Abstract
Drug–drug interaction (DDI) poses a major challenge in clinical pharmacology, often compromising therapeutic efficacy or causing serious adverse events. Traditional detection methods, heavily dependent on experimental assays and expert knowledge, are constrained by high costs and limited scalability. This work explores emerging machine learning (ML)-based strategies for predicting DDIs by leveraging the rapidly expanding biomedical data landscape. Recent advances in deep learning architectures, graph neural networks and sophisticated feature engineering have markedly improved predictive performance, offering scalable and data-efficient alternatives to conventional approaches. We further highlight real-world clinical applications where ML-based models have enhanced drug safety monitoring and informed therapeutic decision-making. Finally, we discuss critical challenges like model interpretability, generalizability and integration with clinical workflows, and outline future directions toward building robust, explainable and clinically actionable DDI prediction systems. This work provides a comprehensive perspective on how AI-driven methodologies are reshaping pharmacovigilance and precision therapeutics.
Similar content being viewed by others
Introduction
Drug-drug interaction (DDI), defined as the pharmacological or clinical modifications induced by the co-administration of multiple drugs, represent a critical and often underestimated determinant of therapeutic efficacy and patient safety1. DDIs are traditionally categorized into pharmaceutical (PC), pharmacokinetic (PK), and pharmacodynamic (PD) interactions2. Although combination therapies are frequently employed to enhance clinical outcomes, particularly in complex diseases such as cancer and cardiovascular disorders3,4,5, the concurrent use of multiple agents exponentially increases the risk of adverse DDIs6,7. Notably, conventional detection strategies including Phase IV trials and post-marketing surveillance, often fail to capture the full spectrum of clinically relevant DDIs prior to drug approval8, underscoring an urgent need for more proactive and systematic identification approaches. Additionally, experimental methods, including in vitro enzyme inhibition assays and in vivo PK studies, remain the gold standard for DDI detection9,10. However, these methods are inherently constrained by limited throughput, high costs and substantial time investment11,12,13. Moreover, given the rapid expansion of available therapeutic agents, exhaustive experimental validation of all potential DDIs is increasingly impractical14,15,16,17.
In this context, computational modeling has emerged as a compelling alternative, offering scalability, speed and the ability to prioritize high-risk interactions18. Artificial intelligence (AI) has further revolutionized DDI prediction by leveraging large-scale biomedical data to enable graph-based reasoning, natural language processing and deep learning, outperforming traditional methods in cost-efficiency and scalability19,20,21,22,23,24. These advancements align with recent progress in computational drug development, where standardized evaluation frameworks have been developed to assess the quality and clinical applicability of predictive resources25. AI methodologies have already demonstrated remarkable success across multiple pharmacological domains, including antiviral drug discovery, optimization of therapeutic regimens, de novo drug design and prediction of protein-ligand binding affinities26,27,28,29,30, further underscoring their transformative potential. These advancements have expanded the scope of computational approaches, which now fall broadly into two categories: text mining-based methods, which extract DDI knowledge from structured or unstructured biomedical data31, and machine learning (ML)-based methods, which predict novel interactions through statistical learning frameworks.
Among these, ML-based strategies have demonstrated remarkable advances in predictive performance and translational potential. By leveraging extensive pharmacological, chemical and clinical datasets, ML algorithms can uncover complex nonlinear dependencies that often elude traditional heuristics32,33. Current ML paradigms for DDI prediction encompass diverse methodological families, including graph- and network-based models34, classification algorithms35, similarity-driven approaches36, multi-task learning frameworks37, structure-based predictions38, matrix factorization techniques39 and deep learning architectures such as convolutional neural networks (CNNs) and graph neural networks (GNNs)40. Each methodology offers distinct advantages in capturing specific aspects of drug interactions, from chemical structure to clinical outcomes. While substantial progress has been made, several key challenges continue to hinder further advancement. For example, existing models often face limitations in interpretability, generalizability across diverse clinical settings and integration with multi-omics and real-world evidence41. Moreover, while ML has significantly expanded the frontier of DDI prediction, systematic evaluation of these models under clinically meaningful conditions remains limited42. Addressing these challenges is crucial for translating computational predictions into actionable clinical insights.
Presently, we provide a comprehensive study of ML-based approaches for DDI prediction, with an emphasis on recent methodological innovations, comparative performance metrics and clinical applicability. We systematically analyze the strengths and limitations of prevailing techniques, discuss exemplar case studies that illustrate the clinical relevance of DDI prediction, and propose future directions, including the integration of causal inference, explainable AI and real-world data analytics. By synthesizing current knowledge and outlining emerging opportunities, this work aims to accelerate the translation of ML-driven DDI prediction into safer, more effective therapeutic strategies.
The general process of machine learning in drug-drug interactions
ML has emerged as a powerful tool for predicting DDIs by learning from large-scale drug-related datasets and automatically extracting latent patterns to enable efficient predictions43. In DDI research, ML methods are uniquely capable of handling complex, multidimensional data, encompassing chemical structures, molecular properties, biological activities, pharmacological mechanisms and clinical information. This ability to process diverse data types significantly enhances the accuracy, interpretability and reliability of DDI predictions44. Compared with traditional experimental approaches, ML not only accommodates larger and more complex datasets but also reveals intricate relationships between drugs, which is crucial for mitigating clinical risks associated with adverse drug interactions45.
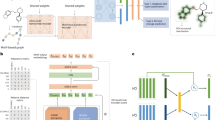
A standardized ML workflow has been widely adopted in DDI research to translate these capabilities into actionable predictions. The overall process for ML-based DDI prediction is illustrated in Fig. 1A. It involves four key stages: data acquisition, model construction, performance evaluation and experimental validation. First, comprehensive datasets are curated from public biomedical databases, with each repository providing specialized data types: (1) DrugBank46,47,48 for targets and drug-target interactions, (2) PubChem49,50,51,52 for chemical structures, bioactivity and chemical molecules, (3) KEGG53,54,55 for genomics, biology and metabolic pathways, and (4) dedicated DDI databases (e.g., TWOSIDES56, DDInter57,58) for known interaction pairs and their clinical effects, etc.
A The workflow of DDI prediction by machine learning method. B Commonly used biomedical databases. C Schematic diagram of drug interaction.
The acquired data serve dual purposes in model development. These datasets provide the foundational features for model development. Subsequently, a range of ML techniques including random forests (RF), support vector machines (SVM) and GNN, are employed to learn complex patterns underlying drug interactions. Model performance is systematically assessed using multiple benchmark datasets and comparative analyses with traditional methods are conducted. Finally, predicted DDIs are subjected to experimental validation through in vitro assays (e.g., cell- and animal-based studies) and clinical data analysis to confirm their reliability and translational relevance59,60,61. This rigorous evaluation phase ensures generalizability before proceeding to biological validation. The commonly used biomedical databases and the schematic diagram of drug interaction are depicted in Fig. 1B, C, respectively.
The efficacy of this workflow hinges on two critical components: high-quality data and adaptable algorithms. High-quality data form the cornerstone of ML-based DDI prediction. Comprehensive databases containing detailed drug information like chemical structures, molecular descriptors, pharmacological mechanisms, as well as the types, intensities and clinical impacts of known DDIs, are essential for training robust models62. A thorough understanding of major biomedical databases, their content, and their specific applications in DDI research is vital for the effective development and validation of predictive models63.
Within this workflow, certain ML approaches have emerged as particularly effective for DDI prediction, including SVM64,65, RF66 and different forms of neural networks. These algorithms analyze drug interactions from multiple perspectives, leveraging features derived from molecular structure, pharmacological properties, and clinical observations. In particular, the advent of deep learning, especially CNNs and GNNs67,68, has markedly advanced the field. Deep learning models excel in automatically extracting high-dimensional, complex features from raw data, significantly improving prediction performance69.
Despite these advances, several challenges persist. Imbalanced datasets can bias model training, reducing prediction reliability. Limited model interpretability often impedes mechanistic understanding of predicted interactions. Moreover, generalization across different datasets remains difficult70. Addressing these challenges has prompted the development of novel algorithms and optimization strategies aimed at improving predictive accuracy, robustness, and model transparency71.
The application of machine learning in drug-drug interactions
To provide a comprehensive overview of current strategies, we systematically summarize the major ML-based DDI prediction methods, which can be categorized into four primary types: (1) learning paradigms-based methods; (2) representation-level-based methods; (3) specific model families-based methods; (4) modeling strategies and training paradigms-based methods. A detailed summary of these approaches is presented in Table 1, facilitating an integrated understanding of methodological advances in the field.
Method based on learning paradigms for predicting drug-drug interactions
The academic community has developed various traditional ML methods to address the increasingly challenging prediction of DDIs72, each with unique advantages and considerations. These methods integrate diverse features and data sources to predict potential drug interactions73,74, thereby improving patient safety and clinical efficacy. These methods integrate diverse features and data sources to predict potential drug interactions, thereby improving patient safety and clinical efficacy. The most widely used methods among them are multi-task learning-based methods, and traditional ML classifiers with handcrafted features-based methods. These traditional ML methods have unique advantages in handling diverse data types and revealing potential associations between drugs75, providing important support for drug safety and pharmacovigilance work.
In detail, method based on multi-task learning leverages the shared knowledge across related tasks, such as predicting different types of drug-drug interactions or jointly predicting DDIs, to enhance the generalization and robustness of the primary DDI prediction task76. This paradigm addresses the common issue of data sparsity in DDI prediction by utilizing auxiliary information from related domains, allowing the model to learn more informative and transferable drug representations. By sharing underlying feature representations and learning a common latent space, multi-task learning-based methods can uncover complex, non-linear relationships that might be overlooked in single-task models77. This approach provides a powerful framework for predicting diverse and high-risk drug combinations, especially when labeled data for specific interaction types is limited. The ability to simultaneously model multiple related pharmacological outcomes makes multi-task learning an increasingly vital strategy for comprehensive drug safety assessment and polypharmacy management.
As mentioned earlier, drug combinations are traditional ML classifiers with handcrafted features-based methods through the extraction of features from large datasets to identify potential DDIs78. This approach provides a reliable means for predicting high-risk drug combinations, especially in cases where clinical trial data is scarce or unavailable. However, when drugs with similar chemical structures are used in combination, traditional ML classifiers with handcrafted features-based methods may generate comparable reactions79. This method can comprehensively evaluate drug interactions by integrating multidimensional drug similarity indicators, such as molecular structure, biological targets, and side effects80. By incorporating ML algorithms to capture the complexity of drug interactions, the prediction accuracy of traditional ML classifiers with handcrafted features-based methods has been significantly improved.
These two methods, i.e., multi-task learning and traditional ML classifiers with handcrafted features, complement each other to construct a complete framework for predicting DDIs. They not only lay the foundation for more advanced technologies, but also provide important support for research to improve drug safety and efficacy. In the next section, we will delve into the application of each method in the field of DDIs.
Method based on multi-task learning
DDI refers to the phenomenon where two drugs synergistically or antagonistically influence each other’s activity when administered in combination. Although some DDIs can be retrieved from existing databases, a substantial portion of information remains embedded within the vast body of biomedical literature81. Thus, there is an urgent need to develop automated methods capable of extracting potential DDI information from textual sources82. The workflow for DDI prediction based on multi-task learning is illustrated in Fig. 2A, demonstrating how different tasks synergistically contribute to DDI prediction.
A Flow chart for predict drug-drug interaction based on Multi-Task Learning. B Flow chart of drug-drug interaction prediction based on Traditional Machine Learning Classifiers with Handcrafted Features.
A notable contribution in this area is the work of Zhou et al.83, who proposed a novel position-aware deep multi-task learning model specifically designed for DDI extraction from biomedical texts. In their approach, sentences are represented as sequences of word embeddings combined with position embeddings. An attention-based bidirectional long short-term memory (BiLSTM) network is used to encode the contextual features of each sentence. The relative positional information of words with respect to the target drug entities is incorporated into the BiLSTM hidden states to generate position-aware attention weights. Furthermore, within the multi-task learning framework, two related tasks—predicting whether two drugs interact and classifying the specific types of interactions—are jointly learned. Experimental results demonstrated that their model achieved an interaction type identification accuracy of 72.99%, surpassing the state-of-the-art method by 1.51%. These findings underscore the effectiveness of incorporating positional information and multi-task learning for DDI extraction from biomedical text.
Potential adverse DDIs can significantly increase the risks associated with combination therapies; however, existing computational methods often fail to detect them effectively. To address this limitation, Wang et al.84 proposed an ensemble-based multi-task neural network, termed DEML, designed for the simultaneous optimization of five synergy regression prediction tasks, a synergy classification task, and a DDI classification task. DEML integrates chemical features and transcriptomics data as inputs. A novel hybrid ensemble layer structure is employed to construct higher-order representations from different perspectives, while a task-specific fusion layer with a gating mechanism links representations for each task. For the Loewe synergy prediction task, DEML achieved a 7.8% reduction in root mean squared error and a 13.2% improvement in the R² correlation coefficient compared to the previous best-performing method. By combining drug synergy prediction and DDI classification within an ensemble multi-task learning framework, DEML significantly enhances prediction accuracy. Nevertheless, while DEML exhibits stable performance across multiple tasks, further refinements—such as improving DDI subtype classification and enhancing adaptability to cross-domain datasets—represent promising directions for future research.
Method based on traditional machine learning classifiers with handcrafted features
Traditional ML classifiers with handcrafted features-based approaches have emerged as a common and effective strategy in DDI prediction research. Although DDIs may occur incidentally during co-administration, their detection through clinical trials remains limited due to small sample sizes85. Consequently, most DDIs are only identified post-marketing, often after causing serious adverse drug reactions86. Traditional ML classifiers with handcrafted features-based methods offer not only high predictive efficiency but also the ability to uncover potential drug interactions by leveraging diverse feature extraction techniques. These methods enable researchers to screen high-risk drug combinations from large datasets, thereby significantly contributing to drug safety research. In recent years, the application of traditional ML classifiers with handcrafted features-based methods in DDI prediction has gained increasing attention. An overview of the general workflow for traditional ML classifiers with handcrafted features-based DDI prediction is illustrated in Fig. 2B.
To address the growing complexity of DDI prediction, Shi et al.87 proposed a novel local classification-based model (LCM), which offers faster training speed, lower memory requirements, and mitigates biases inherent in previous approaches. Building on this foundation, they developed a supervised fusion algorithm based on Dempster-Shafer (DS) theory of evidence, termed LCM-DS. This framework integrates three components to make final predictions: the LCM output, the closeness between the LCM output and a reference output, and a quality classifier assessing the reference output. Experimental results demonstrated that LCM-DS significantly outperformed three state-of-the-art baseline methods as well as classical fusion algorithms in DDI prediction tasks. By integrating multiple classifiers and employing DS evidence theory, LCM-DS enhances prediction performance and robustness. Nevertheless, as the number of drugs increases, the model’s computational and storage demands may become a limiting factor. Additionally, further improvements in fusion method selection and similarity matrix construction could enhance performance. Future research directions include optimizing model scalability, refining fusion strategies, addressing class imbalance issues, and improving interpretability to expand the model’s practical utility.
In contrast, Kastrin et al.88 adopted a different approach by formulating DDI prediction as a binary link prediction task within potential DDI networks. They applied link prediction techniques to infer unknown interactions across five large-scale DDI databases—DrugBank, KEGG, NDF-RT, SemMedDB, and Twosides. Their study evaluated predictive performance using both unsupervised and supervised learning methods, with features derived from topological and semantic similarities. Supervised learning models—including classification trees, k-nearest neighbors, SVM, RF, and gradient boosting machines—consistently outperformed unsupervised approaches. The findings underscore the potential of supervised link prediction as a powerful tool for identifying novel DDIs and advancing clinical research. Although the method demonstrates strong practicality and innovation, its robustness could be further improved through the use of cross-validation and multiple evaluation metrics. Furthermore, incorporating model interpretability techniques such as LIME would enhance transparency and trustworthiness, especially in clinical settings.
Method based on representation-level for predicting drug drug interactions
Beyond the selection of learning paradigms, the effectiveness of DDI prediction critically depends on how drug information is computationally represented. The representation-level approach focuses on transforming raw drug data-such as molecular structures, biological targets, and pharmacological properties—into structured mathematical forms that ML models can process. The design of these representations directly influences a model’s ability to capture complex drug–drug relationships and underlying interaction mechanisms. The most widely used methods among them are similarity-based methods, and structural characteristics-based methods. Similarity-based representations leverage the fundamental principle that similar drugs are likely to share similar interaction profiles, integrating multiple pharmacological and molecular data sources to construct holistic similarity measures89. In contrast, structural characteristics-based representations directly encode the intrinsic physicochemical and topological information of a drug molecule, from atomic composition and functional groups to complete molecular graph structures. Both approaches provide complementary perspectives for quantifying and comparing drug profiles, serving as essential input features for a wide range of ML algorithms. The computational techniques, and application scenarios of these two key representation methods in DDI prediction will be examined in the following sections.
Method based on similarity
The core idea of traditional similarity-based approaches for DDI prediction is that if drug A interacts with drug B to produce a specific effect, then drug A (or drug B) may exhibit similar effects when combined with other drugs. By integrating multiple drug similarity features, these methods facilitate the prediction of potential interactions involving novel drugs90. DDI remains a major contributor to adverse drug reactions in clinical practice91. Particularly, similarity-based methods have been developed with good predictive accuracy, and ML techniques have further extended their applicability. However, the performance of some ML-based models has not met expectations, largely due to limited training datasets and suboptimal similarity measures. An overview of the similarity-based DDI prediction workflow is provided in Fig. 3A.
A Flow chart for predict drug-drug interaction based on Similarity. B Flow chart of drug-drug interaction prediction based on Structural Characteristics.
In recent years, ML methods have shown significant promise in improving the prediction of DDIs, which is critical for pharmacovigilance and drug safety. Various strategies have been proposed to enhance the accuracy and efficiency of DDI prediction by incorporating diverse drug similarity and interaction features. For instance, Song et al.92 developed a SVM-based model utilizing an extensive set of similarity measures derived from well-established databases such as DrugBank and SIDER. The similarity metrics included 2D molecular structure similarity, 3D pharmacophoric similarity, interaction profile fingerprint similarity, target similarity, and adverse drug effect (ADE) similarity. A pairwise kernel was constructed based on these five similarity measures and used as the input vector for the SVM classifier. Ten-fold cross-validation demonstrated excellent predictive performance, achieving an area under the receiver operating characteristic curve (AUROC) greater than 0.97—substantially outperforming previous models with AUROCs around 0.67. This study highlighted the effectiveness of integrating multiple drug similarity features in enhancing DDI prediction. The similarity-based SVM approach offers a robust tool for pharmacovigilance by providing comprehensive and accurate predictions of potential DDIs.
Building on the integration of drug similarity information, Sridhar et al.93 proposed a probabilistic framework for predicting unknown DDIs by leveraging multiple networks of drug similarity and known interactions. They employed probabilistic soft logic (PSL), a highly scalable and extensible probabilistic programming framework, to develop a joint inference model. By combining multiple similarity sources with domain knowledge of network structures, the PSL-based model significantly outperformed existing similarity-based approaches. Evaluation across two datasets containing three types of drug interactions showed that the collective PSL model achieved superior performance on key link prediction metrics, including area under the precision-recall curve (AUPR) and AUROC, exceeding the performance of advanced systems such as INDI by over 50% in AUPR. Furthermore, the model successfully identified five novel DDIs. Although the results are promising, future improvements could focus on enhancing the model’s capability to predict rare or unlabeled interactions and incorporating additional multidimensional data sources—such as patient characteristics—to improve the model’s generalization ability and practical applicability.
In another study, Yan et al.94 introduced a novel method called DDI-IS-SL, which predicts DDIs by integrating similarity information and semi-supervised learning. DDI-IS-SL combines chemical, biological, and phenotypic drug data to calculate feature similarities using cosine similarity. It also constructs Gaussian interaction profile kernel similarities based on known DDIs. A semi-supervised learning algorithm, the Regularized Least Squares classifier, was then employed to predict interaction likelihood scores between drug pairs. Across 5-fold and 10-fold cross-validation studies, as well as de novo drug validation tasks, DDI-IS-SL consistently outperformed comparative methods in prediction performance. While the approach demonstrates strong predictive capabilities, there remains room for further advancement, particularly in improving adaptability to complex real-world scenarios, enhancing prediction accuracy, and increasing the interpretability of the model’s outputs.
Method based on structural characteristics
In the development of prediction models, the extraction of structural characteristics often constitutes a critical step95. Structural characteristic-based prediction methods involve extracting salient features from the data and constructing feature vectors that serve as the foundation for subsequent model training. These features can include numerical, textual, or image-derived information from the original data, as well as abstract features obtained through transformation or dimensionality reduction techniques. Methods based on structural characteristics place particular emphasis on capturing the internal structures and relationships within the data, leveraging complex topological features such as graphs and networks to predict potential associations. The key advantage of these methods lies in their ability to uncover latent patterns and rules within complex datasets, thereby enhancing prediction accuracy. The workflow of DDI prediction based on structural characteristics is summarized in Fig. 3B.
DDIs arise from interactions between various chemical substructures (functional groups) of different drugs. In existing DDI prediction methods based on substructures, each node is typically treated as the center of a substructure pattern, and adjacent nodes are viewed as centers of similar substructures. However, this often introduces redundancy, and significant structural and functional heterogeneity among compounds can result in unrelated pairings, complicating information integration and potentially undermining prediction performance. To address these challenges, Yuan Liang96 proposed a novel DDI prediction method based on Substructure Signature Learning (DDI-SSL). This approach extracts meaningful information from local subgraphs surrounding each drug and effectively utilizes substructures to assist in predicting drug side effects. Furthermore, a deep clustering algorithm aggregates similar substructures, enabling the reconstruction of individual subgraphs based on a set of global substructure signatures. A layer-independent collaborative attention mechanism is also introduced to model the mutual influence between drugs, generating signal strength scores for each drug class to mitigate noise caused by heterogeneity. Experimental evaluation on a comprehensive dataset demonstrated that DDI-SSL outperformed existing state-of-the-art methods in DDI prediction. Notably, the introduction of substructure signature learning and collaborative attention mechanisms significantly improved prediction performance and robustness. Nevertheless, there remains room for improvement in terms of model interpretability, data diversity, computational efficiency, and robustness. Future efforts aimed at optimizing these aspects could further enhance the practical value and applicability of the DDI-SSL model.
Most existing methods focus on predicting interactions based on chemical substructures rather than considering the full chemical structures of drugs. Additionally, many rely on manually designed molecular representations, which are inherently limited by domain knowledge and often lack universality and flexibility. In response, Nyamabo et al.97 proposed the Substructure–Substructure Interaction–Drug–Drug Interaction (SSI-DDI) framework, a knowledge-driven deep learning approach that directly operates on raw molecular graph representations for richer feature extraction. Crucially, SSI-DDI decomposes the DDI prediction task into identifying pairwise interactions between substructures of two drugs. Evaluation on real-world datasets showed that SSI-DDI improved DDI prediction performance compared to state-of-the-art methods. However, a noted limitation is that changes in the ordering of drug pairs during training can affect performance, possibly due to the leakage of noise during substructure extraction. Despite this, the approach highlights the importance of modeling fine-grained interactions between drug substructures.
While traditional experimental methods can predict DDIs, they are often time-consuming and costly, underscoring the need for more efficient computational approaches. In this context, Guo et al.98 proposed MSResG, a deep learning framework that integrates multi-source drug features with a Graph Auto-Encoder for DDI prediction. Specifically, MSResG obtains four types of drug feature representations—chemical substructure, target, pathway, and enzyme information—from databases and calculates pairwise Jaccard similarities. These similarities are then integrated by averaging to balance feature contributions. A comprehensive similarity network, combined with a drug interaction network, is encoded and decoded using a graph auto-encoder built upon a residual graph convolutional network. The encoding phase learns potential drug feature vectors that capture both similarity and interaction information, while the decoding phase reconstructs the network to predict unknown DDIs. Experimental results demonstrated that MSResG outperformed previous state-of-the-art methods. By integrating diverse feature types and utilizing a residual graph convolutional structure, MSResG significantly enhanced prediction performance. However, further optimizations, particularly in feature selection and similarity calculation strategies, are needed to improve the model’s ability to capture complex drug interactions. Additionally, increased experimental validation and testing on larger datasets would help enhance the model’s generalization and robustness.
Similarly, Wang et al.99 proposed StructNet-DDI, a deep learning framework based on SMILES representations of chemical structures for DDI prediction, but utilizing a distinct methodology. Their model extracts Morgan fingerprints and key molecular descriptors, converting them into raw graphical features that are input into a modified ResNet18 architecture. By leveraging a deep residual network enhanced with regularization techniques, the model effectively addresses training challenges such as gradient vanishing and explosion, resulting in superior predictive performance. StructNet-DDI demonstrated excellent results across multiple metrics, including AUC, accuracy, and AUPR. By combining SMILES strings, Morgan fingerprints, and molecular descriptors, StructNet-DDI offers a simple yet robust tool for DDI prediction. Nevertheless, a potential limitation is its reliance on SMILES representations, which may overlook critical three-dimensional structural information. Future research could consider integrating 3D molecular structures or incorporating GNNs to further improve accuracy and generalizability. Moreover, targeted strategies for addressing data imbalance could further enhance model performance.
Method based on specific model families for predicting drug drug interactions
The advancement of DDI prediction has been significantly driven by the development of specialized computational frameworks that are particularly suited for handling complex pharmacological data structures. In recent years, matrix factorization has attracted much attention for its ability to extract potential patterns from sparse drug interaction matrices100. Even in the absence of clear interaction data, these methods can predict unknown interactions by revealing implicit associations between drugs. The matrix factorization technique has shown excellent performance in improving the performance of drug interaction prediction models. The ability to model complex, high-dimensional drug data makes matrix factorization an ideal tool for solving drug development and disease treatment challenges.
To further investigate the application of deep learning approaches in DDI prediction, several innovative methods have been developed, including those based on deep graph embedding, GNNs, as well as attention mechanisms and siamese networks. These methods are designed to leverage the capabilities of advanced deep learning techniques to address the inherent challenges associated with DDI prediction, such as data sparsity, the complexity of biological networks, and the demand for improved predictive accuracy101,102.
Deep graph embedding methods aim to map the nodes or edges of a graph into a low-dimensional vector space103. These vectors preserve the structural information of the graph, enabling them to be used effectively in prediction tasks. By transforming the structural data into continuous vectors, this approach addresses the computational inefficiencies associated with traditional graph algorithms when handling large-scale graphs.
On the other hand, GNNs extend the concept of deep graph embedding. GNNs learn representations of graph-structured data by aggregating and propagating information across nodes104. By leveraging the characteristics of neighboring nodes, GNNs update the node representations to capture complex nonlinear relationships. This process allows GNNs to model higher-order dependencies and intricate interactions between nodes, thereby improving prediction accuracy105.
In the following sections, a detailed exploration will be conducted into three distinct approaches for DDI prediction: methods based on matrix factorization, those leveraging graph embedding, and methods based on GNN. Each of these approaches is recognized for offering unique advantages and addressing specific problems within the field.
Method based on matrix factorization
With the rapid development of the biomedical field, the complexity of drug development and disease treatment is continually increasing106. As one of the key factors affecting drug efficacy and safety, DDIs have become a major focus in pharmacological research107. Traditional DDI prediction methods typically rely on experimental data or expert knowledge. However, these approaches are often limited by the availability of data and the high cost of experiments. In recent years, ML and data mining techniques, particularly matrix factorization methods, have emerged as important tools for DDI research.
Matrix factorization can effectively extract latent rules and relationships from extensive drug interaction data, thereby enhancing prediction accuracy and efficiency. By representing DDI data as a matrix and applying matrix factorization techniques—such as singular value decomposition (SVD)108 and non-negative matrix factorization (NMF)109-potential links between drugs can be identified, enabling the prediction of unknown interactions. Consequently, matrix factorization offers a new perspective and an efficient solution for DDI prediction. The overall flowchart of DDI prediction based on matrix factorization is summarized in Fig. 4.
a SVD schematic diagram. b NMF schematic diagram.
To address challenges in DDI prediction, Shi et al.110 proposed a method based on Balanced Regularized Semi-Nonnegative Matrix Factorization (BRSNMF), aiming to enhance DDI prediction in scenarios involving drug community detection and cold-start problems. By introducing a weak equilibrium relationship, BRSNMF effectively partitions drugs into communities of reasonable size and pharmacological significance. These communities exhibit weak equilibrium characteristics, aiding in the understanding of high-order DDIs. Moreover, for new drugs without known interactions, BRSNMF leverages drug-binding protein features to associate them with existing drugs. Experimental results demonstrated that BRSNMF achieves high performance, with 94% accuracy in predicting the top 50 enhanced DDIs and 86% accuracy for the last 50 inhibitory DDIs. Although BRSNMF significantly improves prediction accuracy and applicability, further optimization may be necessary to better address multi-drug interaction scenarios in complex disease treatments.
Building on matrix factorization, Zhang et al.111 introduced Manifold Regularized Matrix Factorization (MRMF) for DDI prediction. This method treats DDI prediction as a matrix completion task, projecting drugs into a low-dimensional interaction space. MRMF incorporates multiple drug features—such as substructures, targets, enzymes, transporters, pathways, indications, side effects, and unexpected side effects—and embeds them as manifolds within the feature space. By introducing manifold regularization based on drug characteristics, MRMF effectively enhances the predictive capability of matrix factorization. Experiments demonstrated that MRMF achieved an AUPR of 0.7963, outperforming other state-of-the-art methods in cross-validation and case studies, with manifold regularization playing a critical role in accuracy improvement.
Similarly, Rohani et al.112 proposed Integrated Similarity Constrained Matrix Factorization (ISCMF) for DDI prediction. ISCMF integrates multiple drug similarity features, including substructure, targets, side effects, pathways, transporters, enzymes, and indications, and incorporates Gaussian interaction profiles to capture interactive information. Through a nonlinear similarity fusion approach, ISCMF enhances feature expression and projects drugs into a low-rank space for interaction prediction. Although the components of ISCMF had been explored individually in previous studies, their integration for DDI prediction represents a novel contribution. ISCMF achieved substantial improvements, increasing AUPR and F-measure by 10% and 18%, respectively. Further case studies confirmed the accuracy of high-confidence predictions. However, ISCMF may face computational efficiency challenges when scaling to large drug pair datasets. Future improvements could involve optimizing the model’s algorithmic efficiency or adopting parallel computing techniques to accelerate training.
To comprehensively predict enhanced and weakened DDIs, Shi et al.113 proposed a Triple Matrix Factorization Unified Framework (TMFUF). Unlike traditional binary DDI prediction approaches, TMFUF captures pharmacological changes resulting from drug interactions. By utilizing drug side effect information, TMFUF achieved an AUC of 0.842 and an AUPR of 0.526, with AUC and AUPR improvements of ~7% and 20%, respectively, over leading existing methods. Furthermore, TMFUF demonstrated strong performance in various DDI screening tasks, particularly in identifying drug pairings associated with significant side effects, thereby facilitating clinical validation.
TMFUF not only predicts binary DDIs but also captures the pharmacological impact of DDIs, offering a unified solution for multi-drug prescriptions and prediction between drugs lacking prior interaction records. Its strength lies in revealing drug characteristics significantly influencing DDI formation, providing valuable insights for clinical applications. Nonetheless, opportunities remain for further improvement, such as enhancing data diversity, improving model interpretability, boosting generalization ability, and strengthening predictive capabilities for novel drug combinations.
In addition, Jain et al.114 proposed the Graph Regularized Probabilistic Matrix Factorization (GRPMF) method. This method integrates expert knowledge into the matrix factorization paradigm by introducing a novel graph-based regularization term. The regularization term is designed to enforce expected (dis)similarities between drug pairs based on prior knowledge, enhancing the prediction of DDIs. The approach formulates the DDI prediction as a matrix completion problem, where the goal is to predict missing interactions between drugs based on observed ones. GRPMF optimizes this formulation using an efficient alternating optimization algorithm, which solves the non-convex problem effectively. Experimental results on the DrugBank dataset show that GRPMF outperforms state-of-the-art techniques, demonstrating its potential for accurate and scalable DDI prediction. This method is particularly valuable for drug development and repurposing, providing a computational tool for identifying potential adverse drug reactions and interactions at a large scale.
Method based on deep graph embedding
Deep graph embedding methods aim to project a graph into a low-dimensional space while preserving its structural information. In this lower-dimensional space, node representations are learned automatically, making them suitable for subsequent predictive tasks. Compared to shallow embedding techniques, deep graph embeddings have demonstrated superior capability in modeling complex biological data and networks, thereby providing more accurate prediction outcomes115. The complete workflow of DDI prediction using deep graph embedding methods is systematically illustrated in Fig. 5A, which outlines the key steps from graph construction to final interaction prediction.
A Flow chart for predict drug-drug interaction based on Deep Graph Embedding method. B Flow chart of drug-drug interaction prediction based on graph neural network method.
In recent years, knowledge graph (KG) embedding methods have attracted considerable attention in DDI prediction, as they project drug interactions into a low-dimensional feature space for more effective prediction. However, existing approaches often suffer from oversimplification, which limits their ability to train accurate models. To address this limitation, Dai et al.116 proposed a a deep learning-based approach leveraging autoencoders to generate high-quality negative samples. The autoencoder’s hidden vectors serve as plausible candidate drugs, and a discriminator learns embeddings of drugs and interactions based on both positive and negative triplets. To mitigate the vanishing gradient problem in discrete representations, they incorporated Gumbel-Softmax relaxation and Wasserstein distance, which stabilize the training process. Compared to traditional reinforcement learning (RL) strategies and gradient-based methods, this approach proves more effective in optimization tasks and can be applied to enhance the performance of existing models without significant modifications. The deep KG embedding framework introduced by Dai et al. improves performance in DDI tasks by addressing the vanishing gradient issue in generative models and showing significant advancements in link prediction and DDI classification. However, challenges related to insufficient sample diversity persist, and future work could explore more sophisticated negative sample generation techniques or integrate domain-specific knowledge to enhance model generalization and accuracy.
In a similar vein, Wang et al.117 introduced an innovative graph embedding-based method for DDI prediction that addresses data incompleteness and sparsity. This approach enables multi-label predictions for drug pairs by constructing a large-scale drug KG from diverse sources and integrating biomedical text into a shared low-dimensional space. The resulting embeddings are used for efficient computation of DDI information through link prediction. This method not only resolves challenges related to data sparsity and computational complexity but also enables the prediction of a range of pharmacological mechanisms and side effects. Extensive experiments on real-world datasets demonstrate that this framework outperforms existing baseline methods in both performance and accuracy. However, the model still faces issues with data noise, especially when applied to large-scale datasets. Future research could focus on improving the model’s ability to generalize and handle outliers to enhance its reliability in clinical applications.
Additionally, Jiang et al.118 proposed a novel DDI prediction method, RaGSECo, which utilizes relation-aware graph structure embedding with co-contrastive learning. They constructed two heterogeneous graphs—a multi-relational DDI graph and a multi-attribute drug similarity graph—and used these graphs to learn and propagate relation-aware graph structure embeddings (RaGSEs) for drugs. This approach ensures that new drugs can also benefit from effective RaGSEs. The co-contrastive learning module developed in this method learns feature representations for drug pairs from two distinct views (interaction and similarity), promoting mutual supervision between the views to obtain more discriminative drug pair representations. RaGSECo’s multi-graph structure and co-contrastive learning provide an effective solution for the new drug prediction problem. Experimental results show that it outperforms existing methods, although maintaining efficiency and accuracy in large-scale and complex drug networks remains a challenge.
Method based on graph neural network
As specialized deep learning architectures for graph-structured pharmacological data, GNNs extend the concept of CNNs to non-Euclidean spaces, providing a more natural and efficient framework for modeling graph-structured data. Unlike static embedding methods, GNNs dynamically update node representations by iteratively aggregating neighborhood information, thereby eliminating the need for tedious manual feature engineering. In recent years, the application of GNNs at the molecular level has achieved significant progress and demonstrated excellent performance in various drug-related tasks119. The complete workflow of DDI prediction based on GNNs is visualized in Fig. 5B, illustrating the key steps from graph construction to interaction prediction.
A notable advancement in this field is the work by Han et al.120, who proposed a DDI prediction model named SmileGNN. This model aggregates both the structural features of drugs derived from SMILES data and the topological features obtained from KGs using GNNs. Experimental results showed that SmileGNN, by integrating multiple data sources, outperformed existing DDI prediction models in prediction performance. Among the top ten predicted novel DDIs, five were verified using the latest databases, highlighting the model’s credibility. Although SmileGNN demonstrates promising results in integrating multi-source data and enhancing performance, there is still room for improvement, such as better predicting rare drug interactions, expanding to larger drug libraries, and optimizing computational efficiency and interpretability for broader clinical applications.
Currently, most computer-aided DDI prediction methods primarily rely on drug-related features or DDI networks, often neglecting the potential information contained in biological entities such as targets and genes. Furthermore, models based on DDI networks alone tend to perform poorly when predicting interactions for drugs without known DDI records. To address these limitations, Yu et al.121 introduced the attention-based cross-domain graph neural network (ACDGNN) for DDI prediction. ACDGNN constructs a biological heterogeneous network that incorporates various drug-related entities and propagates information across domains. Unlike previous approaches, it employs cross-domain transformations to eliminate the heterogeneity between different types of entities, enabling effective integration of biomedical entity information. ACDGNN can be applied in both transductive and inductive settings. Extensive experiments on real-world datasets demonstrated that ACDGNN outperforms several state-of-the-art methods, effectively addressing the limitations of traditional models and improving prediction accuracy, especially for drugs without prior DDI records.
Another challenge in existing GNN-based methods is that their training processes often fail to directly generate predictive models, resulting in poor performance on unseen data. To overcome this, Peng et al.122 proposed a novel reverse GNN model. This model learns graphs from the intrinsic space of original data points and introduces an innovative out-of-sample extension method. By doing so, the reverse GNN can generate high-quality graphs, enhancing feature learning effectiveness. More importantly, the out-of-sample extension allows the model to perform strongly across both supervised and semi-supervised tasks. Experimental results on real-world datasets showed that the reverse GNN achieved competitive performance in semi-supervised node classification, out-of-sample extension, robustness against random edge attacks, link prediction, and image retrieval tasks. Nonetheless, further improvements are needed in areas such as robustness, computational efficiency, and generalization to make the method more practical.
Taking a different approach, Chen et al.123 proposed the use of signed GNNs to model both assortative and disassortative relationships between drug pairs. Because negative links prevent the direct application of spectral filters designed for unsigned graphs, the authors divided the signed graph into two unsigned subgraphs, each with its specialized spectral filter to capture the commonalities and differences between drug pairs. For drug representation, they developed two signed graph filtering-based neural networks (SGFNNs) that integrate both the signed graph structures and drug node attributes. An end-to-end framework jointly trains the SGFNN and a discriminator under a task-specific loss function. The signed GNN framework effectively distinguishes between homogeneous and heterogeneous drug relationships, overcoming the limitations of traditional unsigned models. Although this approach improves DDI prediction performance, further work is needed to handle larger-scale drug networks, enhance computational efficiency, and improve model interpretability, particularly to facilitate clinical application.
Additionally, Gao et al.124 proposes AutoDDI, an automated method for DDI prediction using GNNs. AutoDDI addresses the challenge of manually designing GNN architectures for DDI prediction by using RL to automatically search and optimize the architecture. The method involves creating a search space for the drug dataset, which adapts the depth of GNN layers and captures the interaction information between drug pairs. AutoDDI outperforms previous handcrafted GNN architectures in DDI prediction tasks on real-world datasets, demonstrating superior performance and providing an effective solution for automatically designing GNN architectures for DDI prediction, ultimately enhancing the accuracy and reliability of drug interaction predictions in clinical applications.
Method based on modeling strategies and training paradigms for predicting drug-drug interactions
Besides, the modeling strategies and training paradigms employed in the learning process play an equally crucial role in determining the performance and robustness of DDI prediction systems. These approaches encompass the high-level design principles and optimization frameworks that govern how models are structured, trained, and regularized, often enabling them to better capture complex interaction patterns, improve generalization, and overcome data-related challenges such as sparsity or class imbalance. Among the most prominent and effective of these advanced strategies are those based on attention mechanisms and Siamese networks. The integration of attention mechanisms and Siamese networks has been demonstrated to provide a promising avenue for DDI prediction. In these models, the relevance of different drug features and interactions is captured through the assignment of adaptive weights to various aspects of the data, resulting in predictions that are both more accurate and interpretable. These techniques are considered to provide complementary insights when combined with graph-based methods, thereby contributing to the advancement of DDI prediction and the improvement of therapeutic decision-making processes.
Method based on attention mechanisms and siamese networks
Faced with the challenges of limited data and high complexity in predicting drug interactions, attention mechanisms and siamese networks based on deep learning models have emerged. These methods demonstrate great potential in extracting effective features and efficiently modeling multimodal drug data. Among them, the attention mechanism significantly improves prediction accuracy by dynamically focusing on key drug attributes and dynamically adjusting feature importance125. Twin networks, on the other hand, provide an ideal framework for identifying interactions between drug combinations through feature contrastive learning. The overall flowchart of DDI prediction using attention mechanisms and siamese networks is summarized in Fig. 6.
a Attention mechanisms diagram. b Siamese networks diagram.
To overcome this limitation, Huang et al.126 proposed a multi-view and multichannel attention deep learning (MMADL) model. MMADL not only extracts rich drug features—including both intrinsic drug attributes and drug-related entity information from multiple databases—but also considers the consistency and complementarity of various drug feature representations to enhance DDI prediction. A single-layer perceptron encoder is used to project multi-source drug information into a common linear space, yielding multi-view drug representation vectors. Subsequently, a multichannel attention mechanism is introduced to adaptively learn the contribution of each feature, generating attention weights that guide the final prediction. These weighted representation vectors are then fed into a deep neural network to predict potential DDIs. Experimental results demonstrated that MMADL effectively integrates multi-source drug information, improving both the accuracy and robustness of DDI prediction. Future work could focus on validating the generalization ability of the model across broader drug datasets to enhance its applicability.
In addition, Huang et al.127 presents a novel method, MDF-SA-DDI, for predicting DDI events using multi-source drug fusion, multi-source feature fusion, and a transformer self-attention mechanism. The approach combines two drugs in four different ways and utilizes various fusion networks, including Siamese networks, CNNs, and two autoencoders, to obtain latent feature vectors for drug pairs. These feature vectors are then fused using transformer blocks with a self-attention mechanism. The method is evaluated on two datasets across three tasks, demonstrating superior performance compared to existing state-of-the-art methods, achieving high precision and recall scores. Additionally, the case studies on specific DDI events also show promising results.
Given the impracticality of experimentally characterizing the vast number of potential DDIs and the superior performance of deep learning on large-scale, high-dimensional datasets128,129, computational approaches based on deep learning are increasingly recognized as essential. In this context, Yang et al.130 introduced CNN-Siam, a deep learning-based model employing a Siamese CNN architecture to learn feature representations from multimodal drug data—including chemical substructures, targets, and enzymes—and predict interaction types. The CNN-Siam model utilizes a dual-network structure, enabling the simultaneous extraction and comparison of drug feature representations, optimized using advanced optimization algorithms such as RAdam and LookAhead.
Experimental evaluations demonstrated that CNN-Siam significantly improves precision–recall (AUPR) and overall prediction accuracy on benchmark datasets, outperforming existing state-of-the-art methods. Ablation studies further confirmed the robustness of CNN-Siam and the effectiveness of the adopted optimizers. The Siamese network design proved particularly advantageous for learning drug pair features compared to traditional single-network architectures. However, CNN-Siam also exhibits certain limitations, such as prolonged training times, limited generalization capabilities in some scenarios, and suboptimal classification performance for specific interaction types. Future improvements could focus on accelerating training, enhancing model generalization, and refining class-specific prediction accuracy.
Recent advances for DDI prediction
In recent years, the field of DDI prediction has witnessed significant advancements, particularly with the advent of large models and diffusion-like generative models. These models have introduced novel approaches for capturing complex patterns and relationships within vast amounts of multimodal data, such as chemical structures, genomic information, and clinical data131. These innovations not only improve prediction accuracy but also address challenges related to data sparsity and the inherent complexity of DDI networks. As these models continue to evolve, they offer great potential for revolutionizing drug discovery, helping to identify safer and more effective drug combinations and reducing the risk of adverse drug reactions in clinical settings132.
The following section will explore how these advanced AI techniques—Large Language Models in DDI and Diffusion-like Generative Models in DDI—are revolutionizing the field of drug interaction prediction. Through the integration of these innovative approaches, the future of DDI prediction looks increasingly promising, with the potential for more personalized and precise medicine.
Large language models in DDI
Large language models (LLMs) have emerged as powerful tools in DDI prediction due to their ability to process and understand vast amounts of unstructured data. These models, originally developed for natural language processing tasks, are now being adapted to the biomedical domain to analyze textual data from scientific literature, clinical notes, drug databases, and other health-related sources133. By fine-tuning LLMs on specialized datasets related to drug interactions, researchers can harness their ability to identify subtle patterns and relationships between drugs that might be overlooked by traditional methods. LLMs can be particularly effective in extracting knowledge from clinical case reports, drug labels, and scientific papers, providing insights into potential DDIs that are not readily available in structured databases134. The ability of LLMs to handle large-scale, multimodal data also allows for more comprehensive and accurate predictions, ultimately contributing to better-informed drug development and safer treatment regimens. As LLMs continue to evolve, their integration with other ML techniques holds the potential to transform the landscape of DDI prediction and drug safety monitoring.
Accurate prediction of DDIs is essential to ensure the safety and efficacy of drugs. Therefore, Li et al.135 introduces a novel model called LLM-DDI, designed to enhance DDI prediction using a combination of LLMs and GNNs. LLM-DDI integrates generative pre-trained transformer (GPT) embeddings to capture diverse molecular information from the biomedical knowledge graph (BKG), then applies a message-passing GNN framework to propagate and refine these embeddings. This approach effectively incorporates both local and global molecular features, leveraging semantic relationships within the BKG for accurate DDI predictions. The model aims to overcome limitations in current DDI prediction methods by offering a comprehensive framework that integrates multiple molecular data types, achieving significant improvements in prediction performance on real-world datasets.
With the rapid growth of data in the field of biomedicine, accurate prediction of DDI is crucial to ensure the safety and effectiveness of drugs. However, traditional prediction methods face certain limitations when dealing with complex biomedical data. In order to solve this challenge, Qi et al.136 proposes a novel model called DDI-JUDGE for DDI prediction, which leverages LLMs enhanced with in-context learning (ICL) and a judging mechanism. This model integrates cosine similarity-based exemplar retrieval for effective ICL and combines multiple LLM predictions through an ensemble discriminator, such as GPT-4, to improve robustness. The DDI-JUDGE model outperforms existing methods in both zero-shot and few-shot settings, demonstrating superior predictive capabilities. It incorporates a modular prompt structure that facilitates better generalization and adaptation to complex biomedical data, offering a scalable approach for DDI prediction and other knowledge-intensive biomedical applications.
Diffusion-like generative models in DDI
Diffusion-like generative models in DDI prediction aim to model the complex relationships between different drugs and their potential interactions. These models leverage the principles of diffusion processes, where information spreads through a network or graph structure, capturing how drug properties and their effects propagate and influence one another137. By employing these models, it is possible to simulate and predict how two or more drugs might interact based on their molecular features, known pharmacological effects, and existing interactions. This approach has gained significant attention due to its ability to handle large, sparse datasets typical in DDI research and its potential to uncover novel, previously unknown drug interactions. Diffusion-like models can be particularly useful in drug repurposing, safety profiling, and personalized medicine, offering a promising tool for enhancing drug development and therapeutic decision-making. As a result, diffusion-like generative models are not only advancing the state of DDI prediction but also paving the way for more efficient and precise drug safety monitoring, clinical decision-making, molecule design, and the identification of new therapeutic avenues138.
Benchmark datasets and evaluation metrics of machine learning models
The following section will explore the importance of benchmark datasets in evaluating ML models for DDI prediction and in-depth comparison of ML models in the field of DDIs described in this article.
Benchmark datasets
In the field of DDI prediction, benchmark datasets are essential for evaluating the performance of ML models. These datasets contain a variety of drug interaction information and serve as the foundation for training and testing models. They are usually curated from diverse sources, including scientific literature, clinical trials, and PC databases139. The use of standardized benchmark datasets allows for the comparison of different ML algorithms under consistent conditions, providing valuable insights into their strengths and limitations140.
Several biomedical databases are commonly used in DDI prediction tasks, each offering unique features and data structures that cater to specific aspects of drug interaction research141,142,14. These datasets vary in terms of the type of drug interaction data they provide, the size of the dataset, and the methods used for data curation. Some of the most widely used biomedical databases for DDI prediction tasks are systematically summarized in Table 2, with their contents and characteristics being comprehensively documented to serve as a selection resource for researchers seeking appropriate data sources.
When leveraging these benchmark datasets, it is critical to address the inherent challenges of data noise and imbalance, which can significantly affect the performance of ML models, and they can lead to reduced accuracy and generalization capabilities of the models143. Data noise refers to errors or inconsistencies in the data that may be caused by experimental bias, literature labeling errors, or inaccuracies in the data collection process, among other reasons. In DDI prediction, the complexity and uncertainty of drug interactions may cause some interaction relationships to present inconsistent labeling in different sources, which threatens the quality of training data. Noisy data makes the patterns learned by the model fuzzy, thus affecting the effectiveness of the model, especially when processing new samples, noisy data may cause the model to make wrong predictions144.
On the other hand, data imbalance is another key factor affecting ML models, especially in drug interaction prediction tasks145. There is no interaction between most drugs, while only a small number of drug pairs have potential interactions. The data imbalance is manifested by a severe imbalance in the proportion of positive and negative samples, where negative samples (i.e., drug pairs without interaction) are usually far more than positive samples (drug pairs with interaction). In this case, traditional ML algorithms may be biased towards predicting negative samples, because most of their data belongs to this class, resulting in the model’s insufficient learning of a few classes (positive samples)146. This bias not only reduces the model’s ability to recognize rare drug interactions, but may also lead to false negative results, that is, the prediction that there is an interaction between drugs is missed.
Solving data noise and imbalance problems usually requires the adoption of some special techniques and strategies. In terms of data noise, commonly used methods include data cleaning, data augmentation, and noise robust algorithms147. These methods can help remove or reduce the impact of noisy data on model training. For example, by using statistical based methods to identify and eliminate obvious outliers, or by enhancing the data (such as generating more samples) to compensate for the bias caused by noise. Common solutions to imbalanced data problems include oversampling, undersampling, and using weighted loss functions. The oversampling method balances the dataset by increasing the number of positive samples, while the undersampling method reduces the number of negative samples. The weighted loss function enhances the model’s focus on minority class samples by assigning different weights to samples of different classes, thereby improving the predictive ability of rare drug interactions. Overall, data noise and imbalance are key issues that need to be addressed in ML, especially in predicting drug drug interactions. Effective preprocessing techniques and model optimization strategies can significantly improve prediction accuracy148, especially when dealing with complex and rare drug interactions.
Evaluation metrics of machine learning models
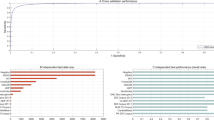
The predictive performance of different DDI prediction models across three key metrics (AUPR, AUC, and F1-score) is systematically evaluated in Table 3. Through a comparative analysis of the performance data of the DDI prediction models introduced in this paper, we can clearly observe the differences in performance and the respective strengths of various methodological categories. GNN-based methods demonstrate the most outstanding overall performance, with ACDGNN and SGFNNs achieving high AUPR scores of 0.9881, while AutoDDI reaches an even more impressive AUPR of 0.9952 and AUC of 0.9953. This fully showcases the powerful capabilities of deep learning in handling complex DDI relationships. In contrast, traditional similarity-based and matrix factorization methods, although still competitive in certain scenarios—such as GRPMF achieving an F1-score of 0.9622—generally exhibit lower AUPR values, reflecting their limitations in balancing precision and recall. Notably, models based on attention mechanisms and Siamese networks, such as MMADL and MDF-SA-DDI, excel in AUC metrics, reaching 0.9980 and 0.9996, respectively, highlighting the advantages of these architectures in overall ranking performance.
From the perspective of methodological advancements, improvements in model performance show a clear positive correlation with architectural innovations. Early classification- and similarity-based methods, such as Link prediction and Similarity-based machine, achieved AUC values of 0.93 and 0.97 but performed modestly in AUPR, indicating their shortcomings in handling class-imbalanced data. With the introduction of matrix factorization methods, such as ISCMF and RaGSEs, AUPR was elevated to 0.864. However, the true breakthrough came with the application of GNNs. SSI-DDI demonstrated the immense potential of structural feature modeling with an AUPR of 0.999 and AUC of 0.9986, while AutoDDI further pushed performance to new heights through the combination of autoencoders and transfer learning. These technological advancements are not only reflected in numerical improvements but also signify a paradigm shift from shallow feature matching to deep relational modeling.
Emerging LLMs approaches, such as LLM-DDI and DDI-JUDGE, have yet to reach the top-tier performance of traditional methods but have already shown unique developmental potential. LLM-DDI, with an AUC of 0.9571 and F1-score of 0.8542, demonstrates the feasibility of language models in DDI prediction. The emergence of these LLM-based methods marks a transition in DDI prediction research from purely data-driven approaches to a new phase combining knowledge guidance with data-driven strategies. The key challenge for future development lies in better integrating the semantic understanding capabilities of language models with domain-specific knowledge while addressing the current shortcomings in AUPR performance. This may require more refined prompt engineering designs and more effective domain knowledge integration strategies.
Clinical application of drug-drug interactions
In clinical treatment, DDIs are critical factors that influence both patient safety and therapeutic efficacy149,150. With the advancement of modern medicine and therapeutic strategies—particularly the widespread adoption of multi-drug combination therapies151—the complexity of DDIs has markedly increased, presenting substantial challenges for physicians in prescription decision-making and for ensuring patient medication safety.
In recent years, AI and ML technologies have introduced innovative approaches for DDI prediction152,153. By analyzing large volumes of drug-related data154, clinical case reports, patient-specific variables, and pharmacological characteristics, these technologies are capable of extracting potential interaction patterns and predicting DDIs with greater accuracy. Compared to traditional experience-based or rule-based methods, data-driven predictive models offer enhanced precision, scalability, and adaptability, thereby providing clinicians with more scientific, real-time decision support.
Importantly, in practical clinical applications, ML models not only assist in identifying well-known DDIs but also uncover novel and previously underreported interaction risks155,156. This capacity significantly enhances the clinician’s ability to anticipate and prevent adverse drug reactions, contributing to improved patient outcomes and medication safety.
The following section will delve into the specific manifestations of DDIs in clinical practice and discuss how advanced ML methods can be leveraged for more accurate DDI prediction and management. In addition, several typical case examples of drug interactions will be presented to illustrate how AI technologies contribute to clinical decision-making, optimize therapeutic efficacy, and safeguard patient health.
Pharmacokinetic interaction
In clinical practice, DDIs are frequently manifested through various mechanisms, with PK interactions being among the most critical157,158. PK refers to the movement of drugs within the body, encompassing absorption, distribution, metabolism, and excretion (ADME) processes159,160. Each stage of the PK pathway plays a pivotal role in determining the efficacy and safety of therapeutic agents161,162.
When two or more drugs are co-administered, they may alter the normal PK processes, leading to changes in drug concentrations that can cause adverse reactions or therapeutic failure163,164. A thorough understanding of PK interaction mechanisms is therefore essential for clinicians to anticipate, manage, and mitigate potential risks associated with polypharmacy165,166,167.
In recent years, the application of advanced ML techniques has enabled more accurate prediction and identification of PK-related DDIs, offering valuable insights into how drugs may influence each other’s PK behavior168.
The following section will focus on the clinical implications of DDIs occurring at different stages of the PK process (ADME). It will examine how drug interactions at each stage affect patient safety and therapeutic outcomes. By exploring these mechanisms, a deeper understanding can be gained of the intricate nature of DDIs and how ML-based approaches can facilitate their management. Furthermore, the clinical applications of DDIs in PK—including the action mechanisms of specific drugs at each ADME stage—are summarized in Fig. 7, providing a visual overview to enhance understanding.
A The interaction diagram of antacids (such as aluminum hydroxide) and ketoconazole in the absorption stage. B The interaction diagram of quinidine and digoxin in the distribution stage. C The interaction diagram of statins (such as Rosuvastatin, Pravastatin or Fluvastatin) and antibiotic clarithromycin in the metabolic stage. D The interaction diagram of Probenecid and penicillin in excretion stage.(Created with bioicons.com, https://smart.servier.com/ is licensed under CC-BY 3.0 Unported https://creativecommons.org/licenses/by/3.0/).
Absorption stage
DDIs at the absorption stage are often mediated by alterations in drug solubility, which directly affect the absorption efficiency. A classic example involves the interaction between antacids (e.g., aluminum hydroxide) and drugs such as ketoconazole169,170. Ketoconazole, a weakly basic drug with a pKa of ~3.0, exhibits higher solubility in acidic environments, such as gastric juice, and significantly reduced solubility when the pH exceeds 4.0.
When antacids are administered, they raise the gastric pH above 4.0, resulting in incomplete dissolution of ketoconazole in the stomach. Consequently, the drug may not fully dissolve before reaching the small intestine, leading to impaired absorption—a phenomenon known as dissolution-limited absorption. Studies have demonstrated that co-administration of antacids with ketoconazole can significantly reduce its area under the concentration–time curve (AUC), with reductions of up to 66%, thereby slowing both the rate and extent of drug absorption110.
Thus, by altering gastric pH and drug solubility, antacids can markedly decrease the absorption of drugs, particularly those with pH-dependent solubility profiles171. Conversely, some weakly acidic drugs, which exhibit poor solubility in acidic conditions but improved solubility at higher pH levels, may experience enhanced absorption when co-administered with antacids.
Therefore, in clinical practice, careful consideration should be given to potential interactions between antacids and other medications to prevent compromised therapeutic efficacy.
Distribution stage
DDIs at the distribution stage are exemplified by the interaction between quinidine and digoxin, which primarily affects both the distribution and clearance of digoxin172,173. Quinidine administration leads to an increase in serum digoxin concentration by reducing its volume of distribution, a phenomenon observed in both healthy volunteers and patients174. A reduced volume of distribution implies that the drug occupies a smaller distribution space within the body, resulting in elevated plasma concentrations.
However, the elevation in plasma digoxin levels is not solely due to changes in distribution volume. Quinidine also impairs digoxin clearance, particularly by inhibiting its non-renal elimination pathways, such as metabolism and biliary excretion175. The interaction between quinidine and digoxin significantly decreases the total body clearance of digoxin, with a pronounced reduction in non-renal clearance mechanisms. This effect is especially notable in patients with renal insufficiency, who rely more heavily on non-renal routes for digoxin elimination176.
Thus, although quinidine decreases the distribution volume of digoxin, the predominant clinical mechanism involves inhibition of digoxin clearance rather than merely altering distribution177. Overall, quinidine increases digoxin serum concentrations primarily by reducing its non-renal clearance, thereby elevating the risk of digoxin toxicity178. Consequently, therapeutic drug monitoring and dose adjustment of digoxin are critical during co-administration with quinidine to ensure patient safety.
Metabolic stage
DDIs at the metabolism stage are critical contributors to adverse clinical outcomes. To further investigate the safety concerns associated with drug combinations, Li et al.179 conducted a large cohort study to assess whether co-administration of statins (such as rosuvastatin, pravastatin, or fluvastatin) and the antibiotic clarithromycin increases the risk of serious adverse events in the elderly population. Clarithromycin is a well-known inhibitor of cytochrome P450 3A4 (CYP3A4) and may also inhibit hepatic organic anion transporting polypeptides (OATP1B1 and OATP1B3).
Using a large health database, the researchers analyzed data from elderly patients between 2002 and 2013, comparing those who received both clarithromycin and statins (n = 51,523) with those who received azithromycin, an antibiotic that does not inhibit CYP3A4 or OATP transporters (n = 52,518). The results showed that, compared with the control group, patients co-treated with clarithromycin and statins exhibited significantly increased risks of hospitalization within 30 days, specifically for acute kidney injury (adjusted relative risk (RR = 1.65), hyperkalemia (RR = 2.17), and all-cause mortality (RR = 1.43)). Additionally, the risk of hospitalization for myolysis (myoglobinuria) was modestly elevated (RR = 2.27). Although the absolute risk increases were relatively small (generally less than 1%), the study demonstrated a clear association between clarithromycin–statin co-administration and an elevated risk of adverse outcomes, particularly in elderly patients.
Mechanistically, as an inhibitor of CYP3A4, clarithromycin affects not only the cytochrome-mediated metabolic pathways but also the hepatic uptake of statins via inhibition of OATP1B1 and OATP1B3 transporters. Although the statins investigated (e.g., rosuvastatin and pravastatin) are not primarily metabolized by CYP3A4, clarithromycin may still alter their PKs by affecting OATP-mediated transport, leading to impaired distribution, excretion, or metabolism of the drugs. Consequently, this interaction increases the risk of adverse events such as acute kidney injury and hyperkalemia.
These findings highlight that drug interactions are not confined to classic metabolic pathways (such as CYP3A4 inhibition) but can also occur through transporter-mediated mechanisms. Therefore, when prescribing clarithromycin to elderly patients taking statins, clinicians should exercise heightened vigilance for potential adverse reactions related to kidney function and electrolyte disturbances.
Excretion stage
DDIs affecting the excretion stage play a crucial role in altering drug PKs and clinical outcomes. A classic example is the combined use of probenecid and penicillin, where probenecid significantly impacts the excretion of penicillin by inhibiting its renal tubular secretion, thereby prolonging its half-life in vivo180. Specifically, probenecid competes with penicillin for the organic anion transport system in the renal tubules, leading to a marked reduction in the renal clearance of penicillin181.
Under normal physiological conditions, penicillin is predominantly excreted through active secretion in the renal tubules. However, probenecid effectively blocks this excretory pathway, resulting in a significant decrease in tubular excretion, leaving glomerular filtration as the primary route for penicillin elimination. In addition to affecting renal clearance, probenecid also inhibits the active transport of penicillin across the choroid plexus, thereby increasing its concentration in the cerebrospinal fluid182. Through these mechanisms, probenecid enhances the systemic exposure and prolongs the efficacy of penicillin, which is particularly beneficial in infections such as meningitis where sustained high drug concentrations are required183.
There is also evidence suggesting that probenecid may slightly increase the serum concentration of penicillin by reducing its volume of distribution, although this effect is less clinically significant. It is important to recognize that while prolonging the half-life of penicillin can enhance therapeutic efficacy, especially for β-lactam antibiotics, it may also increase the risk of adverse drug reactions if not properly monitored184. Moreover, the use of probenecid is not universally beneficial across all drugs. For instance, in the case of nitrofurantoin, which relies heavily on renal tubular secretion for elimination, co-administration with probenecid can reduce therapeutic effectiveness and heighten the risk of side effects185.
In summary, the co-administration of probenecid and penicillin mainly enhances drug efficacy by inhibiting renal tubular secretion, thereby extending the duration of effective drug concentrations in the body. However, careful evaluation is required to balance the therapeutic advantages against the potential for adverse effects, particularly when applied to other medications.
Pharmacodynamic interaction
Following PK interactions, another critical aspect of DDIs is PD interactions186. PDs refers to the effects of drugs on the body, including how therapeutic actions are produced through mechanisms such as receptor binding, enzyme inhibition, or other biological processes187. When two or more drugs interact at the PD level, they can either enhance or diminish each other’s effects, potentially modifying the expected therapeutic outcomes188. Such interactions can profoundly impact efficacy, safety profiles, and overall patient management.
PD interactions are generally categorized into two types: synergistic and antagonistic effects. A synergistic interaction occurs when the combined effect of two drugs exceeds the sum of their individual effects, potentially enhancing therapeutic efficacy189. In contrast, antagonistic interactions arise when one drug reduces or inhibits the effect of another, which may compromise therapeutic goals or even lead to treatment failure.
The following sections will explore these two types of PD interactions in greater detail. A comprehensive understanding of the clinical applications of synergy and antagonism can significantly aid in predicting, managing, and preventing potential risks associated with drug combinations. Moreover, the use of advanced ML techniques holds promise for the early identification of PD interactions, thereby improving clinical decision-making and enhancing patient safety. The mechanisms of PD interactions are systematically illustrated in Fig. 8, providing an intuitive framework for understanding drug interaction pathways in therapeutic contexts190.
A The interaction diagram of aspirin and P2Y12 inhibitors such as clopidogrel, prasugrel or ticagrelor in the pathway synergistic. B The interaction diagram of naloxone and opioid drugs (such as morphine) in the competitive antagonism. C The interaction diagram of β-Blockers and calcium channel blockers in dual-effect interaction. D The interaction diagram of diuretic and ACE inhibitor in complementary interaction. (Created with bioicons.com, https://smart.servier.com/ is licensed under CC-BY 3.0 Unported https://creativecommons.org/licenses/by/3.0/).
Pathway synergism: dual antiplatelet therapy
Building on previous research in antiplatelet therapy, Sinnaeve et al.191 explored strategies for dual antiplatelet therapy (DAPT) following percutaneous coronary intervention or acute coronary syndrome (ACS), with particular focus on de-escalation strategies. The study discussed the conventional use of standard DAPT—combining aspirin with a P2Y12 inhibitor (such as clopidogrel, prasugrel, or ticagrelor)—but noted that prolonged DAPT is associated with an increased risk of bleeding, especially in the era of newer-generation stents that have reduced the risk of stent thrombosis.
Several DAPT de-escalation strategies have been proposed: one approach involves switching from potent P2Y12 inhibitors (such as ticagrelor or prasugrel) to a less potent agent like clopidogrel while continuing aspirin; another involves monotherapy, discontinuing either aspirin or the P2Y12 inhibitor. Multiple clinical trials have demonstrated that these de-escalation strategies can effectively reduce bleeding risk without significantly increasing the incidence of ischemic events. The study found that switching from potent P2Y12 inhibitors to clopidogrel significantly reduced cardiovascular death, emergency revascularization, stroke, and major bleeding events within one year, with no substantial difference in ischemic outcomes.
Furthermore, trials such as TROPICAL-ACS and POPular Genetics tested platelet function- or genotype-guided de-escalation strategies. Their results indicated that using genetic testing and platelet function assays to tailor antiplatelet therapy could further optimize treatment by reducing bleeding risks, although the impact on ischemic events was minimal.
Overall, this evidence highlights that DDIs play a critical role in the synergistic effects observed in PDs, particularly in optimizing antiplatelet therapy strategies. Specifically, the use of P2Y12 inhibitors (clopidogrel, prasugrel, and ticagrelor) allows for individualized treatment adjustment based on patient-specific risk profiles, utilizing genetic testing or platelet function testing. Strategic DDI management and dosage modifications—such as switching from more potent to less potent P2Y12 inhibitors—can effectively minimize bleeding risks while preserving antithrombotic efficacy.
Competitive antagonism: opioid receptor cases
Naloxone, an opioid antagonist, is widely employed to investigate the role of endogenous opioids in various physiological and pharmacological processes192. Its primary mechanism involves antagonizing the effects of opioid drugs, such as morphine, by competitively inhibiting opioid receptors. Research has demonstrated that naloxone not only counteracts the analgesic and other pharmacological effects of morphine193, but in certain circumstances, it also exhibits agonistic effects similar to those of opioids. The antagonistic effects of naloxone typically require a higher dose194, which is attributed to the diversity of opioid receptors, as different receptor subtypes exhibit varying sensitivities to naloxone.
Moreover, naloxone has been shown to produce similar effects to morphine in some behavioral and in vitro experimental models, even demonstrating comparable analgesic effects. Notably, naloxone may also influence non-opioid drugs in certain cases, likely due to interactions with endogenous opioid systems or mechanisms that are unrelated to opioid receptors195. Although naloxone is extensively used in the study of opioid receptors, its complex pharmacological profile suggests that the antagonism it induces is not sufficient on its own to definitively establish the involvement of endogenous opioids in a given physiological process.
To clarify the role of endogenous opioids, additional evidence is required, such as cross-resistance to morphine, similar responses to other opioid antagonists, and the combined use of endogenous opioid peptide degradation inhibitors196. These complementary approaches can provide a more comprehensive understanding of the mechanisms underlying the involvement of endogenous opioids in specific pharmacological processes.
Dual-effect interaction: β-blockers + calcium channel blockers
β-Blockers and calcium channel blockers (CCBs) are both commonly used in the management of hypertension, but their PD interactions are of particular interest when considering the co-administration of these drugs197,198. Both drug classes can affect the heart and vascular tone, but they do so through different mechanisms. β-Blockers work by blocking β-adrenergic receptors, thereby reducing heart rate, myocardial contractility, and the release of renin, which collectively decrease blood pressure.
In contrast, CCBs inhibit the influx of calcium ions through L-type calcium channels, which leads to vasodilation, reduced myocardial contractility, and decreased heart rate. When these two classes are combined, there is a potential for synergistic effects on heart rate and blood pressure. However, the combination may also cause adverse effects, especially in patients with pre-existing heart conditions. For instance, while β-blockers decrease the heart rate, CCBs can also have a similar effect, potentially leading to bradycardia, which in severe cases could result in heart block or other cardiac arrhythmias.
Additionally, there is a concern about hypotension, as both medications can lower blood pressure, potentially leading to dizziness, fainting, or even shock in some patients. Despite these risks, in certain clinical situations, such as in the treatment of angina or heart failure, this combination may be beneficial as it can provide complementary effects, improving both symptoms and outcomes. However, careful monitoring of heart rate, blood pressure, and ECG is essential to prevent any adverse cardiovascular events.
Complementary interaction: diuretic and ACE inhibitor synergy in therapy
Diuretics and angiotensin-converting enzyme (ACE) inhibitors are both widely used in the treatment of hypertension and heart failure, but their PD interactions can have significant implications when used together199. Diuretics work by increasing urine output, thereby reducing blood volume and decreasing blood pressure. They primarily target the kidneys, promoting the excretion of sodium and water, which helps to lower blood pressure and reduce fluid buildup in the body, making them especially effective in conditions like hypertension and edema200.
On the other hand, ACE inhibitors block the ACE, which plays a critical role in the renin-angiotensin-aldosterone system (RAAS). By inhibiting this enzyme, ACE inhibitors reduce the production of angiotensin II, a potent vasoconstrictor, leading to vasodilation and a subsequent reduction in blood pressure. ACE inhibitors also reduce aldosterone secretion, which helps prevent sodium and water retention, further contributing to blood pressure lowering.
When diuretics and ACE inhibitors are used together, their effects on the RAAS and fluid balance can complement each other. Diuretics can enhance the blood pressure-lowering effects of ACE inhibitors by reducing fluid volume, while ACE inhibitors prevent compensatory mechanisms, such as increased aldosterone secretion, which could counteract the diuretic’s effect. However, this combination also carries a risk of hypovolemia and electrolyte imbalances, particularly potassium. Diuretics can cause potassium loss, whereas ACE inhibitors can lead to potassium retention, potentially resulting in hyperkalemia. Close monitoring of electrolyte levels and kidney function is crucial when these drugs are co-administered.
Moreover, in patients with renal impairment, the combination of diuretics and ACE inhibitors can lead to further renal stress, increasing the risk of acute kidney injury. Therefore, while this combination is effective in managing hypertension and heart failure, it requires careful dosing and monitoring to avoid complications like dehydration, electrolyte disturbances, or kidney dysfunction.
Pharmaceutical interactions
The combination of carbenicillin and gentamicin is commonly used in clinical practice to enhance antibacterial efficacy201, particularly in the treatment of severe bacterial infections. This combination therapy is especially effective against Gram-negative bacteria, such as Pseudomonas aeruginosa202,203. The pharmacological mechanisms of carbenicillin and gentamicin differ significantly. Carbenicillin is a β-lactam antibiotic that works by inhibiting bacterial cell wall synthesis, leading to bacterial cell death204,205. Gentamicin, an aminoglycoside antibiotic, interferes with protein synthesis by binding to the 30S ribosomal subunit of bacteria, disrupting their normal physiological processes and causing bacterial death206,207. Since their mechanisms of action do not directly conflict, the two antibiotics can be used in combination to produce a synergistic antibacterial effect208. A flowchart summarizing the clinical application of DDIs in pharmaceutics is provided in Fig. 9A for a clearer understanding of the process.
A The interaction diagram of carbenicillin and carbenicillin in pharmaceutical interaction. B The interaction diagram of grapefruit in drug-food interaction. C The interaction diagram of MAOIs and sympathomimetic amines in absolute contraindications. D The interaction diagram of clarithromycin and amlodipine in relative contraindications. (Created with bioicons.com, https://smart.servier.com/ is licensed under CC-BY 3.0 Unported https://creativecommons.org/licenses/by/3.0/).
However, this combination therapy may also lead to potential drug interactions. First, carbenicillin can alter renal excretion mechanisms, resulting in increased blood concentrations of aminoglycosides like gentamicin, which may elevate the risk of nephrotoxicity209. Gentamicin can cause renal and cochlear toxicity at high concentrations, leading to renal impairment and hearing loss210,211. As a result, when carbenicillin and gentamicin are used together, it is essential to closely monitor renal function and blood drug concentrations to prevent excessive drug accumulation. Second, carbenicillin may compete with gentamicin for entry into the bacterial cell membrane. Although their mechanisms of action are distinct, this competition could affect drug penetration and the overall antibacterial efficacy, particularly at sites with high bacterial concentrations. Moreover, carbenicillin may enhance the sensitivity of bacteria to gentamicin by altering the structure of the bacterial cell wall, thus improving the synergistic effect of the combined therapy212.
In conclusion, while the combination of carbenicillin and gentamicin offers powerful antibacterial effects in clinical practice, it is crucial to be cautious of potential drug interactions, especially in patients with renal insufficiency. Careful use, along with monitoring drug concentrations and adverse reactions, is essential to ensure both the efficacy and safety of the treatment213.
Drug-food interaction
The interaction between grapefruit juice and drugs214,215 is primarily due to the chemical compound furanocoumarins found in grapefruit juice. These compounds are metabolized by the CYP3A4 enzyme, forming reactive intermediates that covalently bind to the enzyme’s active site, resulting in irreversible inhibition of the enzyme (mechanism-based inhibition). This inhibition reduces CYP3A4 activity in the small intestine until newly synthesized enzymes restore the activity to its original level. This process helps explain the significant effect of grapefruit on the PK of drugs, particularly the peak concentration (Cmax) and the area under the plasma concentration-time curve (AUC). These changes indicate an increase in the oral bioavailability of the drug, while the systemic elimination half-life remains unchanged216. A flowchart summarizing the clinical application of DDIs in drug-food science is provided in Fig. 9B to offer a clearer overview of the process.
It is important to note that this interaction is specific to orally administered drugs and does not significantly affect intravenous drugs. All forms of grapefruit, including freshly squeezed juice, frozen concentrated juice, and whole fruit, can have this effect by reducing the activity of CYP3A4. Even the consumption of one complete grapefruit or 200 ml of grapefruit juice is sufficient to cause a significant increase in systemic drug concentrations, potentially leading to adverse reactions217,218. In addition to grapefruit, Sevilla orange (commonly used in jams), lime, and pomelos can also trigger similar interactions, while sweet oranges (e.g., navel and Valencia oranges) do not contain furanocoumarins and thus do not cause this interaction219,220,221,222.
The interaction between grapefruit and drugs is drug-specific, and not all drugs are affected. Drugs most commonly affected are those with low to medium oral bioavailability (ranging from less than 10% to 70%) and those metabolized by CYP3A4196. Additionally, the timing and amount of grapefruit consumed can affect drug PKs. Typically, drug concentrations are most influenced within 4 h of grapefruit intake223. The susceptibility of patients to this interaction can also vary. For instance, individuals with high expression of CYP3A4 in their intestines may be more susceptible to grapefruit’s effects, and adjusting the drug dose may be necessary to achieve the desired therapeutic outcome. Therefore, in clinical practice, it is important to consider whether patients are at risk for grapefruit-related interactions based on these factors and adjust the treatment plan accordingly.
Pharmacological contraindications
Pharmacological contraindications refer to specific circumstances or conditions where the use of a particular medication could cause harm to the patient. These contraindications arise from potential interactions between the drug and the patient’s existing health conditions, other medications, or physiological states. Identifying and understanding these contraindications is essential for healthcare providers to make informed decisions about which drugs are appropriate for individual patients, ensuring both the efficacy of the treatment and patient safety224. Contraindications are generally classified into two categories: absolute and relative, each with distinct implications for patient care.
Absolute contraindications are situations where a particular drug should never be used under any circumstances because its administration would pose a significant risk to the patient’s health. These contraindications are considered critical and non-negotiable. For example, a drug that can cause severe, life-threatening allergic reactions should be absolutely contraindicated for patients with a known history of hypersensitivity to that drug225. Similarly, certain medications may be absolutely contraindicated in patients with specific medical conditions, such as a medication that should not be used in patients with a history of severe liver or kidney failure, as it could further damage these organs226. In these cases, there is no safe alternative for the patient, and using the drug would almost certainly result in harm.
Relative contraindications, on the other hand, refer to circumstances where the use of a drug is not automatically forbidden but should be approached with caution. These contraindications arise in situations where the benefits of the drug may outweigh the risks, but careful consideration and close monitoring are required. For instance, a medication that may cause certain side effects in patients with a pre-existing condition like high blood pressure might still be prescribed, but the patient must be carefully monitored throughout the treatment. In cases of relative contraindications, healthcare providers may adjust the dosage, monitor the patient more frequently, or choose a different treatment if the risks are deemed too high227. Ultimately, the decision is made on a case-by-case basis, considering the patient’s overall health and the therapeutic goals.
In both cases, understanding contraindications—whether absolute or relative—ensures that medications are prescribed safely and effectively, minimizing the risk of adverse outcomes. Furthermore, the clinical applications of pharmacological contraindications—including absolute contraindications and relative contraindications—are systematically illustrated in Fig. 9C, D, providing a visual framework to enhance understanding of these critical safety considerations.
Absolute contraindications: MAOIs + sympathomimetic amines
Monoamine oxidase inhibitors (MAOIs) are a class of drugs used primarily for the treatment of depression and certain other psychiatric disorders. They work by inhibiting the enzyme monoamine oxidase (MAO-A/B), which is responsible for breaking down neurotransmitters such as serotonin, norepinephrine, and dopamine228. However, when MAOIs are combined with sympathomimetic amines—drugs that stimulate the sympathetic nervous system—there is a significant risk of severe, life-threatening reactions. These include hypertensive crises and serotonin syndrome, both of which can be fatal if not promptly treated.
The interaction between MAOIs and sympathomimetic amines leads to a dangerous increase in the levels of neurotransmitters, particularly norepinephrine. This can trigger a hypertensive crisis, marked by a sudden and severe increase in blood pressure, which can lead to headache, sweating, palpitation, tachycardia, bradycardia, nausea, vomiting or dilated pupils. Additionally, the elevated serotonin levels can result in serotonin syndrome, a potentially fatal condition characterized by symptoms such as agitation, hyperthermia, tremors, sweating, and confusion. If left untreated, serotonin syndrome can lead to organ failure or death.
Due to the severity of these reactions, the combination of MAOIs and sympathomimetic amines is considered an absolute contraindication. Healthcare providers must carefully review a patient’s current medications to prevent such dangerous interactions. If a sympathomimetic drug is necessary, alternatives that do not interact with MAOIs should be considered to minimize the risk of these severe adverse reactions, including serotonin syndrome and hypertensive crises.
Relative contraindications: clarithromycin + amlodipine
The pharmacological interaction between clarithromycin and amlodipine primarily occurs through hepatic metabolism. Clarithromycin, as a potent inhibitor of the CYP3A4 isoenzyme, significantly reduces the metabolic clearance of amlodipine, a CCB that undergoes extensive CYP3A4-mediated oxidation229. This metabolic interference leads to elevated systemic exposure to amlodipine, with plasma concentrations increasing ~2–3 fold compared to amlodipine administration alone.
The altered PK profile manifests clinically through amplified PD effects of amlodipine. The drug’s characteristic vasodilatory action becomes exaggerated, resulting in prolonged reduction of peripheral vascular resistance. This manifests most notably as an increased incidence and severity of hypotension, particularly in elderly patients or those with compromised renal function. The interaction also extends amlodipine’s elimination half-life, potentially necessitating dosage adjustments to maintain therapeutic efficacy while minimizing adverse effects.
From a clinical management perspective, this interaction warrants consideration of alternative therapeutic strategies. Azithromycin presents a pharmacologically preferable macrolide option in patients requiring concomitant CCB therapy, as it lacks significant CYP3A4 inhibition properties. When clarithromycin use is unavoidable, close monitoring of blood pressure and renal function is essential, particularly in vulnerable populations such as the elderly or those with renal impairment.
Current challenges and future directions
This paper reviews the current research methods and model construction for DDI prediction. Although significant progress has been made in DDI prediction, demonstrating strong predictive capabilities, reliable results, and broad application prospects, several limitations and challenges remain that need to be addressed.
Optimization of modeling method
As drug development and clinical applications grow more complex, predicting DDIs becomes increasingly difficult and crucial. The occurrence of DDIs can lead to severe adverse reactions, potentially threatening patients’ lives. While existing ML models have addressed many of these challenges, several issues remain unresolved. For instance, traditional similarity indicators may be less effective when dealing with larger, more complex molecules. To address this, alternative or supplementary indicators should be explored, especially for macromolecules, which have not yet been fully examined.
Another challenge lies in extending the attention mechanism to multi-hop structures, which would allow for a more global determination of each entity’s acceptance domain. Additionally, the aggregator currently based on linear changes could be expanded to non-linear structures, enriching neighborhood representations and improving model performance.
Prediction accuracy is also compromised by imbalanced data and noise within the original datasets. To enhance the robustness of prediction models in such conditions, it is necessary to develop methods that reduce the impact of data imbalances and noise.
Moreover, variations in drug order during the occurrence of DDIs can affect model performance, even in the training phase. The substructure extraction phase also faces challenges due to noise information leakage, which can undermine the performance of the model under certain conditions, requiring further improvement.
In conclusion, optimizing modeling methods is a multifaceted challenge that encompasses aspects such as accuracy, efficiency, and interpretability. With the continuous development of algorithms and advancements in hardware technology, improving modeling techniques will remain a vital research focus in the future.
Enhancement of model interpretability
ML models, particularly those based on deep learning, have become essential tools in DDI prediction due to their impressive predictive capabilities and ability to handle large-scale, complex data. However, one of the significant challenges remains the “black box” nature of these deep learning models, where the decision-making process remains unclear. This lack of interpretability not only undermines the credibility of the model but also complicates the verification and pharmacological interpretation of drug interactions, particularly in contexts related to drug safety and clinical applications. To address this, researchers have developed interpretable AI techniques such as GNNExplainer and SHAP-GNN230,231, which provide insights into the decision-making process of GNNs by identifying critical subgraphs or quantifying feature importance through Shapley values. For instance, GNNExplainer generates explanations by highlighting the most influential subgraph structures and node features that contribute to a DDI prediction, while SHAP-GNN leverages game theory to attribute prediction outcomes to specific molecular features, enhancing transparency and trust in model outputs.
Over the past three years, several interpretable DDI prediction models have emerged, each with unique strengths and limitations. DABI-DDI integrates a dual-stage attention mechanism with LSTM networks and Bayesian calibration232, capturing temporal dependencies and reducing false positives. Its active learning strategy improves sample efficiency, but the model’s complexity may hinder real-time deployment. GGI-DDI employs granular computing to identify key molecular substructures responsible for DDIs233, offering biologically interpretable insights by aligning predictions with human cognition patterns. However, its reliance on predefined granulation strategies may limit adaptability to novel drug combinations. KnowDDI enhances interpretability by learning knowledge subgraphs from biomedical KGs, providing explaining paths for DDI predictions234. While it effectively compensates for sparse DDI data, its performance heavily depends on the quality of external knowledge sources, which may introduce biases. Lastly, MVA-DDI introduces a multi-view attention network that combines sequence and graph representations of drugs235, improving prediction accuracy through an interpretable self-attention mechanism. This approach allows clinicians to understand how different molecular perspectives contribute to DDI risks, but its dual-encoder design increases computational overhead. Collectively, these studies demonstrate progress in interpretable DDI prediction, yet challenges remain in balancing model complexity, real-time applicability, and domain-specific explainability. Future work should focus on developing hybrid models that integrate structural, temporal, and knowledge-based explanations while maintaining computational efficiency.
For DDI prediction, understanding the decision-making process is vital, as it directly impacts the safety evaluation and practical application of drug combinations. In the future, expanding research in this direction will enable DDI prediction models to not only provide accurate results but also offer greater transparency in terms of interpretation and validation. By enhancing interpretability, drug developers and clinicians will gain a clearer understanding of drug interactions, enabling them to identify potential risks early and make more informed, scientifically grounded treatment decisions. This, in turn, will improve the reliability of drug safety assessments and foster greater trust in clinical applications.
Multimodal learning and cross-domain integration
DDI prediction plays a crucial role in drug development and clinical practice, with the potential to identify risks associated with drug combinations and enhance drug safety. However, one of the major challenges in this field is the integration of cross-domain data. DDI prediction involves various types of data, including molecular structure, genomic data, clinical information, pharmacological data, and laboratory test results. These data sources often differ in formats, scales, and characteristics, making effective integration a significant technical hurdle in current research.
Traditional methods for predicting drug interactions typically rely on a single data source, which can limit the ability to capture the complexity of drug interactions. Multimodal learning presents a promising solution to this challenge. By integrating data from various domains, such as images, text, and tables, multimodal learning enables the model to understand the relationships between different types of data, thereby enhancing the ability to recognize drug interactions. This approach allows models to process and fuse diverse data types, enabling them to complement each other and mitigate the limitations of any single data source.
Moreover, cross-domain model integration offers another valuable approach for DDI prediction. By combining models from different domains—such as pharmacological, genomic, and clinical prediction models—a more robust and comprehensive prediction framework can be created. This integration improves the model’s adaptability and generalization across varied data environments. Cross-domain integration not only addresses the challenge of handling diverse data types but also overcomes the heterogeneity between data sources. Ultimately, through such integrated approaches, drug safety assessments and personalized treatment strategies can be more accurate, providing patients with safer and more effective treatment options.
Transformation from research to clinical application
The application of ML in DDI prediction heavily relies on a large volume of biological and chemical data. The quality, source, and standardization of these data are critical factors influencing the reliability of predictive models. However, many existing databases suffer from issues such as incomplete data, excessive noise, and inconsistent formats, which can undermine the accuracy and credibility of ML models in real-world applications. Moreover, predicting DDIs involves not only identifying simple interactions between drugs but also understanding complex physiological mechanisms, including factors such as drug metabolism, PKs, and PDs. These complexities pose significant challenges for achieving high prediction accuracy.
While ML models have demonstrated their potential in laboratory settings, translating them into clinical practice requires rigorous validation and standardized processes. This transformation is not only a technical challenge but also involves navigating ethical, legal, and regulatory concerns. Specifically, regulatory frameworks such as the European Union’s Artificial Intelligence Act (EU AI Act) and the U.S. Food and Drug Administration’s Software as a Medical Device (SaMD) guidelines provide critical compliance requirements for the development and deployment of ML-based medical tools. These frameworks emphasize the need for transparency, accountability, and risk management in AI systems used in healthcare, ensuring that they meet stringent safety and efficacy standards before clinical adoption.
In DDI prediction, the model must account for individual patient differences, such as genetic background, pathological conditions, and concurrent medications. These variables make the prediction task more complex and diverse, emphasizing the need for models that are not only accurate but also highly interpretable. Clinicians need to trust the model’s output and understand the reasoning behind its predictions to confidently integrate them into clinical decision-making. Additionally, data privacy is a paramount concern. The model’s reliance on sensitive patient information necessitates strict adherence to data protection laws, such as the General Data Protection Regulation in Europe, which mandates robust measures for data security, consent management, and patient rights.
Furthermore, in clinical settings, patient conditions and drug regimens can change rapidly, necessitating that DDI prediction systems respond quickly to these changes and update predictions in real time. This introduces a need for greater flexibility and dynamic adaptability in the models. Most current ML models are static, with their predictive abilities fixed once training is completed. Given the dynamic nature of clinical drug use, there is a pressing need for models to continuously learn and adapt to new data. This will require ongoing retraining and optimization, as well as integration of real-time data, which in turn demands significant technical resources and infrastructure.
Achieving this transition from research to clinical application will require interdisciplinary collaboration and strong policy and regulatory support. Compliance with established regulatory frameworks, such as the EU AI Act and FDA SaMD guidelines, alongside robust data privacy protections, will be essential to ensure the smooth and responsible implementation of predictive models in healthcare.
Conclusion
ML has become an indispensable tool in the prediction of DDIs, offering powerful capabilities to extract meaningful insights from increasingly complex and large-scale biomedical datasets. In this study, we summarized the major methodological advances, including graph-based models, deep learning frameworks, similarity-driven approaches, and multi-task learning, all of which have significantly improved the accuracy and efficiency of DDI prediction.
While these advances have broadened the horizon of DDI research, key challenges remain. Limitations in model interpretability, difficulties in integrating multimodal and cross-domain data, and barriers to clinical translation continue to constrain the full realization of ML-driven DDI prediction. Addressing these challenges will require the development of more transparent models, robust multimodal fusion techniques, and adaptive systems capable of dynamic learning in real-world clinical environments.
Looking ahead, the convergence of ML, systems pharmacology, and real-time clinical data holds the promise of reshaping how DDIs are predicted, validated, and managed. Future efforts should emphasize not only technical innovation but also interdisciplinary collaboration and regulatory alignment to bridge the gap between computational prediction and clinical application. As ML technologies continue to evolve, their integration into the drug discovery pipeline and clinical decision-making processes is poised to advance personalized medicine and enhance drug safety on a global scale.
Data availability
No datasets were generated or analyzed during the current study.
References
Edwards, I. R. & Aronson, J. K. Adverse drug reactions: definitions, diagnosis, and management. Lancet 356, 1255–1259 (2000).
Beijnen, J. H. & Schellens, J. H. M. Drug interactions in oncology. Lancet Oncol. 5, 489–496 (2004).
van Leeuwen, R. W. F. et al. Drug–drug interactions with tyrosine-kinase inhibitors: a clinical perspective. Lancet Oncol. 15, e315–e326 (2014).
Qato, D. M. et al. Changes in prescription and over-the-counter medication and dietary supplement use among older adults in the United States, 2005 vs 2011. JAMA Intern. Med. 176, 473–482 (2016).
Leape, L. L. et al. Systems analysis of adverse drug events. JAMA 274, 35–43 (1995).
Keith, C. T., Borisy, A. A. & Stockwell, B. R. Multicomponent therapeutics for networked systems. Nat. Rev. Drug Discov. 4, 71–78 (2005).
van Leeuwen, R. W. F. et al. Drug–drug interactions in patients treated for cancer: a prospective study on clinical interventions. Ann. Oncol. 26, 992–997 (2015).
Backman, J. T. et al. Role of cytochrome P450 2C8 in drug metabolism and interactions. Pharmacol. Rev. 68, 168–241 (2016).
Wienkers, L. C. & Heath, T. G. Predicting in vivo drug interactions from in vitro drug discovery data. Nat. Rev. Drug Discov. 4, 825–833 (2005).
Hisaka, A. et al. Prediction of pharmacokinetic drug–drug interaction caused by changes in cytochrome P450 activity using in vivo information. Pharmacol. Ther. 125, 230–248 (2010).
Langness, J. A. & Everson, G. T. Drug–drug interactions in HCV treatment—the good, the bad and the ugly. Nat. Rev. Gastroenterol. Hepatol. 13, 194–195 (2016).
Zhang, T., Leng, J. & Liu, Y. Deep learning for drug–drug interaction extraction from the literature: a review. Brief. Bioinform. 21, 1609–1627 (2020).
Juurlink, D. N. et al. Drug-drug interactions among elderly patients hospitalized for drug toxicity. JAMA 289, 1652–1658 (2003).
Tatonetti, N. P. et al. Data-driven prediction of drug effects and interactions. Sci. Transl. Med. 4, 125ra31 (2012).
Percha, B. & Altman, R. B. Informatics confronts drug–drug interactions. Trends Pharmacol. Sci. 34, 178–184 (2013).
Marzolini, C. et al. Prescribing nirmatrelvir–ritonavir: how to recognize and manage drug–drug interactions. Ann. Intern. Med. 175, 744–746 (2022).
Gallo, K. et al. PROMISCUOUS 2.0: a resource for drug-repositioning. Nucleic Acids Res. 49, D1373–D1380 (2021).
Ai, N., Fan, X. & Ekins, S. In silico methods for predicting drug–drug interactions with cytochrome P-450s, transporters and beyond. Adv. Drug Deliv. Rev. 86, 46–60 (2015).
Singh, S., Kaur, N. & Gehlot, A. Application of artificial intelligence in drug design: a review. Comput. Biol. Med. 179, 108810 (2024).
Wu, Z. et al. A comprehensive survey on graph neural networks. IEEE Trans. Neural Netw. Learn. Syst. 32, 4–24 (2020).
Joshi, A. K. Natural language processing. Science 253, 1242–1249 (1991).
Trajanov, D. et al. Review of natural language processing in pharmacology. Pharmacol. Rev. 75, 714–738 (2023).
Hirschberg, J. & Manning, C. D. Advances in natural language processing. Science 349, 261–266 (2015).
Li, Z. et al. A survey of convolutional neural networks: analysis, applications, and prospects. IEEE Trans. Neural Netw. Learn. Syst. 33, 6999–7019 (2021).
Tanoli, Z. et al. Computational drug repurposing: approaches, evaluation of in silico resources and case studies. Nat. Rev. Drug Discov. 24, 1–22 (2025).
Bess, A. et al. Artificial intelligence for the discovery of novel antimicrobial agents for emerging infectious diseases. Drug Discov. today 27, 1099–1107 (2022).
Schneider, P. et al. Rethinking drug design in the artificial intelligence era. Nat. Rev. drug Discov. 19, 353–364 (2020).
Yang, X. et al. Concepts of artificial intelligence for computer-assisted drug discovery. Chem. Rev. 119, 10520–10594 (2019).
Fleming, N. How artificial intelligence is changing drug discovery. Nature 557, S55–S55 (2018).
Schulman, A. et al. Attention-based approach to predict drug–target interactions across seven target superfamilies. Bioinformatics 40, btae496 (2024).
Chakraborty, S. et al. Artificial intelligence (AI) is paving the way for a critical role in drug discovery, drug design, and studying drug–drug interactions–correspondence. Int. J. Surg. 109, 3242–3244 (2023).
Kanehisa, M. et al. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 38, D355–D360 (2010).
Eloranta, S. & Boman, M. Predictive models for clinical decision making: Deep dives in practical machine learning. J. Intern. Med. 292, 278–295 (2022).
Thafar, M. A. et al. DTiGEMS+: drug–target interaction prediction using graph embedding, graph mining, and similarity-based techniques. J. Cheminform. 12, 1–17 (2020).
Hu, Y. et al. rs1990622 variant associates with Alzheimer’s disease and regulates TMEM106B expression in human brain tissues. BMC Med. 19, 1–10 (2021).
Vilar, S. et al. Similarity-based modeling in large-scale prediction of drug-drug interactions. Nat. Protoc. 9, 2147–2163 (2014).
Moscato, V. et al. Multi-task learning for few-shot biomedical relation extraction. Artif. Intell. Rev. 56, 13743–13763 (2023).
Zhang, W. et al. SFLLN: a sparse feature learning ensemble method with linear neighborhood regularization for predicting drug–drug interactions. Inf. Sci. 497, 189–201 (2019).
Zhu, J. et al. Attribute supervised probabilistic dependent matrix tri-factorization model for the prediction of adverse drug-drug interaction. IEEE J. Biomed. Health Inform. 25, 2820–2832 (2020).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun. ACM 60, 84–90 (2017).
Jang, H. Y. et al. Machine learning-based quantitative prediction of drug exposure in drug-drug interactions using drug label information. NPJ Digit. Med. 5, 88 (2022).
Gottlieb, A. et al. INDI: a computational framework for inferring drug interactions and their associated recommendations. Mol. Syst. Biol. 8, 592 (2012).
Staszak, M. et al. Machine learning in drug design: use of artificial intelligence to explore the chemical structure–biological activity relationship. Wiley Interdiscip. Rev. Comput. Mol. Sci. 12, e1568 (2022).
Jung, S. & Yoo, S. Interpretable prediction of drug-drug interactions via text embedding in biomedical literature. Comput. Biol. Med. 185, 109496 (2025).
Wang, N. N. et al. Comprehensive review of drug–drug interaction prediction based on machine learning: current status, challenges, and opportunities. J. Chem. Inf. Model. 64, 96–109 (2023).
Knox, C. et al. DrugBank 6.0: the DrugBank knowledgebase for 2024. Nucleic Acids Res. 52, D1265–D1275 (2024).
Law, V. et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 42, D1091–D1097 (2014).
Wishart, D. S. et al. DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 36, D901–D906 (2008).
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–D1213 (2016).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51, D1373–D1380 (2023).
Wang, Y. et al. PubChem’s BioAssay database. Nucleic Acids Res. 40, D400–D412 (2012).
Kim, S. et al. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res. 49, D1388–D1395 (2021).
Kanehisa, M. et al. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Kanehisa, M. Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M. et al. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677 (2025).
Zhang, P. et al. Label propagation prediction of drug-drug interactions based on clinical side effects. Sci. Rep. 5, 12339 (2015).
Xiong, G. et al. DDInter: an online drug–drug interaction database towards improving clinical decision-making and patient safety. Nucleic Acids Res. 50, D1200–D1207 (2022).
Tian, Y. et al. DDInter 2.0: an enhanced drug interaction resource with expanded data coverage, new interaction types, and improved user interface. Nucleic Acids Res. 53, D1356–D1362 (2025).
Yoshida, K., Maeda, K. & Sugiyama, Y. Transporter-mediated drug–drug interactions involving OATP substrates: predictions based on in vitro inhibition studies. Clin. Pharmacol. Ther. 91, 1053–1064 (2012).
Vilar, S., Friedman, C. & Hripcsak, G. Detection of drug–drug interactions through data mining studies using clinical sources, scientific literature and social media. Brief. Bioinform. 19, 863–877 (2018).
Huth, F. et al. Prediction of the impact of CYP2C9 genotypes on the drug-drug interaction potential of siponimod with PBPK modeling: a comprehensive approach for drug label recommendations. Clin. Pharmacol. Ther. 106, 1113–1124 (2019).
Wu H. Y., Chiang C. W. & Li L. Text mining for drug–drug interaction. Biomed. Lit. Min. 1159, 47-75 (2014).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic acids Res. 46, D1074–D1082 (2018).
Kumari, A., Akhtar, M., Shah, R. & Tanveer, M. Support matrix machine: a review. Neural Netw. 181, 106767 (2025).
Suthaharan, S. Support vector machine. Machine Learning Models and Algorithms for Big Data Classification: Thinking with Examples for Effective Learning 207–235 (Springer US, 2016).
Liu Y., Wang Y. & Zhang J. New machine learning algorithm: Random forest. International Conference on Information Computing And Applications 246–252 (Springer Berlin Heidelberg, Berlin, Heidelberg, 2012).
Zhang, Q. et al. Recent advances in convolutional neural network acceleration. Neurocomputing 323, 37–51 (2019).
Ruiz, L., Gama, F. & Ribeiro, A. Graph neural networks: architectures, stability, and transferability. Proc. IEEE 109, 660–682 (2021).
Cosgriff, C. V. & Celi, L. A. Deep learning for risk assessment: all about automatic feature extraction. Br. J. Anaesth. 124, 131–133 (2020).
Li, X. et al. Dual-view jointly learning improves personalized drug synergy prediction. Bioinformatics 40, btae604 (2024).
Lin, X. et al. Comprehensive evaluation of deep and graph learning on drug–drug interactions prediction. Brief. Bioinform. 24, bbad235 (2023).
Ren, Z. et al. Predicting rare drug-drug interaction events with dual-granular structure-adaptive and pair variational representation. Nat. Commun. 16, 3997 (2025).
Gao, S., Xie, J. & Zhao, Y. A multi-source drug combination and omnidirectional feature fusion approach for predicting drug-drug interaction events. J. Biomed. Inform. 162, 104772 (2025).
Bai, C. et al. Machine learning-enabled drug-induced toxicity prediction. Adv. Sci. 12, 2413405 (2025).
Wan, Z. et al. Applications of artificial intelligence in drug repurposing. Adv. Sci. 12, 2411325 (2025).
Deng, H. et al. MTMG: a multi-task model with multi-granularity information for drug-drug interaction extraction. Heliyon 9, e16819 (2023).
Zhu, J. et al. DAEM: Deep attributed embedding based multi-task learning for predicting adverse drug–drug interaction. Expert Syst. Appl. 215, 119312 (2023).
Salman, M. et al. Big data management in drug–drug interaction: a modern deep learning approach for smart healthcare. Big Data Cogn. Comput. 6, 30 (2022).
Huang, L. et al. Drug–drug similarity measure and its applications. Brief. Bioinform. 22, bbaa265 (2021).
Takeda, T. et al. Predicting drug–drug interactions through drug structural similarities and interaction networks incorporating pharmacokinetics and pharmacodynamics knowledge. J. Cheminform. 9, 1–9 (2017).
Kim, S. et al. Extracting drug–drug interactions from literature using a rich feature-based linear kernel approach. J. Biomed. Inform. 55, 23–30 (2015).
Bui, Q. C. et al. A novel feature-based approach to extract drug–drug interactions from biomedical text. Bioinformatics 30, 3365–3371 (2014).
Zhou, D., Miao, L. & He, Y. Position-aware deep multi-task learning for drug–drug interaction extraction. Artif. Intell. Med. 87, 1–8 (2018).
Wang, Z. et al. DEML: drug synergy and interaction prediction using ensemble-based multi-task learning. Molecules 28, 844 (2023).
Zurth, C. R. et al. Assessing the clinical relevance of drug-drug interactions (DDI) with darolutamide (DARO). Ann. Oncol. 30, v350 (2019).
Aapro, M. S. & Walko, C. M. Aprepitant: drug–drug interactions in perspective. Ann. Oncol. 21, 2316–2323 (2010).
Shi, J. Y. et al. An integrated local classification model of predicting drug-drug interactions via Dempster-Shafer theory of evidence. Sci. Rep. 8, 11829 (2018).
Kastrin, A., Ferk, P. & Leskošek, B. Predicting potential drug-drug interactions on topological and semantic similarity features using statistical learning. PLoS ONE 13, e0196865 (2018).
Cheng, F. & Zhao, Z. Machine learning-based prediction of drug–drug interactions by integrating drug phenotypic, therapeutic, chemical, and genomic properties. J. Am. Med. Inform. Assoc. 21, e278–e286 (2014).
Yan, X. Y. et al. Prediction of the drug–drug interaction types with the unified embedding features from drug similarity networks. Front. Pharmacol. 12, 794205 (2021).
Mar, P. L. et al. Drug interactions affecting oral anticoagulant use. Circ. Arrhythm. Electrophysiol. 15, e007956 (2022).
Song, D. et al. Similarity-based machine learning support vector machine predictor of drug-drug interactions with improved accuracies. J. Clin. Pharm. Ther. 44, 268–275 (2019).
Sridhar, D., Fakhraei, S. & Getoor, L. A probabilistic approach for collective similarity-based drug–drug interaction prediction. Bioinformatics 32, 3175–3182 (2016).
Yan, C. et al. Predicting drug-drug interactions based on integrated similarity and semi-supervised learning. IEEE/ACM Trans. Comput. Biol. Bioinform. 19, 168–179 (2020).
Yang, Y. H., Yang, J. T. & Liu, J. F. Lactylation prediction models based on protein sequence and structural feature fusion. Brief. Bioinform. 25, bbad539 (2024).
Liang, Y. DDI-SSL: drug–drug interaction prediction based on substructure signature learning. Appl. Sci. 13, 10750 (2023).
Nyamabo, A. K., Yu, H. & Shi, J. Y. SSI–DDI: substructure–substructure interactions for drug–drug interaction prediction. Brief. Bioinform. 22, bbab133 (2021).
Guo, L. et al. MSResG: using GAE and residual GCN to predict drug–drug interactions based on multi-source drug features. J. Interdiscip. Sci. Comput. Life Sci. 15, 171–188 (2023).
Wang, J., Wang, X. & Pang, Y. StructNet-DDI: molecular structure characterization-based ResNet for prediction of drug–drug interactions. Molecules 29, 4829 (2024).
Güvenç Paltun, B., Mamitsuka, H. & Kaski, S. Improving drug response prediction by integrating multiple data sources: matrix factorization, kernel and network-based approaches. Brief. Bioinforma. 22, 346–359 (2021).
Liu, Q. et al. Leveraging network target theory for efficient prediction of drug-disease interactions: a transfer learning approach. Adv. Sci. 12, 2409130 (2025).
Jiang, W. et al. Network-based multi-omics integrative analysis methods in drug discovery: a systematic review. BioData Min. 18, 27 (2025).
Mohamed, S. K., Nounu, A. & Nováček, V. Biological applications of knowledge graph embedding models. Brief. Bioinforma. 22, 1679–1693 (2021).
Li, R. et al. Graph signal processing, graph neural network and graph learning on biological data: a systematic review. IEEE Rev. Biomed. Eng. 16, 109–135 (2021).
Zhang Y., Mastouri M. & Zhang Y. Accelerating drug discovery, development, and clinical trials by artificial intelligence. Med 5, 1050–1070 (2024).
Honap, S. et al. Navigating the complexities of drug development for inflammatory bowel disease. Nat. Rev. Drug Discov. 23, 546–562 (2024).
Prueksaritanont, T. et al. Drug–drug interaction studies: regulatory guidance and an industry perspective. AAPS J. 15, 629–645 (2013).
Stewart, G. W. On the early history of the singular value decomposition. SIAM Rev. 35, 551–566 (1993).
Lee, D. D. & Seung, H. S. Learning the parts of objects by non-negative matrix factorization. nature 401, 788–791 (1999).
Shi, J. Y. et al. Detecting drug communities and predicting comprehensive drug–drug interactions via balance regularized semi-nonnegative matrix factorization. J. Cheminform. 11, 1–16 (2019).
Zhang, W. et al. Manifold regularized matrix factorization for drug-drug interaction prediction. J. Biomed. Inform. 88, 90–97 (2018).
Rohani, N., Eslahchi, C. & Katanforoush, A. ISCMF: integrated similarity-constrained matrix factorization for drug–drug interaction prediction. Netw. Model. Anal. Health Inform. Bioinform. 9, 1–8 (2020).
Shi, J. Y. et al. TMFUF: a triple matrix factorization-based unified framework for predicting comprehensive drug-drug interactions of new drugs. BMC Bioinform. 19, 27–37 (2018).
Jain, S. et al. Graph regularized probabilistic matrix factorization for drug-drug interactions prediction. IEEE J. Biomed. Health Inform. 27, 2565–2574 (2023).
Tang J. et al. Line: large-scale information network embedding. In Proc. 24th International Conference On World Wide Web 1067–1077 (2015).
Dai, Y. et al. Drug–drug interaction prediction with Wasserstein Adversarial Autoencoder-based knowledge graph embeddings. Brief. Bioinforma. 22, bbaa256 (2021).
Wang, M. et al. Drug-drug interaction predictions via knowledge graph and text embedding: instrument validation study. JMIR Med. Inform. 9, e28277 (2021).
Jiang, M. et al. Relation-aware graph structure embedding with co-contrastive learning for drug–drug interaction prediction. Neurocomputing 572, 127203 (2024).
Yu, Z. et al. Predicting drug–disease associations through layer attention graph convolutional network. Brief. Bioinforma. 22, bbaa243 (2021).
Han, X. et al. Smilegnn: drug–drug interaction prediction based on the smiles and graph neural network. Life 12, 319 (2022).
Yu, H. et al. Attention-based cross domain graph neural network for prediction of drug–drug interactions. Brief. Bioinforma. 24, bbad155 (2023).
Peng, L. et al. Reverse graph learning for graph neural network. In IEEE Transactions on Neural Networks and Learning Systems (2022).
Chen, M. et al. SGFNNs: signed graph filtering-based neural networks for predicting drug–drug interactions. J. Comput. Biol. 29, 1104–1116 (2022).
Gao, J. et al. Autoddi: drug–drug interaction prediction with automated graph neural network. IEEE J. Biomed. Health Inform. 28, 1773–1784 (2024).
Zhang, Y. et al. Attention is all you need: utilizing attention in AI-enabled drug discovery. Brief. Bioinform. 25, bbad467 (2024).
Huang, L., Chen, Q. & Lan, W. Predicting drug–drug interactions based on multi-view and multichannel attention deep learning. Health Inf. Sci. Syst. 11, 50 (2023).
Lin, S. et al. MDF-SA-DDI: predicting drug–drug interaction events based on multi-source drug fusion, multi-source feature fusion and transformer self-attention mechanism. Brief. Bioinform. 23, bbab421 (2022).
Erfani, S. M. et al. High-dimensional and large-scale anomaly detection using a linear one-class SVM with deep learning. Pattern Recognit. 58, 121–134 (2016).
Ravì, D. et al. Deep learning for health informatics. IEEE J. Biomed. Health Inform. 21, 4–21 (2016).
Yang, Z. et al. CNN-Siam: multimodal siamese CNN-based deep learning approach for drug‒drug interaction prediction. BMC Bioinform. 24, 110 (2023).
Liu, X. et al. Large language models facilitating modern molecular biology and novel drug development. Front. Pharmacol. 15, 1458739 (2024).
Sarumi, O. A. & Heider, D. Large language models and their applications in bioinformatics. Comput. Struct. Biotechnol. J. 23, 3498–3505 (2024).
Yuan, S. et al. Enhancing herbal medicine-drug interaction prediction using large language models. IEEE Journal of Biomedical and Health Informatics (2025).
Withers, C. A. et al. Natural language processing in drug discovery: bridging the gap between text and therapeutics with artificial intelligence. Expert Opin. Drug Discov. 20, 765–783 (2025).
Li, D. et al. LLM-DDI: leveraging large language models for drug-drug interaction prediction on biomedical knowledge graph. IEEE Journal of Biomedical and Health Informatics (2025).
Qi, H. et al. Improving drug-drug interaction prediction via in-context learning and judging with large language models. Front. Pharmacol. 16, 1589788 (2025).
Kumar, P. Diffusion Models and Generative Artificial Intelligence: Frameworks, Applications and Challenges. Archives of Computational Methods in Engineering, 1–44 (2025).
Yang, L. et al. Diffusion models: a comprehensive survey of methods and applications. ACM Comput. Surv. 56, 1–39 (2023).
Wognum, C. et al. A call for an industry-led initiative to critically assess machine learning for real-world drug discovery. Nat. Mach. Intell. 6, 1120–1121 (2024).
Thiyagalingam, J. et al. Scientific machine learning benchmarks. Nat. Rev. Phys. 4, 413–420 (2022).
Böttiger, Y. et al. SFINX-a drug-drug interaction database designed for clinical decision support systems. Eur. J. Clin. Pharmacol. 65, 627–633 (2009).
Guthrie, B. et al. The rising tide of polypharmacy and drug-drug interactions: population database analysis 1995–2010. BMC Med. 13, 74 (2015).
Altalhan M., Algarni A. & Alouane M. T. H. Imbalanced data problem in machine learning: a review. IEEE Access, (2025).
Gupta, S. & Gupta, A. Dealing with noise problem in machine learning data-sets: a systematic review. Procedia Comput. Sci. 161, 466–474 (2019).
Kaur, H., Pannu, H. S. & Malhi, A. K. A systematic review on imbalanced data challenges in machine learning: applications and solutions. J. ACM Comput. Surv. (CSUR) 52, 1–36 (2019).
Xie W. et al. Extracting Drug-Drug Interactions from Biomedical Texts Using BioBERT with Improved Focal Loss. In Proc. 11th International Conference on Bioinformatics and Computational Biology (ICBCB) 99–104 (IEEE, 2023).
Li, J., Xiong, C. & Hoi, S. C. H. Learning from noisy data with robust representation learning. In Proc. IEEE/CVF International Conference on Computer Vision 9485–9494 (2021).
Chen, W. et al. A survey on imbalanced learning: latest research, applications and future directions. Artif. Intell. Rev. 57, 137 (2024).
Burger, D. et al. Clinical management of drug–drug interactions in HCV therapy: challenges and solutions. J. Hepatol. 58, 792–800 (2013).
Kiang, T. K. L., Ensom, M. H. H. & Chang, T. K. H. UDP-glucuronosyltransferases and clinical drug-drug interactions. J. Pharmacol. Ther. 106, 97–132 (2005).
Jia, J. et al. Mechanisms of drug combinations: interaction and network perspectives. Nat. Rev. Drug Discov. 8, 111–128 (2009).
Tsigelny, I. F. Artificial intelligence in drug combination therapy. Brief. Bioinform. 20, 1434–1448 (2019).
Chen, S. et al. Artificial intelligence-driven prediction of multiple drug interactions. Brief. Bioinform. 23, bbac427 (2022).
Dou, M. et al. Drug–drug interaction relation extraction based on deep learning: a review. ACM Comput. Surv. 56, 1–33 (2024).
Betancur, J. et al. Prognostic value of combined clinical and myocardial perfusion imaging data using machine learning. JACC Cardiovasc. Imaging 11, 1000–1009 (2018).
Dara, S. et al. Machine learning in drug discovery: a review. Artif. Intell. Rev. 55, 1947–1999 (2022).
Prescott, L. F. Pharmacokinetic drug interactions. Lancet 294, 1239–1243 (1969).
Ito, K. et al. Prediction of pharmacokinetic alterations caused by drug-drug interactions: metabolic interaction in the liver. Pharmacol. Rev. 50, 387–411 (1998).
Van De Waterbeemd, H. & Gifford, E. ADMET in silico modelling: towards prediction paradise? Nat. Rev. Drug Discov. 2, 192–204 (2003).
Hodgson, J. ADMET—turning chemicals into drugs. Nat. Biotechnol. 19, 722–726 (2001).
Storelli, F. et al. The next frontier in ADME science: predicting transporter-based drug disposition, tissue concentrations and drug-drug interactions in humans. Pharmacol. Ther. 238, 108271 (2022).
Wang, Y. et al. In silico ADME/T modelling for rational drug design. Q. Rev. Biophys. 48, 488–515 (2015).
Cortes, J. E. et al. Pharmacokinetic/pharmacodynamic correlation and blood-level testing in imatinib therapy for chronic myeloid leukemia. Leukemia 23, 1537–1544 (2009).
Fulton, B. & Markham, A. Mycophenolate mofetil: a review of its pharmacodynamic and pharmacokinetic properties and clinical efficacy in renal transplantation. Drugs 51, 278–298 (1996).
Myung, Y., de Sá, A. G. C. & Ascher, D. B. Deep-PK: deep learning for small molecule pharmacokinetic and toxicity prediction. Nucleic Acids Res. 52, W469–W475 (2024).
Lin, Z. et al. Integration of in vitro and in vivo models to predict cellular and tissue dosimetry of nanomaterials using physiologically based pharmacokinetic modeling. ACS Nano 16, 19722–19754 (2022).
Krewski, D. et al. Applications of physiologic pharmacokinetic modeling in carcinogenic risk assessment. Environ. Health Perspect. 102, 37–50 (1994).
Ota, R. & Yamashita, F. Application of machine learning techniques to the analysis and prediction of drug pharmacokinetics. J. Control. Release 352, 961–969 (2022).
Yang, M. et al. Computational drug repositioning based on multi-similarities bilinear matrix factorization. Brief. Bioinform. 22, bbaa267 (2021).
Ogawa, R. & Echizen, H. Clinically significant drug interactions with antacids: an update. Drugs 71, 1839–1864 (2011).
Maton, P. N. & Burton, M. E. Antacids revisited: a review of their clinical pharmacology and recommended therapeutic use. Drugs 57, 855–870 (1999).
Dacey, R. G. & Sande, M. A. Effect of probenecid on cerebrospinal fluid concentrations of penicillin and cephalosporin derivatives. Antimicrob. Agents Chemother. 6, 437–441 (1974).
Doering, W. Quinidine-digoxin interaction: pharmacokinetics, underlying mechanism and clinical implications. N. Engl. J. Med. 301, 400–404 (1979).
Hager, W. D. et al. Digoxin-quinidine interaction: pharmacokinetic evaluation. N. Engl. J. Med. 300, 1238–1241 (1979).
Bigger, J.rJ. T. & Leahey, J.rE. B. Quinidine and digoxin: an important interaction. Drugs 24, 229–239 (1982).
Fenster, P. E. et al. Digoxin-quinidine interaction in patients with chronic renal failure. Circulation 66, 1277–1280 (1982).
FENSTER, P. E. et al. Digitoxin-quinidine interaction: pharmacokinetic evaluation. Ann. Intern. Med. 93, 698–701 (1980).
George, C. F. Interactions with digoxin: more problems. Br. Med. J. 284, 291 (1982).
Li, D. Q. et al. Risk of adverse events among older adults following co-prescription of clarithromycin and statins not metabolized by cytochrome P450 3A4. CMAJ 187, 174–180 (2015).
Kenwright, S. & Levi, A. J. Impairment of hepatic uptake of rifamycin antibiotics by probenecid, and its therapeutic implications. Lancet 302, 1401–1405 (1973).
Milo, G. & Schwartz, M. A. Apparent effect of probenecid on the distribution of penicillins in man. Clin. Pharmacol. Ther. 9, 345–349 (1968).
WALTERS, N. et al. Penicillin transport from cerebrospinal fluid. Neurology 26, 1008–1008 (1976).
Hieber, J. P. & Nelson, J. D. A pharmacologic evaluation of penicillin in children with purulent meningitis. N. Engl. J. Med. 297, 410–413 (1977).
Wilson, R. C. et al. Addition of probenecid to oral β-lactam antibiotics: a systematic review and meta-analysis. J. Antimicrob. Chemother. 77, 2364–2372 (2022).
Dobrek, L. Lower urinary tract disorders as adverse drug reactions—a literature review. Pharmaceuticals 16, 1031 (2023).
Jonker, D. M. et al. Towards a mechanism-based analysis of pharmacodynamic drug–drug interactions in vivo. Pharmacol. Ther. 106, 1–18 (2005).
Sorkin, E. M. & Heel, R. C. Terfenadine: a review of its pharmacodynamic properties and therapeutic efficacy. Drugs 29, 34–56 (1985).
Jackson, S. H. Pharmacodynamics in the elderly. J. R. Soc. Med. 87, 5 (1994).
Niu, J., Straubinger, R. M. & Mager, D. E. Pharmacodynamic drug–drug interactions. Clin. Pharmacol. Ther. 105, 1395–1406 (2019).
Jawetz, E., Gunnison, J. B. & Speck, R. S. Antibiotic synergism and antagonism. N. Engl. J. Med. 245, 966–968 (1951).
Sinnaeve, P. R. & Adriaenssens, T. Dual antiplatelet therapy de-escalation strategies. Am. J. Cardiol. 144, S23–S31 (2021).
Martin, W. R. Opioid antagonists. Pharmacol. Rev. 19, 463–521 (1967).
Levine, J. D. et al. The narcotic antagonist naloxone enhances clinical pain. Nature 272, 826–827 (1978).
MARTIN, W. R. Drugs five years later: naloxone. Ann. Intern. Med. 85, 765–768 (1976).
Klee, W. A. & NIRENBERG, M. Mode of action of endogenous opiate peptides. Nature 263, 609–612 (1976).
Roques, B. P., Fournié-Zaluski, M. C. & Wurm, M. Inhibiting the breakdown of endogenous opioids and cannabinoids to alleviate pain. Nat. Rev. Drug Discov. 11, 292–310 (2012).
Rothwell, P. M. et al. Effects of β blockers and calcium-channel blockers on within-individual variability in blood pressure and risk of stroke. Lancet Neurol. 9, 469–480 (2010).
Jick, H. et al. Calcium-channel blockers and risk of cancer. Lancet 349, 525–528 (1997).
Lant, A. F. Evolution of diuretics and ACE inhibitors, their renal and antihypertensive actions-parallels and contrasts. Br. J. Clin. Pharmacol. 23, 27S–41S (1987).
Epstein, M. & Calhoun, D. A. Aldosterone blockers (mineralocorticoid receptor antagonism) and potassium-sparing diuretics. J. Clin. Hypertens. 13, 644 (2011).
Thadepalli, H., Niden, A. H. & Huang, J. T. Treatment of anaerobic pulmonary infections: Carbenicillin compared to clindamycin and gentamicin. Chest 69, 743–746 (1976).
KLUGE, R. M. et al. The carbenicillin-gentamicin combination against Pseudomonas aeruginosa: correlation of effect with gentamicin sensitivity. Ann. Intern. Med. 81, 584–587 (1974).
Yuce, K. & Van Rooyen, C. E. Carbenicillin and gentamicin in the treatment of Pseudomonas aeruginosa infection. Can. Med. Assoc. J. 105, 919 (1971).
Staub, I. & Sieber, S. A. β-Lactams as selective chemical probes for the in vivo labeling of bacterial enzymes involved in cell wall biosynthesis, antibiotic resistance, and virulence. J. Am. Chem. Soc. 130, 13400–13409 (2008).
Butler, K. et al. Carbenicillin: chemistry and mode of action. J. Infect. Dis S1–S8 (1970).
Yoshizawa S., Fourmy D. & Puglisi J. D. Structural origins of gentamicin antibiotic action. EMBO J. 17, 6437–6448 (1998).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Andriole, V. T. Synergy of carbenicillin and gentamicin in experimental infection with Pseudomonas. J. Infect. Dis. 124, S46–S55 (1971).
HOFFMAN, T. A., CESTERO, R. & BULLOCK, W. E. Pharmacodynamics of carbenicillin in hepatic and renal failure. Ann. Intern. Med. 73, 173–178 (1970).
El Bakri, F. et al. Ototoxicity induced by once-daily gentamicin. Lancet 351, 1407–1408 (1998).
Modi, N. et al. Gentamicin concentration and toxicity. Lancet 352, 70 (1998).
Lianou, P. E. et al. Increased adherence to human epithelial cells of resistant Pseudomonas aeruginosa strains. J. Infect. 2, 354–359 (1980).
Noone, P. & Pattison, J. R. Therapeutic implications of interaction of gentamicin and penicillins. Lancet 298, 575–578 (1971).
Bailey, D. G., Dresser, G. & Arnold, J. M. O. Grapefruit–medication interactions: forbidden fruit or avoidable consequences?. J. CMAJ 185, 309–316 (2013).
Guo, L. Q. & Yamazoe, Y. Inhibition of cytochrome P450 by furanocoumarins in grapefruit juice and herbal medicines. Acta Pharmacol. Sin. 25, 129–136 (2004).
Ducharme, M. P., Warbasse, L. H. & Edwards, D. J. Disposition of intravenous and oral cyclosporine after administration with grapefruit juice. Clin. Pharmacol. Ther. 57, 485–491 (1995).
Bailey, D. G. et al. Grapefruit-felodipine interaction: effect of unprocessed fruit and probable active ingredients. Clin. Pharmacol. Ther. 68, 468–477 (2000).
Edgar, B. et al. Acute effects of drinking grapefruit juice on the pharmacokinetics and dynamics on felodipine—and its potential clinical relevance. Eur. J. Clin. Pharmacol. 42, 313–317 (1992).
Malhotra, S. et al. Seville orange juice-felodipine interaction: comparison with dilute grapefruit juice and involvement of furocoumarins. Clin. Pharmacol. Ther. 69, 14–23 (2001).
Bailey, D. G., Dresser, G. K. & Bend, J. R. Bergamottin, lime juice, and red wine as inhibitors of cytochrome P450 3A4 activity: comparison with grapefruit juice. Clin. Pharmacol. Ther. 73, 529–537 (2003).
Guo, L. Q. et al. Different roles of pummelo furanocoumarin and cytochrome P450 3A5*3 polymorphism in the fate and action of felodipine. Curr. drug Metab. 8, 623–630 (2007).
Bailey, D. G. et al. Grapefruit juice–drug interactions. Br. J. Clin. Pharmacol. 46, 101–110 (1998).
Lundahl, J. et al. Relationship between time of intake of grapefruit juice and its effect on pharmacokinetics and pharmacodynamics of felodipine in healthy subjects. Eur. J. Clin. Pharmacol. 49, 61–67 (1995).
Guédon-Moreau, L. et al. Absolute contraindications in relation to potential drug interactions in outpatient prescriptions: analysis of the first five million prescriptions in 1999. Eur. J. Clin. Pharmacol. 59, 689–695 (2003).
Broyles, A. D., Banerji, A. & Castells, M. Practical guidance for the evaluation and management of drug hypersensitivity: general concepts. J. Allergy Clin. Immunol. Pract. 8, S3–S15 (2020).
Thuluvath, P. J. When is diabetes mellitus a relative or absolute contraindication to liver transplantation?. Liver Transplant. 11, S25–S29 (2005).
Boudier-Revéret, M., Gour-Provençal, G. & Chang, M. C. Everything is relative,” maybe even absolute contraindications. Pain. Med. 21, 3244–3245 (2020).
Edinoff, A. N. et al. Clinically relevant drug interactions with monoamine oxidase inhibitors. J. Health Psychol. Res. 10, 39576 (2022).
Fraser, L. A. et al. Calcium channel blocker–clarithromycin drug interaction did not increase the risk of nonvertebral fracture: a population-based study. Ann. Pharmacother. 49, 185–188 (2015).
Ying, Z. et al. Gnnexplainer: Generating explanations for graph neural networks. Advances in neural information processing systems. 32. https://doi.org/10.48550/arXiv.1903.03894 (2019).
Uddin, S. H. et al. Interpretable Drug Interaction Forecasting: Leveraging Graph Neural Networks with Explainable Artificial Intelligence. In International Conference on Generative Artificial Intelligence, Cryptography, and Predictive Analytics, 59−72 (Springer Nature, Singapore, 2024).
Li, R. et al. Towards interpretable drug interaction prediction via dual-stage attention and Bayesian calibration with active learning. PeerJ Comput. Sci. 11, e2847 (2025).
Yu, H. et al. GGI-DDI: identification for key molecular substructures by granule learning to interpret predicted drug-drug interactions. Expert Syst. Appl. 240, 122500 (2024).
Wang, Y., Yang, Z. & Yao, Q. Accurate and interpretable drug-drug interaction prediction enabled by knowledge subgraph learning. Commun. Med. 4, 59 (2024).
Lin, X. et al. Interpretable multi-view attention network for drug-drug interaction prediction. In Proc. IEEE International Conference on Bioinformatics and Biomedicine (BIBM) 422–427 (IEEE, 2023).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012).
Zdrazil, B. et al. The ChEMBL Database in 2023: a drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res. 52, D1180–D1192 (2024).
Gaulton, A. et al. The ChEMBL database in 2017. Nucleic Acids Res. 45, D945–D954 (2017).
Altman, R. B. PharmGKB: a logical home for knowledge relating genotype to drug response phenotype. Nat. Genet. 39, 426–426 (2007).
Hewett, M. et al. PharmGKB: the pharmacogenetics knowledge base. Nucleic Acids Res. 30, 163–165 (2002).
Hodge, A.E. et al. The PharmGKB: integration, aggregation, and annotation of pharmacogenomic data and knowledge. J. Clin. Pharmacol. Ther. 81, 21–24 (2007).
Kuhn, M. et al. The SIDER database of drugs and side effects. Nucleic acids Res. 44, D1075–D1079 (2016).
Yang, W. et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–D961 (2012).
Zagidullin, B. et al. DrugComb: an integrative cancer drug combination data portal. Nucleic Acids Res. 47, W43–W51 (2019).
Zheng, S. et al. DrugComb update: a more comprehensive drug sensitivity data repository and analysis portal. Nucleic Acids Res. 49, W174–W184 (2021).
Liu, H. et al. DrugCombDB: a comprehensive database of drug combinations toward the discovery of combinatorial therapy. Nucleic Acids Res. 48, D871–D881 (2020).
Liu, H. et al. Correction to ‘DrugCombDB: a comprehensive database of drug combinations toward the discovery of combinatorial therapy. Nucleic Acids Res. 49, 10801–10802 (2021).
Ianevski, A. et al. RepurposeDrugs: an interactive web-portal and predictive platform for repurposing mono-and combination therapies. Brief. Bioinform. 25, bbae328 (2024).
Acknowledgements
Thanks for the Outstanding Youth Research Project of Anhui, Department ofEducation (No.2022AH020042), the Major Scientific Research Project of Universities in Anhui province (Grant No. 2024AH040143 and No. 2024AH040146), the Anhui Province quality projects (No. 2023sdxx027) and the Training Action Project of Anhui Provincial Education Department of Young and Middle-aged Teachers (No. JNFX2023020 and JWFX2025017), Anhui Provincial Key Laboratoryof Traditional Chinese Medicine Compound Independent Research Project (No. 2025AKLCMF06).
Author information
Authors and Affiliations
Contributions
Yuqing Lu: Validation, Investigation, Data curation, Writing-original draft. Jing Chen: Methodology, Validation, Data curation. Nini Fan: Methodology, Investigation, Data curation. Wenchao Song: Methodology, Data curation. Haiyang Sheng: Investigation. Yinfeng Yang: Conceptualization, Resources, Supervision. Jinghui Wang: Conceptualization, Resources, Supervision, Project administration, Funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, Y., Chen, J., Fan, N. et al. Machine learning models for drug-drug interaction prediction from computational discovery to clinical application. npj Digit. Med. 9, 198 (2026). https://doi.org/10.1038/s41746-026-02400-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-026-02400-3