Abstract
In pancreatic ductal adenocarcinoma (PDAC), the tumor immune microenvironment (TIME) is central to prognosis and therapeutic response, yet predictive biomarkers are still lacking. Here, we used PDAC patients from TCGA, CPTAC, and multiple centers to investigate and validate the relationship between the prognosis and tumor-infiltrating neutrophil-to-T-cell ratio (NTR) in the TIME, including the correlation of NTR with chemotherapy response in PDAC. We developed seven artificial intelligence models to evaluate NTR in the TIME, then evaluated their performance. The multi-modal ROI-model, designated PORCELAIN (Pancreatic Cancer Tumor-Infiltrating Neutrophil-to-T-Cell Ratio Evaluation with Artificial Intelligence), achieved superior external validation performance, demonstrating significant stratification of overall survival and recurrence-free survival. The NTR in the TIME is a potential biomarker for PDAC prognosis and treatment stratification. Importantly, PORCELAIN provides a noninvasive approach to assess NTR in the TIME, offering potential value for clinical management. Trial registration number: NCT06760234.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC), which accounts for approximately 90% of pancreatic cancers1, exhibits the lowest 5-year relative survival rate among all cancer types2. This poor prognosis is attributed to the limited response to multi-modal treatment strategies, including chemotherapy3. Although the TNM staging system remains the most important criterion for selecting multi-modal treatments, patients within the same stage often show heterogeneous clinical outcomes3. Therefore, novel biomarkers are urgently needed to more accurately predict the clinical outcomes.
Although PDAC is typically poorly immunogenic and highly immunosuppressive, the tumor immune microenvironment (TIME) profiling has demonstrated prognostic potential and predictive value for therapeutic response3. Recent evidence shows that tumor-infiltrating neutrophils can disrupt T-cell function, thereby facilitating tumor growth, worsening clinical outcomes, and contributing to therapy resistance4. This highlights the importance of determining whether a combined assessment of neutrophils and T cells is associated with clinical outcomes. Informed by previous studies suggesting that immune cell ratios may hold prognostic value5, our analysis focused on the clinical relevance of the tumor-infiltrating neutrophil-to-T-cell ratio (NTR) in characterizing the TIME.
Current gold-standard TIME assessment relies on histopathological evaluation, yet this approach proves clinically impractical given that more than 80% of patients present with unresectable disease at initial diagnosis6. Furthermore, spatial intra-tumoral heterogeneity and limited biopsy yield fundamentally constrain comprehensive microenvironment characterization7, highlighting the necessity for noninvasive whole-tumor evaluation strategies. Contrast-enhanced computed tomography (CECT), the primary diagnostic modality for PDAC8, provides three-dimensional tumor visualization. Radiomic features extracted from CECT datasets enable AI-driven predictive modeling for clinical decision support9. Successful applications span early detection8, metastases prediction10,11, treatment-response assessment12, and prognosis evaluation13. Emerging evidence suggests radiomic correlates of immune infiltration in PDAC14,15,16,17; however, existing models exhibit critical limitations, including reliance on magnetic resonance imaging data, lack of biological interpretation, and single-center cohort designs, which substantially impede clinical translation.
This multicenter study pioneers a deep-learning-based multi-modal AI model that integrates pretreatment CECT-derived AI-based radiological feature features and clinical parameters to evaluate the tumor-infiltrating NTR, a novel potential TIME biomarker validated through public databases. Our PORCELAIN (Pancreatic Cancer Tumor-Infiltrating Neutrophil-to-T-Cell Ratio Evaluation with Artificial Intelligence) model demonstrates robust prognostic performance across multicenter validation cohorts, establishing a paradigm for noninvasive TIME assessment to advance precision oncology in PDAC.
Results
Patient characteristics
The study framework is schematically presented in Fig. 1. Thirty-three patients (Center A:26, Center B:3, Center C:4) were excluded due to inadequate pretreatment CECT quality, with 14 additional Center A cases excluded for lost follow-up. The final analysis included 402 PDAC patients (176 females, 226 males) from three centers and two public databases. Demographic and clinical characteristics are summarized in Table 1. All patients were divided into four cohorts: TCGA-PAAD cohort (average age: 64.69 ± 10.94; n = 143), CPTAC-3 cohort (average age: 65.00 ± 10.90; n = 59), Center A cohort (average age: 62.99 ± 10.23; n = 158), and external validation cohort (Center B: 36, Center C: 6; total n = 42; average age: 64.90 ± 10.89). Tables S1–S3 provide a detailed overview of the clinical characteristics of patients in both the Center A cohort and external validation cohort. These clinical features were input into the deep learning model, which then automatically assigned weights using its adaptive feature integration mechanism.
a Explore TIME biomarker associated with clinical outcomes. b development and validation of artificial intelligence models; c interpretation and clinical application of PORCELAIN. TIME tumor immune microenvironment, NTR neutrophil-to-T-lymphocyte ratio, IHC immunohistochemical, CECT contrast-enhanced computed tomography, ROI region of interest.
Explore TIME signature associated with clinical outcomes
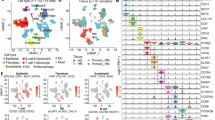
MCP-counter analysis quantified endothelial cells, fibroblasts, and eight immune cell populations in TCGA-PAAD and CPTAC-3 cohorts (Tables S4 and S5). After excluding patients with OS < 30 days or PFS = 0, Cox regression analysis revealed that T-cell abundance showed a protective effect for OS in TCGA cohort (HR = 0.73, 95% CI 0.56–0.96; Fig. S1a) and elevated NTR predicted worse OS (TCGA: HR = 1.51, 95% CI 1.07-2.13; CPTAC: HR = 2.00, 95% CI 1.08–3.70; Fig. S1a, S1c) and PFS (TCGA: HR = 1.55, 95%CI 1.07–2.26; CPTAC: HR = 2.11, 95% CI 1.04–4.28; Fig. S1b, S1d). Multivariate analysis confirmed NTR as an independent predictor of OS (TCGA: HR = 1.52, 95% CI 1.06–2.16; CPTAC: HR = 2.81, 95% CI 1.40–5.65; Fig. S1e, S1g) and PFS (TCGA: HR = 1.47, 95% CI 1.01–2.16; CPTAC: HR = 3.27, 95% CI 1.45–7.34; Fig. S1f, S1h).
To further determine the association between NTR and prognosis, we performed survival prognosis analyses. Using TCGA OS data, the optimal NTR cutoff was determined as 2.47, and patients in both TCGA and CPTAC cohorts were stratified into high-NTR and low-NTR groups (Tables S4, S5). Kaplan–Meier analysis demonstrated patients in high-NTR group had significantly shorter OS and PFS compared to those with low-NTR in both cohorts (p < 0.01, Fig. 2a–d). For chemotherapy response, non-responders (stable/progressive disease; NR) exhibited significantly higher NTR compared to responders (complete/partial response; R) in TCGA cohort (p < 0.05, Fig. S2a). Given the ambiguous documentation of chemotherapy response, CPTAC cohort was excluded in this analysis.
Differential (a) OS and (b) PFS stratified by NTR status in the TCGA cohort; corresponding survival disparities for (c) OS and (d) PFS in the CPTAC validation set; Significant survival disparities in (e) OS and (f) RFS stratified by NTR status within the Center A cohort; consistent prognostic stratification for (g) OS and (h) RFS in the external validation cohort. NTR neutrophil-to-T-lymphocyte ratio, OS overall survival, PFS progression-free survival, DFS recurrence-free survival.
Validation of NTR clinical value in multicenter data
Forest plots validating NTR’s prognostic value across three centers are provided in Fig. S3. Multivariate Cox regression analyses confirmed NTR as an independent predictor of OS and RFS. Since NTR in these patients was determined using immunohistochemistry (IHC), we redefined the thresholds accordingly. The optimal cutoff values for NTR-count were 0.67 (OS) and 0.47 (RFS), and hence we used the midpoint, 0.57 for stratification in Center A and external validation cohorts (Tables S6 and S7). Kaplan–Meier analysis revealed significant differences in OS and RFS between high- and low-NTR groups (p < 0.001, Fig. 2e–h). PDAC recurrence is attributed to chemoresistance, so patients were classified into R and NR according to median RFS (10 months18). Interestingly, the NTR was significantly higher in the NR group than in the R group in both Center A and external validation cohort (p < 0.001, Fig. S2b, S2c). Further Pearson correlation analysis showed no significant association between tumor-infiltrating NTR and peripheral NLR in either cohort (Center A: r = 0.02, p > 0.05; External validation: r = 0.27, p > 0.05; Fig. S4).
Development and validation of AI models
Seven AI models were developed for NTR status prediction: Cuboid-model, ROI-model, ROIex-3mm-model, ROIex-5mm-model, multi-modal Cuboid-model, multi-modal ROIex-model, and multi-modal ROI-model. Table 2 and Fig. 3a–c summarizes the discrimination performance across models. As the ROIex-3 mm model demonstrated more balanced performance relative to the ROIex-5 mm model, we adopted the 3-pixel dilation scheme for constructing the multi-modal ROIex model. The Cuboid-model, ROI-model, ROIex-3mm-model, ROIex-5 mm-model, multi-modal Cuboid-model, multi-modal ROIex-model, and multi-modal ROI-model achieved AUCs of 0.81 (95% CI 0.71–0.91), 0.85 (95% CI 0.76–0.94), 0.83 (95% CI 0.70–0.96), 0.76 (95% CI 0.65–0.87), 0.86 (95% CI 0.77–0.95), 0.84 (95% CI 0.75–0.93), and 0.90 (95% CI 0.84–0.94) in the external validation dataset, respectively (Table 2 and Fig. 3c). DeLong test was performed to assess differences in AUC values between the models (Fig. S5a). Differences did not reach statistical significance, likely owing to the limited external validation sample19,20,21.
Radar plots of model performance in a training, b internal validation, and c external validation datasets; clinical stratification by PORCELAIN showing d OS, e RFS, and f chemosensitivity disparities. OS overall survival, RFS recurrence-free survival.
The evaluation metrics, including ACC, sensitivity, specificity, and F1-score, were detailed in Table 2. In the external validation cohort, the Cuboid model reached an accuracy of 0.77 (95% CI, 0.67–0.87), with a sensitivity of 0.82, a specificity of 0.71, and an F1-score of 0.80. The ROIex-3mm-model showed a comparable accuracy of 0.77 (95% CI, 0.64–0.90), with the same sensitivity (0.82), specificity (0.71), and F1-score (0.80). The ROIex-5mm-model achieved an accuracy of 0.72 (95% CI, 0.61–0.82), with a sensitivity of 0.77, specificity of 0.65, and an F1-score of 0.76. The ROI-model performed slightly better, with an accuracy of 0.79 (95% CI, 0.70–0.89) and a balanced sensitivity (0.82) and specificity (0.76), yielding an F1-score of 0.82. The multimodal Cuboid-model improved performance to an accuracy of 0.82 (95% CI, 0.73–0.91), with a sensitivity of 0.86, specificity of 0.76, and an F1-score of 0.84. The multimodal ROIex-model achieved an accuracy of 0.79 (95% CI, 0.70–0.89), with sensitivity of 0.86, specificity of 0.71, and an F1-score of 0.83. The multimodal ROI-model showed the best overall performance, reaching an accuracy of 0.85 (95% CI, 0.76–0.93), with balanced sensitivity (0.86), specificity (0.82), and an F1-score of 0.86. Figure S5b–S5h shows the confusion matrices of all models. These analyses indicate that, although AUC differences were not significant, the multi-modal ROI-model consistently outperformed comparator models across all other metrics, including accuracy, sensitivity, specificity, and F1-score, demonstrating its reliability and robustness. To facilitate reference to the multi-modal ROI-based model, we designated it as Pancreatic Cancer Tumor-Infiltrating Neutrophil-to-T-Cell Ratio Evaluation with Artificial Intelligence (PORCELAIN).
Clinical value and model interpretation of PORCELAIN
Survival analysis further substantiated the clinical value of PORCELAIN in external validation. Patients stratified by model-predicted NTR status exhibited significant divergence in both OS (p = 0.011, Fig. 3d) and RFS (p = 0.014, Fig. 3e) between high- and low-NTR groups. Furthermore, assessment of chemotherapeutic response revealed marked intergroup disparities, with responders exhibiting a significantly higher proportion of low-NTR status (p = 0.006, Fig. 3f). SHAP analysis revealed the relative importance of clinical features driving predictions in the clinical branch of PORCELAIN. The importance plot identified N-stage as the strongest predictor, followed by serum CA19-9 and CEA levels (Fig. 4a). Directional SHAP effects showed that lower N-stage, lower CA19-9, and lower CEA were consistently associated with low-NTR classification (Fig. 4b). Analyses of the remaining validation folds confirmed these patterns, with N-stage and CA19-9 ranking among the top contributors across models (Fig. S6). Grad-CAM (Fig. 4c) revealed ROI-model focus on the tumor core and margin, whereas the Cuboid-model, ROIex-3mm-model, and ROIex-5mm-model extended attention to non-pancreatic regions, potentially underlying its reduced accuracy. In all three folds, SHAP analysis identified the Deep Learning Signature (DL_Signature) as a top contributor, quantitatively demonstrating the substantial impact of image-derived deep learning features and validating their role in the fusion framework (Figs. 4d, e and S7).
SHAP analysis of clinical features: importance (a) and effect direction (b). c Grad-CAM highlights key imaging regions. Fusion model (PORCELAIN) SHAP analysis: importance (d) and effect direction (e) of clinical features and DL_Signature. SHAP SHapley Additive exPlanations, DL_Signature Deep Learning Signature, N N-stage, Plt Platelet, LMR Peripheral Blood Lymphocyte-to-Monocyte Ratio, Hb Hemoglobin, NLR Peripheral Blood Neutrophil-to-Lymphocyte Ratio, Glb Globulin, AGR Peripheral Blood Albumin-to-Globulin Ratio, Alb Albumin, mono Monocyte, PLR Peripheral Blood Platelet-to-Lymphocyte Ratio, ALP Alkaline Phosphatase, AST Aspartate Aminotransferase.
Discussion
Our multicenter study establishes tumor-infiltrating NTR as a novel potential prognostic biomarker in PDAC through integrative analysis of multi-omics data. Importantly, elevated NTR was consistently associated with reduced overall survival, accelerated disease progression, and greater chemotherapy resistance across all cohorts. Building upon this biological foundation, we developed the PORCELAIN model, which enables noninvasive NTR classification through AI-based CECT features combined with clinical parameters, achieving superior predictive performance. Applied in practice, this multi-modal model may provide clinicians with a decision-support framework for optimizing therapeutic strategies, including the design of personalized neoadjuvant regimens for borderline resectable PDAC.
These findings extend current understanding of pancreatic cancer immunobiology. First, while prior studies primarily focused on peripheral immune indices like NLR22,23,24, our data highlight the superior prognostic value of tumor-infiltrating NTR over circulating biomarkers. The lack of correlation between tumor NTR and peripheral NLR suggests distinct biological mechanisms governing local versus systemic immune responses. Second, although some experimental studies have demonstrated that both tumor-infiltrating neutrophils and T-cells exert distinct effects on tumor prognosis25,26,27,28, our work demonstrates that neither the neutrophil nor T-cell infiltration level within TIME emerge as independent prognostic factors. We propose that combinatorial metrics (NTR) better captures the dynamic interplay within TIME. This aligns with emerging evidence that tumor-associated neutrophils promote T-cell exhaustion by upregulating the expression of Nectin229. Moreover, lactate accumulation in hypoxic niches induces metabolic competition among immune cell subsets30. Third, our results point to a potential link between NTR and chemotherapy resistance. This may drive by both neutrophil-mediated processes and T-cell dysfunction. Chemotherapy has been shown to trigger neutrophil recruitment and the formation of neutrophil extracellular traps (NETs). Two NET-associated proteins, integrin-αvβ1 and MMP9, can activate TGF-β signaling, which promotes epithelial–mesenchymal transition (EMT) in cancer cells and enhances treatment resistance31. In parallel, metabolic reprogramming in the tumor microenvironment leads to lactate accumulation that suppresses histone acetylation, while mitochondrial dysfunction reduces acetyl-CoA availability. Together, these changes limit effector gene expression in T cells and further drive resistance32,33.
Radiomics has advanced noninvasive characterization of tumor immune infiltration34,35,36, but traditional pipelines depend on predefined, formula-driven descriptors (e.g., intensity statistics, texture features). Our convolutional neural network (CNN), in contrast, learns hierarchical representations directly from ROI pixel intensities. Demircioğlu’s systematic review noted that handcrafted features are “generic” and may not capture all relevant patterns, while deep networks adaptively learn features from the data itself37. Thus, our framework differs fundamentally from standard workflows (e.g., radiomics + XGBoost) by replacing human-engineered features with deep CNN-derived representations.
Deep learning has repeatedly been shown to surpass conventional radiomics. Hosny et al. demonstrated that CNN-based features yielded superior prognostic accuracy in lung cancer compared with radiomic signatures38. Similarly, in prostate cancer, a deep learning model significantly outperformed a radiomics-SVM model in predicting clinically significant disease among patients with gray-zone PSA levels39. A systematic review further reinforced this trend, reporting that deep learning achieved higher AUC values than radiomics in nearly three-quarters of studies37. Together, these findings validate our choice of deep CNN-derived representations, enabling the extraction of fine-grained spatial signals beyond the reach of handcrafted descriptors.
EfficientNet-B0 was selected for its balance of accuracy and computational efficiency. It contains 5.3 million parameters, substantially fewer than most modern CNNs, reducing overfitting risks while maintaining strong feature extraction capability. As Sadu et al. highlighted40, this architecture “extracts detailed spatial features while maintaining computational efficiency”. Its effectiveness in small or cropped medical imaging regions makes it well suited to our ROI input dimensions.
We examined how different tumor boundary definitions influenced PDAC model performance, including cuboid, ROI, and 3 mm/5 mm ROI expansions. Performance followed a nonlinear pattern: the ROIex-3 mm model surpassed the cuboid model, whereas the ROIex-5 mm model declined. The intratumoral ROI model consistently achieved the strongest results. The ROIex-3 mm model likely benefited from capturing PDAC’s infiltrative growth. Tissue immediately surrounding the tumor often contains prognostic features, such as lymphovascular invasion, perineural spread, and desmoplastic stroma, and a 3 mm expansion allows these signals to be incorporated41. By contrast, a 5 mm expansion commonly includes adjacent organs or heterogeneous structures, introducing noise that diminishes representational quality. Importantly, the pure intratumoral ROI model performed best overall, indicating that PDAC’s intrinsic texture, marked by hypovascularity and heterogeneous enhancement, remains the most reliable discriminator. Even minimal inclusion of peritumoral tissue may add confounding information, highlighting the value of precise intratumoral segmentation.
It is notable that the Multimodal Cuboid-model outperforms the Multimodal ROIex-model. A plausible explanation is that the ROIex-3mm features overlap with clinical variables, leaving little additional information for the fusion layer to exploit42. In contrast, the Cuboid model generates features that are more distinct from the clinical space, enabling stronger complementarity between modalities43. The 3 mm mask expansion may also introduce boundary-related noise that is difficult to suppress in a small dataset, whereas the simpler Cuboid geometry offers a more stable foundation for multimodal integration44.
The SHAP analysis indicated that within the clinical branch of PORCELAIN, N-stage, CA19-9, and CEA carried the highest feature weights, consistent with previous findings. N-stage, reflecting the degree of lymph node metastasis, is critical given the central role of lymph nodes in immune regulation. Metastatic involvement significantly impairs anti-tumor immune responses, as primary lesions with lymph node metastasis show reduced T-cell activation, cytotoxicity, and proliferation, alongside greater susceptibility to T-cell exhaustion45. Neutrophils also contribute to metastasis by releasing proteases such as MMP9, serine proteases, neutrophil elastase, and cathepsin G, which degrade the extracellular matrix and facilitate tumor spread5. As a well-recognized biomarker in pancreatic cancer, CA19-9 is associated with an immunosuppressive microenvironment; elevated CA19-9 promotes IL-33+ Treg infiltration, leading to T-cell exhaustion46. Similarly, clinical studies have shown that higher serum CEA levels inversely correlate with T-cell infiltration in the tumor microenvironment47. Collectively, these observations provide biological support for the clinical feature weighting identified in PORCELAIN.
In addition to investigating the relationship between deep learning models and TIME, another approach involves applying deep learning analyses to directly predict prognosis or therapeutic response, with numerous studies reporting favorable outcomes in pancreatic cancer and other malignancies48,49,50,51,52. However, the lack of biological interpretation remains a critical challenge of these models, which constitutes a significant barrier to their clinical translation. In contrast, our work is grounded in substantial biological evidence53,54,55,56,57 and bioinformatic analyses from public databases, which collectively demonstrate the role of NTR in OS, disease progression, and treatment response, thereby providing biological rationale for PORCELAIN’s predictions.
Our study has several limitations. First, the retrospective design introduces potential selection bias, though this was mitigated through multicenter enrollment and rigorous exclusion criteria. Second, although this was a multicenter study, the relatively low incidence of PDAC resulted in limited sample sizes at each participating center. To address this, we will prioritize prospective validation of PORCELAIN to verify its generalizability and reproducibility in larger cohorts. Specifically, we plan to conduct a Phase II clinical trial to prospectively evaluate the clinical value of PORCELAIN in two populations: patients with borderline resectable PDAC and those with unresectable PDAC. Third, chemotherapeutic response evaluation in multicenter patients necessitated assessment of postoperative recurrence time due to the requirement for pretreatment tumor specimen acquisition. While this methodology has been validated in prior landmark studies, prospective trials remain imperative to confirm PORCELAIN’s predictive capacity for therapeutic outcomes.
In summary, through comprehensive analysis of public databases and multicenter datasets, we identified tumor-infiltrating NTR as a potential biomarker for predicting survival, progression (recurrence), and chemotherapy response in PDAC. Moreover, our multi-modal AI model, PORCELAIN, demonstrated the highest performance in predicting tumor-infiltrating NTR status. This model enables noninvasive evaluation of NTR status and prediction of clinical outcomes. This is clinically significant for PDAC, given its low surgical resection rates and the difficulty of obtaining tissue samples for pathological evaluation. These findings have the potential to inform therapeutic decision-making, thereby facilitating personalized and precise treatment strategies to improve clinical outcomes in PDAC management.
Methods
Study design
This retrospective multicenter investigation received ethical approval from institutional review boards at the Second Affiliated Hospital, Zhejiang University School of Medicine (ZJU2H, No. 2024-0753), the Fourth Affiliated Hospital, Zhejiang University School of Medicine (ZJU4H, No. K2024147), and Hangzhou Hospital of Traditional Chinese Medicine (HZTCM, No. 2025KLL131), and was prospectively registered at ClinicalTrials.gov (NCT06760234). Informed consent was waived since this was a retrospective study. We rigorously adhered to the TRIPOD + AI guidelines58 for transparent reporting of AI-based prediction models.
Database curation
Our analysis utilized data from two NIH-curated genomic repositories: (1) TCGA-PAAD cohort (n = 143), sourced from “ductal and lobular neoplasms”, was used for TIME biomarker discovery; (2) CPTAC-3 cohort (n = 59), sourced from “pancreas”, was used for TIME biomarker validation. RNA-seq counts and transcripts-per-million (TPM)-normalized expression data, together with clinical metadata, were retrieved from the GDC portals (https://portal.gdc.cancer.gov). Normal tissue specimens were systematically excluded from both cohorts.
Patient cohorts and clinical parameters
A multicenter cohort of 247 treatment-naïve PDAC patients undergoing curative resection with adjuvant chemotherapy was assembled from three centers (Center A: ZJU2H; Center B: ZJU4H; Center C: HZTCM) between March 2013 and May 2024. The inclusion criteria were: (1) histopathological confirmed PDAC; (2) curative resection followed by adjuvant chemotherapy. The exclusion criteria were: (1) suboptimal CT image quality; (2) incomplete clinicopathological data; (3) concurrent or metachronous malignancies; (4) loss to follow-up.
Demographic and clinical parameters encompassed age, gender, BMI, preoperative complete blood count (CBC) results, serum tumor biomarkers, liver function tests. Survival endpoints included overall survival (OS), calculated from the surgery date to last follow-up, and recurrence-free survival (RFS), calculated from the surgery date to event occurrence. Tumor staging was performed according to the American Joint Committee on Cancer (AJCC) 8th edition criteria59.
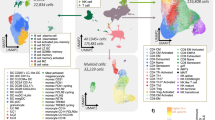
TIME profiling quantification
TPM normalized expression matrices from TCGA-PAAD (discovery cohort) and CPTAC-3 (validation cohort) were analyzed using the Microenvironment Cell Populations-counter (MCP-counter) R package (V1.2.0)60. This algorithm employs lineage-specific gene signatures through weighted linear regression to generate absolute abundance scores for endothelial cells, fibroblasts, and eight immune cell subpopulations within the tumor microenvironment. All abundance scores underwent log2(X + 1) variance-stabilizing transformation prior to subsequent analysis. Prognostic biomarkers were identified by univariate Cox proportional hazards regression (p < 0.05 threshold), followed by multivariate adjustment for clinicopathological covariates. Survival outcomes, including OS and progression-free survival (PFS), were evaluated using Kaplan–Meier estimates, with cohort-stratified survival curves and log-rank testing. Optimal prognostic stratification thresholds for biomarkers were algorithmically determined in the discovery cohort using maximally selected rank statistics, implemented through the “surv_cutpoint” function in the survminer R package.
Immunohistochemical validation of NTR
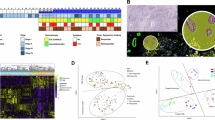
Archival Formalin-fixed paraffin-embedded tumor specimens from participating centers underwent standardized immunohistochemical (IHC) staining protocols. All samples were sectioned at 4-μm thickness and processed for IHC staining as previously described. Each staining run included both negative and positive controls. Following deparaffinization in xylene and rehydration in ethanol, sections were blocked with endogenous peroxidase (3% H2O2 solution) and underwent heat-induced epitope retrieval in citrate buffer (pH 6.0) for 23 min. Sections were blocked with 3% bovine serum albumin (Solarbio Science & Technology) for 30 min. The slides were incubated overnight at 4 °C with antibodies against human CD8 (Cell Signaling Technology, #85336, 1:200) or CD66b (Abcam, #ab300122, 1:10000), followed by incubation with horseradish peroxidase-conjugated secondary anti-rabbit antibodies (Abcam, #ab205718, 1:1000) at room temperature for 50 min. The reaction was visualized using a diaminobenzidine (DAB) substrate kit (Solarbio Science & Technology), and nuclei were counterstained with hematoxylin. DAB staining was monitored under a light microscope and terminated upon achievement of yellowish-brown coloration. All sections were digitized using a Digital Pathology Slide Scanner at 40× magnification (Konfoong Bioinformation Tech, KF-PRO-005). Dual independent evaluation was performed by gastrointestinal pathologists (Y.Z., 9 years’ experience; J.T., 12 years’ experience), blinded to clinical outcomes, with discrepant cases adjudicated by a senior pathologist. Digital whole-slide imaging was conducted using K-Viewer platform (version 1.5.3.1, KFBIO), employing systematic random sampling to select five most representative regions encompassing tumor-stroma interface and core regions. The density of immune cells was then measured at high power (200×) magnification. Neutrophils and T-cells in each area were quantified and expressed as the number of cells per field, and the average of five fields is used to represent the count of immune cells. The NTR was computed by dividing the neutrophil count by the cytotoxic T-cell count. Optimal NTR stratification cutoff was derived through receiver operating characteristic analysis, maximizing Youden’s index in the Center A training cohort.
CECT images acquisition and preprocessing
Pretreatment DICOM-format CECT images were acquired from picture archiving and communication systems. CT acquisition parameters included: 120 kV tube voltage; 170–210 mAs tube current; 0.5–0.6 s rotation time; and slice thickness ranging from 0.75 to 5 mm.
DICOM files were converted to NIfTI format (.nii.gz) using 3D slicer (V5.6.2), preserving affine orientation and voxel spacing for downstream processing. Portal venous phase images were analyzed using a window width of [40, 350] Hounsfield Units (HU), and no intensity interpolation was applied during preprocessing. Region of interest (ROI) segmentation was performed independently by two hepatopancreatobiliary surgeons (Y.F. with 9 years and Y.S. with 4 years of abdominal CT interpretation experience) using 3D Slicer (V5.6.2). To assess segmentation reliability, 50 cases from Center A were independently annotated by two observers, each repeating the task after a two-week interval. This produced four ROI sets for analysis. Intra-observer Dice scores were high for both annotators, 0.87 (95% CI 0.85–0.89) for Y.F. and 0.88 (95% CI 0.86–0.90) for Y.S., while inter-observer Dice values were 0.87 (95% CI 0.83–0.89) and 0.86 (95% CI 0.82–0.88) for the paired sessions (Fig. S6a). PyRadiomics-based extraction of 1409 features followed by ICC analysis showed strong reproducibility: ≥77.9% of features had ICC > 0.75 within observers and ≥72.0% between observers (Fig. S6b–e). These findings demonstrate acceptable consistency in both intra- and inter-observer performance. Discrepancies were resolved through consensus discussion. A senior radiologist (Y.C. with 15 years of abdominal CT experience) verified ROI quality and made necessary adjustments.
From each scan, we extracted the smallest 3D bounding box enclosing the ROI mask and produced two datasets: (i) a Cuboid retaining tumor and peritumoral context; and (ii) a Masked-cuboid in which extra-tumoral voxels were zeroed to isolate the lesion. To standardize inputs for deep learning without intensity resampling, each bounding box volume was center-cropped and/or zero-padded in voxel space to a fixed size of 100 × 100 × 30 (voxels). Intensities were then normalized within each bounding box using statistics computed on non-zero voxels (Eq. 1).
Where \({\mu }_{{bbox}}\) and \({\sigma }_{{bbox}}\) denote the mean and standard deviation within the bounding box non-zero mask.
A fixed-distance dilation strategy, widely used in radiomics, was also applied to define the peritumoral region in this study. Morphological dilation produced peritumoral ROIs that retained the geometry of the tumor. Based on established studies61,62, two distances were examined: 3 mm for the immediate margin and 5 mm for a broader stromal zone. All images were resampled to 1 × 1 × 1 mm prior to dilation to maintain consistent physical dimensions.
Imaging branch (AI-based radiological feature extraction)
The complex imaging characteristics of PDAC and the limitations of CECT reduce the effectiveness of conventional radiomics feature extraction. In contrast, AI-based approaches automatically learn high-level features directly from imaging data, avoiding reliance on hand-crafted descriptors. For our model, we selected EfficientNet-B0 as the backbone due to its strong performance in image classification under resource-constrained conditions and moderate dataset sizes63. Its scalable design offers an optimal balance between accuracy and computational efficiency, making it well-suited for PDAC feature extraction and classification.
To adapt EfficientNet-B0 for CECT analysis, we integrated attention mechanisms at multiple stages, forming the EN-PCT framework. Specifically, EfficientNet-B0 served as the backbone, followed by global average pooling (GAP) to produce scan-level feature vectors, consistent with Network-in-Network64 principles. To enhance tumor-related representation within the ROI, Squeeze-and-Excitation (SE) blocks were added after stages 2 and 365. These modules recalibrate channel responses, amplifying lesion-associated features while minimizing background influence (already reduced by ROI preprocessing). The pooled features were passed through a fully connected (FC) layer with ReLU activation and Batch Normalization to generate the imaging embedding \({z}_{{img}}\), as defined in Eq. (2).
All encoder inputs were tensors of dimension 1 × 100 × 100 × 30 (channels × height × width × slices).
To address data scarcity, we applied on-the-fly augmentation, including random rotations (±15°), horizontal and vertical flips (probability 0.5), and brightness jitter (±10%). Batch Normalization was used across all convolutional and FC layers66, and dropout (0.3) was applied in the imaging head67. Model optimization employed AdamW (\({\beta }_{1}\) = 0.9, \({\beta }_{2}\) = 0.999, weight decay 1 × 10−4) with an initial learning rate of 2 × 10−3, cosine annealing with warm restarts (SGDR)68,69, and early stopping based on validation AUC.
Clinical branch (tabular modeling)
All collected clinical variables were included for analysis. Preprocessing was performed as follows: missing values were imputed using the median for continuous variables and the mode for categorical variables; biomarkers (e.g., CA19-9, CEA) were transformed using log(1 + x); continuous variables were standardized with z-scoring; and categorical variables were one-hot encoded. Structured clinical data were then encoded using TabNet with attentive feature selection70. Model parameters were fixed at 3 decision steps, feature transformer width of 64, relaxation parameter of 1.5, and sparsity regularization coefficient \({\lambda }_{{sparse}}=1\times {10}^{-3}\). The clinical embedding was defined as Eq. (3).
Batch Normalization and dropout (rate = 0.4) were applied throughout the clinical branch66,67.
Multi-modal feature fusion and classification (fixed, feature-level concatenation)
We applied feature-level fusion with modality-specific alignment in a fixed and reproducible manner. Before concatenation, each embedding was normalized with batch normalization, and the higher-dimensional imaging embedding was linearly projected to match the clinical dimension, preventing dominance by a single modality. Concatenation followed a fixed order [clinical; imaging]. Specifically, the imaging embedding was projected to 128 dimensions, yielding the fused vector (Eqs. 4–6).
The fused representation was processed through a single-hidden-layer MLP with ReLU activation, batch normalization, and dropout (0.5), followed by a sigmoid output (Eq. (7)).
Training minimized binary cross-entropy loss (Eq. 8).
This design ensured scale alignment across heterogeneous clinical and imaging embeddings, consistent with established multimodal integration approaches66,67,71.
Model variants
We trained two multimodal model variants within a unified pipeline: (i) the ROI model, which used masked cuboid inputs focusing on the tumor, and (ii) the Cuboid model, which used cuboid inputs encompassing both tumor and peritumoral regions. The clinical branch, fusion module, and classifier were identical across variants; only the imaging input differed. Both models employed EfficientNet-B0 with SE blocks, global average pooling, and a fixed input size of 100 × 100 × 30 voxels63,64,65.
Model evaluation
The models were primarily evaluated using the area under the receiver operating characteristic curve (AUC) with 95% confidence intervals (CI). Additionally, comprehensive assessment of model performance was conducted through calculations of accuracy (ACC), sensitivity, specificity, and F1-score. These metrics were computed on both the training dataset and internal validation dataset to verify the generalizability of the models to unseen data. Three-fold cross-validation was implemented to ensure the robustness of the results and statistical validity. For performance enhancement, hyperparameter tuning was performed using grid search and random search techniques. Evaluated hyperparameters included learning rate, batch size, number of layers, and whether regularization was applied. The optimal set of hyperparameters was determined based on the highest validation accuracy and the best trade-off between sensitivity and specificity.
To assess robustness and generalizability, the final models were tested on an independent external validation dataset encompassing diverse patient populations and imaging conditions. Performance was measured using AUC, accuracy, sensitivity, specificity, F1-score, and confusion matrices to determine their potential clinical utility. To improve interpretability, Grad-CAM72 was applied to visualize image regions that guided predictions, while SHapley Additive exPlanations (SHAP) analysis73 quantified how clinical and imaging features contributed to model outputs.
Statistical analysis
Analyses were executed in R (v4.3.2) and GraphPad Prism (v8.0.1). Continuous variables were analyzed using Student’s t-test (parametric) or Mann–Whitney U test (non-parametric), with distribution normality assessed via Shapiro–Wilk test. Correlation coefficients were calculated using Pearson’s method. Fisher’s exact test was used to analyze categorical variables. Statistical significance was defined as p < 0.05.
Data availability
The datasets generated during and analyzed during the current study are available by the corresponding author, upon reasonable request.
Code availability
The implementation code for these models depends on internal tooling and infrastructure that are currently under patent protection and therefore cannot be publicly released. All experimental protocols and implementation details have been described in detail in the “Methods” section to enable reproducibility using non-proprietary libraries.
References
Stoffel, E. M., Brand, R. E. & Goggins, M. Pancreatic cancer: changing epidemiology and new approaches to risk assessment, early detection, and prevention. Gastroenterology 164, 752–765 (2023).
Siegel, R. L., Kratzer, T. B., Giaquinto, A. N., Sung, H. & Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 75, 10–45 (2025).
Wang, J. et al. Consensus, debate, and prospective on pancreatic cancer treatments. J. Hematol. Oncol. 17, 92 (2024).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Cai, W. et al. Neutrophils in cancer: at the crucial crossroads of anti-tumor and pro-tumor. Cancer Commun. 45, 888–913 (2025).
Nakamura, K. et al. An exosome-based transcriptomic signature for noninvasive, early detection of patients with pancreatic ductal adenocarcinoma: a multicenter cohort study. Gastroenterology 163, 1252–1266.e1252 (2022).
Jiang, Y. et al. Non-invasive tumor microenvironment evaluation and treatment response prediction in gastric cancer using deep learning radiomics. Cell Rep. Med. 4, 101146 (2023).
Cao, K. et al. Large-scale pancreatic cancer detection via non-contrast CT and deep learning. Nat. Med. 29, 3033–3043 (2023).
Sohn, J. H. & Fields, B. K. K. Radiomics and deep learning to predict pulmonary nodule metastasis at CT. Radiology 311, e233356 (2024).
Shi, S. et al. Development and validation of a deep learning radiomics model with clinical-radiological characteristics for the identification of occult peritoneal metastases in patients with pancreatic ductal adenocarcinoma. Int J. Surg. 110, 2669–2678 (2024).
Zhao, B. et al. Development of a radiomics-based model to predict occult liver metastases of pancreatic ductal adenocarcinoma: a multicenter study. Int. J. Surg. 110, 740–749 (2024).
Tomaszewski, M. R. et al. Delta radiomics analysis of Magnetic Resonance guided radiotherapy imaging data can enable treatment response prediction in pancreatic cancer. Radiat. Oncol. 16, 237 (2021).
Wang, K. et al. Delta radiomic features predict resection margin status and overall survival in neoadjuvant-treated pancreatic cancer patients. Ann. Surg. Oncol. 31, 2608–2620 (2024).
Bian, Y. et al. Machine learning for MRI radiomics: a study predicting tumor-infiltrating lymphocytes in patients with pancreatic ductal adenocarcinoma. Abdom. Radio. 46, 4800–4816 (2021).
Bian, Y. et al. Machine learning for computed tomography radiomics: prediction of tumor-infiltrating lymphocytes in patients with pancreatic ductal adenocarcinoma. Pancreas 51, 549–558 (2022).
Li, Q. et al. Prediction of tumor-infiltrating CD20(+) B-cells in patients with pancreatic ductal adenocarcinoma using a multilayer perceptron network classifier based on non-contrast MRI. Acad. Radio. 29, e167–e177 (2022).
Bian, Y. et al. Preoperative radiomics approach to evaluating tumor-infiltrating CD8(+) T cells in patients with pancreatic ductal adenocarcinoma using noncontrast magnetic resonance imaging. J. Magn. Reson Imaging 55, 803–814 (2022).
Daamen, L. A. et al. Detection, treatment, and survival of pancreatic cancer recurrence in the Netherlands: a nationwide analysis. Ann. Surg. 275, 769–775 (2022).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinforma. 12, 77 (2011).
Demler, O. V., Pencina, M. J. & D’Agostino, R. B. Sr Misuse of DeLong test to compare AUCs for nested models. Stat. Med 31, 2577–2587 (2012).
Hanczar, B. et al. Small-sample precision of ROC-related estimates. Bioinformatics 26, 822–830 (2010).
Hackner, D. et al. Neutrophil-to-lymphocyte ratio and prognostic nutritional index are predictors for overall survival after primary pancreatic resection of pancreatic ductal adenocarcinoma: a single centre evaluation. Cancers 16. https://doi.org/10.3390/cancers16162911 (2024).
Li, J. J. X. et al. Neutrophil-lymphocyte ratio reflects tumour-infiltrating lymphocytes and tumour-associated macrophages and independently predicts poor outcome in breast cancers with neoadjuvant chemotherapy. Histopathology 84, 810–821 (2024).
Zheng, H. L. et al. Trajectories of neutrophil-to-lymphocyte ratios during neoadjuvant chemotherapy correlate with short- and long-term outcomes in gastric cancer: a group-based trajectory analysis. BMC Cancer 24, 226 (2024).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Masuda, K. et al. Multiplexed single-cell analysis reveals prognostic and nonprognostic T cell types in human colorectal cancer. JCI Insight 7. https://doi.org/10.1172/jci.insight.154646 (2022).
van der Leun, A. M., Thommen, D. S. & Schumacher, T. N. CD8(+) T cell states in human cancer: insights from single-cell analysis. Nat. Rev. Cancer 20, 218–232 (2020).
Huang, X. et al. Neutrophils in cancer immunotherapy: friends or foes? Mol. Cancer 23, 107 (2024).
Luo, H. et al. Tumor-associated neutrophils upregulate Nectin2 expression, creating the immunosuppressive microenvironment in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 43, 258 (2024).
Heuser, C., Renner, K., Kreutz, M. & Gattinoni, L. Targeting lactate metabolism for cancer immunotherapy−a matter of precision. Semin. Cancer Biol. 88, 32–45 (2023).
Mousset, A. et al. Neutrophil extracellular traps formed during chemotherapy confer treatment resistance via TGF-β activation. Cancer Cell 41, 757–775.e710 (2023).
Zhong, T., Sun, S., Zhao, M., Zhang, B. & Xiong, H. The mechanisms and clinical significance of CD8(+) T cell exhaustion in anti-tumor immunity. Cancer Biol. Med. 22, 460–480 (2025).
Yang, Z. et al. Lactylome analysis suggests lactylation-dependent mechanisms of metabolic adaptation in hepatocellular carcinoma. Nat. Metab. 5, 61–79 (2023).
Khorrami, M. et al. Changes in CT radiomic features associated with lymphocyte distribution predict overall survival and response to immunotherapy in non-small cell lung cancer. Cancer Immunol. Res. 8, 108–119 (2020).
Sun, R. et al. Radiomics to predict outcomes and abscopal response of patients with cancer treated with immunotherapy combined with radiotherapy using a validated signature of CD8 cells. J. Immunother. Cancer 8, e001429 (2020).
Huang, W. et al. Noninvasive imaging of the tumor immune microenvironment correlates with response to immunotherapy in gastric cancer. Nat. Commun. 13, 5095 (2022).
Demircioğlu, A. Are deep models in radiomics performing better than generic models? A systematic review. Eur. Radio. Exp. 7, 11 (2023).
Hosny, A. et al. Deep learning for lung cancer prognostication: a retrospective multi-cohort radiomics study. PLoS Med. 15, e1002711 (2018).
Chen, T. et al. A multimodal deep learning nomogram for the identification of clinically significant prostate cancer in patients with gray-zone PSA levels: comparison with clinical and radiomics models. Acad. Radio. 32, 864–876 (2025).
Sadu, V. B. et al. Optimizing the early diagnosis of neurological disorders through the application of machine learning for predictive analytics in medical imaging. Sci. Rep. 15, 22488 (2025).
Zhang, X., Gao, A., Ma, L. & Yu, N. Integrating intratumoral and peritumoral radiomics with clinical risk factors for prognostic prediction in pancreatic ductal adenocarcinoma patients undergoing combined chemotherapy and HIFU ablation. Int. J. Hyperth. 41, 2410342 (2024).
Baltruschat, I. M., Nickisch, H., Grass, M., Knopp, T. & Saalbach, A. Comparison of deep learning approaches for multi-label chest X-ray classification. Sci. Rep. 9, 6381 (2019).
Huang, S. C., Pareek, A., Seyyedi, S., Banerjee, I. & Lungren, M. P. Fusion of medical imaging and electronic health records using deep learning: a systematic review and implementation guidelines. npj Digit. Med. 3, 136 (2020).
Yip, S. S. & Aerts, H. J. Applications and limitations of radiomics. Phys. Med. Biol. 61, R150–R166 (2016).
Liu, T. et al. Single cell profiling of primary and paired metastatic lymph node tumors in breast cancer patients. Nat. Commun. 13, 6823 (2022).
Pu, N. et al. Intratumoral IL-33 + CD4 + FoxP3 + regulatory T cell infiltration determines poor clinical outcomes and intensive immunoevasive contexture in patients with pancreatic cancer after surgical resection: a cohort study. Int. J. Surg. https://doi.org/10.1097/js9.0000000000002733 (2025).
Guo, G. et al. Immune cell concentrations among the primary tumor microenvironment in colorectal cancer patients predicted by clinicopathologic characteristics and blood indexes. J. Immunother. Cancer 7, 179 (2019).
Lan, T. et al. MRI-based deep learning and radiomics for prediction of occult cervical lymph node metastasis and prognosis in early-stage oral and oropharyngeal squamous cell carcinoma: a diagnostic study. Int. J. Surg. 110, 4648–4659 (2024).
Wei, Z. et al. A CT-based deep learning model predicts overall survival in patients with muscle invasive bladder cancer after radical cystectomy: a multicenter retrospective cohort study. Int. J. Surg. 110, 2922–2932 (2024).
Ye, G. et al. Non-invasive multimodal CT deep learning biomarker to predict pathological complete response of non-small cell lung cancer following neoadjuvant immunochemotherapy: a multicenter study. J. Immunother. Cancer 12. https://doi.org/10.1136/jitc-2024-009348 (2024).
Lee, W. et al. Preoperative data-based deep learning model for predicting postoperative survival in pancreatic cancer patients. Int. J. Surg. 105, 106851 (2022).
Hua, Y. et al. Pretreatment CT-based machine learning radiomics model predicts response in unresectable hepatocellular carcinoma treated with lenvatinib plus PD-1 inhibitors and interventional therapy. J. Immunother. Cancer 12. https://doi.org/10.1136/jitc-2024-008953 (2024).
Tu, M. et al. TNF-α-producing macrophages determine subtype identity and prognosis via AP1 enhancer reprogramming in pancreatic cancer. Nat. Cancer 2, 1185–1203 (2021).
Wang, L. et al. Single-cell RNA-seq analysis reveals BHLHE40-driven pro-tumour neutrophils with hyperactivated glycolysis in pancreatic tumour microenvironment. Gut 72, 958–971 (2023).
Zhang, Y. et al. Interleukin-17-induced neutrophil extracellular traps mediate resistance to checkpoint blockade in pancreatic cancer. J. Exp. Med. 217. https://doi.org/10.1084/jem.20190354 (2020).
Picard, F. S. R. et al. IL-17A-producing CD8(+) T cells promote PDAC via induction of inflammatory cancer-associated fibroblasts. Gut 72, 1510–1522 (2023).
Heiduk, M. et al. TIGIT expression delineates T-cell populations with distinct functional and prognostic impact in pancreatic cancer. Clin. Cancer Res. 29, 2638–2650 (2023).
Collins, G. S. et al. TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. Bmj 385, e078378 (2024).
Allen, P. J. et al. Multi-institutional validation study of the American Joint Commission on Cancer (8th Edition) changes for T and N staging in patients with pancreatic adenocarcinoma. Ann. Surg. 265, 185–191 (2017).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Braman, N. M. et al. Intratumoral and peritumoral radiomics for the pretreatment prediction of pathological complete response to neoadjuvant chemotherapy based on breast DCE-MRI. Breast Cancer Res. 19, 57 (2017).
Tunali, I. et al. Stability and reproducibility of computed tomography radiomic features extracted from peritumoral regions of lung cancer lesions. Med. Phys. 46, 5075–5085 (2019).
Tan, M. & Le, Q. EfficientNet: rethinking model scaling for convolutional neural networks. In Proc. 36th International Conference on Machine Learning Vol. 97 (eds Chaudhuri K. & Salakhutdinov R.) 6105–6114 (PMLR, 2019).
Lin, M., Chen, Q. & Yan, S. Network in network. arXiv preprint https://doi.org/10.48550/arXiv.1312.4400 (2013).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 7132–7141 (IEE2018).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. In Proc. 32nd International Conference on Machine Learning (eds Bach, F. & Blei, D.) Vol. 37, 448–456 (PMLR, 2015)
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Loshchilov, I. & Hutter, F. Decoupled weight decay regularization. arXiv:1711.05101. https://ui.adsabs.harvard.edu/abs/2017arXiv171105101L. (2017).
Loshchilov, I. & Hutter, F. SGDR: stochastic gradient descent with warm restarts. arXiv https://doi.org/10.48550/arXiv.1608.03983 (2016).
Arik, S. Ö. & Pfister, T. TabNet: Attentive Interpretable Tabular Learning. In Proc. AAAI Conference on Artificial Intelligence, Vol. 35, 6679–6687 (2021).
Baltrušaitis, T., Ahuja, C. & Morency, L. P. Multimodal machine learning: a survey and taxonomy. IEEE Trans. Pattern Anal. Mach. Intell. 41, 423–443 (2019).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. In Proc. IEEE International Conference on Computer Vision (ICCV) 618–626 (IEEE, 2017).
Xue, B. et al. Use of machine learning to develop and evaluate models using preoperative and intraoperative data to identify risks of postoperative complications. JAMA Netw. Open 4, e212240 (2021).
Acknowledgements
We extend our gratitude to Professor Li Xue from the School of Public Health, Zhejiang University, for her expert critique and invaluable insights during the scientific refinement of this work. Figure 1 was created with BioGDP.com. This work was a project supported by the National Natural Science Foundation of China under grants No.82272954, No.82573736 (Yulian Wu), No.82572303 (Xiawei Li), No.82202984 (Hongxia Xu), Scientific Research Fund of Zhejiang University under grant No. XY2024034 (Yiqun Fan), and Zhejiang Key R&D Program of China under grant No.2023C03053 (Jian Wu).
Author information
Authors and Affiliations
Contributions
These authors contributed equally: Yiqun Fan, Bang Du, Kaiyue PuY.F., B.D., K.P., Y.W., and X.L. conceived the study. Data curation was performed by Y.F., B.D., K.P., C.L., X.L., S.H., Y.S., and Y.W. Formal analysis was conducted by Y.F., B.D., C.L., H.X., and S.H. Y.F., K.P., C.L., Y.S., and W.B. carried out the investigation. Methodology development was led by Y.F., B.D., C.L., Y.C., J.T., Y.Z., H.X., and X.L. Resources were provided by Y.F., J.W., H.X., and Y.W. Supervision was provided by H.X. and X.L. Validation was conducted by Y.F., K.P., C.L., Y.C., J.T., and Y.Z. Visualization was performed by Y.F. and B.D. The original draft was written by Y.F., B.D., K.P., C.L., R.W., H.X., and X.L. Y.F., B.D., C.L., R.W., H.Z., Y.D., H.X., X.L., J.W., and Y.W. contributed to the review and editing of the manuscript and approved the final version. All authors have approved the content of the manuscript and agreed on publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fan, Y., Du, B., Pu, K. et al. Noninvasive evaluation and clinical value prediction of tumor-infiltrating neutrophil-to-T-cell ratio in pancreatic ductal adenocarcinoma. npj Digit. Med. 9, 123 (2026). https://doi.org/10.1038/s41746-025-02303-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02303-9