Abstract
This systematic review evaluates how decentralized learning (DL) approaches—e.g., federated learning, swarm learning, ensemble–compare with traditional models in healthcare applications. We searched eight databases (01/2012 to 03/2024), screening 165,010 studies with two independent reviewers. Analysis included 160 articles comprising 710 DL models and 8149 performance comparisons across clinical domains, predominantly in oncology, COVID-19, and neurological diagnostics. In paired comparisons, centralized learning (CL) demonstrated advantages in threshold-dependent metrics (78% favourability for accuracy and Dice score with large effect sizes), while DL achieved comparable performance in ranking metrics (51% centralized favourability for AUROC with small effect size). DL consistently outperformed local learning (LL) across all metrics, particularly precision (86% favourability) and accuracy (83% favourability). Clinical threshold analysis (≥0.80 performance) revealed that CL rescued DL viability in up to 18% of comparisons, though when both achieved clinical viability, improvements typically represented “excellent versus acceptable” performance (median difference of 0.7–1.5pp) rather than “acceptable versus inadequate.” DL rescued LL viability with substantial improvements (median difference of 7.6–27pp). These findings demonstrate DL offers clinically acceptable alternatives for privacy-constrained contexts, with implementation decisions balancing marginal performance trade-offs against regulation (e.g., GDPR, AI Act) and application. Future research requires standardized privacy-performance reporting.
Similar content being viewed by others
Introduction
In recent decades, health systems worldwide have been facing unprecedented profound challenges. The epidemiological transition has intensified care needs and costs1,2,3,4, while technological advancements and pharmaceutical innovations have driven increasing expenditures4,5. Simultaneously, these systems still struggle to achieve and ensure universal coverage6,7, while facing increasing critical workforce shortages and limited investments and action across levels of prevention and health determinants8. These problems demand innovative approaches to improve the quality of care, while optimizing resources, particularly in clinical decision-making tasks (e.g., establishing diagnoses, prescribing therapies, offering prognoses).
Against this backdrop, machine learning systems and artificial intelligence (AI) technologies have been proposed as methods to address these demands. The convergence of ubiquitous digital systems, large-scale data, and advanced computational capabilities creates a unique opportunity to address the health domain’s most pressing demands.
However, despite these technological advances, healthcare systems have not yet been successful in developing and implementing effective solutions. As computational capability increases, the primary bottleneck to data science may lie in accessing and utilizing high-quality health data9,10, something particularly challenging in health-related domains. Public and anonymized databases are valuable resources, but they often lack external validity for developing robust healthcare applications. In contrast, real-world data (RWD), the clinical evidence derived from data collected during routine healthcare delivery, may be a more adequate and representative data source11.
While RWD offers potential for developing representative and generalizable models, the implementation of machine learning systems using these data faces multiple barriers. Legal and regulatory frameworks mandate increasingly stringent technical safeguards for data collection, maintenance, analysis, and disposal. Operational challenges include interoperability issues and integration with existing or new clinical workflows, as well as with legacy information systems. In particular, a fundamental tension exists between model performance and privacy protection: more granular data improves performance but increases privacy risks.
In response to these challenges, decentralized learning approaches have emerged as possible solutions. These approaches aim to enable machine learning on distributed healthcare data while maintaining privacy and regulatory compliance. In federated learning (FL), models are trained locally, and only tuned parameters (e.g., weights and biases in a neural network) of participating parties are shared with a central server for aggregation12. In swarm learning (SL), model parameters aggregation occurs peer-to-peer, in a fully decentralized way13. This obviates the need to use a central and authoritative controller. Applications cover a variety of data formats and conditions, including some particularly privacy-sensitive, using FL14,15,16,17,18,19,20 and SL21,22. Complementarily, ensemble methods, such as bagging techniques23, offer simpler aggregation schemes, based on plurality voting to produce the global result. By design, they are more flexible and integrate different learning methods more easily.
However, a fine balance lies between the ambition to maximize model performance and the need to minimize privacy risks and compromises. Still, comparing decentralized and traditional methods requires a nuanced view. Pivotal problems include regulatory compliance, technical feasibility, and privacy guarantees. Depending on the use case, issues such as number of controllers and points of failures can be seen as either positives or negatives. Considering the General Data Protection Regulation (GDPR)24 and the AI Act24, attention has been directed towards the goal of decentralized learning models achieving performance comparable to their traditional counterparts25,26.
Despite the growing interest in decentralized learning for healthcare applications, there is a lack of robust synthesis comparing their performance with traditional, non-decentralized approaches. Such a systematic comparison would provide crucial insights into their relative effectiveness, practical advantages, and limitations across different clinical contexts and tasks. This knowledge gap hampers informed decision-making about implementing these technologies in healthcare settings and highlights the need for a comprehensive literature assessment. Existing systematic reviews have important limitations regarding their size and scope27,28,29, the specificity of health care applications30, and the adequacy and comprehensiveness of query prompts and search strategies31. This review builds on a registered and published protocol32 to provide a comprehensive and replicable analysis.
This systematic literature review seeks to compare the performance of health data models developed using decentralized learning approaches (e.g., federated learning, swarm learning, ensemble) with those developed using traditional centralized or local methods, as the primary objective. The performance comparison is grouped using the metrics reported in the original articles (e.g., accuracy, precision, AUROC), covering a wide range of medical conditions (e.g., COVID-19, breast cancer, type 2 Diabetes), through different clinical tasks (e.g., diagnosis, segmentation, prognosis). Secondary objectives include describing the types of data and datasets used, the nature of the decentralized model architectures, and the reporting of resource demands or privacy impacts.
We expect to help inform policy-making and operational decisions on the applicability of these methods, integrating their reported strengths and shortcomings with the intended use cases. Moreover, this work suggests further research questions and study designs to better understand the privacy protection benefits, as well as challenges and opportunities concerning their validity and implementation.
Results
Study selection
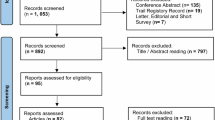
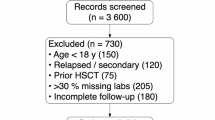
Our systematic review identified a total of 165,010 studies. Figures 1 and 2 describe the phases 1 and 2 of the identification, screening, and selection processes. Before screening, three processes were undertaken to exclude irrelevant or redundant entries. First, exact and apparent duplicates (i.e., only differing in case for the DOI link, title or abstract, or only differing in a whitespace or a full-stop) were removed (43,493 articles). Subsequently, applying the regular expressions filter described in “Search Strategy”, another 111,594 studies were removed. For Phase 2, duplicates of studies already assessed in Phase 1 were excluded from repeated analysis. In the end, 9943 articles were screened, resulting in the exclusion of 8971 articles. The remaining studies were sought for retrieval, with 26 not being able to be retrieved. During the inclusion stage, applying the eligibility criteria to the full text version of the papers, 946 studies were assessed. In the end, 160 primary articles were analysed, comprising 710 decentralized learning models and 8149 comparisons.
The diagram illustrates the study identification and selection process. Boxes detail the number of records at each stage, with arrows indicating the flow between identification, screening, eligibility assessment, and inclusion stages. Side boxes detail reasons for exclusion at each step.
The diagram illustrates the study identification and selection process. Boxes detail the number of records at each stage, with arrows indicating the flow between identification, screening, eligibility assessment, and inclusion stages. Side boxes detail reasons for exclusion at each step.
Study characteristics
The included studies and their main characteristics are presented in Table 1. The most popular broad clinical domains covered were oncological diseases, COVID-19 and neurological conditions, as presented in Table 2. “Diagnosis” was the most common clinical application, as seen in Table 3. A clear trend arises in an increasing number of yearly publications, with 2024 data only including publication until March 28th, as seen in Table 4. The most popular article sources are presented in Table 5.
Regarding the decentralized models, the most common TRIPOD type was 2a (74 articles), followed by types 1b, 2b and 3 (26, 23, and 23 articles respectively), with 11 being unclear and only 3 classified as type 1a. Concerning code access, 106 articles did not report code availability, 43 provided all code, 6 provided some code, 4 explicitly stated no code would be available, and 1 had pending code access requests. For data access, 64 articles provided all data, 36 did not report data access, 32 had pending data access requests, 20 provided some data, and 8 provided no data access.
Regarding the model architectures used, Federated Learning was the most common (557 models), followed by Fully Decentralized approaches, including Swarm Learning (111 models), Ensemble methods (21), Split or Transfer Learning (14), and Secure Multi-Party Computation (4). Most models (687) use real data, while only 16 use synthetic data and 5 use both types.
In terms of the data used, Electronic Health Records/Text is the most commonly used data type (121 models). Image-based data was used very frequently, namely MRI (101 models), X-Ray (96 models), Pathology/Whole slide images (67 models), CT Scans (65 models), retina fundus images (37 models), ECG/EKG (27 models), dermoscopic images (21 models), EEG (5 models) and ultrasound (5 models). Genome data was used in 33 models. Some models use combinations of data types, such as Electronic Health Records/Text + X-Ray (12 models) and MRI + Pathology/Whole slide images (6 models). Less frequent data types were endoscopic videos (17 models), laparoscopic cholecystectomy videos (16 models), and wireless capsule endoscopy videos (4 models). The least common data types, used by only 1 or 2 models, include Mammography, Microscopy, Intra-Oral Mesh Scans, OCTA + OCT and PET images.
Using the PROBAST + AI tool, we appraised the 25 most cited articles of the TRIPOD Types 2a, 2b and 3, considering up to two models for each article (Figs. 3–6).
Red segments, yellow segments and green segments represent the proportion of classifications of high concern level, unclear concern level and low concern level.
Red segments, yellow segments and green segments represent the proportion of classifications of high risk of bias, unclear risk of bias and low risk of bias.
Red segments, yellow segments and green segments represent the proportion of classifications of high concern level, unclear concern level and low concern level.
Red segments, yellow segments and green segments represent the proportion of classifications of high concern level, unclear concern level and low concern level.
Performance comparison
We grouped our findings by the main types of model evaluations conducted (i.e., combinations of broader clinical domain, clinical application and performance metric). Regarding centralized learning models, as presented in Table 6, the most common evaluation was for diagnostic accuracy in Oncology (53 models and 189 comparisons). Other frequent clinical domains include COVID-19 and Cardiology. Regarding local learning models, as presented in Table 7, the most common evaluation was for diagnostic accuracy in COVID-19 (31 models and 195 comparisons). Similarly, Oncology and Cardiology were also frequently explored.
In terms of performance comparisons, the paired differences between decentralized and non-decentralized learning approaches are summarized in Fig. 7. Supplementary Figs. 2–15 present the distribution these differences for each performance metric and type of non-decentralized model, alongside additional data (e.g., 25th and 75th percentiles of differences, 95% confidence intervals). To describe the magnitude of these differences, the effect sizes for these comparisons are provided in Fig. 8.
Higher percentage corresponds to increase favourability. Row 1 represents the proportion of paired comparisons in which Decentralized Learning overperforms Local Learning. Row 2 represents the proportion of paired comparisons in which Centralized Learning overperforms Decentralized Learning.
The table presents effect sizes calculated using the Wilcoxon two-sample paired signed-rank test across seven performance metrics (AUROC, Accuracy, Precision/Positive Predictive Value, Sensitivity/Recall, F1 score, Specificity, and Dice score). Each metric is compared against both Local Learning and Centralized Learning approaches using the corresponding Decentrralized Learning values. Columns display the estimate of effect size, magnitude classification, and number of comparisons.
Centralized learning
Overall, centralized learning presents a clear performance advantage across different metrics. This approach is particularly superior to decentralized learning in accuracy and Dice score, with a higher performance in 78% of 1089 comparisons and 78% of 856 comparisons respectively, with narrow interquartile ranges and large effect sizes (≥0.5).
In contrast, AUROC and specificity comparisons present a more balanced distribution, with centralized learning performing better in 51% of 1063 comparisons and 54% of 160 comparisons, respectively. For both metrics, the effect sizes estimated are small (<0.3). The remaining metrics also favor centralized learning, with less pronounced favourability (from 63% to 69%), wider interquartile ranges and moderate effect size.
Local learning
In contrast, decentralized learning models consistently outperform their local counterparts. In particular, accuracy and precision/positive predictive value metrics, performing better in 83% of 1023 comparisons and 86% of 440 comparisons respectively, featuring large effect sizes. Similarly, strong preference is observed in AUROC and F1 score comparisons, with decentralized learning models being superior in 82% and 79% of cases, respectively, in association with large effect sizes.
The decentralized learning advantage is smaller for Dice score and specificity metrics (72% and 76% of comparisons, respectively), while still featuring large effect sizes. Sensitivity/recall shows decentralized learning performing better in 71% of cases, with a wider interquartile range and a moderate effect size.
Performance difference significance
Considering the direction, magnitude and statistical relevance of the performance differences, we subsequently explored the clinical relevance of these findings using a clinical viability threshold (≥0.80).
When comparing centralized and decentralized models, centralized approaches frequently rescued clinical viability from underperforming DL models (Fig. 9, Panel A). Specifically, centralized models provided clinically valid alternatives to DL in up to 18% of comparisons, particularly for sensitivity (median difference of 16pp) and accuracy metrics. Conversely, even when both models achieved clinical viability (≥0.80), centralized approaches typically demonstrated only marginal advantages (Panel B), with median performance differences ranging from 0.7 to 1.5 percentage points across metrics. DL models rarely offered superior alternatives to viable centralized counterparts (Panel C).
Blue = Decentralized Learning; Green = Centralized Learning; Gray line = Difference. Clinical viability threshold set at 80%. Lines connect DL to centralized model performance. Δ indicates median performance difference (only comparisons with n ≥ 20 shown). A Centralized model rescues clinical viability. B Both models are viable but centralized performs better. C Centralized model loses viability compared to viable DL model.
In comparisons with local models, DL consistently rescued clinical viability across substantial proportions of cases (Fig. 10, Panel A). The rescue effect was most pronounced for threshold-dependent metrics, particularly sensitivity (median difference of 27pp for 100 comparisons) compared to ranking metrics like AUROC (median difference of 7.6pp). DL provided clinically acceptable alternatives across metrics with median improvements ranging from 7.6 to 27 percentage points. When both approaches met the viability threshold (Panel B), DL maintained performance advantages, with accuracy superior in 53% of comparisons (median difference of 1.9pp). DL underperformance against viable local models was rare (Panel C) and associated with significant losses.
Blue = Decentralized Learning; Red = Local Learning; Gray line = Difference. Clinical viability threshold set at 80%. Lines connect DL to local model performance. Δ indicates median performance difference (only comparisons with n ≥ 20 shown). A Decentralized model rescues clinical viability. B Both models are viable but DL performs better. C Centralized model loses viability compared to viable DL model.
Detailed metric-specific analyses are provided as Supplementary Material.
Additional privacy-preserving techniques and secondary aims
On the technical side, we report details of the methods used (e.g., encryption, GAN, homomorphic encryption) strictly based on the original manuscripts. On the resources demand side, we report details on various metrics–ranging from energy consumption, server and client memory requirements to time needed for model training and model inference. This information is presented as Supplementary Material.
Supplementary analyses
For further granularity, we developed an online dynamic dashboard to produce different graphical representations of data collected. Available filters are performance metrics, non-decentralized learning approach, clinical applications, larger clinical domains and data type. Additionally, it is possible to restrict decentralized model comparisons to only federated learning models.
To complement the analyses of the performance distributions, we provide histogram representations by nature of data collection used by the models compared (“primary”, “secondary” and “both”) in Supplementary Figs. 23–36.
Additionally, we produced sub-analyses on the distribution of the absolute differences (i.e., the difference between non-decentralized approach and decentralized approach), as well as on distribution of the relative differences (i.e., the quotient of the absolute differences with the difference between 1 and the non-decentralized approach). These representations correspond to Supplementary Figs. 37–64.
Discussion
This systematic review provides the most comprehensive analysis to date comparing decentralized learning approaches with traditional methods in healthcare, examining 160 studies comprising 710 decentralized models and 8149 performance comparisons.
The rapid growth in research output, particularly since 2020 and multidisciplinary scope reflects increasing recognition of decentralized learning’s potential in healthcare applications.
Considering the paired comparisons between decentralized and centralized methods, performance differences present low magnitude median values and reduced interquartile ranges. This demonstrates that decentralized approaches can broadly achieve comparable performance, although moderately inferior. In particular, strong relative performance in AUROC (51% centralized favourability, small effect size) suggests that the observation ranking ability is preserved through decentralized learning processes. In turn, threshold-dependent metrics—such as accuracy and Dice score– show increased centralized relative advantages, with mostly moderate or large effect sizes. These findings reveal calibration challenges and spatial feature averaging difficulties, respectively. However, DL seems to overperform in terms of specificity (54% centralized favourability, small effect size), suggesting that aggregation processes differentially affect error types. In particular, multi-site learning may filter site-specific false positive patterns while simultaneously diluting rare positive case signals, given case presentation variation and uneven distribution of rare cases across sites.
Focusing on the application viability of these models, centralized models can offer clinically useful alternatives to underperforming decentralized counterparts, in up to 18% of the cases. Sensitivity and accuracy are particularly benefited by the centralized approach, aligning with DL limitations to identify true positive cases.
Regarding the differences with local approaches across all metrics, decentralized performance is dominant, despite some heterogeneity. While, decentralized models benefit from more and often more diverse data, they are less tuned to the specific distribution of a local dataset. In particular, DL demonstrates the strongest advantage in precision (86% favourability), substantially exceeding gains in other metrics.
This likely reflects multi-site models’ ability to filter out site-specific artifacts (e.g., differences in imaging protocols, scanner calibration). Local models overfit to these artifacts, leading to overconfident predictions that inflate false positive rates when encountering variation. In turn, sensitivity shows the smallest DL advantage (70%), with only a moderate effect size, likely reflecting challenges in aggregating rare or subtle pathological patterns across heterogeneous sites. Specificity shows greater improvement (76% favourability), as normal imaging features are more consistent across sites than disease presentations, and DL models learn to avoid falsely flagging benign site-specific variations. This asymmetry reflects a fundamental trade-off: local models can optimize to site-specific patterns—potentially overfitting—at the expense of external validity, whereas DL prioritizes features robust across heterogeneous sites. A similar pattern emerges when comparing DL to centralized models, where challenges in aggregating rare signals similarly constrain sensitivity improvements.
With our sensitivity analysis, excluding observations from articles with the most comparisons, variations in favourability ratios were generally within single-digit percentage points. This strengthens the validity of the data presented and our conclusions.
Focusing on clinical applicability, the threshold-stratified analysis (≥0.80) reveals important patterns for implementation decisions. Centralized models can rescue clinical viability from underperforming DL in up to 18% of cases, primarily for sensitivity and accuracy. This aligns with DL’s documented limitations in identifying true positive cases, particularly rare or subtle pathological patterns across heterogeneous sites.
Importantly, the clinical threshold analysis demonstrates that when both centralized and DL approaches achieve clinical viability, centralized superiority typically represents “excellent versus acceptable” performance rather than “acceptable versus inadequate.” While centralized improvements occur frequently, their magnitude is limited (median difference ranging from 0.7pp to 1.5pp). This suggests that when DL models achieve clinically acceptable performance, centralized alternatives provide only modest incremental gains. This positions DL as a viable alternative for contexts where centralized approaches are prohibited by privacy regulations or data sharing constraints.
Regarding differences with local approaches, DL demonstrates dominant performance across all metrics. The clinical rescue effect is substantial, with median improvements of 7.6–27pp depending on metric. The disproportionate improvement in threshold-dependent metrics (27pp for sensitivity) compared to ranking metrics (7.6pp for AUROC) reveals that local models suffer from overfitting to site-specific patterns and class distributions. Decentralized learning mitigates this by learning features robust across heterogeneous clinical settings, resulting in more generalizable decision boundaries. Notably, even when local models achieve clinical viability, DL frequently offers performance increases that should be considered alongside potentially superior external validity.
Regarding additional privacy-preserving techniques and the secondary aims of this study, these data points are reported infrequently and not in a standardized fashion. Even when reported, key variables (e.g., noise levels) are often fixed, making it impossible to assess their impact in each study. Due to differences in datasets, clinical domains, clinical applications or different computational set-ups, cross-study comparisons would not provide reliable insights. Overall, decentralized models are more resource demanding than their counterparts, especially when privacy-preserving methodologies are added.
A qualitative synthesis of the evidence presents some notable patterns. Noise levels of 0.001 provides a superficial level of protection with negligible impacts on performance. Memory and data transmission requirements, outside of resource scarce environments, should not cause significant hardship for model development. While some techniques can increase development time, these rarely duplicate the duration for their standard counterparts. In real-world settings inference time may be a more relevant constraint. Depending on the techniques used, this can lead to compounded increases and may function as an effective bottleneck to the deployment of larger and more complex models.
The findings from this systematic review enable evidence-based decision-making for healthcare AI implementations balancing privacy preservation with clinical performance requirements. To allow actionable application of these insights we propose a simple decision framework.
We start by highlighting when decentralized learning can be recommended. DL represents the optimal approach in three primary scenarios. First, when data sharing is legally prohibited or institutionally restricted (e.g., under GDPR constraints, cross-border regulations or institutional data governance policies), DL enables model development that would otherwise be impossible. Our analysis demonstrates DL achieves clinically acceptable performance (≥0.80) in the majority of applications, with 83% favourability over local approaches for accuracy and 82% for AUROC. Second, when local data alone yields insufficient performance, DL rescues clinical viability in 12% to 15% of cases with substantial improvements (median difference of 7.6–27pp depending on metric). Third, when external validity is prioritized over maximal performance, DL’s multi-site learning reduces site-specific overfitting, particularly valuable for precision metrics where DL shows 86% favourability over local models.
In turn, centralized approaches should be selected when privacy constraints are manageable and maximal performance is required. Centralized models demonstrate advantages in threshold-dependent metrics, particularly accuracy (78% favourability) and Dice score (78% favourability), with large effect sizes. Clinical threshold analysis reveals these advantages typically represent mostly “excellent versus acceptable” rather than “acceptable versus inadequate” performance. When both approaches achieve clinical viability, centralized improvements average only 0.7–1.5pp, in 16% to 44% of comparisons. However, centralized approaches still provide clinically meaningful rescue in 6% to 17% of comparisons. Therefore, centralized learning is justified primarily when: (1) marginal performance improvements are clinically critical, (2) working with rare pathological patterns requiring maximum sensitivity or (3) privacy-preserving infrastructure is unavailable.
Alternatively, local-only approaches should be avoided for deployment across multiple sites or generalizable applications. Local models systematically underperform DL across all metrics, with particularly poor precision (14% favourability) due to overfitting to site-specific artifacts. The 27pp sensitivity improvement from DL versus local models in rescue scenarios indicates local approaches risk missing true positive cases when applied beyond their training environment. Local models may only be appropriate for strictly site-specific applications where external validity is not required and privacy or technical constraints prevent any data collaboration.
Decision-makers should consider that DL’s primary trade-off is not clinical inadequacy but rather marginal performance concessions (typically 1–2pp) for privacy preservation. The resource overhead—while measurable—rarely doubles development time, though inference latency may constrain deployment of complex models. Organizations should prioritize DL when regulatory compliance, institutional policies, or ethical considerations prohibit centralized data aggregation, accepting that performance will be clinically acceptable rather than optimal in most scenarios.
Despite the robustness of this work, some limitations may have affected these results. Publication bias, reporting bias, and selection bias could influence which results are available for inclusion, potentially skewing the aggregated findings. No specific efforts were made to assess or address these. In addition, gray literature or publications outside primary scientific articles were not examined. Our focus on peer-reviewed publications prioritized methodological rigor and clinical applicability, although this approach may have introduced a temporal lag in capturing the most recent developments and reduced the breadth of included results. We mitigated this by searching for published versions of identified preprints and conducting updated searches through March 2024, to balance evidence quality with timeliness. However, a single moment for evidence retrieval and classification would have been preferable. While we aimed for a clear selection and definition of decentralized learning approaches considered, we recognize other interpretations may be valid. However, the majority of data concerns well established methods (e.g., Federated Learning, Swarm Learning). In addition, we recognize some mistakes (i.e., random errors) may have occurred during our extensive process. During our peer-review process, a small number of otherwise eligible papers33,34 were by mistake not considered.
Regarding data quality of the included studies, many included articles relied on secondary data or inadequately detailed primary data collection. Both private and public datasets featured instances of insufficient number of participants, observations or predictors, as well as the poor quality of reporting of eligibility criteria, outcome definitions and methods used. In practice, these challenges, alongside inconsistent reporting formats, made identifying different health data models, their characteristics, and performance comparisons more difficult.
Therefore, our evidence appraisal document issues related to the primary studies used. Due to the broad scope of our research question and the comparability of the decentralized and non-decentralized model development and evaluation processes, we believe evidence used to be of low concern for this purpose. Additionally, while clinical applicability performance thresholds vary by application and context, 0.80 provides a standardized benchmark across heterogeneous domains.
Considering the main implications of the study, this systematic review makes three novel contributions to the field: (1) quantification of favourability ratios between traditional and decentralized learning approaches across performance metrics, (2) identification of performance ranges where variations are most pronounced, and (3) clinical significance assessment through threshold-stratified analysis.
This is the first study that presents a quantitative evaluation of the difference between decentralized and non-decentralized approaches at a paired comparison level and grouped by clinical application characteristics. This work demonstrates the ability for DL to present robust ranking assessments, while still struggling to retain positive and rare signals, especially when compared to their centralized counterparts. When considering clinically relevant performance ranges, centralized learning superiority is deepened. Compared to local learning, DL advantages are significant, especially in AUROC, accuracy and precision, and present sizable performance increases, when considering clinical applicability. Therefore, decentralized learning represents a clear superior alternative to local-only approaches, centralized learning continues to be the gold standard. However, DL offers a viable alternative for contexts in which centralized learning is not possible.
As the AI Act advocates for performance parity between traditional and privacy-preserving techniques, the quantitative synthesis of the evidence provides an objective insight for monitoring the state of art and evolution of these approaches. In parallel, our limited findings on privacy-performance trade-off support the need for increase adoption of standardized privacy evaluation metrics. In particular, we recommend more rigorous comparative studies, better documentation of implementation details and focus on practical deployment in healthcare settings. Heterogeneous and infrequent reporting does not allow for an adequate study of dynamics between privacy-preserving guarantees and performance cost.
Considering the issues raised during the evidence appraisal of the most cited, and the variety of specific clinical use cases, these results cannot validate particular implementations for widespread deployment. Problems related to reporting of sampling processes, target population definition and data collection methods compromise external validity of the studies considered. In addition, small variations in performance metrics even for a specific disease can have different clinical and operational impacts (e.g., screening versus diagnosis application). Nonetheless, we encourage the exploration of different sub-analyses in our online dashboard to identify promising research fields.
Comparing this study with similar recent reviews, this work provides a detailed and quantitative assessment of the results from the primary articles. Contemporary research mostly focuses on reporting the article and model characteristics, commonly using narrative syntheses of the primary articles35,36,37,38. In addition, these works do not provide actionable information on the added benefit of using decentralized approaches in contrast with traditional methods already being used, nor valuable syntheses of the evidence. Moreover, to the best of our knowledge, no published review on the topic was preceded by the respective protocol publication or registry.
In this domain, future research should focus on the impact of local adaptation processes on decentralized learning performance. A two-step paradigm including local calibration learning followed by local calibration may balance privacy preservation, feature generalization and clinically relevant performance. New studies on the topic should present higher methodological quality, with clearer reporting of eligibility criteria, data collection strategies, outcome definitions and model performance comparisons. For privacy-preserving reporting, guiding references—including quantitative and qualitative dimensions – are needed for comparability. While GDPR and AI Act intentionally do not offer specific metrics, there are alternatives39,40 from experts on the field.
Other topics regarding the adoption of decentralized learning methods require further discussion. From data distribution challenges to considerable technical overheads and machine “unlearning”41 requirements, data collaboration still faces foundational constraints that may limit its widespread adoption. Meanwhile, novel methods such as local fine-tuning pre-trained models, the advent of AI-capable personal devices and normative AI approaches42 can help leverage the development of decentralized learning models.
Methods
Eligibility criteria
The inclusion criteria were: (1) original and published primary research scientific journal articles; (2) studies addressing clinical decisions regarding specific human medical conditions (e.g., diagnosis, segmentation, prognosis); (3) application of decentralized learning methods for model development; (4) comparison against centralized or local methods; (5) numeric reporting of model performance using at least one relevant metric (e.g., accuracy, precision). While unpublished studies (e.g., pre-prints) and other presentations (e.g., conference proceedings) were retrieved, they were only considered insofar as to search for their corresponding version matching this criterion. Performance metrics were not considered during the screening phase but were used in the appraisal phase. Articles excluded were marked with the first unmet eligibility criterion. Regarding the exclusion criteria, papers published before 2012 were not considered for this analysis.
Information sources
Eleven databases were queried—covering biomedical scientific research (namely, SpringerLink, Lippincott Williams & Wilkins), computer science and informatics engineering (namely, the Association for Computing and Machinery Digital Library or Guide to Computing Literature and IEEE Xplore), and more general sources (namely, Wiley Online Library, Scopus, Web of Science, and Lens [including the PubMed database]). Additional databases were consulted, including those containing not peer-reviewed papers or unpublished research papers (such as medRxiv and arXiv). The Cochrane Database of Systematic Reviews and PROSPERO registers were consulted on the same dates to identify other ongoing or finished systematic reviews on the topic.
For every listed source, searches were conducted in two moments, retrieving article meta-data from both databases and registries. The first moment concerned articles from January 1st, 2012 until the query dates: April 6th (for all sources, except ACM DL) and April 7th (for ACM DL) 2023. The second moment targeted articles from April 6th, 2023 to those available on March 28th, 2024.
In this stage, articles of different natures (i.e., unpublished, conference proceedings, pre-prints) were considered for retrieval, but only peer-reviewed articles were included when available.
Search strategy
For searching evidence in this recent and multidisciplinary domain, it was necessary to devise a broad search strategy, considering a large pool of databases and advanced query techniques. A representation of the intended query is seen in Table 8. Due to the popularity of some terms and heterogeneous search engine features, a filtration process was applied, using regular expressions (RegEx) code. Detailed information is available in Supplementary Material.
The intended search query results were to include terms from the first and second group separated by no more than two other terms. The order in which they appeared in the title or abstract was not considered relevant.
Selection process
During the screening phase, titles and abstracts were reviewed. During the appraisal phase, the papers were evaluated using their full-text versions. For each exclusion, the unmet eligibility criterion was registered. For these tasks, the Rayyan43 platform was used.
To assess eligibility: (1) reviewers verified publication type using the DOI link or, if unavailable, the title and abstract information and the source to evaluate its nature; (2) clinical applications on specific human medical conditions were verified by identifying specific health targets; (3) application of decentralized learning methods to develop health data models was confirmed when models were trained on data that remained local to each party; (4) comparisons between the decentralized learning models performance and their non-decentralized counterparts supported by the presence of local (i.e., data from a single silo) and centralized learning strategies (i.e., combined data from multiple parties); (5) model performance comparisons were gathered based on written numeric data in the manuscript text or within tables, graphs, and figures, if the numeric information and the model they represent were clear. Efforts were made to also include data from the Supplementary Material.
To classify different model development approaches, we adopted an operational framework we used, based on two core dimensions. Approaches were classified based on data movement (i.e., whether raw, primary data leaves its original source) and participation of parties (i.e., whether one or multiple parties contribute to model development). Using these criteria, we define the categories as follows:
-
Local Learning: Model development is carried out by a single entity using only its own data. No data sharing or coordination with external parties occurs.
-
Centralized Learning: Multiple parties contribute to model development by sharing raw data with a central aggregator, where training is conducted.
-
Decentralized Learning: Multiple parties participate in model development without exchanging raw data. Instead, models, parameters, or privacy-preserving computations are shared to enable collaborative learning.
Within decentralized learning, we distinguish the following approaches:
-
Federated Learning: A central server coordinates the training of local models on distributed data. Only model updates (e.g., weights or gradients) are shared; raw data remains local.
-
Swarm Learning: A fully decentralized version of federated learning with no central server; model updates are aggregated peer-to-peer.
-
Ensemble Methods: Independent models are trained locally by each party and later combined (e.g., via voting or stacking) without creating a unified global model or sharing data. These are considered decentralized as model combination occurs without raw data exchange.
-
Split or Transfer Learning: Due to their similarity and reduced number of observations, we group split learning and transfer learning approaches, using the following definitions. In split learning, the model is partitioned into segments, with early layers trained locally and intermediate outputs (e.g., activations) passed to another party for further training. In transfer learning, a model trained by one party is fine-tuned or extended by another using local data. As long as only model components, intermediate representations, or parameters are exchanged—and no primary data is shared—these methods are considered decentralized under our operational framework.
-
Secure Multi-Party Computation (SMPC): Parties collaboratively compute a shared model using cryptographic protocols that ensure privacy of inputs. While SMPC is a privacy-enhancing technology rather than a learning paradigm per se, under our framework it qualifies as a decentralized learning approach when used to support joint model training without data exposure.
To resolve classification ambiguity—especially for hybrid or multi-stage training setups—we applied the following rule: If no raw (primary) data is shared between parties throughout the model development process, the approach is classified as decentralized, regardless of whether models, parameters, or representations are exchanged.
For the selection process, papers retrieved through the search strategy were evaluated by researchers acting independently and blinded for each other’s decisions. Each paper was classified by two researchers, with a total of seven reviewers. Whenever there was not a complete agreement on the decision, the researchers reviewed their decisions and discussed them to achieve a consensus. A third senior researcher was identified to resolve potential remaining conflicts. No automation tools were used during the selection process.
Due to a longer than expected initial article selection and the fast-moving research field, it was decided to update and apply the search strategy a second time, using the same methodology. The complete selection process is summarized using the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) 2020 flow diagram in accordance with the corresponding guidelines44.
Data collection process
Data were collected from the full-text version of the selected articles by two researchers, using a prepared online document piloted before its implementation. Researchers worked on different articles and discussed any doubts regarding the process to produce a harmonized data collection. The first author conducted a subsequent data collection verification looking for wrong, unclear, or missing records. Both researchers agreed by consensus on the version of the database reported. Data collection was organized in three ordered steps: general article information, models information and performance comparison information.
Regarding the model demands, we extracted reported data on multiple dimensions. First, for time-based metrics, we identified the following variables: training time, computation time, communication time, execution/inference/prediction time, encryption/decryption time, searching time and latency measurements. As far as resources consumption goes, the following metrics were covered: memory consumption (server and client memory), energy consumption and battery capacity requirements, power consumption and bandwidth consumption. Communication and data transfer variables considered were data upload volumes and total moved/transferred data. Specific privacy budget (ε-differential privacy parameters) and ξ, ζ-differential privacy impacts on performance metrics were collected.
Effect measures
The primary effect measures were the performance metrics values of the decentralized learning models and their non-decentralized counterparts. These values were extracted directly from the included studies. To explore non-parametric effect sizes the Wilcoxon two-sample paired signed-rank test were used, comparing the distributions of the individual performance comparisons. Estimates of effect sizes and their respective magnitude are presented.
Synthesis methods
Data collected were grouped by each performance metric and divided in the classes of the following variables: decentralized learning architecture, larger clinical domain and clinical application. Individual performance metrics with at least 30 comparisons collected were explored. An online dashboard was produced to allow for a customized search of relevant performance comparisons, using the Shiny R package – https://jmdiniz.shinyapps.io/phdiniz_systematic_review_analysis/.
The distribution of individual model performance differences between decentralized and non-decentralized alternatives across the difference performance metrics is presented using histograms and calculating their median difference, the 25th percentile and the 75th percentile, as well as the bootstrapped 95% Confidence Intervals, based on 10.000 simulations.
For sensitivity analysis, variations of these histograms are produced without the contributions of the study with the most observations – available in the Supplementary Material in Supplementary Figs. 43–56.
Specific detailed syntheses were produced for performance metrics-larger clinical domain-clinical application combinations, for instances with at least 10 comparisons and featuring at least 5 different studies.
Given the heterogeneity of clinical domains and applications, to assess clinically acceptable performance we set a threshold value of 0.80 (80%). Using this standard, we examined scenarios where local models failed to achieve clinical viability (<0.80) but DL achieved acceptable performance (≥0.80). Moreover, we identified the cases in which both local and decentralized models are clinically viable, but DL is superior, as well as instances in which local performance is clinically viable, but DL are not. The symmetric analysis was conducted considering centralized and decentralized models.
The only data processing concerned the conversion of values presented in percentages in some instances. Due to the heterogeneity in the data collected, no meta-analysis was conducted.
For each comparison and metric pairing, data were segmented into 10 equal-width intervals based on the range of the decentralized model performance. Within each segment, decentralized models were compared to their counterparts, based on the paired performance comparisons. In each facet, both the decile and the corresponding decentralized model performance range are showcased.
Evidence appraisal
We applied the PROBAST + AI tool and the TRIPOD checklist for model type45,46 to the 25 most cited included research papers. For each paper, up to two models were considered, in order of presentation. Due to their inherent limitations, we opted to exclude TRIPOD Type 1a (i.e., all data used for model development without validation) and Type 1b articles (i.e., all data used for model development, evaluation using resampling). Using an approximation of the relative prevalence of the remaining TRIPOD types, 15 Type 2a articles, 5 Type 2b articles and 5 Type articles were included. Each article and its corresponding appraisals were conducted by a single reviewer.
Registration and protocol
The research protocol for this study was published32, on June 6th, 2023. It was previously registered with PROSPERO, under the number 393126, on February 3rd, 2023, and accessible through https://www.crd.york.ac.uk/prospero/display_record.php?ID=CRD42023393126.
Details about the changes made to the protocol, and the rationale used, are presented in the Supplementary Material.
Data availability
A dashboard for select metrics is made available. Detailed data extracted from the included studies, including data used for analyses, the data processing and analytic code, is made available upon request. Moreover, documentation is provided regarding the specific queries used, tailored to each source, including their adapted formulation and filters, to ease reproducibility. Whenever possible, a direct URL link to the query is included.
References
Omran, A. R. The epidemiologic transition: a theory of the epidemiology of population change. Milb ank. Meml. Fund. Q. 49, 509–538, https://doi.org/10.2307/3349375 (1971).
The World Bank. World Bank Open Data - Current health expenditure (% of GDP). World Bank Open Data. Accessed 29 January 2024. https://data.worldbank.org
World Health Organization. Global spending on health: Weathering the storm. December 10, 2020. accessed 29 January 2024. https://www.who.int/publications-detail-redirect/9789240017788
OECD. Fiscal Sustainability of Health Systems: How to Finance More Resilient Health Systems When Money Is Tight? Organisation for Economic Co-operation and Development; Accessed January 29, 2024. https://www.oecd-ilibrary.org/social-issues-migration-health/fiscal-sustainability-of-health-systems_880f3195-en 2024.
Licchetta M. & Stelmach M. Fiscal Sustainability Analytical Paper: Fiscal Sustainability and Public Spending on Health. Office for Budget Responsibility (United Kingdom) Accessed 13 September 2023. https://obr.uk/docs/dlm_uploads/Health-FSAP.pdf
World Health Organization. World Health Statistics 2024: Monitoring Health for the SDGs, Sustainable Development Goals. World Health Organization; 2024:96. Accessed 4 April 2025. https://iris.who.int/bitstream/handle/10665/376869/9789240094703-eng.pdf?sequence=1
World Health Organization, World Bank. Tracking Universal Health Coverage 2023 Global Monitoring Report. 2023:160. https://www.who.int/publications/i/item/9789240080379
Galea, G. et al. Quick buys for prevention and control of noncommunicable diseases. Lancet Reg. Health Eur. https://doi.org/10.1016/j.lanepe.2025.101281 (2025).
Villalobos P, et al. Position: will we run out of data? Limits of LLM scaling based on human-generated data. In Proceedings of the 41stInternational Conference on Machine Learning. Accessed January 8, https://openreview.net/forum?id=ViZcgDQjyG (2024)
Udandarao V, et al. No "Zero-Shot" Without Exponential Data: Pretraining Concept Frequency Determines Multimodal Model Performance. Adv Neural Inf Process Syst. 38. Accessed January 8, https://proceedings.neurips.cc/paper_files/paper/2024/file/715b78ccfb6f4cada5528ac9b5278def-Paper-Conference.pdf (2024)
Liu, F. & Panagiotakos, D. Real-world data: a brief review of the methods, applications, challenges and opportunities. BMC Med. Res. Methodol. 22, 287 (2022).
McMahan B, et al. Communication-efficient learning of deep networks from decentralized data. In Proceedings of the 20th International Conference on Artificial Intelligence and Statistics. 1273-1282. Accessed January 8, https://proceedings.mlr.press/v54/mcmahan17a.html (2017)
Tajabadi, M. & Heider, D. Fair swarm learning: improving incentives for collaboration by a fair reward mechanism. Knowl.Based Syst. 304, 112451 (2024).
Linardos, A., Kushibar, K., Walsh, S., Gkontra, P. & Lekadir, K. Federated learning for multi-center imaging diagnostics: a simulation study in cardiovascular disease. Sci. Rep. 12, 3551 (2022).
Lee, H. et al. Federated learning for thyroid ultrasound image analysis to protect personal information: validation study in a real health care environment. JMIR Med. Inform. 9, e25869 (2021).
Lo, J. et al. Federated learning for microvasculature segmentation and diabetic retinopathy classification of OCT Data. Ophthalmol. Sci. 1, 100069 (2021).
Soltan, A. A. S. et al. A scalable federated learning solution for secondary care using low-cost microcomputing: privacy-preserving development and evaluation of a COVID-19 screening test in UK hospitals. Lancet Digit Health 6, e93–e104 (2024).
Haggenmüller, S. et al. Federated learning for decentralized artificial intelligence in melanoma diagnostics. JAMA Dermatol. 160, 303–311 (2024).
Kundu, D. et al. Federated deep learning for monkeypox disease detection on GAN-augmented dataset. IEEE Access. 12, 32819–32829 (2024).
Nguyen, T. P. V. et al. Lightweight federated learning for STIs/HIV prediction. Sci. Rep. 14, 6560 (2024).
Warnat-Herresthal, S. et al. Swarm Learning for decentralized and confidential clinical machine learning. Nature 594, 265–270 (2021).
Saldanha, O. L. et al. Swarm learning for decentralized artificial intelligence in cancer histopathology. Nat. Med. 28, 1232–1239 (2022).
Breiman, L. Bagging predictors. Mach. Learn. 24, 123–140 (1996).
European Parliament. Regulation (EU) 2024/1689 of the European Parliament and of the Council of 13 June 2024 Laying down Harmonised Rules on Artificial Intelligence and Amending Regulations (EC) No 300/2008, (EU) No 167/2013, (EU) No 168/2013, (EU) 2018/858, (EU) 2018/1139 and (EU) 2019/2144 and Directives 2014/90/EU, (EU) 2016/797 and (EU) 2020/1828 (Artificial Intelligence Act) (Text with EEA Relevance).; 2024. Accessed 3 November 2024. http://data.europa.eu/eli/reg/2024/1689/oj/eng
Brauneck, A. et al. Federated machine learning, privacy-enhancing technologies, and data protection laws in medical research: scoping review. J. Med. Internet Res. 25, e41588 (2023).
Woisetschläger, H. et al. Federated Learning Priorities Under the European Union Artificial Intelligence Act. Preprint at https://doi.org/10.48550/arXiv.2402.05968 (2024).
Zerka, F. et al. Systematic review of privacy-preserving distributed machine learning from federated databases in health care. JCO Clin. Cancer Inform. 4, 184–200 (2020).
Agbo, C. C., Mahmoud, Q. H. & Eklund, J. M. Blockchain technology in healthcare: a systematic review. Healthcare 7, 56 (2019).
Crowson, M. G. et al. A systematic review of federated learning applications for biomedical data. PLOS Digital Health 1, e0000033 (2022).
Qammar, A., Karim, A., Ning, H. & Ding, J. Securing federated learning with blockchain: a systematic literature review. Artif. Intell. Rev. Published online September 16, 1–35. https://doi.org/10.1007/s10462-022-10271-9 (2022).
Antunes, R. S., André da Costa, C., Küderle, A., Yari, I. A. & Eskofier, B. Federated learning for healthcare: systematic review and architecture proposal. ACM Trans. Intell. Syst. Technol. 13, 54:1–54:23 (2022).
Diniz, J. M. et al. Comparing decentralized learning methods for health data models to nondecentralized alternatives: protocol for a systematic review. JMIR Res. Protoc. 12, e45823 (2023).
Souza, R. et al. A multi-center distributed learning approach for Parkinson’s disease classification using the traveling model paradigm. Front. Artif. Intell. 7, 1301997 (2024).
Qu, L., Balachandar, N., Zhang, M. & Rubin, D. Handling data heterogeneity with generative replay in collaborative learning for medical imaging. Med. Image Anal. 78, 102424 (2022).
Khalil, S. S., Tawfik, N. S. & Spruit, M. Exploring the potential of federated learning in mental health research: a systematic literature review. Appl. Intell. 54, 1619–1636 (2024).
Teo, Z. L. et al. Federated machine learning in healthcare: A systematic review on clinical applications and technical architecture. CR Med. 5, 101419 (2024).
Hiwale, M., Walambe, R., Potdar, V. & Kotecha, K. A systematic review of privacy-preserving methods deployed with blockchain and federated learning for the telemedicine. Healthc. Anal. 3, 100192 (2023).
Sohan, M. F. & Basalamah, A. A systematic review on federated learning in medical image analysis. IEEE Access. 11, 28628–28644 (2023).
Wagner, I. & Eckhoff, D. Technical privacy metrics: a systematic survey. ACM Comput. Surv. 51, 57:1–57:38 (2018).
Kaabachi, B. et al. A scoping review of privacy and utility metrics in medical synthetic data. npj Digit Med. 8, 60 (2025).
Xu, J., Wu, Z., Wang, C. & Jia, X. Machine unlearning: solutions and challenges. IEEE Trans. Emerg. Top. Comput. Intell. 8, 2150–2168 (2024).
Bercea, C. I., Wiestler, B., Rueckert, D. & Schnabel, J. A. Evaluating normative representation learning in generative AI for robust anomaly detection in brain imaging. Nat. Commun. 16, 1624 (2025).
Ouzzani, M., Hammady, H., Fedorowicz, Z. & Elmagarmid, A. Rayyan—a web and mobile app for systematic reviews. Syst. Rev. 5, 210 (2016).
Page, M. J. et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372, n71 (2021).
Moons, K. G. M. et al. PROBAST+AI: an updated quality, risk of bias, and applicability assessment tool for prediction models using regression or artificial intelligence methods. BMJ 388, e082505 (2025).
Collins, G. S., Reitsma, J. B., Altman, D. G. & Moons, K. G. M. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD statement. Ann. Intern. Med. 162, 55–63 (2015).
Acknowledgements
Some authors (J.M.D., J.S., R.R., C.A., A.F.) were researchers of the “Secur-e-Health: Privacy preserving cross-organizational data analysis in the healthcare Sector” (ITEA 20050), cofinanced by the North Regional Operational Program (NORTE 2020) under the Portugal 2020 and European Regional Development Fund, with the reference NORTE-01-0247-FEDER-181418. This participation only occurred during part of the research work on this paper. The funding agency did not have a role in either the study design, the data analysis, the manuscript preparation, or the submission of this work.
Author information
Authors and Affiliations
Contributions
Design experimental methodology—J.M.D., H.V., R.R., C.A., J.S., and A.F. Collect data–J.M.D., H.V., R.R., C.A., D.R., P.R., A.T., Y.G., and J.S. Interpret data and results—J.M.D. and A.F. Create figures, charts, and visualizations—J.M.D. Review and edit the complete manuscript—All authors Format submission according to Nature guidelines—J.M.D. Prepare supplementary materials and data availability statements—J.M.D. Review and approve final proofs before publication—All authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diniz, J.M., Vasconcelos, H., Rb-Silva, R. et al. Comparing decentralized machine learning and AI clinical models to local and centralized alternatives: a systematic review. npj Digit. Med. 9, 174 (2026). https://doi.org/10.1038/s41746-025-02329-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02329-z