Abstract
Future pandemic threats may be caused by novel coronaviruses and influenza A viruses. Here we show that when directly added to a cell culture, 12mer guanine RNA (G12) and its phosphorothioate-linked derivatives (G12(S)), rapidly entered cytoplasm and suppressed the propagation of human coronaviruses and influenza A viruses to between 1/100 and nearly 1/1000 of normal virus infectivity without cellular toxicity and induction of innate immunity. Moreover, G12(S) alleviated the weight loss caused by coronavirus infection in mice. G12(S) might exhibit a stable G-tetrad with left-handed parallel-stranded G-quadruplex, and inhibit the replication process by impeding interaction between viral nucleoproteins and viral RNA in the cytoplasm. Unlike previous antiviral strategies that target the G-quadruplexes of the viral genome, we now show that excess exogenous G-quadruplex-forming small RNA displaces genomic RNA from ribonucleoprotein, effectively inhibiting viral replication. The approach has the potential to facilitate the creation of versatile middle-molecule antivirals featuring lipid nanoparticle-free delivery.

Similar content being viewed by others
Introduction
Although medical advances, accessible health care, and sanitation improvements over the last two decades have reduced mortality from common infectious diseases, severe infectious disease outbreaks have impacted lives in the 21st century. Recent examples include outbreaks of severe acute respiratory syndrome (SARS) in 2003, swine-derived influenza A virus (IAV) disease in 2009, Middle East respiratory syndrome (MERS) in 2012, Ebola virus disease in West Africa between 2013 and 2016, zika virus disease in 2015, and the ongoing coronavirus infectious disease outbreak that began in 2019 (COVID-19). Changes in the global climate and human activities enhance the morbidity and mortality linked to emerging and reemerging infectious diseases1. Pandemic threats from emerging zoonotic diseases are particularly caused by novel coronaviruses (CoVs)2 and novel IAVs3.
CoVs infect humans, livestock, and wildlife (including mammals and birds), resulting in respiratory and enteric diseases. Human coronaviruses (HCoVs) that primarily infect upper respiratory tracts include HCoV-229E (229E), HCoV-OC43 (OC43), HCoV-NL63, and HCoV-HKU14,5. One report indicated that these HCoVs were causative agents of 15–29% of common colds6. In contrast, SARS-CoV-1 (2002), MERS-CoV (2012), and SARS-CoV-2 (2019) are causative agents of severe pneumonia. Most of these CoVs are thought to have originated from the spread of bat CoVs, transferred to intermediate animal hosts, and then spread through human-to-human transmission2.
IAV is a constantly evolving pathogen that evades host immunity by altering its antigenicity via two mechanisms. Antigenic drift is the process of antigenic changes by the amino acid substitution of the viral surface antigens hemagglutinin (HA) and neuraminidase (NA) due to replication error. The 2009 A(H1N1) pandemic IAV was driven by antigenic shifts caused by the co-infection of swine airway epithelial cells with different serotypes of IAVs, resulting in segmental genomic reassertment in different types of HA and/or NA7. The establishment of such human-to-human transmission of reassortant IAV—never experienced by humans—could lead to a new epidemic of this IAV3,8.
Considering these circumstances, it will be difficult to prevent the outbreak of novel zoonoses such as IAV and CoV infections in the future. Thus, the development of multiple effective preventive and therapeutic methods to respond to such outbreaks is required. Effective anti-IAV drugs that target the replication steps of IAV have been used. However, mutants often raise viruses resistant to antivirals9,10. It is possible that middle molecule drugs not only interact with broader regions of viral proteins to inhibit their function11 but also resist drug evasion through adaptive mutations. The NPs of CoV and IAV interact with single-stranded genomic RNA to form viral ribonucleoprotein (vRNP), which is the core of the enveloped viral particles. The NP-RNA interaction is required for replication, viral RNA (vRNA) resistance to cellular RNase, and the assembly of the viral genome as vRNP within the particles. Therefore, drugs targeting vRNPs may serve as potential antiviral drug candidates12.
We aimed to develop the antiviral agents of middle-molecular-weight nucleic acids that inhibit the interaction between vRNA and NP—a common property against coronavirus and IAV. We previously found that a 12mer guanine RNA (G12) inhibited NP-RNA-induced liquid-liquid phase separation (LLPS), droplet formation in vitro13. Here, we demonstrate that G12 and phosphorothioate (PS)-modified G12 (G12(S)) directly enters the cell and can inhibit the propagation of the three different HCoVs (OC43, 229E, and SARS-CoV-2) and two IAV variants (H3N1 and H1N1). In the case of CoV, G12(S) is likely to promote the release of non-infectious vRNA outside the cells and reduces the infectious titer. On the other hand, G12(S) inhibits intracellular transport of IAV vRNPs. Multiple intranasal administrations of G12(S) were found to significantly reduce the weight loss induced by intranasal infection with OC43. Guanine-rich nucleotide sequences are known to form a four-guanine plane (G-tetrad) via Hoogsteen hydrogen bonds, as well as multiple G-tetrads stacks as a G-quadruplex that is stabilized by alkali metal ions (e.g., K+ or Na+). We found that in response to K+, G12(S) formed a stable structure that might be similar to recently identified parallel-stranded G-quadruplexes with left-handed helicity14,15. An aptamer identified as anti-Human immunodeficiency virus (HIV) integarase inhibition called 93del (16mer DNA) has been shown to reduce infectivity of HIV at multiple steps including direct action to the HIV particle16. Interestingly, we showed that PS-modified RNA (PS-RNA) forms of several 12–18mer nucleotide sequences, including 93del, known to form G-quadruplexes, had anti-IAV activity, although G12(S) had the strongest antiviral effect among the PS-RNAs. Indeed, G-Quadruplex formation level in cytoplasm correlated anti-viral activity. Our results suggest that cytoplasmic vRNP assembly and/or vRNP transport may be inhibited by the G12(S) G-quadruplex.
Since putative G-quadruplex forming sequences are present in all viral genomes, antiviral effects of G-quadruplex binding small molecules (G-quadruplex binders) have been investigated17, and their effect on viral replication were observed with Zika virus18,19. Unlike the previous idea of targeting the G-quadruplex on the viral genomes by G-quadruplex binder, the mechanism herein is that excess G-quadruplex forming small-RNA itself binds to NP that originally interacts with genomic RNA, and replaces with the genomic RNA away. The approach presented herein could represent an alternative modality for broad-spectrum antivirals.
Results and discussion
Based on the fact that the absence of a consecutive guanosine sequence of 6mer or longer nucleotide sequences in a ~30 k-base genome sequence of HCoV, we have previously found and 12mer guanine RNA (G12; GGGGGGGGGGGG) strongly inhibited the in vitro interaction between SARS-CoV-2 NPs and vRNA13. Therefore, we investigated the effects of G12 on viral propagation in cells.
To examine the direct incorporation of the guanine RNA oligomers, G12 was fluorescently labeled with 5-TAMRA (5-Carboxytetramethylrhodamine) to observe its uptake and behavior using human lung cancer A549 cells. The results presented in Fig. 1a indicate that the fluorescence of TAMRA-G12 was observed within the cells. Since the PS modification of oligonucleotides increases nuclease resistance and uptake by cells20, we modified G12 by using PS linkages (G12(S)). Fluorescence of TAMRA-G12(S) was also observed, and, additional ten-fold unlabeled G12(S) competed with the fluorescence of TAMRA-G12(S) (Fig. 1b). This is because the cellular uptake of G12(S) and TAMRA-G12(S) might be a similar mechanism. The time-lapse experiment suggested that TAMRA-G12(S) was rapidly taken up within seconds and the level was increased in a time-dependent manner within the cells (Fig. 1c). Fluorescence recovery after photobleaching (FRAP) experiments (Fig. 1d) showed gradual recovery of the fluorescence in the photobleached region from 1 min, and the rate is likely to be similar to the uptake rate. Additionally, TAMRA-G12(S) fluorescence was excluded from the nucleus and endoplasmic reticulum (Fig. 1e). These results demonstrate that G12(S) rapidly entered cells without using lipid nanoparticles (LNP) and was distributed in the cytoplasm in a soluble state.
Images of the TAMRA-G12, TAMRA-G12(S), and nuclei (Hoechst 33342) in A549 cells were observed at the times indicated below and are shown in green (TAMRA) and blue (Hoechst 33342) pseudocolors, respectively (a–e). a Cellular uptake of TAMRA-G12 (2 μM, 30 min). b Uptake of TAMRA-G12(S) (0.2 μM) in the absence or presence of a 10-fold concentration of unlabeled G12(S) (2 μM) for 1 h. c Time-dependent incorporation of TAMRA-G12(S) signals from 0 to 15 min after the addition of 2 μM TAMRA-G12(S). d FRAP analysis of TAMRA-G12(S). Fluorescence was bleached for 5 min at 100% laser power. The photobleached area is marked with a white circle in the leftmost image. Images were acquired over time for the times (min) shown in the figure. e Distribution of TAMRA-G12(S) (green) and ER-Tracker Red (red). Cells were treated with 2 μM TAMRA-G12(S) in the medium for 1 h and further incubated with 1 μM ER-Tracker Red and NucBlue (Hoechst 33342). Scale bars are shown in (a–e). f Cytotoxicity of G12(S) on A549. The cell viabilities were examined after 24 h of culture in the presence of 0–10 μM G12(S) using AlamerBlue. One-way analysis of variance (ANOVA) was used for the statistical analysis (n = 3, P-values = 0.3582, F-value = 1.230). There is no significant difference between the combinations. g Effect of G12(S) on protein synthesis in A549 were determined using the puromycin uptake assay SuNSET method21. Cells were cultured as a condition of the virus infection in the presence of 1 and 2 μM G12(S) for 24 h. Cycloheximide (10 μg/mL CHM) instead of G12(S) was used to inhibit protein synthesis as a control. See Supplementary Fig. 1f for western blotting data for the SuNSET. Reporter gene assays for IFN-β promoter (h) and a promoter containing the NFκB binding sites (i) in HEK293T cells. The values shown are relative values when the control (without G12(S) and poly I:C) is set as “1”. The reporter genes activated by polyI:C were not induced by 2 μM G12(S), but rather suppressed. Statistical analysis of the significance for (g–i) were carried out by One-Way ANOVA (g n = 3, F(3,8) = 43.26, P < 0.0001; h n = 3, F(3, 8) = 67.30, P < 0.0001; i n = 3, F (3, 8) = 290.1, P < 0.0001), and Dunnett’s multiple comparison test (g) or Tukey’s multiple comparison test (h, i). Data are shown by Box-and-whisker plot are used with P-values for the multiple comparison (f–i).
G12(S) and G12 do not affect cellular viability, or protein synthesis
Next, we examined whether G12(S) shows cytotoxicity. As shown in Fig. 1f and Supplementary Fig. 1a, the cell proliferation of A549, HCT-8, LLC-MK2 and MDCK cells was not reduced by 1–10 μM G12(S) contained in the culture medium. We then examined the effects of G12(S) on cellular protein synthesis using the puromycin uptake assay SuNSET21. As shown in Fig. 1g and Supplementary Fig. 1b–g, G12(S) did not inhibit protein synthesis.
Cells recognize internal and external nucleic acids with toll-like receptors (TLRs) and RIG (Retinoic acid inducible gene-I)-like receptors (RLRs) and induce interferon-β and cytokines22. We then examined whether interferon-β promoter and NFκB-dependent promoter activities were affected by G12(S) using a reporter gene assay. These reporter genes were not activated by G12(S) (Fig. 1h, i). Furthermore, G12(S) even reduced the reporter gene activities that were induced by the inducer polyI:C, suggesting that the antiviral activities of G12(S) (see following sessions) are unlikely due to the induction of innate immunity driven by signaling from TLRs and RLRs. Collectively, neither cytotoxicity including innate immunity nor the downregulation of protein synthesis was observed due to G12(S).
Inhibition of β-coronavirus OC43 replication in infected cells by G12 and G12(S)
The β-coronavirus genome is a single-stranded positive-sense RNA encoding four structural proteins: spike (S) protein, envelope (E) protein, membrane (M) protein, and NP23,24. The NP is the most abundant viral protein in infected cells25 and binds to genomic RNA to form vRNP for packaging. The architecture of the SARS-CoV-2 RNA–NP ultrastructure is likely based on protein–protein interaction and the interaction of the NP and vRNA26.
We first examined the effect of G12 on β-coronavirus OC43 replication. The addition of naked G12 (0.2 and 2 μM) to a culture medium of human ileocecal adenocarcinoma HCT-8 cells infected with OC43 strongly reduced the expression of both cellular OC43 NP and RNA (Supplementary Fig. 2a–c) in a concentration-dependent manner. A time-of-addition experiment revealed that G12 has no direct effect on viral particles to reduce the infectivity; however, the constitutive presence of G12 in the culture medium was required for suppression of the NP level (Supplementary Fig. 2d–f). Therefore, we investigated the antiviral effects of G12 or G12(S) after washing and the medium exchange of post-virus-infected cells in all experiments in this paper unless otherwise noted.
As shown in Fig. 2a–c, OC43 NP levels were significantly reduced in the presence of 2 µM G12(S) over 5 days. Upon comparing Supplementary Figs. 2a to 2c and Fig. 2a to 2d, it was found that G12(S) exhibited stronger anti-OC43 activity than G12. The reduction of OC43 NP-RNA in the infected cells (Fig. 2d left) and the infectivity (plaque forming unit (pfu)) of OC43 produced in the medium (Fig. 2e) were nearly 10−3-fold in the presence of G12(S). However, the RNA in the medium was reduced by less than 10-fold (Fig. 2d right). Ratios of the medium RNA to the virus infectivity (RNA/pfu) were enhanced in the presence of G12(S) (Fig. 2f). The effect of G12(S) was also exhibited in other cell line: the cytopathic effect (CPE) of OC43 on human embryonic lung fibroblast MRC-5 cells27 was strongly suppressed by the presence of 2 µM G12(S) in the medium (Fig. 2g). Additionally, the fluorescently-labeled G12(S) (TAMRA-G12(S)), which can enter the cytoplasm also reduced OC43 replication in A549 cells (Supplementary Fig. 2g).
OC43 NP levels in HCT-8 cells infected with OC43 (multicity of infection (MOI) = 0.01, n = 3) in the absence (a) and presence of 2 μM G12(S) in the medium (b). Cell lysates were prepared at 2, 3, 4, and 5 days post-infection (dpi) in the absence (a) and presence of 2 μM G12(S) in the medium (b), and determined by western blotting with anti-OC43 NP and anti-actin antibodies. c Changes in NP/actin ratio (n = 3, normalized with the ratio of (a) at 2 dpi as 1) without (−, from a) and with (+, from b) G12(S). d OC43 RNA levels in OC43-infected (MOI = 0.01) cells (left) and culture medium (right) in the absence (control) and in the presence of 2 μM G12(S). e Effect of G12(S) on OC43 virus titers (pfu) of the medium. f OC43 RNA levels in the culture medium. Statistical analyses of the significance of G12(S) effect were performed by Two-Way ANOVA and data are shown by box-and-whisker plots including all data plots (c–f) (c n = 3, F(3,8) = 25.28, P = 0.0002; d left, n = 3, F(1,4) = 250.8, P < 0.0001; d right, n = 3, F(1,4) = 1800, P < 0.0001; e n = 3, F(1,4) = 5.245, P = 0.0838; f n = 3, F(1,4) = 77.53, P = 0.0009). Sidak’s multiple comparison test was used and the P-values are indicated. g Effect of 1 and 2 μM G12(S) on the CPE of OC43 (MOI = 0.1)-infected MRC5 cells (3 dpi). h OC43 infection in nasal cavities and olfactory bulbs of newborn mice. Immunofluorescence images of OC43 NP (yellow) and nucleus (DAPI, blue) are shown. Scale bars indicate 1 mm (i) and 20 μm (iii, iv) lengths. See Supplementary Fig. 3 for entire images. i, j Prevention of mice toxicity due to OC43 mouse infection by G12(S). OC43 infection and single-dose (i, n = 22) and three-dose (j, n = 13) schedules of G12(S) are shown. The survival rate is shown as Kaplan–Meier curves. P-values were determined by Logrank test.
Previous reports have indicated that a mouse-adapted OC43 strain is infective to infant mice via nasal infection28,29. Thus, we examined the effect of G12(S) in the nasal infection system of OC43 in suckling mice. We found that 7-day-old C57BL/6 mice were also susceptible to OC43 (VR-1558) from American Type Culture Collection (ATCC). As shown in Fig. 2h, NP signals (yellow) were observed in the nasal cavity (i, ii), the olfactory bulb (i), the brain (i), and the surface of the nasal cavity (ii–vi). These observations were consistent with those previously described by Dubé et al.28. Nasal OC43 infection stunted mouse development, as indicated by Kaplan–Meier curves, which were plotted with 20% weight loss of the endpoints (Fig. 2i, j). We performed the intranasal pretreatment of G12(S) on OC43-infected mice and additional repeated nasal treatments of G12(S) for 2 days. G12(S) treatments significantly restored the stunting of mice due to OC43 infection (Fig. 2i), and a more pronounced effect was observed with repeated dosing (Fig. 2j).
G12(S) reduces α-coronavirus 229E and β-coronavirus SARS-CoV-2 replication
CoV is genetically subdivided into four groups: alfa, beta, delta, and gamma. The NP is structurally and functionally homologous among the genetic groups of CoV and is responsible for forming vRNP. The seven HCoVs belong to the alpha and beta groups30. To investigate whether the effect of G12(S) occurs across the groups and strains, we examined the effect of G12(S) on an alpha CoV 229E and SARS-CoV-2 as a beta coronavirus other than OC43. As shown in Fig. 3a, b left, 229E intracellular viral protein (NP) and vRNA decreased. Even though vRNA levels in the medium had a significant but relatively small effect (Fig. 3b, right), G12(S) reduced medium infectivity by nearly two orders of magnitude at 3 days post-infection (dpi) (Fig. 3c). Ratio of the medium RNA to the pfu at 3 dpi was enhanced by G12(S) (Fig. 3d).
a 229E NP levels in 229E-infected LLC-MK2 cells after 2, 3, and 4 dpi were determined by western blotting. The graph indicates changes in NP/actin ratio (n = 3, normalized with the ratio of G12(S) (−) at 2 dpi as 1) of western blotting experiments. b 229E RNA in the infected cells (left) and in the culture medium (right). c Reduction of 229E virus titers in the presence of G12(S) in the medium. LLC-MK2 cells were infected by 229E for 3 and 4 dpi. d 229E RNA number in the culture media per viral infectious titer are shown. Reduction of SARS-CoV-2 virus RNA (e) and titers (f) in the medium of the virus-infected VeroE6/TMPRSS2 cells (MOI = 0.01) in the presence of 1 and 2 μM G12(S) at 24 and 48 hpi. g SARS-CoV-2 medium RNA per infection titer (pfu). Data are shown box-and-whisker plots including all data plots with the P-values. Statistical analyses of the G12(S) effect by Two-Way ANOVA (a n = 3, F(1,4) = 116.5, P = 0.0004); b left, n = 3, F(1, 2) = 1015, P < 0.0001; b right, n = 3, F(1,4) = 60.59, P = 0.0015; c n = 3, F(1,4) = 13.62, P = 0.0210; d n = 3, F(1, 4) = 818.5, P < 0.0001) or by REML model (e n = 3, F(2, 6) = 6.227, P = 0.0344; f n = 3, F(2, 6) = 11.20, P = 0.0094; g n = 3, F(2, 12) = 24.12, P < 0.0001). Sidak’s multiple comparison test (a–c) and Dunnett’s multiple comparison test (e–g) were used and the P-values are indicated.
Furthermore, G12(S) also significantly suppressed SARS-CoV-2 proliferation. The virus infection titer (pfu) was suppressed in a concentration-dependent manner and decreased to about 1/1000 at 2 μM for 24 h (Fig. 3f). However, effect of G12(S) on vRNA level in the culture medium was small at 24–48 hours post infection (hpi) (Fig. 3e). Our results indicated that, although G12(S) suppressed the production of infectious viruses in the culture media, its efficiency in suppressing the release of vRNA is low—a phenomenon common to OC43 (Fig. 2f), 229E (Fig. 3d), and SARS-CoV-2 (Fig. 3g). These results suggest that G12(S) likely inhibits the production of the infectious viruses of HCoV family in a similar manner.
G12(S) affects the localization of NP in HCoV-OC43-infected cells
We hypothesized that G12(S) directly affects the CoV replication process in the cytoplasm resulting in inhibition of infectious coronavirus production (see Supplementary Fig. 4). CoV genomic RNA replicates within double-membrane vesicles (DMVs) originating from the endoplasmic reticulum (ER)31,32,33. The DMVs are present near the ER31. Newly synthesized genomic RNA is transported from the DMVs into the cytoplasm through pores embedded into the DMV membrane, and subsequently associates with NP to form vRNP. The vRNP is then imported into the ER-Golgi intermediate (ERGIC) for viral packaging. We examined the distribution of NP and an ER marker (Lys-Asp-Glu-Leu (KDEL) proteins) as well as an ERGIC marker (ERGIC53). A549 cells were susceptible to OC43 (Supplementary Fig. 5a, b). OC43-infected A549 cells were cultured for 48 h and then treated with G12(S) for 24 h. OC43 RNA and NP levels were attenuated by this condition (Supplementary Fig. 5c, d). Cells were fixed and immunostained with KDEL (Fig. 4a, b) and ERGIC53 (Fig. 4c, d) for ER and ERGIC markers, respectively. In OC43-infected A549 cells, NP was distributed in the cytoplasm and had non-overlapping distribution with the ER (Fig. 4a) and the ERGIC (Fig. 4c) markers. However, in the presence of G12(S), there were multiple dots overlapping NP and ER (Fig. 4b) or ERGIC (Fig. 4d), suggesting that G12(S) may retard the viral production process associated with NP.
a–d Subcellular localization of OC43 NP and EIGIC-53 or ER (KDEL) proteins. A549 cells infected with OC43 (MOI = 1) were cultured for 48 h. The cells were treated with 2 μM G12(S) for the next 24 h of the culture, and fixed. Immunostaining was performed for OC43 NP with KEDL proteins (a and b) or ERGIC53 (c and d). See Supplementary Fig. 5e for the specificities of the antibody used for OC43 NP, ERGIC-53, and ER protein (anti-KDEL antibody). e Experimental overview of LLPS formation and G12(S) treatment on the LLPS. Droplets of SARS-CoV-2 NP were formed by LLPS in the presence of 1 µM FITC-32 base of the 3’-untranslated region of SARS-CoV-2 RNA and 40 µM SARS-CoV-2 NP overnight. f Effect of TAMRA-G12(S) on droplets of SARS-CoV-2 NP formed by LLPS in the presence of 1 µM FITC-leveled vRNA. Fluorescence and differential interference contrast (DIC), TAMRA (magenta) and FITC (Green) and merged (Merge) images are shown. TAMRA-G12(S) (final 1, 3, and 6 µM) were added and observed at 25 °C using fluorescence and DIC microscopy at the indicated time. See Supplementary Fig. 6 for the time-dependent image data (10 min, 20 min, 30 min, 1 h, 2 h, 3 h, 4 h, 5 h, 6 h and 7 h). g The supernatant (soluble fraction) was collected from the wells forming LLPS 1 h after addition of TAMRA-G12(S) (final 0, 3, and 6 µM), and the adsorbed fraction was collected by suspending the wells in LLPS buffer containing 0.1%SDS. These samples were separated in 7.5 M urea/10 mM KCl containing 15% PAGE. Fluorescent images were taken with a ChemiDoc Touch MP (BioRad) using filter sets for Alexa546 (left) and fluorescein (right). Tamura and FITC fluorescence were observed using the Aelxa546 mode. FITC fluorescence was observed in fluorescein mode (center). An overlay image is shown (right). The positions of ssRNA marker lengths are indicated.
Effect of G12(S) on SARS-CoV-2 NP-RNA interaction in vitro
Previously, we showed that G12(S) detached the vRNA from droplets formed by SARS-CoV-2 NPs and vRNA through LLPS while also destroying the droplets at high concentrations, suggesting that G12(S) might replace the preexisting genomic RNA in the vRNP13. Here, we examined whether TAMRA-labeled G12(S) could replace the fluorescein isothiocyanate (FITC)-labeled vRNA within the droplets formed by the LLPS of SARS-CoV-2 NP-RNA (Fig. 4e). As shown in Fig. 4f, at low concentrations (1 and 3 μM; nucleotide concentrations lower and comparable to FITC-vRNA), TAMRA-G12(S) was incorporated into droplets over a longer period of time (3–5 h) (1 and 3 μM). At even higher concentrations (6 μM), TAMRA-G12(S) itself formed droplets and soaked into the FITC droplets within 20 min. Finally, within 1 h, TAMRA-G12(S) surpassed the levels of FITC-vRNA in the droplets. Both TAMRA-G12(S) and FITC-vRNA were stable in the NP containing mixture (Supplementary Fig. 7). TAMRA-G12(S) (6 μM) was found to be incorporated into a smear band in 7.5 M urea-containing PAGE and to aggregate at the migration start of 15% urea PAGE (Fig. 4g, lane 6, adsorbed (LLPS) fraction). TAMRA-G12(S) (3 and 6 μM) reduced FITC-vRNA levels in the adsorbed fractions (Fig. 4g, lanes 5 and 6). Taken together, TAMRA-G12(S) might be preferentially incorporate within the LLPS droplets and replace with NP-bound vRNA.
In all three tested HCoVs (OC43, 229E, and SARS-CoV-2), G12(S) can repress infectious virus production despite its inefficient suppression of vRNA levels in the culture medium. One possibility is that G12(S) prohibits accurate vRNP formation, resulting in the production of non-infectious vRNA-containing CoV particles. The other possibility is that the export of vRNA itself—without NP—was facilitated by the inhibitory activity of G12(S).
Inhibition of IAV propagation by G12 and G12(S)
We predicted that the inhibitory activity of G12 is not limited to CoVs but also affects other RNA viruses that retain vRNP in the cytoplasm during the replication process. Although IAV has a different replication cycle than CoV, they share the presence of vRNPs in the cytoplasm before virus packaging. After the formation of each of the eight segmented genomic vRNPs constituting the vRNA-NP complex with RNA polymerase subunits transported from the nucleus, the vRNPs were transported to the plasma membrane site, where viral packaging occurs (see Supplementary Fig. 8a; for a review, see Dou et al.34). We noticed that consecutive guanine sequences of 6mer or longer were also absent in the negative strand segmented genomic RNA (the total length of 8 segments genome RNA is 13.6 k bases), as well as in A/Aichi/2/68 (H3N2) and A/California/04/09 (H1N1pdm09).
Interestingly, from 15 h to 24 h post infection, intracellular IAV NP and HA proteins (Fig. 5a) and NP and HA RNAs (Fig. 5b) were reduced up to 10-fold by G12(S). Time-of-addition experiments showed that intracellular HA protein and infectious IAV production were efficiently suppressed only in the constitutive presence of G12 during IAV propagation, but not by pretreatment of cells or direct treatment of IAV stocks (Supplementary Fig. 8b–f). Furthermore, extracellular NP and HA RNAs were more efficiently reduced by approximately 100-fold by G12(S) (Fig. 5c). In addition, infectious virus in the medium were reduced to a similar extent in the presence of G12(S) (Fig. 5g). These result suggested that IAV production in the medium might be efficiently suppressed by G12(S). Because the conditions used (MOI = 0.01, 24 h) resulted in repeated IAV infectious cycles, we next performed experiments under single-replicative cycle conditions (Fig. 5d). Both intracellular positive and negative strand RNA were unaffected by the presence of G12(S) (Fig. 5e and Supplementary Fig. 8g), whereas extracellular viral RNA ((-)RNA) production at 6 hpi was suppressed in the presence of 0.5 and 2 μM G12(S) (Fig. 5f). Therefore, it was suspected that G12(S) might inhibit cytoplasmic event such as vRNP transport and/or viral formation resulting in the reduction of infectious IAV.
a, b Effect of IAV protein and RNA levels (NP and HA) in IAV (A/Aichi/2/68 (H3N2))-infected MDCK cells by 2 μM G12(S). The time-dependent levels at 15, 18, 21, and 24 hpi were determined. Relative NP and HA levels were quantified from the western blotting images (normalized by actin). The RNA copy numbers in cells were determined as described in the Materials and Methods (b). c Level of IAV RNA (NP and HA) in the culture medium of each well at 15, 18, 21 and 24 hpi. d To avoid repeated infections, MDCK cells were infected with IAV at an MOI of 1 in medium lacking trypsin, which is required for nascent IAV infectivity. e Effect of 0.5 and 2 μM of G12(S) on (+) strand and (−) strand NP RNA levels in cells at 4 and 6 hpi were determined. f Inhibitory effect of G12(S) on (−) strand NP RNA in the culture medium at 6 hpi was determined. g Pfu of the culture media of MDCK infected IAV (H3N2 Aich) (MOI = 0.01) at 24 hpi was determined in the presence of 2 µM G12(S). h Pfu of the culture media of MDCK infected (MOI = 0.01) with mouse-adapted A/California/04/09 (H1N1pdm09) and mouse-adapted A/Aichi/2/68(H3N2) (24 hpi). i Dose-response (inhibition of CPE) curve obtained by treating IAV (A/Aichi/2/68 (H3N2))-infected MDCK cells (MOI = 0.01, N = 3) with oseltamivir (Tamiflu), laninamivir (Inavir), and G12(S) at 20 μM and six 4-fold dilutions for 27 h are shown. Because IAV infection caused cells to detach from the plate surface, cells remaining on the plate were quantified by crystal violet staining and their absorbance. The dose-response curves (n = 4, +/−SD) were fitted by nonlinear regression (curve fit) using log (drugs) vs. response variable slope (four parameters). The results and EC50 were obtained by three independent experiments (See Supplementary Fig. 9). j Pfu of the culture medium of MDCK infected IAV (H3N2 Aich) (MOI = 0.01) in the presence of 0.5, 1.0 and 2.0 µM G12(S), Tamiful and Inavir at 22 hpi was determined. Data (a–c, e–h) are shown by box-and-whisker plots including all data plots with the P-values. Statistical analyses of the G12(S) effect by Two-Way ANOVA (a left, n = 3, F(3, 12) = 36.68, P < 0.0001; a right, n = 3, F (1, 4) = 38.59, P = 0.0034; b left, n = 3, F(1, 4) = 2025, P < 0.0001; b right, n = 3, F(1, 4) = 712.1, P < 0.0001; c left, n = 3, F(1, 4) = 528.9, P < 0.0001; c right, n = 3, F(1, 4) = 410.5, P < 0.0001; e left, n = 6, F(2,15) = 1.239, P = 0.3177; e right, n = 6, F(2, 15) = 0.2740, P = 0.2740), or by One-Way ANOVA (f n = 6 or 8, F(2, 17) = 21.92, P < 0.0001; h left, n = 3, F(2, 6) = 18.06, P = 0.0029; h right, n = 3, F(2, 6) = 16.68, P = 0.0035). Two-tailed t-test was used for g (F(2, 2) = 2006, P-value = 0.0010). Sidak’s multiple comparison test (a–c), Tukey’s multiple comparison test (e) or Dunnett’s multiple comparison test (f, h) were used and the P-values are indicated.
Moreover, the propagation of infectious viruses of other IAV strains (H1N1 and H3N2) was also suppressed by G12(S) in a concentration-dependent manner (Fig. 5h), suggesting that G12(S) has broad specificity on the anti-IAV effects. We compared anti-IAV activity of G12(S) with used anti-influenza drugs35,36. Dose-CPE curve in 26 h infection cycle (MOI = 0.01) by G12(S) indicated EC50 of G12(S) was 0.243+/−0.0473 μM (n = 3, +/−SD), whereas that of Tamiful (Oseltamivir) and Inavir (Laninamivir) showed 2.142 μM (2.359 μM) and 3.844 μM (2.358 μM), respectively (Fig. 5i, see Supplementary Fig. 9 for the raw data and replicates). In addition, the reduction in infectious virus production during 28 h infection cycle (MOI = 0.01) by 0.5, 1.0 and 2.0 uM of G12(S) tended to be stronger than that of Tamiflu and Inavir (Fig. 5j).
Inhibitory effect of G12 and G12(S) on the cytoplasmic transport of IAV vRNPs
Next, we focused on the IAV vRNP transport in the cytoplasm. Upon export from the nucleus, neo-synthesized RNPs are transiently accumulated in a prenuclear region near the microtubule organizing center (MTOC) and endocytic recycling compartment (ERC) where Rab11A GTPase is present37,38,39,40 (see Supplementary Fig. 8a). Rab11A is the master regulator of ERC, and is responsible for RNP transport in IAV-infected cells via the Rab11A-dependent recycling endosome to reach the cytoplasmic side of the plasma membrane where viral packaging occurs40. Additionally, a recent finding indicating the coexistence of Rab11A and IAV vRNAs on modified, tubulated ER (called irregularly coated vesicles (ICVs)) throughout the cells (including plasma membrane) suggests that Rab11A-dependent ICVs are responsible for IAV vRNP transport41. In any case, cytoplasmic IAVs interact with Rab11A during the transport step of the vRNP.
We then examined the effects of G12 and G12(S) on the cellular colocalization of IAV NP and Rab11A via live-cell imaging. mCherry-Rab11A and EGFP-NP were simultaneously expressed from a bicistronic gene in which an internal ribosome entry site (IRES) sequence was inserted between these two coding regions (Fig. 6a). mCherry-Rab11A showed a membranous reticulated structure throughout the cytoplasm in uninfected cells, whereas the EGFP-NP was localized in the nucleus (Fig. 6b (i) and Supplementary Fig. 10 (i). As previously reported42, upon infection with IAV, nuclear EGFP-NP is exported from the nucleus and interacts with mCherry-Rab11A in the perinuclear region (MTOC), resulting in punctuated colocalization throughout the cytoplasm (Fig. 6b (ii) and Supplementary Fig. 10 (ii)). This indicates that EGFP-NP is incorporated in vRNPs and exhibits the characteristic colocalization of NP and Rab11A in IAV-infected cell cytoplasm. Interestingly, colocalization of EGFP-NP and mCherry-Rab11A was withdrawn in the presence of G12 for 24 h (Fig. 6b (iv) and Supplementary Fig. 10 (iv), suggesting that G12 can disrupt the transport of vRNPs in the cytoplasm. mCherry-Rab11A localization in uninfected cells was not affected by G12 (Fig. 6b (iii) and Supplementary Fig. 10 (iii)). This result was supported by a time-lapse observation upon exposure to G12(S) for 85 min in the late replication stage (24 h) (Fig. 6c). We found that NP-Rab11A colocalization in the MTOC was strongly reduced by G12(S) treatment for 85 min (compare G12(S) exposure times of 15 and 85 min in Fig. 6c). Furthermore, G12(S) treatment for 85 min withdrew punctate mCherry-Rab11A with rapid movements found in the IAV-infected cells (see the upmost panels in Fig. 6c and Supplementary Movie 1) as well as in the infected cells with G12(S) treatment for 15 min (Fig. 6c, middle panels and Supplementary Movie 2), and reverted mCherry-Rab11A to the membranous reticulated structure throughout the cytoplasm (see Fig. 6c, lower panels and Supplementary Movie 3). These results indicate that G12(S) rapidly reduces the colocalization of Rab11A and NPs, reverting the punctate distribution, rapid migration, and structures of Rab11A to those of the uninfected cell state (Fig. 6d and Supplementary Movie 4).
a Construction of a dicistronic plasmid that expresses mEGFP and mCherry-Rab11A simultaneously. The gene was driven by a CMV promoter and an EMCV IRES was inserted before the second cistron in mCherry-Rab11A. At 1 h after IAV infection (MOI = 1), the plasmid was transfected to the A549 cells. The cells were further cultured for 23 h. The medium was changed to phenol red-free MEM containing glutamine and NucBlue (Hoechst 33342) for staining the nucleus. b Effect of 2 μM G12 on the colocalization of mEGFP-NP (green) and mCherry-Rab11A (red). Non-infected cells (i and iii), IAV-infected cells (ii and iv), and each mCherry, EGFP, and the superimposed images of all including NucBlue (blue) are shown. Multiple images taken by same condition are shown in Supplementary Fig. 10. c, d Effect of 85 min treatment of G12(S) on IAV-induced punctate appearance and movement of mCherry-Rab11A. Superimposed images of mEGFP-NP and mCherry-Rab11A are shown in the left panels. Images at 1-s intervals for mCherry–Rab11A and its movement are shown under the conditions of IAV infection (c) uninfected cells (d) after treatments of 2 μM G12(S) for 15 and 85 min (c) and 40 min (d). The parts of images of the mCherry-Rab11A are 200% magnified. Scale bars indicate 10 and 2.5 μm, respectively. Corresponding movie files (Supplementary Movies 1–4) are indicated in c and d.
A previous report43 has shown that the interaction of Rab11A with NP—but not with RNA polymerase complex—is sensitive to RNase A treatment, indicating that vRNA-NP interaction is required for Rab11A-dependent transport of intact vRNP transport. Therefore, G12 and G12(S) might inhibit the interaction between NP and vRNA followed by the dissociation of vRNP-Rab11A interaction during cytoplasmic transport. Inhibition of the vRNP transport step is consistent with our results that G12(S) simultaneously reduced vRNA levels (Fig. 5c) and the release of infectious virus into the medium (Fig. 5g).
G12(S) forms a left-handed G-quadruplex-like structure
Guanine-rich nucleotide sequences are known to form G-tetrads through Hoogsteen-type base pairing and a centrally coordinating alkali metal ion (Na+ or K+) (see Fig. 7a). Several G-tetrad planes consisting of single or two to four multiple G-rich oligonucleotides result in the formation of a stable G-quadruplex structure (for a review, see Zhang et al.44). We used circular dichroism (CD) spectroscopy, electrophoretic mobility shift, the fluorescence detection of G-tetrad-specific probe N-methyl mesoporphyrin IX (NMM)45 and G-tetrad-specific absorbance at 295 nm46 to demonstrate that G12(S) could form G-tetrads and G-quadruplexes in response to K+ and/or Na+. As shown in Fig. 7b, upon the addition of Na+, the CD spectra of G12(S) (ca. 264 nm max, 245 nm min) showed a G-quadruplex formed with a typical parallel topology (with four strands oriented in the same 5’ to 3’ direction; see Fig. 7a)47,48, whereas KCl induced a completely different spectrum (ca. 247 nm max, 276 nm min). Interestingly, the K+-induced CD spectrum dominated in culture medium (minimal essential medium, MEM) (Supplementary Fig. 11a) and a phosphate buffer (pH 6.8) containing 131.2 mM NaCl and 5.3 mM KCl (Na-K phosphate; the condition similar to MEM; Supplementary Fig. 11b). These K+-induced CD spectrum was emphasized for long-term incubation (24 h). Our results suggested that G12(S) forms the K+-induced type of G-quadruplex in culture medium and cells.
a The four guanine form a planar G-tetrad structure through Hoogsteen hydrogen bonds and are stabilized by alkali metal ions (e.g., K+ and Na+). Guanosine-rich RNA oligomers form several G-tetrads with a parallel orientation to form G-quadruplex structures. The structure of Block2Δ15 (GTGGTGGTGGTG) consists of two subunits of two-layered parallel-strand G-quadruplex structure with left-handed helicity. b CD spectra of G12(S) in 10 mM Tris/HCl (pH 7.0) (black curve), 30 mM NaCl (blue curve) or 30 mM KCl (yellow curve). G12(S) (4 μM) was incubated overnight in 10 mM Tris/HCl, and then further incubated overnight with 30 mM NaCl or 30 mM KCl. c Nucleotide sequences of G12(S) and BL2(S). The Tm of G12(S) in 30 mM KCl or 30 mM NaCl in 10 mM Tris/HCl (pH7.0) obtained for the first and second temperature increases are shown (see Supplementary Fig. 12). Helicity of G-quadruplexes determined by CD spectroscopy are shown as left-handed parallel (L) and right-handed parallel (R). d CD spectra of BL2(S). BL2(S) was incubated in H2O overnight (blue curve), and then further incubated in Na-K phosphate buffer (orange curve), 30 mM KCl-10 mM Tris/HCl (pH 7.0; yellow curve) or 30 mM NaCl-10 mM Tris/HCl (pH 7.0) (gray curve) for 1 h. e, f Mobility shift of G12(S) and BL2(S) in native PAGE. G12(S) and BL2(S) at 4 μM (20 pmols) pretreated with 0, 5, 10, 20, or 40 μM of NMM and then with 10 mM KCl-Tris/HCl for 20 min were separated on a 15% PAGE at a constant voltage of 120 mV for 100 min using Tris/borate containing 10 mM KCl (e) or 10 mM NaCl (f). The gels were stained with NMM. The dsRNA marker in the rightmost lane of each gel was pretreated with 200 pmol of NMM prior to PAGE, whereas the second lane from the left was not, which was detected by subsequent DAPI staining (see Supplementary Fig. 11c). g Effects of MNN on CD spectra of G12(S), BL2(S) and ADAM10(S). These PS-RNA were incubated overnight in H2O, and then incubated with 0, 10 and 20 μM NMM in 10 mM Tris/HCl (pH 7.0) for 30 min and further incubated with 30 mM KCl for 1 h. CD spectra of 0 μM NMM (blue curve), 10 μM NMM (orange curve) and 20 μM NMM (gray curve) are shown. h Possible structural changes induced by NMM and NaCl in the G-quadruplex dimer structure of BL2(S), which may be a structural model for G12(S). NMM and/or NaCl convert the left-handed parallel dimer of BL2(S) formed in the presence of K+ into a right-handed parallel monomer.
We determined the Tm of G12(S) by monitoring changes in G-tetrad-specific absorbance at 295 nm46 using a spectrometer. G12(S) in the K+ buffer indicated that the Tm of two denaturing processes were 90.5 and 96.5 °C, which were markedly higher than that in the Na+ condition (63.0 and 62.0 °C). The Tm of G12(S) in Na-K phosphate was approximately 10 °C lower than that in KCl (86.5 and 83.5 °C) (Fig. 7c and Supplementary Fig. 12).
The K+-induced CD spectrum was similar to that of a G-quadruplex structure with left-handed helicity such as Z-G4 (28mer)14 and it’s a minimal sequence Block2Δ (12mer)15, which exhibit atypical CD spectra with a positive peak at ~250 nm and a negative peak at ~275 nm. We compared G12(S) and a PS-modified RNA form of the minimal Block2Δ (BL2(S): GUGGUGGUGGUG). BL2(S) in KCl and Na-K-phosphate exhibited a CD spectrum (Fig. 7d) similar to that of Block2Δ DNA in KCl15. However, NaCl induced no specific CD spectra.
In the presence of KCl, Block2Δ DNA is known to stack at the 5’-5’ ends of two-plane G-quadruplexes15 (see Fig. 7a). To investigate the structural similarity between BL2(S) and G12(S), we used NMM, which can bind G-tetrad surfaces of the parallel conformation49. We speculated that NMM could inhibit such a noncovalent interface between the two subunits. As shown in Fig. 7e, pretreatment of increasing the amount of NMM shifted the mobility of G12(S) and BL2(S) by approximately half the size (from 60b to 30b of ssRNA marker). These mobility shifts were enhanced in NaCl-containing PAGE (Fig. 7f). These data indicated that NMM contact and/or the substitution of K+ with Na+ during PAGE reduced the stability of the G-quadruplex dimeric structure of G12(S) and BL2(S). The structural changes of G12(S) and BL2(S) were also demonstrated by CD spectra. As shown in Fig. 7g, NMM inhibited formation of the left-handed G-quadruplex structure of BL2(S). ADAM10(S) (see Fig. 8 for sequence) was used as a control for a monomer G-quadruplex50, and as expected, both its mobility in native PAGE (Supplementary Fig. 11d) and CD spectrum (Fig. 7g) were unchanged by NMM, demonstrating the validity of the experimental conditions. The CD spectra of G12(S) suggested that NMM enhanced right-hand spectra (Fig. 7g). Therefore, although G12(S) does not exhibit a homogeneous structure, our results suggested that G12(S) in KCl exhibited a BL2(S)-like dimer structure. Furthermore, the disruption of G12(S) dimer induced by NaCl and NMM generated a faster-migrating structure of the right-handed helix, however the precise molecular structure was not revealed in this assay.
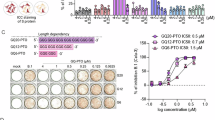
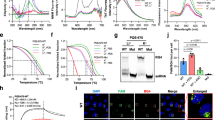
a Nucleotide sequences of left-handed parallel DNA Block2, Block2Δ and corresponding BL2-RNA and PS-modified BL2 (BL2(S)). b Anti-IAV activity was examined by the Block2 and Block2Δ DNAs (2 μM). c Nucleotide sequences of PS-RNA version of known sequences (12mer to 18mer). Gq(S) is a 12-nt sequence consisting of UUU flanked by two GGGs, with U and UU at the 5’ and 3’ ends, respectively. MTA2 and ADAM10 are found in the 5’-untranslated region of human cellular mRNAs that can form G-quadruplex structures50. 93del is derived from an aptamer DNA isolated as HIV-1 integrase inhibitor and inhibits HIV-1 replication16. 93del forms a parallel-fold G-quadruplex structure51. G15 oligo DNA has been found to be the shortest poly-G sequence that form a strict G-quadruplex structure52. All PS-RNA showed right-hand parallel CD spectra (R) (see Supplementary Fig. 13b). d Anti-IAV activities of PS-oligonucleotide (2 μM) examined. e, f A549 cells were treated with 2 μM PS-RNAs (indicated in the figures) and stained with QUMA-1. We measured G-quadruplex levels in cells (7-μm thick) by overlaying eight slices acquired with a super-resolution confocal laser scanning microscope (LSM900 with Airyscan 2) using the G-quadruplex-specific fluorescent dye QUMA-1. Shown are separate images (e) overlay images (f) of eight planes (z-direction) of QUMA-1 (yellow) and DAPI staining (blue). AF610 (Excitation = 612 nm, Emission = 630 nm, Detection = 590–700 nm) used for QUMA-1 observation. Black and white of AF610 set at approximately 4000 and 12,000 (f). g PS-RNA-dependent QUMA-1 fluorescence by adjusting the brightness threshold to 4000 to exclude nucleus autofluorescence and fluorescence of cells untreated with PS-RNA (see Supplementary Fig. 13d, original data were available at https://doi.org/10.5061/dryad.8931zcs1s). The total QUMA-1 brightness value for each overlay image was divided by the number of nuclei to obtain the QUMA-1 brightness value per cell. To average out variability due to cell size, the QUMA-1 brightness values were divided by the average size of nuclei in each overlay image. Independent data sets (2~6 overlay images containing 36–99 cells) are shown by box-and-whisker plots including all data plots. Statistical analyses of the G12(S) effect by One-Way Repeated measures ANOVA (b F(2, 6) = 2.263, P-value = 0.1852; d F(7, 19) = 551.1, P < 0.0001; g F(7, 20) = 6.133, P = 0.0006), and Tukey’s multiple comparison test was used. “ns” indicates P-value > 0.5 (b). P-values of d and g are indicated in Supplementary Figs. 13c and 13e, respectively. The letters shown in d and g are classifications created using Compact Letter Display (CLD) in GraphPad Prism 10, which is another way of grouping based on significant differences, e.g., between A and B, the P-value is less than 0.05, but between A and AB or AB and B, the P-value is greater than 0.05. h Correlation between IAV antiviral effect and cytoplasmic G-quadruplex level was examined using Pearson’s correlation coefficient. The average IAV reduction value and QUMA-1 fluorescence intensity are plotted, and the color of each point indicates the PS-RNA treatment corresponding to the color used in graphs d and g.
Anti-IAV activity of G-quadruplex PS-RNAs
We found BL2-RNA and its phosphorothioate form BL2(S) had anti IAV activity (Supplementary Fig. 13a), but Block2 (DNA) and Block2Δ (DNA corresponding BL2-RNA, Fig. 8a) did not (Fig. 8b). Thus, RNA structure, not DNA, is important for anti-IAV activity. We next examined the anti-IAV viral activity of various PS-modified G-rich PS-RNAs (Fig. 8c). When these five sequences were modified into PS-RNA, they showed CD spectra characteristic of right-handed parallel folding in response to KCl (Supplementary Fig. 13b), which is consistent with the results of previously reported DNA oligomers50,51,52.
All PS-RNAs except Gq(S) showed anti-IAV activity. In the case of MTA2(S), ADAM10(S), G15(S), and 93del(S), the activity was approximately 1/10 that of G12(S) (Fig. 8d). However, Gq(S) has no anti-IAV activity. BL2(S) exhibited anti-IAV activity equivalent to G12(S) (Fig. 8d). Next, we attempted to quantify intracellular G-quadruplex levels. Cells treated overnight with these PS-RNAs at a concentration of 2 μM were fixed and stained with the RNA G-quadruplexes-specific fluorescent dye QUMA-1 ((E)-2-(2-(7-(diethylamino)-2-oxo-2H-chromen-3-yl)vinyl)-6-fluoro-1-methyl-7-(4-methyl piperazin-1-yl)quinolin-1-ium iodide)53. The detection of QUMA-1 was set up to exclude cellular autofluorescence in the nucleus (Fig. 8e and Supplementary Fig. 13d), and PS-RNA-dependent punctate fluorescence was observed in the cytoplasm but not in the nucleus (Fig. 8f). All PS-RNA except Gq(S) showed high levels of QUMA-1 fluorescence (Fig. 8f, g). As shown in Fig. 8h, we found a correlation between the levels of cytoplasmic QUMA-1 fluorescence (Fig. 8g) and the anti-IAV activity (Fig. 8d). The cytoplasmic QUMA-1 fluorescence of G15(S) was similar to that of BL2(S) (Fig. 8g), but the anti-IAV activity by G15(S) was lower than that of BL2(S) (Fig. 8d). G15 (DNA) forms three-plane G-tetrad with three guanines (G6, G10 and G14) pointing outward like a propeller52. Among the PS-RNAs examined, G12(S) and BL2(S) had higher cytoplasmic QUMA-1 fluorescence and anti-IAV activity. Thus, it is possible that the structures of G12(S) and BL2(S) are suitable for NP-RNA inhibition.
Requirement of cytoplasmic G-quadruplex formation in anti-IAV PS-RNA
To investigate whether G-quadruplex formation in PS-RNA is required for anti-IAV activity, we examined the modification of BL2(S) (Fig. 9a). Two G-U substitutions introduced into BL2-14(S) to generate BL2-UU25(S) disrupted the left-handed parallel CD spectrum of BL2-14(S) (Fig. 9b) and reduced the anti-IAV activity (Fig. 9c) and intracellular G-quadruplex (Fig. 9d, e). On the other hand, although the KCl-induced CD profile of 5U-BL2(S) was similar to that of BL2-14(S) (Fig. 9b right), 5U-BL2 failed to induce intracellular G-quadruplex (Fig. 9d, e) and anti-IAV activity. These results suggest that efficient anti-IAV activity requires the formation and stability of G-quadruplex structures in the cytoplasm.
a Nucleotide sequences of PS-RNAs examined. BL2-14(S) and 5U-BL2-14(S) correspond to Block 2 and 5’T-Block 2 DNA oligomer, respectively15. BL2-UU25(S) is the 5’-teminal half of 2xBlock 2 T2,5,16,19TT DNA15. b CD spectra of BL2-14(S), BL2-UU25(S) and 5U-BL2(S) in H2O or 10 mM Tris/HCl (pH 7.0) containing 30 mM KCl. c Anti-IAV activity was examined by the PS-RNAs (2 μM). d, e QUMA-1-stained cells were observed as described legend to Fig. 8. Independent data sets (3 overlay images containing 30–65 cells) were shown by box-and-whisker plots including all data plots with the P-values. Statistical analyses of the G12(S) effect by One-Way Repeated measures ANOVA (c F(3, 26) = 63.85, P < 0.0001; e F(3, 13) = 14.38, P = 0.0002) and Tukey’s multiple comparison test was used. P-values of c and e are indicated in Supplementary Figs. 14a and b, respectively. The letters shown in (c and e) are classifications created using CLD (see legend to Fig. 8).
Docking simulation of SARS-CoV-2 NP and BL2 RNA
In order to gain insight into the binding mode of G-quadruplex RNA to SARS-CoV-2 NP, 3D models of each component were constructed using AlphaFold3 server54. Since crystal structure of Block2Δ has been determined15, the corresponding RNA, BL2-RNA, was used for docking. We found that BL2(S) had also anti-coronavirus (OC-43) activity (Fig. 10a), and that the G-quadruplex structure of BL2(S) was only weakly affected in the presence of equimolar SARS-COV-2 NP (Supplementary Fig. 15). The NP formed a dimer, utilizing the C-terminal dimerization domains as observed in the crystal structure (PDB ID: 6WJI)55. The N-terminal domains are facing toward a same direction. Two BL2-RNAs were constructed to include three K+ ions, and were modeled to form a G-quadruplex. Docking of the BL2-RNA to N protein was performed using HADDOCK 2.4 server56, and the active site was set to the N-terminal domain of N protein (aa48-aa160), which has been identified as the RNA binding region57. The putative BL2-RNA-binding site of the NP located outside the basic finger, a positively charged region composed of lysine and arginine residues. Since vRNA-binding site is predicted to be located in a positively charged canyon between the basic finger and the palm of the N-terminal domain of SARS-CoV-2 NP (PDB ID: 7ACS and 7ACT)57, BL2-RNA binding may affect the interaction of vRNA with the basic fingers. Further experimental evidence is needed to validate the current complex model and to specify in detail the intermolecular interactions.
a Anti-coronavirus (OC43) activity of BL2(S). MRC-5 cells were infected with OC43 (MOI = 0.01) and the culture medium was collected at 4 pdi. Pfu was determined by plaque assay using MRC-5 cells. b Docking models of the SARS-CoV-2 N protein dimer in complex with BL2-RNA molecules. Docking was performed using HADDOCK server and the three docking poses were superimposed. The proteins are depicted in gray with ribbon model, and the unstructured regions were omitted for clarity. The bound BL2-RNAs are shown in orange. c The replication and transcription of CoV vRNAs occur in DMVs derived from the ER membrane. Viral vRNA are transported to the cytoplasm, where they perform NP binding and form RNP complexes. This step may be inhibited by G12(S) and the resulting putative immature RNPs may be then transported to the ER-Golgi intermediate compartment (ERGIC). Virus particles are formed within the ERGIC and secreted as noninfectious coronaviruses (RNAs) outside the cell. d Replication cycle for IAV. Upon the entry and shedding of IAVs, eight vRNPs are transported into the nucleus, where mRNA and complementary RNA (cRNA) are synthesized. The newly synthesized NPs and IAV RNA polymerase subunits (PA, PB1, and PB2) are imported into the nucleus for vRNA replication and vRNP formation. The assembled vRNPs are exported to the cytoplasm and attached to the recycling endosome-specific G-protein Rab11A (red circle). The Rab11A-vRNPs are transported to the plasma membrane, where the viral is packaged and secreted outside the cells. G12(S) may inhibit vRNA-NP interaction, which is required for Rab11-dependent transport.
In this study, we demonstrated that G12(S) forms G-quadruplex structure in the cytoplasm inhibit cytoplasmic viral replication event (Fig. 10c, d). This finding may be interest to other viruses with cytoplasmic vRNPs in their replication process. Future studies will reveal that other viruses with similar vRNPs in the cytoplasm may be affected by G12(S). Recent findings have shown that a portion of tRNA-derived intracellular G-quadruplex (tiRNAs) bind to eIF4G and suppress translational repression, leading to the formation of stress granules58. G12(S) is unlikely to have such activity because it cannot repress protein synthesis (Fig. 1). The binding specificity of viral G-quadruplex RNAs to cellular proteins appears to be used to evade the cellular antiviral stress response. An intriguing finding is that the yellow fever virus vRNA G-quadruplex structure hijacks hnRNPH1 to prevent tiRNA-induced translation inhibition59. Our data indicated that high concentration (>5 uM) of G12(S) were not toxic but tended to increase the numbers of MDCK cells specifically (Supplementary Figs. 1a and 9b). To understand the effect of G12(S) on cell proliferation, it is necessary to examine not only the biochemical specific binding ability of G12(S) but also its interaction and localization in the cells under physiological conditions60. Identification of G12(S)-specific binding cytoplasmic proteins may provide insight into not only NP-vRNA interaction but also their alternative antiviral effects.
G12(S) has promising results against IAV, demonstrating better anti-IAV activity than two approved antivirals, and is expected to progress as a potential antiviral drugs. In animal studies, multiple dosing improved anti-OC43 toxicity, thus improving stability of G12(S) or modifying the PS-RNA sequence may be beneficial for antiviral drugs. Some modifications to the RNA backbone or covalent attachment of carrier molecules may be beneficial for intracellular delivery and stabilization61. Improved drug delivery methods and formulations, as well as further studies using animal models of infection, are needed to achieve this goal.
Methods
Ethics statement
We have complied with all relevant ethical regulations for animal use. The mouse experiments were conducted with their content approved by the President of Tohoku Medical and Pharmaceutical University (TMPU) after examination by the Animal Experimentation Committee of TMPU. The experiment complied with the “Standards for the Care and Management of Laboratory Animals and Alleviation of Pain” under the “Act on Welfare and Management of Animals.” Additionally, this research was conducted in accordance with the “Guidelines for Proper Conduct of Animal Experiments” formulated by the Science Council of Japan based on the “Basic Guidelines for Conducting Animal Experiments in Research Institutions” formulated by the Ministry of Education, Culture, Sports, Science and Technology. The mouse breeding and infection experiments were performed in specific pathogen-free conditions in the Center for Laboratory Animal Science.
Reagents
Guanine RNA (12mer), PS-modified guanine RNA (12mer) and other RNA oligomers were purchased from Fasmac (Kanagawa, Japan) and Aji Bio-Pharma (Osaka, Japan). Guanine RNA (12mer) and PS-modified guanine RNA (12mer) labeled with the fluorescent dye TAMRA (5-Carboxytetramethylrhodamine) at their 5’-end (TAMRA-G12(S)) were purchased from Fasmac (Kanagawa, Japan). These RNAs were dissolved in H2O (1 mM and 0.1 mM concentration) and stored at −80 °C. MEM and Roswell Park Memorial Institute (RPMI)-1640, Dulbecco’s phosphate-buffered serine (PBS; 143.5 mM Na+ and 1.47 mM K+), trypsin-ethylenediaminetetraacetic acid (EDTA) (PS, 2.5 g/L trypsin-1 mM EDTA solution), 100x penicillin-streptomycin solution, acetate, and ethanol were purchased from Nacarai Tesque (Kyoto, Japan). Fetal carf serum (FCS; Nichirei Biosciences, Tokyo, Japan and (Biosera, Nuaillé, France), MEM (05900, Shimazu, Tokyo, Japan) for plaque assay, phenol red-free MEM (Gibco 51200-038, ThermoFisher, MA, USA), 1 mol/L 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) buffer (Dojindo, Kumamoto, Japan), Cellstain DAPI (4’,6-Diamidino-2-phenylindole, dihydrochloride) solution (Dojindo, diethylaminoethyl cellulose (DEAE)-dextran (Cytiva, Tokyo, Japan), Bacto agar (Difco, BD Life Sciences, New Jersey, USA), trypsin (T1426, Merk, Darmastadt, Germany), NucBlue (Hoechst 33342, Invitrogen, ThermoFisher), ER-Tracker Red (BODIPY TR Glibenclamide, ThermoFisher), N-methyl mesoporphyrin IX (Cayman, Ann Arbor, MI, USA), QUMA-1 (Merck Millipore, New Jersey, USA) and SYBRTMGold (S11494, ThermoFisher) were used.
Antibodies and their concentrations
Primary antibodies:
The following antibodies were purchased from Sino Biological (Beijing, China), GeneTex (CA, USA), Roche Diagnostics (Basel, Switzerland), and Santa Cruze Biotechnology (Texas, USA):
HCoV-OC43 nucleocapsid antibody, rabbit polyclonal, Sino Biological, 40643-T62, 2000x (western), 500x (immunofluorescence, IF)
HCoV-229E nucleocapsid antibody, rabbit polyclonal, Sino Biological, 40640-T62, 5000x (western)
IAV nucleoprotein antibody, rabbit polyclonal, GeneTex, GTX125989, 10,000x (western)
IAV HA antibody, rat monoclonal (clone 3F10), Roche, 100 ng/mL (western)
KDEL antibody, mouse monoclonal, MBL, M181-3, 400x (IF), 10,000x (western)
ERGIC-53 antibody, mouse monoclonal (C-6), Santa Cruz, sc-365158, 400x (IF), 1000x (western)
Actin antibody, goat polyclonal, Santa Cruz, 2000x (western)
Puromycin antibody, mouse monoclonal, Millipore MABE343, 1000x (western)
Secondary antibodies:
Polyclonal goat anti-mouse IgG-horse radish peroxidase (HRP) (Dako, Agilent, CA, USA), 5000x (western)
Polyclonal rabbit anti-goat IgG-HRP (Dako, Agilent), 5000x (western)
Polyclonal goat anti-rabbit IgG-HRP (Dako, Agilent), 5000x (western)
Goat anti-rat IgG mouse antibody HRP (SouthernBiotech, AL, USA), 5000x (western)
Donkey anti-mouse IgG H&L (Alexa Fluor 488) (abcom, Cambridge, UK), 1000x (IF)
Donkey anti-rabbit IgG H&L (Alexa Fluor 594) (abcom), 1000x (IF)
Cell culture
A human lung cancer cell line A549 (RCB3677) and a human embryonic lung fibroblast cell line MRC-5 (RCB0218) were provided by the RIKEN bioresource center (BRC) through the National BioResource Project of the Ministry of Education (MEXT), Japan). A549 and MRC-5 cells were cultured in MEM containing 10% FCS. Human adenocarcinoma cell line (HCT-8; American Type Culture Collection, ATCC, CCL-244) cells were cultured in RPMI 1640 containing 10% horse serum (HS, GIBCO, ThermoFisher). Monkey kidney cell line LLC-MK2 cells62 were cultured in 5% FCS containing MEM. Epitherial-like cell line established from normal kidney of dog (MDCK)62 were culture in 10% FCS containing MEM. All culture mediums used 1x penicillin-streptomycin. Cell viability was determined with CellTiter-Blue (Promega, Wisconsin, USA). VeroE6/TMPRSS2 cells63 provided by the JCRB (Japanese Collection of Research) cell bank were cultured in Dulbecco’s MEM containing 5% FCS.
Virus infection and detection
HCoV-OC43
HCT-8 cells were infected with HCoV-OC43 virus (ATCC, VR-1558) at 33 °C. To determine HCoV-OC43 infectivity, MRC-5 cells cultured in 24-well plates were infected with serial dilution, and plaques were observed using the overlay of MEM containing 2% FBS and 1.5% carboxymethyl cellulose or 1% methyl cellulose at 4 dpi27.
HCoV-229E
LLC-MK2 cells were infected with HCoV-229E virus (2004-1948 strain isolated in Sendai64) at a MOI of 0.01–0.1 in MEM containing 1–1.5 μg/ml trypsin at 33 °C. To determine the virus titer, LLC-MK2 cells (96-well plates) were infected with a serial dilution of the virus, the appearance of syncytium was observed for 3–5 days, and the TCID50 was calculated.
SARS-CoV-2
HCoV-19/USA/WI-UW-5250/2021 (delta variant) was infected to VeroE6/TMPRSS2 at MOI of 0.001. Virus titers were determined by a standard plaque assay on VeroE6/TMPRSS2 cells.
IAV
The following IAV strains were used: A/Aichi/2/68 (H3N2), mouse-adapted A/California/04/09 (H1N1pdm09), and mouse-adapted A/Aichi/2/68 (H3N2). After washing once with PBS and then with MEM, MDCK cells (confluent) were infected with IAV (H3/Aichi) at an MOI of 0.01 in MEM containing 2.5 µg/ml trypsin at 37 °C. To determine the virus titer (pfu/mL), confluent MDCK cells (6-well plates) were infected with a 10-fold serial dilution of the virus. The appearance of virus plaques was then observed under an agar overlay (0.8% bacto agar, 5 µg/ml trypsin, 0.2% BSA, 10 mM HEPES, and 0.01% DEAE-dextran in MEM). The cells were fixed with a layer of 50% acetic acid-50% ethanol and then stained with 2% crystal violet to count the number of plaques.
Virus RNA produced in the medium was prepared using the QIAamp Viral RNA Mini kit (Qiagen, Venlo, Netherlands). The total cellular RNA from infected cells was prepared using TRI (Trizol) Reagent (Molecular Research Center, Ohio, USA). The real-time quantitative PCR (qRT-PCR) of viral RNAs was performed using the Thunderbird Probe One-step qRT-PCR kit (Toyobo, Osaka, Japan) with each viral primer set. RT-PCR was conducted using LightCycler (Roche, Basel, Switzerland).
HCoV-OC43 NP primer set:
OC43 left primer (GGAAGGTCTGCTCCTAATTCCA)
OC43 right primer (TGATCAGCCATGTCAGGTGTTA)
OC43 detection probe (5’-/56-FAM/ACTTCGCGC/ZEN/ACATCCAGCAGAGCCT/3IABkFQ/-3’
HCoV-229E NP primer set:
229E left primer (GCCACGGTGGAAAAGACAGC)
229E right primer (CCATTGGCCACAACACCTGC)
229E detection probe (5’-/56-FAM/TGGCCCCAG /ZEN/AGACCTTGACCACAACT/3IABkFQ/-3’
IAV H3/Aichi NP primer set:
NP left primer (TGGCTACGACTTCGAAAAAGA)
NP right primer (ACCAGCTGACTCTTGTGTGCT)
NP detection probe (5’-/56-FAM/TATACAGCC/ZEN/TAATCAGACCGAACGAG/3IABkFQ/-3’
IAV H3/Aichi HA primer set:
HA left primer (CCAGTGCTGAACGTGACTATG)
HA right primer (ACTCTCCCTGATGCTTGAACA)
HA detection probe (5’-/56-FAM/AGCACGAAC/ZEN/CAAGAACAAACCAG/3IABkFQ/-3’
These probe sequences were selected using the Primer3 website. The unlabeled primer oligonucleotides were purchased from Fasmac. The detection probes were PrimeTime Probes purchased from Integrated DNA Technologies (Iowa, USA). The genomic RNA samples were also isolated from the stock viruses and used as the quantitative controls. The viral RNA numbers were calculated by correlating the stock virus RNA levels to their infectious titers.
SARS-CoV-2 RNA produced in the medium was prepared using the QIAamp Viral RNA Mini kit (Qiagen). The real-time quantitative PCR (qRT-PCR) of SARS-CoV-2 RNAs was performed using the SARS-CoV-2 direct detection RT-qPCR kit (RC300A; Takara Bio, Shiga, Japan) and LightCycler.
For western blotting, infected cells were lysed with sodium dodecyl sulfate (SDS) sample buffer (50 mM Tris/HCl pH 6.8, 2% SDS, 10% glycerol, and 0.01% bromophenol blue) containing a Complete Mini protease inhibitor cocktail (Roche) and boiled at 96 °C for 5 min. The protein concentration was estimated using a DC protein assay kit (BioRad, California, USA). After electrophoresis in 10 or 12% SDS-PAGE, proteins were electro-blotted to an Immobilon-P PVDF membrane (Merck Millipore) using a Criterion cell (BioRad). Blocking was performed in 5% skimmed milk in TBST (10 mM Tris/HCl, pH 7.5, 0.15 M NaCl, and 0.1% Tween20). The primary antibodies were prepared in BockingOne (Nacalai). TBST containing 0.5% skimmed milk was used for the washing buffer and the secondary antibody solution. Chemiluminescence was detected with ChemDoc (BioRad) using the Chemi-Lumi One L (Nacalai) or Immobilon HRP substrate (Merck Millipore) as the substrate for HRP. Image Lab (BioRad) was used for image quantification.
Treatment of virus infected cells with PS-RNA
Cells were infected with coronavirus or IAV for 1 or 1.5 h in the serum-free medium used to grow the cells. After washing the cells, PS-RNA was added directly to the same serum-free medium and incubated for the indicated times.
Detection and analysis of fluorescently-labeled G12 or G12(S) by microscopy
A549 cells were cultured in a 96-well plate (ViewPlate black, PerkinElmer) with phenol red-free MEM supplemented with 0.03% L-glutamine. The cells were treated with 2 μM of TAMRA-G12 or TAMRA-G12(S). Fluorescent images were observed using LSM900 with Airyscan 2 equipped with Axio Observer Z1/7 using Plan-Apochromat 63x/1.4 Oil DIC M27 (Zeiss, Oberkochen, Germany).
FRAP analysis of TAMRA-G12(S) was performed as follows: After A549 cells were incubated with 2 μM TAMRA-G12(S) for 1 h, photobleaching was performed by maximum laser power for 5 min. Time-dependent recovery for TAMRA-G12(S) fluorescence was chased for 3.75 min.
Distribution of TAMRA-G12(S): A549 cells were treated with 2 μM TAMRA-G12(S) for 1 h, and with 1 μM ER-Tracker Red (1/1000) and Hoechst 33342 for the last 30 min.
Immunofluorescence microscopy
A549 cells were cultured in MEM containing 10% FCS and PS on glass cover slips (15 mm, 0.22 mm, Matsunami, Osaka, Japan). The cells were infected with OC43 (MOI = 1) and cultured for 48 h in MEM without FCS at 33 °C, as described above. Cells were fixed on the glass cover slips with 4% paraformaldehyde/PBS at 4 °C overnight. Cells were washed with PBS three times before following each step. To enhance permeability, cells were treated with 0.5% Triton-X100/PBS at room temperature for 10 min. Additionally, 1% HS/PBS was used to reduce nonspecific antibody binding. Cells were treated with anti-HCoV-OC43 NP rabbit antibody and anti-KDEL monoclonal mouse antibody or anti-EREGIC-53 in 2% HS/PBS at 4 °C overnight. For the secondary antibody, donkey anti-rabbit IgG-Alexa Fluor 594 and donkey anti-mouse IgG-Alexa Fluor 488 in 2% HS/PBS were used at room temperature for 1 h. DNA was stained with Cellstain DAPI solution (1/1000, Dojindo, Kumamoto, Japan) in PBS. Finally, cells were embedded in a fluorescence mounting medium (Dako). Fluorescence images were observed under confocal microscopy (LSM 900 with Airyscan 2, Zeiss, Oberkochen, Germany).
HCoV-OC43 infection and detection in mice
The infection of mice with HCoV-OC43 was performed as described by Niu et al.29. The HCoV-OC43 (ATCC, VR-1558) was amplified by one passage in HCT-8 cells to produce the viral stock. C57BL/6JJcl mice were purchased from CLEA Japan (Tokyo, Japan). The purchased mice were then bred. PBS (5 uL) containing HCoV-OC43 (2.2 × 104 TCID50) was intranasally inoculated 7 days after birth, raised together with mother mice, and observed until the end point of a 20% reduction in weight. G12(S) (8.16 µg) in PBS was intranasally administered once before HCoV-OC43 inoculation or repeatedly three times. Mice at the endpoint were anesthetized by sevoflurane (Maruishi Pharmaceutical, Osaka, Japan) through the airway. For the detection of HCoV-OC43 RNA by RT-PCR, total RNA from olfactory bulbs and the whole brain was isolated by TRI Reagent as described above.
Immunofluorescence for HCoV-OC43 NP detection was performed as described by Dubé et al.28. Five days after infection by OC43, mice were perfused with PBS and perfusion-fixed with 250 mL of 4% paraformaldehyde (Nacarai Tesque)-PBS for 4 consecutive days. The whole head region was isolated. After decalcification in 6% EDTA (4 days at 4 °C) and dehydration in 30% sucrose (3 days at 4 °C), the whole-head sections (100 µm) were prepared in PBS using a vibratome (VT1200S, Leica Biosystems, Wetzlar, Germany). The sections were treated with 3% H2O2 for 10 min to disrupt the erythrocyte. For permeabilization, the sections were treated with 0.1% TritonX100-PBS for 2 h at room temperature. The sections were then treated with 0.05% TritonX100-PBS containing 1/400 of anti-HCoV-OC43 NP antibody overnight at 4 °C. After washing with 0.05% TritonX100-PBS, the sections were incubated with 0.05% TritonX100-PBS containing 1/500 of the secondary antibody. Finally, the cell nucleus in the section was stained with 1 μg/ml DAPI for 30 min. The sections were then mounted in mounting media (Dako) on glass slides. Fluorescent images were taken with an LSM900 (Carl Zeiss) equipped with Axio Observer Z1/7 and Plan-Apochromat 10X/0.45 M27 as an objective using the following conditions:
DAPI: 353 nm laser; 410–546 nm for wavelength detection.
Alexia Fluore594 (HCoV-OC43 NP): 561 nm laser; 590–700 nm for wavelength detection.
Construction and transfection of the mCherry-Rab11A-IRES-mEGFP-IAV_NP plasmid and observation of IAV transport by confocal microscopy
The construction of mEGFP-IAV(H3N2)NP-IRES-mCherry-Rab11A was achieved by PCR and the in vitro recombination system InFusion (Takara Bio). PrimeScript II and PrimSTAR GXL (Takara Bio) were used for cDNA synthesis and PCR, respectively. Rab11A and IAV (H3N2) Aichi cDNA were amplified cDNAs from Huh-7 total RNA and the virus stock, respectively. mEGFP (pEGFP, Takara Bio), mCherry (pmCherry, Takara Bio), and IRES (pIRES, Takara Bio) were PCR amplified. To fuse these fragments, PCR reactions were conducted to join IRES, mCherry, and Rab11A. The mEGFP was cloned into the BamHI-XhoI sites of pcDNA5-FRT, and the IAV(H3N2)NP was fused to the mEGFP by InFusion. The resulting pcDNA5-mEGFP-IAV(H3N2)NP was digested with ApaI, and the IRES-mCherry-Rab11A was inserted by InFusion.
Primers used for the construction:
GFP-NP-InFusion GCTGTACAAGACCGGCATGGCGTCCCAAGGCACC
NP-XhoI InFusion GCCCTCTAGACTCGATTAATTGTCGTACTCCTCTGCATTG
Rab11A-F-mCherry GGCATGGACGAGCTGTACAAGATGGGCACCCGCGACGA
Rab11A-R TTAGATGTTCTGACAGCACTGCA
Inf-ApaI IRES TAATCGAGTCTAGAGGGATCCGCCCCTCTCCCT
mCherry-R-Rab11A TCGTCGCGGGTGCCCATCTTGTACAGCTCGTCCATGCC
Rab11A-InfApaI CAGCGGGTTTAAACGTTAGATGTTCTGACAGCACTGCA
The mEGFP-IAV(H3N2)NP-IRES-mCherry-Rab11A plasmid was transfected into A549 cells by Lipofectamin 3000, and then IAV(H3N2) was infected (MOI = 1).
Construction of IFNβ and NFκB-luciferase reporter genes and the assay
To construct pGL4.24-NFκB, the following oligonucleotides were annealed and inserted into NheI-BglII sites of pGL4.24[luc2P minP].
(3’-5’) CTAGCGGGAATTTCCGGGGACTTTCCGGGAATTTCCGGGGACTTTCCGGGAATTTCCA
GATCTGGAAATTCCCGGAAAGTCCCCGGAAATTCCCGGAAAGTCCCCGGAAATTCCCG
To construct pGL4.1-IFNβ-x2, two IFNβ promoter regions were amplified by PCR using Primer 1 and Primer 2, and Primer 3 and Primer 4, respectively, with 293 T DNA as a template. Each was digested with NheI-EcoRI or EcoRI-HindIII, and cloned into NheI-HindIII-digested pGL. IFNbeta promoter linked to two tandem repeats were PCR amplified using the following oligos from 293 T DNA and coned into NheI-HindIII of pGL4.1.
Primer1: gtca GCTAGC TCAGGTCGTTTGCTTTCCTT
Primer2: gtca GAATTC GAAAGGTTGCAGTTAGAATGTCC
Primer3: gtca GAATTC TCAGGTCGTTTGCT
Primer4: gtca AAGCTT GAAAGGTTGCAGTTAGAATGTCC
HEK293 cells were transfected with the reporter plasmid using polyethylenimine MAX (Polysciences, PA, U.S.A.), and cultured for 24 h. pGL4.1-IFNβ-x2-transfected cells were cultured with poly I:C (40 μg/mL) and G12(S) (2 μM) for 24 h. pGL4.24-NFκB-transfected cells were cultured with G12(S) (2 μM) for 24 h, and treated with 40 μg/ml poly I:C (+Lipofectamin 3000) in the last 5 h. Cell viability were determined by CellTiter-Blue. Luciferase activity was determined by Steady-Glo (Promega, WI, U.S.A.) using Varioskan (ThermoFisher).
CoV genome analysis
The homology of genome sequences between SARS-CoV-2 and other CoV strains and variants was determined by CLUSTALW from Kyoto University’s Bioinformatics Center (https://www.genome.jp/tools-bin/clustalw). The selected variants of SARS-CoV-2 correspond to variants of concern indicated by the World Health Organization.
LLPS assay
Droplets of SARS-CoV-2 NP were formed by LLPS in the presence of1µM FITC-leveled 32 base RNA from the 3’-untranslated region of SARS-CoV-2 RNA (CGAG GCCACGCGGAGUACGAUCGAGUGUACAG) and 40 µM SARS-CoV-2 NP overnight13. E. coli BL21 CodonPlus (DE3)-RIL (Agilent) harboring pET-15-SARS-CoV-2 was grown and expression was induced in the presence of 1 mM isopropyl β-D-1-thiogalactopyranoside in 400 ml LB (OD600 = 0.4) for 6 h at 25 °C. NP was purified by Ni-NTA agarose (Qiagen) from E. coli lysate prepared in 50 mM HEPES pH 7.5, 500 mM NaCl, 10% glycerol, 20 mM imidazole, and 6 M urea. NP was dialyzed against 50 mM HEPES pH 7.5, 50 mM NaCl, and 10% glycerol. The reaction buffer included 50 mM NaCl, 20 mM HEPES-Na (pH 7.5), 10% glycerol, 4.6 U/µL RNase inhibitor, and 2.5 mM KCl (LLPS buffer). Then, TAMRA-G12(S) (final 1, 3, and 6 µM) were added and observed at 25 °C using fluorescence and differential interference contrast (DIC) microscopy at the indicated time. Images were taken using an RFP filter for the detection of TAMRA-G12(S) (1 msec) and a GFP filter for FITC (100 msec) using a Leica DMR equipped with a Zyla 4.2 P sCMOS (Andor Technology, Belfast, UK) using µManager.
Comparison of anti-IAV activity ofG12(S) with that of Tamiful and Inavir
IAV (A/Aichi/2/68 (H3N2)) was infected with MDCK cells (MOI = 0.01, n = 4) cultured in 96 well culture plates. One hour after infection oseltamivir phosphate (Tamiflu), laninamivir octanoate hydrate (Inavir), and G12(S) were added at 20 uM and six 4-fold dilutions for 27 h. Excipient amounts were corrected and dissolved in water (Tamiful) or DMSO (Inavir). Because IAV infection caused cells to detach from the plate surface, cells remaining were fixed with 50% acetate in ethanol and stained with 0.2% Cristal Violet. After washing and drying the plates, crystal violet was eluted with 70% ethanol and the absorbance at OD590 was measured. Values were calculated by dividing by the maximum value for each concentration series. Dose-response curves were fitted by nonlinear regression (curve fit) using log (drugs) vs. response variable slope (four parameters). EC50 was calculated from the best fit curve for the slope by Prism. Results were obtained by three independent experiments (See Supplementary Fig. 9).
CD analysis
A CD spectrometer J-720 (JASCO) and a quartz optical cuvette with an optical path length of 0.5 cm were used to analyze CD spectra. Samples for CD spectroscopy were prepared in the indicated buffers with a 4 μM concentration. CD spectra were monitored as an averages of 20 scans at room temperature (25 °C) using the following conditions: scan speed 50 nm/min, response time 2.0 s, band width 1.0, band width 1.0 nm, resolution 0.2 nm, sensitivity 100 mdeg. Sample RNAs were incubated overnight at room temperature in 10 mM Tris/HCl (pH 7.0) or H2O and further incubated with 30 mM NaCl or 30 mM KCl or Na-K phosphate for 1 h or overnight. CD spectrum of each sample was used after subtracting the CD spectrum of the solvent.
Native PAGE
15% polyacrylamide gels (acrylamide: bisacrylamide = 29:1) prepared as 8.5 × 8.5 × 0.1 cm glass plates (BioCraft, Tokyo) in 45 mM Tris/45 mM boric acid containing 10 mM KCl or 10 mM NaCl. The buffers contained in each of the gels were used as the electrode buffers. G-tetrad bands were detected by staining with ~20 μM NMM for 5 min and using ChemiDoc Touch (SYBR Gold filter setting, BioRad). Prestaine marker for small RNA (DynaMarker, BioDynamics Laboratory, Tokyo, Japan; DM253) and dsRNA (New England BioLabs, MA, U.S.A.; N0363S). The gels were then washed and further stained with SYBRTM Gold for 5 min to detect total RNA. The two lowest ssRNA 30 and 20 base markers are fluorescent and the fluoresce is quenched by SYBRTM Gold. The dsRNA marker was only stained by SYBRTM Gold.
Tm analysis
G12(S) (4 μM) was incubated overnight at room temperature in 10 mM Tris/HCl containing 30 mM KCl (K) or 30 mM NaCl (Na), as well as in Na-K phosphate (Na-K). Tm was determined using UV-Vis spectrophotometer Model UV-1900i (Shimadzu, Kyoto, Japan). Optical density at 295 nm was monitored every 0.5 °C from 20 to 110 °C.
QUMA-1 stain for the cells
A549 cells were cultured in 96-well plates (ViewPlate). After being washed with MEM, cells were cultured in the presence of 2 μM of each PS-RNA for 17 h. Cells were then washed with MEM and fixed with 4% paraformaldehyde in PBS for 15 min on ice. After washing three times with PBS, the cells were observed after 7 h or later by adding PBS containing 50 nM QUMA-1 and 1 μg/mL DAPI. Confocal imaging was performed using LSM900 (Zeiss) equipped with Airyscan 2 using Plan-Apochromat 63x/1.40 Oil DIC M27 (objective). Excitation and detection wavelengths were 353 nm laser and 465 nm for DAPI, 612 nm laser and 630 nm (AF610) for QUMA-1. Images of 8 slices (7 µm thick) were processed by Airyscan-2 were superimposed by ZEN (Zeiss). TIFF files (16 bit) were exported and intensity values of QUMA-1 fluorescence were quantitated using Fiji (ImageJ 2.14.0). To quantify PS-RNA specific QUMA-1 fluorescence, Image-J (Fiji) was used to set the threshold at approximately 4000 to exclude autofluorescence (lower panel). The DAPI stained area of each image was quantified and divided by the number of cells to calculate the area per cell. The brightness value per cell was divided by this value to correct for the cell size between the images.
Construction of a docking model
Amino acid sequence of the SARS-CoV-2 N protein was obtained from the UniProt database (ID: P0DTC9). BL2-RNA (12 nucleotides) is the RNA form of Block2∆15, and its sequence is 5’-GUGGUGGUGGUG-3’. 3D structure of the BL2-RNA dimer in the presence of three K+ ions and were generated by AlphaFold3 server54 to form a G-quadruplex15. The SARS-CoV-2 N protein dimer was also generated by AlphaFold3 server. Then docking simulation of N protein (dimer) and BL2 (dimer) was performed using HADDOCK 2.456. In the docking simulation, the active site of N protein was set to the N-terminal domain (48-160)57 and the amino acid residues whose Cα pLDDT is lower than 70 were removed. From the resulting 22 clusters, two clusters (four docking poses in total) were selected with HADDOCK scores of 10.3 and 0.2. The resulting structures were visualized using the PyMOL Molecular Graphics System (Version 2.5, Schrödinger, LLC).
Statistics and Reproducibility
Statistical analysis was performed with GraphPad Prism 10 software. Data were presented as box-and-whisker plots (min to max, show all points) with the P-values. The statistical analysis protocol used is indicated in the figure legends. Each experiment was performed at least twice with triplicate independent experimental samples (most experiments were performed more than three times); however, due to animal welfare considerations and limited access to research facilities, only the animal experiments and SARS-CoV-2 experiments shown in the data were performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Baker, R. E. et al. Infectious disease in an era of global change. Nat. Rev. Microbiol. 20, 193–205 (2022).
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Neumann, G. & Kawaoka, Y. Predicting the Next Influenza Pandemics. J. Infect. Dis. 219, S14–S20 (2019).
Liu, D. X. L. & Fung, J. Q. T.S. Human Coronavirus-229E, -OC43, -NL63, and -HKU1 (Coronaviridae). Encycl. Virol. 2, 428–440 (2021).
Su, S. et al. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol. 24, 490–502 (2016).
Matoba, Y. et al. Detection of the human coronavirus 229E, HKU1, NL63, and OC43 between 2010 and 2013 in Yamagata, Japan. Jpn J. Infect. Dis. 68, 138–141 (2015).
Kim, H., Webster, R. G. & Webby, R. J. Influenza Virus: Dealing with a Drifting and Shifting Pathogen. Viral Immunol. 31, 174–183 (2018).
Urbaniak, K. & Markowska-Daniel, I. In vivo reassortment of influenza viruses. Acta Biochim. Pol. 61, 427–431 (2014).
Hussain, M., Galvin, H. D., Haw, T. Y., Nutsford, A. N. & Husain, M. Drug resistance in influenza A virus: the epidemiology and management. Infect. Drug Resist 10, 121–134 (2017).
Imai, M. et al. Influenza A variants with reduced susceptibility to baloxavir isolated from Japanese patients are fit and transmit through respiratory droplets. Nat. Microbiol. 5, 27–33 (2020).
Fukase, K. Middle Molecular Strategy, Flow Synthesis to Functional Molecules (eds. Fukase, K. & Doi, T.) v-vii (Springer, 2021).
Amorim, M. J., Kao, R. Y. & Digard, P. Nucleozin targets cytoplasmic trafficking of viral ribonucleoprotein-Rab11 complexes in influenza A virus infection. J. Virol. 87, 4694–4703 (2013).
Sekine, R. et al. Inhibition of SARS-CoV-2 nucleocapsid protein-RNA interaction by guanosine oligomeric RNA. J. Biochem. 173, 447–457 (2023).
Chung, W. J. et al. Structure of a left-handed DNA G-quadruplex. Proc. Natl Acad. Sci. USA 112, 2729–2733 (2015).
Bakalar, B., Heddi, B., Schmitt, E., Mechulam, Y. & Phan, A. T. A Minimal Sequence for Left-Handed G-Quadruplex Formation. Angew. Chem. Int. Ed. Engl. 58, 2331–2335 (2019).
Faure-Perraud, A. et al. The guanine-quadruplex aptamer 93del inhibits HIV-1 replication ex vivo by interfering with viral entry, reverse transcription and integration. Antivir. Ther. 16, 383–394 (2011).
Ruggiero, E., Zanin, I., Terreri, M. & Richter, S. N. G-Quadruplex Targeting in the Fight against Viruses: An Update. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms222010984 (2021).
Majee, P. et al. Inhibition of Zika virus replication by G-quadruplex-binding ligands. Mol. Ther. Nucleic Acids 23, 691–701 (2021).
Zou, M. et al. G-quadruplex binder pyridostatin as an effective multi-target ZIKV inhibitor. Int. J. Biol. Macromol. 190, 178–188 (2021).
Eckstein, F. Phosphorothioates, essential components of therapeutic oligonucleotides. Nucleic Acid Ther. 24, 374–387 (2014).
Schmidt, E. K., Clavarino, G., Ceppi, M. & Pierre, P. SUnSET, a nonradioactive method to monitor protein synthesis. Nat. Methods 6, 275–277 (2009).
Hartmann, G. Nucleic Acid Immunity. Adv. Immunol. 133, 121–169 (2017).
V’Kovski, P., Kratzel, A., Steiner, S., Stalder, H. & Thiel, V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 19, 155–170 (2021).
Kim, D. et al. The Architecture of SARS-CoV-2 Transcriptome. Cell 181, 914–921.e910 (2020).
Bouhaddou, M. et al. The Global Phosphorylation Landscape of SARS-CoV-2 Infection. Cell 182, 685–712 e619 (2020).
Yao, H. et al. Molecular Architecture of the SARS-CoV-2 Virus. Cell 183, 730–738.e713 (2020).
Schirtzinger, E. E., Kim, Y. & Davis, A. S. Improving human coronavirus OC43 (HCoV-OC43) research comparability in studies using HCoV-OC43 as a surrogate for SARS-CoV-2. J. Virol. Methods 299, 114317 (2022).
Dube, M. et al. Axonal Transport Enables Neuron-to-Neuron Propagation of Human Coronavirus OC43. J. Virol. 92, https://doi.org/10.1128/JVI.00404-18 (2018).
Niu, J. et al. Non-invasive bioluminescence imaging of HCoV-OC43 infection and therapy in the central nervous system of live mice. Antivir. Res. 173, 104646 (2020).
Vakulenko, Y., Deviatkin, A., Drexler, J. F. & Lukashev, A. Modular Evolution of Coronavirus Genomes. Viruses 13, https://doi.org/10.3390/v13071270 (2021).
Klein, S. et al. SARS-CoV-2 structure and replication characterized by in situ cryo-electron tomography. Nat. Commun. 11, 5885 (2020).
Knoops, K. et al. SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol. 6, e226 (2008).
Malone, B., Urakova, N., Snijder, E. J. & Campbell, E. A. Structures and functions of coronavirus replication-transcription complexes and their relevance for SARS-CoV-2 drug design. Nat. Rev. Mol. Cell Biol. 23, 21–39 (2022).
Dou, D., Revol, R., Ostbye, H., Wang, H. & Daniels, R. Influenza A Virus Cell Entry, Replication, Virion Assembly and Movement. Front. Immunol. 9, 1581 (2018).
Kim, C. U. et al. Influenza neuraminidase inhibitors possessing a novel hydrophobic interaction in the enzyme active site: design, synthesis, and structural analysis of carbocyclic sialic acid analogues with potent anti-influenza activity. J. Am. Chem. Soc. 119, 681–690 (1997).
Yamashita, M. et al. CS-8958, a prodrug of the new neuraminidase inhibitor R-125489, shows long-acting anti-influenza virus activity. Antimicrob. Agents Chemother. 53, 186–192 (2009).
Amorim, M. J. et al. A Rab11- and microtubule-dependent mechanism for cytoplasmic transport of influenza A virus viral RNA. J. Virol. 85, 4143–4156 (2011).
Eisfeld, A. J., Kawakami, E., Watanabe, T., Neumann, G. & Kawaoka, Y. RAB11A is essential for transport of the influenza virus genome to the plasma membrane. J. Virol. 85, 6117–6126 (2011).
Momose, F., Kikuchi, Y., Komase, K. & Morikawa, Y. Visualization of microtubule-mediated transport of influenza viral progeny ribonucleoprotein. Microbes Infect. 9, 1422–1433 (2007).
Pohl, M. O., Lanz, C. & Stertz, S. Late stages of the influenza A virus replication cycle-a tight interplay between virus and host. J. Gen. Virol. 97, 2058–2072 (2016).
de Castro Martin, I. F. et al. Influenza virus genome reaches the plasma membrane via a modified endoplasmic reticulum and Rab11-dependent vesicles. Nat. Commun. 8, 1396 (2017).
Han, J. et al. Host factor Rab11a is critical for efficient assembly of influenza A virus genomic segments. PLoS Pathog. 17, e1009517 (2021).
Momose, F. et al. Apical transport of influenza A virus ribonucleoprotein requires Rab11-positive recycling endosome. PLoS One 6, e21123 (2011).
Zhang, Z., Dai, J., Veliath, E., Jones, R. A. & Yang, D. Structure of a two-G-tetrad intramolecular G-quadruplex formed by a variant human telomeric sequence in K+ solution: insights into the interconversion of human telomeric G-quadruplex structures. Nucleic Acids Res. 38, 1009–1021 (2010).
Li, M., Zhao, A., Ren, J. & Qu, X. N-Methyl Mesoporphyrin IX as an Effective Probe for Monitoring Alzheimer’s Disease beta-Amyloid Aggregation in Living Cells. ACS Chem. Neurosci. 8, 1299–1304 (2017).
Mergny, J. L., Phan, A. T. & Lacroix, L. Following G-quartet formation by UV-spectroscopy. FEBS Lett. 435, 74–78 (1998).
Del Villar-Guerra, R., Trent, J. O. & Chaires, J. B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angew. Chem. Int. Ed. Engl. 57, 7171–7175 (2018).
Zhang, S., Wu, Y. & Zhang, W. G-quadruplex structures and their interaction diversity with ligands. ChemMedChem 9, 899–911 (2014).
Nicoludis, J. M., Barrett, S. P., Mergny, J. L. & Yatsunyk, L. A. Interaction of human telomeric DNA with N-methyl mesoporphyrin IX. Nucleic Acids Res. 40, 5432–5447 (2012).
Binas, O., Bessi, I. & Schwalbe, H. Structure Validation of G-Rich RNAs in Noncoding Regions of the Human Genome. Chembiochem 21, 1656–1663 (2020).
Ngo, K. H., Liew, C. W., Lattmann, S., Winnerdy, F. R. & Phan, A. T. Crystal structures of an HIV-1 integrase aptamer: Formation of a water-mediated A*G*G*G*G pentad in an interlocked G-quadruplex. Biochem. Biophys. Res. Commun. 613, 153–158 (2022).
Sengar, A., Heddi, B. & Phan, A. T. Formation of G-quadruplexes in poly-G sequences: structure of a propeller-type parallel-stranded G-quadruplex formed by a G(1)(5) stretch. Biochemistry 53, 7718–7723 (2014).
Chen, X. C. et al. Tracking the Dynamic Folding and Unfolding of RNA G-Quadruplexes in Live Cells. Angew. Chem. Int. Ed. Engl. 57, 4702–4706 (2018).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Vandervaart, J. P. et al. Serodominant SARS-CoV-2 Nucleocapsid Peptides Map to Unstructured Protein Regions. Microbiol Spectr. 11, e0032423 (2023).
Honorato, R. V. et al. The HADDOCK2.4 web server for integrative modeling of biomolecular complexes. Nat. Protoc. https://doi.org/10.1038/s41596-024-01011-0 (2024).
Dinesh, D. C. et al. Structural basis of RNA recognition by the SARS-CoV-2 nucleocapsid phosphoprotein. PLoS Pathog. 16, e1009100 (2020).
Lyons, S. M. et al. eIF4G has intrinsic G-quadruplex binding activity that is required for tiRNA function. Nucleic Acids Res. 48, 6223–6233 (2020).
Schult, P., Kummerer, B. M., Hafner, M. & Paeschke, K. Viral hijacking of hnRNPH1 unveils a G-quadruplex-driven mechanism of stress control. Cell Host Microbe 32, 1579–1593.e1578 (2024).
Kharel, P. & Ivanov, P. Identification of bona fide RNA G-quadruplex binding proteins. Methods Enzymol. 695, 255–274 (2024).
Roberts, T. C., Langer, R. & Wood, M. J. A. Advances in oligonucleotide drug delivery. Nat. Rev. Drug Discov. 19, 673–694 (2020).
Moriuchi, H. et al. Human malignant melanoma cell line (HMV-II) for isolation of influenza C and parainfluenza viruses. J. Clin. Microbiol. 28, 1147–1150 (1990).
Matsuyama, S. et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc. Natl Acad. Sci. USA 117, 7001–7003 (2020).
Shirato, K. et al. Differences in neutralizing antigenicity between laboratory and clinical isolates of HCoV-229E isolated in Japan in 2004-2008 depend on the S1 region sequence of the spike protein. J. Gen. Virol. 93, 1908–1917 (2012).
Acknowledgements
This research was carried out based on the suggestions of the late A. Nomoto, with the continuing advice of Shinnyo. We thank Prof. G.-W. Hwang for the use of his RT-PCR equipment. We also thank Y. Nishimura for the vibratome instruction and H. Irokawa for laboratory maintenance and management. We thank K. Watashi and S. Nakajima (National Institute of Infectious diseases) for initial analysis for effect of RNA oligomers on SARS-CoV-2 RNA production in culture medium. Additionally, we thank D. Miyoshi, K. Kawauchi and M. Tsuruta for their advices on the G-quadruplex analyses, and H. Yoshikura and Y. Yoshimura for reading the manuscript and for his kind comments. This work was supported by JSPS (Japan Society for the Promotion of Science) KAKENHI (grant numbers: 18K06630 and 22K06616) and the Japan Agency for Medical Research and Development (JP223fa627001 and JP22wm0125002).
Author information
Authors and Affiliations
Contributions
R.S. performed in vivo and in vitro OC43 experiment studies; S.K. performed most of 229E and IAV experiments and G-quadruplex experiment studies; S.T., T.S., T.Kaiya and K.T. contributed to experiments; M.K., S.Y. and Y.K. performed SARS-CoV-2 experiments; S.O. and Y.Y performed docking simulation of SARS-CoV-2 NP and BL2 RNA; Y.T. and S.N. performed Tm analyses; K.S., K.I., T.Kanda, Y.K. and H.N. advised on all aspects of the study; S.K. designed the study and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Wen Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christina Karlsson Rosenthal. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sekine, R., Takeda, K., Suenaga, T. et al. G-quadruplex-forming small RNA inhibits coronavirus and influenza A virus replication. Commun Biol 8, 27 (2025). https://doi.org/10.1038/s42003-024-07351-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-07351-7