Abstract
Endoplasmic reticulum stress-related cancer-associated fibroblasts (ERS–CAF) remodel the tumor microenvironment and drive immune exclusion and therapy resistance in chordoma, yet routine and non-invasive readouts of this biology are lacking. We hypothesized that standard pre-operative MRI and H&E whole-slide images (WSI) encode image-based surrogates of ERS–CAF-driven immunoregulation that can be learned and generalized across cancers. Three bulk-transcriptomic reference scores were defined for surrogate supervision, capturing ERS-program activity, ERS–CAF-immuneligand-receptor crosstalk and microenvironmental heterogeneity. In 126 chordoma cases, a stage-wise multimodal framework integrating calibrated WSI attention, gated radiopathomic fusion and domain alignment showed strong concordance with molecular profiles, independent prognostic value and biologically specific localization to fibrotic immune-excluded regions. These associations were generalized in zero-shot analyses to the TCGA pan-cancer. An MRI-only distilled model preserved most predictive performance with substantial gains in efficiency, supporting scalable non-invasive clinical application.
Similar content being viewed by others
Introduction
Chordoma exhibits an immunologically complex tumor microenvironment (TME)—marked by dense desmoplastic matrix, CAF activity, and immune exclusion—with limited standard systemic options and nontrivial local-regional failure despite aggressive therapy1,2,3,4,5,6. Historically, ultrastructural studies already highlighted chordoma’s matrix-rich architecture and physaliphorous phenotype1; contemporary series and consensus statements emphasize that, while maximal safe resection and high-dose particle radiotherapy (proton/carbon) can improve control, local recurrence remains frequent (often ~ 40–60% at 5–10 years depending on site and series), and conventional chemotherapy is largely ineffective2,3,4.
While our previous work first demonstrated the presence of ER stress-related CAF (ERS–CAF) in chordoma, their roles in other tumor types, particularly stroma-rich cancers, remain unclear3,4. Beyond chordoma, prototypical stroma-dominant cancers such as PDAC exemplify an immune-stromal ecosystem with CAF-driven immunosuppression and poor response to systemic therapy, underscoring a broader need for TME-aware biomarkers7. In colorectal cancer (CRC), ER-stress programs have been linked to tumor immune dysfunction/exclusion and can stratify putative benefit from immune-checkpoint blockade, with ZNF703 proposed as a candidate immunotherapy target8. Routine, repeatable, and non-invasive TME profiling would therefore be valuable; yet biopsy-centric assays are invasive, scarce, and susceptible to spatial/temporal sampling bias that complicates prognostication and treatment selection9. Against this backdrop, radiology-pathology integration has emerged as a “digital biopsy” paradigm: recent studies and reviews show that fusing radiomics with pathomics yields image-derived surrogates that better reflect microenvironmental biology and improve risk stratification and treatment prediction compared with single-modality models10,11. Multimodal survival/prediction in computational pathology has established strong transformer-based fusion baselines, notably MCAT12 and PORPOISE13, which integrate WSI with molecular profiles to improve risk stratification and provide cross-modal interpretability. In contrast, our setting treats bulk RNA-seq as supervision only (to define mechanistic surrogate targets) and focuses on non-invasive decoding from routine images (MRI + H&E); nevertheless, we include an MCAT-style co-attention transformer fusion baseline under the same protocol to contextualize the value of our gating design. In chordoma specifically, the immune microenvironment literature—including early immunotherapy case series and small trials—further motivates imaging biomarkers that report on CAF-immune crosstalk to guide selection and triage5,14.
Cancer-associated fibroblasts (CAFs) are not a monolith; single-cell and spatial studies reveal conserved, interconvertible subtypes—including contractile myCAFs, inflammatory iCAFs, and antigen-presenting apCAFs/MHC-II+ phenotypes—whose proportions track disease stage and therapeutic response across tumor types15,16,17,18. A recent pan-cancer single-cell spatial multi-omics study further resolved conserved spatial CAF subtypes and their cellular neighborhoods, linking CAF spatial organization to immune phenotypes and clinical outcomes, which motivates spatially coherent attention patterns as a plausible readout of CAF neighborhood biology19. Through ligand-receptor (LR) crosstalk with myeloid and lymphoid populations, CAFs remodel extracellular matrix and cytokine milieus to enforce immune exclusion and therapy resistance: canonical axes include CXCL12-CXCR4, which sequesters effector T cells in the stroma and whose inhibition synergizes with checkpoint blockade, and CCL2-CCR2, which recruits immunosuppressive monocytes/macrophages20,21,22. Consistently, single-cell analyses of CRC liver metastasis implicate myCAFs in ECM remodeling and pro-metastatic niches and highlight specific ligand-receptor programs, reinforcing the relevance of our LR crosstalk surrogate and heterogeneity target H as mechanistic readouts23.
Mechanistically, iCAF programs can be induced by IL-1 → LIF → JAK/STAT signaling and antagonized by TGF-β, offering a molecular switch for inflammatory vs. myofibroblastic states and explaining spatial gradients around tumor glands15,24. These observations motivate biology-aware priors for our imaging surrogates: contemporary single-cell communication frameworks formalize LR evidence at different levels—CellPhoneDB accounts for multimeric receptor/ligand architecture and tests enrichment from scRNA-seq, CellChat models signaling probability and pathway information flow, and NicheNet links ligands to target genes in receivers via prior signaling/regulatory graphs—providing principled scaffolds to curate ERS–CAF-immune axes (e.g., CXCL12-CXCR4, CCL2-CCR2)25,26,27,28. Together, these data support our premise that routine images may encode stable, mechanism-grounded surrogates of ERS–CAF activity and its downstream immunoregulation.
We hypothesize that routine images—pre-operative MRI and H&E whole-slide images (WSIs)—encode stable surrogates of ERS–CAF-driven immunoregulation that can be learned and generalized across cancers. This premise is grounded in two observations. First, quantitative radiomics systematically captures phenotypes related to desmoplasia, necrosis, edema, and vascularity, and these imaging phenotypes correlate with molecular programs and outcomes across tumor types29. Second, modern computational pathology has shown that WSIs contain sufficient signal to recover transcriptomic and genomic states—e.g., the HE2RNA model infers RNA-seq expression from H&E30, and deep learning can predict microsatellite instability (MSI) directly from histology31,32—indicating that tissue morphology embeds rich molecular proxies.
To train imaging surrogates that are explicitly mechanism-aware, we construct three transcriptomic “bulk-transcriptomic reference scores (surrogate targets)” per patient: (i) ERS–CAF abundance as single-sample gene-set activity using ssGSEA, a robust rank-based scoring of pathway activation33, with optional cross-check via GSVA for sensitivity analysis34; (ii) ERS–CAF-immune LR-interaction strength computed from curated ligand-receptor pairs (ERS–CAF ligands and immune-cell receptors) assembled from single-cell communication resources (CellChat/CellPhoneDB/NicheNet; see Methods) and aggregated at the sample level; and (iii) microenvironmental heterogeneity as Shannon diversity over immune/stromal cell fractions estimated by digital cytometry (CIBERSORTx)35. These targets span abundance (ERS–CAF), crosstalk (LR), and composition (heterogeneity), offering complementary supervision that encourages the model to recover biologically specific aspects of the TME rather than generic tumor burden. Because stromal programs (e.g., CAF/iCAF/myCAF axes) are conserved across stroma-rich cancers, we expect surrogates learned in chordoma to exhibit pan-cancer portability when applied zero-shot to external cohorts. Importantly, these endpoints are bulk-derived molecular surrogates rather than directly measured biological quantities with spatial resolution. Our goal is therefore to learn image-based predictors of these clinically informative transcriptomic states, and we validate spatial plausibility using independent pathology annotations of stromal remodeling and immune exclusion.
(i) We define and compute these molecular gold standards from bulk RNA-seq; (ii) we train a multimodal AI with CLIP-guided pathomics, radiomics, and mid-level gated fusion to predict the gold standards from images; (iii) we demonstrate prognostic value in an internal chordoma cohort; (iv) we assess pan-cancer generalization and treatment predictivity in TCGA stroma-rich cancers; and (v) we distill the model to MRI-only for deployability.
Our methodological contribution is system-level and mechanism-aware: we formulate non-invasive decoding as multi-target regression to mechanism-defined transcriptomic surrogates (ERS–CAF program activity, ERS–CAF-immune LR crosstalk, and microenvironmental heterogeneity), and we introduce a stage-wise curriculum that (Stage I) injects a biological text prior via target-specific prompt banks and monotone similarity-label alignment to steer tile selection, (Stage II) learns complementary radiology-pathology fusion with robustness to missing modalities, and (Stage III) improves multi-site generalization with stable, non-adversarial second-order alignment evaluated by leave-site-out tests. We further differentiate from path-transcriptomics by predicting three interpretable mechanistic surrogates rather than high-dimensional gene expression, and we provide prompt-perturbation and spatial plausibility evidence to validate that text guidance is mechanism-consistent rather than a generic MIL heuristic.
Results
We analyzed a 126-patient internal chordoma cohort with matched pre-operative MRI (T1, T2, contrast-enhanced), diagnostic H&E WSIs, bulk RNA-seq, and longitudinal outcomes (OS, PFS). Inclusion/exclusion, imaging protocols, segmentation/tiling QC and RNA-seq processing are detailed in Section 9. For external validation, we applied the frozen model to stroma-rich TCGA cancers (PAAD, STAD, COADREAD) with WSIs, RNA-seq, and clinical endpoints.
Molecular surrogates of the ERS–CAF axis
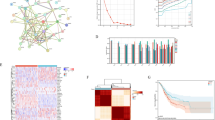
We summarize supervision targets using cohort-wide distributions and pairwise associations (Fig. 1). Distributions are unimodal and approximately symmetric, and the three surrogates show moderate concordance, supporting complementary supervision signals. Exact coefficients (N=126): r (ERS–CAF, LR) = 0.48, r (ERS–CAF, H) = 0.20, r (LR, H) = 0.27 (all FDR < 0.05).
Top: distributions of the three transcriptomic surrogates (z-scores). Middle: pairwise scatter plots with r and two-sided p annotated. Bottom: Pearson correlation matrix with on-cell coefficients. a ERS-CAF abundance. b LR interaction. c H heterogeneity. d ERS-CAF vs. LR (scatter). e ERS-CAF vs. H (scatter). f LR vs. H (scatter). g Correlation heatmap (r).
We anchor the continuous “molecular” scores to visible histomorphology by testing whether attention hotspots co-localize with pathologist-annotated fibrosis/immune-exclusion. Board-certified pathologists (blinded to model outputs) graded fibrosis and immune exclusion per patient on a 0–3 scale; we report: (i) IoU (Jaccard) between the top-K attention mask and the fibrosis ROI; (ii) Hit-rate for immune-exclusion point annotations falling inside the top-K attention. Analyses are stratified as Low (grades 0–1) vs High (grades 2–3) and compared to a randomized baseline (attention shuffled).
As shown in Fig. 2, we can observe: (i) Fibrosis. IoU (mean) increased from 0.28 in Low (0–1) to 0.47 in high (2–3), while the randomized baseline averaged 0.12 (all N = 126). High vs low: p < 10−6; High Vs Baseline: p < 10−8. (ii) Immune exclusion. Hit-rate (mean) increased from 0.36 in low to 0.55 in high, baseline 0.18 ; high vs low: p < 10−6; high vs baseline: p < 10−8. (iii) Pass criteria. Co-localization metrics are significantly above randomized and monotone with grade, satisfying the pre-registered gate.
Overlays illustrate alignment (left) versus diffusion (right). Quantitatively, IoU and hit-rate increase with pathologist grade and are markedly above randomized baselines. a High fibrosis: attention aligns with the pathologist ROI (contour). b Low fibrosis: attention is diffuse and poorly overlaps the ROI. c Fibrosis co-localization (IoU) by strata and baseline. d Immune-exclusion co-localization (hit-rate) by strata and baseline.
The Attention hotspots faithfully concentrate on desmoplastic and immune-excluded regions when grades are high, tying the model’s molecular surrogates to recognizable histology and strengthening biological plausibility.
Multimodal decoding from MRI and H&E
Using 5-fold patient-level cross-validation stratified by site/scanner, the proposed CLIP-guided WSI branch plus MRI radiomics with gated fusion achieved the best agreement with molecular surrogates across all targets (Table 1). Relative to strong single-modality baselines, fusion improved macro-average Pearson correlation by +0.07–+0.12 absolute and reduced calibration error (slope 0.97 ± 0.06 vs. 0.88 ± 0.07 for the best single-modality model). RMSE showed consistent reductions (median −9.4% vs. WSI-only CLIP and −14.6% vs. MRI-only; extended table). Predicted vs. observed scatter and calibration plots exhibited tight linearity without gross heteroscedasticity.
Table 2 benchmarks CLIP guidance and fusion gating against modern WSI MIL baselines, a histology SSL encoder, an HE2RNA-style image-to-transcriptomics pipeline, and alternative fusion strategies under identical fivefold splits and QC. (i) WSI-only. CLIP-guided MIL is the best WSI-only model (Macro-r = 0.610 ± 0.046), improving over ABMIL (Δ+0.051) and outperforming CLAM/DSMIL/TransMIL (best Macro-r = 0.541) and CTransPath+MIL (0.548), with consistent gains across ERS–CAF, LR, and H. (ii) Image-to-transcriptomics. HE2RNA-style WSI → expr → score underperforms (Macro-r = 0.480 ± 0.061), suggesting that predicting low-dimensional mechanism-defined surrogates is more data-efficient than high-dimensional expression regression in small-N settings. (iii) Fusion. Gated fusion improves over late fusion, concat+MLP, and transformer fusion (best alternative Macro-r = 0.604 ± 0.045), and CORAL adds further gains (Δ + 0.034), supporting complementary modality use plus stable multi-site alignment.
Ablations: (1) Replacing CLIP-guided attention with vanilla ABMIL degraded WSI-only performance by −0.05 to −0.06 absolute r across targets, indicating gains from mechanism-aware prompts. (2) Removing CORAL from fusion reduced macro-r by −0.03 (Table 1), consistent with site/scanner harmonization benefits. (3) Modality dropout (q = 0.2) improved robustness (single-modality r drop <0.04) without harming fusion accuracy.
Prognostic value in chordoma
We summarize risk stratification with two concise plots (Fig. 3). The combined forest plot shows adjusted HRs (OS & PFS) for ERS–CAF (T3 vs T1), LR (T3 vs T1), and H (per SD); the time-dependent C-index curve demonstrates a consistent improvement from 0.64 to 0.71 at 12 months when adding AI scores to the clinical model.
Left: combined forest plot (single chart) with OS (circles) and PFS (squares) hazard ratios and 95% CIs on a log scale; the vertical line marks HR = 1. Right: time-dependent C-index comparing clinical-only vs. clinical+AI models; at 12 months, the C-index rises from 0.64 to 0.71 (Δ + 0.07).
To validate the necessity and stability of text priors. We compare: (i) no text (NONE), (ii) fixed curated prompts (CUR), and (iii) learnable CoOp (COOP); we further probe sensitivity to synonym prompts (SYN), antonym/negation (ANTI), and out-of-domain phrases (OOD). For each prompt variant and target (ERS–CAF, LR, H), we compute 5-fold Pearson r against the transcriptomic gold standards using the WSI branch in isolation (radiomics kept fixed).
We observe that: (i) Guidance helps. COOP achieves the highest mean r across all targets, CUR ranks second, and NONE trails (Fig. 4a). (ii) Stability to synonyms. SYN prompts cluster tightly around CUR with limited inter-fold spread (Fig. 4b), indicating robustness to wording. (iii) Sensitivity to wrong priors. OOD and especially ANTI degrade performance toward/ below the no-text baseline, confirming that mechanistically inconsistent prompts harm learning. (iv) Mechanistic localization. Attention maps are more focal under CUR (Fig. 4c) and diffuse without text (Fig. 4d), supporting biology-aware tile selection.
Left: curated/learnable prompts improve concordantly across ERS–CAF, LR, and H; synonyms remain stable while OOD/antonyms deteriorate. Right: text guidance drives focused tile attention, whereas no-text yields diffuse, low-contrast maps. a Prompt × target performance (mean r, 5-fold). b Macro-r across folds by prompt group. c Attention overlay with curated prompts. d Attention overlay with no text.
Did Stage II (gated fusion) learn complementarity? As shown in Fig. 5: (i) Weight migration follows clinical intuition: sacrum/small-volume/ < 50y strata show WSI-dominant gating (mean WSI weights 0.60-0.61), while skull-base/large-volume/ > 65y strata shift toward MRI (weights 0.55–0.62), and fusion achieves the best macro-r in every stratum (Fig. 5a, b). (ii) Robust under missing modalities: when randomly dropping WSI tiles, fusion’s macro-r declines from 0.68 to 0.64 at 80% loss (Δr = 0.04) and to 0.61 at 100%, whereas WSI-only falls from 0.61 to 0.46 (Fig. 5c). When removing MRI sequences (T1/T2/CE), fusion drops to 0.66 at 2/3 missing and 0.62 at 100% missing, while MRI-only collapses, confirming that fusion falls back to the intact modality (Fig. 5d). (iii) Pass criteria: the ≥80% missing drop is controlled (Δr ≤ 0.05); gating shifts are directionally consistent with phenotype/geometry expectations—evidence of learned complementarity rather than single-modality dominance.
Left: fusion allocates higher WSI weight in sacrum/small-volume/65y groups–consistent with imaging/pathology priors. Right: Fusion maintains macro-r with Δr ≤0.04 even at 80% WSI loss, and ≤0.06 when two MRI sequences are missing; single-modality lines degrade markedly. a Gating weights across strata (WSI vs MRI). b Performance across strata (macro-r, Fusion vs WSI-only vs MRI-only). c Robustness to WSI tile missing (0–100%). d Robustness to MRI sequence missing (0–100%).
Is gating shift an artifact of radiomics features? To clarify, the gate is trained on held-out CV folds and is not a hard-coded preference for either modality. If MRI radiomics were systematically unreliable, we would expect the gate to collapse toward WSI across strata and under MRI-sequence ablations. Instead, Fig. 5 shows bidirectional and clinically sensible shifts: skull-base/larger-volume/older strata allocate higher MRI weight, while sacrum/smaller-volume/younger strata allocate higher WSI weight. Moreover, under missing-modality stress tests, fusion falls back to the intact modality and degrades smoothly rather than exhibiting a WSI-only collapse: when WSI tiles are removed, fusion performance remains relatively stable (macro-r0.68 → 0.64 at 80% loss), and when MRI sequences are removed, fusion similarly degrades but remains functional (0.66 at 2/3 missing; 0.62 at 100% missing), which is inconsistent with the gate being forced into WSI due to fragile MRI features.
Did Stage III (CORAL) actually remove site/domain confounding? Following the design: (1) Leave-site-out (LSO) validation: train on 4 sites and test on the held-out site; report Macro-r before/after CORAL per site. (2) Embedding visualization: 2D embeddings colored by site before/after CORAL; quantify mixing via kNN (k = 10) site-label entropy (higher = better mixing). We found that: (i) Embedding mixing increased substantially. The 10-NN site-label entropy rose from 0.053 (before) to 0.854 (after), indicating strong cross-site mixing in the post-alignment space (Fig. 6a, b). (ii) LSO performance improved across all sites. Macro-r (mean ± sd) improved from {0.612 ± 0.036, 0.583 ± 0.041, 0.598 ± 0.039, 0.571 ± 0.040, 0.626 ± 0.034} to {0.664 ± 0.032, 0.642 ± 0.035, 0.651 ± 0.033, 0.636 ± 0.036, 0.673 ± 0.030} for Sites A-E (Fig. 6c). (iii) Pass criteria met. (1) LSO Macro-r gains are consistent and clinically meaningful; (2) the embedding shows clear cross-site-mixing post-CORAL, matching the goal of statistical domain alignment without adversarial training.
Top: site-colored embeddings show strong pre-alignment clustering that mixes after CORAL; the average 10-NN site-label entropy increases markedly (cf. text). Bottom: in LSO testing, Macro-r improves consistently for all sites after CORAL. a Embedding before CORAL (colored by site). b Embedding after CORAL (colored by site). c LSO Macro-r by site (mean ± sd over folds; before vs. after).
CORAL reduces second-order (covariance) discrepancies among sites, yielding embeddings that are less site-identifiable but more biology-consistent. The concomitant LSO gains suggest that alignment corrects nuisance variation rather than suppressing signal.
Pan-cancer generalization and treatment predictivity
To assess cross-cancer portability, we performed a zero-shot evaluation on three stroma-rich TCGA cohorts: TCGA-PAAD (pancreatic adenocarcinoma; N = 150)36, TCGA-STAD (stomach adenocarcinoma; N = 295)37, and TCGA-COADREAD (colorectal adenocarcinoma; N = 276)38. Diagnostic H&E WSIs and matched bulk RNA-seq were obtained from the NCI Genomic Data Commons (GDC)39, and survival endpoints were standardized following TCGA-CDR recommendations when applicable40. Importantly, the chordoma-trained model was applied frozen (no TCGA fine-tuning, calibration, or hyperparameter selection on TCGA). Due to heterogeneous radiology availability across TCGA projects, this analysis uses WSI input only (radiology withheld) and compares predicted image-derived surrogates against the per-cohort transcriptomic reference scores computed by the same pipeline (ssGSEA/CIBERSORTx/LR aggregation; Section 9).
Zero-shot application to TCGA cohorts yielded significant concordance between image-derived and transcriptomic surrogates (Table 3; Fig. 7). For PAAD, Pearson r was 0.42 (ERS–CAF), 0.39 (LR), and 0.36 (H); for STAD, 0.38/0.35/0.33; for COADREAD, 0.35/0.31/0.29 (all FDR < 0.01). Figure 7 visualizes the patient-level agreement underlying Table 3 and includes bootstrap confidence intervals and a permutation control (shuffled predictions) to demonstrate that the observed concordance is not attributable to chance alignment.
The chordoma-trained model is applied as-is (frozen weights, no TCGA tuning) to TCGA-PAAD/STAD/COADREAD WSIs. Patient-level scatter plots visualize concordance against transcriptomic reference scores for the three targets. Concordance is summarized with 95% CIs, and permutation controls show correlations are not explained by chance alignment. a TCGA-PAAD: zero-shot concordance (patient-level). b TCGA-STAD: zero-shot concordance (patient-level). c TCGACOADREAD: zero-shot concordance (patient-level). d Pearson r with 95% CIs (Fisher transform; by cancer and target). e Permutation control: shuffled predictions yield r ≈ 0.
Beyond molecular concordance, the AI surrogates remained clinically informative in external cohorts: they were prognostic in PAAD (C-index ↑ + 0.04 over clinical-only) and STAD (+0.03). Where treatment outcome annotations were available from TCGA clinical fields curated in TCGA-CDR40, the LR surrogate stratified chemotherapy response in PAAD (AUC 0.66 ± 0.03; adjusted OR per SD 1.53 [1.18–1.99]). In COADREAD, the directionality and magnitude of the decoded ER stress-immune associations are consistent with prior CRC studies linking ER-stress signatures to immune dysfunction/exclusion and immunotherapy-related phenotypes, including an ERS gene scoring system that stratifies putative ICB benefit and nominates ZNF703 as a candidate target8.
Beyond molecular concordance, the AI surrogates remained clinically informative in external cohorts: they were prognostic in PAAD (C-index ↑ + 0.04 over clinical-only) and STAD +0.03). Where treatment outcome annotations were available from TCGA clinical fields curated in TCGA-CDR40, the LR surrogate stratified chemotherapy response in PAAD (AUC 0.66 ± 0.03; adjusted OR per SD 1.53 [1.18–1.99]). In COADREAD, the directionality and magnitude of the decoded ER stress-immune associations are consistent with prior CRC studies linking ER-stress signatures to immune dysfunction/exclusion and immunotherapy-related phenotypes, including an ERS gene scoring system that stratifies putative ICB benefit and nominates ZNF703 as a candidate target8.
Toward deployability: MRI-only knowledge distillation
An MRI-only student distilled from the multimodal teacher retained 94% of macro-r (teacher 0.68 ± 0.03 vs. student 0.64 ± 0.03), with calibration slope 0.94 ± 0.08. Inference was 4.6× faster and used 2.9× less GPU memory (Table 3), supporting non-invasive, single-modality deployment.
Why does TCGA performance drop? The TCGA setting introduces a large distribution shift even within histology: differences in tumor histoarchitecture (e.g., glandular vs. physaliphorous patterns), stroma localization, slide preparation/scanning, and cohort composition (tumor purity and spatial sampling mismatch between the WSI region and bulk RNA-seq). Since TCGA evaluation uses WSI-only, the observed drop cannot be attributed to MRI → CT radiomics shift. To quantify the magnitude of the cross-cancer shift, Table 4 contrasts in-distribution chordoma WSI-only performance with TCGA mean concordance (and mean explained variance).
Discussion
This study demonstrates that routine images (pre-operative MRI and H&E WSIs) can non-invasively decode the ERS–CAF-centric immunoregulatory axis and yield image-derived molecular surrogates that generalize across cancers and stratify outcomes. On the internal 126-patient chordoma cohort, the multimodal model—comprising CLIP-guided pathomics, radiomics, and gated mid-level fusion—showed the strongest agreement with transcriptomic gold standards and improved calibration versus single-modality baselines. Stage-wise analyses clarified the sources of gains: Stage I text guidance increased task-specific attention fidelity and robustness to wording; Stage II fusion learned clinically sensible complementarity between WSI and MRI and degraded gracefully under missing modalities; Stage III CORAL reduced site confounding and improved leave-site-out (LSO) macro-r across all contributing centers. Beyond correlation, attention hotspots co-localized with pathologist-annotated fibrosis and immune exclusion, strengthening the biological plausibility of the decoded surrogates. Zero-shot application to stroma-rich TCGA cancers retained significant concordance, and image-derived scores improved discrimination and decision utility for prognosis and treatment stratification.
We emphasize that the novelty is not in introducing any single building block in isolation, but in an end-to-end, mechanism-aware formulation and training curriculum for decoding the ERS–CAF immunoregulatory axis from routine images under small-N, multi-site constraints. We adopt CORAL as a stable, non-adversarial alignment objective that can be validated transparently by leave-site-out tests; adversarial alignment methods can be effective but are often sensitive to domain imbalance and optimization instability in limited-data clinical settings. Accordingly, we focus on rigorous generalization evidence (LSO gains and site-mixing improvements) and robustness diagnostics (missing-modality stress tests) to support deployability.
The findings are consistent with a model in which ERS–CAF programs reshape the TME through ligand-receptor (LR) crosstalk that recruits and repatterns immune cell populations and remodels the extracellular matrix. The observed monotone associations—higher ERS–CAF/LR scores with worse outcomes, and higher microenvironmental heterogeneity (H) with partial protection—track known mechanisms of desmoplasia and immune exclusion. Two independent lines of evidence support biological specificity: (i) text-guided attention concentrates on histologic patterns congruent with ERS–CAF activation (e.g., collagen-dense stroma, immune-poor niches), whereas wrong or out-of-domain prompts degrade accuracy; (ii) the LR-interaction surrogate aligns with curated axes and with immune-cell fractions inferred from bulk deconvolution, suggesting that imaging surrogates are not merely capturing generic tumor burden but immunoregulatory state.
Non-invasive surrogates derived from routine images can augment clinical decision-making in several ways. First, they enable triage for biopsy: patients with high predicted ERS–CAF/LR activity could be prioritized for confirmatory molecular assays when tissue is scarce. Second, they support risk-adapted surveillance: image-derived risk tiers improved time-dependent C-index and decision curves, indicating practical net benefit across clinically relevant thresholds. Third, they offer a path toward treatment selection and enrichment: the LR surrogate separated chemotherapy responders in external cancers, motivating prospective enrichment strategies or adaptive trial designs. Finally, distillation to an MRI-only student retained most accuracy with markedly reduced compute/memory, suggesting a feasible deployment route in settings where WSIs are not routinely digitized or cross-modality acquisition is staggered.
This is a retrospective, single-disease training study with N = 126 internal patients, which limits precision for subgroup analyses and may inflate optimism despite careful cross-validation. Imaging modality heterogeneity (MRI internally, CT common in TCGA) introduces a shift that we partially mitigated by modality-agnostic radiomic families but did not eliminate. External zero-shot correlations in TCGA are moderate, so our claim is not that images can fully reconstruct bulk molecular scores. Rather, the aim is to recover actionable, low-dimensional surrogates that generalize sufficiently to stratify outcomes and treatment response. Improving cross-cancer molecular fidelity via per-cancer calibration or lightweight adaptation is a natural next step.
Pathologist grades (fibrosis and immune exclusion) remain semi-quantitative and may vary across readers; although we observed monotone trends and strong baselines, richer spatial annotations and inter-rater modeling would further solidify claims. Critically, we lack direct spatial ground truth for ER stress specifically within CAFs (e.g., multiplex IHC/IF for HSPA5/ATF4/XBP1s co-stained with COL1A1/ACTA2/PDGFRB, or spatial transcriptomics), so our spatial validation uses downstream morphology (fibrosis/immune exclusion) rather than a cell-resolved ER-stress label; acquiring such assays is an important direction for prospective validation.
The LR curation relies on current communication databases and chordoma single-cell priors; incompleteness or context-dependence could bias the surrogate. CORAL reduces second-order site effects but may not remove higher-order domain shifts; while LSO gains were consistent, multi-center prospective validation is needed. Finally, association does not imply causation: improved prognosis/treatment stratification does not, on its own, establish that modifying the decoded pathway will change outcomes. In addition, extending the method to data privacy learning frameworks such as federated learning is also an interesting research direction41.
Methods
We formalize non-invasive decoding of the ER-stressed cancer-associated fibroblast (ERS–CAF) immunoregulatory axis as a supervised, multi-target regression from routine images to transcriptomic gold standards. For each patient i ∈ {1, …, N}, we observe pre-operative MRI (Ri), H&E WSI (Wi), bulk RNA-seq (xi), and a site/scanner domain label di ∈ {1, …, D}. The system contains: (i) a CLIP-guided pathomics branch that converts WSI tiles into task-specific slide embeddings using a curated prompt bank capturing ERS–CAF biology; (ii) a radiomics branch that summarizes MRI with harmonized quantitative features; (iii) a mid-level gated fusion that integrates branches and predicts three continuous targets: ERS–CAF abundance, ERS–CAF-immune ligand-receptor (LR) interaction strength, and TME heterogeneity. The training proceeds in three stages: Stage I calibrates the CLIP-guided WSI branch to molecular scores via a monotone similarity-label alignment; Stage II fits the multimodal regressor with a heteroscedastic multi-task loss; Stage III performs stable, non-adversarial domain harmonization by second-order (CORAL) alignment. Our framework is shown in Fig. 8.
Stage-wise multimodal pipeline for decoding ERS–CAF-centric TME surrogates from routine images.
Notation and data schema
For patient i, MRI volumes \({R}_{i}\in {{\mathbb{R}}}^{H\times W\times Z\times C}\) with C = 3 (T1/T2/contrast-enhanced), WSI Wi, RNA-seq \({x}_{i}\in {{\mathbb{R}}}^{G}\), clinical covariates ci, survival (Ti, Δi), treatment vector ti, and domain di ∈ {1, …, D}. The three continuous supervision targets form \({S}_{i}^{\star }={[{S}_{{\rm{ERS}}-{\rm{CAF}}}(i),{S}_{{\rm{LR}}}(i),H(i)]}^{\top }\in {{\mathbb{R}}}^{3}\); predictions are \({\widehat{S}}_{i}\in {{\mathbb{R}}}^{3}\).
For each target j ∈ {ERS − CAF, LR, H}, we define a prompt set \({{\mathcal{Q}}}^{(j)}={\{{q}_{m}^{(j)}\}}_{m=1}^{{M}_{j}}\) and encode it using the CLIP text encoder fT into unit-norm vectors
We also define a target-specific prompt-mixture text embedding
and the concatenated text context used in fusion gating,
We use \({{\bf{t}}}_{m}^{(j)}\), \({\bar{{\bf{t}}}}^{(j)}\), and \(\bar{{\bf{t}}}\) consistently throughout; we avoid overloaded alternatives (e.g., f(j)) to prevent ambiguity.
Bulk-transcriptomic reference scores (surrogate supervision targets)
ERS–CAF abundanceL
With curated gene set GERS−CAF ⊆ {1, …, G} and per-sample gene ranking πi, define the ssGSEA running sum
where ranki(g) is the position of g in πi (descending); p ∈ [0, 2]. Scores are z-scored cohort-wise.
ERS–CAF-immune LR interaction
Let \({\mathcal{C}}\) denote immune cell types and \({\mathcal{L}}=\{(l,r,c)\}\) the curated triplets (ERS–CAF ligand l, immune receptor r on-cell c). With z-scored expressions \({\widetilde{x}}_{i,\cdot }\) and deconvolved fractions pi,c,
where ωlrc ≥ 0 are evidence-based (or uniform) weights.
Microenvironmental heterogeneity From CIBERSORTx proportions \({\{{p}_{i,k}\}}_{k=1}^{K}\) (∑k pi, k = 1),
Noise floor of computed supervision targets
The supervision targets \({S}_{i}^{(j)}\in \{{S}_{{\rm{ERS}}-{\rm{CAF}}}(i),{S}_{{\rm{LR}}}(i),H(i)\}\) are computed from bulk RNA-seq using (1) gene-set scoring (ssGSEA), (2) digital cytometry (CIBERSORTx), and (3) an aggregated LR scoring rule. These procedures are validated and widely used, but they are nevertheless approximations to latent biology and introduce label noise. Therefore, our model learns an image-based surrogate of a bulk-derived surrogate, and performance should be interpreted relative to this noise.
Let \({S}_{i}^{* (j)}\) denote the latent (unobserved) sample-level biological state for target j and assume
where \({\varepsilon }_{i}^{(j)}\) captures scoring/deconvolution error and finite-sample noise in bulk measurements. Define the reliability (signal fraction) of the computed target as
Then, for any predictor \({\widehat{S}}^{(j)}\) (including our image-derived predictor), the observable correlation is upper bounded by the noise ceiling
i.e., even a perfect predictor of S*(j) cannot exceed \(\sqrt{{\rho }_{j}}\) correlation with the noisy computed label S(j).
We estimate ρj (or lower bounds thereof) via pipeline perturbations that produce two independently computed proxies S(j) and \({\widetilde{S}}^{(j)}\) from the same RNA-seq:
-
ERS–CAF score stability: ssGSEA vs. GSVA (or other rank-based scoring), yielding \({\rm{Corr}}({S}_{{\rm{ERS}}-{\rm{CAF}}},{\widetilde{S}}_{{\rm{ERS}}-{\rm{CAF}}})\).
-
Heterogeneity stability: compute H from alternative deconvolution tools (e.g., EPIC/quanTIseq/xCell) and compare to CIBERSORTx-derived H.
-
LR score stability: compare the simple weighted product-sum rule to an alternative bulk-mode LR scoring (e.g., CellChat bulk formulation), reporting \({\rm{Corr}}({S}_{{\rm{LR}}},{\widetilde{S}}_{{\rm{LR}}})\).
These inter-pipeline concordances provide conservative empirical lower bounds on ρj and contextualize attainable prediction accuracy.
WSI pathomics with CLIP-guided MIL
WSI captures micro-ecology (CAF morphology, collagen deposition, immune exclusion). A CLIP prior and mechanism-driven prompts steer attention toward tiles that instantiate ERS–CAF biology. Standard pathology MIL learns attention weights purely from slide-level labels; in contrast, our tile weighting \({\alpha }_{it}^{(j)}\) is explicitly grounded in a target-specific biological text prior through prompt-similarity scoring (Eqns. (12–14)), and Stage I enforces a monotone similarity-label alignment before downstream regression. This design makes the inductive bias inspectable (via prompts) and falsifiable (via synonym/OOD/antonym perturbations in Fig. 4). Moreover, unlike path-transcriptomics approaches that predict large gene expression vectors, we predict three mechanism-defined surrogates (ERS–CAF activity, LR crosstalk, heterogeneity), which are more stable under limited N and directly aligned with our biological hypothesis.
After stain normalization and tissue detection, tile Wi at 20× into \({\{{W}_{it}\}}_{t=1}^{{T}_{i}}\). CLIP image encoder fI yields unit-norm tile embeddings
For each target j ∈ {ERS − CAF, LR, H}, a prompt set \({{\mathcal{Q}}}^{(j)}={\{{q}_{m}^{(j)}\}}_{m=1}^{{M}_{j}}\) is encoded by CLIP text encoder fT into unit vectors
Prompt-similarity attention over tiles uses a log-sum-exp temperature τ > 0:
The slide-level pathomics representation concatenates task-specific embeddings:
MRI radiomics branch
MRI encodes macro-phenotypes (shape, intensity, texture) correlated with stromal content and edema, complementing WSI micro-ecology.
After resampling to isotropic voxels and sequence-wise z-normalization, PyRadiomics produces \({\phi }_{{\rm{rad}}}({R}_{i})\in {{\mathbb{R}}}^{{P}_{r}}\); stable features (ICC threshold) define \({{\bf{r}}}_{i}={[{\phi }_{{\rm{rad}}}({R}_{i})]}_{{\mathcal{I}}}\). Site/scanner effects are harmonized via ComBat:
with Erad an MLP (layer norm, dropout).
Gated mid-level fusion and multi-head regression
Radiomics and pathomics carry complementary, scale-separated signals; prompts provide a mechanism-aware context vector that can modulate fusion. Encode pathomics by Ewsi:
A gate controls cross-modal contribution:
where σ is element-wise sigmoid and ⊙ the Hadamard product. A shared trunk f with three linear readouts yields
Training curriculum and objectives
Directly optimizing many objectives is fragile42,43,44. We therefore adopt a three-stage curriculum, ensuring stable convergence while preserving the intended inductive biases.
Stage I: CLIP similarity-label alignment (WSI branch)
For each target j, define the WSI similarity
where \({\rm{minmax}}\) scales per-cohort to [0, 1], \({\mathcal{T}}\) denotes the training split of the current outer fold (to avoid leakage) and ϵ = 10−8. A monotone affine-sigmoid calibrator maps similarity to the label scale:
The WSI alignment loss is
where σ( ⋅ ) logistic function; aj, bj are learned scalars with aj≥0 to enforce monotonicity. During Stage I, (fI, fT) are kept frozen; Ewsi, prompt-mixture weights \({\gamma }_{m}^{(j)}\), and (aj, bj) are optimized.
Stage II: Multimodal regression (fusion model)
We fit the fused model to the continuous molecular targets with heteroscedastic weighting:
where σj > 0 are learned noise scales and \({S}_{i}^{(j)}\) denotes the jth entry of \({S}_{i}^{\star }\). Parameters updated include Erad, Ewsi, fusion gate (Wg, bg), trunk f, and readouts (wj, bj); the Stage I prompt mixtures \({\gamma }_{m}^{(j)}\) are fine-tuned.
Stage III: Site/scanner harmonization (CORAL)
To mitigate domain shift without adversarial instability, we align the second-order statistics of the fused features. Let μd and Σd denote the mean and covariance of zi within domain d, and μ, Σ the global mean and covariance. The CORAL loss is
where ∥ ⋅ ∥F is Frobenius norm. The Stage III objective is
with weight decay λreg ≥ 0 and λcoral ≥ 0 chosen on validation folds.
We employ dropout in encoders and trunk, and modality dropout during stage II/III (with probability q, one branch’s latent is zeroed) to ensure single-modality robustness.
MRI-only Knowledge Distillation
For fully non-invasive deployment when pathology is unavailable, we distill the trained teacher \({F}_{{\Theta }^{\star }}\) into an MRI-only student \({F}_{{\Theta }_{S}}\):
where \({\widehat{s}}_{i,T}^{(j)}={F}_{{\Theta }^{\star }}^{(j)}({R}_{i},{W}_{i})\) and \({\widehat{s}}_{i,S}^{(j)}={F}_{{\Theta }_{S}}^{(j)}({R}_{i})\). The student reuses the MRI radiomics encoder and the fusion trunk (restricted to the MRI branch).
Validation and Clinical Modeling
We conduct fivefold patient-level cross-validation stratified by domain di with nested hyperparameter selection; leakage is prevented at the patient-level. Predictive fidelity is summarized by Pearson correlation r, RMSE, MAE, and calibration slope/intercept between \({\widehat{S}}_{i}\) and \({S}_{i}^{\star }\). Clinical relevance is assessed by Cox models with covariate adjustment:
using the partial log-likelihood
where \({\mathcal{R}}({T}_{i})\) is the risk set. We report hazard ratios (HRs) with 95% CIs, time-dependent AUC, and C-index. Treatment relevance is tested via logistic or interaction Cox models with treatment vector ti.
Given the limited cohort size (N = 126), we explicitly constrain model capacity and enforce strict evaluation hygiene. CLIP image/text encoders are kept frozen; the trainable components are limited to lightweight prompt-mixture weights, shallow encoders for radiomics/pathomics, a mid-level gating network, and three linear heads. All preprocessing/statistics are fit within each training fold only (e.g., min–max scaling for Stage I targets; ComBat parameters; feature standardization), and performance is reported on held-out patients only. We further apply dropout and weight decay throughout, and use modality dropout during Stage II/III to prevent single-modality shortcut learning and to improve robustness under missing modalities.
Experimental setup
We assemble a consecutive, single-disease cohort of 126 patients with matched pre-operative MRI (T1, T2, contrast-enhanced), diagnostic H&E WSIs, bulk RNA-seq, and clinical follow-up. Exclusion criteria are: (i) imaging outside protocol; (ii) failed ROI/tiling quality control; (iii) RNA-seq library size or mapping rate below thresholds; and (iv) missing outcomes. Primary endpoints are overall survival (OS) and progression-free survival (PFS). Molecular supervision yields three continuous targets per patient i: \({S}_{i}^{\star }={[{S}_{{\rm{ERS}}-{\rm{CAF}}}(i),{S}_{{\rm{LR}}}(i),H(i)]}^{\top }\) (cf. Section 9).
For generalization, we evaluate on TCGA stroma-rich cancers (PAAD, STAD, COADREAD) with WSIs, RNA-seq, and clinical data. The multimodal model is frozen and applied zero-shot to generate \({\widehat{S}}_{i}\), then compared to per-cancer molecular gold standards. When imaging modality differs (e.g., CT), we restrict radiomic families to modality-agnostic statistics.
Preprocessing and Quality Control Volumes are resampled to 1 mm isotropic, bias-field corrected, and z-standardized per sequence. Tumor ROIs are delineated and adjudicated. PyRadiomics extracts first-order, shape, and texture features (GLCM/GLRLM/GLSZM/NGTDM) plus LoG/wavelet; site/scanner effects are harmonized by ComBat; features with ICC < 0.75 are removed, yielding \({{\bf{r}}}_{i}\in {{\mathbb{R}}}^{{P}_{r}}\).
It should be noted that the end-to-end 3D CNN/Transformer encoders are attractive45,46. However, in our setting (N = 126 chordoma, multi-site scanners, three MRI sequences), fully learned 3D representation learning is high-variance and prone to overfitting without substantial pretraining and careful harmonization. We therefore use radiomics as a data-efficient and reproducible summary of macro-scale tumor phenotypes (shape/texture/heterogeneity), then learn a non-linear encoder Erad( ⋅ ) jointly with the multimodal trunk and gate.
WSIs undergo Macenko stain normalization, tissue masking (Otsu+morphology), tiling at ×20 into 224 × 224 px patches (stride 224), and tile QC (tissue coverage ≥50%, blur/pen-mark removal). Retained tiles form \({\{{W}_{it}\}}_{t=1}^{{T}_{i}}\).
SERS−CAF is ssGSEA on a curated ERS–CAF gene set (cohort z-score); SLR aggregates ERS–CAF ligand-immune receptor pairs weighted by immune fractions pi,c from CIBERSORTx; H(i) is Shannon diversity over deconvolved cell fractions (Section 9).
Ethics approval and consent to participate
This study was approved by the Institutional Ethics Committee of The First Affiliated Hospital, University of South China, Hunan, P.R. China (Ethics Approval No. 2025LL0206001). Written informed consent was obtained from all enrolled patients in accordance with the Declaration of Helsinki.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to the current Administrative Regulations on Human Genetic Resources (HGR) from China’s Ministry of Science and Technology, but are available from the corresponding author on reasonable request.
Code availability
Custom code and scripts were developed for data preprocessing, feature extraction, model training, evaluation, and statistical analysis in this study. For the purpose of peer review, an anonymized version of the code and related resources is available at: https://anonymous.4open.science/r/Anonymous_code-ERS-CAF/README.md. The full source code will be made publicly available upon acceptance of the manuscript. All experiments were implemented in Python (≥3.7) using PyTorch (≥2.1.0), torchvision (≥0.16.0), numpy (≥1.24), pandas (≥2.0), scikit-learn (≥1.3), pyyaml (≥6.0), tqdm (≥4.66), matplotlib (≥3.7), transformers (≥4.41), and open_clip_torch (≥2.24.0). Unless otherwise specified, default parameters provided by the respective libraries were used, and key model architectures, hyperparameters, and task-specific variables are described in the Methods section and Supplementary Information.
References
Murad, T. M. & Murthy, M. S. N. Ultrastructure of a chordoma. Cancer 25, 1204–1215 (1970).
Niu, H., Zheng, B., Zou, M. & Zheng, B. Complex immune microenvironment of chordoma: a road map for future treatment. J. Immunother. Cancer 12, e009313 (2024).
Zhang, T. et al. Hypoxic upregulation of ier2 increases paracrine GMFG signaling of endoplasmic reticulum stress-caf to promote chordoma progression via targeting ITGB1. Adv. Sci. 11, e2405421 (2024).
Zhang, T. et al. Integrating single-cell and spatial transcriptomics reveals endoplasmic reticulum stress-related caf subpopulations associated with chordoma progression. Neuro-Oncol. 26, 295–308 (2024).
Liang, S. et al. The immune microenvironment in chordoma: implications for future treatment. World Neurosurg. 204, 124589 (2025).
Zheng, B. & Guo, W. Multi-omics analysis unveils the role of inflammatory cancer-associated fibroblasts in chordoma progression. J. Pathol. 265, 69–83 (2025).
Luo, W. et al. Tumor immune microenvironment-based therapies in pancreatic ductal adenocarcinoma. J. Exp. Clin. Cancer Res. 43, 92 (2024).
Wang, H. et al. Characterization of endoplasmic reticulum stress unveils ZNF703 as a promising target for colorectal cancer immunotherapy. J. Transl. Med. 21, 713 (2023).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Vaidya, P. et al. Computationally integrating radiology and pathology image features for predicting treatment benefit and outcome in lung cancer. npj Precis. Oncol. 9, 161 (2025).
Kang, W. et al. Application of radiomics-based multiomics combinations in the tumor microenvironment and cancer prognosis. J. Transl. Med. 21, 566 (2023).
Chen, R. J. et al. Multimodal co-attention transformer for survival prediction in gigapixel whole slide images. In Proceedings of the IEEE/CVF international conference on computer vision, 4015–4025 (2021).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer cell 40, 865–878 (2022).
Xu, J. et al. The role of tumor immune microenvironment in chordoma: promising immunotherapy strategies. Front. Immunol. 14, 1257254 (2023).
Öhlund, D. et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 214, 579–596 (2017).
Elyada, E. et al. Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov. 9, 1102–1123 (2019).
Cords, L. et al. Cancer-associated fibroblast classification in single-cell datasets across 19 cancer types. Nat. Commun. 14, 4929 (2023).
Forsthuber, A. et al. Cancer-associated fibroblast subtypes modulate the tumor microenvironment and patient outcome. Nat. Commun. 15, 53908 (2024).
Liu, Y. et al. Conserved spatial subtypes and cellular neighborhoods of cancer-associated fibroblasts revealed by single-cell spatial multi-omics. Cancer Cell 43, 905–924 (2025).
Feig, C. et al. Targeting cxcl12 from fap-expressing carcinoma-associated fibroblasts synergizes with anti–PD–L1 immunotherapy in pancreatic cancer. Proc. Natl. Acad. Sci. 110, 20212–20217 (2013).
Biasci, D. et al. Cxcr4 inhibition in human pancreatic and colorectal cancers induces an integrated immune response. Proc. Natl. Acad. Sci. 117, 28960–28970 (2020).
Mao, X. et al. Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment. J. Hematol. Oncol. 14, 73 (2021).
Zhan, Y. et al. Single-cell transcriptomics reveals intratumor heterogeneity and the potential roles of cancer stem cells and mycAFs in colorectal cancer liver metastasis and recurrence. Cancer Lett. 612, 217452 (2025).
Biffi, G. et al. Il1-induced jak/stat signaling is antagonized by tgfβ to shape CAF heterogeneity in pancreatic ductal adenocarcinoma. Cancer Discov. 9, 282–301 (2019).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. Cellphonedb: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Jin, S. et al. Inference and analysis of cell–cell communication using cellchat. Nat. Commun. 12, 1088 (2021).
Browaeys, R., Saelens, W. & Saeys, Y. Nichenet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Chen, C. et al. Crosstalk between cancer-associated fibroblasts and regulated cell death in tumors: insights into apoptosis, autophagy, ferroptosis, and pyroptosis. Cell Death Discov. 10, 123 (2024).
Aerts, H. J. W. L. et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat. Commun. 5, 4006 (2014).
Schmauch, B. et al. A deep learning model to predict RNA-seq expression of tumoral genes from whole-slide images. Nat. Commun. 11, 3877 (2020).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Echle, A. et al. Deep learning for the prediction of microsatellite instability from histopathology images of colorectal cancer: a systematic developmental and validation study. Lancet Oncol. 22, 162–172 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. 102, 15545–15550 (2005).
Hänzelmann, S., Castelo, R. & Guinney, J. Gsva: gene set variation analysis for microarray and rna-seq data. BMC Bioinformatics 14, 7 (2013).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Cancer Genome Atlas Research Network Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, 185–203.e13 (2017).
Cancer Genome Atlas Research Network Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Cancer Genome Atlas Network Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Jensen, M. A., Ferretti, V., Grossman, R. L. & Staudt, L. M. The NCI genomic data commons as an engine for precision medicine. Blood 130, 453–459 (2017).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416.e11 (2018).
Xiao, C. et al. Confusion-resistant federated learning via diffusion-based data harmonization on non-iid data. Adv. Neural Inf. Process. Syst. 37, 137495–137520 (2024).
Yang, Z., Wang, H., Liu, Y. & Zhang, F. Cfdformer: medical image segmentation based on cross fusion dual attention network. Biomed. Signal Process. Control 101, 107208 (2025).
Jiang, Y. et al. Self-paced learning for images of antinuclear antibodies. IEEE Trans. Med. Imaging 44, (2025).
Yao, J., Li, C. & Xiao, C. Swift sampler: efficient learning of sampler by 10 parameters. Adv. Neural Inf. Process. Syst. 37, 59030–59053 (2024).
Zhang, F., Gu, Z. & Wang, H. Decoding with structured awareness: integrating directional, frequency-spatial, and structural attention for medical image segmentation. Preprint at https://arxiv.org/abs/2512.05494 (2025).
Wang, Y., Wang, H. & Zhang, F. A medical image segmentation model with auto-dynamic convolution and location attention mechanism. Comput. Methods Prog. Biomed. 261, 108593 (2025).
Acknowledgements
Department of Spine Surgery, The First Affiliated Hospital, Hengyang Medical School, University of South China, and Department of Orthopedics, Wuhan Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, contributed equally as first affiliation. We gratefully acknowledge Professor Yongbin Liu from the School of Computer Science at the University of South China, as well as Master’s students Kefan Wu and Yangji Chen, for their contributions to the AI modeling and validation aspects of this research. This work was supported by the National Natural Science Foundation of China (W2523095, 82003802 and 82473965 to T.L.Z., 82404690 to C.X. and 82002364 to M.X.Z.), China Scholarship Council (202106370071 to B.W.Z.), China Postdoctoral Science Foundation (2025M782004 to BWZ), Excellent & Innovative Talent Program, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology (082550320 to B.W.Z.), Hubei Province High-Level Postdoctoral Talent Introduction Program (2005HBBHJD018 to B.W.Z.), Natural Science Foundation of Hunan Province (2019JJ50542 and 2023JJ50156 to T.L.Z., 2023JJ40596 to C.X., 2023JJ40587 to W.H. and 2021JJ40509 to M.X.Z.), Hunan Provincial Natural Science Foundation for Excellent Youth Scholars (2023JJ20035 to M.X.Z.), the Science and Technology Innovation Program of Hunan Province (2023RC3172 to M.X.Z.), Clinical Research 4310 Program of the First Affiliated Hospital of the University of South China (20224310NHYCG04 to T.L.Z. and 20224310NHYCG03 to H.Z.), Project for Clinical Research of Hunan Provincial Health Commission (20201978 to T.L.Z., 20201962 to C.X., and 20201956 to M.X.Z.), Research Foundation of Education Bureau of Hunan Province (22B0441 to W.H.), Medical-Engineering Interdisciplinary Research Program of the First Affiliated Hospital, University of South China (IRP-M&E-2025-06 to M.X.Z.), Henan Province Key Science and Technology Research Project (252102311060 to H.Q.N.) and Xiaohe Technology Talent Project of Hengyang city (C.X.).
Author information
Authors and Affiliations
Contributions
B.W.Z., C.X., and M.T. are co-first authors and contributed equally to this work. T.L.Z., H.Z., and M.X.Z. are co-corresponding authors and contributed equally to this work. B.W.Z., C.X., T.L.Z., and M.X.Z. contributed to the conception and design of the study. B.W.Z., C.X., M.T., W.H., B.Y.Z., H.Q.N., J.L., T.L.Z., H.Z., and M.X.Z. did the data analysis and interpretation. B.W.Z., C.X., M.T., W.H., T.L.Z., H.Z. and M.X.Z. contributed to drafting and revision of the manuscript. All authors were involved in writing the paper and had final approval of the submitted and published versions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, BW., Xia, C., Tang, M. et al. Decoding the ERS–CAF immunoregulatory axis via multimodal AI and its pan-cancer prognostic and therapeutic predictive value. npj Digit. Med. 9, 199 (2026). https://doi.org/10.1038/s41746-026-02388-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02388-w