Abstract
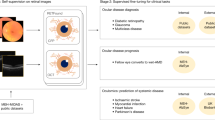
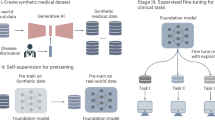
Foundation models (FMs) enable generalizable medical AI, but existing retinal FMs perform best on cross-sectional classification and detection and are less effective for predicting disease incidence and progression. We present RETFound Plus, a CFP-based FM trained with temporal modeling on 1,304,292 fundus photographs from 304,345 participants across multiple visits to learn progression-aware representations. Compared with RETFound, RETFound Plus improved calibration and 5-year risk prediction across systemic and ocular diseases, with larger gains for systemic outcomes (stroke, myocardial infarction, diabetes and hypertension; +4–10% c-index) than ocular outcomes (diabetic retinopathy and glaucoma; +3–7% c-index), and improved risk stratification for systemic diseases (1.2–2.1-fold higher hazard-ratio trend). Results were consistent across external multi-regional, multi-ethnic datasets from the UK, US, Singapore, Hong Kong, and Denmark.
Similar content being viewed by others
Data availability
The datasets used for pretraining, fine-tuning, and validation in this study contain sensitive human participant information, including retinal images with potentially identifiable retinal vascular patterns, as well as linked clinical and follow-up records. To protect participant privacy and comply with the data governance requirements and ethics approvals of each contributing cohort and institution, these datasets are not publicly available. Access to the data may be granted for research purposes on a reasonable request to the corresponding author(s), subject to approval by the relevant data custodians and/or institutional review boards, completion of applicable data use agreements, and any additional restrictions imposed by the originating cohorts (including for externally sourced datasets).
Code availability
The code used to train, fine-tune, and evaluate the model in this study is available at https://github.com/jaranwayne/RETFound-Plus.git. The environment was configured with the following dependencies: Python v3.8.2, Torch v.1.9.1, Torchvision v.0.10.1, Scikit-Image v.0.19.3, Scikit-Learn v.1.3.2, seaborn v.0.11.2, timm v.0.5.4, SciPy v1.10.1, opencv-python v.4.7.0.72, Pillow v9.5.0, setuptools v.59.4.0, Matplotlib v3.7.1, NumPy v1.24.2, Pandas v2.0.0, openpyxl v.3.1.2, and pycm v.4.0.
References
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Azizi, S. et al. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nat. Biomed. Eng. 7, 756–779 (2023).
Krishnan, R., Rajpurkar, P. & Topol, E. J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 6, 1346–1352 (2022).
Zhou, Y. et al. A foundation model for generalizable disease detection from retinal images. Nature 622, 156–163 (2023).
Wang, J. et al. Self-improving generative foundation model for synthetic medical image generation and clinical applications. Nat. Med. https://doi.org/10.1038/s41591-024-03359-y. (2024).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. https://doi.org/10.1038/s41591-024-02857-3 (2024).
Lu, M. Y. et al. A visual-language foundation model for computational pathology. Nat. Med. https://doi.org/10.1038/s41591-024-02856-4 (2024).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nat. Med. https://doi.org/10.1038/s41591-024-03141-0 (2024).
Han, T. et al. Image prediction of disease progression for osteoarthritis by style-based manifold extrapolation. Nat. Mach. Intell. 4, 1029–1039 (2022).
Mukkavilli, S. K. et al. AI Foundation Models for Weather and Climate: Applications, Design, and Implementation. Preprint at http://arxiv.org/abs/2309.10808 (2023).
Nguyen, T., Brandstetter, J., Kapoor, A., Gupta, J. K. & Grover, A. ClimaX: A foundation model for weather and climate. In Proc. 40 th International Conference on Machine Learning (PMLR, Honolulu, Hawaii, USA, 2023).
Wang, X., Feng, M., Qiu, J., Gu, J. & Zhao, J. From News to Forecast: Integrating Event Analysis in LLM-Based Time Series Forecasting with Reflection. In 38th Conference on Neural Information Processing Systems (NeurIPS 2024).
Wang, X., Sontag, D. & Wang, F. Unsupervised learning of disease progression models. in Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 85–94 (ACM, New York, New York, USA). https://doi.org/10.1145/2623330.2623754 (2014).
Ting, D. S. W. et al. Development and Validation of a Deep Learning System for Diabetic Retinopathy and Related Eye Diseases Using Retinal Images From Multiethnic Populations With Diabetes. JAMA 318, 2211 (2017).
Gunasekeran, D. V. et al. National use of artificial intelligence for eye screening in Singapore. NEJM AI. 1, (2024).
Wagner, S. K. et al. Insights into systemic disease through retinal imaging-based oculomics. Transl. Vis. Sci. Technol. 9, 6–6 (2020).
Wang, J. et al. Artificial intelligence-enhanced retinal imaging as a biomarker for systemic diseases. Theranostics 15, 3223–3233 (2025).
Poplin, R. et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat. Biomed. Eng. 2, 158–164 (2018).
Ong, J. et al. Development of oculomics artificial intelligence for cardiovascular risk factors: A case study in fundus oculomics for HbA1c assessment and clinically relevant considerations for clinicians. Asia-Pac. J. Ophthalmol. 13, 100095 (2024).
Holste, G. et al. Harnessing the power of longitudinal medical imaging for eye disease prognosis using Transformer-based sequence modeling. Npj Digit. Med. 7, 216 (2024).
Qiu, J. et al. Deep representation learning for clustering longitudinal survival data from electronic health records. Nat. Commun. 16, 2534 (2025).
Dai, L. A deep learning system for predicting time to progression of diabetic retinopathy. Nat. Med.
He, K. et al. Masked Autoencoders Are Scalable Vision Learners. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 15979–15988 (IEEE, New Orleans, LA, USA, 2022). https://doi.org/10.1109/CVPR52688.2022.01553.
Radford, A. et al. Learning transferable visual models from natural language supervision. in Proceedings of the 38th International Conference on Machine Learning 8748–8763, PMLR, (2021).
Zhai, X., Mustafa, B., Kolesnikov, A. & Beyer, L. Sigmoid Loss for Language Image Pre-Training. in 2023 IEEE/CVF International Conference on Computer Vision (ICCV) 11941–11952. https://doi.org/10.1109/ICCV51070.2023.01100 (2023).
Wang, Z., Wu, Z., Agarwal, D. & Sun, J. MedCLIP: Contrastive Learning from Unpaired Medical Images and Text. Proc. Conf. Empir. Methods Nat. Lang. Process. Conf. Empir. Methods Nat. Lang. Process. 2022, 3876–3887 (2022).
Qiu, J., Wu, J., Wei, H. et al. Development and validation of a multimodal multitask vision foundation model for generalist ophthalmic artificial intelligence[J]. NEJM AI 1(12), AIoa2300221 (2024).
He, K., Zhang, X., Ren, S. & Sun, J. Deep Residual Learning for Image Recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778. https://doi.org/10.1109/CVPR.2016.90 (2016).
Liu, Z. et al. A ConvNet for the 2020s. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 11966–11976 (2022). https://doi.org/10.1109/CVPR52688.2022.01167.
Dosovitskiy, A. et al. An Image is Worth 16x16 Words: transformers for image recognition at scale. in (2020).
Lee, C., Zame, W., Yoon, J. & Schaar, M. van der. DeepHit: A deep learning approach to survival analysis with competing risks. Proc. AAAI Conf. Artif. Intell. 32, (2018).
Allen, A. et al. A digital twins machine learning model for forecasting disease progression in stroke patients. Appl. Sci. 11, 5576 (2021).
Young, A. L. et al. Data-driven modelling of neurodegenerative disease progression: thinking outside the black box. Nat. Rev. Neurosci. 25, 111–130 (2024).
Chen, R. & Wang, H. Time-to-event endpoints in imaging biomarker studies. J. Magn. Reson. Imaging JMRI.29446 (2024) https://doi.org/10.1002/jmri.29446.
Dai, L. et al. A deep learning system for detecting diabetic retinopathy across the disease spectrum. Nat. Commun. 12, 3242 (2021).
Krittanawong, C. Future physicians in the era of precision cardiovascular medicine. Circulation 136, 1572–1574 (2017).
Lau, E. & Wu, J. C. Omics, big data, and precision medicine in cardiovascular sciences. Circ. Res. 122, 1165–1168 (2018).
Dziopa, K., Chaturvedi, N., Asselbergs, F. W. & Schmidt, A. F. Identifying and ranking non-traditional risk factors for cardiovascular disease prediction in people with type 2 diabetes. Commun. Med. 5, 77 (2025).
DeGroat, W. et al. Discovering biomarkers associated and predicting cardiovascular disease with high accuracy using a novel nexus of machine learning techniques for precision medicine. Sci. Rep. 14, 1 (2024).
Eastwood, S. V. et al. Algorithms for the capture and adjudication of prevalent and incident diabetes in UK Biobank. PLOS ONE 11, e0162388 (2016).
Burton, M. J. et al. The Lancet Global Health Commission on Global Eye Health: vision beyond 2020. Lancet Glob. Health 9, e489–e551 (2021).
Flaxman, S. R. et al. Global causes of blindness and distance vision impairment 1990-2020: a systematic review and meta-analysis. Lancet Glob. Health 5, e1221–e1234 (2017).
Loewenstein, A. et al. Save our Sight (SOS): a collective call-to-action for enhanced retinal care across health systems in high income countries. Eye 37, 3351–3359 (2023).
Ye, X. et al. Osteocalcin and Risks of Incident Diabetes and Diabetic Kidney Disease: A 4.6-Year Prospective Cohort Study. Diab. Care 45, 830–836 (2022).
Tang, Z. et al. Relationship of OCT-based diabetic retinal neurodegeneration to the development and progression of diabetic retinopathy: a cohort study. Invest. Ophthalmol. Vis. Sci. 66, 32 (2025).
Majithia, S. et al. Cohort Profile: The Singapore Epidemiology of Eye Diseases study (SEED). Int. J. Epidemiol. 50, 41–52 (2021).
Thinggaard, B. S. et al. The I-OPTA Questionnaire: A National Assessment of Patients with Neovascular Age-Related Macular Degeneration. Ophthalmol. Ther. 13, 3035–3046 (2024).
Vitale, S. et al. Association of 2-year progression along the AREDS AMD scale and development of late age-related macular degeneration or loss of visual acuity: AREDS Report 41. JAMA Ophthalmol. 138, 610 (2020).
Fu, H. et al. Evaluation of Retinal Image Quality Assessment Networks in Different Color-Spaces. in Medical Image Computing and Computer Assisted Intervention – MICCAI 2019 (eds Shen, D. et al.) 48–56 (Springer International Publishing, Cham, 2019).
Zhou, J. et al. IBOT : IMAGE BERT PRE-TRAINING WITH ONLINE TOKENIZER. In International Conference on Learning Representations (ICLR, 2022).
Grill, J.-B. et al. Bootstrap your own latent a new approach to self-supervised learning. In Proc. 34th International Conference on Neural Information Processing Systems (Curran Associates Inc., Red Hook, NY, USA, 2020).
Zhang, H. et al. DINO: DETR with Improved DeNoising Anchor Boxes for End-to-End Object Detection. The Eleventh International Conference on Learning Representations (2023).
Loshchilov, I. & Hutter, F. Decoupled Weight Decay Regularization. In International Conference on Learning Representations (ICLR, 2019).
Popescu, D. M. et al. Arrhythmic sudden death survival prediction using deep learning analysis of scarring in the heart. Nat. Cardiovasc. Res. 1, 334–343 (2022).
Acknowledgements
This study was supported by the National Key R&D Program of China (2022YFC2502800), the National Natural Science Foundation of China (82388101) and the Beijing Natural Science Foundation (IS23096) to T.Y.W.; the National Medical Research Council of Singapore (NMRC/MOH/HCSAINV21nov-0001) to Y.-C. T.; the National Natural Science Foundation of China (T2525004 & 62272298), the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2023ZD0509202 & 2023ZD0509201), the National Key Research and Development Program of China (2022YFC2407000) to B.S. and Z.W.; the UK Research & Innovation Future Leaders Fellowship (MR/T019050/1), Moorfields Eye Charity with The Rubin Foundation Charitable Trust (GR001753), and an Alcon Research Institute Senior Investigator Award to PAK; an Alcon Research Institute Senior Investigator Award to Y.W.; Wellcome Award 318987/Z/24/Z to Y.Z.; and the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2023ZD0509202 & 2023ZD0509201), the Clinical Special Program of Shanghai Municipal Health Commission (20224044) and the Three-Year Action Plan to Strengthen the Construction of the Public Health System in Shanghai (2023-2025 GWVI-11.1-28) to T.C.
Author information
Authors and Affiliations
Contributions
Y-C.T., B.S. and T.Y.W. conceptualized this work. Z.W. developed the algorithm. Z.W., Y.Z. conducted the experiments. Z.W., Y.W. analyzed the data, prepared the figures and tables, and drafted the manuscript. J.H.L.G., K.Z., Y.C., Z.G., Y.C., G.D.Y., P.Z., C.Y., A.R.R., M.L.C., C.X., Z.S., S.Y., D.F., X.L., B.T., J.G., H.L., Y.J., N.J., H.L., J.S., T.L., T.C., X.W., Q.W., C.S., S.K.W., C.Y.C., C-Y.C., and P.A.K. contribute with the validation datasets. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
P. A. K. is a cofounder of Cascader Ltd. and has acted as a consultant for Retina Consultants of America, Roche, Boehringer-Ingleheim, and Bitfount and is an equity owner in Big Picture Medical. He has received speaker fees from Zeiss, Thea, Apellis, and Roche. He has received travel support from Bayer and Roche. He has attended advisory boards for Topcon, Bayer, Boehringer-Ingelheim, and Roche. T. Y. W. is a consultant for AbbVie Pte Ltd, Aldropika Therapeutics, Bayer, Boehringer-Ingelheim, Carl Zeiss, Genentech, Iveric Bio, Novartis, Opthea Limited, Plano, Quaerite Biopharm Research Ltd, Roche, Sanofi, and Shanghai Henlius. He is an inventor, holds patents, and is a co-founder of start-up companies EyRiS and Visre, which have interests in and develop digital solutions for eye diseases. All potential conflicts of interest for consultancy, advisory boards, and positions in the start-up companies, and financial remuneration, if any, are managed by institutional policies under SingHealth and Tsinghua University. The other authors declare no financial or non-financial competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Z., Zhou, Y., Wu, Y. et al. Time and person sensitive foundation model for disease prediction and risk stratification. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02524-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02524-6