Abstract
The complex spatiotemporal dynamics of neurons encompass a wealth of information relevant to perception and decision-making, making the decoding of neural activity a central focus in neuroscience research. Traditional machine learning or deep learning-based neural information modeling approaches have achieved significant results in decoding. Nevertheless, such methodologies require the vectorization of data, a process that disrupts the intrinsic relationships inherent in high-dimensional spaces, consequently impeding their capability to effectively process information in high-order tensor domains. In this paper, we introduce a novel decoding approach, namely the Least Squares Sport Tensor Machine (LS-STM), which is based on tensor space and represents a tensorized improvement over traditional vector learning frameworks. In extensive evaluations using human and mouse data, our results demonstrate that LS-STM exhibits superior performance in neural signal decoding tasks compared to traditional vectorization-based decoding methods. Furthermore, LS-STM demonstrates better performance in decoding neural signals with limited samples and the tensor weights of the LS-STM decoder enable the retrospective identification of key neurons during the neural encoding process. This study introduces a novel tensor computing approach and perspective for decoding high-dimensional neural information in the field.
Similar content being viewed by others
Introduction
Neural information acquisition allows for the detection of spatiotemporal activity of neurons in biological processes such as perception and decision-making1,2. Organisms perceive a vast amount of external information daily and make decisions based on that information, during which the neurons in the brain encode complex information. There are various established methods for collecting this neural information, including electrical, optical, and chemical approaches, which enable the acquisition of neural data at different temporal and spatial scales3,4,5,6,7. Specifically, invasive microelectrodes are used to record neuronal cell activity, providing high temporal resolution3,8. Non-invasive EEG (electroencephalography) also offers high temporal resolution but lacks cellular-level precision6,9,10. fMRI (functional magnetic resonance imaging) is a commonly used non-invasive method, indirectly measuring neural activity by detecting changes in blood oxygenation, and it covers the largest acquisition area among all methods3,11,12. Calcium imaging, utilizing electron microscopy, records the position and fluorescence changes of neurons, allowing for the capture of activity information from specific types of neurons13,14,15. Different neural information acquisition methods yield data with distinct data structures, signal types, and spatial and temporal resolutions, which have implications for designing decoders to decode neural information. With the advancement of tools and techniques, there is an increasing availability of high-order tensor data in the field of neural information decoding. Neural information is often characterized by high dimensionality and limited sample sizes, which presents challenges in effectively extracting meaningful insights and decoding the neural information. The utilization of high-order tensor data provides an opportunity to capture and analyze complex relationships and interactions across multiple dimensions, potentially enhancing the decoding capabilities and facilitating a deeper understanding of neural processes.
With the advancement of computer science, numerous algorithms have been applied to extract information from complex neural data. For instance, Yargholi et al. successfully classified fMRI signals generated during handwritten digit recognition using an augmented naive Bayesian classifier16. Ali R. Rezai et al. employed support vector machines to decode cortical EEG signals associated with brain motor intention1. De Luca et al. utilized convolutional neural networks to classify calcium imaging signals in visual stimulus tasks17. Neural signals encompass multiple dimensions such as time, space, and frequency, which provide valuable structural information. In practice, neural data often exhibits tensor patterns, while these decoders transform tensorized data into vectorized data for computation. However, the current approach of vectorization in decoding neural signals can lead to the loss of this inherent structural information, potentially compromising its effectiveness in more complex decoding tasks. Therefore, there is a need to explore methods that utilize tensors to preserve and utilize the multidimensional nature of neural signals.
Neural signals often contain tensorized data, with fMRI data being a typical example of tensor data, represented as a multidimensional matrix3,8. Unfortunately, tensor data in neural signals is often characterized by small sample sizes and high dimensions. For instance, Stereoelectroencephalography (sEEG) is a type of intracranial electroencephalography (iEEG), which is an important technique used in the diagnosis of epilepsy and the field of neural decoding. SEEG data exhibits a rapid increase in dimensions as the number of channels and sampling rate increase6,9,10. While deep learning techniques have shown promising performance in handling high-dimensional data, they may lack robustness when faced with small sample sizes. In the field of machine learning, an increasing number of researchers have proposed computational models for tensorized data to address the challenges of small-sample high-dimensional data. These models not only tackle the issue but also offer good interpretability, making them valuable in various applications18,19,20,21.
In the field of tensor-based models, researchers have proposed robust approaches for various tasks. For instance, Shengli Xie et al. introduced a more robust tensor PCA method based on Tensor Train(TT) and Tucker tensor decompositions22. Selin Aviyente et al. enhanced the efficiency of tensor networks by introducing a multi-branch structure with lower time and space complexity23.Xindong Wu et al. improved support vector machines by proposing Support Tensor Machines24, while Ngai Wong et al. combined Tensor Train decomposition with SVM to develop STTM (Tensorized SVM)25. Cheng Dai et al. further enhanced STTM by introducing weighted relaxation factors26. By transforming the vector computation framework of machine learning algorithms into a tensor computation framework, superior performance has been achieved. This is primarily due to the fact that tensor algorithms can leverage the structural information inherent in tensor data, which is particularly effective in handling image and temporal data27,28,29. Moreover, the rapid increase in dimensionality after vectorization can lead to overfitting, especially for small-sample data. In contrast, tensor-based models exhibit stronger stability, requiring fewer model parameters. Methods like sliceTCA30 further highlight the advantages of tensor approaches, capturing task-relevant neural data with fewer components compared to traditional methods. This not only helps mitigate overfitting but also saves computational and storage resources18,31,32,33.
Tensor models have advantages in computing high-dimensional, small-sample neural data. We have extended the Support Tensor Machine (STM) to develop a new model called the LS-STM. LS-STM utilizes least squares optimization34 and can obtain tensor decoding models using various tensor decomposition methods. We tested LS-STM on neural data from humans and mice and found that its decoding performance surpasses that of traditional machine learning decoders across different species. Additionally, we observed that the least squares optimized support tensor machine decoder exhibits better decoding performance, higher accuracy, and lower time and space complexity across varying data sizes. The LS-STM decoder also allows traceability and selection of key neurons based on tensor weights. We conducted tests to demonstrate the effectiveness of neuron selection through tensor weighting. LS-STM represents the first traceable and scalable decoder based on tensor computing, offering a novel approach to neural data analysis. The term “traceable" refers to LS-STM’s capability to identify and track the key neural channels that significantly influence decoding accuracy. The term “scalable" indicates that LS-STM can be expanded and enhanced through modifications in tensor decomposition methods and kernel functions, allowing for potential improvements and adaptability to various tasks.
Main
We employed LS-STM as a neural information decoder, enabling the decoding of high-dimensional, small-sample neural data and unveiling crucial neural information channels.
Workflow
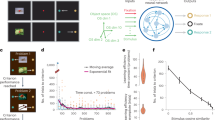
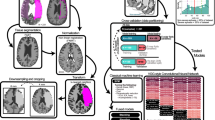
We utilized LS-STM as a neural information decoding framework. This enabled us to decode high-dimensional, small-sample neural data and facilitate the traceability of critical neural information channels. The neural traceability using LS-STM involves two steps. Firstly, a tensor construction is performed considering the temporal and spatial characteristics of the neural data. During tensor construction, prior information such as spatial positions, temporal sequences, and frequency densities of cell responses can be incorporated, which significantly impacts the decoding performance and traceability results. Secondly, LS-STM, a tensor-based neural information decoding model, is applied to trace the influential neural channels. Prior to these two steps, a standard set of criteria is used to filter the neural data, removing redundant and ineffective neural channels based on specific selection criteria and the requirements of the decoding task. The overall process is illustrated in Fig. 1, which outlines the two crucial steps, while detailed methodology can be found in the Method section. We evaluated the decoding and traceability capabilities of LS-STM using high-throughput neural data from Allenbrain Neuropixel recordings in mice (N = 30) and human iEEG recordings (N = 51).
A Overview of visual encoding in mice using Neuropixel recording. Mice were stimulated with movies and light spots, and neuronal activity was recorded from layers L1-L6 of the primary visual cortex using Neuropixel probes. (N = 30) (B) iEEG recordings of visual encoding in humans. Movies and speech stimuli were used, and different recording methods such as ECoG, HD ECoG, and SEEG were employed across different individuals. The physiological recordings encompassed a range of coverage and types, including non-mutually exclusive data (e.g., the same subject may have recordings from both the left and right hemispheres). (N = 51) (C) Overview of the decoding process using the LS-STM model.
The first step involved constructing tensors from high-throughput neural electrical data to facilitate LS-STM decoding. While LS-STM has the potential to utilize up to seven modalities (time, frequency, space, trial, condition, subject, and grouping), in this study, we focused on three key features: time, channel (space), and frequency. Specifically, for neural electrical data acquired from iEEG and Neuropixel recordings, we transformed the time series data into the time-frequency domain, constructing a three-way tensor of time, channel, and frequency (see details in the Methods section under Data Preprocessing). This approach preserved the spatial relationships between channels and the temporal order of the time dimension. The tensor construction used in this study can be extended to incorporate the other four features in future applications, but for the purposes of this research, we prioritized the three features most relevant to our analysis. In the second step, the constructed tensor was used to perform LS-STM model computations as outlined in Fig. 2. Further computational details are provided in the Methods section.
A Tensor construction: High-dimensional neural signals collected are transformed into a tensor data structure. B Tensor decomposition, using Tucker decomposition as an example. C Tensor inner product: The tensor inner product is estimated by approximating the core tensor and factor matrices obtained from tensor decomposition, which is then incorporated into a kernel functionϕ. Here, Xi represents the ith tensor, \({A}_{1r1}^{i}\) denotes the ith row of factor matrix \({A}_{1}^{i}\). D Tensor mapping: The mapping of an inseparable tensor space to a linearly separable tensor space. E Least Squares Support Tensor Machine: A model that utilizes least squares regression for tensor classification. Here, yi represents the label (y=1 or −1) of the ith tensor Xi, and ei denotes the tensor error. Further details can be found in the Methods.
LS-STM: superior decoding performance in mice and human brain datasets compared to ML decoders
We performed comprehensive evaluations of LS-STM on two decoding tasks, and the results are depicted in Fig. 3 and Supplementary 1–2 shown detail. To illustrate the efficacy of tensor construction in the decoding process, we compared the performance of LS-STM with tensor input against STM and other machine learning decoders that utilized vector input. The evaluation was based on accuracy. Our findings indicated that LS-STM with tensor input consistently outperformed the other models, achieving significant accuracy improvements, with 70% and 82.4% of cases showing the highest accuracy in the mice and human brain datasets, respectively. In the following sections, we provide a detailed analysis of these results, highlighting the decoding improvements gained from using LS-STM with tensor input, supported by statistical evidence.
A Decoding performance of different decoders in the mouse brain (N = 30), where each scatter represents the decoding performance of the model in different mouse brains. Compared with other models, the average accuracy of LS-STM is beyond models and passes the test of significance, the Wilcoxon signed-rank test. Error bars represent mean with SD. B Difference in decoding performance between LS-STM and STM in different sessions. (pass the Wilcoxon signed-rand test, < 0.0001). C Difference in decoding performance between LS-STM and RF in different sessions. (pass the Wilcoxon signed-rand test, < 0.001). D Number of occurrences where each model performed optimally in decoding tasks across different mouse brains. For example, LS-STM exhibited the best decoding performance in 21 out of 30 mouse brains. E−H Test results in the human brain.
When comparing the decoding performance of tensor-based and vector-based decoders, we observed that the tensor-based models consistently outperformed their vector-based counterparts. In the mouse dataset, the tensor decoder achieved the highest accuracy in 21 out of 30 cases (N = 30). Similarly, in the human brain dataset, the tensor decoder exhibited superior performance in all 51 cases (N = 51), significantly outperforming the vector decoder, as shown in Fig. 3 (Wilcoxon signed-rank test, p < 0.0001). Across both mouse and human neural data, the tensor-based decoders consistently outperformed the vector-based decoders. The findings are theoretically sound conceptually. The process of constructing the tensor incorporates a greater amount of prior information. On the other hand, the input format of the tensor decoder itself preserves more of the original structural information when compared to the vector-based decoder.
Overall, LS-STM demonstrated superior decoding performance across both datasets, as detailed in Supplementary Tables 1 and 2. In the mouse dataset, LS-STM achieved the highest performance in 70% of the cases (N = 30), as shown in Fig. 3D. While tree models demonstrated relatively good performance with an average accuracy of 89%, their stability was weaker compared to LS-STM. LS-STM, in contrast, exhibited superior stability and performance, surpassing STM and RF significantly, as evidenced by the results of the Wilcoxon test, as shown in Fig. 3B and C. In the human brain dataset, although tree models did not exhibit outstanding performance as in the mouse dataset, LS-STM still maintained stability, achieving the highest performance in 82.4% of the cases (N = 51), as shown in Fig. 3H. Machine learning models, on the other hand, nearly lost their decoding capability, with an average accuracy of around 50%, as shown in Fig. 3E. Nevertheless, the decoding capability of the tensor decoder remained intact, with LS-STM demonstrating a performance improvement over the alternative method, as substantiated by the results of the Wilcoxon test, as shown in Fig. 3A, E. The results from the two datasets collectively demonstrate that LS-STM outperforms traditional machine learning decoders, offering superior and robust decoding performance.
Determining channel decisive for decoding
The overall understanding of the LS-STM structure is provided through the description in Fig. 2, with detailed explanations available in the Methods section. Notably, the weights in LS-STM are in tensor form, where wijk represents the tensor weight corresponding to the strength of the jth time and kth frequency in the ith channel. In contrast to vector weights obtained from vector computations, tensor weights contain more structural information within each channel. Furthermore, during the LS-STM computation process, tensor decomposition enables the extraction of effective information from each of the three dimensions, thereby preserving valuable structural information. During LS-STM decoding, calculations are performed for each time and frequency within a channel, which means that the tensor weights corresponding to a channel are not unique. To determine the key channels, it is necessary to calculate and sort the tensor weights within the channel. We provide a method for computing the weights of key channels, which is detailed in the Methods section.
The magnitudes of the weight tensors impact the decoding results. Larger weights indicate a greater influence of the information within that channel in the decoding process, reflecting the crucial role of the neuronal information recorded in that channel during brain encoding. Conversely, the weight tensors also contain numerous small weights, indicating that these channels do not play a significant role in decoding.
We calculated decoding channels across sessions using the data. Each session’s data was divided into five segments, comprising both a training set and a test set, ensuring that the data in each split was distinct from the others. This was done to address the issue of tensor weights used in identifying key channels. The tensor weights were obtained by training on the training set. Using the same set would result in identical weights and fail to demonstrate channel selection stability. By employing various sessions, we have observed that the utilization of a determining channel can effectively enhance the decoding performance, as depicted in Fig. 4.
A Decoding performance comparison using selected channels versus randomly chosen channels, with different channel proportions (25%, 50%, 75%, 100%) of sorted channels based on importance (as described in the Method section under channel selection) in the AllenBrain dataset. The percentages represent the proportion of the top-ranked channels used for decoding. The Wilcoxon test shows significant differences at 25% (p < 0.01) and 50% (p < 0.001) of the selected channels. Error bars represent mean with SD. B Decoding performance comparison between selected and random channels across different proportions of sorted channels. C Statistical comparison of the two channel selection methods, showing the number of better performances across sessions when using selected channels. D–F Test results in the human brain dataset, following the same methodology.
We developed a computational approach to evaluate the contributions of individual channels to the decoding process, as explained in the Methods section. We used this algorithm to determine how each channel contributes to the decoding process. Considering the distinct contributions of channels, we sorted them based on their respective contributions and subsequently employed different ratios to evaluate the decoding performance. To evaluate the impact of key channels on decoding performance, we exclusively selected key channels for decoding and excluded non-key channels. We performed decoding tests using LS-STM and randomly selected an equal number of channels from non-key channels for comparison. Fig. 4 provides a comprehensive illustration of the results.
When selecting channels at different ratios, a notable trend emerges, with significant improvements observed at the 50% and 75% ratios in both datasets, as shown in Fig. 4A and D. These improvements, confirmed by the Wilcoxon test, suggest that the selected channels contain more informative data compared to non-key channels, making them more effective for the decoding process. This highlights LS-STM’s traceability in identifying key channels that significantly influence decoding accuracy, as shown in Supplementary 3–5. A more detailed analysis, depicted in Fig. 4B and E, reveals that improvements were observed in the majority of sessions and were statistically significant according to the Wilcoxon test. Despite some variations in decoding performance across the 50% to 80% range, the model consistently demonstrates its ability to trace and leverage the most impactful neural channels. The session count analysis in Fig. 4C and F shows a decreasing trend from 25% to 75%, reaching a more stable state due to the increased utilization of key channels. In summary, the traceability and selection of channels based on their contribution proves to be a viable approach, ensuring stable and improved decoding performance across both datasets. We present the contribution of specific tensor channels in Supplementary Figs. 3, 4, and 5.
Increasing number of data improves decoding
In Fig. 3, we compared two tensor decoders, LS-STM and STM. LS-STM outperformed STM in decoding both mouse and human brain data, demonstrating superior robustness across a large number of samples. We aimed to examine if tensor decoders maintain consistent decoding performance with varying data volumes. We selected different percentages (20%, 40%, 60%, 80%, 100%) of the dataset, ensuring the labels were balanced. In both datasets, we observed that the performance of both tensor decoders improved as the data volume increased, as shown in Supplementary Fig. 1. This growth can be attributed to the increased encoding of information brought by larger data volumes, thereby enhancing the robustness of the decoders. Even with small sample sizes, tensor decoders displayed stable performance. This highlights their ability to decode high-dimensional data with limited samples. Furthermore, we found that LS-STM consistently outperformed STM in decoding across various data volumes.
From a theoretical standpoint, the LS-STM model represents a more streamlined variant of the traditional STM, answering the question of how much simplification can be achieved without sacrificing any advantages of the STM model. By inheriting the strengths of STM, the LS-STM model modifies the insensitive loss function in STM to include a second-norm error term and replaces the inequality constraints with equality tensor constraints. This transformation allows the quadratic programming problem of solving STM to be directly converted into a system of linear equations with a tensor structure, thereby obtaining optimal solutions. As a result, LS-STM exhibits enhanced robustness to outliers in nonlinear decoding problems and demonstrates strong resistance to the interference of significant noise in neural data.
LS-STM is an extensible tensor decoder that allows for the expansion of tensor decomposition, tensor computations, and tensor construction processes during the decoding process. We specifically expanded the tensor decomposition aspect to illustrate the potential development of tensor decoders in the context of biological decoding. We utilized Tucker decomposition, TensorTrain decomposition, and CP decomposition in LS-STM and evaluated their performance on previous datasets. The results from both sets of data, as shown in Supplementary Fig. 2, demonstrate the superior and robust performance of LS-STM across different decomposition methods, far exceeding the decoding performance of SVM.
Discussion
We propose the LS-STM model, an expandable tensor computational decoder that enables the traceability of key neurons. Our main innovation lies in utilizing tensor computation models for decoding and providing a neuron/channel selection method based on tensor decoders. In neuroscience research, most decoding tasks rely on vector computation models, which result in the loss of tensor structural information in biological data. By employing tensor computation models, we not only preserve this structural information but also leverage the spatial relationships inherent in the tensor structure. As demonstrated in Fig. 4, the model’s traceability is evident in the trend where decoding accuracy increases up to 50% of selected channels in the human brain dataset and 75% in the mice dataset before declining, indicating the identification of key channels that influence decoding accuracy.
We propose the LS-STM model, an expandable tensor computational decoder that enables the traceability of key neurons. Our main innovation lies in utilizing tensor computation models for decoding and providing a neuron/channel selection method based on tensor decoders. In neuroscience research, most decoding tasks rely on vector computation models, which result in the loss of tensor structural information in biological data. By employing tensor computation models, we not only preserve this structural information but also leverage the spatial relationships inherent in the tensor structure. As demonstrated in Fig. 4, the model’s traceability is evident from the trend of decoding accuracy with varying percentages of selected channels. Initially, the accuracy increases as more channels are selected, reaching a peak before declining. Specifically, in the human brain dataset, this trend reverses after 50% of channels, while in the mice brain dataset, the turning point occurs after 75%. This trend underscores LS-STM’s capability to trace and identify the critical channels that most significantly influence decoding performance, highlighting the model’s utility in preserving and exploiting biological data structure.
The computational process of LS-STM involves two main steps: tensor construction and tensor decomposition. Firstly, tensor construction enables the incorporation of additional prior information, such as temporal relationships, spatial positions, frequency features, and more. Secondly, tensor decomposition can effectively extract meaningful information embedded within the tensors. Tucker decomposition, in particular, captures interactions between dimensions and reveals multidimensional structures. Neural information is often characterized by the firing of neurons located in different spatial positions with specific temporal sequences. Therefore, accounting for the interactions between different dimensions is particularly meaningful for neural data as it captures the interplay between temporal and spatial dimensions.
During practical testing, LS-STM exhibited stable decoding performance across neural signal datasets from different species. LS-STM achieved a decoding accuracy of 82.4% on 51 human iEEG recordings. It is important to note that almost all vector decoders produced an insignificant decoding performance for this task, with an accuracy rate of around 50%. LS-STM achieved an average decoding accuracy of 90% on a dataset of 30 mouse Neuropixel recordings. LS-STM demonstrated superior decoding stability compared to other decoders and significantly outperformed vector decoders with an accuracy of around 75%.
LS-STM, a tensorized decoder, offers two advantages over vector input decoders due to its specialized data structure. Vectorizing tensors for vector decoders often leads to overfitting, especially with small datasets and large feature numbers, as seen in Fig. 3, where vector models significantly worse than tensor models. In contrast, tensor decoders LS-STM avoid this issue by directly performing tensor calculations, resulting in higher accuracy. As shown in Supplementary Fig. 1, LS-STM maintains stable accuracy even when trained on only 20% of the data, confirming its robustness to overfitting. Additionally, LS-STM improves interpretability by calculating tensor weights across different dimensions, which quantify the contribution of each feature and guide key neuron selection. This interpretability is demonstrated in Fig. 4, where decoding performance varies depending on the selected channels. The variations in performance highlight how channels identified by tensor weights contribute more effectively to decoding, showcasing LS-STM’s ability to prioritize and select the most informative neural features for the task.
There are still unanswered questions regarding the use of various data collection tools such as fMRI, calcium imaging, EEG, etc., and how to construct tensors. In different decoding tasks, introducing more prior information, such as the spatial location and functional information of brain regions, could enhance the tensor construction process. Additionally, further research is required on tensor decomposition, specifically how different decomposition methods can be applied and analyzed across various dimensions. Fortunately, with the development of tensor computing, more and more related tensor models have been developed, which will aid in the future of neural signal decoding. While this paper primarily focuses on the tensor model decoder, it is worth noting that LS-STM can be extended to multiclass classification tasks using machine learning techniques similar to SVM, such as one-vs-one or one-vs-all strategies, showcasing its versatility beyond binary classification.
Method
Data introduction
Human Brain Data: Open multimodal iEEG-fMRI dataset from naturalistic stimulation with a short audiovisual film. The human brain data used in this study were obtained from Mariana P. Branco et al. who collected iEEG data from 51 participants aged 5–55 years. Among them, 46 patients were implanted with intracranial ECoG electrode grids with an exposed diameter of 2.3 mm and an electrode spacing of 10 mm, with the number of contact points ranging from 48 to 128. Additionally, 6 patients were implanted with high-density (HD) ECoG electrode arrays, with an exposed diameter of 1.3 mm, electrode spacing of 3–4 mm, and 32, 64, or 128 contact points. Furthermore, 16 patients were implanted with SEEG electrodes, with the number of contact points ranging from 4 to 173. The electrode coverage in most patients included peri-Sylvian areas, with electrodes predominantly located in the frontal and motor cortices. The stimulation for the participants was derived from film clips of the 1970 movie “Pippi on the Run," resulting in a 6.5 min short film that maintained a coherent narrative and limited the duration of the task. As a clinical task used for language localization, the movie consisted of 13 alternating blocks of speech and music, each lasting 30 s (with seven blocks of music and six blocks of speech).
Mouse Brain Data: Allenbrain Visual Coding - Neuropixels. The mouse brain data used were obtained from Allenbrain and comprised recordings from 58 mice during visual encoding experiments. Four types of mice were used: wild type, Sst-IRES-Cre; Ai32 (ChR2), Pvalb-IRES-Cre; Ai32 (ChR2), and Vip-IRES-Cre; Ai32 (ChR2). Each mouse had six neuropixel recordings of electrical signals from the cortex, hippocampus, and thalamus. Standardized visual stimuli were presented to each mouse to map visual responses across multiple brain regions simultaneously. The stimulus set was designed to characterize the tuning properties of individual neurons. It included classical visual stimuli such as Gabors, full-field drifting gratings, and moving dots, as well as a set of natural images and movies.
Data preprocessing
Data preprocessing consists of two main parts: raw data processing and tensor construction. In this study, two datasets were used, collected from human brains and mice, with different data collection methods and tasks, requiring separate processing approaches. For the human brain data, intracranial EEG (iEEG) was recorded at a sampling rate of 512 or 2048 Hz, and 1-second segments were selected as individual data points. This resulted in 180 data points labeled as speech and 210 labeled as music. To balance the data, we randomly selected 180 data points from the music category for the decoding task. For the mouse brain data, Neuropixel recordings were collected from multiple brain regions in response to visual stimuli. Channels from the VISP region were specifically selected for the visual decoding task, and 600 seconds of data were randomly sampled for each stimulus category: natural images and Gabors. Our decoding tasks involved using LS-STM to classify whether the human brain was processing music or speech, and to determine whether mice were viewing Gabors or natural images.
The construction of tensors involved three main steps: (1) filtering the raw data in each channel within the 0–40 Hz frequency range, (2) performing Fourier transforms on the filtered data to obtain time-frequency matrices for each channel, and (3) combining all the time-frequency matrices from different channels, aligning them in terms of time and frequency. This process resulted in a 3-way tensor suitable for tensor-based computations.
Least square support tensor machine
Firstly, the training tensor dataset is considered as follows:
where for any \(i\in \{1,2,\ldots ,l\},{{{\bf{X}}}}_{i}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\) is the third order tensor input, \({y}_{i}\in {\mathbb{Y}}=\{1,-1\}\) is the binary output indicator.
Then, a least squares support tensor machine (LS-STM) optimization model is designed for the binary recognition problem in tensor space \({{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\), i.e.,
where \({{\bf{W}}}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\) is the unknown normal tensor (i.e., a third order tensor weight), \(b\in {\mathbb{R}}\) is the unknown bias variable, ei(i = 1, 2, …, l) are unknown error variables, c > 0 is a regularization parameter, and \(\parallel \cdot {\parallel }_{{\mathfrak{F}}}\) is the Ferbennius norm.
In order to solve the unknown W, b and \({e}_{i}{| }_{i = 1}^{l}\) in (2), the Lagrangian function of model (2) is given as follows:
where for any i = 1, 2, …, l, γi is the Lagrangian multiplier.
Next, let the partial derivatives of \({L}^{{\prime} }({{\bf{W}}},b,{e}_{i}{| }_{i = 1}^{l},{\gamma }_{i}{| }_{i = 1}^{l})\) (3) with respect to \({{\bf{W}}},b,{e}_{i}{| }_{i = 1}^{l}\) and \({\gamma }_{i}{| }_{i = 1}^{l}\) be 0, respectively. That is, we have the following Karush-Kuhn-Tucker condition:
Therefore, according to condition (4) and the tensor decomposition method, model (2) is transformed into the LS-STM algebraic equation with tensor decompositions, i.e.,
where for any i = 1, 2, ⋯ , l and j = 1, 2, ⋯ , l, the inner product of tensors Xi and Xj under tensor-Tucker decomposition is derived as follows:
where \({\widehat{{{\bf{X}}}}}_{i}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\) is the approximate tensor of the tensor \({{{\bf{X}}}}_{i}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\), \({{{\bf{G}}}}^{i}\in {{\mathbb{R}}}^{{n}_{1}\times {n}_{2}\times {n}_{3}}\) is the core tensor of the tensor \({{{\bf{X}}}}_{i}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}},{{{\bf{A}}}}_{p}^{i}\in {{\mathbb{R}}}^{{I}_{p}\times {n}_{p}}(p=1,2,3)\) are factor matrices of the tensor \({{{\bf{X}}}}_{i}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\). For any k1 = 1, 2, …, n1, k2 = 1, 2, …, n2, k3 = 1, 2, …, n3, \({{{\bf{G}}}}_{{k}_{1}{k}_{2}{k}_{3}}^{i}\) is the \({({k}_{1}{k}_{2}{k}_{3})}^{th}\) element of the third-order core tensor Gi. For any p = 1, 2, 3 and \({k}_{p}=1,2,\ldots ,{n}_{p},{{{\bf{A}}}}_{p,{k}_{p}}^{i}\) is the \({({k}_{p})}^{th}\) column vector of the matrix \({{{\bf{A}}}}_{p}^{i}\), the symbol ” × k” is the k-mode product of a tensor and a matrix, and some approximate expressions of the tensor Xj are similar to those of Xi.
In addition, \({\gamma }_{i}{| }_{i = 1}^{l}\) and b can be obtained by solving algebraic equation (5), and the optimal tensor weight W is derived by Karush-Kuhn-Tucker condition (4), i.e.,
where the symbol “∘” represents the outer product of vectors.
Finally, according to (7) and b, a real-valued decision LS-STM recognizer with Tucker decomposition for the new test tensor \({{\bf{X}}}\in {{\mathbb{R}}}^{{I}_{1}\times {I}_{2}\times {I}_{3}}\) is designed as follows:
Select key channels
Channel selection was based on the tensor weights Wijk computed by LS-STM, which reflect the importance of neural channels in terms of frequency and time during encoding. We calculated the contribution level Ci of each channel by analyzing their respective weights. This allowed us to assess the significance of each channel in the encoding process.
Key channels were selected based on their contribution levels Ci, and the top n% channels were chosen after sorting Ci. The model was then retrained and tested for decoding tasks using these selected channels. An equal number of non-top n% channels were randomly chosen for decoding as well.
Baseline model
In the baseline comparison, we employed four widely used classifiers: logistic regression, AdaBoost, random forest, and support vector machine (SVM). Logistic regression was implemented with an L2 regularization term to prevent overfitting, while AdaBoost utilized decision trees as base learners and was set to iterate over 50 estimators. Random forest was constructed with 100 decision trees to ensure robust performance, and the SVM used a radial basis function (RBF) kernel to capture non-linear relationships in the data. All baseline models were optimized using grid search for hyperparameter tuning, ensuring a fair comparison with our proposed LS-STM model.
Data availability
All data used in this study are publicly available. The Human Brain data used in this study are from Mariana P. Branco etc. work.3 Online Database is available at https://openneuro.org/datasets/ds003688. The data of Mouse Brain were downloaded from Allen Institute Accession: https://portal.brain-map.org/.
Code availability
The source code of LS-STM is publicly available on GitHub (https://github.com/weightwater/LS-STM).
Change history
12 June 2025
In the Acknowledgment the following funding information sentences “the National Key Research and Development Project of China under Grant 2022YFF1202900” has been moved at the beginning of the Acknowledgment section.
References
Bouton, C. E. et al. Restoring cortical control of functional movement in a human with quadriplegia. Nature 533, 247–250 (2016).
Chang, C. et al. Tracking brain arousal fluctuations with fMRI. Proc. Natl. Acad. Sci. USA 113, 4518–4523 (2016).
Berezutskaya, J. et al. Open multimodal iEEG-fMRI dataset from naturalistic stimulation with a short audiovisual film. Sci. Data 9, 91 (2022).
Gründemann, J. et al. Amygdala ensembles encode behavioral states. Science 364, eaav8736 (2019).
Weber, S. C., Kahnt, T., Quednow, B. B. & Tobler, P. N. Frontostriatal pathways gate processing of behaviorally relevant reward dimensions. PLoS Biol. 16, e2005722 (2018).
Günseli, E. et al. Eeg dynamics reveal a dissociation between storage and selective attention within working memory. Sci. Rep. 9, 13499 (2019).
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364, eaav7893 (2019).
Musall, S., Kaufman, M. T., Juavinett, A. L., Gluf, S. & Churchland, A. K. Single-trial neural dynamics are dominated by richly varied movements. Nat. Neurosci. 22, 1677–1686 (2019).
Brazier, M. A. A history of the electrical activity of the brain: The first half-century. (1961).
Roach, B. J. & Mathalon, D. H. Event-related eeg time-frequency analysis: an overview of measures and an analysis of early gamma band phase locking in schizophrenia. Schizophrenia Bull. 34, 907–926 (2008).
Keren, H. et al. Reward processing in depression: a conceptual and meta-analytic review across fMRI and EEG studies. Am. J. Psychiatry 175, 1111–1120 (2018).
Radua, J. et al. Ventral striatal activation during reward processing in psychosis: a neurofunctional meta-analysis. JAMA psychiatry 72, 1243–1251 (2015).
Garasto, S., Bharath, A. A. & Schultz, S. R. Visual reconstruction from 2-photon calcium imaging suggests linear readout properties of neurons in mouse primary visual cortex. bioRxiv 300392 (2018).
Yoshida, T. & Ohki, K. Natural images are reliably represented by sparse and variable populations of neurons in visual cortex. Nat. Commun. 11, 872 (2020).
Ellis, R. J. & Michaelides, M. High-accuracy decoding of complex visual scenes from neuronal calcium responses. BioRxiv 271296 (2018).
Yargholi, E. & Hossein-Zadeh, G.-A. Brain decoding-classification of hand written digits from fmri data employing bayesian networks. Front Hum. Neurosci. 10, 351 (2016).
De Luca, D. et al. Convolutional neural network classifies visual stimuli from cortical response recorded with wide-field imaging in mice. J. Neural Eng. 20, 026031 (2023).
Chen, C., Batselier, K., Yu, W. & Wong, N. Kernelized support tensor train machines. Pattern Recognit. 122, 108337 (2022).
Ma, Z., Yang, L. T., Lin, M., Zhang, Q. & Dai, C. Weighted support tensor machines for human activity recognition with smartphone sensors. IEEE Trans. Industr. Info. (2021).
Nguyen, V.-D., Abed-Meraim, K. & Linh-Trung, N. Fast tensor decompositions for big data processing. In 2016 International Conference on Advanced Technologies for Communications (ATC), 215–221 (IEEE, 2016).
Ma, Z., Yang, L. T. & Zhang, Q. Support multimode tensor machine for multiple classification on industrial big data. IEEE Trans. Ind. Inf. 17, 3382–3390 (2020).
Qiu, Y., Zhou, G., Huang, Z., Zhao, Q. & Xie, S. Efficient tensor robust pca under hybrid model of tucker and tensor train. IEEE Signal Process Lett. 29, 627–631 (2022).
Sofuoglu, S. E. & Aviyente, S. Multi-branch tensor network structure for tensor-train discriminant analysis. IEEE Trans. Image Process 30, 8926–8938 (2021).
Tao, D., Li, X., Hu, W., Maybank, S. & Wu, X. Supervised tensor learning. In Fifth IEEE International Conference on Data Mining (ICDM’05), 8–pp (IEEE, 2005).
Xu, L., Cheng, L., Wong, N. & Wu, Y.-C. Tensor train factorization under noisy and incomplete data with automatic rank estimation. Pattern Recognit. 141, 109650 (2023).
Ma, Z., Yang, L. T., Lin, M., Zhang, Q. & Dai, C. Weighted support tensor machines for human activity recognition with smartphone sensors. IEEE Transactions on Industrial Informatics. (IEEE, 2021).
Bai, Z., Li, Y., Woźniak, M., Zhou, M. & Li, D. Decomvqanet: Decomposing visual question answering deep network via tensor decomposition and regression. Pattern Recognit. 110, 107538 (2021).
Tangwiriyasakul, C. et al. Tensor decomposition of tms-induced eeg oscillations reveals data-driven profiles of antiepileptic drug effects. Sci. Rep. 9, 17057 (2019).
Armingol, E. et al. Context-aware deconvolution of cell–cell communication with tensor-cell2cell. Nat. Commun. 13, 3665 (2022).
Pellegrino, A., Stein, H. & Cayco-Gajic, N. A. Dimensionality reduction beyond neural subspaces with slice tensor component analysis. Nat. Neurosci. 1–12 (2024).
Ahmadi, S. & Rezghi, M. Generalized low-rank approximation of matrices based on multiple transformation pairs. Pattern Recognit. 108, 107545 (2020).
Koniusz, P., Wang, L. & Cherian, A. Tensor representations for action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 44, 648–665 (2021).
Liu, J., Zhu, C., Long, Z., Huang, H. & Liu, Y. Low-rank tensor ring learning for multi-linear regression. Pattern Recognit. 113, 107753 (2021).
Suykens, J. A. & Vandewalle, J. Least squares support vector machine classifiers. Neural Process Lett. 9, 293–300 (1999).
Acknowledgements
This work was supported in part by the National Key Research and Development Project of China under Grant 2022YFF1202900, in part by the National Natural Science Foundation of China under Grant 62401331, 62303262, 62088102, and U21B2013, in part by the China Postdoctoral Science Foundation under Grant 2022TQ0179 and Grant 2023M741991, in part by the Beijing Nova Program (Grants No. 2021B00003292, 20220484216), in part by the National Key Research and Development Project of China (Grant No. 2023YFC2415600), and in part by the MOST under Grants 2020AA0105500 and 2022YFF1202904.
Author information
Authors and Affiliations
Contributions
B.Y.Z. and T.S. designed the methodology and wrote the manuscript. B.Y.Z. and T.S. wrote software and performed formal analysis. T.S., B.Y.Z., and Y.H.Z. performed data curation and pre-processing. T.S., B.Y.Z., Y.H.Z., and S.W. conceptualized the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks David Halliday and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Marta Vallejo and Benjamin Bessieres. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zang, B., Sun, T., Lu, Y. et al. Tensor-powered insights into neural dynamics. Commun Biol 8, 298 (2025). https://doi.org/10.1038/s42003-025-07711-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07711-x
This article is cited by
-
Distinguishing task-evoked dynamic brain networks from intrinsic activity with tensor component analysis
Brain Imaging and Behavior (2026)