that by exposing the detection model to a wider and more diverse range of clinically plausible examples during training, our GAN-based approach enables the model to learn more fundamental and invariant features of polyps. Consequently, the resulting model is less susceptible to superficial variations in imaging conditions and is better equipped to generalize its knowledge to new, unseen clinical environments, thereby affirming our central hypothesis.
The bars represent the drop in mAP from the internal test set to the average of the external test sets for each experimental group. A smaller bar indicates a smaller generalization gap and superior robustness.
To specifically assess the impact of our augmentation strategy on the detection of the most clinically significant and difficult-to-detect lesions, we performed a sub-analysis on a curated challenging subset of the data. This subset was compiled by pooling all instances of flat, depressed, and serrated polyps from both the internal and external test sets. The primary goal of this analysis was to determine if the increased diversity from GAN-generated data translates into a tangible improvement in identifying these endoscopically occult neoplasms, which are a major cause of post-colonoscopy colorectal cancers.
The results on this challenging subset, presented in Table 6, are particularly revealing. The baseline model’s recall of 0.72 indicates that it missed nearly 28% of these critical lesions. While traditional augmentation offered a modest improvement to 0.76, the GAN-augmented models demonstrated a substantial leap in performance. The hybrid model (Group D) achieved a recall of 0.87, representing a 15 percentage point improvement over the baseline. This significant increase in sensitivity directly addresses one of the core clinical limitations of human-led colonoscopy that was identified in the introduction. The qualitative examples in Fig. 10 provide further insight, illustrating cases where the baseline model failed to identify a subtle flat lesion that was successfully detected with high confidence by the model trained with our diverse, synthetic data. This strongly suggests that by exposing the model to a richer variety of morphological features during training, our GAN-based augmentation strategy specifically enhances its ability to recognize these clinically crucial, invisible polyps.
The top row shows examples of flat or subtle polyps (blue boxes are ground truth) missed by the baseline model. The bottom row shows the same images with successful, high-confidence detections (green boxes) by our hybrid GAN-augmented model (Ours a and b).
Ablation studies
To isolate the contribution of our conditional framework, we trained an additional model (Group C-Unconditional) using an unconditional StyleGAN. This model was trained on the same data but without any class labels. The generated images were then used to augment the detector (same 1:1 ratio as Group C). The results, summarized in Table 7, show that while unconditional generation does improve generalization over the baseline, it significantly underperforms our conditional approach. This is most evident in the challenging lesion subset, where the unconditional model provides minimal recall improvement. This confirms that the conditional mechanism, which allows us to direct the synthesis of rare lesion types, is the key driver of our model’s improved sensitivity.
To better understand the sensitivity of the downstream model to the quantity of synthetic data, we conducted an ablation study on the ratio of synthetic to real images shown in Fig. 11 used during training. We trained several instances of the hybrid augmentation model (Group D), varying the ratio of synthetic images added to the real training set from 1:0.5 (one synthetic image for every two real ones) to 1:2 (two synthetic images for every one real one). The performance of each resulting model was then evaluated on the combined external validation set.
The line graph plots the mAP on the combined external validation set as a function of the ratio of synthetic to real images in the training data.
Discussion
The comprehensive evaluation presented in the previous section yields two central findings. First, our proposed StyleGAN-based framework is capable of generating high-fidelity, diverse, and clinically plausible endoscopic images of colorectal neoplasms, as validated by both state-of-the-art quantitative metrics and blinded expert evaluation. Second, and most importantly, we have demonstrated that leveraging these synthetic images as a form of data augmentation provides a substantial and statistically significant improvement to the performance of a downstream polyp detection model. This benefit is not only observed in in-distribution test data but, crucially, also translates to a marked improvement in generalization performance on unseen external datasets, effectively narrowing the generalization gap. The most pronounced improvements were seen in the detection of challenging, clinically significant lesions.
The superior performance of GAN-based augmentation over traditional methods can be attributed to its ability to expand the feature space of the training data in a meaningful way. While traditional techniques create variations in pose, color, and scale, they do not create novel instances of pathology. Our generative model, having learned the underlying data distribution, can synthesize entirely new examples of polyps with unique combinations of texture, morphology, and lighting that, while plausible, are not present in the original dataset.
Our ablation study (Table 7) confirms that the conditional nature of our framework is the key mechanism for this, as it alone enabled the targeted synthesis of rare, subtle lesions that drove the crucial gains in recall. Foundational choices, such as the high resolution 512 × 512 format and the Projection Discriminator, were deliberately selected to ensure these fine-grained and class-specific morphological features were captured with high fidelity. This forces the detection model to learn more robust and fundamental features of what constitutes a polyp, rather than overfitting to the specific instances in a limited real dataset. The success of the hybrid approach (Group D) further suggests that the feature space explored by GANs is complementary to that of traditional augmentation, and their combined use leads to the most robust and generalizable models.
We addressed the fundamental challenge of data scarcity in the development of robust AI systems for colonoscopy. We demonstrated that a state-of-the-art conditional Generative Adversarial Network can synthesize a diverse library of high-resolution, clinically realistic images. The central contribution of this work lies in the empirical validation of these synthetic data, which revealed two critical, clinically translatable advantages.
First, our approach markedly improves clinical robustness. By training on a more diverse, GAN-augmented dataset, our model demonstrated significantly smaller performance degradation on unseen external datasets (Fig. 9). This reduced generalization gap is crucial for real-world deployment, suggesting the model is more reliable and less likely to fail when encountering the domain shift of new hospital systems or endoscopy hardware.
Second, and most importantly, our method directly targets the primary cause of post-colonoscopy colorectal cancers (PCCRCs). Our hybrid model achieved a 15 percentage point increase in recall for challenging, subtle lesions (Table 6), the very flat, depressed, and serrated polyps that are most often missed by endoscopists and are responsible for the majority of interval cancers. This enhanced sensitivity to invisible high-risk precursors provides a direct pathway to reducing the adenoma miss rate, improving the quality of colonoscopy, and ultimately lowering the incidence of preventable interval cancers. This generative strategy is therefore a powerful tool for overcoming the data bottleneck and developing more robust and clinically effective AI.
Limitations include reliance on publicly available datasets and evaluation restricted to still images rather than full colonoscopy video. Furthermore, while our results strongly suggest the gains from GAN augmentation stem from feature novelty rather than data volume, a future ablation study comparing our hybrid model against a traditional augmentation model trained for a matched number of total iterations (e.g., double the epochs) could further isolate this effect. Future work will explore the generation of full video sequences and the synthesis of images from different imaging modalities, such as narrow band imaging, to further enhance the realism and utility of the synthetic data.
Methods
This section systematically delineates the comprehensive technical framework proposed herein to address the critical issue of data scarcity in endoscopic imaging. We commence by detailing the origins, curation criteria, and preprocessing pipeline for the datasets utilized in this study. Following this, the core of our methodology—the architecture, innovative components, and training strategy of our bespoke Generative Adversarial Network (GAN)—is thoroughly explicated. Subsequently, we define the rigorous qualitative and quantitative methodologies employed to assess the quality of the synthesized images. Finally, the section outlines the experimental design for applying these synthetic data to a downstream polyp detection task, detailing the protocols and performance metrics established to validate their efficacy in a clinically relevant context.
Data acquisition and preprocessing
The foundation of any robust deep learning model is a large-scale, high-quality, and diverse dataset. To this end, we adopted a multi-source data fusion strategy, aggregating multiple internationally recognized, public colonoscopy datasets to construct a comprehensive training corpus that captures a wide spectrum of clinical variability. The specifics of each source dataset are detailed in Table 8. To ensure the fidelity of this aggregated dataset, we implemented a stringent, two-stage data curation and annotation protocol executed by a team of board-certified gastroenterologists. First, every image was meticulously screened, with inclusion restricted to frames demonstrating adequate bowel preparation (e.g., Boston Bowel Preparation Scale score ≥2 per segment), clear lesion visibility, and minimal artifacts; images compromised by poor focus, extensive artifacts, or ambiguous findings were systematically excluded65. Following this expert-led curation, the retained high-quality images were annotated using specialized software66. This process generated two forms of ground truth: precise pixel-wise segmentation masks for training the generative model and tight-fitting bounding boxes for the downstream detection task. To ensure the highest accuracy and inter-rater consistency, annotations were cross-validated by at least two experts, with any discrepancies resolved by a senior gastroenterologist. This quality control process was periodically validated using metrics such as Cohen’s Kappa coefficient to ensure a robust and reliable ground truth dataset67. High definition NBI images with histopathological labels. For our study, these labels were used only as class conditions during GAN training to generate specific polyp types, not for training a diagnostic classifier.
To prepare the data for our conditional GAN framework, a unified labeling taxonomy was required. Instead of performing de novo annotation, we leveraged the existing expert-provided labels from the source datasets (e.g., histopathological labels from PICCOLO, morphological data from others). We reconciled these heterogeneous labels by mapping all neoplasms into three broad, visually distinct morphological categories relevant to the detection task: (1) Pedunculated Polyps (lesions with a clear stalk), (2) Sessile Polyps (protruding, broad-based lesions), and (3) Subtle Lesions. This third category grouped the clinically critical non-polypoid types, including flat, depressed, and serrated lesions. This pragmatic classification allowed us to condition the GAN on visual appearance. The aggregated training corpus, prior to balancing, exhibited a natural imbalance, with Subtle Lesions being the least common (approx. 25% of positive frames).
To prepare the curated dataset for model ingestion, a standardized preprocessing pipeline was applied to convert each image into a uniform tensor format. This pipeline involved three key steps: first, all images and their corresponding masks were resized to a uniform resolution of 512 × 512 pixels, balancing the preservation of fine-grained detail with the computational demands of high-resolution GAN training45. Second, pixel values were normalized from the original [0, 255] integer range to the [−1, 1] floating point range to stabilize training and accelerate model convergence23. Finally, to mitigate the domain shift inherent in multi-source data, a structure-preserving color normalization technique based on histogram matching was employed to standardize the color profile across the entire dataset, encouraging the model to learn fundamental morphological features over superficial color variations41,68.
Proposed generative adversarial network framework
The selection of the GAN architecture is paramount for generating high-fidelity medical images. While early architectures like DCGAN established the viability of image synthesis, they often struggled with training stability and were limited to low-resolution outputs44. To meet the stringent quality demands of clinical-grade endoscopic images—characterized by complex textures and subtle features—we required a more advanced foundation. Therefore, we selected the StyleGAN family of architectures as the backbone for our generative model, specifically building upon the principles of StyleGAN2 and its alias-free successor, StyleGAN369,70. This choice was motivated by the architecture’s proven excellence in generating photorealistic, high-resolution images; its use of a mapping network to create a disentangled latent space for intuitive and fine-grained stylistic control; and its sophisticated regularization techniques that ensure stable training. This combination of high fidelity output, feature disentanglement, and training stability makes the StyleGAN framework the ideal choice for this work45.
The generator in our framework is a two-stage network, comprising a Mapping Network (f) and a Synthesis Network (g) shown in Fig. 12. The Mapping Network, an 8-layer MLP, transforms an input latent vector z into an intermediate, disentangled latent vector w. This unwarps the input distribution, allowing elements of w to correspond more directly to semantic attributes of the final image45. The Synthesis Network then uses this w to produce the final image. It begins with a learned 4 × 4 constant tensor and progressively upsamples it through a series of convolutional blocks. The key mechanism is the Adaptive Instance Normalization (AdaIN) layer within each block, which modulates the feature maps using style information derived from w. This allows the model to control distinct visual features at different scales, from coarse attributes like lesion morphology to fine-grained details like mucosal texture and vascular patterns.
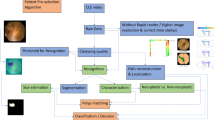
This diagram illustrates the complete pipeline of our study, beginning with data acquisition and preprocessing, followed by the training of the generative adversarial network, the generation of synthetic images, the augmentation of the training set, and culminating in the training and comparative performance evaluation of the downstream detection models.
The discriminator, D, acts as the adversary to the generator, performing the binary classification task of distinguishing between real images from the training set and synthetic images from the generator. Architecturally, the discriminator mirrors the multi-resolution structure of the synthesis network, taking a high-resolution image as input and processing it through a series of convolutional blocks that progressively downsample the feature maps. This design allows the discriminator to analyze the image at various scales, identifying inconsistencies in both fine-grained textures and coarse-level structures69. The network’s final layers aggregate these features to output a single scalar score representing the image’s authenticity, which is then used to calculate the adversarial loss that drives the competitive training dynamics of the entire framework.
To enable the targeted generation of specific lesion types (e.g., flat adenomas), we extend the architecture to a conditional framework by providing a class label vector y as an additional input to both networks. This label y corresponds to one of the three morphological categories (Pedunculated, Sessile, or Subtle) defined in Section 6. This transforms the model’s objective from learning the general data distribution p(x) to learning the class conditional distributions p(x∣y). To integrate this class information effectively, we employ a Projection Discriminator71. Instead of simply concatenating the label to the image features, this technique incorporates the conditionality directly into the discriminator’s final output calculation. It computes an inner product between the image feature vector and a learned, class-specific embedding of the label y. This formulation enforces a strong relationship between image features and their corresponding class, compelling the generator to produce images that are not only realistic but also highly consistent with the target class, a crucial capability for our augmentation goals.
The training of our GAN framework is guided by a composite objective function that combines an adversarial loss with a set of crucial regularization terms designed to ensure training stability and enhance the quality of the generated images. The overall loss function can be expressed as:
where Ladv is the adversarial loss, LR1 is the R1 gradient penalty, and Lpl is the path length regularization term. We set the weighting coefficients λR1 to 10.0 and λpl to 2.0, consistent with standard StyleGAN2-based training protocols with λR1 and λpl serving as their respective weighting coefficients.
Each component of the loss function plays a distinct role. The adversarial loss, Ladv, is based on the non-saturating logistic loss formulation, which has been shown to provide better gradient dynamics and prevent the generator’s gradients from vanishing early in training compared to the original minimax loss43. To further stabilize the adversarial training, we employ the R1 gradient penalty, LR1, which penalizes the discriminator’s gradients only with respect to real data72. This discourages the discriminator from learning a function with overly large gradients, which can destabilize the training process. The final component, path length regularization, Lpl, is applied to the generator. It encourages a smoother and more disentangled latent space \({\mathcal{W}}\) by penalizing deviations from a fixed length step in \({\mathcal{W}}\), thereby improving the predictability and consistency of the image generation process69. Together, this combination of adversarial loss and advanced regularization techniques provides a robust framework for training a high fidelity, stable generative model. The overall research framework, including data acquisition, GAN training, and downstream evaluation, is illustrated in Fig. 12.
The conditional StyleGAN framework was trained using the Adam optimizer (β1 = 0.0, β2 = 0.99) with a learning rate of 0.0025 for both the generator and discriminator, and a batch size of 64. Training was performed on four NVIDIA V100 GPUs for approximately 7 days. The primary metric for model selection was the Fréchet Inception Distance (FID), computed every 10,000 iterations. The final model checkpoint was selected after the FID score had plateaued, indicating stable convergence (as reflected in Table 2).
Evaluation of synthetic image quality
To objectively assess the quality of the generated endoscopic images, we employ a suite of established quantitative metrics, each designed to capture different aspects of image fidelity and diversity. Representative comparisons between real and GAN-synthesized images are shown in Fig. 13.
This grid provides a direct visual comparison between real endoscopic images (left column) and synthetic images (middle, right columns) generated by providing specific class labels to the generator. The examples cover the three morphological categories used for conditioning: (top row) tubulovillous adenoma, (middle row) pedunculated polyp, and (bottom row) sessile polyp. This qualitatively demonstrates the model’s fidelity and its ability to control the generated morphology based on the input condition.
Our primary metric is the Fréchet Inception Distance (FID), which has become a de facto standard for evaluating GAN performance73. The FID score measures the similarity between two sets of images by comparing the statistics of their features as extracted by a pre-trained InceptionV3 network. Specifically, it computes the Wasserstein 2 distance between the multivariate Gaussian distributions fitted to the feature representations of the real and synthetic image sets. By considering both the mean and covariance of these feature vectors, FID simultaneously evaluates the fidelity (realism) and diversity of the generated images. A lower FID score indicates that the distribution of synthetic images is closer to that of the real images, signifying higher quality.
Secondly, we utilize the Inception Score (IS) as a complementary metric74. The IS is designed to measure two desirable properties of generated images: clarity (each image should clearly belong to a single class) and diversity (the model should generate images from all classes equally). It calculates the Kullback-Leibler (KL) divergence between the conditional class distribution p(y∣x) for a generated image x and the marginal class distribution p(y) over the entire generated dataset. A high IS indicates that the generated images are both individually distinct and collectively diverse.
Finally, to specifically evaluate perceptual similarity in a manner that aligns with human vision, we employ the Learned Perceptual Image Patch Similarity (LPIPS) metric75. Unlike traditional metrics such as PSNR or SSIM that operate on raw pixel values, LPIPS computes the distance between the deep feature representations of two images, extracted from a pre-trained neural network (e.g., AlexNet or VGG). This approach has been shown to correlate much better with human judgments of image similarity. A lower LPIPS score signifies that two images are more perceptually similar. A summary of these quantitative evaluation metrics is provided in Table 8.
Experimental design for downstream detection task
To empirically validate the utility of our synthetically generated data, we selected a state-of-the-art, real-time object detection model, YOLOv5m (medium)76, as the downstream task architecture. YOLOv5 is a single-stage detector renowned for its exceptional balance of high accuracy and fast inference speed, making it highly suitable for clinical applications like real-time video analysis in colonoscopy. The architecture is composed of three primary components: a CSPDarknet53 backbone for efficient feature extraction, a Path Aggregation Network (PANet) neck for multi-scale feature fusion, and a YOLO detection head that performs the final bounding box regression and class prediction. We selected YOLOv5 not only for its strong performance as reported in numerous computer vision benchmarks but also for its widespread adoption in the research community, which establishes it as a robust and well-understood baseline for our data augmentation experiments Table 9.
To rigorously evaluate the impact of our GAN-based data augmentation, a carefully designed set of controlled experiments was conducted. We established a strict data separation protocol to ensure rigorous validation and prevent data leakage. Our aggregated corpus (Table 8) was first divided. The CVC-ClinicDB and ETIS-LaribPolypDB datasets were immediately sequestered in their entirety to serve as independent external validation sets. These datasets were never used for training, validation, or hyperparameter tuning of any model (neither the GAN nor the YOLOv5 detector), ensuring a true test of generalization. The remaining datasets (e.g., Kvasir-SEG, CVC-ColonDB, HyperKvasir, PICCOLO, SUN, ASU-Mayo) formed our internal corpus for development.
This internal corpus was then partitioned at the patient and video level, ensuring that all frames from a single procedure or patient were assigned exclusively to one of three splits: a training set (70%), a validation set (15%), or an internal test set (15%). This protocol explicitly prevents identical or sequential frames from contaminating the test sets. To further mitigate the risk of near duplicates, we applied a perceptual hash-based de-duplication algorithm, identifying and removing any images from the validation/test splits that were found to be near identical (hash distance < 5) to images in the training set. This multi-level separation protocol ensures an unbiased evaluation of model performance on both in-distribution and out-of-distribution data.
We then trained four instances of the YOLOv5 detection model, each with an identical architecture and hyperparameter configuration but using a different training data strategy. The experimental groups are defined as follows: Group A (Baseline): The detector was trained using only the original, curated real training images, without any form of data augmentation. This group serves as the absolute baseline to measure the performance of the model on the raw data. Group B (Traditional Augmentation): The detector was trained on the same real images as Group A. This group represents the strong traditional augmentation baseline inherent to the YOLOv5 architecture, which includes Mosaic augmentation, MixUp, and standard geometric/photometric transformations (e.g., rotations, flips, scaling, and color jittering). This represents the current Group C (GAN-based Augmentation): The real training set was augmented by adding a balanced set of our GAN-generated synthetic images, mixed at a 1:1 ratio with the real images. No traditional augmentations were applied to this combined dataset. Group D (Hybrid Augmentation): This group represents a combined approach. The training set consisted of both the original real images and the GAN-generated synthetic images (at a 1:1 ratio), and traditional on-the-fly augmentations were applied to this entire hybrid dataset during training.
All four experimental groups were trained using identical hyperparameters for 300 epochs to ensure a fair comparison of training compute and time. By comparing the performance of these four groups on the internal and external test sets, we can systematically isolate and quantify the specific benefits of our proposed GAN-based augmentation strategy against both a no-augmentation baseline and the standard augmentation methodology.
To quantitatively evaluate and compare the performance of the polyp detection models trained under different augmentation strategies, we adopted a comprehensive set of standard object detection metrics. These metrics are designed to assess various aspects of detection accuracy, including the model’s ability to correctly identify polyps (precision) and its capacity to find all existing polyps (recall). The evaluation was performed on both the internal and external test sets to rigorously assess model performance and generalization. The primary metrics are detailed in Table 10.
Ethical and moral statement
This study used only public, de-identified images; no new human data were collected. Original approvals were obtained by dataset providers as applicable.
Data availability
All datasets used in this study are publicly available from their official sources: Kvasir-SEG, CVC-ClinicDB, CVC-ColonDB, ETIS-Larib, HyperKvasir, PICCOLO, SUN, and ASU-Mayo. Details and download links are provided in the cited references within the manuscript. No private or patient-identifiable data were collected in this study.
Code availability
The custom source code developed for this study, encompassing the data preprocessing pipeline, the Conditional StyleGAN training framework, and the downstream detection experiments, is publicly available for peer review and reproducibility at https://anonymous.4open.science/r/Polyp-StyleGAN-4463/README.md. Access is open and requires no authentication. The models were implemented using the PyTorch deep learning framework (version 1.9.0) and Python (version 3.8). All experiments were conducted on NVIDIA V100 GPUs using CUDA 11.1. Specific configuration files defining the hyperparameters utilized to generate the current datasets are included in the repository. Key training parameters include: • GAN Training: Resolution 512 × 512, batch size 64, Adam optimizer (β1 = 0.0, β2 = 0.99), learning rate 0.0025, λR1 = 10.0, and λpl = 2.0. • Detection Training (YOLOv5): SGD optimizer, learning rate 0.01, momentum 0.937, and weight decay 0.0005, trained for 300 epochs. Full environment requirements (e.g., requirements.txt) and training scripts are provided in the root directory of the repository.
References
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–67 (1990).
Brenner, H., Kloor, M. & Pox, C. P. Colorectal cancer. Lancet 383, 1490–1502 (2014).
Boland, C. R. & Goel, A. Molecular basis of colorectal cancer. Gastroenterology 134, 1279–1290 (2008).
Grady, W. M. & Carethers, J. M. Genomic and epigenetic instability in colorectal cancer. Gastroenterology 135, 1079–1099 (2008).
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2024. CA: A Cancer J. Clin.74, 12–49 (2024).
Davidson, K. W. et al. Screening for colorectal cancer: US Preventive Services Task Force recommendation statement. JAMA 325, 1965–1977 (2021).
Rex, D. K. et al. Colorectal cancer screening: recommendations for physicians and patients from the U.S. Multi-Society task force on colorectal cancer. Am. J. Gastroenterol. 112, 1016–1030 (2017).
Ferlitsch, M. et al. Colorectal polypectomy and emr: European society of gastrointestinal endoscopy (esge) clinical guideline. Endoscopy 49, 270–297 (2017).
Winawer, S. J. et al. Prevention of colorectal cancer by colonoscopic polypectomy. N. Engl. J. Med. 329, 1977–1981 (1993).
Zauber, A. G. et al. Colonoscopic polypectomy and long-term prevention of colorectal-cancer deaths. N. Engl. J. Med. 366, 687–696 (2012).
Schoen, R. E. et al. Colorectal-cancer incidence and mortality with screening flexible sigmoidoscopy. N. Engl. J. Med. 366, 2345–2357 (2012).
Lee, T. H. & Lee, S. K. Factors influencing the adenoma detection rate. Clin. Endosc. 48, 192 (2015).
Kaminski, M. F. et al. Quality indicators for colonoscopy and the risk of interval colorectal cancer. N. Engl. J. Med. 362, 1795–1803 (2010).
Asmad, K. et al. Endoscopist fatigue and its impact on polyp detection and other colonoscopy quality indicators: a systematic review and meta-analysis. Gastroenterol. Res. 11, 398 (2018).
Rex, D. K. et al. Quality indicators for colonoscopy. Gastrointest. Endosc. 81, 31–53 (2015).
Van Rijn, J. et al. Polyp miss rate determined by tandem colonoscopy: a systematic review. Am. J. Gastroenterol. 101, 343–350 (2006).
Soetikno, R. M. et al. Prevalence of nonpolypoid (flat and depressed) colorectal neoplasms in asymptomatic and symptomatic adults. JAMA 299, 1027–1035 (2008).
Chung, G. E. et al. A prospective comparison of two computer-aided detection systems with different false positive rates in colonoscopy. npj Digit. Med. 7, 366 (2024).
Kudo, S.-e et al. Flat and depressed types of early colorectal cancer. Gastroenterology 134, 1581–1590 (2008).
Leggett, B. & Whitehall, V. Colorectal cancer: serrated lesions. Gastroenterology 138, 2054–2064 (2010).
Pohl, H. & Robertson, D. J. Postcolonoscopy colorectal cancers: are they preventable? Annu. Rev. Med. 66, 85–96 (2015).
Hetrick, S. P. et al. Miss rate for colorectal serrated polyps. J. Clin. Gastroenterol. 45, 146–151 (2011).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Poon, C. C. et al. Ai-doscopist: a real-time deep-learning-based algorithm for localising polyps in colonoscopy videos with edge computing devices. NPJ Digit. Med. 3, 73 (2020).
Litjens, G. et al. A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017).
Erickson, B. J., Korfiatis, P., Akkus, Z. & Kline, T. L. Machine learning for medical imaging. Radiographics 37, 505–515 (2017).
Bera, K., Schalper, K. A., Rimm, D. L., Harbhajanka, A. & Madabhushi, A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Hassan, C. et al. New-generation artificial intelligence in gastroenterology. Gut 69, 1655–1663 (2020).
Berzin, T. M., Parasa, S. & Wallace, M. B. Can artificial intelligence solve the grand challenge of colonoscopy? Gastroenterology 157, 25–27 (2019).
Wang, P. et al. Real-time automatic detection system for colorectal polyps during colonoscopy: a multicenter, prospective, randomized controlled study. Gastrointest. Endosc. 90, 657–666 (2019).
Repici, A. et al. Efficacy of a real-time computer-aided detection system in improving adenoma detection rate during colonoscopy: a multicenter, randomized controlled trial. Gastroenterology 159, 512–520 (2020).
Hassan, C. et al. Impact of artificial intelligence on colonoscopy outcomes: a systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 6, 468–478 (2021).
Mori, Y. et al. Computer-aided diagnosis for colonoscopy. Gastrointest. Endosc. 88, 645–657 (2018).
Byrne, M. F. et al. Real-time computer-aided detection of colonic polyps by using a deep-learning model (with video). Gastrointest. Endosc. 89, 529–541 (2019).
Rex, D. K. et al. The asge pivi (preservation and incorporation of valuable endoscopic innovations) on real-time endoscopic assessment of the histology of diminutive colorectal polyps. Gastrointest. Endosc. 73, 419–422 (2011).
Ignjatovic, A., Repici, A. & Hassan, C. Artificial intelligence for the real-time assessment of diminutive colorectal polyps: is the technology ready for prime time? Gastrointest. Endosc. 90, 667–669 (2019).
Berzin, T. M. & Wallace, M. B. Can artificial intelligence move beyond polyp detection to characterization? Gastroenterology 160, 1002–1004 (2021).
Rajpurkar, P., Chen, E., Banerjee, O. & Topol, E. J. AI in health and medicine. Nat. Med. 28, 31–38 (2022).
Willemink, M. J. et al. Preparing medical imaging data for machine learning. Radiology 295, 4–15 (2020).
Cheplygina, V., de Bruijne, M. & Pluim, J. P. Not-so-supervised: a survey of semi-supervised, multi-instance, and transfer learning in medical image analysis. Med. image Anal. 54, 280–296 (2019).
Koh, P. W. et al. Understanding and mitigating the effects of domain shift in machine learning. arXiv preprint arXiv:2104.12933 (2021).
Shorten, C. & Khoshgoftaar, T. M. A survey on image data augmentation for deep learning. J. Big Data 6, 1–48 (2019).
Goodfellow, I. et al. Generative adversarial nets. Adv. Neural Inf. Process. Syst. 27 (2014).
Radford, A., Metz, L. & Chintala, S. Unsupervised representation learning with deep convolutional generative adversarial networks. arXiv preprint arXiv:1511.06434 (2015).
Karras, T., Laine, S. & Aila, T. A style-based generator architecture for generative adversarial networks 4401–4410 (2019).
Yi, X., Walia, E. & Babyn, P. Generative adversarial network in medical imaging: A review. Med. Image Anal. 58, 101552 (2019).
Han, C. et al. Infinite brain MR images: PGGAN-based data augmentation for tumor detection. In Smart Biological Medicine, 21–33 (Springer, 2019).
Frid-Adar, M. et al. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing 321, 321–331 (2018).
Bissoto, A., Perez, F., Fornaciali, M., Avila, S. & Valle, E. (de) constructing bias in skin lesion datasets. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, 3968–3977 (2021).
Wei, J. et al. Stain-adaptive GCN for robust, accurate, and interpretable classification of different-stain histopathology images. Med. Image Anal. 58, 101562 (2019).
Kazeminia, S. et al. Gans for medical image analysis. arXiv preprint arXiv:2009.06267 (2020).
Li, A., Li, Y., Wang, Y. & Zhang, Y. A deep learning-based framework for automated diagnosis of colorectal polyps from endoscopic images. BMC Gastroenterol. 21, 1–11 (2021).
Zhang, R. et al. Polyp-yolov3: a real-time polyp detection system for colonoscopy. Knowl. -Based Syst. 209, 106451 (2020).
Azenfar, A., Balamoun, O., El Adib, B. & El Kettani, C. Artificial intelligence in gastrointestinal endoscopy: a review of the current state and future directions. J. Med. Syst. 44, 1–12 (2020).
Song, E.-M. et al. Real-time optical diagnosis of diminutive colorectal polyps using a deep learning model: a multicentre study. Lancet Digit. Health 2, e516–e526 (2020).
Wallace, M. B., Parasa, S. & Berzin, T. M. Ai in gi: a guide to the current state of artificial intelligence in gastroenterology and hepatology. Gastroenterology 159, 1675–1680 (2020).
Ahmad, O. F. et al. Deep learning in colorectal cancer: a review of the literature. Endosc. Int. Open 9, E729–E738 (2021).
Spadaccini, M. et al. Real-time artificial intelligence for colorectal polyp characterization: a multicenter, international study. Endoscopy 53, 1023–1031 (2021).
Taylor, G. W. & Krishnan, D. Generalization in deep learning. arXiv preprint arXiv:1807.03848 (2018).
Chlap, P. et al. A review of deep learning techniques for medical image segmentation. Med. Phys. 48, 5258–5288 (2021).
Hassan, C. & Wallace, M. B. Artificial intelligence in colonoscopy: a review of the state of the art. Gastrointest. Endosc. 93, 281–292 (2021).
Bargsten, T. et al. Synthetic data augmentation for improved polyp detection in colonoscopy. Gastrointest. Endosc. 91, AB36–AB37 (2020).
Majurski, M. S. et al. Generative adversarial networks for improved classification of colorectal polyps on endoscopic images. J. Med. Imaging 8, 054502 (2021).
Besar, S. N. A., Mohd-Yusof, K. & Ahmad, M. A. Generative adversarial networks for synthetic medical image generation: a review. IEEE Access 10, 57778–57796 (2022).
Calderwood, A. H. & Jacobson, B. C. Validation of the Boston bowel preparation scale. Gastrointest. Endosc. 72, 347–352 (2010).
Dutta, A. & Zisserman, A. Vgg image annotator (via). In Proceedings of the 27th ACM international conference on multimedia, 2586–2589 (2019).
Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 20, 37–46 (1960).
Reinhard, E., Adhikhmin, M., Gooch, B. & Shirley, P. Color transfer between images. IEEE Computer Graph. Appl. 21, 34–41 (2001).
Karras, T. et al. Analyzing and improving the image quality of StyleGAN 8110–8119 (2020).
Karras, T. et al. Alias-free generative adversarial networks. Adv. Neural Inf. Process. Syst. 34, 852–863 (2021).
Miyato, T. & Koyama, M. CGANs with projection discriminator. arXiv preprint arXiv:1802.05637 (2018).
Mescheder, L., Geiger, A. & Nowozin, S. Which training methods for GANs do actually converge? In International Conference on Machine Learning, 3481–3490 (PMLR, 2018).
Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B. & Hochreiter, S. Gans trained by a two-time-scale update rule converge to a local Nash equilibrium. Adv. Neural Inf. Process. Syst. 30 (2017).
Salimans, T. et al. Improved techniques for training GANs. In Adv. Neural Inf. Process. Syst. 2234–2242 (2016).
Zhang, R., Isola, P., Efros, A. A., Shechtman, E. & Wang, O. The unreasonable effectiveness of deep features as a perceptual metric. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 586–595 (2018).
Jocher, G., Chaurasia, A. & Qiu, J. Yolov5 by ultralytics. https://github.com/ultralytics/yolov5 (2020).
Barclay, R. L., Vicari, J. J., Doughty, A. S., Johanson, J. F. & Greenlaw, R. L. Colonoscopic withdrawal times and adenoma detection during screening colonoscopy. N. Engl. J. Med. 355, 2533–2541 (2006).
Johnson, D. A. et al. Optimizing adequacy of bowel cleansing for colonoscopy: recommendations from the US Multi-Society Task Force on Colorectal Cancer. Gastroenterology 147, 903–924 (2014).
Jha, D. et al. Kvasir-seg: A segmented polyp dataset. In International Conference on Multimedia Modeling, 487–498 (Springer, 2020).
Bernal, J. et al. Wm-dova maps for accurate polyp highlighting in colonoscopy: a new validation database. Comput. Med. Imaging Graph. 43, 99–111 (2015).
Tajbakhsh, N., Gurudu, S. R. & Liang, J. Automated polyp detection in colonoscopy videos using shape and context information. IEEE Trans. Med. imaging 35, 630–644 (2015).
Silva, J., Histace, A., Romain, O., Dray, X. & Granado, B. Toward automated polyp detection in colonoscopy videos: a new dataset and a comparison of methods. Comput. Med. Imaging Graph. 38, 101–111 (2014).
Borgli, H. et al. Hyperkvasir, a comprehensive multi-class gastrointestinal dataset for computer-aided diagnosis. Sci. Data 7, 283 (2020).
S’anchez-Peralta, L. F. et al. Piccolo: a multi-centre, specialist-validated dataset of high-quality NBI colonoscopy images. Sci. Data 7, 348 (2020).
Bernal, J. et al. Asu-mayo clinic polyp challenge 2015. In MICCAI 2015 Challenge on Automatic Polyp Detection in Colonoscopy Videos (2015).
Everingham, M., Van Gool, L., Williams, C. K., Winn, J. & Zisserman, A. The Pascal Visual Object Classes (VOC) challenge. Int. J. Comput. Vis. 88, 303–338 (2010).
Acknowledgements
The authors sincerely thank the developers and maintainers of the open source datasets (Kvasir-SEG, CVC-ClinicDB, CVC-ColonDB, ETIS-Larib, HyperKvasir, PICCOLO, SUN, and ASU-Mayo) for making their data publicly available, which enabled the conduct of this research. We also thank the clinical experts who assisted in data annotation and model evaluation for their constructive suggestions. We are grateful to Beijing Qichuang Era Technology Co., Ltd. for providing technical support. This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.L., C.H., X.L. Methodology: Y.L., H.W., X.L. Software: H.T., H.W. Validation: H.T., T.D., B.Y. Formal Analysis: C.H., B.Y. Investigation: Y.L., C.H., T.D. Resources: Y.P., X.L. Data Curation: H.T., T.D. Writing – Original Draft: Y.L., C.H. Writing – Review & Editing: Y.P., H.W., X.L. Visualization: Y.L., T.D. Supervision: Y.P., X.L. Project Administration: Y.P., X.L. Funding Acquisition: X.L. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lin, Y., Huang, C., Tian, H. et al. Improving generalization of polyp detection via conditional StyleGAN augmented training. npj Digit. Med. 9, 113 (2026). https://doi.org/10.1038/s41746-025-02293-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02293-8