Abstract
Accurate medical image segmentation continues to pose significant challenges, as existing methods often struggle to concurrently achieve efficient global context modeling, precise boundary delineation, and robust generalization. To address these issues, a novel framework named Contextual and Frequency-Guided Mamba Network (CFG-MambaNet) is presented. Specifically, a variable-scale state space block based on Mamba is employed so that long-range dependencies can be captured with linear complexity, efficiently addressing the inefficiency of Transformer-based models in high-resolution medical imaging. Moreover, a frequency-guided representation module is incorporated to explicitly separate global low-frequency structures from high-frequency boundary details, which significantly alleviates the difficulty of segmenting lesions with blurred contours or weak textures. Furthermore, an adaptive context aggregation mechanism is introduced to integrate heterogeneous semantic cues and to consistently highlight clinically critical regions, substantially improving robustness across diverse anatomical scales and morphologies. To further stabilize training and improve boundary adherence, a composite loss combined with deep supervision is employed. Extensive experiments were conducted on four publicly available datasets, including ACDC, Kvasir-SEG, ISIC, and SEED, covering cardiac MRI, endoscopy, dermoscopy, and pathology images.
Similar content being viewed by others
Introduction
Medical image segmentation is a pivotal research direction in the field of medical image computing, aiming to structurally analyze complex and variable medical imaging data1. It precisely extracts and annotates specific anatomical regions (such as organs, tissues, or lesions) from original images, providing a reliable visual foundation for downstream clinical diagnosis, treatment planning, intraoperative navigation, and efficacy assessment2. In traditional medical image analysis, manual segmentation is both time-consuming and labor-intensive, and is limited by the subjective experience of the operator, resulting in significant variability and irreproducibility3,4.
In recent years, convolutional neural networks (CNNs) have become the mainstream technology in the field of medical image segmentation. Utilizing an end-to-end training approach, CNNs can automatically learn multi-scale features from complex medical images, enabling precise pixel-level segmentation5. The U-Net architecture, as a pioneering model in this field, employs a symmetric encoder-decoder structure and skip connections to effectively integrate high-level semantic information with low-level details, making it the foundational method for two-dimensional medical image segmentation6. Building upon this foundation, numerous variants such as U-Net++7 and U-Net 3+8 have been developed, which further boost segmentation performance through multi-level skip structures and richer context fusion. To accommodate the inherent characteristics of volumetric medical imaging modalities such as MRI and CT, researchers introduced 3D CNN and V-Net to enable the model to capture the spatial continuity of body data, immensely improving the segmentation accuracy of three-dimensional structures9,10,11. With the growing success of attention mechanisms in visual recognition tasks, their integration into medical image segmentation has emerged as a compelling trend. Attention U-Net incorporates attention gates to guide the network to automatically focus on target structures while suppressing irrelevant background information12. Furthermore, Gu et al. proposed the Comprehensive Attention Network (CA-Net), which integrates spatial, channel, and scale attention modules13. Nevertheless, due to the limited receptive field of convolutional neural networks, CNNs struggle to effectively capture global information, which plays a key role in fine-grained segmentation of medical images. In particular, modeling long-range dependencies between background and distant pixels is essential for accurately delineating tumor morphology and size, where subtle contextual cues often govern critical diagnostic boundaries.
Transformer is naturally equipped with a self-attention mechanism that can break through the limitations of CNNs global modeling, and their introduction into the field of image segmentation is catalyzing a profound shift in research paradigms14. TransUNet introduces the Transformer encoding module into U-Net, leveraging its global context modeling capabilities to compensate for the shortcomings of CNN in detail recovery, achieving significant improvements in the segmentation performance of multiple organs and cardiac structures15. Swin-Unet builds a pure Transformer U-shaped structure, using a hierarchical mechanism to take into account both global features and spatial restoration effects in encoding and decoding16. In addition, the hybrid CNN-Transformer architectures, which extract local texture features via convolutional modules and then leverage Transformer-based self-attention to model long-range semantic dependencies, have demonstrated remarkable efficacy in simultaneously preserving fine-grained anatomical boundaries and ensuring global structural consistency through the integration of both frameworks and complementary strengths17,18,19. Despite the promising performance of Transformer-based approaches in medical image segmentation, their clinical deployment remains constrained by substantial computational overhead, primarily due to the quadratic relationship between computational cost and input sequence length.
With the widespread adoption of deep neural networks in medical image segmentation, the demand for a balanced trade-off between long-range dependency modeling and computational efficiency has become increasingly prominent. Traditional CNNs excel at capturing local structural details, yet their inherently limited receptive fields hinder the acquisition of global contextual information. In contrast, Transformer-based models offer powerful long-range modeling capabilities, but their quadratic computational complexity with respect to input size poses a significant barrier to their application in high-resolution medical imaging tasks. To address this bottleneck, the recently proposed Mamba architecture is based on a linear state space model (SSM) and utilizes a selective state update mechanism and hardware-friendly parallel operations to significantly reduce computational and storage overhead while maintaining modeling capabilities20. Mamba-based segmentation models can be broadly categorized into three types of architectural design approaches. First, some approaches attempt to integrate Mamba with CNNs at the structural level to achieve complementary representation of local and global information. A representative example is U-Mamba, which employs convolutional modules to capture fine-grained local features while leveraging Mamba for long-range dependency modeling21. Furthermore, some studies have focused on constructing pure Mamba encoding-decoding structures that completely eliminate convolutional and self-attention operations. For example, Mamba-UNet22 and Swin-UMamba23 have constructed state-driven context abstraction paths and enhanced spatial restoration capabilities through deep residual and skip connection mechanisms, increasing their ability to distinguish low-contrast boundaries and weak texture areas. Moreover, researchers integrated the Mamba module into the attention mechanism or context fusion channel to enhance the model’s responsiveness to critical structural regions24,25,26. Mul-VMamba is a VMamba based multimodal semantic segmentation network with shared backbones, incorporating the Mamba Spatial consistency Selective Module for spatially consistent multimodal feature selection and the Mamba Cross Fusion Module with Cross S6 to achieve linear complexity cross modal fusion27. SegMamba is a Mamba based model for general 3D medical image segmentation that adopts a four stage convolution Mamba hybrid architecture, integrating the Tri-orientated Spatial Mamba block to model global dependencies across three orthogonal planes via multi direction scanning28. Despite these promising attempts, several critical challenges remain unresolved. A key limitation of current Mamba-based networks lies in their difficulty to maintain global structural consistency while accurately capturing boundary details, especially in scenarios involving low-contrast or morphologically diverse lesions. Another open issue is the lack of effective strategies for adaptive multi-scale context aggregation, which is crucial for representing anatomical structures of varying sizes. In addition, the robustness of existing designs is often undermined in high-noise clinical environments, where small lesions or irregular textures can be easily misclassified or overlooked. To address the challenge of balancing global modeling and boundary characterization in existing methods, this paper proposes an innovative medical image segmentation framework. A Mamba-based visual state space (VSS) module is introduced to efficiently capture long-range dependencies while preserving computational scalability. To further enhance structural fidelity, a frequency-guided representation (FGR) module explicitly disentangles global morphology from boundary details, improving robustness in low-contrast and weak-texture regions. In addition, a multi-scale adaptive context aggregation (MSACA) module integrates heterogeneous contextual cues and emphasizes critical lesion areas, enabling the model to adapt effectively to diverse anatomical scales and morphological variations. Our main contributions can be summarized as follows:
-
A novel medical image segmentation framework is proposed, where a visual state space (VSS) module based on Mamba is introduced to achieve long-range dependency modeling while maintaining linear complexity, effectively balancing global context modeling capabilities and computational efficiency.
-
A frequency-guided representation (FGR) module is designed to significantly improve the model’s adaptability to blurred contours and complex textures by separating global structure information from boundary detail information in the frequency domain.
-
The multi-scale adaptive context aggregation (MSACA) module is constructed to integrate cross-scale semantic information and highlight critical regional features, which dramatically enhanced the model’s ability to handle lesions of different sizes and complex morphologies.
-
Extensive experiments on ACDC, Kvasir-SEG, ISIC, and SEED datasets demonstrate that the proposed framework consistently surpasses state-of-the-art segmentation models. The results highlight substantial gains in segmentation accuracy and boundary delineation, together with enhanced robustness and generalization across diverse imaging modalities.
Results
On four publicly available medical imaging datasets, the proposed method demonstrated advanced performance and robust generalization capabilities. On the ACDC dataset for cardiac MRI, the model achieved a Dice score of 92.74% and an IoU of 86.22%, reducing ASD to 0.72, and accurately delineated ventricular boundaries and myocardial thickness, providing reliable support for clinical functional parameter measurements. On the Kvasir-SEG dataset, the method achieved 92.84% Dice and 88.10% IoU under complex and variable polyp boundaries, significantly reducing boundary offset issues and demonstrating its potential in early screening for colorectal cancer. On the ISIC dataset, the model achieved 94.46% Dice and 97.71% recall, which effectively increased the recognition rate of blurred and irregular skin lesions. In the SEED pathology slide task, the model achieved 86.52% Dice and 91.69% ACC, with optimal boundary accuracy, validating its robustness in complex tissue structures. Ablation experiments further demonstrated that the designed VSS, FGR, and MSACA modules play a crucial role in long-range dependency modeling, boundary detail preservation, and multi-scale feature fusion, while the composite loss function and deep supervision mechanism enhance overall stability and boundary fitting performance.
Datasets
The Automated Cardiac Diagnosis Challenge (ACDC) dataset29 is based on real clinical examinations of cardiac Cine-MRI scans conducted at the University Hospital of Dijon. It has been anonymized and ethically reviewed, and serves as a standard data source for segmentation and diagnostic evaluation in cardiac imaging research. The entire dataset includes 150 subjects, evenly distributed across five categories: Healthy (NOR), heart failure due to previous myocardial infarction (MINF), dilated cardiomyopathy (DCM), hypertrophic cardiomyopathy (HCM), and right ventricular abnormalities (RV).
Kvasir-SEG30 is a publicly available benchmark dataset for pixel-level segmentation of colorectal polyps, curated by the Norwegian Vestre Viken Health Trust and the Simula team from Kvasir v2. It comprises 1000 colonoscopy polyp images and their corresponding precise annotations.
The International Skin Imaging Collaboration (ISIC) Dermoscopy Challenge Dataset31 provides structured benchmark resources for three core tasks in dermoscopy imaging: lesion segmentation, dermoscopy feature detection, and melanoma classification. The training set contains 900 de-identified images, and the test set contains 379 homologous images, all of which are sourced from multi-center clinical settings and annotated by a panel of dermoscopy experts.
The SEED Pathology Dataset originates from the Jiangsu Provincial People’s Hospital and has been carefully constructed for pathological analysis and gastric cancer research. The dataset consists of 1770 strictly selected pathological section images covering normal tissue and cancerous tissue at different stages. The lesion areas in the images have been carefully annotated by professional pathologists, with cancerous areas clearly marked to ensure the clinical accuracy of the labels.
Experimental settings
All experiments were conducted using Python 3.10 and PyTorch 2.1.0 on an NVIDIA A6000 GPU with 48 GB of RAM. The AdamW optimizer was employed in conjunction with the Cosine Annealing Learning Rate scheduler. The minimum learning rate was set to 1 × 10−6, and the network was trained for 300 epochs. In the MSACA module, η1 is set to 0.4, η2 is 0.3, and η3 is 0.3. The loss function weights were set to λCE = 0.3 and λDice = 0.7 across all experiments. All datasets used an initial learning rate of 1 × 10−4, and the weight decay was uniformly set to 5 × 10−2. For the Kvasir dataset, the image and label sizes were adjusted to 256 × 256, and the batch size was set to 16. For the ISIC and SEED datasets, the image and label sizes were uniformly adjusted to 224 × 224, and the batch sizes were set to 16 and 8, respectively. For the ACDC dataset, 3D data were converted into 2D slices, and the 2D slices and their corresponding labels were resized to 256 × 256, with a batch size of 16. Comprehensive data augmentation strategies were applied during training, including rotation, scaling, mirroring, noise addition, blurring, brightness adjustment, contrast enhancement, gamma correction, and simulated low resolution, which greatly improved the model’s generalization ability.
Performance comparison
Results on ACDC dataset: the quantitative results on the ACDC dataset are shown in Table 1. We compared classic convolutional architecture models such as U-Net32, nnUNet33, AttUNet12, and Rolling-unet34, as well as representative Transformer-based architecture methods such as UCTransNet35, H2Former36, and MISSFormer37. We also compared Hetero-UNet38 and Swin-UMamba39 based on the Mamba method, as well as recently proposed improved networks such as FSCA-Net40, EMCAD41, and GH-UNet42. Traditional convolutional networks still exhibit clear limitations in cardiac MRI segmentation tasks. For example, U-Net yields a Dice score of 77.92% and an ASD of 1.71, indicating that it often produces discontinuous or distorted boundaries between the ventricles and myocardium. Even with enhancements such as attention mechanisms in AttUNet, which achieves 81.13% Dice, or edge-preserving strategies in Rolling-unet, which reaches 81.94% Dice, these models struggle to balance global consistency with fine structural preservation. The proposed method improves the Dice score to 92.74% while reducing the ASD to 0.72. This notable performance gain stems from the incorporation of the Mamba-based VSS Block, which captures long-range dependencies with linear computational complexity and ensures structural coherence even along anatomically complex and curved boundaries. H2Former and GH-UNet achieved Dice scores of 92.37% and 92.57%, respectively, and recall scores exceeding 90%, significantly improving overall morphology preservation compared to CNNs. However, they still tend to blur the boundary between the ventricle and the myocardium in short-axis slices, resulting in inaccurate boundary determination. Our method further improves recall to 92.43%, which is closely related to our designed FGR module. This module effectively avoids the underestimation of myocardial thickness by traditional methods by explicitly separating low-frequency overall structure from high-frequency boundary details in the frequency domain. MISSFormer yields a Dice score of only 73.04%, while Hetero-UNet, despite reaching 87.38%, suffers from boundary shrinkage in regions with low contrast, leading to underestimation of myocardial thickness. The MSACA module proposed in this paper demonstrates clear advantages in addressing this issue. By integrating parallel multi-scale feature extraction with a sparse attention mechanism, the module effectively accommodates cardiac structures of varying sizes and shapes. As a result, the model achieves the IoU of 86.22%, outperforming leading methods such as H2Former with 84.68% and GH-UNet with 85.58%.
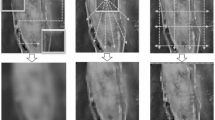
Figure 1 shows visual segmentation results of different models on representative ACDC cases. Traditional convolution-based networks, such as U-Net and its variants (e.g., AttUNet, Rolling-unet), tend to produce discontinuous boundaries or morphological deviations in segmenting the left ventricular blood pool and myocardium. This issue is particularly evident in the long-axis slice (b), where broken or irregular blood pool edges lead to abnormal fluctuations in myocardial thickness, which in turn may introduce significant errors in clinically important metrics such as left ventricular volume and ejection fraction. Transformer-based models such as UCTransNet and H2Former exhibit better global structural consistency compared to CNNs. However, in short-axis slices (c and d), the boundaries between the myocardium and right ventricle remain indistinct, often showing adhesion artifacts that hinder accurate measurement of ventricular wall thickness and right ventricular functional parameters. While MISSFormer and Hetero-UNet produce smoother overall contours, they suffer from inward shrinkage or blurring along myocardial boundaries, resulting in underestimated myocardial thickness, which is detrimental for quantitative analysis in conditions such as hypertrophic cardiomyopathy. The proposed method demonstrates superior boundary adherence and structural consistency across varying slice orientations and individual anatomical differences. Both the blood pool and myocardial regions closely align with the ground truth (GT) annotations, particularly in regions with highly curved myocardial edges, where the model accurately captures irregular shapes without over-smoothing or boundary overextension. In the long-axis view (b), our approach clearly distinguishes the myocardium from the blood pool, ensuring accurate measurements of ventricular diameter and myocardial thickness.
Each row a–d shows different cases, where the ground truth (GT) is compared with U-Net, AttUNet, Rolling-unet, GH-UNet, UCTransNet, nnUNet, H2Former, MISSFormer, EMCAD, FSCA-Net, and the proposed method. The proposed network produces results closer to the ground truth with more accurate lesion boundaries and fewer mis-segmented regions.
Results on Kvasir-SEG dataset: the experimental results on the Kvasir-SEG dataset are shown in Table 2, which demonstrates the performance variations among different methods in colorectal polyp segmentation. Most convolutional-based models perform weakly, with U-Net and AttUNet achieving Dice scores of 80.46% and 83.92%, respectively, and the ASD remaining above 7.27, indicating significant shortcomings in handling complex polyp boundaries. In contrast, some models incorporating fusion attention or multi-scale mechanisms such as Hetero-UNet achieves some improvement in accuracy, but the IoU metric remains around 82%, which reflects that the utilization of multi-scale semantic information is still insufficient. The proposed method achieves 92.84% Dice and 88.10% IoU under the same conditions, while significantly reducing ASD to 3.56, indicating that it effectively controls boundary errors while maintaining overall overlap. The Dice scores for GH-UNet and Swin-UMamba both exceed 91%, and ASD is also at a relatively low level, demonstrating the advantage of global modeling capabilities in preserving the overall contour of polyps. However, the Dice score for H2Former is only 72.56%, and UCTransNet is only 78.61%, which suggests that relying solely on global attention is insufficient to address the high diversity in scale and morphology of colorectal polyps.
As illustrated in the Fig. 2, we present the visual segmentation results of different models on the gastrointestinal endoscopy polyp segmentation task. Traditional methods such as U-Net and its improved variants can roughly delineate the contours of polyps with regular morphology. However, they tend to exhibit boundary shifts or discontinuities, resulting in segmentation regions that do not fully align with the true lesion, which may adversely affect clinical assessments of polyp size. Transformer-based approaches, including UCTransNet and H2Former, demonstrate better preservation of overall shape. Nevertheless, their performance degrades in the presence of specular reflections, vascular textures, or complex backgrounds, where they are prone to noise-induced false positives or blurred boundaries. MISSFormer and Hetero-UNet, on the other hand, often produce over-fragmented or irregular segmentations. In particular, their predictions near mucosal folds frequently misclassify normal tissues as lesion regions, which may mislead clinical interpretation. In contrast, the proposed method exhibits superior robustness and accuracy across varying levels of visual complexity. For polyps with smooth and well-defined boundaries, the segmentation outputs closely match the ground truth, accurately capturing both the size and shape of the lesion. In challenging cases involving strong reflections and complex backgrounds, the proposed model effectively suppresses noise, avoids over- or under-segmentation, and preserves the complete morphology of the lesion. This capability not only improves the model’s adaptability to diverse clinical scenarios but also provides a reliable foundation for downstream tasks such as adenoma detection rate (ADR) evaluation, polyp grading, and malignancy risk prediction.
Each row a–d shows representative polyp cases predicted by U-Net, AttUNet, Rolling-unet, GH-UNet, UCTransNet, nnUNet, H2Former, MISSFormer, EMCAD, FSCA-Net, and the proposed method. The green contours denote the ground truth annotations, while the blue contours indicate the predicted boundaries. The proposed method produces results closer to the ground truth, yielding smoother and more accurate segmentation in challenging cases.
Results on ISIC dataset: Table 3 shows the results on the ISIC dataset. U-Net achieved a Dice score of 91.56, indicating relatively stable overall segmentation. However, its large ASD value indicates insufficiently refined boundaries. AttUNet and MISSFormer, with their attention mechanism, improved Dice scores to 92.20 and 92.52, respectively. Despite this improvement in overall accuracy, they were still insufficient for skin lesions with blurred edges and complex morphology. nnUNet and EMCAD performed relatively consistently, with Dice scores exceeding 93%. EMCAD achieved a score of 3.30 on ASD, the best among all compared methods. It’s worth noting that EMCAD’s Recall score of 96.41 is still slightly lower than the 97.71 achieved by our method. This advantage stems from MSACA, whose parallel multi-scale branches capture large-scale skin lesions while maintaining sensitivity to small or poorly defined lesions, resulting in a leading Recall score. GH-UNet and Swin-UMamba achieved the closest overall performance to our method, with Dice scores of 93.74% and 93.99% respectively, and IoU scores exceeding 89%, demonstrating their strong global modeling capabilities. Nevertheless, these methods still exhibit slight deviations in boundary processing, with ASD remaining above 3.55. The visualization results on the ISIC dataset are presented in Fig. 3, which provides an intuitive comparison of the performance differences among various models in the skin lesion segmentation task. Traditional convolutional networks such as U-Net, AttUNet, and Rolling-unet can portray the general outline of lesions, but often appear offset or jaggedly discontinuous at boundary details, especially when lesion edges are blurred or pigmentation is diffuse, leading to over-segmentation or under-segmentation and failing to accurately depict the true morphology of lesions. Transformer-based methods such as GH-UNet, UCTransNet, and H2Former perform well in maintaining the overall contour, but in areas with complex textures or uneven pigment transitions, the predicted results are often overly smooth, lacking precise depiction of subtle boundary structures. EMCAD and FSCA-Net have improved in terms of boundary fitting, but there is still a certain degree of false negatives in the detection of small lesions. In contrast, the proposed method demonstrates higher stability and accuracy in segmentation across different types of lesions. Whether for small lesions with regular borders or large lesions with irregular shapes, the predicted results highly align with the actual annotations, accurately capturing the true extent and details of the lesions while avoiding common issues such as edge blurring and excessive smoothing.
Each row a–d shows representative skin lesion cases predicted by U-Net, AttUNet, Rolling-unet, GH-UNet, UCTransNet, nnUNet, H2Former, MISSFormer, EMCAD, FSCA-Net, and the proposed method. The green contours denote the ground truth annotations, while the blue contours indicate the predicted boundaries. The proposed network generates predictions closer to the ground truth, achieving more accurate lesion localization and smoother boundary delineation.
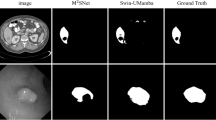
Results on SEED dataset: as shown in Table 4 segmentation results on the SEED dataset reveal significant performance differences across competing methods in terms of Dice, Intersection over Union (IoU), specificity (SP), and accuracy (ACC). U-Net achieves a Dice score of only 75.67% and an average surface distance (ASD) of 12.98, indicating considerable boundary deviations when processing pathological slides. AttUNet and UCTransNet also show limited performance, with Dice scores around 79% and IoU values below 71%, reflecting insufficient structural consistency in complex lesion regions. Although MISSFormer and Rolling-unet improve global shape consistency to some extent, their ASD values remain above 11, suggesting unstable boundary delineation. nnUNet yields relatively stable performance, achieving a Dice score of 82.98% and an accuracy of 89.00%. EMCAD attains a better ASD of 10.69 and reaches a specificity of 91.60%, indicating improvements in boundary convergence and classification reliability. However, none of these methods consistently lead across all evaluation metrics. The model proposed in this paper achieves the best overall performance across multiple metrics. It improves the Dice score to 86.52%, raises IoU to 79.34%, and reduces ASD to 8.69. Additionally, specificity and accuracy reach 92.90% and 91.69%, respectively. These results demonstrate that the proposed approach achieves a well-balanced trade-off among region overlap, boundary precision, and classification reliability, making it particularly suitable for robust segmentation in pathological image analysis.
The visualization results of the pathological slice dataset are shown in Fig. 4, where the green contours denote the ground truth and the blue contours represent the segmentation results of different models. Overall, traditional convolutional networks such as U-Net can roughly delineate the lesion structures, but in regions with complex glandular boundaries or dense cellular distribution, the predicted contours show noticeable deviations, with discontinuities or excessive smoothing. AttUNet and Rolling-unet enhance edge representation to some extent, yet in noisy or irregular tissue regions, the blue contours still deviate significantly from the ground truth. Transformer-based methods such as GH-UNet and UCTransNet demonstrate advantages in preserving the overall tissue morphology, but often suffer from contour over-expansion at the gland-stroma interface, leading to blue contours exceeding the green annotations. H2Former and MISSFormer tend to produce fragmented segmentations in complex slices, where the blue contours appear jagged or incomplete, particularly evident in challenging regions such as those shown in (b) and (d). Hetero-UNet and EMCAD provide relatively smoother results in some cases, but still show inaccuracies in boundary detail preservation. Conversely, the proposed method exhibits superior stability and accuracy across various pathological slices. Both in regions with well-defined glandular boundaries and in tissues with densely packed or highly complex structures, the blue contours align closely with the green annotations. Notably, in cases such as (a) and (c), our method achieves precise boundary fitting, avoiding discontinuities, ensuring both the completeness of the lesion region and the accuracy of boundary delineation.
Each row a–d shows representative tissue samples predicted by U-Net, AttUNet, Rolling-unet, GH-UNet, UCTransNet, nnUNet, H2Former, MISSFormer, EMCAD, FSCA-Net, and the proposed method. The green contours denote the ground truth annotations, while the blue contours indicate the predicted boundaries. The proposed network yields predictions that better align with the ground truth, achieving more precise delineation of pathological regions and clearer boundary localization compared with competing methods.
Ablation study
Ablation experiments on components: as shown in Table 5, ablation experiments on four medical imaging datasets explicitly demonstrate the effectiveness of the proposed VSS, FGR, and MSACA modules. In the ISIC dataset, when the three modules are used together, the Dice score reaches 94.46%, the IoU reaches 89.83%, and the ASD is reduced to 3.42, all of which outperform the performance after removing a single module. In the Kvasir-SEG dataset, the complete model achieves an optimal Dice of 92.84% and an ASD of 3.56. When FGR is removed, the IoU is slightly higher at 88.12%. This result revealed that global relationship modeling and other modules were complementary, but the joint configuration still performed better in terms of Dice score and boundary stability. In cardiac MRI segmentation, when all three modules are enabled simultaneously, the Dice and IoU are 92.74% and 86.22% respectively, reaching the highest level, and the ASD is 0.72, which is close to the optimal level. When MSACA is removed, the ASD is slightly lower at 0.71. This shows that the boundary effect of multi-scale attention in tasks with regular anatomical structures may not be as prominent as in complex lesions, but overall the full module still has higher accuracy. Furthermore, in pathological section experiments, the complete model achieved a Dice score of 86.52% and an ASD of 8.69, both of which outperformed other combinations. Although the IoU was slightly higher at 79.36% when FGR was removed, the Dice and ASD performance better met the requirements of pathological analysis for accuracy and boundary reliability.
Ablation experiments on the loss functions
A comparative analysis of the combination of cross-entropy and Dice in the loss function is shown in Table 6, which clearly illustrates the impact of different loss configurations on segmentation performance. In the ISIC dataset, when only cross-entropy was used, Dice and IoU were 92.40% and 86.90%, respectively. However, when only Dice was used, Dice improved to 93.10%, and IoU reached 89.90%, indicating that the Dice loss function has a significant advantage in terms of region overlap accuracy. When both are combined, Dice further improves to 94.46% and IoU to 89.83%. On Kvasir-SEG, training with only BCE loss yields a Dice of 90.20%, whereas training with only Dice loss yields a Dice of 91.10%. When combined, Dice reached 92.84%, approaching the optimal value, while IoU was 88.10%, slightly lower than the 88.12% achieved by Dice alone, but with a more balanced overall performance. In the ACDC dataset, the Dice score for cross-entropy alone is 90.52% and the IoU is 82.40%, while the Dice score for Dice loss alone increases to 91.05% and the IoU is 83.66%. When combined, Dice and IoU improve to 92.74% and 86.22%, respectively, achieving optimal performance, demonstrating that composite loss optimizes both global and local features simultaneously. In the SEED pathological slice dataset, when cross-entropy is used alone, Dice is 83.70% and IoU is 75.70%, while when Dice loss is used alone, Dice and IoU improve to 84.50% and 76.90%, respectively. When combined, Dice reached 86.52% and IoU reached 79.34%, both of which were the highest.
Ablation experiments on the loss weights: as shown in Table 7, a systematic ablation study was conducted by varying the weight distribution between cross-entropy loss (λCE) and Dice loss (λDice), and the results indicate that an appropriate balance between the two terms has a clear impact on segmentation accuracy and boundary quality (ASD). On the ACDC dataset, the configuration λCE = 0.3 and λDice = 0.7 achieves the best overall performance, yielding the highest Dice score of 92.74% and the lowest ASD of 0.72. When using equal weights (λCE = 0.5, λDice = 0.5), Dice slightly drops to 92.40% and ASD increases to 0.78, suggesting that a Dice-dominant weighting is more favorable for this dataset. On the Kvasir-SEG dataset, λCE = 0.3 and λDice = 0.7 provides the best Dice (92.84%), while the lowest ASD is obtained at λCE = 0.5 and λDice = 0.5 (ASD = 3.54), showing a mild trade-off between overlap and boundary distance. On the ISIC dataset, the highest Dice is achieved at λCE = 0.5 and λDice = 0.5 (Dice = 94.48%), whereas the minimum ASD occurs at λCE = 0.3 and λDice = 0.7 (ASD = 3.48). On the SEED dataset, λCE = 0.3 and λDice = 0.7 again delivers the best results, achieving the highest Dice of 86.52% and the lowest ASD of 8.69.
An ablation experiment on the deep supervision: an ablation experiment on the deep supervision mechanism was conducted on the ISIC dataset, as illustrated in Fig. 5a. The results demonstrate that this mechanism has a positive impact on the overall performance of the model. Numerically, enabling the deep supervision mechanism increased the Dice score from 94.00% to 94.46%, the IoU from 89.20% to 89.83%, and the Recall from 97.20% to 97.71%. It can be observed that deep supervision has brought stable gains in all core metrics, particularly in metrics such as Dice and IoU that measure region overlap, which indicates that the model has achieved better performance in overall lesion segmentation accuracy.
a Effect of deep supervision on Dice, IoU, and Recall. b Effect of Top-k sparse attention on Dice, IoU, and Recall. Ablation results on a deep supervision and b Top-k sparse attention, illustrating the improvements in Dice, IoU, and Recall, when the corresponding mechanisms are enabled. Both modules contribute to higher segmentation accuracy and boundary quality across evaluation metrics.
An ablation experiment on the Top-k: an ablation study was conducted to evaluate the effectiveness of introducing the Top-k sparse attention mechanism, as shown in Fig. 5b. The results revealed that enabling Top-k improved the Dice score from 93.95% to 94.46%, increased the IoU from 88.85% to 89.83%, and slightly improved Recall.
Ablation results on the weighting factors of MSACA: as shown in Table 8 and Fig. 6, we evaluate different settings of the MSACA weighting factors η1, η2, and η3 on four benchmark datasets. Balanced allocations consistently outperform single-branch dominated settings. On ACDC, setting η1 to 0.4, η2 to 0.3, and η3 to 0.3 yields a Dice of 92.74% and an ASD of 0.72, compared with a Dice of 90.36% and an ASD of 1.06 when η1 is set to 1.0. The highest ACDC IoU of 86.24% appears at η1 set to 0.3, η2 set to 0.4, and η3 set to 0.3, while the lowest ACDC ASD of 0.71 is achieved at η1 set to 0.5, η2 set to 0.3, and η3 set to 0.2. On Kvasir-SEG, the default setting increases Dice to 92.84% and reduces ASD to 3.56, while the minimum ASD of 3.53 is obtained at η1 set to 0.5, η2 set to 0.5, and η3 set to 0.0. On ISIC, the default setting achieves the best IoU of 89.83% with a Dice of 94.46%, whereas the highest Dice of 94.54% is observed at η1 set to 0.3, η2 set to 0.4, and η3 set to 0.3. On SEED, the default setting delivers the highest Dice of 86.52% and maintains a low ASD of 8.69, while the minimum ASD of 8.65 is achieved at η1 set to 0.3, η2 set to 0.3, and η3 set to 0.4. These results indicate that high-performing solutions concentrate in the near-balanced region, and we therefore adopt η1 set to 0.4, η2 set to 0.3, and η3 set to 0.3 as the default setting due to its consistently strong and stable performance across datasets.
a ACDC dataset with Dice scores under varying (η1, η2, η3) settings, showing the optimal performance at (0.4, 0.3, 0.3). b Kvasir-SEG dataset with Dice scores, similarly highlighting that balanced weight allocation yields the most stable and accurate segmentation results.
Discussion
The proposed segmentation framework has demonstrated consistently superior performance across four benchmark datasets, highlighting its strong generalization capability in diverse medical imaging scenarios. From a methodological perspective, the integration of the Mamba-based visual state space (VSS) block effectively alleviates the computational bottleneck of Transformer models while retaining the ability to capture long-range dependencies. The Frequency-Guided Representation (FGR) module explicitly decomposes global structural information and boundary details in the frequency domain, enhancing robustness in low-contrast and irregular lesion regions. Furthermore, the Multi-Scale Adaptive Context Aggregation (MSACA) module facilitates effective multi-scale feature integration and emphasizes critical regions, enabling the model to adapt to significant anatomical variability.
From the clinical perspective, the improvements observed in cardiac MRI indicate that the proposed method can provide reliable delineation of ventricular boundaries, supporting precise measurement of clinically relevant parameters such as ejection fraction and myocardial thickness. The results on endoscopy and dermoscopy datasets further suggest its potential for early tumor screening and skin cancer diagnosis, where accurate boundary detection is crucial for risk stratification. In pathology, the robustness of our method under noisy and structurally complex tissue environments underscores its value for fine-grained histological analysis.
Nevertheless, several limitations remain. Although the proposed model demonstrates encouraging cross dataset generalization, further validation on larger scale multi center clinical cohorts is necessary to confirm robustness and broaden clinical applicability. While the Mamba based design improves computational efficiency, processing ultra high resolution inputs such as whole slide pathology images remains challenging and may require dedicated tiling strategies and memory efficient inference. In addition, the interpretability and controllability of the framework in real world clinical workflows warrant deeper investigation, particularly under interactive use cases involving radiologists and pathologists.
In summary, this paper presents a segmentation framework that advances both methodological rigor and clinical applicability. Future work will explore multi-modal integration, self-supervised pretraining, and seamless incorporation into clinical pipelines, aiming to further bridge the gap between algorithmic innovation and practical deployment in intelligent medical imaging.
Methods
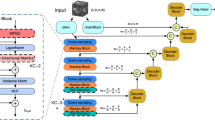
As illustrated in Fig. 7, the proposed framework adopts an encoder-decoder architecture enhanced with three core modules, namely the visual state space (VSS) block, the Frequency-Guided Representation (FGR) module, and the Multi-Scale Adaptive Context Aggregation (MSACA) module. In the encoder, successive patch embedding and VSS blocks progressively capture hierarchical representations, where long-range dependencies are efficiently modeled while maintaining scalability. At multiple stages, the FGR module projects features into the frequency domain and disentangles global low-frequency information from high-frequency boundary cues, which enhances the delineation of weakly contrasted or blurred structures. In parallel, the MSACA module aggregates contextual information from multiple receptive fields and applies dual sparse Top-k attention to emphasize critical lesion regions, improving adaptability to scale variations and morphological diversity. In the decoder, features are gradually upsampled through patch expanding and fused with encoder representations by skip connections, ensuring recovery of fine-grained spatial details while preserving semantic consistency. A final CNN block refines the reconstructed features and produces the segmentation output.
The left part illustrates the encoder-decoder framework with integrated VSS blocks, FGR modules, and MSACA modules for feature extraction and aggregation. The right part provides detailed structures of the VSS block, the FGR module, and the MSACA module, showing how variable-scale sampling, global relation modeling, and multi-scale adaptive channel attention are jointly employed to enhance segmentation performance.
Mamba-based VSS block
The Mamba architecture, built upon the State Space Model (SSM) paradigm, demonstrates remarkable capability in modeling long-range dependencies for medical image segmentation tasks. Leveraging an efficient state-space recurrence mechanism, Mamba achieves global context modeling with linear computational complexity, enabling superior perception of anatomical relationships across spatial regions and improved delineation of complex lesion boundaries. Mamba models the sequence processing process as a linear dynamic system in continuous time:
where h(t) means the hidden state, x(t) is the input feature, y(t) is the output feature, and A, B, C are learnable parameters. Through discretization, the recursive form is obtained:
where \(\bar{A},\bar{B},C\) are the state transition matrix, input projection matrix, and output projection matrix respectively. This process can model long-range dependencies along the spatial scanning direction, laying the foundation for subsequent two-dimensional extension (S6)43.
To further enhance the model’s capacity for spatial feature extraction, the S6 Block is employed to perform directional unfolding and fusion modeling on 2D medical images as shown in Fig. 8. Given an input feature map denoted as \({\bf{X}}\in {{\mathbb{R}}}^{H\times W\times C}\), the process begins with:
where Πr denotes the Scan Expand. For each directionally unfolded sequence, a state-space recurrence is applied to model dependencies as follows:
where \({{\bf{x}}}_{t}^{(r)}\) is the input at time step t along direction r, \({{\bf{h}}}_{t}^{(r)}\) is the hidden state, and A(r), B(r), C(r) are learnable direction-specific transition, input, and output matrices, respectively. Then, the sequences in different directions are restored to two-dimensional space and weighted fused:
where \({\Pi }_{r}^{-1}\) is the Scan Merge, and γr is the learnable directional attention weight. The consistent modeling of boundary sensitivity and long-range dependencies is especially essential in medical image segmentation tasks. To this end, we designed the VSS Block, which organically combines local convolution with the global scanning characteristics of S6. Given input \({\bf{X}}\in {{\mathbb{R}}}^{H\times W\times C}\), perform layer normalization and linear projection:
where \({W}_{1}\in {{\mathbb{R}}}^{C\times d}\). Subsequently, deep convolutions are used to extract boundary texture features:
The two-dimensional state space modeler SS2D43 was introduced to capture global dynamic correlations across locations:
To achieve channel selectivity and interaction enhancement, we construct a parallel gating path and obtain the gating feature through another linear mapping. We perform element-wise weighted fusion with the main path feature. The final output is expressed as:
where ⊙ denotes element-wise interaction, and W2, W3 are linear mapping matrices.
The scan expand operation decomposes the input into directional sequences for feature modeling, which are then processed by the S6 block. The scan merge operation aggregates the outputs back into the original spatial layout, ensuring both global context and local detail are preserved.
Frequency-guided representation module
Medical image segmentation tasks not only require models to accurately depict the overall morphology of lesions or organs, but also to preserve their boundaries and detailed structures in order to meet the dual requirements of accuracy and interpretability in clinical practice. However, traditional convolutional neural networks (CNNs) are limited by their local receptive field and can only indirectly expand their perception range through multi-layer stacking, which often leads to insufficient global consistency in cases such as low-contrast boundaries, cross-scale structural changes, and blurred lesion contours. Medical images naturally lend themselves to frequency-domain decomposition, where low-frequency components represent the global shape and structure of anatomical regions, while high-frequency components capture edge information and fine-grained textures. Integrating frequency-domain modeling with spatial feature processing thus offers a promising pathway to enhance fine detail representation without compromising the integrity of global structural understanding. Based on this, we propose the Frequency-Guided Representation Module (FGRM), which maps spatial features to the frequency domain for learnable frequency-selective modulation, then remaps them back to the spatial domain for detail restoration and residual fusion, thereby achieving cross-domain complementary modeling. This approach enhances the model’s robustness and generalization capabilities under complex imaging conditions while maintaining segmentation accuracy.
In medical image segmentation, low-frequency components often contain global structural information about organs or tissues, while high-frequency components contain boundary contours and lesion details. In order to explicitly separate and utilize different frequency components in feature modeling, the spatial domain features of the feature map \({\bf{X}}\in {{\mathbb{R}}}^{H\times W\times C}\) are mapped to the frequency domain, which can be expressed as:
where Fc(u, v) denotes the frequency domain representation of channel c at frequency domain coordinates (u, v), Xc(x, y) is the pixel value of the spatial domain input feature map, and H, W are the height and width of the image. Some high-frequency components in original medical images may be noise (such as imaging artifacts and scan noise), and directly participating in segmentation may lead to boundary jitter and mis-segmentation.
In order to enhance frequency components that are beneficial to segmentation and suppress irrelevant frequencies, a learnable complex weight matrix is introduced:
where ρc(u, v) controls the amplitude strength of different frequency components, and ϕc(u, v) controls the fine-tuning of the phase to optimize the structural alignment. The modulation process is:
where ⊙ is element-wise multiplication. This process enhances the global contours of low frequencies to maintain organ shape consistency, highlights high-frequency details related to lesion boundaries, and suppresses ineffective high-frequency components caused by noise.
Subsequently, the frequency modulated features are mapped back to the spatial domain through a two-dimensional inverse transform. The specific formula is as follows:
In the spatial domain, pointwise convolution is used to achieve cross-channel feature interaction, and BN and ReLU are applied to stabilize training and enhance nonlinear expression capabilities:
where σ(⋅) represents the activation function, PWConv is the pointwise convolution operation, and BN is Batch Normalization.
Multi-scale adaptive context aggregation module
In medical image segmentation tasks, objects often exhibit significant scale and morphological diversity. That is, the same image may contain both large structures that occupy the majority of the field of view and tiny lesions measuring only a few pixels. Furthermore, lesion morphology varies significantly between cases, with boundaries varying from regular to highly irregular or even fuzzy. Feature extraction methods that rely solely on a single scale or fixed morphology struggle to simultaneously account for these variations, leading to errors in depicting large object structures or identifying small lesions. To address this challenge, we propose the Multi-Scale Adaptive Context Aggregation (MSACA) module, which efficiently models multi-scale structural information through parallel multi-scale feature extraction branches and a dual Top-k sparse attention mechanism.
The MSACA module first extracts semantic information at different scales through three parallel branches. Specifically, given the input feature \({{\bf{F}}}_{{\rm{in}}}\in {{\mathbb{R}}}^{H\times W\times C}\), k × k average pooling is performed to suppress local texture fluctuations and preserve the global intensity trend, and then feature enhancement is performed through batch normalization and nonlinear activation function:
where ϕ(⋅) represents the ReLU activation function and k denotes the kernel size. The smooth background features provided by this branch can supplement global context information in subsequent fusion, helping to improve the segmentation model’s ability to recognize low-contrast lesions.
Organs and lesions in medical images often exhibit irregular shapes and variable boundaries, and standard convolution with fixed sampling positions may fail to align with the most informative regions. To address this issue, deformable convolution is adopted to adaptively adjust the sampling locations, which is formulated as:
where \({{\bf{F}}}_{{\rm{in}}}\in {{\mathbb{R}}}^{H\times W\times C}\) denotes the input feature map, ϕ(⋅) is a nonlinear activation, and BN(⋅) is batch normalization. The offset field \({\mathbf{\Delta }}\in {{\mathbb{R}}}^{H\times W\times 2K}\) predicts a 2D displacement \(\Delta {p}_{k}(p)\in {{\mathbb{R}}}^{2}\) for each of the K kernel sampling points at spatial location p, and the modulation mask \({\bf{M}}\in {{\mathbb{R}}}^{H\times W\times K}\) provides a learnable confidence weight for each sampled position. Specifically, let \({\mathcal{R}}={\{{r}_{k}\}}_{k=1}^{K}\subset {{\mathbb{Z}}}^{2}\) be the kernel index set. For an output location p, the deformable sampling position is given by p + rk + Δpk(p). The deformable convolution can be written as:
where Wk is the k-th kernel weight. Since p + rk + Δpk(p) can be fractional, Fin(⋅ ) is sampled via differentiable bilinear interpolation. Both Δ and M are predicted by lightweight convolution layers and optimized end-to-end via backpropagation. This branch therefore adapts the receptive field to lesion morphology, improving boundary delineation and structural adaptability.
To expand the receptive field without substantially increasing the parameter count, the third branch incorporates dilated convolution to capture mid-scale contextual information. Given a dilation rate of d, the operation in this branch is formulated as:
where \({{\rm{DilatedConv}}}_{3\times 3}^{d}\) denotes a 3 × 3 convolution with dilation rate d. The outputs of the three branches are weighted fused:
Here, (η1, η2, η3) is a fixed hyper-parameter vector with non-negative components satisfying η1 + η2 + η3 = 1, balancing the three terms. After multi-scale feature fusion, we further introduce a dual Top-k sparse attention mechanism to mitigate the computational burden and noise propagation commonly associated with fully connected attention in high-resolution medical images. The input feature map \({\bf{X}}\in {{\mathbb{R}}}^{B\times C\times H\times W}\) is flattened and linearly mapped to:
where B is the batch size, C is the number of channels, and N = H × W is the pixel sequence length. The key and value are calculated as follows:
where SplitHeads(⋅) divides the channel into h attention heads to enhance the diversity expression capability of features. To highlight key anatomical regions in attention calculation, the MSACA module introduces a dual Top-k strategy. In traditional scaled dot-product attention, the attention weight matrix is:
where dk is the dimension of each attention head. Based on this, we select two Top-K index sets \({{\mathcal{I}}}_{1}\) and \({{\mathcal{I}}}_{2}\) with different proportions, retaining the largest \(\frac{N}{{k}_{1}}\) and \(\frac{N}{{k}_{2}}\) attention connections, respectively, and setting the remaining positions to − ∞ to zero after Softmax:
Here, the Mask operation refers to the selective suppression of attention connections outside the Top-k range, a key step in the Top-k attention mechanism to ensure computational sparsity. The masked attention is formally defined as:
The final output O is obtained by weighting and combining the results of two Top-k attention branches, where α and β control the importance of different branches, which takes into account both global dependencies and local details in feature representation:
Loss function
As the foreground region typically occupies only a small portion of the image in medical image segmentation tasks, the category distribution is highly imbalanced. Additionally, small targets and blurry boundaries often pose challenges for accurate identification. Consequently, a single loss function struggles to balance global structural consistency with local boundary accuracy. To address this, this paper employs a composite optimization objective combining cross-entropy loss with Dice loss to simultaneously constrain pixel-level classification and region-level overlap. The overall loss function is defined as:
where λCE and λDice are weighting coefficients. The cross-entropy loss is used to measure pixel-level classification consistency, which is defined as:
where N represents the total number of pixels, C is the number of categories, yi,c ∈ {0, 1} indicates the true label of the ith pixel in category c, and \({\widehat{y}}_{i,c}\in [0,1]\) denotes the corresponding predicted probability. The Dice loss is used to measure the degree of overlap between the predicted area and the actual area, which is formalized as:
where yi ∈ {0, 1} represents the true label of the pixel, \({\widehat{y}}_{i}\in [0,1]\) denotes the predicted probability, and ε is the smoothing factor.
To further enhance the effectiveness of multi-scale feature learning, this paper introduces a deep supervision mechanism at the multi-layer outputs of the decoder. Specifically, the composite loss mentioned above is calculated for the predicted outputs at each scale, and a weighting factor that decays layer by layer is assigned. The overall optimization objective is:
where S denotes the number of scales in depth supervision, \({{\mathcal{L}}}_{\,{\rm{CE}}}^{(s)}\) and \({{\mathcal{L}}}_{\,{\rm{Dice}}}^{(s)}\) is the cross-entropy loss and Dice loss of the sth scale, respectively. The weight ωs is distributed according to an exponential decay:
By leveraging the complementary constraints of CE and Dice, the model can simultaneously ensure the integrity of the overall anatomical structure and the accuracy of boundary details. Incorporating deep supervision further promotes the effective learning of multi-scale features, resulting in more stable and robust performance in medical image segmentation tasks.
Data availability
All datasets used in this study are publicly available or available upon request: ISIC: International Skin Imaging Collaboration challenge dataset for skin lesion segmentation. Available at: https://challenge.isic-archive.com/data. Kvasir-SEG: Public polyp segmentation dataset collected from colonoscopy videos. Available at: https://datasets.simula.no/kvasir-seg/. ACDC: Automated Cardiac Diagnosis Challenge dataset for cardiac MRl segmentation. Available at: https://www.creatis.insa-lyon.fr/Challenge/acdc/databases.html. SEED: Seed: the challenge of intelligent diagnosis of cancer risk. Available at: https://www.jseedata.com/.
Code availability
Our code is publicly available at: https://github.com/gaidi1/CFG-MambaNet.
References
Ma, J. et al. Segment anything in medical images. Nat. Commun. 15, 654 (2024).
Huang, S.-C., Pareek, A., Seyyedi, S., Banerjee, I. & Lungren, M. P. Fusion of medical imaging and electronic health records using deep learning: a systematic review and implementation guidelines. NPJ Digital Med. 3, 136 (2020).
Asgari Taghanaki, S., Abhishek, K., Cohen, J. P., Cohen-Adad, J. & Hamarneh, G. Deep semantic segmentation of natural and medical images: a review. Artif. Intell. Rev. 54, 137–178 (2021).
Cui, C., Li, Y., Liu, S., Wang, P. & Huang, Z. The unsupervised machine learning to analyze the use strategy of statins for ischaemic stroke patients with elevated transaminase. Clin. Neurol. Neurosurg. 232, 107900 (2023).
Salpea, N., Tzouveli, P. & Kollias, D. Medical image segmentation: a review of modern architectures. In European Conference on Computer Vision, 691–708 (Springer, 2022).
Azad, R. et al. Medical image segmentation review: the success of u-net. In IEEE Transactions on Pattern Analysis and Machine Intelligence, Vol. 12, 10076–10095 (2024).
Zhou, Z., Rahman Siddiquee, M. M., Tajbakhsh, N. & Liang, J. Unet++: a nested u-net architecture for medical image segmentation. In International Workshop on Deep Learning in Medical Image Analysis, 3–11 (Springer, 2018).
Huang, H. et al. Unet 3+: a full-scale connected unet for medical image segmentation. In ICASSP 2020-2020 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 1055–1059 (IEEE, 2020).
Milletari, F., Navab, N. & Ahmadi, S.-A. V-net: fully convolutional neural networks for volumetric medical image segmentation. In 2016 Fourth International Conference on 3D Vision (3DV), 565–571 (IEEE, 2016).
Zhang, L. et al. Block level skip connections across cascaded v-net for multi-organ segmentation. IEEE Trans. Med. Imaging 39, 2782–2793 (2020).
Chen, J., Yang, L., Zhang, Y., Alber, M. & Chen, D. Z. Combining fully convolutional and recurrent neural networks for 3d biomedical image segmentation. In Advances in Neural Information Processing Systems 29 (Curran Associates, 2016).
Oktay, O. et al. Attention u-net: learning where to look for the pancreas. Preprint at arXiv https://doi.org/10.48550/arXiv.1804.03999 (2018).
Gu, R. et al. Ca-net: comprehensive attention convolutional neural networks for explainable medical image segmentation. IEEE Trans. Med. Imaging 40, 699–711 (2020).
Xiao, H., Li, L., Liu, Q., Zhu, X. & Zhang, Q. Transformers in medical image segmentation: a review. Biomed. Signal Process. Control 84, 104791 (2023).
Chen, J. et al. Transunet: rethinking the u-net architecture design for medical image segmentation through the lens of transformers. Med. Image Anal. 97, 103280 (2024).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In European Conference on Computer Vision, 205–218 (Springer, 2022).
Tang, H. et al. Htc-net: a hybrid cnn-transformer framework for medical image segmentation. Biomed. Signal Process. Control 88, 105605 (2024).
Tragakis, A., Kaul, C., Murray-Smith, R. & Husmeier, D. The fully convolutional transformer for medical image segmentation. In Proc. IEEE/CVF Winter Conference on Applications of Computer Vision, 3660–3669 (IEEE, 2023).
Kuang, H. et al. Hybrid cnn-transformer network with circular feature interaction for acute ischemic stroke lesion segmentation on non-contrast ct scans. IEEE Trans. Med. Imaging 43, 2303–2316 (2024).
Gu, A. & Dao, T. Mamba: linear-time sequence modeling with selective state spaces. Preprint at arXiv https://doi.org/10.48550/arXiv.2312.00752 (2023).
Ma, J., Li, F. & Wang, B. U-mamba: enhancing long-range dependency for biomedical image segmentation. Preprint at arXiv https://doi.org/10.48550/arXiv.2401.04722 (2024).
Wang, Z., Zheng, J.-Q., Zhang, Y., Cui, G. & Li, L. Mamba-unet: Unet-like pure visual mamba for medical image segmentation. Preprint at arXiv https://doi.org/10.48550/arXiv.2402.05079 (2024).
Liu, J. et al. Swin-umamba: Mamba-based unet with imagenet-based pretraining. In International Conference on Medical Image Computing and Computer-assisted Intervention, 615–625 (Springer, 2024).
Khan, A., Asad, M., Benning, M., Roney, C. & Slabaugh, G. Cams: convolution and attention-free mamba-based cardiac image segmentation. In 2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), 1893–1903 (IEEE, 2025).
Lv, C. et al. Ecm-transunet: edge-enhanced multi-scale attention and convolutional mamba for medical image segmentation. Biomed. Signal Process. Control 107, 107845 (2025).
Xu, J. et al. Cda-mamba: cross-directional attention mamba for enhanced 3d medical image segmentation. Sci. Rep. 15, 21357 (2025).
Ni, R. et al. Mul-vmamba: multimodal semantic segmentation using selection-fusion-based vision-mamba. Knowl. Based Syst. 334, 115119 (2025).
Xing, Z. et al. Segmamba-v2: long-range sequential modeling mamba for general 3d medical image segmentation. In IEEE Transactions on Medical Imaging, Vol. 45, 4–15 (2025).
Bernard, O. et al. Deep learning techniques for automatic mri cardiac multi-structures segmentation and diagnosis: is the problem solved?. IEEE Trans. Med. Imaging 37, 2514–2525 (2018).
Jha, D. et al. Kvasir-seg: a segmented polyp dataset. In International Conference on Multimedia Modeling, 451–462 (Springer, 2019).
Gutman, D. et al. Skin lesion analysis toward melanoma detection: a challenge at the International Symposium on Biomedical Imaging (ISBI) 2016, hosted by the International Skin Imaging Collaboration (ISIC). Preprint at arXiv https://doi.org/10.48550/arXiv.1605.01397 (2016).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-assisted Intervention, 234–241 (Springer, 2015).
Isensee, F., Jaeger, P. F., Kohl, S. A., Petersen, J. & Maier-Hein, K. H. nnu-net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211 (2021).
Liu, Y. et al. Rolling-Unet: revitalizing MLP’s ability to efficiently extract long-distance dependencies for medical image segmentation. In Proc. AAAI Conference on Artificial Intelligence, Vol. 38, 3819–3827 (AAAI Press, 2024).
Guo, X. et al. Uctnet: uncertainty-guided CNN-transformer hybrid networks for medical image segmentation. Pattern Recognit. 152, 110491 (2024).
He, A. et al. H2former: an efficient hierarchical hybrid transformer for medical image segmentation. IEEE Trans. Med. Imaging 42, 2763–2775 (2023).
Huang, X., Deng, Z., Li, D., Yuan, X. & Fu, Y. Missformer: an effective transformer for 2d medical image segmentation. IEEE Trans. Med. Imaging 42, 1484–1494 (2022).
Yan, Z., Liu, Y., Li, X. & Sun, L. Hetero-unet: heterogeneous transformer with mamba for medical image segmentation. In Advancements in Medical Foundation Models: Explainability, Robustness, Security, and Beyond. (Curran Associates, 2024)
Liu, J. et al. Swin-umamba†: adapting mamba-based vision foundation models for medical image segmentation. In IEEE Transactions on Medical Imaging, Vol. 44, 3898–3908 (2024).
Tan, D. et al. A novel skip-connection strategy by fusing spatial and channel wise features for multi-region medical image segmentation. IEEE J. Biomed. Health Inform. 28, 5396–5409 (2024).
Rahman, M. M., Munir, M. & Marculescu, R. Emcad: efficient multi-scale convolutional attention decoding for medical image segmentation. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 11769–11779 (IEEE, 2024).
Wang, S. et al. Gh-unet: group-wise hybrid convolution-vit for robust medical image segmentation. npj Digital Med. 8, 426 (2025).
Liu, Y. et al. Vmamba: visual state space model. Adv. Neural Inf. Process. Syst. 37, 103031–103063 (2024).
Acknowledgements
This work was supported by China Postdoctoral Science Foundation [2024M762614], the Natural Science Foundation of Jiangxi [Grant Nos. 20232BAB212008 and 20242BAB25078], the 2023-SZZ Key Specialized Construction Project in Cardiology Department of Zhejiang Province [Grant No. 2023-SZZ], the Ningbo Key Research and Development Program [Grant No. 2024Z232], the Medical and Health Science and Technology Projects of Zhejiang Province [Grant No. 2024KY281], the Medical Science and Technology Innovation Project of Xuzhou Municipal Health Commission [Grant No. XWKYHT2024112].
Author information
Authors and Affiliations
Contributions
G.R. and D.G. led the experimental investigation and visualization and drafted the manuscript; Z.C. and P.S. carried out validation and contributed to investigation and critical revision; D.L., P.L., and H.Y. directed methodology and project administration, secured funding, and oversaw manuscript revisions; X.Y., W.X., and X.W. developed the methodology and executed validation, with X.W. additionally leading data curation and management; Y.M. provided overall supervision and formal analysis; H.C. organized data acquisition from public sources, handled preprocessing and quality control, and contributed to revisions; X.Z. guided engineering implementation and verified algorithmic results; X.F. performed statistical analysis, consolidated results and figures, and executed substantial manuscript updates.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ren, G., Chen, Z., Su, P. et al. CFG-MambaNet: Contextual and Frequency-Guided Mamba Network for medical image segmentation. npj Digit. Med. 9, 202 (2026). https://doi.org/10.1038/s41746-026-02393-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02393-z