Abstract
The regulatory mechanism between N6-methyladenosine (m6A) RNA methylation and histone modification in endometrial receptivity remains poorly understood. In this study, we depict that RIF induced m6A and Mettl3 level restrain, affecting H3K27me3 modification and chromatin accessibility. We show that Mettl3 deletion in the endometrium alters mRNA m6A methylation via Eed interaction. This reduces m6A recognized by Ythdc1, which recruits Eed to suppress H3K27me3 modification co-transcriptionally. The reduction of H3K27me3 disrupts chromatin accessibility and impairs transcription of genes critical for endometrial receptivity. Collectively, these results shed light on a Mettl3-Eed-m6A-Ythdc1 axis that links m6A and histone modification in regulating local chromatin state and gene expression, advancing our understanding of the epigenetic crosstalk between RNA and DNA modification in infertility disease.
Similar content being viewed by others
Introduction
Endometrial receptivity is crucial for embryo-attachment, invasion, and subsequent development, which are essential for individual formation and species survival. In the majority of healthy women, the implantation window typically lasts 3–6 days during the secretory phase1. Endometrial dysfunction can lead to severe conditions such as infertility, endometriosis, and recurrent implantation failure (RIF)2. RIF is typically characterized by the inability to achieve a clinical pregnancy despite the transfer of at least 4 high-quality embryos over a minimum of 3 frozen or fresh in vitro fertilization cycles. Insufficient and impaired decidualization of the endometrium is regarded as the primary underlying cause of RIF3. Recent studies have analyzed of human endometrial menstrual cycle samples at single-cell resolution, providing a more precise estimation of the embryo implantation window4. The spatiotemporal dynamic atlas of single-cell endometrial profiling has also revealed insights into the molecular and cellular abnormalities occurring in endometrial diseases5. However, the precise epigenetic mechanisms regulating decidualization and endometrial function remain unclear.
Emerging evidence suggests that epigenetic aberrations, such as alterations in histone acetylation6, contribute to endometrial disorders. Histone modifications, such as methylation and acetylation, play critical roles in transcriptional regulation7. In particular, H3K27me3, the repressive mark deposited by the Polycomb Repressive Complex 2 (PRC2), is associated with transcriptional repression8. PRC2 consists of key components, including Embryonic Ectoderm Development (EED), Enhancer of Zeste 1 and 2 (EZH1/2), and Suppressor of Zeste 12 (SUZ12)9. EZH2 acts as the methyltransferase for H3K27me3, with full activity requiring SUZ12 and EED10. Dynamic regulation of H3K27me3 levels in decidual stromal cells is crucial for pregnancy and parturition11. During decidualization, EZH2 in the endometrium suppresses cell-cycle-related genes via H3K27 trimethylation, and EZH2-deficient mice in the uterine epithelium and stroma exhibit significant subfertility12. Moreover, EZH2-deficient mice often experience mid-gestation failure due to ectopic myofibroblast formation13. Although some reports have explored histone modifications in the endometrium and embryonic development, further investigation is needed to explore the crosstalk between histone modifications and N6-methyladenosine (m6A) RNA modifications.
m6A, the most prevalent RNA modification in eukaryotes, regulates RNA splicing, translocation, stability, and translation14. Its functions are orchestrated by proteins known as writers, erasers, and readers15. The METTL3-METTL14 complex serves as the primary writer of m6A, with METTL3 as the catalytic core and METTL14 as an RNA-binding platform16. The catalytic cavity of METTL3 contains three primary loops: two near the perimeter of the SAM binding site and a larger third loop that interacts with METTL1417. METTL3 is essential for decidualization in human endometrial stromal cells, with Mettl3-deficient mice failing to respond to uterine artificial decidualization18. And Mettl3 coordinates transcriptomic and epitranscriptomic regulation during developmental pausing, impacting stem cell dormancy19. Our previous study revealed that the RNA-modifying enzyme NSUN2 upregulates H3K9ac by suppressing SIRT4 expression, thereby significantly influencing endometrial receptivity20. Resistance to progesterone signaling and activation of estrogen signaling are associated with decreased levels of METTL3 in the endometrium21. m6A is also recognized by reader proteins, such as YTH domain-containing proteins (YTHDC1/2 and YTHDF1/2/3)15. As a critical m6A reader, YTHDC1 recruits the H3K9me2 demethylase KDM3B to m6A-associated regions, promoting the demethylation of H3K9me2 in the corresponding chromatin regions and facilitating the transcription of relevant genes22. And YTHDC1 is essential for early embryonic development, maintaining stem cell identity by repressing retrotransposons and Dux via SETDB1-mediated H3K9me3 modifications23. However, the crosstalk between RNA modifications and DNA epigenetics, particularly their mechanisms in endometrial receptivity, remains unexplored.
In this study, we performed m6A RIP sequencing and found low methylation levels in the RIF endometrium, linking this to histone modification and chromatin remodeling. Using METTL3 deletion models, we demonstrated that METTL3 recruits EED and interacts with YTHDC1 to modify EED m6A, suppressing EED transcription and inhibiting H3K27me3. This disruption leads to impaired endometrial function. This is the first study to uncover the METTL3-EED-YTHDC1 axis in regulating endometrial receptivity, offering new insights into infertility-related epigenetics. This study represents a substantial contribution to the field of reproductive epigenetics. The insights into the METTL3/EED/YTHDC1 axis offer a promising avenue for understanding and potentially treating RIF.
Results
Low m6A levels cause RIF
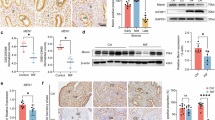
The immunofluorescence results indicated that the positive cells of endometrial function genes (LIF, HOXA10, HAND2, CK18) were decreased in endometrial tissue with RIF (Fig. 1A-D, Supplementary Fig. 1A, B). The m6A level was inhibited in the RIF group as indicated by dot blot (Fig. 1E, F), and m6A-RIP-seq revealed differences in m6A modification between the RIF and NC groups by principal component analysis (PCA) (Fig. 1G). The number of m6A modified genes and peaks decreased in RIF groups (Fig. 1H), While the transcriptional level moderately increased (Fig. 1I, J). Differential gene expression analysis between the RIF and NC groups showed that 1314 m6A genes were down-regulated whereas 7 m6A genes were up-regulated (Fig. 1K). GSEA analysis linked these m6A modificatory differential genes to important biological function (Adhesion, Epithelial-mesenchymal transition, Stem cell, Apoptosis) and histone regulation (H3K27me3, EED, H3K4me3, ESR1) in Fig. 1L-O and Supplementary Fig. 1C-F. Besides, GO analysis exhibited that m6A modificatory differential genes were enriched in the pathways of Histone modification, Chromatin remodeling and DNA damage (Fig. 1P). Immunofluorescence results found that RIF could lead to the reduction of m6A RNA-methyltransferase METTL3 and METTL14 positive cells (Fig. 1Q-S). In summary, the reduction in m6A levels correlates with increased expression of endometrial function genes, suggesting an epigenetic regulatory mechanism.
A–D Immunofluorescence method tested the endometrial function (LIF, HOXA10, HAND2) in the NC and RIF groups. E, F Dot Blot detected the m6A level in the NC and RIF groups. G PCA method analyzed the inter-group differences between the NC and RIF groups. H The number of m6A modificatory genes and peaks were calculated. I, J The m6A modification and mRNA levels were compared in the NC and RIF groups. K. The volcano plot showed the m6A modificatory difference genes. L-O. GSEA analysis was carried out for m6A modificatory difference genes. P GO analysis was carried out for m6A modificatory difference genes. Q-S. Immunofluorescence method to test the positive cell numbers of METTL3 and METTL14 in the RIF group. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. For all IF, n = 2. NC Normal control, RIF Recurrent implantation failure.
METTL3 Knockdown promoted apoptosis, inhibited proliferation and affected histone modification
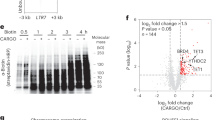
To confirm whether METTL3 affected the endometrial function, METTL3 knockdown (METTL3KD) Ishikawa cell lines were established. METTL3KD resulted in reduced protein levels of METTL3 and METTL14 (Fig. 2A, B). and decreased m6A levels (Fig. 2C, D). Meanwhile, the adherence rate of the BeWo sphere to the METTL3KD cells was reduced (Fig. 2E,F). Increased apoptosis level (BAK and TUNEL) and decreased proliferation level (CCK8 and KI67) were observed after METTL3 knockdown by western blot and immunofluorescence experiments (Supplementary Fig. 2A-G). To investigate the regulatory genes targeted by METTL3, m6A-RIP-seq was performed. The results revealed that the overall m6A-modified genes displayed high similarity within the same groups, while were more diverse across groups (Fig. 2G). The METTL3KD reduced the number of m6A modificatory genes and peaks (Fig. 2H). Following METTL3 knockdown, the m6A level decreased, while the mRNA level increased slightly (Fig. 2I-L). A set of 1754 common genes was identified between different m6A modificatory and transcriptional genes (Fig. 2M, N). GO-BP and GO-MF analyses showed that these common genes were involved in histone and chromatin modification (Fig. 2O). In addition, 8 common genes were found to overlap between different m6A modificatory genes, transcriptional genes and endometrial function markers, and a positive correlation was observed between the m6A and mRNA levels of these common genes (Supplementary Fig. 2H, I). Meanwhile, m6A levels declined while mRNA levels increased in the common genes (Fig. 2P, Q). Besides, METTL3 deletion inhibited H3K27me3 and elevated the H3K27ac level significantly (Fig. 2R-U). In sum, METTL3 knockdown disrupts endometrial function by reducing m6A modifications and altering histone marks, providing insight into its role in regulating gene expression and endometrial receptivity.
A-B. Western Blot tested the expression levels of METTL3 and METTL14 after METTL3KD. C, D. Dot Blot method tested the m6A level in the METTL3KD cell line. E-F. The adhesion rate of BeWo to the METTL3KD cells. G. The correlation analysis was carried out in the METTL3KD and EV groups. H. m6A modificatory genes and sites were calculated. I, J. The m6A modification and mRNA level were compared. K-L. The m6A modificatory difference genes were analyzed. M. The difference genes with mRNA levels were analyzed. N. The venn analysis was carried out for m6A and mRNA common genes. O. GO analysis was carried out for the common genes. P, Q The comparison of m6A modification and mRNA level was carried out for the common genes. R, S. Western Blot tested the H3K27me3 levels in the METTL3KD cell line. T, U. Western Blot tested the H3K27ac levels in the METTL3KD cell line. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. EV = Empty vector.
The Mettl3 conditional knockout mouse model established and influenced endometrial function
To investigate the role of Mettl3 in regulating endometrial receptivity, Pgr-Cre transgenic mice were crossed with Mettl3f/f mice (Fig. 3A, B). Immunofluorescence and western blot analysis at gestation day 4 (GD 4) revealed significant reduction in the Mettl3 and Mettl14 expression in endometrial tissue, but not in ovary tissue (Supplementary Fig. 3A-E). Phenotypic study found that Mettl3 conditional knockout (Mettl3d/d) resulted in smaller and thinner endometrial tissue compared to the M3f/f groups (Fig. 3C). Besides, the Mettl3 deletion decreased the uterine weight and inhibited the litter size significantly (Fig. 3D, E). The hematoxylin and eosin staining of M3d/d groups showed thinner epithelial layers and fewer endometrial glands than in the M3f/f groups (Fig. 3F-I). The endometrial function test found that Mettl3 lack reduced the percentage of Lif, Hoxa10 and Vimentin positive cells in the epithelial layer and stromal layer (Fig. 3J-M, Supplementary Fig. 3F, G). Furthermore, Mettl3d/d promoted the number of positive cells expressing apoptotic activating genes (Caspase3, Bim and p-Bad) in Fig. 3N-Q and Supplementary Fig. 3H, I, while decreasing the number of positive cells with suppressive apoptotic genes (Bcl2) in Supplementary Fig. 3J, K. In addition, Mettl3d/d accelerated the process of endometrial aging, as evidenced by the number of positive cells expressing p27 and p21 (Supplementary Fig. 3L-O). In total, Mettl3 knockout in mice impaired endometrial structure and function, promoted apoptosis, accelerated aging, and reduced fertility, highlighting its crucial role in endometrial receptivity.
A Diagram demonstrating the strategy used for conditional Mettl3 deletion. Conditional Mettl3 deletion (Mettl3d/d) mice were generated by crossing Mettl3f/f mice with mice of the Pgr-Cre driver line. B Genotyping analysis showing the Mettl3 knockout efficiency at the DNA level in the endometrium. C Phenotyping analysis showing the change of endometrial tissue after Mettl3 deletion. D The weight of the uterus was estimated. E The litter size was estimated. F, G The endometrium of epithelial thickness. Scale bars, 50 μm. H, I The number of endometrial glands was tested after Mettl3d/d. Scale bars, 500 μm. J-M IF tested the influence of endometrial function (Lif and Hoxa10) in the Mettl3d/d group. Scale bars, 50 μm. N-Q IF tested the influence of endometrial apoptosis (Caspase 3 and Bim) in the Mettl3d/d group. Scale bars, 50 μm. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. For all IF and HE, n = 3.
m6A RIP-seq for Mettl3d/d endometrial tissue and Mettl3 deletion affected the levels of H3K27me3 and H3K27ac
In order to explore the regulatory mechanism of Mettle3 in endometrial function, m6A RIP-seq was performed on Mettl3-deficient endometrial tissue, and the difference analysis revealed that 835 up-regulated m6A modification genes and 13 down-regulated m6A modification genes (Fig. 4A). Correlation analysis of the differential m6A genes revealed strong similarity within each group and greater diversity between the M3f/f and M3d/d groups (Fig. 4B). The GO analysis performed on these m6A differential m6A genes, demonstrated enrichment in epithelial morphogenesis, epithelial cell proliferation and the histone modification signal pathway (Fig. 4C). Meanwhile, both m6A and mRNA levels of differential genes were significantly reduced after Mettl3d/d (Fig. 4D, E). And the GSEA method was performed on m6A differential m6A genes, indicated that the differential genes were related to epithelium development and organ morphogenesis (Fig. 4F, G). IGV demonstrated that the m6A and transcription levels were inhibited in endometrial functional genes (Hoxa10, Vim, Hand2 and Esr1) in Fig. 4H, I and supplementary Fig. 4A, B. In addition, in order to find out whether Mettl3d/d induced the endometrial receptivity loss by affecting histone modification, the immunofluorescence revealed that the H3K27me3 positive cells were reduced and H3K27ac positive cells were increased in RIF and Mettl3d/d endometrium tissue (Fig. 4J-O). In sum, Mettl3 deletion in endometrial tissue altered m6A and mRNA levels of key genes, with reducing H3K27me3 and increasing H3K27ac, indicating its role in regulating endometrial receptivity through epigenetic mechanisms.
A. Volcano plot showed the m6A modificatory difference genes after Mettl3 deletion. B. The correlation analysis was carried out for m6A modificatory difference genes. C. GO analysis was carried out for m6A modificatory difference genes in the Mettl3d/d group. D, E. The m6A modification and mRNA level were compared in the Mettl3d/d group. F, G. GSEA analysis was carried out for m6A modificatory difference genes. H, I. IGV demonstrated the m6A and mRNA levels of endometrial receptivity genes (Hoxa10 and Vim). J-L. IF tested the positive cell numbers of H3K27me3 and H3K27ac in the RIF group. Scale bars, 50 μm. M-O. IF tested the positive cell numbers of H3K27me3 and H3K27ac in the Mettl3d/d group. Scale bars, 50 μm. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. For all IF, n = 3.
Mettl3d/d affected the transcriptional level through inhibiting H3K27me3 levels and chromatin accessibility
In order to confirm whether Mettl3d/d inhibited endometrial function via affecting H3K27me3 modification and chromatin stability, the CUT&Tag-seq and ATAC-seq were carried out for Mettl3d/d endometrial tissues. CUT&Tag-seq found that H3K27me3 (transcriptional suppressive histone) enriched statistically different sites were divided into increased and decreased groups, and the transcriptional level was accelerated after Mettl3 deletion (Fig. 5A, B). GO and KEGG analyses indicated that the up-regulated pathway related to epithelial cell proliferation, stem cell differentiation, cell adhesion, PI3K-AKT and MAPK, while the down-regulated pathways include autophagy, oxidative stress, mTOR (Fig. 5C). GSEA analysis of these statistically different genes identified increased genes associated with cellular phenotype, female genitalia, inflammatory response and RNA polymerase II, whereas decreased genes were related to p53 targets, chromosome, mitotic cell cycle and muscle weakness (Fig. 5D-G, Supplementary Fig. 5A-D). Meanwhile, the K-mean method was used to evaluate the CUT&Tag-seq data for H3K27me3 and H3K27ac, and identified Cluster 6 (C6) as having inhibited H3K27me3 signal intensity and increased transcriptional levels after Mettl3 loss (Fig. 5H-J). IGV showed distinct enrichment in H3K27me3 and H3K27ac of the endometrial receptivity genes Muc1 and Lif (Fig. 5K, L). Besides, the statistically different sites from ATAC-seq were divided into increased and decreased group, and GO analysis showed that genes derived from both closed and open chromatin sites were enriched in epithelial cell migration, cell adhesion, apoptosis signal pathway (Fig. 5M, N). IGV further demonstrated that chromatin closed sites decreased the mRNA level of Esr1, and chromatin opened sites elevated transcriptional level of Lifr (Fig. 5O, P). All in all, Mettl3 deletion in the endometrium disrupted H3K27me3 modification and chromatin accessibility, leading to altered gene expression and impaired endometrial function.
A The statistically differential sites from CUT&Tag-seq for H3K27me3 were divided into increased and decreased group. B The transcription level analysis was carried out for statistically differential sites. C GO method was used to analyze the statistically differential sites derived genes. D-G GSEA method was used to analyze the statistically differential sites derived genes. H K-mean method was employed to analyze the statistically differential sites from CUT&Tag-seq for H3K27me3 and H3K27ac. I, J The statistically differential sites from cluster 6 (H) were analyzed with signal intensity and transcription rate. K, L The enriched difference of endometrial receptivity genes Muc1 and Lif in H3K27me3 and H3K27ac. M The statistically differential sites from ATAC-seq were divided into closed and opened groups. N GO method was used to analyze the closed and opened sites derived genes. O, P The comparison of transcriptional levels in the closed and opened sites derived genes (Esr1 and Lifr).
Mettl3/Eed/Ythdc1 regulated axis influenced the function of endometrium
To investigate the regulatory relationship between Mettl3 and Eed/Ythdc1, the IF was performed on the RIF samples and Mettl3d/d endometrial tissues. We found that the number of EED, SUZ12 and EZH2 positive cells were reduced (Fig. 6A-D). Meanwhile, Mettl3d/d reduced the percentage of Eed, Suz12 and Ezh2 positive cells in endometrial tissues (Fig. 6E-H). Co-IP analysis of uterine tissues from GD 4 verified a direct interaction between Mettl3 and Eed, but not with Ezh2 or Suz12 (Fig. 6I). Moreover, Co-IP found that Eed has the protein interact with Ezh2, Suz12, Mettl3 and Ythdc1 (Fig. 6J). Interestingly, m6A reader Ythdc1 also directly interacted with Eed, but not with Ezh2 or Suz12 (Fig. 6K). To assess the functional impact, EED overexpression (EEDOE) endometrial epithelium cells were generated (Supplementary Fig. 6A-B). Dot Blot found that EEDOE rescued the m6A level in METTL3KD cells (Supplementary Fig. 6C). Meanwhile, EEDOE elevated the protein level of METTL3 partially in METTL3KD cells, and restored the protein levels of endometrial functional genes (FOXA2 and MUC1) to normal level (Fig. 6L-P). In addition, EEDOE elevated the protein level of H3K27me3 in METTL3KD cells (Fig. 6Q, R). In all, the Mettl3/Eed/Ythdc1 axis regulated endometrial function through coordinated regulation of m6A modification and histone modifications H3K27me3, demonstrating the communication and interaction at different epigenetic levels within the endometrium.
A–D IF tested the positive cell numbers of EED, EZH2 and SUZ12 in the RIF group. Scale bars, 50 μm. E–H IF tested the positive cell numbers of Eed, Ezh2 and Suz12 in the Mettl3d/d group. Scale bars, 50 μm. I Co-IP in mouse endometrial tissue at GD 4 showed that Nsun2 interacted with Eed, except for Ezh2 and Suz12. J Co-IP in mouse endometrial tissue at GD 4 showed that Eed interacted with Ezh2, Suz12, Mettl3 and Ythdc1. K Co-IP in mouse endometrial tissue at GD 4 showed that Ythdc1 interacted with Eed, Ezh2 and Suz12. L-P Western blot tested the protein levels of EED, METTL3, FOXA2 and MUC1 in the EV, METTL3KD and METTL3KD/EEDOE groups. Q, R Western blot tested the protein level of H3K27me3 in the EV, METTL3KD and METTL3KD/EEDOE groups. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. For all IF in RIF group, n = 3, For all IF in Mettl3d/d group, n = 3.
METTL3-mediated m6A affected EED and H3K27me3 modification
To prove whether METTL3 loss decreased the EED and H3K27me3 via regulating m6A modification, the common m6A motif (UUACA) was found in the NC, RIF, EV and METTL3KD groups (Fig. 7A). Therefore, YTHDC1 overexpression (YTHDC1OE) endometrial epithelium cells were generated (Fig. 7B, C). Furthermore, YTHDC1OE elevated the protein level of EED significantly in METTL3KD cells, and rescued the protein expression of endometrial functional genes (FOXA2 and CK18) to normal level (Fig. 7D-G). Besides, YTHDC1OE elevated the protein level of H3K27me3 in METTL3KD cells (Fig. 7H, I), and increased the m6A modification level after METTL3 deletion (Fig. 7J). In order to conduct an in-depth mechanism study, the exogenous mutation of METTL3 functional domain (S-adenosyl methionine binding site) cells was produced. Dot Blot showed that the METTL3 mutation (METTL3mut) restrained the m6A level (Fig. 7K), reversed the inhibiting effect of METTL3OE on EED (Fig. 7L-N), and abolished the inhibiting effect of METTL3OE on H3K27me3 (Fig. 7O, P). In sum, METTL3 loss reduced EED and H3K27me3 via m6A modification, while YTHDC1 overexpression rescued these effects, demonstrating m6A’s role in regulating endometrial function.
A The common m6A motifs at EED were performed by Homer in the NC, RIF, EV and METTL3KD groups. B, C Western blot tested the protein level of YTHDC1 after YTHDC1 overexpression (YTHDC1OE). D-G Western blot tested the protein levels of EED, CK18 and FOXA2 in the YTHDC1OE. H, I Western blot tested the protein level of H3K27me3 in the YTHDC1OE. J Dot Blot detected the m6A level in the EV, METTL3KD and METTL3KD/YTHDC1OE groups. K Dot Blot detected the m6A level in the EV, METTL3OE and METTL3mut groups. L–N Western blot tested the protein levels of METTL3 and EED in the EV, METTL3OE and METTL3mut groups. O, P Western blot tested the protein level of H3K27me3 in the EV, METTL3OE and METTL3mut groups. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD.
Inhibition of EED affected H3K27me3 and the endometrial receptivity
To confirm the impact of EED deletion on endometrial function, endometrial epithelial cells were treated with the EED inhibitor (MAK683) at concentrations of 0, 20, and 40 nM. At 40 nM, MAK683 significantly decreased the protein levels of EED, SUZ12 and EZH2, and altered the endometrial receptivity genes (HOXA10, LIF and FOXA2) significantly (Fig. 8A-H). Meanwhile, 40 nM MAK683 reduced the H3K27me3 while increasing H3K27ac modification obviously (Fig. 8I-K). In vivo, injection of 100 μl MAK683 (2.7 mg/ml) into cornua uterus at GD 3.5, followed by HE staining at GD 5.5 showed. that the EED inhibitor reduced the thickness of endometrial epithelium (Fig. 8L, M). In addition, MAK683 treatment restrained the number of Eed and H3K27me3 positive cells (Fig. 8N, O). while decreasing the number of Ck18 positive cells and dramatically increasing the number of Foxa2 and Muc1 positive cells (Fig. 8P-R). In a word, the interaction between METTL3 and Eed suggests a novel regulatory axis for histone modifications in endometrial receptivity, warranting further investigation in other reproductive disorders.
A-H. Western Blot tested the protein level of PRC2 complex genes (EED, SUZ12, EZH2) and endometrial receptivity genes (HOXA10, LIF, FOXA2) after 0, 20 and 40 nM MAK683 treated with endometrial epithelium cells. I-K Western Blot tested the protein levels of H3K27me3 and H3K27ac after 0, 20 and 40 nM MAK683 treatment with endometrial epithelium cells. L,M HE method detected the thickness of endometrial epithelium after 100 μl MAK683 (2.7 mg/ml) treatment in vivo. Scale bars, 50 μm. N, O. IF tested the positive cell numbers of Eed and H3K27me3 after MAK683 treatment. Scale bars, 50 μm. P-R IF tested the positive cell numbers of Ck18, Foxa2 and Muc1 after MAK683 treatment. Scale bars, 50 μm. S The mechanism schematic of this article. Significance was calculated using t test in all panels. *p < 0.05, **p < 0.01, ***p < 0.001. All error bars are ± SD. For all IF and HE, n = 3.
Discussion
To explore how m6A regulates chromatin and associated enzymes, we conduct a thorough analysis of the m6A transcriptome and other epigenomic factors, along with METTL3 deletion models both in vivo and in vitro. In the endometrium, we identify for the first time that METTL3 recruits EED and interacts with YTHDC1 to identify the m6A modification of EED. This interaction suppresses EED transcription, inhibiting H3K27me3 and ultimately leading to subfertility by affecting endometrial receptivity METTL3 and METTL14 constitute the central heterodimeric core of the primary RNA m6A methyltransferase complex (MTC) that responsible for m6A deposition16. Our findings revealed that METTL3 depletion reduces H3K27me3 and increases H3K27ac levels, indicating distinct regulatory mechanisms for repressive and activating histones in endometrial function. Notable changes in histone modifications, especially increased H3K27ac levels, have been observed in both the proximal and distal regions during uterine decidualization24. Moreover, elevated H3K27ac levels in the distal upstream region of IGFBP1 significantly enhance IGFBP1 expression in endometrial stromal cells during decidualization25. Meanwhile, in the mESC cell line, reducing m6A methylation via METTL3 depletion leads to increased H3K27ac supporting our conclusion26.Our findings suggest that METTL3 modulates both repressive and activating histones, likely influenced by upstream enzymes. The histone-modifying enzymes and associated substrates that regulate H3K27ac and H3K27me3 are completely distinct27. Although we primarily discuss the mechanism of m6A modification in regulating H3K27me3 in the endometrium. Future research should elucidate the upstream and downstream mechanisms of H3K27ac in endometrial function.
The m6A reader YTHDC1 exhibits diverse functions, promoting both transcriptional activation and repression22,28. YTHDC1-mediated transcriptional activation involves direct interaction with chromatin remodelers, recruiting KDM3B to remove repressive H3K9me2 mark22. Besides, YTHDC1-dependent retrotransposon transcript silencing is an important mechanism for early embryonic stem cells (ESCs) to maintain heterochromatin integrity and genomic stability26. In general, the binding of YTHDC1 to m6A-modified mRNAs promotes transcriptional activation22, while its interaction with lncRNA or retroviral-associated RNAs leads to transcriptional repression through heterochromatin silencing26,28. Our findings support YTHDC1’s role in transcription activation, but we propose it may not be the only m6A reader regulating epigenetic mechanisms in the endometrium. Among numerous m6A readers, YTHDC1 uniquely displays sequence selectivity29, predominantly recognizing the DRACH motif in mES cell line30. Yet its motif specificity in the endometrium requires further investigation. Furthermore, current research lacks any in vivo or in vitro investigations on the role of YTHDC1 in the endometrium. The establishment of YTHDC1 deletion models in the future would provide strong support for further elucidating the epigenetic mechanisms that regulate endometrial function.
Recent studies have highlighted the significant role of transposable elements (TEs) in endometrial function. For instance, TEs such as Alu, LINE-1 are integrated into the regulatory framework of decidualizing stromal cells during the implantation window31. Moreover, ancient TEs have been found to play a crucial role in the cis-regulatory mechanisms involved in decidualization32. m6A methylation may indirectly impact the endometrium by silencing or activating TEs, which are involved in histone modification regulation. For instance, H3K27 hyperacetylation increase chromatin accessibility and enhancer RNA expression during endometrial invasion33. H3K27ac and H3K4me1 promote the interaction between enhancers and promoters, resulting in optimal Hand2 transcription during regular stromal-epithelial communication34. Our results proved that the METTL3/EED axis affects H3K27me3 levels and thereby modulates endometrial receptivity, which may be related to regulating TEs. In addition, m6A methylation directly impact ERVs, through the METTL3-METTL14 complex, resulting in elevated levels of IAPs and related ERVK elements, eventually controlling the fate of ERVs35. Meanwhile, FTO-mediated m6A demethylation of LINE1 RNA affects transcription levels, thereby influencing the chromatin state36. Based on these articles, we propose that the dual modifications of histones and m6A methylation in TEs have an indirectly or directly effect on in the endometrium function. However, the crosstalk between histone modifications and RNA modifications such as m6A in TEs need to be explored, and advanced RNA or histone epigenetic tools will boost progress in this field.
In summary, our study provides key insights into the role of the METTL3-EED-YTHDC1 axis in regulating endometrial receptivity. Our study indicates that METTL3 recruits EED and interacts with YTHDC1 to identify the m6A modification on EED. This interaction suppresses EED transcription, inhibiting H3K27me3 and ultimately disrupting endometrial function (Fig. 8S). Targeting the METTL3/EED axis could offer promising novel therapeutic strategies for improving endometrial receptivity and addressing RIF. More importantly, this study provides groundbreaking insights into the Mettl3/EED/Ythdc1 axis as a key regulator of endometrial receptivity, unveiling a novel layer of epigenetic regulation that could be targeted for therapeutic benefit. By elucidating how m6A modification impacts histone modifications and chromatin remodeling, we have identified potential therapeutic avenues for improving implantation success in patients with RIF. Moreover, our findings pave the way for future studies exploring the complex interactions between RNA modifications and histone modifications in reproductive health, opening new doors for personalized treatments in infertility and related disorders.
Materials and Methods
Sex as a biological variable
Our study exclusively examined female mice because the disease modeled is only relevant in females.
Ethical Statement
The research was approved by the Ethical Committee of the Nanjing Medical University (Approval no. 2019921). All informed consent was obtained based on the Declaration of Helsinki.
Human endometrial sample collection
Human endometrial tissue was collected from IVF/ICSI patients under 40 years old. Patients with Recurrent Implantation Failure (RIF) were identified based on two or more failed embryo transfers using high-quality embryos. Fertility control patients were defined as those who achieved clinical pregnancy in the subsequent cycle after an endometrial biopsy. The study protocol using human specimens was approved by the Institutional Review Committee of Suzhou Hospital affiliated with Nanjing Medical University, and all participants provided informed consent for sample usage.
Generation of Mettl3f/f and Mettl3d/d mice
Mettl3f/f and PgrCre/+ mice were generated from previous study18. Mettl3d/d mice were bred by crossing Mettl3f/f mice with PgrCre/+ mice. Mettl3f/f littermates were used as control mice. Female mice aged 8 to 12 weeks were utilized for the experiments, while male C57BL/6 J mice aged 8 weeks to 6 months were used for mating with the females, marking vaginal plug observation as GD 0.5. Uterine tissue was collected at GD 4, and detailed animal genotype information is provided in Table S1.
Wounding experiments
For injury experiments on pregnant animals, mice were mated as described earlier. At GD 3.5, mice were anesthetized, and a 1 cm surgical incision was made in their abdomen. The uterus was then exposed, and a 26-gauge needle connected to a syringe was used to inject 100 μl (2.7 mg/ml) MAK683 (MedChemExpress, HY-103663) into the uterine horn, 0.5 cm below the ovary. The control group received an equivalent volume of DMSO (Sigma, D2650). After injection, the incision was closed, and the mice received routine post-anesthesia care. Uterine tissue samples were collected at GD 5.5 for verification experiments. This procedure followed strict institutional guidelines and the IACUC Animal Welfare Protocol, ensuring proper anesthesia, analgesia, and post-operative monitoring for animal welfare.
HE staining
Uterine tissues from mice at GD 4 and GD 5.5 were processed as follows: one side was frozen in liquid nitrogen, while the other was fixed in 4% paraformaldehyde overnight at 4°C. The fixed tissue was washed, dehydrated, made transparent with xylene, and embedded in paraffin wax. Sections of 5 μm thickness were then prepared, dewaxed, rehydrated, and stained with hematoxylin and eosin (HE) before examination under an Olympus microscope.
Immunohistochemistry and image analysis
The dissected uteri were fixed in 4% paraformaldehyde/PBS overnight at 4 °C, embedded in paraffin, and sectioned into 5 μm slices. Dewaxing was done conventionally. For immunofluorescence, dewaxed slides were treated with methanol containing 3% H2O2 to block endogenous peroxidase activity. Antigen retrieval was performed using either 1 mg/ml trypsin in PBS at 37°C for 13 minutes or boiling in 0.01 M citric acid (pH 6.0), depending on the primary antibody (Key antibody related information details provided in Table S3). Slides were then incubated with the primary antibody overnight at 4 °C, followed by secondary antibody incubation. DAPI was used for counterstaining, and slides were sealed with an anti-fluorescence quencher. Imaging was done using an Olympus microscope, and Image J software was used for image processing.
Cell line culture and plasmid transfection
ISHIKAWA cells from Pricella Biotechnology were cultured in DMEM medium with serum and antibiotics at 37°C and 5% CO2. Regular mycoplasma testing was done. For experiments, cells were transfected using Lipofectamine 2000 with specific plasmids (Details provided in Table S2). Transfection occurred when cells reached 80% confluence, with collection done 72 hours later.
Immunocytochemistry
Cells were implanted in 96-well plates, CCK-8 solution (Beyotime, C0039) was added to the wells and the absorbance was measured at 450 nm. The In Situ Cell Death Detection Kit TMR red (Roche, 12156792910) Labeled apoptotic cells according was used with manufacturer’s instructions. KI67 (Abcam, ab15580) was used to label the cell proliferative, Nuclei were stained with Hoechst 33258 (Beyotime, C1017).
BeWo cells adhesion experiment
BeWo cells were inoculated in 10 mg/ml poly-HEMA (Sigma, 041M0024U) solution and routinely cultured in cell culture dishes. BeWo cell spheres were filtered through 150 mesh screens to obtain 150–200 μm simulated blastocysts of the same size, with 50 cell spheres per well, and transplanted into treated monolayer Ishikawa cells. Culture for 1-2 h. After washing with PBS twice, the cells with poor adhesion were removed, and fixed with 4% paraformaldehyde at room temperature for 30 min. The adhesion efficiency of simulated embryo was observed and calculated under microscope.
RNA isolation
Total RNA was extracted from uterine tissues and cell lines using RNAiso plus (Takara, 9109). To eliminate DNA contamination, TURBO DNase (Invitrogen, AM2239) was employed. The concentration of total RNA was quantified using the Qubit RNA HS assay kit (Thermo Fisher Scientific, Q32855).
m6A Dot Blot
RNA was transferred onto Hybond N+ membranes and crosslinked twice with UV. The membrane was sealed in 5% skim milk and incubated overnight with anti-m6A antibody (Cell Signaling Technology, 56593S). After washing and incubation with secondary antibodies (ProteinTech, RGAR001), methylene blue staining was used for normalization. The imprints were then analyzed using Image J software.
Western Blotting
Samples were lysed using RIPA buffer (Beyotime, P0013B) with protease inhibitor (Roche, 11873580001). And then mixed with sample buffer before separation on a 12% SDS-PAGE gel. Proteins were transferred to a PVDF membrane, incubated with TBST containing skimmed milk, and probed with primary and secondary antibodies (Key antibody related information details provided in Table S3). The bands described in the diagram as loading controls were true loading controls, and the membranes were washed through the Stripping buffer (Beyotime, P0025B) for 30 min and incubated with TBST containing skimmed milk, then re-incubated for antibody detection, that is, they run on the same gel as the other proteins detected in the experiment. Protein signals were detected using an enhanced chemiluminescence kit and scanned. Experiments were triplicated, and results are presented as fold change ± SD.
Co-immunoprecipitation and protein expression
GD 4 mouse uterine tissue was lysed in buffer containing 1% NP-40, 150 mM NaCl, 20 mM Tris-HCl pH 8, 1 mM DTT, and protease inhibitor for 1 hour. The cleavage products were collected and normalized. A portion of the lysate was saved as Input and stored at -80 °C. The rest was used for immunoprecipitation (IP) with Protein G-Agarose (Roche, 11243233001) and specific antibody (Key antibody related information details provided in Table S3) in IP buffers. After overnight incubation at 4 °C, immunoprecipitated complexes were washed and separated on a gel. Protein signals were detected using an enhanced chemiluminescence kit.
m6A-meRIP-seq
After digesting the ribosomal RNA in total RNA using Thermostable RNase H (Beyotime, R7090M), total RNA was treated with TURBO DNase (Thermo Fisher Scientific, AM2239) for 20 minutes at 37°C to remove DNA contamination, the RNA was precipitated using glycogen (25 μg/ml final) (Thermo Fisher Scientific, AM9510) and isopropanol at -80°C for 2 hours. The precipitated RNA was then washed with 70% ethanol. The final pellet was resuspended in ultrapure H2O. The concentration of total RNA was measured by Qubit RNA HS Assay Kit (Thermo Fisher Scientific, Q32855). The MeRIP sequencing process adapted from a low-input m6A-seq protocol began with fragmenting total RNA into 200 nt at 70 °C for 5-6 min, followed by ethanol precipitation. For m6A input, 10 ng fragmented RNA was used, and the remaining RNA was allocated to m6A-seq. 1 μg anti-m6A antibody (Millipore, ABE572-I) coated 10 μl of protein A magnetic beads (Thermo Fisher Scientific, 10002D) and 10 μl of protein G magnetic beads (Thermo Fisher Scientific, 10004D) underwent IP buffer washes (150 mM NaCl, 10 mM Tris-HCl [pH 7.5], 0.1% IGEPAL CA-630 in nuclease-free H2O) before binding RNA, eluted with N6-methyladenosine. Eluted RNA underwent RNeasy MiniElute spin columns and library preparation with SMARTer Stranded Total RNA-Seq Kit v3-Pico Input Mammalian (TaKaRa, 634488), differing in PCR cycles between input RNA (14 cycles) and m6A RNA (16 cycles). Sequencing was performed on an Illumina NovaSeq with PE 150 bp read length.
ATAC and CUT&Tag sequencing
All steps should be performed on ice or at 4°C. Pre-chill a centrifuge to 4°C. Frozen tissue fragments from mouse endometrial tissue was used in this protocol. Briefly, Prepare a 4 × homogeneous buffer (1.28 M Sucrose, 40 mM Tris-HCl [pH 7.8], 400 nM EDTA, 20 mM CaCl2, 12 mM Mg(Ac)2, 668 μM β-mercaptoethanol) in advance, Transfer frozen tissue sample to a 1 ml micronic tube and add in 600 μl of cold 1× homogeneous buffer (with 1× Protease Inhibitor Cocktail), Add three 2.8-mm ceramic beads to the micronic tube. Vortex micronic tube thrice for 5-10 seconds and keep it on ice. Homogenize tissue sample in a mill homogenizer cooling unit using 2 cycles of the following settings: speed: 1.0, time: 20 sec and speed: 2.1, time: 20 sec, dwell time: 20 sec at both speeds. Following homogenization, filter tissue homogenate through a mini 20 μm pluriStrainer (pluriSelect, NC1423042). Count 50000 nuclei in the filtrate using an Automated Cell Counter such as Countess II or Cellometer K2 Image cytometer according to the manufacturer’s instructions. ATAC-Seq and CUT&Tag sequencing were carried out using the ATAC-Seq Kit (Vazyme, TD711) and the CUT&Tag Assay Kit (Vazyme, TD903), respectively, following the protocols provided by the manufacturer, (Key antibody related information details provided in Table S3). Subsequently, sequencing was conducted on a NovaSeq6000 platform using PE 150 bp read length.
Identify m6A peaks of mRNAs
Raw sequencing reads underwent quality control and adapter trimming using fastp (v0.12.4) and Trim_galore with specific parameters. Trimmomatic (v0.36) further processed reads to remove low-quality bases. Bowtie2 (v2.4.1) aligned cleaned reads to rRNA (hg38), retaining unmapped reads. STAR (v2.7.8) aligned remaining reads to hg19, with output converted to bam format using BEDTools (v2.29.2). MACS2 (v2.2.7) identified putative m6A sites using control input samples. Annotated peak files were generated using GenomicFeatures and ChIPseeker. Gene-level m6A methylation analysis considered peaks with 50% m6A modifications within genes, calculating methylation levels based on peak and transcript lengths. GO enrichment analysis used clusterProfiler and ggplot2, with read coverage of m6A peaks visualized using IGV software (v2.4.15).
Quantification of transcripts
To assess transcript levels, we utilized the TPM (Transcripts Per Million) and FPKM (Fragments Per Kilobase Million) quantification methods. These analyses were conducted using the Salmon tool (version 0.60.0), which facilitated the estimation of mapped reads within the transcriptome. Specifically, we employed the GENCODE (GRCh37) annotation or an augmented GENCODE annotation when referring to transcripts as the basis for our analysis.
CUT&Tag data processing
To ensure data quality, we initially removed low-quality reads using Trim Galore and Trimmomatic. The remaining reads were aligned to the mouse genome (mm10) with Bowtie 2. SEACR (v1.3) was used for peak calling with relaxed parameters, enabling the identification of enriched peaks. Differential analysis of these peaks was conducted using DiffBind in R, and ChIPseeker provided annotation of these differentially enriched peaks. Heatmaps generated with deepTools (v3.5.0) visually represented the results, facilitating data interpretation by revealing patterns and clustering within the analyzed peaks.
ATAC-seq data processing
Low-quality reads were removed using Trim Galore and Trimmomatic. The remaining reads were aligned to the mouse genome (mm10) using Bowtie 2. Duplicate reads were then eliminated with Sambamba (v1.0.1). Subsequent filtration with SAMtools (v1.6) removed mitochondrial reads and any remaining low-quality data. Peak calling was performed using SEACR (v1.3) with relaxed parameters, and read counts were normalized by TPM for further analysis. Differential analysis was conducted using DiffBind in R to identify enriched peaks. These peaks were annotated using ChIPseeker to provide insights into their genomic locations and functional relevance.
Statistical analysis and Reagent information
The details of important reagent consumables in this study were provided in Table S4. The sequencing data for mouse and human samples were shown in Supplementary Data 1. Each experiment was triplicated for robustness, and data are expressed as mean ± SD. Statistical analysis was conducted using GraphPad Prism software, including the two-group t-test and the three-group one-way ANOVA. The significant difference is indicated by a p value < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request. The source data behind the graphs in the paper can be found in Supplementary Data 1. Numerical source data for all graphs in the manuscript can be found in Supplementary Data 1 file. The sequencing reads for mouse and human sample information that supports the findings of this study have been deposited, the accession codes are PRJCA035031 and PRJCA035196.
References
Lessey, B. A. & Young, S. L. What exactly is endometrial receptivity? Fertil. Steril. 111, 611–617 (2019).
Franasiak, J. M. et al. A review of the pathophysiology of recurrent implantation failure. Fertil. Steril. 116, 1436–1448 (2021).
Valdes, C. T., Schutt, A. & Simon, C. Implantation failure of endometrial origin: it is not pathology, but our failure to synchronize the developing embryo with a receptive endometrium. Fertil. Steril. 108, 15–18 (2017).
Wang, W. et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle. Nat. Med. 26, 1644–1653 (2020).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genet. 53, 1698–1711 (2021).
Kim, T. H. et al. Loss of HDAC3 results in nonreceptive endometrium and female infertility. Sci. Transl. Med. 11, (2019).
Millan-Zambrano, G., Burton, A., Bannister, A. J. & Schneider, R. Histone post-translational modifications - cause and consequence of genome function. Nat. Rev. Genet. 23, 563–580 (2022).
Husmann, D. & Gozani, O. Histone lysine methyltransferases in biology and disease. Nat. Struct. Mol. Biol. 26, 880–889 (2019).
Blackledge, N. P. & Klose, R. J. The molecular principles of gene regulation by Polycomb repressive complexes. Nat. Rev. Mol. Cell Biol. 22, 815–833 (2021).
Piunti, A. & Shilatifard, A. The roles of Polycomb repressive complexes in mammalian development and cancer. Nat. Rev. Mol. Cell Biol. 22, 326–345 (2021).
Nancy, P. et al. H3K27me3 dynamics dictate evolving uterine states in pregnancy and parturition. J. Clin. Invest. 128, 233–247 (2018).
Fukui, Y. et al. The EZH2-PRC2-H3K27me3 axis governs the endometrial cell cycle and differentiation for blastocyst invasion. Cell Death Dis. 14, 320 (2023).
Osokine, I. et al. Gene silencing by EZH2 suppresses TGF-beta activity within the decidua to avert pregnancy-adverse wound healing at the maternal-fetal interface. Cell Rep. 38, 110329 (2022).
Murakami, S. & Jaffrey, S. R. Hidden codes in mRNA: Control of gene expression by m(6)A. Mol. Cell. 82, 2236–2251 (2022).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Wang, X. et al. Structural basis of N6-adenosine methylation by the METTL3–METTL14 complex. Nature (London). 534, 575–578 (2016).
Wang, P., Doxtader, K. A. & Nam, Y. Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol. Cell. 63, 306–317 (2016).
Zheng, Z. et al. METTL3 is essential for normal progesterone signaling during embryo implantation via m6A-mediated translation control of progesterone receptor. Proc. Natl. Acad. Sci. USA120, 1–2080283176 (2023).
Collignon, E. et al. m(6)A RNA methylation orchestrates transcriptional dormancy during paused pluripotency. Nat. Cell Biol. 25, 1279–1289 (2023).
Lu, J. et al. NSUN2-Mediated m(5)C Methylation Impairs Endometrial Receptivity. Lab. Invest. 104, 100327 (2024).
Wan, S. et al. METTL3-dependent m(6)A methylation facilitates uterine receptivity and female fertility via balancing estrogen and progesterone signaling. Cell Death Dis. 14, 349 (2023).
Li, Y. et al. N (6)-Methyladenosine co-transcriptionally directs the demethylation of histone H3K9me2. Nat. Genet. 52, 870–877 (2020).
Chen, C. et al. Nuclear m(6)A reader YTHDC1 regulates the scaffold function of LINE1 RNA in mouse ESCs and early embryos. Protein Cell. 12, 455–474 (2021).
Tamura, I. et al. Genome-wide analysis of histone modifications in human endometrial stromal cells. Mol. Endocrinol. 28, 1656–1669 (2014).
Tamura, I. et al. The distal upstream region of insulin-like growth factor-binding protein-1 enhances its expression in endometrial stromal cells during decidualization. J. Biol. Chem. 293, 5270–5280 (2018).
Liu, J. et al. N (6)-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367, 580–586 (2020).
Lavarone, E., Barbieri, C. M. & Pasini, D. Dissecting the role of H3K27 acetylation and methylation in PRC2 mediated control of cellular identity. Nat. Commun. 10, 1679 (2019).
Patil, D. P. et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Xu, C. et al. Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat. Chem. Biol. 10, 927–929 (2014).
Liu, J. et al. The RNA m(6)A reader YTHDC1 silences retrotransposons and guards ES cell identity. Nature 591, 322–326 (2021).
Vrljicak, P. et al. Dynamic chromatin remodeling in cycling human endometrium at single-cell level. Cell Rep. 42, 113525 (2023).
Lynch, V. J. et al. Ancient transposable elements transformed the uterine regulatory landscape and transcriptome during the evolution of mammalian pregnancy. Cell Rep. 10, 551–561 (2015).
Wilson, M. R. et al. ARID1A Mutations Promote P300-Dependent Endometrial Invasion through Super-Enhancer Hyperacetylation. Cell Rep. 33, 108366 (2020).
Xin, Q., Feng, I., Yu, G.& Dean, J. Stromal Pbrm1 mediates chromatin remodeling necessary for embryo implantation in the mouse uterus. J. Clin. Invest. 134, (2024).
Chelmicki, T. et al. m(6)A RNA methylation regulates the fate of endogenous retroviruses. Nature 591, 312–316 (2021).
Wei, J. et al. FTO mediates LINE1 m(6)A demethylation and chromatin regulation in mESCs and mouse development. Science 376, 968–973 (2022).
Acknowledgements
This research was funded by the National Natural Science Foundation of China (92168104, 82071720, 32370913), the National Key Project of Research and Development Program (2022YFC2702901, 2021YFC2700705), Suzhou key clinical diseases funding (LCZX202109), Suzhou Talent Training Program (GSWS2022117, GSWS2020057, GSWS2023051), Jiangsu Provincial Medical Key Discipline (Laboratory) Cultivation Unit (JSDW202214).
Author information
Authors and Affiliations
Contributions
B.H. and Q.M. conceptualized the study, X.H. and J.J. performed most of the experiments and analyzed the data, J.L. and J.-F.L. performed the analysis for CUT & Tag-seq, ATAC-seq and m6A-RIP-seq, X.H. and W.X. cultured cell line, X.H. and H.S constructed the plasmids, M.Z. prepared mouse samples, C.D. prepared the figures, B.H., J-L.L. and Y.L. wrote the paper, B.H. and H.L. edited the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, X., Li, J., Ding, C. et al. Mettl3/Eed/Ythdc1 regulatory axis controls endometrial receptivity and function. Commun Biol 8, 215 (2025). https://doi.org/10.1038/s42003-025-07667-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07667-y