Abstract
The pig is an important model for studying human diseases and is also a significant livestock species, yet its testicular development remains underexplored. Here, we employ single-cell RNA sequencing to characterize the transcriptomic landscapes across multiple developmental stages of Bama pig testes from fetal stage through infancy, puberty to adulthood, and made comparisons with those of humans and mice. We reveal an exceptionally early onset of porcine meiosis shortly after birth, and identify a distinct subtype of porcine spermatogonia resembling transcriptome state 0 spermatogonial stem cells identified in humans, which were previously thought to be primate specific. We also discover the persistent presence of proliferating progenitors for myoid cells in postnatal testes. The regulatory roles of Leydig cell steroidogenesis and estrogen synthesis in supporting cell lineages are also explored, including the potential impact of estrogen on Sertoli cell maturation and spermatogenesis. Overall, this study offers valuable insights into porcine testicular development, paving the way for future research in reproductive biology, advancements in agricultural breeding, and potential applications in translational medicine.
Similar content being viewed by others
Introduction
Animal models play a crucial role in exploring human diseases including infertility. Recent research has highlighted substantial disparities between rodent models, such as mice, and humans in terms of cellular composition, molecular features, and developmental trajectories1,2,3. This disparity emphasizes the need to investigate alternative animal models that more accurately replicate human testis development. Pigs (Sus scrofa) closely resemble humans in various anatomical aspects, including heart percentage of body weight, brain white matter composition, length of intestine per kilogram, and epidermal thickness4. Due to these similarities, pigs have become increasingly valuable as translational medical models in recent years, particularly in areas like organ grafting and drug toxicological testing5,6,7,8. Besides that, pigs are the second most important source of meat world-wide and male pigs play a crucial role in pig production. Pioneering research studies have provided histological and morphological descriptions that elucidate key biological events occurring during spermatogenesis of boar9,10. However, these studies did not examine the transcriptional aspects of this process that hinder its broader interpretation. Consequently, it is of great importance to study the cellular composition and the molecular signatures of pig testicular cells.
The testes undergo a complex developmental process initiated during embryonic stages, progressing through neonatal, pubertal, and adult periods11,12. This process is intricately regulated, involving communication between the germline and various somatic cell types. In humans, abnormal testes development or malfunction is linked to susceptibility to various male reproductive disorders, including disorders of sex development, infertility, and testicular cancer. Understanding the cell fate-transition process in the testes poses challenges due to intricate molecular mechanisms within and across lineages. Knowledge of testicular cell types in pigs remains limited13. Descriptive studies exploring testis development at high resolution are scarce, creating a significant knowledge gap. This gap hinders not only the extensive modeling of human diseases but also the improvement of pig reproduction efficiency.
Single-cell RNA sequencing (scRNA-seq) has become a powerful tool for unraveling the molecular intricacies underlying developmental, physiological and pathological conditions, providing valuable clues for the creation of transgenic and knockout animals for disease modeling and animal breeding. In this study, we employ scRNA-seq to characterize, at a single-cell resolution, the gene expression and lineage changes during the testis development of Bama pig—an important human disease model. This comprehensive mapping not only reveals the developmental trajectories of major cell types constituting the testes but also elucidates the cell-to-cell interactions governing spermatogenesis. To discern similarities and heterochronic differences among species, we reanalyze previously published datasets of testes from humans and mice14,15,16, providing an overview of molecular and cellular patterns across mice, pigs, and humans.
Results
Single-cell transcriptomic profiles of the developing porcine testes
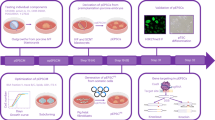
To delineate the developmental process of the porcine testis, we conducted single-cell transcriptomic profiling on testicular cells isolated from Bama pigs at various postnatal days (PD), including PD9, PD30, PD60, PD90, and PD150, covering the entire period from newborn to adult (Fig.1a). To enhance the representation of testicular somatic cells, additional biological replicates were collected at PD90 and PD150. Following the filtering out of multiplets and low-quality cells, a total of 41,548 cells were retained for downstream analysis (Supplementary Fig. 1a). Cluster analysis enabled the annotation of nine major cell types based on the expression of established cell markers14,17,18,19,20, constructing a temporal gene expression landscape of porcine testis development (Fig. 1b). Each cell exhibited a median expression of 2025 detected genes (Supplementary Fig. 1b). The dataset’s robustness was supported by the specific expression of marker genes for spermatogenic and somatic cell types. Germ cells encompassed spermatogonia (S’gonia, UCHL1+), spermatocytes (S’cyte, SYCP2+), and spermatids (S’tid, TNP1+) (Fig.1c). Testicular niche cells (VIM+) represented a heterogeneous mix of Sertoli cells (SOX9+), myoid/stromal cells (COL1A1+ and/or ACTA2+), Leydig cells (CYP11A1+), vascular cells (VWF+), and immune cells (PTPRC+) (Fig. 1c and Supplementary Fig. 1c). Visualization on the UMAP plot illustrated consistent agreement among samples regarding different postnatal ages, biological replicates, and cell cycle phases (Fig. 1d and Supplementary Fig. 1d, e). The distribution of testicular cell types reflects the developmental progression across the sampled time points (Fig.1e). Notably, the proportion of germ cells increased significantly during spermatogenesis, accompanied by a gradual reduction in the percentage of somatic cells (Fig. 1f and Supplementary Fig. 1f). Furthermore, in contrast to human testes, a limited number of lymphocytes (marked by CCL5 and GZMB) were identified in porcine testes (Supplementary Fig. 1g). In summary, our scRNA-seq datasets effectively captured the major cell types and their dynamic diversity during the testis development of Bama pigs.
a Schematic diagram of the experimental workflow. b UMAP visualization of combined single-cell transcriptome data from porcine testes. Each dot represents a single cell that is color-coded by cell type. c Dot plot depicting selected differentially expressed genes for all cell types identified. d UMAP visualization of distribution of single cells collected from different timepoints. Cells are colored based on the sample of origin. e UMAP visualization of annotated cell types from individual timepoints. f Fraction of cell type per timepoint, exhibiting a progressive increase in cellular complexity throughout development. Source data are provided in Supplementary Data 1.
Stepwise establishment of porcine spermatogenesis and comparison with humans and mice
To gain a deeper understanding of the molecular characteristics throughout spermatogenesis, we isolated porcine germ cells from PD9 to PD150, and subsequently categorized them into eight subtypes (Fig. 2a and Supplementary Fig. 2a). By utilizing known markers, we organized two sub-clusters of spermatogonia, four sub-clusters of spermatocytes, and two sub-clusters of spermatids, sequentially representing mitotic, meiotic, and postmeiotic stages (Supplementary Fig. 2b). Through an extensive analysis, we identified 952 genes exhibiting highly dynamic changes along the germ cell differentiation path. This allowed us to explore their associated Gene Ontology (GO) terms (Fig. 2b). As anticipated, biological processes such as ‘ribosome biogenesis’ and ‘cellular respiration’ were notably enriched in spermatogonia, concomitant with upregulation of proliferative genes (e.g., TOP2A) during the transition to differentiating spermatogonia, which may be related to the balance between self-renewal and differentiation. Notably, this analysis unveiled FANCI and FANCD2, components of the core complex in the Fanconi anemia (FA) pathway21, as prominent markers for porcine undifferentiated spermatogonia, suggesting potential avenues for future research (Supplementary Fig. 2c). For instance, the confirmed role of FA component in the maintenance of mouse undifferentiated spermatogonia supports the probability of the involvement of this conserved pathway in porcine undifferentiated spermatogonia as well22. Furthermore, in spermatocytes to spermatids, genes implicated in meiosis and spermatogenesis exhibited high expression levels, signifying the most characteristic transcriptomic changes in this phase (Fig. 2b).
a Left: Highlight of germ cells in the UMAP from Fig. 1b, showing the cells selected for focused analysis. Right: UMAP visualization of porcine germ cell types. Each dot represents a single cell that is color-coded by cell type. b Heatmap showing average expression of DEGs for each spermatogenic cell type. Each row represents a gene. The representative gene and enriched GO term of each gene cluster are listed on the right. c Pie plot showing the proportion of spermatogenic cell types at the indicated developmental stages of pigs, humans, and mice. Colors indicate cell types. Source data are provided in Supplementary Data 1. (Data from Guo et al.15,32, and Zhao et al.16) d Hematoxylin and eosin (H&E) staining of sections of porcine testes. Arrowhead indicated a specific cell type; SG spermatogonia, SC spermatocyte, rST round spermatid, eST elongated spermatid, ST Sertoli cell, LC Leydig cell, MC myoid cell, BV Blood vessel. Scale bars, 50 μm. e UMAP visualization of sub-clusters of spermatogonia. Each dot represents a single cell that is color-coded by cell type. Arrowhead indicated pseudo-developmental trajectory for spermatogonia. f Dot plot depicting selected differentially expressed genes for each sub-cluster identified in spermatogonia. g Fraction of each sub-cluster of spermatogonia at per timepoint. Source data are provided in Supplementary Data 1. h Heatmap showing the Pearson correlation of the normalized average gene expression between spermatogonia from pigs and humans.
Next, we compared germ cell composition across pigs, humans, and mice throughout the infancy to adulthood period. Previous studies have systematically integrated single-cell transcriptomic atlases covering the entire process of male germ cell development in humans and mice14,15,16. To ensure consistency, we compiled and re-analyzed published data using canonical markers with the same analysis pipeline on our pig data (Supplementary Fig. 2d–g). Remarkably, with sex-maturity normally at 6-months old age in pigs, at PD9, transcriptome-based clustering revealed the emergence of pachytene spermatocytes which is observed only up to the postnatal day (PND) 14 in mice, indicating an earlier onset of meiotic initiation in pigs compared to mice (Fig.2c). In humans, germ cells consist solely of spermatogonia at the period from birth to 7 years afterward, during which the proportion of differentiating spermatogonia increases, and a small number of spermatids appear in the 11-year-old juvenile15,23. However, histological examination revealed small testis cords lacking apparent spermatocytes in PD9 pig testes (Fig. 2d). This appearance resembles the testis morphology observed in human childhood and fetal/early postnatal mice. Nonetheless, we identified spermatocytes in the seminiferous tubules of porcine testes at PD15 (Fig. 2d). The incomplete consistency between the analysis and histological findings could be due to cells undergoing transcriptional changes preceding the occurrence of morphological transformation. By PD30, a distinct tubular structure with a cavity became progressively apparent, and spermatids were observed in a small proportion of pig tubules (Fig. 2d). Unexpectedly, we discovered that spermatogenesis commences at a very early stage in Bama pigs.
Discovery of a porcine spermatogonial subtype resembling transcriptome state 0 SSCs in humans
Spermatogonia maintain the life-long process of spermatogenesis by balanced self-renewal and differentiation. Our focus turned to Bama pig spermatogonia, given divergent findings between humans and model animals in previous studies24. Through unsupervised clustering, we identified four distinct clusters in porcine Undiff. S’gonia and differentiating spermatogonia: SPG1-SPG4 (Fig. 2e). Pseudotime analysis, as indicated by the arrow, further defined the developmental routine from SPG1 to SPG4 (Fig. 2e and Supplementary Fig. 2g). Here, SPG1 represents the most naïve developmental state of spermatogonia, while SPG4 aggregates at the end of this developmental trajectory. Notably, further analysis of gene expression and cell cycle revealed unique molecular signatures for each subtype (Fig. 2f and Supplementary Fig. 2h). SPG3 expressed markers such as CDK2, CCND1 and MKI67, indicating that this population exhibits active proliferation and appears to share molecular similarities with differentiating spermatogonia in humans and mice. Whereas SPG4 represented the transition to preleptotene cells by upregulating meiotic genes including SYCE3, REC8, and MEIOC (Fig. 2f). Of special note are the porcine SPG1 subgroup, marked by the high expression levels of PIWIL4, EGR4, and FGFR3, resembles human transcriptome state 0 SSCs, which represent undifferentiated and quiescent reserves maintained throughout human male’s lifetime14. Statistical analysis of germ cell composition supported the stable presence of SPG1 in pig testes from infancy to adulthood (Fig. 2g). We also found a strong correlation in gene expression between porcine SPG1 and human state 0 spermatogonia, with subsequent subtypes showing some degree of similarity (Fig. 2h). Overall, we have uncovered a spermatogonial subtype within porcine testes closely resembling human state 0 SSCs, suggesting their potential as the most naïve germline stem cells. Notably, this marks the first instance of identifying such cells in a species other than primates1.
Developmental landscape of supporting and interstitial cell lineages
As pivotal constituents of the testicular somatic niche, a substantial number of cells from the supporting lineage (Sertoli cells) and interstitial lineage (myoid/stromal cells and Leydig cells) were captured in the developing testes of Bama pigs (Fig. 1f). To delineate the trajectory of supporting cell development, our focus turned to Sertoli cells across five postnatal stages (Fig. 3a). The clustering plot depicted cellular continuity across ages, initiating from PD9 and progressing towards a uniform cluster after PD60. This observation suggests that Sertoli cells in porcine testes likely remain in uniformed state around three months after birth. Pseudotime analysis revealed the progression of immature Sertoli cells into two distinct states, turning into one from PD30 onwards, ultimately forming terminal Sertoli cells along the trajectory (Supplementary Fig. 3a). Furthermore, we examined the expression dynamics of genes associated with the cell cycle (GO:0007049) and the number of transcripts along the pseudotime of Sertoli cells (Supplementary Fig. 3b). As expected, cell cycle-related genes (e.g., CDC34, GAB1, CDC16) are predominantly expressed in PD9 cells, supporting the more proliferative condition of immature Sertoli cells. We identified differentially expressed genes and GO terms in each sample, offering an additional paradigm for characterizing cell identity (Fig. 3b). At PD30 and PD60, we revealed an absence of SOX9/KI67+ cells, indicating that Sertoli cell proliferation has ceased (Supplementary Fig. 3c). Notably, PD30 marked a potential metabolic shift for Sertoli cells, indicated by the upregulation of genes associated with fatty acid metabolism, including ELOVL2 and FADS2 (Fig. 3c). This metabolic adaptation likely contributes to meeting the energy and morphological requirements for Sertoli cells necessary to support spermatogenesis. Additionally, the synthesis enzyme ALDH1A1, responsible for producing retinoic acid (RA) that induces spermatogonia differentiation, is partially expressed in Sertoli cells since PD9, and significantly elevated from PD30, contributing to the subsequent progression of germ cell development (Fig. 3c). The immunofluorescence staining for ALDH1A1 and SOX9 confirmed this event, demonstrating the specific expression and localization of ALDH1A1 within supporting cells. It also indicated a tendency for ALDH1A1 signals to move towards the basal part of the seminiferous tubule as testis development progresses (Fig. 3d). Also, the expression patterns of genes associated with tight junctions also mirror the developmental timing of Sertoli cells (Fig. 3c). In summary, a focused analysis of porcine Sertoli maturation showcased dynamic changes in molecular features and functional interpretation from newborn to adult.
a Left: Highlight of Sertoli cells in the UMAP from Fig. 1b, showing the cells selected for focused analysis. Right: PCA map visualization of porcine Sertoli cells. Each dot represents a single cell that is color-coded by timepoints. b Heatmap showing average expression of DEGs for timepoints. Each row represents a gene. The representative gene and enriched GO term of each gene cluster are listed on the right. c The gene-normalized dynamics of selected genes along the pseudotime trajectories. Each dot represents a single cell that is color-coded by timepoints. d Immunofluorescence images of ALDH1A1 (green) and SOX9 (red) in porcine testis sections from different timepoints. Nuclei were counterstained with DAPI (blue). Scale bars, 50 μm. e Immunofluorescence images of ACTA2 (red) and HSD3β (green) in porcine testis sections from different timepoints. Nuclei were counterstained with DAPI (blue). Scale bars, 50 μm. f Left: Highlight of interstitial cells in the UMAP from Fig. 1b, showing the cells selected for focused analysis. Right: PCA map visualization of porcine interstitial cells. Each dot represents a single cell that is color-coded by timepoints. g Expression of representative marker genes identifying interstitial lineages cast on the PCA plot from Fig. 4f Normalized expression is plotted on a low-to-high scale (gray-purple).
We next investigated interstitial cell lineage differentiation during testis development. Immunofluorescence staining revealed the presence of myoid cells (marked by ACTA2) and Leydig cells (marked by HSD3β) within the porcine testis from PD9 to PD150 (Fig. 3e). Upon re-clustering these two cell types, distinct lineages were evident as early as PD9 in Bama pigs (Fig. 3f). The expression of marker genes and enriched pathways affirmed the identity and functionality of myoid cells and Leydig cells in porcine testes, aligning with findings in humans and mice25,26,27 (Fig. 3g and Supplementary Fig. 3d). However, the origin of these two cell types in Bama pigs from a common interstitial progenitor, as previously demonstrated in human and mouse testes15,28, remains unknown. This led us to address this question using testicular samples from embryonic pigs.
Identification of a testicular myoid progenitor persisting from the embryonic period to adulthood
To gain a better understanding of interstitial cell lineage specification, we profiled testicular cells from fetal pigs at embryonic day 72 (E72). Additionally, we reanalyzed previously published single-cell transcriptomic data of embryonic testes at E24, E27, E30, and E35 of Bama pigs29 (Supplementary Fig. 4a). After integration and annotation, we successfully isolated somatic cells belonging to the myoid/stromal and Leydig lineages from all samples (Supplementary Fig. 4b, c). Further analysis revealed a notable clustering at the early stage (~E24), followed by transcriptional segregation into two separate lineages around E30, as depicted in the three-dimensional UMAP plot (Fig. 4a). By examining the expression of canonical markers, we identified the upper trajectory characterized by CYP17A1 expression as representing the Leydig lineage, while the lower trajectory marked by ACTA2 expression corresponded to the myoid/stromal lineage (Fig. 4b).
a 3D UMAP visualization of distribution of interstitial cells collected from different developmental stages. Each dot represents a single cell and is colored according to its donor of origin. b Expression patterns of known interstitial marker genes projected onto the UMAP plot from a. c UMAP visualization of reclustered myoid/stromal cells, colored by assignment to four cell types. d Dot plot depicting marker genes for 4 cell types identified from c. e Bar plot showing the cell number of myoid/stromal cell types for each developmental stage (outer). percentage of cell types for myoid/stromal cells from PD60-PD150 testes (inner). Source data are provided in Supplementary Data 1. f Pseudotime trajectory of myoid/stromal cells. Each dot represents a single cell and is colored according to its annotated cell type. g Developmental potency score of each cell projected onto the UMAP plot from c. h Immunofluorescence images of a proliferation marker, KI67 (green) with a myoid cell marker, ACTA2 (red) in porcine testis sections from different timepoints. Nuclei were counterstained with DAPI (blue). Scale bars, 50 µm. i The timeline and proposed model for Bama pig, human and mouse testicular somatic niche cell development at embryonic, fetal, and postnatal stages.
This integrated dataset enabled us to explore the origins, emergence, dynamics, and specialization of the interstitial lineage. We isolated myoid/stromal cells for focused clustering analysis (Supplementary Fig. 4d). Due to the expression of genes associated with mesenchymal identity like ACTA2 and COL3A1, we observed the differentiation of myoid cells and stromal cells after birth, respectively (Fig. 4c, d). As development progressed, differentiated cell types eventually occupied the entire proportion within the lineage (Fig. 4e). Notably, we identified one actively proliferating cluster, marked by high expression of TOP2A and MKI67, associated with promoting cellular proliferation (Fig. 4d). Combining this with pseudotime analysis, we distinguished four cell types: proliferating progenitor (Prolif progenitor, marked by TOP2A), the intermediate cluster (Transient, marked by POSTN), and two terminal types mentioned above (Fig. 4c, d and Supplementary Fig. 4e). The resulting branched trajectory tree accurately reflected the differentiation status and expression of known markers (Fig. 4f and Supplementary Fig. 4f). To validate the differentiation potential of the progenitor cells, we compared them with mouse in vitro testis interstitial cell populations30. We found that the progenitor cells in pig testes closely resemble the mice interstitial progenitors distinguished before (Supplementary Fig. 4g)30. Further, using CytoTRACE, we predicted a higher developmental potency for the Prolif progenitors compared to Myoid cells and ACTA2-stromal cells (Fig. 4g).
To further provide functional insights into their characteristics, we calculated the myoid/stromal signatures of these cell types. As anticipated, peritubular myoid cells exhibited higher muscle signatures, while ACTA2- cells showed higher expression of stromal signatures (Supplementary Fig. 4h). Correlation analysis and hierarchical clustering further supported that myoid and stromal cells represent two distinct cell populations with unique functional traits (Supplementary Fig. 4i,j). In contrast, proliferating progenitor cells exist in a more primitive and early state (Fig. 4f and Supplementary Fig. 4j). Considering the known plasticity of the testicular interstitial lineage and the potential proliferative capacity of these cells, we investigated whether the proliferating progenitors annotated were present in the adult testes of Bama pigs. Immunofluorescence staining revealed the presence of a potential proliferating progenitor within the basal lamina of seminiferous tubules, characterized by co-expression of MKI67 and ACTA2 (Fig. 4d, h). These findings demonstrate the existence of actively dividing myoid progenitors in postnatal porcine testes. Despite the prevailing notion suggesting that testicular somatic cells arise post-mitotically, our analysis identified such rare cell subpopulations in Bama pigs, holding the potential to differentiate into either stromal or myoid cells.
The regulatory role of Leydig cell steroidogenesis in developing porcine testes
The sex hormone secreted by Leydig cells plays a pivotal role in male gonadal differentiation and maturation26. We proceeded to analyze the Leydig lineage from Fig. 4a. Re-clustering analysis revealed two major clusters displayed in the PCA plot after regressing out batch effects (Fig. 5a). In line with the recognized roles of fetal Leydig cells in androgen synthesis in humans and mice31, we observed that fetal Leydig cells in Bama pigs also exhibited upregulation of terms linked to the steroid biosynthetic process. Conversely, porcine adult Leydig cells showed more significant enrichment of cellular catabolic processes (Supplementary Fig. 5a). As of current, we have delineated the trajectory of testicular somatic differentiation of pigs, and summarized its developmental patterns in relation to human and mouse (Fig. 4i).
a Single cell of Leydig lineage cast on PCA plot, colored by stages of development. b Line plots showing the expression levels of steroidogenic genes in each stage of development. c Box plots showing the concentration of Testosterone (top) or Estrogen (bottom) in serum of postnatal Bama pigs. Boxes represent the interquartile range with the median as a horizontal line, and points are shown as each replicate. Source data are provided in Supplementary Data 1. d Expression patterns of CYP17A1 and CYP19A1 projected onto the PCA plot from (e) GO terms enriched in up-regulated genes of estrogen synthesizing Leydig cells compared to other Leydig cells (log2foldchange > 0.25). f Line plots showing the expression levels of ESR1 and ESRRA in each stage of development. g Clustering analysis of dynamic gene expression during development of supporting lineage. Four modules of genes form two distinct categories according to the expression patterns. Median of normalized gene-expression level of each module (orange line) was shown. h Correlation analysis of ESR1 and ESRRA expression with up-regulated ER genes (left) or down- regulated ER genes (right) using Pearson’s correlation coefficients. Points represent the cells in supporting lineage, colored by developmental stages. AveExpr: average expression. i GO terms enriched in up-regulated and down-regulated ER genes with -log10(p value) colored according to the color key at the bottom.
To further explore the steroidogenic activity of Leydig cells, we monitored the expression dynamics of steroid biosynthetic enzymes during development (Fig. 5b). As expected, we observed the upregulation of genes responsible for producing testosterone after PD60, including STAR, CYP11A1, and CYP17A1. Unexpectedly, the expression of estrogen synthase was also found in porcine Leydig cells, reaching a level comparable to testosterone synthase in adulthood, unlike in humans or mice16,32 (Fig. 5b and Supplementary Fig. 5b). We measured serum testosterone and estrogen levels in Bama pigs aged 0–6 months after birth and observed a gradual rise in estrogen content (Fig. 5c), consistent with previous reports in boar testes33,34. Notably, the expression of estrogen synthase was concentrated in a subset of Leydig cells termed as “Leydig_type2”, rather than being widely expressed (Fig. 5d and Supplementary Fig. 5c). Compared to other Leydig cells (Leydig_type1), Leydig_type2 exhibits similar LHCGR expression, but higher Androgen Receptor (AR) expression (Supplementary Fig. 5d). Moreover, upregulated GO terms included cholesterol metabolic processes, represented by enzymes functioning in lipid metabolism upstream of sex hormone biosynthesis (e.g., HMGCR) (Fig. 5e and Supplementary Fig. 5d). The downregulated genes included DLK1 (Supplementary Fig. 5d), which is specifically expressed in human fetal Leydig cells32.
Sertoli cells serve as the primary targets of androgens in testes. The androgen receptor signaling pathway governs Sertoli cell proliferation, maturation, and the maintenance of the blood-testis barrier formed between Sertoli cells35,36,37,38,39. Considering this, we hypothesized that estrogen may also play a role in the development of the porcine testicular supporting cell lineage. Indeed, our integrated dataset of supporting cells (from Supplementary Fig. 4b) revealed a significant upregulation of Estrogen Receptor 1 (ESR1) and Estrogen Related Receptor Alpha (ESRRA) at E35 and postnatal stages (Fig. 5f and Supplementary Fig. 5e). Both have been reported as transcription factors responsible for activating or inhibiting the expression of target genes40,41,42. To identify their target genes, we utilized Cluster-Buster to locate direct binding sites of ESR1 and ESRRA in the promoter regions of all genes in the pig, considering regions with a score greater than 8 (indicating a turning point in the number of regions) as estrogen signaling target genes (Supplementary Fig. 5f). In supporting cells, we identified four clusters of genes exhibiting similar (cluster 1 & 2) or opposite (cluster 3 & 4) expression trends to ESR1 and ESRRA. Among these clusters, 57 and 50 genes, respectively, overlapped with estrogen signaling target genes (Fig. 5g), termed estrogen-regulated genes (ER genes) in supporting cells. The expression of these ER genes positively or negatively correlated with the expression of ESR1 or ESRRA, confirming their reliability as target genes (Fig. 5h). Moreover, upregulated ER genes were enriched in immunity-related GO terms, which were also upregulated in human Sertoli cells during puberty15, leading to speculation that they may contribute to protecting the testis from infections43,44,45(Fig. 5i). Conversely, downregulated GO terms enriched in ER genes, such as ‘regulation of cell differentiation’ and ‘cell cycle process’, suggest that estrogen has the potential to regulate porcine supporting cells, guiding them to exit proliferation and commence differentiation into immature Sertoli cells (Fig. 5i). Additionally, estrogen receptors and estrogen-related receptors also exhibited stage-specific expression during germ cell development (Supplementary Fig. 5g–i). Further investigation is needed to determine whether estrogen directly regulates porcine spermatogenesis. In summary, we discovered that a subset of porcine Leydig cells possesses the ability to synthesize estrogens, activating estrogen-mediated signaling pathways in the supporting cell lineage and aiding in the proliferation and maturation of porcine Sertoli cells.
The testicular signaling niche of porcine germ cells
The communication between the niche and germline is crucial for testis development, and understanding how these interactions trigger puberty and spermatogenesis is currently an area of significant interest. Through ligand-receptor analysis using CellChat, we identified cell types actively interacting with porcine germ cells, including myoid/stromal cells, vascular cells, followed by Sertoli cells and Leydig cells (Fig. 6a, b, and Supplementary Fig. 6a). We observed cell type-specific expression of genes encoding ligands and receptors involved in various signaling pathways, such as Pleiotrophin (PTN), KIT, Granulin (GRN), and Laminin (Fig. 6c). Furthermore, the comparative landscape revealed the dynamics of intercellular signaling activity with age, with the strongest cell-cell communication observed at PD9, maintaining nearly consistent levels after PD30 (Supplementary Fig. 6b). In terms of specific signaling pathways, spermatogonia at PD9 receive signals from various somatic cells, such as LAMB1/LAMA1/LAMC3 - DAG1, JAG1/DLL3/DLL4 - NOTCH2, INHABB - ACVR2B, INHABA - ACVR2A, etc (Supplementary Fig. 6c). The interactions gradually diminish as the testes develop (Supplementary Fig. 6c). This finding suggests that regulatory influences from the niche are highly active during the maturation process of porcine testes.
a Circle plots depict the number of interactions (left) or interaction weights/strength (right) in the cell-cell communication network among major cell types. Colors of dot/edges indicated the particular cell type. b Bubble diagram showing ligand-receptor communications between germline (target) and somatic niche (source) of porcine testes. The color of bubbles indicates the means of the communication probability. c Expression of ligands (red) and receptors (blue) of indicated signaling pathways cast on the UMAP plot from Fig. 1b. Normalized expression is plotted on a low-to-high scale (gray-purple). d Dot plot depicting expression of ligands and receptors of GDNF signaling for major cell types of porcine testes. e Schematic model of selected signaling pathways involved in germling-somatic niche interactions in porcine testes.
Our data also provides valuable insights into the conservation and diversity of the roles of glial cell line-derived neurotrophic factor (GDNF) signaling in SSCs maintenance across species. Mouse single-cell RNA sequencing (scRNA-seq) data reproduced the expression of the GDNF ligand, Gdnf, primarily in Sertoli cells and myoid/stromal cells, with the receptor Gfra1 highly expressed in spermatogonia (Supplementary Fig. 6d, e). While GFRA1 marks undifferentiated spermatogonia in humans as well, the ligand of GFRA2, NRTN, was detected in human Sertoli cells (Supplementary Fig. 6f, g). In contrast, GDNF was scarcely expressed in the human testicular niche. In light of this, we re-examined the regulatory logic of GDNF signaling in Bama pig using our dataset (Fig. 6d). Enrichment of ligand-receptor genes led us to speculate that NRTN in Sertoli cells, rather than GDNF, plays a central role as a paracrine factor, controlling the fate of porcine spermatogonia. In summary, we outlined the germline-niche interactions in porcine testes, emphasizing the need for further detailed functional investigation (Fig. 6e).
Discussion
Although porcine serve as an important agricultural resource and useful animal model for human diseases, its testicular development has not been extensively studied. In this study, we conducted a single-cell transcriptomic survey of germline and testicular somatic cells in the Bama pig, aiming to characterize similarities and differences in gene expression, cellular composition, and cell type trajectory across species.
While most spermatogenic cell types are shared among mice, pigs, and humans, we observed an exceptionally early initiation of meiosis in Bama pigs, comparable to mice in terms of natural days after birth. Additionally, at PD30, the emergence of mature spermatozoa was concurrently observed at both histological and transcriptomic levels. We propose that this event likely signifies the ‘first-wave’ of porcine spermatogenesis, a previously undefined aspect. Previous studies in mice have indicated that the first round of spermatogenesis during puberty is less efficient than in adults, accompanied by notable apoptosis46. Considering the sexual maturity at a later stage, i.e. 4–6 months old of pigs47 and 6–8 weeks old of mice, there is a prevailing suspicion that the first-wave of spermatogenesis might not yield fertile spermatozoa capable of spontaneous conception. Interestingly, a late emergence of spermatozoa in PD90 Guanzhong pig was previously reported in a study examining the histology of seminiferous tubules, indicating inter-species difference in the timing of spermatogenesis13. These findings provide new insights into the reproduction and breeding of Bama pigs, as the ability to generate functional sperm from the first-wave spermatogenesis allows males to attain fertility at an earlier stage.
Our investigation included insights relevant to optimizing humanized models. Comparing findings in rodents and humans, our study suggests a comparable continuum of spermatogonial differentiation in Bama pigs and humans, supported by transcriptomic profiles and cellular distribution. Particularly noteworthy is the identification of the early quiescent spermatogonia subtype in adult pigs, SPG1, which transcriptionally resembles the human state 0 SSCs, a transcriptome state not defined in adult mice14. In comparison to other cells, the transcriptome and abundance of state 0 cells remain stable during testis aging in humans48. However, due to the difficulty in obtaining samples, the characterization of such cells can only rely on observational studies, lacking extensively functional interpretation. Although prior work on Guanzhong pig provided knowledge of porcine spermatogenesis, it did not define this subtype of undifferentiated spermatogonia49. Hereby, the identification of heterogeneous spermatogonial subpopulations in Bama pigs provides clues for optimizing SSCs maintenance under regenerative conditions, potentially enhancing human fertility preservation technologies. Moreover, there is currently significant interest in exploring the communication between the niche and spermatogonia. In this study, we present new findings that illustrate potential differences among humans, mice, and pigs, as well as changes during development, setting the stage for future functional exploration. Specifically, we observed that for GDNF signaling, NRTN was predominantly expressed at the RNA level by Sertoli cells in pigs, as opposed to GDNF, expressed by Sertoli cells in mice. These findings open the door for future research into functional species differences in SSC maintenance.
This developmental map of testicular cells made in the present study serves as a valuable resource for understanding testicular interstitial lineage specification, addressing the low efficiency in unbiased capture of such cells by scRNA-seq in both adult humans and mice. We identified a small group of potential myoid progenitors that retain proliferative capacity within the testes of adult pigs. This challenges the traditional belief that testis interstitial cells are mitotically inactive. It is important to test whether these progenitors act as a reservoir of stem cells capable of differentiating into mature cell types in response to specific physiological simulations in future studies.
Our exploratory analysis is currently limited in several ways. Firstly, we are unable to incorporate more functional evidence in this work due to the unavailability of commercial antibodies and genetic modification techniques suitable for pigs. In addition, the reference genome of the pig is not as well annotated as that of humans and mice, which makes evolutionary comparisons impractical. By tackling these conceptual challenges, we can improve our ability to use comprehensive analyses to uncover fundamental principles of gonad development. Before pigs can be established as the preferred model for studying human diseases and gametogenesis, extensive inter-species comparison and functional validation are essential. Our work provides novel insights into the timing and strategic approach of porcine testis development, offering a valuable resource for future hypothesis-driven studies. The demonstrated porcine features, particularly in spermatogonial development, provide not only a detailed characterization of an experimental model organism closely mimicking human physiology, but also lay the theoretical groundwork for porcine reproduction, breeding and husbandry.
Methods
Porcine testicular tissues
All animal experiments were performed in accordance with the guidelines for the care and use of laboratory animals established by the Beijing Association for Laboratory Animal Science, and approved by the Animal Ethics Committee of the Institute of Zoology, Chinese Academy of Sciences. The Bama pigs were bred and reared at the Beijing Farm Animal Research Center. We have complied with all relevant ethical regulations for animal use. Before sacrifice, pigs were anesthetized (5 kg/mL, intravenous injection) with xylazine. Then, we collected morphologically intact and viable testicular tissue from euthanized pig samples.
Tissue isolation for single cell RNA sequencing
Porcine testes at different developmental stages were placed in ice-cold PBS buffer for transportation, which takes 1–1.5 h. Upon these samples’ arrival to cell culture room at ISCR, testes were washed with 1× PBS twice, followed by removing the epididymides, the visible connective tissue and tunica albuginea aseptically. Testes were carefully dissected into 0.4–0.6 g of tissue blocks, each block was placed in dissociation buffer containing 5 mL of freshly-prepared enzymatic mixture of (1 mg/mL) Collagenase I (Gibco, 17100-017) supplemented with (1 mg/mL) DNase I (AppliChem, A3778) in 1× PBS for digestion in a 37 °C water bath, this process needs constant gentle shaking at 60 rpm/min for 6 min until the mixture became uniform and turbid, gentle pipetting for several times using a 5-mL pipette was conducted to dissociate the cell clumps every 2 min while shaking. Afterwards, the mixture was firstly filtered through a 100 μm strainer to eliminate undigested tissue fragment, then centrifuged at 600 g for 5 min and pellets were washed with 1× PBS twice. Further digestion in 2.5 mL of 0.25% Trypsin-EDTA (Gibco, 25200072) supplemented with (1 mg/mL) DNase I for 3–5 min and collected the single cells by centrifugation after stop of digestion. Besides, the red blood cells were removed by lysing in red blood cell lysis buffer (Beyotime, C3702) and the pellet obtained was resuspended in 1× PBS with 0.04% BSA and finally strained through 40 μm strainer for single cell RNA sequencing.
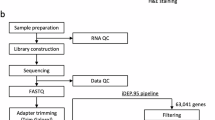
Single-cell transcriptomic library construction and sequencing
The procedures were instructed by the user guide of Chromium Next GEM Single Cell 3ʹ Reagent Kits v3.1, as provided by 10X Genomics. Briefly, in the step of GEM Generation and Barcoding, cells were diluted for recovery of ∼10k cells per lane and then loaded with master mix on Chromium Next GEM Chip G. After Post GEM-RT Cleanup, 12 cycles were used for cDNA Amplification. The resulting libraries were then subjected to sequencing on Illumina Novaseq 6000 instrument with following setting: 28 cycles for Read 1, 10 cycles for i5 index, 10 cycles for i7 index, and 90 cycles for Read 2.
H&E and immunofluorescence staining of testicular tissues
Fresh porcine testis tissue blocks (0.4–0.6 g) were fixed in 15 mL of 4% PFA for 24 h at 4 °C, followed by three times wash with 1× PBS for 10 min each wash, and then processed by conventional paraffin embedding procedure. 4–5-μm thick paraffin sections were made with a microtome (Leica RM2235) and mounted onto glass slides. Before conducting the H&E and immunofluorescence staining, sections were deparaffinized with xylene twice for 15 min each, then rehydrated through a graded ethanol series (100%, 100%, 95%, 95%, 85%, 85%, 75%, 75%, 50% and water) respectively.
For H&E staining, hematoxylin staining (Baso Biotech, BA4041) for 2 min, followed by treatment of freshly-prepared differentiator, acid alcohol (70% ethanol containing 1% HCl) for 5 s, then tap water was used for bluing. Eosin staining (Baso Biotech, BA4022) for 45 s to 1 min each sample and washed with tap water, then dehydrate and render the tissue completely transparent with several baths of ethanol and xylene, 30 s for 95% ethanol, 1 min for 100% ethanol, and 3 min for xylene, each step was conducted twice. Finally, neutral balsam mounting medium was added dropwise onto tissues and covered with coverslips carefully. Images were acquired on slide scanner (Leica Aperio VESA8) and analyzed using Aperio ImageScope (Leica company).
For immunofluorescence staining, antigen retrieval was performed using microwave-based method with sodium citrate solution (pH 6.0). Slides with tissues were immerged into solution, followed by being heated in microwave at 100 °C for around 4 min until slightly boiling, then kept at 30–40 °C for 15 min, and finally cooled down to room temperature. Sections were washed with 1× PBS and blocked with 5% BSA for 1 h at room temperature in a humidity chamber. Incubate the slides horizontally overnight at 4 °C, with the primary antibody (anti-DDX4, ab13840, 1:500; anti-ALDH1A1, A22531,1:500-1:2000; anti-ACTA2, ab124964, 1:500; anti-HSD3β, sc515120, 1:50; anti-Ki67, ab15580, 1:500; anti-SOX9, ab185966, 1:100) diluted in blocking solution in humidity chamber. After three washes in 1× PBS (10 min each), tissues were incubated with corresponding secondary antibodies at room temperature for 1 h shielded from light. After three washes in 1× PBS (10 min each), DAPI was added onto tissue sections for 15 min, then washed three times in 1× PBS (10 min each). Anti-fade reagent DABCO was finally added dropwise to the sections and covered with coverslips, then the edges were sealed with nail polish. Slides were stored at 4 °C until ready to microscopic image. Images were acquired on a Zeiss LSM 880 Confocal microscope and analyzed with software ZEN 2.6 (blue edition) (Zeiss company).
Measurement of serum hormone
Serum levels of testosterone and estradiol from Bama pig were quantified by Beijing Lawke Health Laboratory using ELISA method. Specifically, venous blood was collected from porcine veins at the particular time point, and was allowed to clot at room temperature for 2 h. Serum was separated by centrifugation (1503 × g, 20 min, 4 °C) and stored at −80 °C prior to the ELISA procedure. ELISA was performed using the manufacturer’s instructions.
scRNA-seq data analysis
We analyzed the scRNA-seq data of porcine testes using the Seurat package50 (https://satijalab.org/seurat/). The generated Unique molecular identifier (UMI) count matrix for each sample were loaded into R using the Read10X function. After adding their sample information, all of the matrices were merged and the Seurat object was created. According to the developer’s vignettes, the data were filtered. Specifically, cells were retained with >800 expressed genes, >2000 detected UMIs, and <25% of the reads mapped to the mitochondrial genome. The Seurat object was normalized by default setting and underwent linear dimensional reduction via principal component analysis (PCA), and then Harmony was used to remove batch effect between samples51. The uniform manifold approximation and projection (UMAP) technique was used to visualize the dataset by choosing 1–15 PCs in the cluster analysis. We subsequently identified cell types by visualizing marker genes using the FeaturePlot function. Contaminating red blood cell genes and sperm genes were detected in unspecific stage. As these non-biological genes were deemed irrelevant, they were subsequently excluded from the final Seurat object.
When conducting focused analysis on particular cell types, the cell barcodes of the designated cells were extracted and the corresponding UMI count table from the total matrix was utilized to create a new Seurat object.
For the combined analysis of published data of embryonic testes and this work, external datasets from Chen et al. (https://doi.org/10.5281/zenodo.6918355) was acquired and loaded into R using Read10X_h5 function, and then integrated with ours.
Differentially expressed genes (DEGs) between given clusters or samples were identified by the FindMarkers function in Seurat. The genes with expression level difference greater than 1 and padjust values less than 0.001 were set as the threshold. All GO enrichment analysis on DEGs was performed using enrichGO function of clusterProfiler and org.Ss.eg.db52.
The Monocle2 package was used for trajectory construction and pseudotime inference following the default settings53. The Seurat object of particular cell type was converted to a CellDataSet object for importing into orderCells function to construct the trajectory.
To infer the cell cycle stage of individual cells, the CellCycleScoring function from Seurat was utilized with default parameters, utilizing marker genes for G2/M and S phase. Cells were subsequently categorized into distinct cell cycle phases, namely G1, S, and G2M.
To quantify the functionality of interstitial cell types, the AddModuleScore function from Seurat was used by calculating myoid (GO: 0042692) or stromal (GO: 0062023) signature genes. Correlation analysis of given cell types were conducted by corr function in R, with correlation coefficient was calculated with the Pearson method. Hierarchical clustering with Lance-Williams dissimilarity distance was generated by hclust function in R.
The estrogen signaling target genes were calculated by Cluster-Buster (https://github.com/weng-lab/cluster-buster), while using the motif matrices (MA0112.3, MA0592.3) from JASPAR2022 and UCSC.susScr11 promoter sequence as input. To identify estrogen target genes, 6000 genes in supporting linage were clustered along development stage using the R package Mfuzz. Genes involved in selected clusters were then intersected with estrogen signaling target genes calculated above.
The ligand-receptor pairs were defined based on the UMI count matrix using CellChat54. Due to the unavailability of a pig-specific database, the CellChatDB.human was employed as a ligand-receptor interaction database for subsequent cell-cell communication analysis using default configurations. The Comparation of a total number of interactions and interaction strength were obtained via merging the CellChat objects of each sample by function mergeCellChat.
Statistics and reproducibility
Eight Bama pig testis samples were collected across six different timepoints. Two biological replicates were included at the PD90 and PD150 timepoints. In the computational analysis, the statistical analysis outlined in the specific packages was utilized.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences55 (GSA: CRA014793) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa/. Source data has been provided in Supplementary Data 1. Any additional information will be available from the Jingtao Guo (jingtao.guo@ioz.ac.cn) upon request.
References
Shami, A. N. et al. Single-Cell RNA Sequencing of Human, Macaque, and Mouse Testes Uncovers Conserved and Divergent Features of Mammalian Spermatogenesis. Dev. Cell 54, 529–547.e512 (2020).
Murat, F. et al. The molecular evolution of spermatogenesis across mammals. Nature 613, 308–316 (2023).
Tharmalingam, M. D., Jorgensen, A. & Mitchell, R. T. Experimental models of testicular development and function using human tissue and cells. Mol. Cell Endocrinol. 468, 95–110 (2018).
Lunney, J. K. et al. Importance of the pig as a human biomedical model. Sci. Transl. Med. 13, eabd5758 (2021).
Montgomery, R. A. et al. Results of Two Cases of Pig-to-Human Kidney Xenotransplantation. N. Engl. J. Med. 386, 1889–1898 (2022).
Anand, R. P. et al. Design and testing of a humanized porcine donor for xenotransplantation. Nature 622, 393–401 (2023).
Camacho, P., Fan, H., Liu, Z. & He, J. Q. Large Mammalian Animal Models of Heart Disease. J. Cardiovasc. Dev. Dis. 3, https://doi.org/10.3390/jcdd3040030 (2016).
Schelstraete, W., Devreese, M. & Croubels, S. Comparative toxicokinetics of Fusarium mycotoxins in pigs and humans. Food Chem. Toxicol. 137, 111140 (2020).
Frankenhuis, M. T., Kramer, M. F. & de Rooij, D. G. Spermatogenesis in the boar. Vet. Q 4, 57–61 (1982).
Okwun, O. E., Igboeli, G., Ford, J. J., Lunstra, D. D. & Johnson, L. Number and function of Sertoli cells, number and yield of spermatogonia, and daily sperm production in three breeds of boar. J. Reprod. Fertil. 107, 137–149 (1996).
Koskenniemi, J. J., Virtanen, H. E. & Toppari, J. Testicular growth and development in puberty. Curr. Opin. Endocrinol. Diabetes Obes. 24, 215–224 (2017).
Makela, J. A., Koskenniemi, J. J., Virtanen, H. E. & Toppari, J. Testis Development. Endocr. Rev. 40, 857–905 (2019).
Zhang, L. et al. Single-cell RNA-seq analysis of testicular somatic cell development in pigs. J. Genet Genomics 49, 1016–1028 (2022).
Guo, J. et al. The adult human testis transcriptional cell atlas. Cell Res. 28, 1141–1157 (2018).
Guo, J. et al. The Dynamic Transcriptional Cell Atlas of Testis Development during Human Puberty. Cell Stem Cell 26, 262–276 e264 (2020).
Zhao, J. et al. Cell-fate transition and determination analysis of mouse male germ cells throughout development. Nat. Commun. 12, 6839 (2021).
Wang, M. et al. Single-Cell RNA Sequencing Analysis Reveals Sequential Cell Fate Transition during Human Spermatogenesis. Cell Stem Cell 23, 599–614.e594 (2018).
Green, C. D. et al. A Comprehensive Roadmap of Murine Spermatogenesis Defined by Single-Cell RNA-Seq. Dev. Cell 46, 651–667.e610 (2018).
Zheng, Y. et al. Sertoli cell and spermatogonial development in pigs. J. Anim. Sci. Biotechnol. 13, 45 (2022).
Lee, Y. A. et al. Cryopreservation of porcine spermatogonial stem cells by slow-freezing testis tissue in trehalose. J. Anim. Sci. 92, 984–995 (2014).
Knipscheer, P. et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 326, 1698–1701 (2009).
Kato, Y. et al. FANCB is essential in the male germline and regulates H3K9 methylation on the sex chromosomes during meiosis. Hum. Mol. Genet 24, 5234–5249 (2015).
Chemes, H. E. Infancy is not a quiescent period of testicular development. Int. J. Androl. 24, 2–7 (2001).
Boitani, C., Di Persio, S., Esposito, V. & Vicini, E. Spermatogonial cells: mouse, monkey and man comparison. Semin Cell Dev. Biol. 59, 79–88 (2016).
Maekawa, M., Kamimura, K. & Nagano, T. Peritubular myoid cells in the testis: their structure and function. Arch. Histol. Cytol. 59, 1–13 (1996).
Chen, H., Ge, R. S. & Zirkin, B. R. Leydig cells: From stem cells to aging. Mol. Cell Endocrinol. 306, 9–16 (2009).
Svingen, T. & Koopman, P. Building the mammalian testis: origins, differentiation, and assembly of the component cell populations. Genes Dev. 27, 2409–2426 (2013).
Ademi, H. et al. Deciphering the origins and fates of steroidogenic lineages in the mouse testis. Cell Rep. 39, 110935 (2022).
Chen, M. et al. Integration of single-cell transcriptome and chromatin accessibility of early gonads development among goats, pigs, macaques, and humans. Cell Rep. 41, 111587 (2022).
Shen, Y. C. et al. TCF21(+) mesenchymal cells contribute to testis somatic cell development, homeostasis, and regeneration in mice. Nat. Commun. 12, 3876 (2021).
Shima, Y. et al. Fetal Leydig Cells Persist as an Androgen-Independent Subpopulation in the Postnatal Testis. Mol. Endocrinol. 29, 1581–1593 (2015).
Guo, J. et al. Single-cell analysis of the developing human testis reveals somatic niche cell specification and fetal germline stem cell establishment. Cell Stem Cell 28, 764–778.e764 (2021).
Raeside, J. I. & Renaud, R. L. Estrogen and androgen production by purified Leydig cells of mature boars. Biol. Reprod. 28, 727–733 (1983).
Raeside, J. I. The formation of unconjugated and sulfoconjugated estrogens by Leydig cells of the boar testis. Can. J. Biochem. Cell Biol. 61, 790–795 (1983).
Legacki, E., Conley, A. J., Nitta-Oda, B. J. & Berger, T. Porcine sertoli cell proliferation after androgen receptor inactivation. Biol. Reprod. 92, 93 (2015).
Shah, W. et al. The Molecular Mechanism of Sex Hormones on Sertoli Cell Development and Proliferation. Front. Endocrinol. 12, 648141 (2021).
Martins, A. D. et al. Control of Sertoli cell metabolism by sex steroid hormones is mediated through modulation in glycolysis-related transporters and enzymes. Cell Tissue Res. 354, 861–868 (2013).
Toocheck, C. et al. Mouse Spermatogenesis Requires Classical and Nonclassical Testosterone Signaling. Biol. Reprod. 94, 11 (2016).
Li, M. W. et al. Tumor necrosis factor alpha reversibly disrupts the blood-testis barrier and impairs Sertoli-germ cell adhesion in the seminiferous epithelium of adult rat testes. J. Endocrinol. 190, 313–329 (2006).
Kumar, R. et al. The dynamic structure of the estrogen receptor. J. Amino Acids 2011, 812540 (2011).
Giguere, V. Transcriptional control of energy homeostasis by the estrogen-related receptors. Endocr. Rev. 29, 677–696 (2008).
Xia, H., Dufour, C. R. & Giguere, V. ERRalpha as a Bridge Between Transcription and Function: Role in Liver Metabolism and Disease. Front Endocrinol. (Lausanne) 10, 206 (2019).
Sangeeta, K. & Yenugu, S. Male reproductive tract antimicrobial expression in the extremes of ages of rats. Gene 710, 218–232 (2019).
Hu, S. G. et al. Androgenic regulation of beta-defensins in the mouse epididymis. Reprod. Biol. Endocrinol. 12, 76 (2014).
Com, E. et al. Expression of antimicrobial defensins in the male reproductive tract of rats, mice, and humans. Biol. Reprod. 68, 95–104 (2003).
Mori, C. et al. Morphological analysis of germ cell apoptosis during postnatal testis development in normal and Hsp 70-2 knockout mice. Dev. Dyn. 208, 125–136 (1997).
Hemsworth, P. H. & Tilbrook, A. J. Sexual behavior of male pigs. Horm. Behav. 52, 39–44 (2007).
Nie, X. et al. Single-cell analysis of human testis aging and correlation with elevated body mass index. Dev. Cell 57, 1160–1176.e1165 (2022).
Zhang, L. et al. Single-cell RNA-sequencing reveals the dynamic process and novel markers in porcine spermatogenesis. J. Anim. Sci. Biotechnol. 12, 122 (2021).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Members, C.-N. & Partners. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2023. Nucleic Acids Res. 51, D18–D28 (2023).
Acknowledgements
This project was funded by the National Key R&D Program of China (2022YFA1104100 to J.G., 2021YFA0805902 to J.Z.), the National Natural Science Foundation of China (NSFC) (31925036, 32230100, 82301816), the Science and Technology Innovation 2030 - Major Projects (2023ZD0407503 to C.H), the Initiative Scientific Research Program supported by the Institute of Zoology, Chinese Academy of Sciences (2023IOZ0102 and 2023IOZ0306), and China Postdoctoral Science Foundation (2022M713135 to X.W.). We thank members of Guo laboratory, Han laboratory, and Zhao laboratory for discussion. Some of the schematic illustrations were modified and/or created with images from https://clipart-library.com/ or https://www.10xgenomics.com/.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.G., J.Z., and C.H.; computational analysis, X.W., Y.G. and J.Yue; experiments and sample preparation, Yang W., Yu W., R.Z., S.H. and J.Yao; writing – original draft, X.W., Yang W., Yu W., and Y.G.; and supervision, J.G., J.Z., and C.H. All authors contributed to manuscript revision, read, and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Jon Oatley and the other, anonymous, reviewer for their contribution to the peer review of this work. Primary Handling Editors: Simona Chera and David Favero.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Wang, Y., Wang, Y. et al. Single-cell transcriptomic and cross-species comparison analyses reveal distinct molecular changes of porcine testes during puberty. Commun Biol 7, 1478 (2024). https://doi.org/10.1038/s42003-024-07163-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-07163-9
This article is cited by
-
METTL3 regulates Leydig cell proliferation via miR-145-PCK1 mediated gluconeogenesis in goats
Journal of Animal Science and Biotechnology (2026)
-
Single-nucleus RNA sequencing provides insights into the genetic mechanisms underlying reproductive adaptability in Tibetan sheep (Ovis aries)
Communications Biology (2026)
-
Unique Sertoli cell adaptations support enhanced spermatogenesis in chickens
Journal of Animal Science and Biotechnology (2025)
-
Comprehensive analysis of long non-coding RNA and mRNA expression patterns during seminiferous tubules maturation in Guanzhong dairy goats
BMC Genomics (2025)
-
Single-cell RNA sequencing uncovers dynamic roadmap during chicken spermatogenesis
BMC Genomics (2025)