Abstract
DIAPH1 is a member of the family of Diaphanous Related Formins (DRFs) implicated in cell migration and cytokinesis. DRFs are maintained in an autoinhibited state by the intramolecular association between diaphanous inhibitory (DID) and diaphanous autoregulatory (DAD) domains. Actin polymerization requires the binding of activated RhoA to the GTPase binding domain (GBD) of DIAPH1 and the dissociation of DAD. In the presence of excess RhoA, actin polymerization is only partially activated. Using monomeric domain constructs of DIAPH1, the sequential binding affinities of RhoA and DAD to GBD-DID were characterized. Binding of RhoA and DAD were negatively cooperative requiring a 100-fold greater concentration of DAD to achieve saturation when RhoA binding site was occupied. The unimolecular architecture of full length DIAPH1 establishes an effective concentration of DAD in the micromolar range, which is 100-fold larger than the intrinsic affinity of DAD for DID. The effective concentration is large enough to maintain DIAPH1 autoinhibition, yet small enough to permit partial activation of DIAPH1 after RhoA binding. By exploiting negative cooperativity, DIAPH1 maintains a reserve of inactivated molecules enabling gradual responses to cellular processes that require prolonged and sustained regulation. The proposed mechanism is extended to other DIAPH1 activating ligands and broadly applicable to all DRFs.
Similar content being viewed by others
Introduction
Many cellular responses to stimuli are mediated through the formation of membrane bound complexes whose activity is regulated by the binding of effector molecules1,2,3. These complexes contain multimeric proteins capable of binding different ligands. Linkages between ligand binding can alter the affinity of other ligands to modulate the specificity and level of activity4,5,6. The diaphanous related formins (DRFs) are dimeric multidomain proteins that facilitate cytoskeletal dynamics relating to cell migration, phagocytosis, molecular trafficking and cytokinesis by regulating the assembly and dynamics of actin filaments7,8,9. Aberrant DRF activation in humans has been implicated in numerous disease pathologies including inflammation and diabetic complications through interactions with the receptor for advanced glycation end products (RAGE)10,11,12,13,14, cortical blindness15, autosomal dominant deafness16,17,18, moya-moya disease19, thrombocytopenia20,21, focal segmental glomerulosclerosis, and Charcot-Marie-Tooth disease22.
DRFs are distinct from other formin homology proteins in that they contain a highly conserved N-terminal GTPase-binding domain (GBD), a Diaphanous Inhibitory Domain (DID), and a C-terminal Diaphanous autoregulatory domain (DAD) connected by long unstructured linkers (Fig. 1A). DRFs are maintained in an autoinhibited (inactive) state by the intramolecular association of the N-terminal DID and C-terminal DAD domains. Binding of GTP-bound Ras homolog gene family A (RhoA) GTPase to the GBD of an autoinhibited DRF induces actin polymerization by promoting the dissociation of the DID-DAD domains, relieving the autoinhibited state and allowing the Formin Homology Domain 2, FH2, to bind to globular G-actin creating filamentous F-actin. A theoretical ternary complex of RhoA, GBD-DID (GDID), and DAD created by superimposing crystallographic and NMR structures shows that Rho binding reorients the GBD so that it sterically and electrostatically clashes with the N-terminus of the amphipathic DAD helix (Fig. 1B)12,23,24,25,26.
A Schematic representation of the domain structure of DRFs. N-terminal domain (N, black); GTPase binding domain (GBD, red); Diaphanous Inhibitory Domain (DID, orange); Interhelical Domain (IH, yellow); Dimerization Domain (DD, green); Intrinsically Disordered Region (IDR, cyan); Coiled Coil Domain (CC, brown); Formin Homology Domains 1 and 2 (FH1, gray, and FH2, pink); Diaphanous Autoinhibitory Domain (DAD, blue); C-Terminal Domain (C, gold). Diaphanous homologs 1, 2, and 3 (DIAPH1, 2, and 3); Formin Like Protein 1 (FMNL1); Inverted Formin 2 (INF2); Formin Homology Domain Containing Protein 1 (FHOD1); Disheveled Associated Activator of Morphogenesis 1 (DAAM1); Constitutively active DIAPH1 (DIAPH1ΔDAD); DIAPH 1 mimic of DIAPH 2 (DIAPH1/2); GBD-DID construct (GDID); DAD construct (DAD). B Ribbon diagram showing two views of a superposition of the DID-DAD structure (PDB entry 8FG1) and the GBD-DID-RhoC structure (PDB entry 1Z2C); RhoC amino acid sequence is 92% identical to RhoA and backbone RMSD between RhoA (PDB entry 1A2B) and RhoC (PDB entry 2GCO) is ~1 Å. The residues of DAD that sterically clash with the GBD because of Rho binding are shown in cyan (box). The color scheme is the same as in panel A and RhoC is in purple. C Thermodynamic cycle of sequential DAD, D, and RhoA, R, binding to GBD-DID, G, and the corresponding equilibrium expressions39 where [R] and [D] are the concentrations of RhoA and DAD, KdR and KdD, are the intrinsic dissociation constants for RhoA and DAD binding to GDID, and KdR/D and KdD/R are the apparent dissociation constants for RhoA binding to GDID-DAD and DAD binding to RhoA-GDID, respectively. α is a cooperativity factor where α > 1 denotes positive cooperativity and α < 1 denotes negative cooperativity.
The functional linkages between RhoA and DAD binding to GDID and actin polymerization have been investigated using monomeric domains derived from mouse Diaphanous 1 (DIAPH1), also called mDia1. Isothermal calorimetry and fluorescence polarization experiments showed RhoA binds tightly to the GDID of mouse DIAPH1 and competes with DAD by direct displacement. Dramatic decreases in fluorescence polarization suggested that RhoA displaces DAD to just above baseline levels26,27. RhoA binds to GDID with nanomolar affinity but requires micromolar concentrations to activate actin polymerization, and then only partial activation is achieved23,28,29,30,31. The only known RhoA-regulated effect on DRFs is the release of DAD from DID23,26,27,28,29,30,31,32,33,34, which cannot explain the partial activation by RhoA or whether DRF activation by RhoA is modulated by other proteins. Why is such a high concentration of RhoA required to activate Diaphanous-induced actin polymerization? And why is only partial activation achieved? This is the enigma that has puzzled researchers31.
A thermodynamic cycle describing the sequential binding of RhoA and DAD binding to GDID similar to that described by Rosen35,36 for Wiskott-Aldrich Syndrome Protein regulation offers insight as to how regulation of actin polymerization by DIAPH1 may work (Fig. 1C). Previous estimates for the intrinsic binding affinity of RhoA, KdR, and DAD, KdD, for GDID were ~6 nM25,27,34,37 and ~250 nM25,26,28,34, respectively, and ~29 μM for DAD binding to a Rho-GDID complex, KdD/R 26. Conservation of energy predicts an apparent dissociation constant for Rho binding to the GDID-DAD complex, KdR/D, of ~1 μM. Assigning binding of the second ligand as the sum of the intrinsic free energy and an interaction free energy predicts a positive interaction free energy of ~8–16 kJ/mol (2–4 kcal/mol). These numbers reveal an ~100-fold decrease in the binding affinity of the second ligand and are compatible with negative cooperative binding. Cooperativity occurs when a bound ligand affects the intrinsic binding affinity of other ligands. If subsequent binding affinity is greater due to a bound ligand, it is referred to as positive cooperativity, as observed in hemoglobin and described by the Monod-Wyman-Changeux model5. If the subsequent binding affinity is reduced, it is referred to as negative cooperativity, as observed with CTP synthetase38 and described by the Koshland-Nemethy-Filmer model4. Cooperative binding enhances or diminishes the response to a binding event by altering the bound fraction at a given ligand concentration providing tunability to increase or decrease the corresponding activity.
To investigate the mechanism of RhoA-activated actin polymerization by DIAPH1 the intrinsic and sequential binding affinities of RhoA and DAD for GDID (Fig. 1A) were measured and evaluated using a thermodynamic cycle (Fig. 1C), and the activation potential of several full-length DIAPH1 constructs (Fig. 1A) were examined for the ability to activate actin polymerization in vitro and in vivo. A model is proposed to explain how negative cooperative binding of effector ligands promotes actin polymerization activity. Given the homologous domain structure of DRFs (Fig. 1A) the model is generally applicable to all DRFs.
Results
DAD, GDID, and RhoA form a tripartite complex
To determine whether a stable ternary complex can exist between GDID, DAD and RhoA, the interaction between GDID and RhoA was assessed by using size exclusion chromatography, SEC, SDS-PAGE analysis and NMR spectroscopy. An equimolar mixture of ~35 μM GDID and RhoA eluted as a single peak at 14.4 mL, consistent with the formation of a protein complex (Supplementary Fig. 1A). SDS-PAGE gel analysis of the single eluted peak confirmed the presence of both GDID and RhoA (Supplementary Fig. 1A). 1H-15N Heteronuclear Single Quantum Coherence, HSQC, NMR spectroscopy of uniformly labeled DAD, [U-15N]-DAD, was used to identify any interactions between GDID, RhoA and DAD. The 1H-15N HSQC spectrum of 50 µM [U-15N]-DAD in the presence of 60 µM RhoA is identical to the spectrum of free [U-15N]-DAD indicating there is no interaction between RhoA and DAD (Supplementary Fig. 1B). The 1H-15N HSQC spectra for 50 µM [U-15N]-DAD in the presence of 60 µM GDID (Fig. 2A) or 60 µM DID (Supplementary Fig. 1C) exhibited identical large chemical shift changes relative to free [U-15N]-DAD indicating a highly similar structure for both complexes and that the GBD domain did not alter the DID-DAD interaction surface.
A Superimposed 1H-15N HSQC spectra of 50 µM [U-15N]-DAD in the absence (red) and presence (black) of 60 µM GDID (black). The DAD Q1207 cross peak, which is part of the DID-DAD binding interface, undergoes a large change in chemical shift between the bound and free states. B Superimposed 1H-15N HSQC spectra of 5 µM [U-15N]-DAD with 60 µM of RhoA and 50 µM GDID (black) and 5 µM [U-15N]-DAD with 19 µM of RhoA and 13 µM GDID (red). Free DAD is evident at the lower concentration of the RhoA-GDID complex. C NMR-generated isothermal titration of 5 µM [U-15N]-DAD binding to the RhoA-GDID complex and fit using a ‘total binding, accounting for ligand depletion’ model to estimate KdD/R of 27 ± 5 μM for DAD binding to the RhoA-GDID complex. D ELISA-generated isotherm of RhoA binding to 10 nM GDID fit to a “one site-total and nonspecific binding” model to estimate KdR of 8.9 ± 2.4 nM. E ELISA-generated isotherm of RhoA binding to 10 nM GDID-DAD fit to a ‘one site-total and nonspecific binding’ model to estimate KdR/D of 840 ± 200 nM. All values for ELISA experiments are represented as mean ± SD and were performed as N = 3 independent experiments. Values from individual experiments are shown as gray dots. Note that the decrease in the plateau values observed in the ELISA titrations are due to the Hook or Prozone effect in which high ligand concentrations result in a decrease in signal strength61.
No further chemical shift changes were observed for 50 μM [U-15N]-DAD when titrated with 70 μM of a RhoA-GDID complex suggesting that the binding sites for DAD and RhoA do not directly overlap in solution and that the tripartite complex in Fig. 1B represents a low population state unobserved by NMR.
The NMR spectra of samples of 5 μM [U-15N]-DAD in the presence of 50 μM of the RhoA-GDID complex showed only bound DAD, however at 15 μM RhoA-GDID, cross peaks consistent with both free and bound DAD became evident (Fig. 2B). The NMR sample containing 50 µM [U-15N]-DAD with 70 µM of RhoA-GDID was loaded onto an SEC column. A single peak eluted at 14.2 mL corresponding to a complex larger than that observed for RhoA or GDID alone (Supplementary Fig. 1D). SDS-PAGE analysis of the peak confirmed the presence of both RhoA and GDID (Supplementary Fig. 1E). Note that the ~3 kDa DAD would not be visible on the gel. The eluted peak was re-examined by NMR where the 1H-15N HSQC spectrum exhibited a subset of bound DAD peaks showing that DAD co-eluted with the RhoA-GDID complex (Supplementary Fig. 1F). Collectively these data confirm that DAD can form a stable tripartite complex with RhoA and GDID.
RhoA and DAD form a negatively cooperative complex with GDID
To assess the stability of the interacting components of the tripartite complex, SEC, NMR spectroscopy and enzyme linked immunosorbent saturation assays, ELISA, were used to measure binding affinities between GDID, RhoA and DAD. Thirty-five µM complexes of RhoA-GDID were mixed with increasing amounts of lyophilized DAD and subjected to SEC separation. Two peaks were observed, a “main peak” at ~14.2 mL for the ternary complex and a “shoulder” peak at 16.5 mL corresponding to displaced RhoA (Supplementary Fig. 1G). Increasing the DAD concentration led to a dose-dependent increase in the “shoulder” peak and an asymmetric broadening of the “main peak”, consistent with a dissociation process. The ratio of the absorbance of the “shoulder“ peak relative to the “main peak” was plotted as a function of DAD concentration and yielded a stoichiometric curve with a break point of ~35 µM, showing a 1:1 binding stoichiometry of DAD to the RhoA-GDID complex. Further analysis of the data using the ‘one site–total, accounting for ligand depletion’ regression model in GraphPad Prism, yielded a KdD/R of 10 ± 3.5 μM for DAD binding to the RhoA-GDID complex (Supplementary Fig. 1H).
To estimate the binding affinity of DAD to GDID in the presence of RhoA, 5 μM of [U-15N]-DAD was titrated against the RhoA-GDID complex, and changes were monitored using 1H-15N HSQC NMR. The magnetically inequivalent NεH amides of Q1207 of DAD exhibit distinct cross peaks for both free and bound states and were used to quantify the concentration of free and bound DAD (Fig. 2A). The data were fit to a model of ‘total binding, accounting for ligand depletion’ to yield a KdD/R of 27 ± 5 μM for DAD binding to the RhoA-GDID complex (Fig. 2C). This result is in general agreement with the SEC data and in agreement with a previous estimate of 29 µM at 20 °C for the release of DAD from DID titrated with RhoA26.
The affinities of RhoA for GDID and for the GDID-DAD complex were measured at room temperature by ELISA. An intrinsic dissociation constant, KdR, of 8.9 ± 2.4 nM was resolved for RhoA binding to GDID (Fig. 2D) in good agreement with the affinity determined for the binding of activated RhoA to mouse DIAPH1 at 20 °C, ~10 nM25,27,34. An apparent dissociation constant, KdR/D, of 840 ± 200 nM was resolved for the binding of RhoA to the GDID-DAD complex (Fig. 2E). The affinity of DAD for DID, KdD, was previously determined to be 270 ± 100 nM12.
When the binding affinities are examined in the context of the thermodynamic cycle (Fig. 1C) it is apparent that the binding of the second ligand is ~100-fold weaker than its intrinsic binding. Energetically, this amounts to a positive interaction free energy, consistent with negative cooperativity. To quantitate the formation of the ternary complex, a cooperativity factor, α, was incorporated into the binding equations (Fig. 1C and Supplementary Appendix 1)39. α is defined as the ratio of the intrinsic dissociation constant of a ligand for its binding site over the dissociation constant for subsequent binding to the binary complex:
where α > 1 denotes a positive cooperativity and α < 1 denotes negative cooperativity (Supplementary Appendix 1). Note that equality of the expression only occurs under saturation conditions, i.e. [DAD] » KdR or [RhoA] » KdD. The data show that the RhoA-GDID-DAD complex is characterized with a negative cooperativity factor of α = 0.01 ± 0.003 (Fig. 1C and Supplementary Appendix 1).
The DID-DAD interdomain persistent length modulates DIAPH1 activity
Only one study has analyzed RhoA activation and the DID-DAD interaction of DIAPH1 using full length constructs31. Studies to elucidate the structural and regulatory properties of Rho GTPases bound to DRFs used subdomains that do not accurately represent regulatory interactions between the N-terminal DID and C-terminal DAD domains23,24,26,27,28,29,31,34,40. Full-length DRFs contain both domains but vary in composition and length of the polypeptide chain between them (Fig. 1A). For example, the CC and FH1 domains of DIAPH1, DIAPH2 and DIAPH3 differ by ~100 residues. The unimolecular architecture of DRFs constrains the interacting domains to a maximum distance that places a lower limit on the effective concentration of unbound DAD, [DAD]eff, available for DID-DAD interactions. Shortening the polypeptide chain increases [DAD]eff which increases the population of the autoinhibited state and thus will decrease the potency of DRF activation by RhoA. To investigate if variations in the persistence length of the unstructured regions between DID and DAD modulate the activity of Diaphanous, a homolog of DIAPH1 (UniProt: O60610) called DIAPH1/2, containing CC/FH1 domains equal in length to that of DIAPH2 (UniProt: O60879) was created. The amino acid sequence of the CC/FH1 domains in DIAPH1/2 is ~140 residues shorter than in DIAPH1.
Using the method outlined by Hofmann and Schuler41 the radius of gyration, RG, was calculated for polypeptide chains from 100 to 300 amino acids long with a compactness variable, ν, ranging from 0.30 for the most compact globule state to 0.60 for an expanded coil state (Supplementary Appendix 2). The calculations showed an increase in RG as the number of amino acids increased and that compact globule states are less dependent on amino acid length than expanded coils (Fig. 3A). Monte-Carlo simulations were performed to calculate the average DID-DAD distance in DIAPH1 and DIAPH1/2 using a compactness variable of ν = 0.60 (Fig. 3B and Supplementary Appendix 2). DIAPH1 has a CC/FH1 length of ~280 residues while that of DIAPH1/2 is ~140 residues. The simulations showed a decrease of ~20 Å in persistence length for DIAPH1/2 corresponding to ~50% reduction in volume. These observations imply that shorter polypeptide chains have a larger [DAD]eff resulting in a greater population of autoinhibited Diaphanous and a corresponding decrease in RhoA induced actin polymerization.
A Plot of the radius of gyration, RG, versus the number of amino acids in a theoretical polypeptide. The parameter ν characterizes the dimensions of the amino acid chain where 0.60, red, represents an expanded coil and 0.30, maroon, represents the most compact globule state. B Monte-Carlo simulation using ν = 0.60 showing the theoretical persistence length between DID and DAD domains as a function of the number of amino acids: ~109 Å for the 285 amino acid DIAPH1 and ~90 Å for the 140 amino acids of DIAPH1/2. C Rates of actin polymerization for Diaphanous constructs DIAPH1, D1, DIAPH1/2, D1/2 and DIAPH1ΔDAD, D1ΔD, in the presence of RhoA, +R, compared to actin alone, A. Initial rates of polymerization, dF/dt, were determined by fitting the first 11 data points to a linear equation yielding slopes of 62 ± 6 AFU*s−1 for A (red), 66 ± 6 AFU*s−1 for D1 (green), 62 ± 7 AFU*s−1 for D1/2 (purple), 85 ± 5 AFU*s−1 for D1 + R (white), 75 ± 4 AFU*s-1 for D1/2 + R (gray) and 104 ± 7 AFU*s−1 for D1 ΔD (orange). D1ΔD represents 100% activation and D1 or D1/2 represent 0% activation, respectively. Statistical p-values at 95% confidence are shown on the graph and all values are represented as mean ± SD derived from N = 3 independent experiments.
To examine the effect of DIAPH1 architecture and [DAD]eff on RhoA activation in vitro, actin polymerization assays were performed using pyrene-labeled actin, DIAPH1, DIAPH1/2 and a DIAPH1 construct lacking the DAD domain, DIAPH1ΔDAD. Purification of the DIAPH1 constructs revealed that DIAPH1ΔDAD has the largest hydrated radius due to the inability to form the autoinhibited state followed by full length DIAPH1 and DIAPH1/2 (Supplementary Fig. 2). The rate of actin polymerization increased with increasing RhoA for DIAPH1 (Supplementary Fig. 3A, B) and showed no significant dose dependence for DIAPH1/2 (Supplementary Fig. 3C, D). DIAPH1 showed a dose-dependence that reflects a smaller [DAD]eff than that of DIAPH1/2, and neither activated actin polymerization to the level of constitutively active DIAPH1ΔDAD. Activation by 10 μM RhoA significantly increased both DIAPH1 and DIAPH1/2 induction of actin polymerization by 51 ± 25% and 36 ± 20%, respectively (Fig. 3C and Supplementary Fig. 3E), which correspond to a [DAD]eff of 30 ± 20 µM and 50 ± 30 µM for DIAPH1 and DIAPH1/2, respectively (Supplementary Appendix 3). This observation is consistent with the hypothesis that the shorter polypeptide chain of DIAPH1/2 results in a larger [DAD]eff that reduces RhoA activation of polymerase activity by maintaining a higher population of autoinhibited DIAPH1.
Autoinhibition is required for normal actin distributions
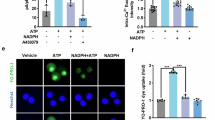
The dissociation constant for sequential binding of activated RhoA and DAD for GDID increases 100-fold for the second ligand. Since the dissociation constant for DAD binding to the RhoA-GDID complex is approximately the same as [DAD]eff, 27 μM vs 30 μM, about half of the intracellular population of DIAPH1 will be in the autoinhibited state requiring a concentration of 840 nM RhoA to bind to GDID. Since the typical cellular concentration of RhoA is ~3 µM42, it is expected that RhoA can bind to DIAPH1 under physiological conditions. To examine the effect of autoinhibition on the localization and activation of DIAPH1 by RhoA, the intracellular distribution of F-actin and GFP-DIAPH1 or GFP-DIAPH1ΔDAD was assessed in AD293 cells. RhoA was activated in vivo using the RhoA activator ACTII, also known as CN03, which abolishes GTPase activity resulting in constitutively active RhoA43. Under basal conditions, the distribution of DIAPH1 was primarily cytosolic with a membrane to cytosol ratio of 0.45 ± 0.01 (Fig. 4A and Supplementary Fig. 4) and the distribution of F-actin was primarily membranous with a ratio of 1.33 ± 0.06 (Fig. 4B). CN03 activation of RhoA significantly increased the membranous distribution of both DIAPH1, 0.53 ± 0.02, and actin, 1.71 ± 0.06, as expected since DIAPH1 is primarily activated when membrane bound. Induction of actin polymerization by DIAPH1 changed the morphology of the cells from ellipsoidal with some filopodia to elongated with extensive filopodia (Fig. 4C).
A Membrane to cytosol ratio of CFP-DIAPH1 fluorescence. The sample sizes for DIAPH1 (D1, green circles), DIAPH1 + CN03 (D1 + CN03, red circles), DIAPH1ΔDAD (D1ΔD, cyan squares) and DIAPH1ΔDAD + CN03 (D1ΔD + CN03, purple squares) are 31, 44, 51 and 105, respectively. B Membrane to cytosol ratio of phalloidin fluorescence, which binds to F-actin. The sample size for D1 (green circles), D1 + CN03 (red cricles), D1ΔD (cyan squares) and D1ΔD + CN03 (purple squares) are 31, 44, 51 and 105, respectively. The analysis uses the ratio of intensities of membrane and cytosol, which decreases the bias caused by alternative actin filaments staining62. C Confocal microscope images of AD293 cells transfected with CFP-labeled D1 and D1ΔD. RhoA activator CN03 was at 1 mg/mL. Cells were stained with Alexa Fluor568 Phalloidin to image filamentous actin and Hoechst 33258 to image the nuclei. Scale bar is 10 µm. The significant cytosolic fraction of overexpressed DIAPH1 after CN03 activation of RhoA or of constitutively active DIAPH1ΔDAD suggests that there are limited membrane binding sites for DIAPH1 as observed previously30. The height of the bar represents the mean and the error bars are ± SEM and statistical p-values at 95% confidence are shown on the graph.
The membrane to cytosol ratio of DIAPH1ΔDAD was significantly greater than that of DIAPH1, 0.60 ± 0.02, but membrane-associated F-actin was significantly reduced, 0.82 ± 0.02. The presence of CN03 had no effect on the membrane to cytosol ratio of either DIAPH1 or F-actin as expected since DIAPH1ΔDAD is fully activated (Fig. 4A, B). Cells containing GFP- DIAPH1ΔDAD exhibited the same elongated morphology in both the absence and presence of CN03 (Fig. 4C). Although both DIAPH1 and DIAPH1ΔDAD are cytosolic with DIAPH1 almost entirely in an autoinhibited conformation, the affinity of activated RhoA, KdR, for DIAPH1ΔDAD is 100 times greater than for autoinhibited DIAPH1, thus there is more membrane-bound DIAPH1ΔDAD than DIAPH1. In addition, DIAPH1ΔDAD binds to the plasma membrane in a RhoA-independent manner further increasing its localization to the plasma membrane44,45. Because DIAPH1ΔDAD is constitutively active, actin polymerization can occur at the plasma membrane as well as in the cytosol in the absence of RhoA resulting in more cytosolic F-actin and a subsequent decrease in the membrane/cytosol ratio of F-actin as compared to CN03-activated DIAPH1 (Fig. 4B, C). These observations are consistent with the idea that the autoinhibited state is critical for proper membrane localization of DIAPH1 and F-actin structures, and that the negatively cooperative RhoA-DIAPH1 complex provides a mechanism for maintaining the autoinhibited state.
ctRAGE increases the affinity of RhoA for DIAPH1
Localization and regulation of DIAPH1 activity is accomplished in part by interactions with cytoplasmic and membrane bound factors other than RhoA37,46,47,48,49. Examples of such heterotropic regulation are the binding of Flightless-I, Fli-I, to the DAD domain of DIAPH1 and the binding of IQ motif containing GTPase activating protein-1, IQGAP, to the DID domain, both of which enhance RhoA-mediated activation of actin filament nucleation in vitro37,47 by weakening the DID-DAD interaction 10- to 30-fold37. RhoA binding to Fli-I- or IQGAP-bound DIAPH1 restores the intrinsic affinity between N- and C-terminal constructs37. Conversely, binding of liprin-α3 to mouse DIAPH1 decreases both RhoA and DAD binding affinity and modulates activation of actin polymerization suggesting that a compensatory regulation mechanism is also active in cells49. The linked functionality of these heterotropic effectors to regulate DIAPH1 activity can be described by extending the thermodynamic cycle to include additional ligands (Supplementary Fig. 5 and Supplementary Appendix 4).
Another example of heterotropic regulation of DIAPH1 activity is the C-terminal cytoplasmic domain of the receptor for advanced glycation end products, ctRAGE. The binding of ctRAGE to the FH1 domain of DIAPH1 enhances RAGE signaling; the FH1 domain of DIAPH1 is far removed from the RhoA-GDID-DAD binding sites14,50. To examine whether ctRAGE affects RhoA binding and activation of DIAPH1, in vitro actin polymerization assays were performed. 3 µM of ctRAGE had an insignificant effect on actin polymerization, but 3 µM of RhoA significantly increased actin polymerization vs. that of DIAPH1 alone (Fig. 5A and Supplementary Fig. 6). A significant dose dependent increase was observed when equimolar amounts of ctRAGE and RhoA were used (Fig. 5A and Supplementary Fig. 6). To determine if ctRAGE binding alters the affinity of RhoA for DIAPH1, ELISA titrations of RhoA binding to DIAPH1 in the absence and presence of 3 μM ctRAGE were performed. RhoA binding to DIAPH1 yielded a KdR/D of 450 ± 40 nM (Fig. 5B), which is twice as strong as RhoA binding to the GDID-DAD monomeric subunit. This datum yields a cooperativity factor of α = 0.01 ± 0.005 for the RhoA-DIAPH1 complex, which is identical to the cooperativity factor determined for the monomeric constructs (Supplementary Appendix 5). RhoA binding to DIAPH1 in the presence of ctRAGE yielded a KdR/D of 160 ± 20 nM (Fig. 5C), suggesting that ctRAGE binding increased the affinity of RhoA for DIAPH1 by threefold. This change in affinity corresponds to a decrease in [DAD]eff to 6 ± 3 µM (Supplementary Appendix 3) and provides a functional linkage between ctRAGE, RhoA binding and DIAPH1 polymerase activity.
A Rates of actin polymerization for Diaphanous constructs DIAPH1, D1, in the presence of 3 µM ctRAGE, 3 C, 3 µM RhoA, 3 R, or 1, 2, or 3 µM of both ctRAGE and RhoA, 1CR, 2CR, 3 CR, respectively, compared to actin alone, A. Initial rates of polymerization, dF/dt, were determined by fitting the first 13 data points to a linear equation yielding slopes of 11.1 ± 0.4 AFU*s-1 for A (red), 11.6 ± 0.3 AFU*s−1 for D1 (beige), 13.1 ± 1.0 AFU*s−1 for D1 + 3C (light blue), 14.6 ± 0.8 AFU*s-1 for D1 + 3 R (dark blue), 17.8 ± 0.7 AFU*s−1 for D1 + 1RC (light green), 20.6 ± 0.8 AFU*s-1 for D1 + 2RC (green), and 22.0 ± 0.9 AFU*s−1 for D1 + 3RC (dark green). Statistical p-values at 95% confidence are shown on the graph. B ELISA-based isotherm of RhoA binding to DIAPH1. The data were fit to “one site-specific binding” curve to estimate the dissociation constant, KdR/D, for the interaction. C ELISA-based binding isotherm of RhoA binding to DIAPH1 in the presence of ctRAGE. The data were fit to a “one site-specific binding” curve to estimate the dissociation constant, KdR/D, for the interaction in the presence of ctRAGE. All values are represented as the mean ± SD and derived from N = 3 independent experiments. Values from individual experiments are shown as gray dots.
Discussion
Spatiotemporal control of actin dynamics is essential for proper cellular function. The formin subfamily of DRFs uses autoinhibition that is relieved by Rho GTPases to modulate activity. Earlier studies suggested that RhoA binds to mouse DIAPH1 with a Kd of ~10 nM and overlaps with the DAD binding site resulting in direct displacement and no tripartite complex. It was demonstrated that RhoA, GDID and DAD form a ternary complex at saturating concentrations by using NMR spectroscopy, SEC and ELISA (Fig. 1 and Supplementary Figs. 1 and 2). Analysis of the thermodynamic linkage between the sequential binding affinities of RhoA and DAD to GDID resolved a cooperativity factor of ~0.01 confirming that the ternary complex is negatively cooperative.
Applying the binding affinities from this work and the RhoA and DAD concentrations from the fluorescence polarization experiments of Rose et al.27 and Lammers et al.26, our model predicts that 2–4% of DAD should remain bound to the GDID-RhoA complex in agreement with previous observations. In the context of full-length DIAPH1 and 10 µM RhoA, the model predicts an activation of ~51%, comparable to the level of activation observed here and by Maiti et al.31 (Fig. 3C and Supplementary Appendix 3). Pathological mutations of DIAPH1 observed in autosomal hearing loss16,17,18,20 and thrombocytopenia20,21, e.g. p.Arg1213X (KdD ~1.5 µM26), p.Ala1210Glyfs*31 (KdD ≥ 160 µM26) and p.Ala1210Serfs*31 (KdD ≥ 160 µM26), are predicted to have DIAPH1 activities of ~80% and ≥99% at 10 µM RhoA, consistent with the observed abnormalities in stereocilia in hair cells and platelet budding in proplatelets.
A general RhoA based regulatory mechanism that incorporates negative cooperativity for DIAPH1 activation of actin polymerization is proposed (Fig. 6A). Following GDP to GTP exchange, i.e. RhoA activation (i), the membrane-bound RhoA-GTP complex binds to autoinhibited DIAPH1 with low affinity, KdR/D, to partially activate DIAPH1 and localize it to the membrane (ii). Activated DIAPH1 relieves the negative cooperative interaction via DAD dissociation allowing RhoA to bind to GDID with high affinity, KdR (iii), while concurrently inducing actin polymerization (iv). Induction of actin polymerization is subject to further modulation by heterotropic regulatory molecules such as Fli-1, IQGAP, and ctRAGE (v). The linked functionality of these effectors enhances RhoA activation of DIAPH1 by increasing the affinity of RhoA binding or decreasing [DAD]eff (Fig. 5). The strength of the unimolecular interaction that sustains bound DAD is determined by [DAD]eff, which is dependent on the persistence length of the linker between DID and DAD (Fig. 3). [DAD]eff is sufficiently high to maintain a population of autoinhibited DIAPH1, even at saturating concentrations of activated RhoA resulting in less than full induction of actin polymerization. The hydrolysis of GTP by RhoA (vi) facilitates its dissociation from DIAPH1(vii) allowing the formation of an encounter complex12 between DID and DAD, and enabling DAD to bind to DID with high affinity, KdD, restoring the autoinhibited state (viii). Because of the homologous domain structure of Diaphanous Related Formins (Fig. 1A) this mechanism should be applicable to all DRFs.
A Mechanism of DIAPH1-RhoA localization and activation (i-viii). Color scheme of the DIAPH1 domains is as in Fig. 1A. RhoA-GDP is pink and RhoA-GTP is purple. To localize at the membrane RhoA is farnasylated and unphosphorylated. Other cytoskeleton remodeling proteins, such as profilin, cofilin1/ADF, or cross-linkers that contribute to the described mechanism are omitted for clarity; their roles in this model are a subject for future investigations. B Three-dimensional plot showing KdD/R as a function of α and [RhoGTPase] for DRFs. α and [RhoGTPase] are on a logarithmic scale. The line represents α = 1 and has zero curvature in the [RhoGTPase] dimension. The plot was constructed using KdD and KdR derived from the RhoA-DIAPH1 interaction. Derivations are in Supplementary Appendix 1. C Three-dimensional plot of unbound DAD, 1-FDB, as a function of RhoGTPase concentration and [DAD]eff. [DAD]eff and [RhoGTPase] are on a logarithmic scale. The gradient from deep blue to yellow shows increasing DIAPH1 activation. Derivations are in Supplementary Appendix 5. The plot was constructed using KdR, KdD and α derived from the RhoA-DIAPH1 interaction.
A three-dimensional plot showing the thermodynamic surface for DAD binding to a RhoGTPase-bound DRF as a function of α and [RhoGTPase] is shown in Fig. 6B. KdD/R values increase hyperbolically as RhoGTPase concentrations increase for α < 1, which is consistent with greater DAD concentrations yielding stable RhoA-GDID-DAD complexes for DIAPH1. Thus, negative cooperativity ensures gradual activation over a broad range of RhoGTPase concentrations. A three-dimensional plot showing the fraction of activated DRF (unbound DAD) as a function of [RhoGTPase] and [DAD]eff when α = 0.01 is shown in Fig. 6C. The population of activated DRF increases with increasing [RhoGTPase] and decreasing [DAD]eff, as expected. DRF activation begins to decrease when [DAD]eff > KdD/R, which is governed by the intrinsic affinity of the DID-DAD interaction and linker length. The magnitudes of KdD and [DAD]eff set the maximum limit of activation that can be attained by increasing amounts of RhoGTPase. Importantly, full activation would require [DAD]eff to decrease to values not permitted by the intramolecular constraints imposed by the structure of the DRF.
The model proposed for DRF activation entails multifarious specificity14,37,47,49 by combining complex linkages between ligand binding and the inherent configuration of unstructured linkers to afford tight regulation of formin activity (Fig. 6 and Supplementary Fig. 5). By exploiting negative cooperativity, DRFs maintain a reserve of inactivated DIAPH1 for other cellular processes. And unlike the sharp, switch-like functional transitions that accompany small changes in ligand concentration in allosteric systems that make use of positive cooperative binding, negative binding cooperativity in DRFs enables more gradual responses to cellular processes that require prolonged and sustained regulation, such as cell migration51, phagocytosis52, and cytokinases53.
Methods
Chemicals and reagents
All chemicals used were of molecular biology grade or better. Rho Activator II CN03 was obtained from Cytoskeleton, Inc. (Denver, CO). Alexa Fluor™ 568 Phalloidin was purchased from Fisher Scientific. Guanosine triphosphate mimic, guanosine 5′-[β,γ-imido]triphosphate, GPPNP, was purchased from Sigma-Aldrich. The Hoechst 33258 nuclear stain was purchased from Invitrogen.
RhoAQ63L expression and purification
Plasmid pGEX-4T3-RhoA-Q63L, which confers ampicillin resistance and expresses N-terminal GST tagged RhoAQ63L (21,753 Da) was purchased from Addgene (Plasmid #12961). The Q to L mutation inhibits GTPase activity to maintain RhoA in an active GTP-bound state. This construct is used throughout and referred as RhoA. The plasmid was transformed into E. coli strain BL21(DE3) and cultured overnight at 37 °C in Luria Broth, LB, containing 35 µg/mL of carbenicillin, LB-Cn, and transferred to 1 L of LB-Cn and incubated at 37 °C until the A550 reached ~0.70. Overexpression of the GST-RhoA fusion protein was induced with 100 µM isopropyl-β-D-thiogalactopyranoside, IPTG, and incubated for 20 h at 18 °C. Cells were centrifuged at 4000 g, resuspended in lysis buffer, 25 mM Tris-HCl, pH 7.5, 1 M NaCl, 5 mM MgCl2 and 2 mM β-mercaptoethanol containing an EDTA-free protease inhibitor tablet (Roche) and sonicated with a Model 250 Digital Sonifier (Branson) at 40% amplitude using 2.5-min cycles of 0.3 s pulses interrupted with 1.0 s pauses. The lysate was centrifuged for 30 min at 20,000 g and 4 °C. The supernatant was loaded into a glutathione Sepharose affinity column (Cytiva) pre-equilibrated with lysis buffer. The column was washed with ten column volumes of lysis buffer and the GST-RhoA fusion protein was eluted using elution buffer, 50 mM Tris-HCl, pH 7.5, 150 mM NaCl and 10 mM reduced glutathione. The eluted protein was dialyzed in thrombin cleavage buffer, 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 10 mM MgCl2 and 2.5 mM CaCl2, and cleaved for 1 h at room temperature using a thrombin cleavage kit in which the thrombin is bound to beads (Sigma). The resulting slurry was washed repeatedly with thrombin cleavage buffer to recover the cleaved proteins. RhoA was separated from GST by using a SuperdexTM 200 Increase 10/300 GL size exclusion column (Cytiva) calibrated using ~1 mg/mL Gel Filtration Standards (Bio-Rad). Samples were run in 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl and 0.01% NaN3 at 0.5 mL/min. Two peaks eluted: residual GST at 15.0 mL (51 kDa) and monomeric Rho at 16.8 mL (~19 kDa). EDTA was added to the eluted fractions to a final concentration of 1 mM to reduce thrombin activity. The fractions were pooled, concentrated using 10 kDa MWCO centrifugal filters (Millipore) and stored in SEC buffer, 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl and 0.01% NaN3, at 4 °C. The purity was estimated at >95% on 12% SDS-PAGE and the concentration of RhoA was determined by absorbance at 280 nm using a molar extinction coefficient of 21,805 M−1 cm−1 54. To prepare activated RhoA, samples were incubated for 1 h at 4 °C with a 100 molar excess of GPPNP, a non-hydrolyzable GTP analog, in SEC buffer containing 2 mM MgCl2.
GDID expression and purification
Expression plasmid pET-28a(+)-GDID, which confers kanamycin resistance and expresses N-terminal His-tagged GDID residues 83–380 from human DIAPH1 (Uniprot O60610; 35,420.71 Da), was purchased from Genscript. pET28a(+)-GDID was transformed into E. coli strain BL21(DE3) and grown overnight at 37 °C in 100 mL of LB containing 35 µg/mL of kanamycin (LB-Kn). The culture was transferred to 1 L of LB-Kn and incubated at 37 °C until the OD550 reached ~0.90. Over-expression of GDID was induced with 100 µM IPTG and incubated for 4 h at 37 °C. Cells were centrifuged at 4000 g, resuspended in lysis buffer, 20 mM HEPES, pH 8.0, 1 M NaCl, 12.5% w/v sucrose, 4 M urea and 10 mM β-mercaptoethanol, containing an EDTA-free protease inhibitor tablet (Roche) and sonicated with a Model 250 Digital Sonifier (Branson) at 40% amplitude using 2.5-min cycles of 0.3 s pulses interrupted by 1.0 s pauses. The lysate was centrifuged for 40 min at 20,000 g and 4 °C. The supernatant was loaded into a nickel-nitriloacetic acid agarose, Ni-NTA, affinity column (Qiagen) pre-equilibrated with lysis buffer. The partially denatured protein bound to the column was refolded by washing with five column volumes of refolding buffer, 10 mM HEPES, pH 8.0, 300 mM NaCl and 10 mM imidazole. The column was washed with five column volumes of wash buffer, 10 mM HEPES, pH 8.0, 300 mM NaCl and 20 mM imidazole, and the refolded GDID was eluted with elution buffer, 10 mM HEPES, pH 8.0, 300 mM NaCl and 250 mM imidazole. EDTA was added to a final concentration of 10 mM and the eluant was dialyzed into ion exchange chromatography buffer A, 20 mM NaH2PO4/Na2HPO4, pH 8.1 and 100 mM NaCl, prior to loading onto a Hi-Trap Q-XL anion exchange column (Cytiva). A linear gradient of buffer B, 20 mM NaH2PO4/Na2HPO4, pH 8.1 and 1 M NaCl, was used to elute the protein at a conductivity of ~40 mS/cm. The eluted GDID was further purified using a SuperdexTM 200 Increase 10/300 GL size exclusion column (Cytiva) calibrated using ~1 mg/mL Gel Filtration Standards (Bio-Rad). The sample was run in SEC buffer at 0.5 mL/min and eluted as a monomer at 14.8 mL (~56 kDa). EDTA was added to the eluted fractions to a final concentration of 1 mM. The fractions were pooled, concentrated using conical centrifugation tubes and stored in SEC buffer at 4 °C. The purity was estimated at >95% by 12% SDS-PAGE, and the concentration of GDID was determined by absorbance at 280 nm using a molar extinction coefficient of 13,200 M−1 cm−1 54.
DAD cloning, expression, and purification
Creation of the plasmid used for the DAD peptide was previously described12. This DAD peptide has a conserved substitution, M1199L, that does not affect DID-DAD binding. The plasmid was transformed into E. coli BL21(DE3) for over-expression and purification of both labeled and unlabeled peptide. To purify DADM1199L, the bacterial cell pellet was resuspended in 50 mM Tris-HCl, pH 7.2, 1% w/v Triton-X-100 and 1 mM EDTA, sonicated as described above and centrifuged for 30 min at 10,000 g and 4 °C. The pellet was successively washed, sonicated and centrifuged using 50 mM Tris-HCl, pH 7.2, 2% w/v Triton-X-100 and 1 mM EDTA, followed by 25 mM Tris-HCl, pH 7.2 1 M NaCl and 0.5 mM EDTA, and finally 50 mM Tris-HCl, pH 7.2. The final pellet was dissolved in denaturing buffer, 50 mM NaH2PO4/Na2HPO4, pH 8.3 containing 6 M guanidinium chloride, and clarified by centrifugation for 40 min at 10,000 g and 4 °C. The supernatant was incubated with Ni-NTA beads overnight for batch binding. The beads were packed into a column and washed once with denaturing buffer. The column was washed with 5 volumes of 50 mM NaH2PO4/Na2HPO4, pH 7.0 containing 6 M urea followed by 50 mM NaH2PO4/Na2HPO4, pH 6.4 and 6 M urea. Elution was carried out using 50 mM NaH2PO4/Na2HPO4, pH 3.6 and 6 M urea. Fractions containing TrpL-DADM1199L were pooled, dialyzed into water, and lyophilized. Cleavage of the TrpL leader sequence was performed by dissolving the TrpL-DADM1199L fusion protein in 70% formic acid with a 100 molar excess of cyanogen bromide and incubated for 1.5 h at 25 °C. The cleaved products were dried in vacuo, loaded onto a C18 column (Agilent ZORBAX 300SB-C18) for high performance liquid chromatography (HPLC). Cleaved products were resolved using a gradient of 0%-90% acetonitrile in 0.1% trifluoroacetic acid. DADM1199L eluted first and was collected, lyophilized and stored at −20 °C. The concentration of DADM1199L was determined by integrating HPLC chromatogram traces at 260 nm, the approximate wavelength of maximal absorption for phenylalanine residues, and comparing them to wild-type DAD peptide reference solutions.
ctRAGE expression and purification
A plasmid construct expressing human TrpL-ctRAGE was used50. pTM7-ctRAGE was transformed into E. coli strain BL21(DE3) and grown overnight at 37 °C in 100 mL of LB-Kn. The culture was transferred to 1 L of LB-Kn and incubated at 37 °C until the OD550 reached 0.90. To overexpress unlabeled TrpL-ctRAGE, isopropyl β-D-1-thiogalactopyranoside, IPTG, was added to a final concentration of 1 mM and the cells were incubated for 20 h at 37 °C. For isotopic labeling, cells were harvested by centrifugation at 4000 g and resuspended in minimal, M9, medium, 5 mM Na2HPO4, 2.5 mM KH2PO4, pH 7.0, 1 mM NaCl, 2 mM MgSO4, 0.1 mM CaCl2 and 1 mg/L thiamine hydrochloride, supplemented with 35 µg/mL of kanamycin, M9-Kn. For [U-15N] labeling, 1 g/L of 15N-ammonium chloride (Sigma Aldrich) was used as the sole nitrogen source. For [U-15N,13C] labeling 2 g/L of 13C-glucose (SigmaAldrich) was used as the sole carbon source. Overexpression of labeled TrpL-ctRAGE constructs was induced with 1 mM IPTG and proceeded for 20 h at 37 °C. Cells were centrifuged at 4000 g, resuspended in denaturing lysis buffer, 20 mM NaHPO4/Na2HPO4, pH 8.0, 8 M urea, and 1 mM DTT containing an EDTA-free protease inhibitor tablet (Roche), and sonicated with a Model 250 Digital Sonifier (Branson) at 40% amplitude using 2.5-min cycles of 0.3 s pulses interrupted by 1.0 s pauses. The lysate was centrifuged for 40 min at 20,000 g and 4 °C. The supernatant was loaded into a nickel-nitriloacetic acid agarose, Ni-NTA, affinity column (Qiagen) pre-equilibrated with equilibration buffer, 20 mM NaHPO4/Na2HPO4, pH 8.0, 8 M urea. The denatured constructs bound to the column were washed with five column volumes of equilibration buffer. The column was washed with five column volumes of wash buffer, 20 mM NaHPO4/Na2HPO4, pH 6.5, 8 M urea, and eluted with elution buffer, 20 mM NaHPO4/Na2HPO4, pH 3.6, 8 M urea. The eluant was dialyzed into water and lyophilized. Cleavage of the TrpL sequence was accomplished by dissolving the fusion protein in 70% formic acid with a 100 molar excess of cyanogen bromide and incubating for 1.5 h at 25 °C. The mixture was rotary evaporated to dryness, resuspended in minimal water, dialyzed and lyophilized. The lyophilized powder was dissolved in buffer A, 20 mM NaHPO4/Na2HPO4, pH 7.0, centrifuged and clarified prior to loading onto a Hi-Trap Q-XL anion exchange column (Cytiva). A linear gradient of buffer B, 20 mM NaH2PO4/Na2HPO4, pH 7.0, 1 M NaCl, was used to elute the protein at a conductivity of 30 mS/cm. The purity of ctRAGE was found to be >99% by mass spectrometry and the concentration was determined by absorbance using a molar extinction coefficient of 5,500 M−1 cm−1 at 280 nm54.
DIAPH1, DIAPH1/2, and DIAPH1ΔDAD expression and purification
Human Embryonic Kidney cells, HEK293T, were purchased from American Type Culture Collection (ATCC, Cat. No. CRL-3216) from clone 17 at passage 3 and tested for mycoplasma by the manufacturer. Cells were grown in DMEM, low glucose, GlutaMAX Supplement, pyruvate medium (Gibco), non-essential amino acids (Gibco, Cat. No. 11140035), and 10% horse serum. Cultures were grown on batches of ten 10 cm plates (Corning, Millipore) to 65% confluence before transfecting. Cells were transfected using 1 μg of the desired DIAPH1 construct per 1 mL of cell culture. Transfection cocktails were prepared by dissolving plasmids in Tris-Borate-EDTA pH 8.3 at 1 mg/mL and added to Opti-MEM Reduced Serum Medium (Gibco) to a final concentration of 10 µg/mL. Polyethyleneimine was added to a final concentration of 30 µg/mL and incubated for 10 min at 37 °C. One-hundred µL of transfection cocktail was slowly added to each mL of cell culture and the cells allowed to express for 40–48 h. Cells were harvested via centrifugation for 2 min at 200 g and 4 °C, washed twice with phosphate buffered saline PBS, 10 mM Na2HPO4/KH2PO4, pH 7.4, 137 mM NaCl, 2.7 mM KCl, and stored with 1 tablet of mini-Roche EDTA-free protease inhibitor at −80 °C. For purification, cells were thawed at 25 °C, resuspended in 3 mL of PBS pH 8.0 per gram of wet cell pellet, WCP, and adjusted to a final composition of 1% Triton X-100, 2 mM MgCl2. One uL of 26.5 units of Benzonase Nuclease (Sigma Aldrich) was added per gram of WCP. Lysis was accomplished via five freeze-thaw cycles alternating between 37 °C and liquid nitrogen. The lysate was adjusted to a final volume of 10 mL in 10 mM Na2HPO4, KH2PO4, pH 8.0, 300 mM NaCl, 2.7 mM KCl, 10 mM imidazole and 1 mM EDTA, and clarified in a Beckman Coulter Optima LE-80k ultracentrifuge in a 90Ti rotor for 30 min at 99,000 g at 4 °C. The clarified lysate was filtered through a 0.45 um polyethersulfone membrane (Thermo Scientific), loaded onto a 5 mL PureCube 100 Ni-INDIGO immobilized metal affinity column (Cube Biotech) and washed with buffer A, 50 mM NaH2PO4/Na2HPO4, pH 8.0, 300 mM NaCl, 2 mM EDTA and 0.01%NaN3, until the absorbance at 260 nm and 280 nm exhibited no changes followed by washing with 20% buffer B, 50 mM NaH2PO4/Na2HPO4, pH 7.0, 500 mM imidazole, 300 mM NaCl, 2 mM EDTA and 0.01% NaN3, until the absorbance at 260 nm and 280 nm baselined. All DIAPH1 constructs were eluted using 15 mL of 100% buffer B. The eluted protein was concentrated using 100 kDa MWCO centrifugal filters (Millipore) to ≤500 µL. DIAPH1, DIAPH1/2, and DIAPH1ΔDAD were further purified by diluting to a final volume of 500 µL in SEC running buffer, 20 mM HEPES, pH 7.4, 50 mM KCl, 10% glycerol, 0.01% sodium azide, 1 mM EDTA, and run on a Sepharose 6 10/300 column (Cytiva) size exclusion column at 0.5 mL/min. DIAPH1, DIAPH1/2, and DIAPH1ΔDAD eluted at 13.3 mL, 14.4 mL, and 12.7 mL, respectively. Apparent masses were determined using ~1 mg/mL Gel Filtration Standards (Bio-Rad). The purity of DIAPH1 (141.3 kDa), DIAPH1/2 (114.2 kDa), and DIAPH1ΔDAD (133.0 kDa) were found to be >95% on 8% SDS-PAGE. The concentrations of DIAPH1, DIAPH1/2, and DIAPH1ΔDAD were determined by absorbance at 280 nm using molar extinction coefficients of 42580 M−1 cm−1, 42455 M−1 cm−1, and 42455 M−1 cm−1, respectively54.
Size exclusion chromatography
The activated RhoA-GDID complex was prepared by mixing RhoA, GDID and GPPNP in 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl, 1 mM EDTA, 2 mM MgCl2 and 0.5 mM DTT to final concentrations of ~40 μM RhoA, ~35 μM GDID and 4 mM GPPNP. Five hundred µL samples were loaded onto a Superdex 200 10/30 column (Cytiva) column and eluted using 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl and 0.5 mM DTT at a flow rate of 0.5 mL/min. 100 µL aliquots of ~40 μM of the RhoA-GDID complex were mixed with lyophilized DAD to yield final DAD concentrations ranging from 0-100 μM. The samples were loaded onto a Superdex 200 10/30 column (Cytiva) and eluted using, 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl and 0.5 mM DTT at a flow rate of 0.5 mL/min.
NMR spectroscopy
Heteronuclear Single Quantum Coherence, 1H-15N HSQC, NMR spectra were acquired using a 600 MHz Bruker Ascend spectrometer at 298 K equipped with an ultrasensitive cryoprobe. Experiments were performed with Watergate water suppression55, processed using TopSpin version 3.2 or 3.6 (https://www.bruker.com/en/products-andsolutions/mr/nmr-software/topspin.html), and assigned using Computer Aided Resonance Assignment, CARA56.
To determine KdD/R, solutions of 80 μM RhoA and 70 μM GDID were mixed with 1 mM GPPNP in 20 mM NaH2PO4/Na2HPO4, pH 7.0, 100 mM NaCl, 1 mM EDTA, 2 mM MgCl2 and 0.5 mM DTT. Lypohilized [U-15N]-DAD was added to a final concentration of 5 μM and analyzed via 1H-15N HSQC. The solution was serially diluted with 5 μM [U-15N]-DAD in the same buffer. The ratio of Q1207 Nε for bound DAD relative to the total was plotted versus the concentration of the GDID-RhoA complex. Since the concentration of DAD was comparable to the measured Kd, the data were fit to a “total binding, accounting for ligand depletion” model using GraphPad Prism, v 9.0 (GraphPad), to estimate the dissociation constant.
Chemical Exchange Saturation Transfer, CEST, NMR spectroscopy was performed using 40 µM [U-15N]-DAD with 60 µM DID and 65 µM RhoA in 20 mM NaH2PO4/Na2HPO4, pH 7, 100 mM NaCl, 1 mM EDTA, 2 mM MgCl2, 0.5 mM DTT. Experiments were performed on a 600 MHz Bruker Ascend spectrometer using an 15N saturation pulse of 400 ms at both 10 Hz and 20 Hz. The saturation frequency ranged from 105 ppm to 131 ppm in 0.10 ppm intervals. All residues observed exhibited a single dip indicating no low population “dark” states.
Actin polymerization assays
Actin polymerization assays were performed using a commercially available Actin Polymerization Biochem Kit (Cytoskeleton). Samples contained 2 μM of 5% pyrene-labeled actin and 10 nM of SEC purified DIAPH1, DIAPH1/2 or DIAPH1ΔDAD (Supplementary Fig. 2), in the absence and presence of 10 μM RhoA57. Profilin was not included in the assay to measure basal formin activity9,31. The assays were performed in a Greiner Black Round Bottom Polypro 96-well plate in 60 μL volumes. Reactions were induced using 6 μL of 10x actin polymerization buffer, 500 mM KCl, 20 mM MgCl2, 50 mM guanidine carbonate and 10 mM ATP, and the fluorescence was collected at 50 s intervals using a Synergy H1 plate reader (BioTek) at 25 °C with 365 nm and 405 nm for the excitation and emission wavelengths, respectively. Assay data were plotted and analyzed using GraphPad Prism version 10.4.1 for Windows. The rates of the actin polymerization were analyzed using pseudo-first order kinetics due to the ~200 molar excess of G-actin relative to DIAPH1. Thus, the reaction velocity is proportional to the concentration of the DIAPH1 construct present during the initial linear portion of the data. Velocities were calculated from the slope of a linear regression analysis of the first 11 data points in the absence and presence of ligands.
ELISAs
Purified proteins were dissolved in 100 mM sodium bicarbonate, pH 8.3. Four 100 μL volumes of 10 nM GDID, 150 nM DAD, a mixture of 100 nM DID and 150 nM DAD, 10 nM full length DIAPH1 and a positive RhoA control, at a concentration of the midpoint of the titration, were loaded into a 96-well MaxiSorp plate and incubated overnight at 4 °C. Excess solution was removed, the wells were washed once with deionized water, twice with PBS, pH 7.4, containing 0.05% Tween-20, PBST, blocked with 200 μL of 1% BSA in PBS and incubated overnight at 4°C. Excess solution was removed and the wells were washed once with PBST, treated with 100 μL of RhoA containing a 100 molar excess of GPPNP over a range of concentrations and incubated at 4 °C overnight. The wells were treated with 100 μL of a 500:1 dilution of horseradish peroxidase, HRP, -conjugated mouse monoclonal primary RhoA-antibody in PBS ((26C4): sc-418, Santa Cruz Biotechnology) and incubated at 4 °C overnight. Excess solution was removed and the wells were washed once with PBST and treated with 100 μL of 1-Step Ultra 3,3′,5,5′-tetramethylbenzidine-ELISA substrate solution, TMB, (ThermoFisher) to detect HRP activity. Changes in absorption at 650 nm over time were monitored by using a Synergy H1 plate reader at 25 °C until the RhoA positive control reached an absorbance of 2.5. The wells were quenched with 100 μL of 0.1 M H2SO4 and the absorption at 450 nm was recorded. All absorption data were normalized to samples containing no RhoA. Dissociation constants were estimated by fitting the data to the “one-site specific binding” of “one site-total and nonspecific binding” models in GraphPad Prism (https://www.graphpad.com/) version 9.0 software (GraphPad).
Cell culture
Plasmid constructs expressing human DIAPH1 or DIAPH1ΔDAD with a C-terminal monomeric enhanced CFP fluorescent tag were utilized58. AD293 cells (Agilent Inc) were used for transfection and slide preparation13. AD293 was tested by PCR for mycoplasma infection by Agilent, Inc. Briefly, transfection of the DIAPH1-eCFP construct was performed using the ViaFect reagent (Promega) according to manufacturer’s recommendations. After 16 h, the transfection medium was replaced with Opti-MEM/0.1% FBS and the cells were incubated for an additional 24 h. CN03 was prepared by dissolving the reagent in doubly distilled water and added at a concentration of 1 μg/mL to cells in DMEM for 4 h and control cells were untreated. Cells were fixed by replacing the medium with 4% formaldehyde in PBS, pH 7.4, for 20 min at room temperature. Cells were washed twice with PBS, permeabilized in PBS containing 0.1% Triton X-100 for 15 min and stained with Alexa Fluor568 Phalloidin and Hoechst 33258 nuclear stain following the manufacturer’s protocols.
Fluorescence and confocal microscopy
Coverslips were mounted on slides using Fluoromount G (Electron Microscopy Sciences). Images were taken with a ZEISS LSM 980 confocal imaging system using a Plan-Apochromat objective 63×/1.40 Oil DIC. Lasers at 405 nm (0.2% power), 445 nm (4.5% power), and 561 nm (0.9% power) were used to excite the Hoechst 33258, eCFP, and the Alexa Fluor568, respectively. The emission windows for Hoechst 33258, eCFP, and the Alexa Fluor568 were 411–453 nm, 472–579 nm and 588–632 nm, respectively. Since RhoA activity in HEK293 cells varies with cell density, showing a 40% decrease in confluent cultures compared to sub-confluent ones, we employed cell growth protocols that yield sub-confluent cultures59. Images of sub-confluent cells were processed using CellProfiler 4 software60. Cell contours were determined based on the staining intensity of eCFP-DIAPH1 using nuclei images as seeds. Contributions to eCFP and AlexaFluor568 membrane fluorescence were defined as 12-pixel thick regions along cell boundaries. Contributions to eCFP and AlexaFluor568 cytosolic fluorescence were defined as cell areas excluding the nuclear and membrane area. Data were processed using Prism 9.0.2 software (GraphPad). Column heights represent mean values, error bars represent the standard deviation. The statistical significance of comparisons was estimated using the One-way ANOVA subroutine and Fisher’s Least Significant Difference test with a single pooled variance displayed.
Statistics and reproducibility
All data, unless otherwise mentioned in the text, were obtained from a minimum of three independent experiments performed at different times. The Shapiro–Wilk test for normality was performed for each group of data before performing any pairwise comparisons. Statistical analysis was performed using the unpaired, two-tailed t-test in Prism 6. For figure generation, we used Adobe Illustrator 2020.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Hilger, D., Masureel, M. & Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 25, 4–12 (2018).
Peurois, F., Peyroche, G. & Cherfils, J. Small GTPase peripheral binding to membranes: molecular determinants and supramolecular organization. Biochem. Soc. Trans. 47, 13–22 (2019).
Call, M. E. & Wucherpfennig, K. W. The T cell receptor: critical role of the membrane environment in receptor assembly and function. Annu. Rev. Immunol. 23, 101–125 (2005).
Koshland, D. E. Jr., Nemethy, G. & Filmer, D. Comparison of experimental binding data and theoretical models in proteins containing subunits. Biochemistry 5, 365–385 (1966).
Monod, J., Wyman, J. & Changeux, J. P. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965).
Di Cera, E., Gill, S. J. & Wyman, J. Canonical formulation of linkage thermodynamics. Proc. Natl. Acad. Sci. USA 85, 5077–5081 (1988).
Hall, A. Rho GTPases and the actin cytoskeleton. Science 279, 509–514 (1998).
Breitsprecher, D. & Goode, B. L. Formins at a glance. J. Cell Sci. 126, 1–7 (2013).
Kovar, D. R., Harris, E. S., Mahaffy, R., Higgs, H. N. & Pollard, T. D. Control of the assembly of ATP- and ADP-actin by formins and profilin. Cell 124, 423–435 (2006).
Manigrasso, M. B. et al. Small-molecule antagonism of the interaction of the RAGE cytoplasmic domain with DIAPH1 reduces diabetic complications in mice. Sci. Transl. Med. 13, eabf7084 (2021).
Ramasamy, R., Shekhtman, A. & Schmidt, A. M. RAGE/DIAPH1 axis and cardiometabolic disease: from nascent discoveries to therapeutic potential. Arterioscler. Thromb. Vasc. Biol. 44, 1497–1501 (2024).
Theophall, G. G. et al. Disruption of the productive encounter complex results in dysregulation of DIAPH1 activity. J. Biol. Chem. 299, 105342 (2023).
Yepuri, G. et al. DIAPH1-MFN2 interaction regulates mitochondria-SR/ER contact and modulates ischemic/hypoxic stress. Nat. Commun. 14, 6900 (2023).
Hudson, B. I. et al. Interaction of the RAGE cytoplasmic domain with diaphanous-1 is required for ligand-stimulated cellular migration through activation of Rac1 and Cdc42. J. Biol. Chem. 283, 34457–34468 (2008).
Ercan-Sencicek, A. G. et al. Homozygous loss of DIAPH1 is a novel cause of microcephaly in humans. Eur. J. Hum. Genet. 23, 165–172 (2015).
Palander, O. & Trimble, W. S. DIAPH1 regulates ciliogenesis and trafficking in primary cilia. FASEB J. 34, 16516–16535 (2020).
Ueyama, T. et al. Constitutive activation of DIA1 (DIAPH1) via C-terminal truncation causes human sensorineural hearing loss. EMBO Mol. Med. 8, 1310–1324 (2016).
Lynch, E. D. et al. Nonsyndromic deafness DFNA1 associated with mutation of a human homolog of the Drosophila gene diaphanous. Science 278, 1315–1318 (1997).
Kundishora, A. J. et al. DIAPH1 variants in non-East Asian patients with sporadic Moyamoya disease. JAMA Neurol. 78, 993–1003 (2021).
Westbury, S. K. et al. Phenotype description and response to thrombopoietin receptor agonist in DIAPH1-related disorder. Blood Adv. 2, 2341–2346 (2018).
Stritt, S. et al. A gain-of-function variant in DIAPH1 causes dominant macrothrombocytopenia and hearing loss. Blood 127, 2903–2914 (2016).
Labat-de-Hoz, L. & Alonso, M. A. Formins in human disease. Cells 10, 2554 (2021).
Otomo, T., Otomo, C., Tomchick, D. R., Machius, M. & Rosen, M. K. Structural basis of Rho GTPase-mediated activation of the formin mDia1. Mol. Cell 18, 273–281 (2005).
Nezami, A., Poy, F., Toms, A., Zheng, W. & Eck, M. J. Crystal structure of a complex between amino and carboxy terminal fragments of mDia1: insights into autoinhibition of diaphanous-related formins. PLoS One 5, https://doi.org/10.1371/journal.pone.0012992 (2010).
Nezami, A. G., Poy, F. & Eck, M. J. Structure of the autoinhibitory switch in formin mDia1. Structure 14, 257–263 (2006).
Lammers, M., Rose, R., Scrima, A. & Wittinghofer, A. The regulation of mDia1 by autoinhibition and its release by Rho*GTP. EMBO J. 24, 4176–4187 (2005).
Rose, R. et al. Structural and mechanistic insights into the interaction between Rho and mammalian Dia. Nature 435, 513–518 (2005).
Li, F. & Higgs, H. N. Dissecting requirements for auto-inhibition of actin nucleation by the formin, mDia1. J. Biol. Chem. 280, 6986–6992 (2005).
Li, F. & Higgs, H. N. The mouse Formin mDia1 is a potent actin nucleation factor regulated by autoinhibition. Curr. Biol. 13, 1335–1340 (2003).
Seth, A., Otomo, C. & Rosen, M. K. Autoinhibition regulates cellular localization and actin assembly activity of the diaphanous-related formins FRLalpha and mDia1. J. Cell Biol. 174, 701–713 (2006).
Maiti, S. et al. Structure and activity of full-length formin mDia1. Cytoskeleton 69, 393–405 (2012).
Watanabe, N. et al. p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein for Rho small GTPase and is a ligand for profilin. EMBO J. 16, 3044–3056 (1997).
Watanabe, N., Kato, T., Fujita, A., Ishizaki, T. & Narumiya, S. Cooperation between mDia1 and ROCK in Rho-induced actin reorganization. Nat. Cell Biol. 1, 136–143 (1999).
Lammers, M., Meyer, S., Kuhlmann, D. & Wittinghofer, A. Specificity of interactions between mDia isoforms and Rho proteins. J. Biol. Chem. 283, 35236–35246 (2008).
Buck, M., Xu, W. & Rosen, M. K. A two-state allosteric model for autoinhibition rationalizes WASP signal integration and targeting. J. Mol. Biol. 338, 271–285 (2004).
Padrick, S. B. & Rosen, M. K. Physical mechanisms of signal integration by WASP family proteins. Annu. Rev. Biochem. 79, 707–735 (2010).
Chen, A. et al. The scaffold-protein IQGAP1 enhances and spatially restricts the actin-nucleating activity of Diaphanous-related formin 1 (DIAPH1). J. Biol. Chem. 295, 3134–3147 (2020).
Levitzki, A. & Koshland, D. E. Jr. Negative cooperativity in regulatory enzymes. Proc. Natl. Acad. Sci. USA 62, 1121–1128 (1969).
Velazquez-Campoy, A., Goni, G., Peregrina, J. R. & Medina, M. Exact analysis of heterotropic interactions in proteins: Characterization of cooperative ligand binding by isothermal titration calorimetry. Biophys. J. 91, 1887–1904 (2006).
Otomo, T., Tomchick, D. R., Otomo, C., Machius, M. & Rosen, M. K. Crystal structure of the Formin mDia1 in autoinhibited conformation. PLoS One 5, https://doi.org/10.1371/journal.pone.0012896 (2010).
Hofmann, H. et al. Polymer scaling laws of unfolded and intrinsically disordered proteins quantified with single-molecule spectroscopy. Proc. Natl. Acad. Sci. USA 109, 16155–16160 (2012).
Cytoskeleton, Inc. Total RhoA ELISA Biochem Kit - Manual, https://www.cytoskeleton.com/pdf-storage/datasheets/bk150.pdf (2025).
Flatau, G. et al. Toxin-induced activation of the G protein p21 Rho by deamidation of glutamine. Nature 387, 729–733 (1997).
Gorelik, R., Yang, C., Kameswaran, V., Dominguez, R. & Svitkina, T. Mechanisms of plasma membrane targeting of formin mDia2 through its amino terminal domains. Mol. Biol. Cell 22, 189–201 (2011).
Ramalingam, N. et al. Phospholipids regulate localization and activity of mDia1 formin. Eur. J. Cell Biol. 89, 723–732 (2010).
Chen, A., Arora, P. D., McCulloch, C. A. & Wilde, A. Cytokinesis requires localized beta-actin filament production by an actin isoform specific nucleator. Nat. Commun. 8, 1530 (2017).
Higashi, T. et al. Flightless-I (Fli-I) regulates the actin assembly activity of diaphanous-related formins (DRFs) Daam1 and mDia1 in cooperation with active Rho GTPase. J. Biol. Chem. 285, 16231–16238 (2010).
Sakamoto, S. et al. Liprin-alpha controls stress fiber formation by binding to mDia and regulating its membrane localization. J. Cell Sci. 125, 108–120 (2012).
Brenig, J. et al. Structural and Biochemical Basis for the Inhibitory Effect of Liprin-alpha3 on Mouse Diaphanous 1 (mDia1) Function. J. Biol. Chem. 290, 14314–14327 (2015).
Rai, V. et al. Signal transduction in receptor for advanced glycation end products (RAGE): solution structure of C-terminal rage (ctRAGE) and its binding to mDia1. J. Biol. Chem. 287, 5133–5144 (2012).
Scholz, J. et al. Decisive role of mDia-family formins in cell cortex function of highly adherent cells. Sci. Adv. 10, eadp5929 (2024).
Junemann, A. et al. A Diaphanous-related formin links Ras signaling directly to actin assembly in macropinocytosis and phagocytosis. Proc. Natl. Acad. Sci. USA 113, E7464–E7473 (2016).
Zimmermann, D. et al. Mechanoregulated inhibition of formin facilitates contractile actomyosin ring assembly. Nat. Commun. 8, 703 (2017).
Wilkins, M. R. et al. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 112, 531–552 (1999).
Piotto, M., Saudek, V. & Sklenar, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR 2, 661–665 (1992).
Masse, J. E. & Keller, R. AutoLink: automated sequential resonance assignment of biopolymers from NMR data by relative-hypothesis-prioritization-based simulated logic. J. Magn. Reson. 174, 133–151 (2005).
Kouyama, T. & Mihashi, K. Fluorimetry study of N-(1-pyrenyl)iodoacetamide-labelled F-actin. Local structural change of actin protomer both on polymerization and on binding of heavy meromyosin. Eur. J. Biochem. 114, 33–38 (1981).
Xue, J. et al. Change in the molecular dimension of a RAGE-ligand complex triggers RAGE signaling. Structure 24, 1509–1522 (2016).
Dubash, A. D. et al. The small GTPase RhoA localizes to the nucleus and is activated by Net1 and DNA damage signals. PLoS One 6, e17380 (2011).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinforma. 22, 433 (2021).
Butch, A. W. Dilution protocols for detection of hook effects/prozone phenomenon. Clin. Chem. 46, 1719–1721 (2000).
Nagasaki, A. et al. https://doi.org/10.1101/2025.02.05.636544 (2025).
Acknowledgements
The project described was supported by Award Numbers 1P01HL146367-01, 5R01DK122456 and R01GM085006 from the U.S. National Institute of Health. Mass spectrometry analyses were performed by the RNA Epitranscriptomics & Proteomics Resource located at the Life Science Research Building, University at Albany, supported by the SUNY Research Foundation and NIH Award 5R01GM143749.
Author information
Authors and Affiliations
Contributions
G.G.T., A.P., S.R., E.O., P.N. investigation; G.G.T., S.R. formal analysis. R.R., A.M.S., and A.S. conceptualization; G.G.T., D.S.B., R.R., A.M.S., and A.S. writing-original draft; G.G.T., D.S.B., R.R., A.M.S., and A.S. writing and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Kien Xuan Ngo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Kaliya Georgieva.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Theophall, G.G., Premo, A., Reverdatto, S. et al. Negative cooperativity regulates ligand activation of DIAPH1 and other diaphanous related formins. Commun Biol 8, 776 (2025). https://doi.org/10.1038/s42003-025-08222-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08222-5