Abstract
Acinetobacter baumannii (A. baumannii) is an important pathogen causing various nosocomial infections. CRISPR-Cas system is the adaptive immune system of bacteria, which is also closely related to the drug resistance and virulence of bacteria. However, the effect and mechanism of cas3 (type I-Fa) in A. baumannii is still unclear. In this study, we successfully constructed a cas3 deletion mutant (19606Δcas3) and complemented strain (19606Δcas3/pcas3) to study the regulatory mechanism of type I-Fa cas3 on bacterial virulence. Our results showed that deletion of cas3(type I-Fa) significantly reduced the biofilm formation, virulence and pathogenicity to mice. The organ bacterial load of mice infected with cas3 deletion strain was significantly reduced, the lung inflammation was slightly changed, and the serum cytokine level was also decreased. All results demonstrated that cas3 enhanced the virulence and pathogenicity of A. baumannii. Mechanism analysis showed that deletion of cas3 can lead to the down-regulation of virulence factors such as biofilm formation related factors and outer membrane protein A(ompA). In addition, cas3 was also involved in the regulation of carbon metabolism and oxidative phosphorylation pathway of A. baumannii. Altogether, our study may provide cas3 as a therapeutic target in the future because of the close link to the virulence of A. baumannii.
Similar content being viewed by others
Introduction
A cinetobacter baumannii (A. baumannii) is an important gram-negative non-fermenting opportunistic pathogen that causes hospital-acquired pneumonia and ventilator-associated pneumonia easily in hospital patients with low immunity1. Due to the rapid development of its antibiotic resistance, the World Health Organization has listed carbapenem-resistant A. baumannii as one of the pathogens in urgent need of exploring new drugs2. In addition to studying its drug resistance mechanism, more and more virulence-related studies on A. baumannii have been conducted in recent years. For example, the major virulence factors found include those that relate to adherence (Ata, TFP)3,4, effector delivery systems (T2SS, T6SS)5,6, exotoxin (phospholipase C, phospholipase D)7,8, exoenzyme (CpaA)9, immune modulation (capsule, LPS, ompA, pbpG)10,11,12,13, biofilm (adeFGH efflux pump, Bap, Csu fimbriae, PNAG, Quorom sensing)14,15,16,17,18, and nutritional/metabolic factors (Acinetobactin, HemO cluster)19,20. With the absence of some virulence factors, the pathogenicity of the highly virulent strain in the mouse infection model is remarkably reduced21. Therefore, the regulation of different virulence factors may point to a new direction in the treatment of A. baumannii infection.
The clustered regularly interspaced short palindromic repeats (CRISPR)-Cas system is an acquired immune system widely present in genomes of archaea and bacteria, which acts as a defense mechanism against the invasion of exogenous nucleic acids such as phages and plasmids22. CRISPR-Cas systems consist of the CRISPR arrays, leader sequences, and CRISPR-associated (cas) genes. Type I CRISPR-Cas system is the most widely distributed in bacteria, and Type I-F CRISPR-Cas system is the main type existing in A. baumannii23. Type I-F CRISPR-Cas system is characterized by the fusion of Cas2 and Cas3 (Cas2/3), as well as the integration of the Cas1-mediated spacer to the CRISPR site24. The I-Fa subtype of the CRISPR-Cas system is composed of the cas genes: cas1, cas3, csy2, csy3, and cas6 (Fig. 1). Cas3 is the marker of type I CRISPR-Cas system, which encodes a protein that may form cascade-like complexes with different compositions, in addition, it encodes a large protein with independent helicase and deoxyribonuclease activities25.
Composition of the type I-Fa CRISPR-Cas system.
Apart from its role in adaptive immunity, CRISPR-Cas system also plays an important role in regulating gene expression, especially in the regulation of bacterial virulence and population behavior26. Previous studies have shown that the CRISPR-Cas system can regulate the synthesis of outer membrane proteins of Salmonella typhi by regulating the outer membrane protein synthesis gene OmpR27. The biofilm formation, virulence, and pathogenicity of Salmonella are reduced after cas3 deleted in Salmonella28. Jose Solbiati et al.29 found that the virulence of the pathogen Porphyromonas gingivalis is controlled by the CRISPR-Cas protein Cas3. These all indicated that CRISPR-Cas system might be a potential virulence regulatory factor. Our previous research mainly focused on the effect of type I-Fb CRISPR-Cas system on the biological traits in A. baumannii30,31, and conducted the whole genome analysis of the related strains32. We found that different subtypes (I-Fa and I-Fb) cas genes had low homology and identity. The results showed that a complete I-Fb CRISPR-Cas system could inhibit the acquisition of antimicrobial resistance and reduce the virulence of A. baumannii31. However, the role of the type I-Fa CRISPR-Cas system in the acquisition of antimicrobial resistance and virulence of A. baumannii is still unclear. Cas3 is a key protein of I-Fa CRISPR-Cas system24. To further understand the role of cas3 gene in the virulence of A. baumannii, we constructed a type I-Fa cas3 gene knockout strain, and complemented the deletion strain in A. baumannii ATCC19606. We explored the effect of cas3 gene deletion on bacterial growth, biofilm formation, and measured adhesion and invasion efficiency on A549 cells. We also established a bacteremia model of A. baumannii infection in mice to evaluate the role of cas3 in A. baumannii infection. In conclusion, our results indicated that type I-Fa cas3 is essential for biofilm formation and virulence of A. baumannii.
Results
Deletion of cas3 gene does not affect growth
We successfully constructed the 19606Δcas3 mutant, and 19606Δcas3/pcas3 complement strains, which were verified by PCR and sequencing (Figs. S1 and S2). The ATCC19606, 19606Δcas3, and 19606Δcas3/pcas3 strains were cultured in a shaking table, and the standard growth curves were measured for 24 h. According to the OD600 value and the number of viable bacteria, the growth of ATCC19606, 19606Δcas3, and 19606Δcas3/pcas3 showed no significant difference (Fig. 2).
A OD600 value for 24-hour bacterial growth test. B Bacterial calculate after dilution plating. Values represent mean ± SD (n = 3).
Deletion of cas3 gene reduces bacterial biofilm formation
Bacterial biofilm is defined as a structured microbial community adherent to surfaces and encased within a self-produced extracellular polymeric matrix. Strong biofilm-forming bacterial strains can exhibit significantly enhanced resistance to both antimicrobial agents and host immune defense33. In order to evaluate the effect of cas3 gene on bacterial biofilm formation, the biofilm formation ability was quantified by crystal violet staining, and the results showed that both ATCC19606 strain and the 19606Δcas3 mutant strain could form biofilm. However, the ability to form biofilms after cas3 gene knockout was significantly reduced, and the complementary strain restored the biofilm-forming ability (Fig. 3A, B). The bacterial biofilm matrix is composed of a complex consortium of extracellular polymeric substances, including exopolysaccharides (EPS), structural and functional proteins, lipids, extracellular DNA (eDNA), lipoteichoic acids, and lipopolysaccharides. To comprehensively characterize the three-dimensional architecture and spatial organization of these biofilm components, we employed confocal laser scanning microscopy (CLSM) with appropriate fluorescent staining protocols. Specifically, the extracellular polysaccharide (EPS) matrix was labeled using Alexa Fluor 647-conjugated dextran (emission: 668 nm), while bacterial cells within the biofilm were counterstained with SYTO9 green fluorescent nucleic acid stain (emission: 498 nm)34. This staining strategy enables distinct visualization of biofilm architecture, with EPS components exhibiting red fluorescence and bacterial cells displaying green fluorescence, allowing for clear spatial differentiation between the extracellular matrix and cellular constituents. As shown in Fig. 3C, D, the fluorescence intensities of ATCC19606 and 19606Δcas3/pcas3 strains were similar under CLSM. In addition, the fluorescence intensity of 19606Δcas3 strain biofilm was significantly lower, and the thickness of the biofilm exhibited a significant reduction in the ∆cas3 strain. Therefore, cas3 gene of the type I-Fa CRISPR-Cas system is an important factor affecting the biofilm formation of A. baumannii.
A Quantification of 19606WT, 19606Δcas3, and 19606Δcas3/pcas3 strains biofilm formation by crystal violet staining. B Observation of biofilm growth of A. baumannii strains incubated at 37 °C for 48 hours by crystal violet staining in sterile 24-well plates. C Images of A. baumannii biofilm (EPS (red) and bacterial cells (green)) under a confocal microscope. D Thickness of the formed biofilm using ZEN software. Data are presented as mean ± SD from three independent biological replicates. P values were determined using a two-tailed Student’s t test, *p < 0.05, **p < 0.01.
Deletion of cas3 gene decreases virulence and pathogenicity of A. baumannii
As the knockout of cas3 gene resulted in a decrease in the biofilm formation ability of the strain, and biofilm is also a virulence-related factor that can avoid immune-mediated clearance and establish persistent infection, we speculated that the virulence of 19606Δcas3 may also be changed. The virulence of bacteria is closely related to the adhesion and invasion of bacteria into host cells. Epithelial cells form a layer of cells that cover the internal and external surfaces of the human body, constituting the protective barrier of the skin and various organs. A. baumannii employs multiple molecular mechanisms to invade epithelial cells, with its adhesive and invasive capabilities being closely associated with bacterial virulence. The deletion of the cas3 gene has been shown to significantly alter the pathogen’s virulence profile. To systematically investigate these pathogenic characteristics, we conducted in vitro infection assays using A549 human alveolar epithelial cells as a model system3,35. Specifically, confluent monolayers of A549 cells were infected with three isogenic strains: 19606WT, 19606Δcas3, and 19606Δcas3/pcas3 at a standardized multiplicity of infection (MOI) of 100, and their adhesion and invasion ability to mammalian cells were observed. As shown in Fig. 4, compared with 19606WT and 19606Δcas3/pcas3, the adhesion rate and invasion rate of strain 19606Δcas3 were significantly reduced.
A Adhesion rate to A549 cells. B Invasion rate to A549 cells. Values represent mean ± SD (n = 3). P values were determined by two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001.
To evaluate the effect of cas3 gene on the virulence of A. baumannii, an infection model of Galleria mellonella was established, and the survival was observed. An equal bacterial inoculum (1.0 × 106 CFU in 100 μL PBS) was administered to each Galleria mellonella. Results were shown in Fig. 5A, as we can see that a total of 90% larvae infected with ATCC19606 strain died within 12 hours, while only 20% of the larvae infected with 19606Δcas3 strain died. After 24 hours, all larvaes infected with ATCC19606 strain and the complementary strain were dead, while those infected with 19606Δcas3 strain maintained a 50% survival rate after 96 hours.
A Survival rates of G. mellonella larva. Infected larvae (n = 10) with 10 μl (~1.0 × 106 CFU) 19606WT, 19606Δcas3, and 19606Δcas3/pcas3 strains at the right posterior gastropoda. B Bacterial load of organs in mouse bacteremia model (n = 8). Bacterial load determinations for 19606WT, 19606Δcas3 and 19606Δcas3/pcas3 in murine infected organs at 24 h post infection. P values were determined by two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001.
To further evaluate the impact of cas3 gene deletion on the virulence of A. baumannii, we established a murine infection model that closely mimics human disease progression. Based on the above findings, we hypothesized that the cas3 knockout strain would exhibit attenuated infectivity and reduced bacterial burden in vivo compared to the wild-type and complemented strains. The mice were intraperitoneally injected with the bacterial suspension to construct a systemic infection model and further understand the virulence changes. After 24 h infection, we observed that the bacterial load of each organs in infected 19606Δcas3 mice was significantly lower than that in infected ATCC19606 and 19606Δcas3/pcas3 strains (Fig. 5B). The changes of pneumonic inflammation and injury were observed by hematoxylin and eosin (H&E) staining. Representative pictures were shown in Fig. 6B, the alveolar structure in the lung of ATCC19606-infected mice was filled with neutrophil infiltration and edema, and the alveolar structure was incomplete. However, the injury was milder in mice infected with 19606Δcas3 and was not observed in non-infected mice. The H&E staining results in the 19606Δcas3/pcas3 strain infection group were consistent with the observation and analysis results in the ATCC19606 infection group. In addition, enzyme-linked immunosorbent assay (ELISA) was used to determine the levels of cytokines (interleukin -1β [IL-1β], IL-6, and tumor necrosis factor-α [TNF-α]) in mice serum. Results showed that IL-1β, IL-6, and TNF-α in the 19606Δcas3-infected mice were reduced significantly compared to the ATCC19606 or 19606Δcas3/pcas3-infected mice (Fig. 6A). In conclusion, our results demonstrate that the presence of cas3 gene could enhance the virulence and pathogenicity of A. baumannii.
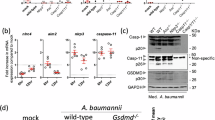
A Detection of TNF-α, IL-6, IL-1β in serum by ELISA, values represent mean ± SD (n = 8), P values were determined by two-tailed Student’s t test, **p < 0.01, ***p < 0.001. B Right lung tissue was stained with hematoxylin and eosin. Magnification, ×100. The arrow indicates the site of alveolar destruction and inflammatory cell infiltration.
RNA-seq to explore the potential regulatory mechanism of cas3
In order to further understand the mechanism of cas3 gene in regulating biofilm and virulence, we performed transcriptome sequencing analysis. After cas3 gene knockout, other related cas genes in CRISPR-Cas system were upregulated except for cas3 and cas6 genes, as shown in Table 1. Compared with the ATCC19606 strain, 539 genes in 19606Δcas3 strain were differentially expressed, including 272 genes upregulated and 267 genes downregulated (Fig. 7A and Supplementary Data 1), suggesting that cas3 gene was associated with multiple activities that are essential for bacteria. The differential expressed genes (DEGs) between ATCC19606 and 19606Δcas3 strains were evaluated according to Gene Ontology (GO) classification to investigate potential biological functions in which DEGs might be involved (Fig. 7C), differentially expressed genes were mainly present in biological process, cellular component and molecular function. In the classification of biological processes, differentially expressed genes are mainly enriched in redox, fatty acid metabolism, cation transport, monocarboxylic acid transport, metal ion transport, and other processes. In the classification of cell composition, it is mainly enriched in protein complexes, the cell membrane, the mitochondrial membrane, and other components. In molecular function, it is mainly enriched in oxidoreductase activity, adenine flavin dinucleotide binding, 3-hydroxyacyl-CoA dehydrogenase activity, ion transmembrane transporter activity, and other functions.
A Differential gene volcano map. B Venn diagram of gene co-expression. C GO analysis of different expression genes. D KEGG enrichment analysis of differentially expressed genes; the color of the dot represents the size of the Q value, and the size of the dot represents the number of differentially genes.
Different genes usually coordinate with each other to exercise their biological functions in microbiology. Through the significant pathway enrichment in the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, we can identify the most important biochemical metabolic pathways and signal transduction pathways involved in the differentially expressed genes. As shown in Fig. 7D, the main pathway changes were microbial metabolism in diverse environments, carbon metabolism, pyruvate metabolism, and benzoate degradation, and the top 20 pathways are shown in the figure. We also found that genes related to the oxidative phosphorylation pathway are significantly downregulated, and oxidative phosphorylation is the pathway of ATP synthesis, so we detected the intracellular ATP of ATCC19606, 19606Δcas3, and 19606Δcas3/pcas3. The result is consistent with our conjecture; the intracellular ATP content of 19606Δcas3 decreased significantly (Fig. S3).
Our results showed that the deletion of cas3 gene had a strong impact on the biofilm and virulence. Analysis of transcriptome results also indicated that the expression of biofilm and virulence genes was downregulated after cas3 gene knockout, as shown in Table 2. The expression levels of genes associated with Csu pili (specifically csuA, csuB, csuC, and csuE) were observed to be significantly downregulated. It is noteworthy that the majority of A. baumannii strains have the ability to encode and produce type I chaperone-usher pilus, known as Csu pili. Csu pili play a critical role in the formation and maintenance of biofilms on abiotic surfaces36. Notably, the expression levels of OmpA family proteins and two-component regulatory system BfmSR were also downregulated. BfmR is essential for cell attachment and initiation of biofilm formation by A. baumannii 19606 cells37, and the results were verified by qRT-PCR (Fig. 8). Therefore, RNA-seq further indicated that cas3 gene could regulate the bacterial virulence and biofilm formation by affecting the expression of related genes.
Values represent mean ± SD (n = 3). P values were determined by two-tailed Student’s t test,*p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
In recent years, multi-drug resistant rates of A. baumannii have gradually increased, and MDR bacteria can tolerate a variety of adverse stimuli to adapt to the environment and survive easily36. CRISPR-Cas system plays an important role in the adaptive immune activity of bacteria, its main function is defending against bacteriophage, plasmids, and other foreign DNA22. Recent studies have shown that CRISPR-Cas system not only protects the host against mobile genetic elements, but also regulates other physiological functions of bacteria, such as virulence, biofilm formation, drug resistance, and quorum sensing26. Jianjian Jiao et al. found that Cas proteins were secreted virulence factor, which can activate the host immune response in Mycobacterium tuberculosis38. Our research discovered the unconventional function of A. baumannii type I-Fa CRISPR-Cas system, cas3 significantly enhanced the biofilm formation capacity of the strain, potentially through the regulation of Csu-related genes. This regulatory mechanism facilitates bacterial adhesion and host invasion, ultimately contributing to enhanced virulence.
In this study, we studied the effect of cas3 gene on the biological activity of A. baumannii by knocking out the cas3 gene. Results first showed that the deletion of cas3 gene did not affect its growth, indicating that cas3 gene was not necessary gene for bacterial growth. Biofilm formation is a collective behavior of bacteria regulated by quorum sensing, which helps bacteria stand up to external environmental pressure39,40. Through the biofilm formation assay, the biofilm formation ability of 19606Δcas3 strain was decreased compared with ATCC19606. Similar to a previous study showed that biofilm formation of the Δcas3 Streptococcus strain (type I-C CRISPR-Cas system) was reduced41. Also, type I-E CRISPR-Cas Salmonella typhimurium wild strain has a stronger biofilm formation ability compared with the Δcas3 strain28.
The effect of cas3 gene on the virulence of A. baumannii was explored through the infection model of Galleria mellonella and mice. Results showed that the virulence and pathogenicity of A. baumannii decreased after cas3 gene was knocked out. Previous studies have also shown that CRISPR-Cas can enhance or reduce the virulence of bacteria. For example, cas9 could enhance the virulence by inhibiting its transcription regulatory factor regR in Streptococcus agalactiae42. Type I-F CRISPR-Cas system of Pseudomonas aeruginosa PA14 can change its virulence by targeting and inhibiting LasR, and cas3 gene knockout increases the virulence of PA14 and the host’s innate immune response43.
After the deletion of cas3 gene, the expression of many genes was upregulated or downregulated, indicating that cas3 might be involved in the regulation of other bacterial life activities. Cas3 has an important influence on the expression of several Outer membrane protein A (OmpA) family protein. OmpA is a major component of outer membrane proteins (OMPs) in gram-negative bacteria, which is a key virulence factor. Overexpression of OmpA was strongly associated with severe hospital infection, bacteremia, and pneumonia infection44. Similarly, Medina-Aparicio L et al.27 studied the CRISPR-Cas system in Salmonella spp. and showed that the CRISPR-Cas system could participate in the regulation of OmpR gene synthesis of Salmonella typhi. It suggested to us that the regulation of bacterial virulence by cas3 could be targeted to the OmpA family protein. We also found that the largest number of gene changes was in the oxidation-reduction process in GO terms (Fig. 7C). KEGG analysis showed that the oxidative phosphorylation pathway was inhibited after cas3 gene deletion. The bacterial oxidative phosphorylation (OxPhos) system plays an essential role in energy production in the form of ATP, which has been considered to be a potential antibacterial drug target45. KEGG analysis showed that the oxidative phosphorylation pathway of A. baumannii was inhibited, and ATP production was decreased after cas3 gene deletion, which may be related to the weakening of the virulence of 19606Δcas3. What is also worth mentioning is that A. baumannii 17978 and some newly detected clinical A. baumannii strains do not have CRISPR-Cas system at all, this implies that the function of cas3 limited in certain A. baumannii strains.
In our previous study, we found that the resistance, biofilm formation ability, and virulence of the strain were upregulated after any cas gene knocked out in the type I-Fb CRISPR-Cas system46. However, the opposite situation appeared after the cas3 gene knocked out in the type I-Fa CRISPR-Cas system in this study, which indicated that the two subtypes of CRISPR-Cas system in A. baumannii played different regulatory roles in resistance and virulence. It is still necessary to continue to study how the CRISPR-Cas system regulates these functions. Exceedingly, all these evidences indicated that cas3 gene and CRISPR-cas system are closely linked to the virulence of A. baumannii, which maybe a therapeutic target in the future.
Methods
Bacterial strains, plasmids, and growth conditions
Bacterial strains, plasmids and primers used in this study are listed in Supplementary Table S1. A. baumannii and E.coli strains were grown in lysogeny broth (LB) at 37 °C. To select colonies during the construction of mutant and complementary strains, antimicrobial agents, tetracycline (50 μg/ml), carbenicillin (75 μg/ml), kanamycin (50 μg/ml), were added to the growth medium.
Construction of cas3 gene deletion strain 19606∆cas3
The Δcas3 mutant strain was constructed by a RecET recombinase system in A. baumannii ATCC19606 as previously described30,47. First, the kanamycin resistance gene fragment with FRT locus was amplified using pKD4 as the template, upstream(165 bp) and downstream(142 bp) fragments of cas3 were amplified using ATCC19606 as the template. All primer sequences and corresponding product sizes in this study were listed in Supplementary Table S2. The polymerase chain reaction (PCR) products were identified by agarose gel electrophoresis and purified using the Fast Pure gel DNA Extraction Mini Kit (Vazyme). Products of three stages were fused by fusion PCR, and the recombinant product was recovered by gel cutting and sequencing by Beijing Tsingke Biotech Co., Ltd. Then electrotransform the sequenced correct 1 μg fusion PCR product into ATCC19606-pAT04(plasmid pAT04 with RecAb system) competent cells (100 µL) in a 2 mm cuvette at 1.8 kV, transfer it to 4 mL liquid LB containing 2 mM IPTG culture for 4 hours, centrifuge at 4000 rpm for 10 minutes to collect bacteria. Incubate bacteria overnight on LB agar medium containing 50 μg/mL kanamycin and 10 μg/mL tetracyline at 37 °C. Positive clones confirmed the insertion of the kanamycin resistance gene by screening primers.
The kanamycin resistance gene in mutant 19606∆cas3::kan was then lost by the following procedure. First, the pAT03 (pMMB67EH with flippase recombinase) plasmid was introduced by electroporation into the ATCC19606∆cas3::kan mutant. Then, the bacteria were incubated with liquid LB containing 2 mM IPTG for 1 hour. Positive clones were selected on LB agar plates containing carbenicillin (75 μg/mL) and incubated overnight at 37 °C. cas3 gene knockout mutant was confirmed by PCR and sequencing of the cloned PCR fragment, primer sequences were listed in Table S2.
The single colonies verified by PCR as successful recombinant deletion mutants were subcultured on antibiotic-free LB solid plates and incubated statically at 37 °C. After three passages, several single colonies were selected for enrichment, with each colony simultaneously streaked onto both a dual-antibiotic plate (containing tetracycline and carbenicillin) and an antibiotic-free plate. If a colony grew only on the antibiotic-free plate but not on the antibiotic-containing plates, this indicated the loss of both the pAT04 and pAT03 plasmids.
Construction of 19606∆cas3/pcas3 and 19606∆cas3/empty vector
Complementation of cas3 gene in the gene knockout mutant was performed through constructing a recombinant plasmid by the pMMB67EH vector. A 3236 bp fragment of cas3 was amplified from ATCC19606 genome DNA and ligated to the pMMB67EH vector with the tac promoter. Upstream cas3 fragment with EcoR I enzymatic restriction site, and downstream cas3 fragment with BamH I enzymatic restriction site. Then, the recombinant plasmid was introduced by electroporation into 19606∆cas3 competent cells. We also constructed the 19606Δcas3 strain with an empty vector as a negative control, named 19606∆cas3/empty vector, the empty vector (pMMB67EH) was electro transformed into 19606∆cas3 competent cells. The ∆cas3 mutant transformed with the complementary vector was selected on LB agar plates containing 50 μg/mL kanamycin and 50 μg/mL tetracycline. Positive clones were verified by PCR and sequencing.
Determination of growth curve
ATCC19606, 19606∆cas3 and 19606∆cas3/pcas3 cultured overnight were inoculated into fresh liquid LB broth at a ratio of 1:1000, and cultured by shaking at 37 °C. The optical density at 600 nm (OD600) was measured every 2 hours after inoculation, bacteria counts were performed on LB plates after fold dilution in 96-well plates. Growth curve was drawn according to the results.
Biofilm formation assay
The bacteria were cultured in a shaking table at 37 °C until the OD600 value was 0.5 (the bacterial count was 1.0 × 108 CFU/mL), inoculated in 200 μL to 96-well plates at a ratio of 1:100, and placed in an incubator at 37 °C for 48 hours with no shaking. Then the bacterial solution was washed three times with PBS. After the 96-well plate was dried, it was dyed with 4% crystal violet (Solaribo) for 30 min in dark. Discard the staining solution and wash three times with PBS. Finally, the crystal violet dye adsorbed on the biofilm was dissolved with 95% ethanol, and the absorbance at 595 nm was measured. In this experiment, liquid LB medium without bacteria was used as the negative control.
Confocal scanning laser microscopy
The bacteria were cultured until the OD600 value was 1 and diluted 100-fold. Inoculate 500 μl of the bacterial liquid with the Alexa Fluor 647 (Thermo Fisher) and inoculated into a 24-well plate. A round coverslip, untreated with poly-glycine, was placed at the bottom of each well and placed in an incubator at 37 °C in the dark for about 24h. Discard the bacterial liquid and rinse it twice with physiological salt solution. According to the manufacturer’s instructions, the biofilm was dyed with SYTO9(Invitrogen) for 15 minutes in the dark, observed under a laser confocal scanning microscope and analyzed by ZEN software34.
Adhesion and invasion
The A549 cells were cultured in a 24-well cell culture plate to a cell density of ~3.0 × 105 cells/well. A. baumannii strains were infected with a bacterium: cell ratio of infection multiple (MOI) of 100:1. The adhesion and invasion of strains to A549 cells were measured by colony-forming unit (CFU) counts. After the bacteria strains and cells were co-cultured for 2 h, the extracellular free bacteria were removed by washing with PBS three times, and treated with freshly prepared sterile 0.1% Triton X-100 for 15 minutes. The cell culture medium containing 32 μg/mL tigecycline was replaced in the remaining wells and incubated for a further 90 min at 37 °C to kill bacteria adhering to the outside of the cells. Then discard the cell culture medium and wash the cells three times with PBS. Finally, 0.1% Triton X-100 was added to each well for incubation 10 minutes to release intracellular bacteria, and the bacterial CFUs assay was performed on agar plates. The results are expressed as the percentage of adherent and invasive bacterial counts relative to the number of infected bacteria for adhesion frequency and invasion frequency, respectively35.
Infection model of Galleria mellonella
All the Galleria mellonella larvae used in the experiment were purchased from the Organisms of Family Origin Company in Yunnan Province. We randomly selected four groups of healthy larvae (10/group) with no evidence of blacking, weighing between 200 ~ 300 mg, for subsequent infection. The Galleria mellonella larvae were infected by right posterior gastropoda injection of 10 μl (~1.0 × 106 CFU) each, control group was injected with equivalent PBS. The Galleria mellonella larvae, dead or not, were determined by two distinct phenotypic indicators: complete melanization of the cuticle and cessation of movement upon physical stimulation48. The number of dead larvae was observed and recorded every 12 hours.
Mouse bacteremia model and cytokines determination
Eight-week-old female C57 mice were raised under conventional conditions. Mice were handled following the Guidelines for the Care and Use of Laboratory Animals, and procedures were approved by the Ethical Committee of Yangzhou University. We have complied with all relevant ethical regulations for animal use. Cyclophosphamide was used to induce neutropenia in mice, as described previously31. The neutropenic mice were infected with 5 × 108 CFU of A. baumannii. The mice were sacrificed 24 hours after infection. Blood from the eyeball and organs were taken. The left lung and other organs were used for bacterial load count on LB agar plates. Right lung tissues were fixed in formalin solution and stained with H&E.
Cytokines in the serum of infected mice were detected by ELISA. Blood was taken from eyeball of each aforementioned mice. After blood was naturally coagulated at room temperature for 30 minutes, it was centrifuged at 1000 × g for 10 min at 4 °C. The supernatant(serum) was aliquoted into EP tubes and stored at −20 °C for future use. ELISA were completed by Solarbio’s kit.
RNA transcriptome sequencing and qRT-PCR
The bacteria RNA samples were subjected to quality detection, cDNA synthesis, cDNA fragment modification and sorting, library amplification, and the constructed cDNA library was sequenced by Illumina HiSeqTM. Technical services for this process and subsequent bioinformatics analysis were provided by Shanghai Shenggong Biotechnology Limited liability Company.
Bacteria used in qRT-PCR were cultured overnight to 109 CFU/ml, 1 ~ 3 ml of bacterial culture was centrifuged, and the total RNA of bacteria was extracted by RNAprep Pure Bacteria Kit (TIANGEN, Beijing) according to the instructions. cDNA was obtained by reverse transcription of RNA with reverse transcription kit (Vazyme, Nanjing, China), which was packaged and stored at −20 °C for future use, and qRT-PCR was performed with ChamQ SYBR qPCR Master Mix (Vazyme, Nanjing, China). The fold change in mRNA expression was calculated by 2−ΔΔCt method using 16S rRNA as reference gene. Primers used are listed in Supplementary Table 1.
Statistics and reproducibility
Data were presented as the mean standard deviation of replicates(n ≥ 3). Mean comparisons between groups were performed using the two-tailed Student’s t test. All statistical analyses were performed using GraphPad Prism version 8.4.2 (GraphPad Software Inc., San Diego, CA, USA), and a significant P value of <0.05 was considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-sequencing data have been deposited in the National Center for Biotechnology Information (NCBI) sequence read archive (SRA) database (PRJNA1209644). Source data for the main figures are provided in Supplementary Data 2. Uncropped gel images are provided in Supplementary Fig. 4 of Supplementary materials. All other data are available from the corresponding authors.
References
Ibn Saied, W. et al. A comparison of the mortality risk associated with ventilator-acquired bacterial pneumonia and nonventilator ICU-acquired bacterial pneumonia. Crit. Care Med. 47, 345–352 (2019).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Weidensdorfer, M. et al. The Acinetobacter trimeric autotransporter adhesin Ata controls key virulence traits of Acinetobacter baumannii. Virulence 10, 68–81 (2019).
Piepenbrink, K. H. et al. Structural diversity in the type IV Pili of multidrug-resistant acinetobacter. J. Biol. Chem. 291, 22924–22935 (2016).
Elhosseiny, N. M., El-Tayeb, O. M., Yassin, A. S., Lory, S. & Attia, A. S. The secretome of Acinetobacter baumannii ATCC 17978 type II secretion system reveals a novel plasmid encoded phospholipase that could be implicated in lung colonization. Int. J. Med. Microbiol.306, 633–641 (2016).
Fitzsimons, T. C. et al. Identification of novel Acinetobacter baumannii type VI secretion system antibacterial effector and immunity pairs. Infect. Immun. 86, e00297 (2018).
Camarena, L., Bruno, V., Euskirchen, G., Poggio, S. & Snyder, M. Molecular mechanisms of ethanol-induced pathogenesis revealed by RNA-sequencing. PLoS Pathog. 6, e1000834 (2010).
Jacobs, A. C. et al. Inactivation of phospholipase D diminishes acinetobacter baumannii pathogenesis. Infect. Immun. 78, 1952–1962 (2010).
Haurat, M. F. et al. The glycoprotease CpaA secreted by medically relevant acinetobacter species targets multiple O-linked host glycoproteins. mBio 11, e02033 (2020).
Russo, T. A. et al. The K1 capsular polysaccharide of Acinetobacter baumannii strain 307-0294 is a major virulence factor. Infect. Immun. 78, 3993–4000 (2010).
Boll, J. M. et al. Reinforcing lipid A acylation on the cell surface of acinetobacter baumannii promotes cationic antimicrobial peptide resistance and desiccation survival. mBio 6, e00478–00415 (2015).
Moon, D. C. et al. Acinetobacter baumannii outer membrane protein A modulates the biogenesis of outer membrane vesicles. J. Microbiol.50, 155–160 (2012).
Russo, T. A. et al. Penicillin-binding protein 7/8 contributes to the survival of Acinetobacter baumannii in vitro and in vivo. J. Infect. Dis. 199, 513–521 (2009).
He, X. et al. Biofilm formation caused by clinical acinetobacter baumannii isolates is associated with overexpression of the AdeFGH efflux pump. Antimicrob. Agents Chemother. 59, 4817–4825 (2015).
De Gregorio, E. et al. Biofilm-associated proteins: news from Acinetobacter. BMC Genomics 16, 933 (2015).
Tomaras, A. P., Dorsey, C. W., Edelmann, R. E. & Actis, L. A. Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology 149, 3473–3484 (2003).
Flannery, A., Le Berre, M., Pier, G. B., O’Gara, J. P. & Kilcoyne, M. Glycomics microarrays reveal differential in situ presentation of the biofilm polysaccharide poly-N-acetylglucosamine on acinetobacter baumannii and Staphylococcus aureus cell surfaces. Int. J. Mol. Sci. 21, 2465 (2020).
Niu, C., Clemmer, K. M., Bonomo, R. A. & Rather, P. N. Isolation and characterization of an autoinducer synthase from Acinetobacter baumannii. J. Bacteriol. 190, 3386–3392 (2008).
Giardina, B. J., Shahzad, S., Huang, W. & Wilks, A. Heme uptake and utilization by hypervirulent Acinetobacter baumannii LAC-4 is dependent on a canonical heme oxygenase (abHemO). Arch. Biochem. Biophys. 672, 108066 (2019).
Gaddy, J. A. et al. Role of acinetobactin-mediated iron acquisition functions in the interaction of Acinetobacter baumannii strain ATCC 19606T with human lung epithelial cells, Galleria mellonella caterpillars, and mice. Infect. Immun. 80, 1015–1024 (2012).
Wang, J. et al. The role of the type VI secretion system vgrG gene in the virulence and antimicrobial resistance of Acinetobacter baumannii ATCC 19606. PloS one 13, e0192288 (2018).
Burley, K. M. & Sedgley, C. M. CRISPR-Cas, a prokaryotic adaptive immune system, in endodontic, oral, and multidrug-resistant hospital-acquired Enterococcus faecalis. J. Endod. 38, 1511–1515 (2012).
Karah, N. et al. CRISPR-cas subtype I-Fb in Acinetobacter baumannii: evolution and utilization for strain subtyping. PloS ONE 10, e0118205 (2015).
He, L., St John James, M., Radovcic, M., Ivancic-Bace, I. & Bolt, E. L. Cas3 protein-a review of a multi-tasking machine. Genes11, 208 (2020).
Sinkunas, T. et al. Cas3 is a single-stranded DNA nuclease and ATP-dependent helicase in the CRISPR/Cas immune system. EMBO J. 30, 1335–1342 (2011).
Westra, E. R., Buckling, A. & Fineran, P. C. CRISPR-Cas systems: beyond adaptive immunity. Nat. Rev. Microbiol. 12, 317–326 (2014).
Medina-Aparicio, L. et al. The CRISPR-Cas system is involved in OmpR genetic regulation for outer membrane protein synthesis in salmonella typhi. Front. Microbiol. 12, 657404 (2021).
Cui, L. et al. CRISPR-cas3 of salmonella upregulates bacterial biofilm formation and virulence to host cells by targeting quorum-sensing systems. Pathogens 9, 53 (2020).
Solbiati, J., Duran-Pinedo, A., Godoy Rocha, F., Gibson, F. C. 3rd & Frias-Lopez, J. Virulence of the pathogen porphyromonas gingivalis is controlled by the CRISPR-Cas protein Cas3. mSystems 5, e00852–20 (2020).
Guo, T. et al. The involvement of the csy1 gene in the antimicrobial resistance of acinetobacter baumannii. Front. Med. 9, 797104 (2022).
Wang, Y. et al. CRISPR-Cas in Acinetobacter baumannii contributes to antibiotic susceptibility by targeting endogenous AbaI. Microbiol. Spectr. 10, e0082922 (2022).
Guo, T. et al. Whole-genome analysis of Acinetobacter baumannii strain AB43 containing a type I-Fb CRISPR-Cas system: insights into the relationship with drug resistance. Molecules 27, 5665 (2022).
Djebbi-Simmons, D., Xu, W., Janes, M. & King, J. Survival and inactivation of Salmonella enterica serovar Typhimurium on food contact surfaces during log, stationary and long-term stationary phases. Food Microbiol. 84, 103272 (2019).
Xiao, J. et al. The exopolysaccharide matrix modulates the interaction between 3D architecture and virulence of a mixed-species oral biofilm. PLoS Pathog. 8, e1002623 (2012).
Rumbo-Feal, S. et al. Contribution of the A. baumannii A1S_0114 gene to the interaction with eukaryotic cells and virulence. Front. Cell Infect. Microbiol 7, 108 (2017).
Harding, C. M., Hennon, S. W. & Feldman, M. F. Uncovering the mechanisms of Acinetobacter baumannii virulence. Nat. Rev. Microbiol. 16, 91–102 (2018).
Tomaras, A. P., Flagler, M. J., Dorsey, C. W., Gaddy, J. A. & Actis, L. A. Characterization of a two-component regulatory system from Acinetobacter baumannii that controls biofilm formation and cellular morphology. Microbiology 154, 3398–3409 (2008).
Jiao, J. et al. M. tuberculosis CRISPR/Cas proteins are secreted virulence factors that trigger cellular immune responses. Virulence 12, 3032–3044 (2021).
Flemming, H. C. et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575 (2016).
Álvarez-Fraga, L. et al. Analysis of the role of the LH92_11085 gene of a biofilm hyper-producing Acinetobacter baumannii strain on biofilm formation and attachment to eukaryotic cells. Virulence 7, 443–455 (2016).
Tang, B. et al. Deletion of cas3 gene in Streptococcus mutans affects biofilm formation and increases fluoride sensitivity. Arch. Oral. Biol. 99, 190–197 (2019).
Ma, K. et al. cas9 enhances bacterial virulence by repressing the regR transcriptional regulator in streptococcus agalactiae. Infect. Immun. 86, e00552–17 (2018).
Li, R. et al. Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity. Cell Res. 26, 1273–1287 (2016).
Sánchez-Encinales, V. et al. Overproduction of outer membrane protein a by acinetobacter baumannii as a risk factor for nosocomial pneumonia, bacteremia, and mortality rate increase. J. Infect. Dis. 215, 966–974 (2017).
Rubin, H. et al. Acinetobacter baumannii OxPhos inhibitors as selective anti-infective agents. Bioorg. Med. Chem. Lett. 25, 378–383 (2015).
Wang, Y. H. et al. CRISPR-Cas in contributes to antibiotic susceptibility by targeting endogenous AbaI. Microbiol. Spectr. 10, e0082922 (2022).
Tucker, A. T. et al. Defining gene-phenotype relationships in Acinetobacter baumannii through one-step chromosomal gene inactivation. mBio 5, e01313–e01314 (2014).
Tao, Y., Duma, L. & Rossez, Y. Galleria mellonella as a good model to study acinetobacter baumannii pathogenesis. Pathogens 10, 1483 (2021).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82373637, 82073611, 82002186), the Jiangsu Provincial Natural Science Foundation of China (BK20231241), and the Open Project Program of Jiangsu Key Laboratory of Zoonosis (Grant number: R2201).
Author information
Authors and Affiliations
Contributions
G.L., J.H., and T.G. designed and supervised the project. T.G., J.Y., and N.Z. performed all experiments. J.Y. and X.S. analyzed the data and prepared all figures. T.G., J.Y., C.H., T.L., and G.B. wrote the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks K Prashanth and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Jonathan Perreault and Mengtan Xing. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, T., Yang, J., Zhou, N. et al. Cas3 of type I-Fa CRISPR-Cas system upregulates bacterial biofilm formation and virulence in Acinetobacter baumannii. Commun Biol 8, 750 (2025). https://doi.org/10.1038/s42003-025-08124-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08124-6