Abstract
The mitochondria-associated degradation pathway (MAD) mediates removal and elimination of damaged, unfolded mitochondrial proteins by the ubiquitin-proteasome system (UPS). Previous studies revealed that MAD is critical for mitochondrial protein quality control and that MAD function extends beyond mitochondrial outer membrane (MOM) to proteins within the organelle. Here, we reconstitute retrotranslocation of MAD substrates from the mitochondrial matrix across mitochondrial inner and outer membranes in cell-free systems. This retrotranslocation is ATP-dependent but membrane potential-independent. We also identify a role for the TOM complex, the protein import channel in the MOM, in this process. Inhibition of protein translocation across the Tom40p channel reduces the retrotranslocation of MAD substrates. Our studies support the model that the TOM complex is a bidirectional protein channel in the MOM: it mediates retrotranslocation of damaged mitochondrial proteins across the MOM in the MAD pathway for mitochondrial protein quality control in addition to its function in import of proteins into the organelle.

Similar content being viewed by others
Introduction
Mitochondria have critical cellular functions including energy mobilization, calcium regulation, metabolite production and redox homeostasis1,2. However, mitochondrial proteins are challenged by proteostasis stressors that are not found in many other organelles. Mitochondrial proteins are encoded by two genomes (nuclear and mitochondrial DNA), targeted to four submitochondrial compartments, assembled into some of the largest membrane protein complexes found in eukaryotic cells, and subject to damage by mitochondrial reactive oxygen species (ROS)2,3. Indeed, defects in folding or assembly of the human respiratory chain megacomplex I2III2IV2, which contains 140 integral and peripheral membrane proteins4, play a role in the pathology of proteinopathies including Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis5.
Cells utilize different quality control mechanisms to maintain mitochondria, from degradation of individual unfolded mitochondrial proteins to recycling of whole organelles by mitophagy6. Here, we studied the mechanism underlying the mitochondria-associated degradation pathway (MAD). MAD is a member of a family of quality control pathways in which unfolded and damaged proteins are identified, ubiquitinated, removed from organelles and degraded by the ubiquitin-proteasome system (UPS)7,8. The ER-associated degradation pathway (ERAD) is the founding member of these pathways. Similar pathways were later identified in mitochondria (e.g., MAD), chloroplasts (chloroplast-associated degradation, CHLORAD) as well as endosomes and golgi (Endosome and Golgi-associated degradation, EGAD)8,9,10,11,12,13,14. There are also multiple MAD-related, UPS-dependent pathways for mitochondrial proteostasis, including mitochondrial protein translocation-associated degradation (mitoTAD), mitochondrial compromised protein import response (mitoCPR) and mitochondria-localized ribosome-associated quality control (mitoRQC)15,16,17,18.
In most of these proteostasis pathways, unfolded, damaged proteins are marked by ubiquitination19. The ubiquitin serves as a signal for removal of those proteins from organelles and transfer of those proteins to the proteasome by the segregase complex20. The segregase complex contains conserved, essential proteins and is a target for mutation in human disease. The core subunit of this complex is Cdc48p (VCP/p97 in mammals)20,21,22,23,24,25, an abundant, essential AAA-ATPase with protein unfolding activity that uses the energy of ATP hydrolysis to extract proteins from membranes and protein complexes. Cdc48p associates with cofactors, including Npl4p and Ufd1p, that mediate substrate recognition by binding to ubiquitin on unfolded proteins. It also binds to organelle-targeting proteins, which are critical for recruiting the segregase complex to its site of action within cells. For example, Doa1p (aka Ufd3p, PLAA or PLAP in mammals) binds to ubiquitin and Cdc48p, and is required for targeting of the complex to mitochondria in MAD21,22,23,24,25,26.
MAD and its associated pathways, CHLORAD or EGAD, were initially identified as pathways that target unfolded proteins that localize to the cytosolic surface of mitochondria, chloroplast, endosome or golgi, respectively9,10,11,21,23. In contrast, degradation of unfolded, damaged or misassembled proteins within these organelles was believed to be carried out exclusively by proteases within these organelles27,28. For example, Lon, ClpXP, and m-AAA ATPase, are conserved proteases that mediate degradation of misfolded proteins in the mitochondrial matrix27. However, it is clear that MAD and its related pathways can target unfolded proteins within organelles for proteasomal degradation. For example, early studies revealed that ERAD substrates are found not only on the ER surface, but within the organelle. ERAD components, including mammalian Derlin and yeast Hrd1p, function as channels for retrotranslocation of unfolded proteins from the ER lumen or lumenal side of the ER membrane across the ER membrane to the ER surface14,29.
Our previous studies revealed that mitochondrial matrix and inner membrane proteins undergo oxidative stress-induced ubiquitination and are substrates for the MAD pathway. Two of the mitochondrial matrix proteins identified in these studies, Kgd1p (a subunit of the α ketoglutarate dehydrogenase complex) and Pim1p (the Lon protease of yeast) are known targets for oxidative damage, are ubiquitinated under basal conditions and exhibit increased ubiquitination in response to mitochondrial oxidative stress. Moreover, these proteins interact with Cdc48p in a Doa1p-dependent manner and are degraded by the proteasome30. Similarly, recent studies indicate that Cdc48p mediates ubiquitin-dependent degradation of damaged intra-chloroplast proteins in CHLORAD31. Thus, there is evidence that proteins within organelles are clients for proteasomal degradation not just in ERAD, but also in MAD and CHLORAD.
Since MAD substrates localize to the mitochondrial matrix, they must be retrotranslocated across both mitochondrial membranes prior to interaction with the segregase complex and proteasome in the cytosol. However, there is no direct evidence for retrotranslocation of MAD substrates across the mitochondrial outer membrane (MOM) or mitochondrial inner membrane (MIM) or for a mechanism underlying those membrane translocation events. Interestingly, several findings have linked the TOM complex, which serves as a channel for import of proteins across the MOM, to mitochondrial protein quality control. Specifically, the segregase complex interacts with the TOM complex to target translocation-stalled precursor proteins for degradation by the UPS in the MAD-related pathways mitoTAD and mitoCPR16,17,18. Moreover, the TOM complex functions in retrotranslocation of conformationally destabilized proteins from the mitochondrial intermembrane space (IMS) to the cytosol32,33. However, since the fate of retrotranslocated IMS proteins is not known, it is not clear whether this retrotranslocation event is linked to MAD or MAD-related pathways. Here, we reconstituted retrotranslocation of MAD substrates from the matrix across both mitochondrial membranes using isolated mitochondria. We also identified a role for the TOM complex as a retrotranslocation channel for MAD in the MOM.
Results
Reconstitution of the retrotranslocation of MAD substrates from the mitochondrial matrix in a cell-free system
Here, we tested whether MAD substrates that localize to the matrix can be retrotranslocated from mitochondria using the budding yeast Saccharomyces cerevisiae as a model organism. Specifically, isolated mitochondria are incubated in “assay buffer”. Thereafter, mitochondria are separated from the assay buffer by centrifugation, and western blot analysis is used to test whether Kgd1p, a MAD substrate in the mitochondrial matrix, is released from the organelle (Fig. 1). In these studies, the KGD1 gene was tagged at its chromosomal locus with GFP and expressed at endogenous levels. We confirmed that GFP-tagged Kgd1p localized exclusively to mitochondria by visualization of Kgd1p-GFP and MitoTracker Red-stained mitochondria in living yeast cells (Fig. 1A) and by subcellular fraction studies in which Kgd1p-GFP co-fractionated with the mitochondrial marker protein Tom40p (Fig. 1B). In addition, the GFP tag had no detectable effect on mitochondrial morphology and distribution (Fig. 1A) or on cell growth rates (data not shown). Thus, the GFP tag has no obvious negative effect on Kgd1p or mitochondria.
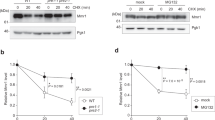
A Representative images of the localization of Kgd1p-GFP in control (WT) or doa1Δ in SUB592 background. Mitochondria were visualized with MitoTracker Red. Scale bar, 2 µm. B Representative western blots of Kgd1p-GFP in the isolated mitochondrial (Mito) and cytosolic (Cyto) fraction from WT (+) and doa1Δ (−) cells in the SUB592 background. The blot was probed with antibodies against GFP for detection of Kgd1p, and antibodies against Tom40p and hexokinase. Total protein load was assessed using TCE. C Representative western blots of mitochondrial pellet (Pellet) recovered in the release assay and proteins released (Released) from isolated mitochondria from WT (+) and doa1Δ (−) cells in the SUB592 background after incubation for 10, 30, and 60 min in assay buffer. The blot was probed with antibodies against GFP for detection of Kgd1p, and antibodies against Tom40p and Cyb2p. Total protein load was assessed using TCE. D Quantification of Kgd1p recovered in the mitochondria pellet in (C). Kgd1p signals were normalized to TCE and then normalized to WT (+) at 10 min (n = 3, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01). E Quantification of Kgd1p released from isolated mitochondria in (C). Kgd1p signals were normalized to TCE and then normalized to Kgd1p at 10 min (n = 3, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; ***p < 0.001; ****p < 0.0001) F Western blot of crude mitochondria (input) and total ubiquitinated proteins pulled down with Ni2 + -NTA magnetic beads from isolated mitochondria (IP). Blots were probed with antibodies against GFP to detect GFP-tagged Kgd1p, against Cit1p and against 6xHis for ubiquitin detection. G Representative western blots as in (C). The blot was probed with antibodies against GFP for detection of Kgd1p, and antibodies against Cit1p. H Quantification of Cit1p in mitochondria in (G). I Quantification of Cit1p in released fraction from isolated mitochondria in (G). Cit1p signals were normalized to mitochondrial signals. For panels H, I, n = 3, 1-way ANOVA with Sidak’s multiple comparison test, **p < 0.01; ***p < 0.001; ns no significant difference.
To assess whether the release of Kgd1p-GFP from mitochondria is MAD-dependent, the assay was conducted using mitochondria isolated from yeast bearing a deletion in DOA1. Deletion of DOA1 disables MAD by inhibiting transfer of MAD substrates from the mitochondrial surface to the proteasome for degradation. Since Doa1p also modulates mono-ubiquitin levels, which can affect MAD substrates through effects on the UPS34, ubiquitin levels were maintained at wild-type (WT) levels by ectopic, plasmid-borne ubiquitin expression30,35,36. Thus, under these experimental conditions, the phenotype observed upon deletion of DOA1 is due to defects in Doa1p function in MAD and not maintenance of mono-ubiquitin.
We find that Kgd1p-GFP is released from mitochondria isolated from wild-type (WT) and doa1∆ cells. In both cases, the release of Kgd1p-GFP is time-dependent: Kgd1p-GFP release from mitochondria is evident after incubation for 10 min and increases upon incubation for 30–60 min (Fig. 1C, G). Equally important, the observed release of Kgd1p-GFP from mitochondria is linked to MAD. First, deletion of DOA1 results in a 1.3-fold increase in the steady state levels of Kgd1p-GFP in mitochondria and a 2.5- to 4-fold increase in release of the Kgd1p-GFP from isolated mitochondria (Fig. 1C–E). Second, we find that Kgd1p-GFP is ubiquitinated in a MAD-dependent manner. We immunoprecipitated ubiquitinated proteins from mitochondria isolated from WT and doa1∆ cells found that Kgd1p is ubiquitinated, and deletion of DOA1 results in increased the steady state levels of ubiquitinated Kgd1p in mitochondria. Moreover, MAD-linked ubiquitination of Kgd1p appears to be specific: there is no detectable ubiquitination of Cit1p, another abundant mitochondrial matrix protein, in mitochondria from WT or doa1∆ cells (Fig. 1F).
Under these conditions, we find that deletion of DOA1 results in an increase in release of Kgd1p from isolated mitochondria compared to that observed from mitochondria isolated from WT cells (Fig. 1C–E). Again, the effect appears to be specific for Kgd1p: there is no detectable release of Cit1p from mitochondria isolated from WT or doa1∆ cells (Fig. 1G–I). Finally, only a small percent of the total mitochondrial Kgd1p-GFP is released from the organelle: during the 60 min time-course, only 1–2% and 2–7% of total mitochondrial Kgd1p-GFP is released from mitochondria isolated from WT or doa1∆ cells, respectively. A similar small percentage of mitochondrial Kgd1p is ubiquitinated. Thus, we detect MAD-dependent release of Kgd1p-GFP from isolated yeast mitochondria.
To determine whether the observed release is due to defects in mitochondrial integrity, we monitored the behavior of three mitochondrial proteins: Tom40p, a subunit of the TOM complex in the MOM; Cyb2p, Cytochrome b2 of yeast, a peripheral membrane protein associated with the outer leaflet of the MIM; and Cit1p, mitochondrial citrate synthase, a soluble matrix protein. None of these proteins have been identified as MAD substrates23,30. Indeed, there is no detectable ubiquitination of Cit1p in mitochondria isolated from WT or doa1∆ yeast. Under conditions where we observe MAD-linked release of Kgd1p from isolated mitochondria, there is no detectable release of Tom40p, Cyb2p, or Cit1p from mitochondria after incubation for 10-60 min or and no detectable change in release of Tom40p, Cyb2p, or Cit1p from mitochondria isolated from WT compared to doa1∆ cells. Thus, the release of Kgd1p from isolated mitochondria, is linked to MAD and not due to leakiness of the MOM or MIM.
To exclude the possibility that the observed release is due to genetic background or the GFP tag on Kgd1p, we performed the assay using KGD1 that was tagged with the Myc epitope in the yeast from a different background (BY4741). Here too, we observe release of Myc-tagged Kgd1p from isolated mitochondria under conditions where Tom70p and Cit1p are quantitatively retained within the organelle. In addition, deletion of DOA1 results in an increase in Kgd1p-Myc release but has no effect on the integrity of the organelle (Fig. S1). Thus, the observed release of Kgd1p-GFP from the mitochondrial matrix is not due to the genetic background or the protein tag employed.
Next, we tested whether the observed Kgd1p-GFP release is linked to mitoTAD (mitochondrial protein translocation-associated degradation). If release of Kgd1p-GFP is due to mitoTAD, then there should be defects in import of Kgd1p-GFP into the organelle. However, as described above, visualization of Kgd1p-GFP in living yeast cells and subcellular fractionation studies revealed that Kgd1p-GFP localizes exclusively to mitochondria in WT and doa1∆ cells (Fig. 1A, B). Thus, the import of the protein into mitochondria in WT or doa1∆ cells is not severely compromised in vivo. In addition, if the released Kgd1p-GFP is due to MitoTAD and not MAD, then Kgd1p-GFP should be mislocalized to the MOM as a result of defective transport of the protein at the TOM complex. To test this, we studied that the effect of proteinase K treatment of isolated mitochondria on the recovery of Kgd1p-GFP with the organelle, and release of Kgd1p-GFP from the organelle in vitro. Here, the proteinase K conditions used degraded MOM marker proteins without affecting MOM or MIM integrity. Specifically, proteinase K degrades the MOM marker, Tom70p, but did not result in degradation of Cyb2p, an IMS protein, or Cit1p, a matrix protein (Fig. 2A, C, D, F, G). Importantly, proteinase K treatment has no detectable effect on mitochondrial Kgd1p-GFP levels (Fig. 2A, B, D, E). Thus, there is no widespread mistargeting of Kgd1p-GFP to the MOM. Equally important, proteinase K-mediated degradation of mitochondrial surface proteins has no effect on the release of Kgd1p-GFP from isolated mitochondria, and does not result in non-specific release of Tom70p or Cyb2p from the organelle (Fig. 2D, H–J). Overall, our findings indicate that the observed release of Kgd1p-GFP from isolated mitochondria is not due to MitoTAD.
A Representative western blot of isolated mitochondria from WT (+) and doa1Δ (−) cells in the SUB592 background, +/− proteinase K (ProK) or 2% Triton X for 30 min at 0 °C. B, C Quantification of Kgd1p (B) and Tom70p (C) levels in mitochondria in (A). Kgd1p and Tom70p signals were normalized to TCE, and then these normalized signals were normalized to WT (+) (For panels B, C, n = 4, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ****p < 0.0001; ns, no significance). D Isolated mitochondria were incubated in the presence or absence of proteinase K (10 µg/mL) for 30 min at 0 °C. After inactivation of proteinase K by addition of protease inhibitors, mitochondria were separated from the proteinase K-containing solution and release of Kgd1p from isolated mitochondria was assessed after 30 min in assay buffer. Representative western blots of the mitochondrial pellet after release (Pellet) and proteins released (Released) from mitochondria isolated from WT (+) and doa1Δ (−) cells in the SUB592 background, +/− proteinase K (ProK). E–G Quantification of Kgd1p (E), Tom70p (F) and Cyb2p (G) levels in mitochondria in (D). Kgd1p, Tom70p and Cyb2p signals were first normalized to TCE, and then these normalized signals were normalized to WT (+). H–J Quantification of Kgd1p (H), Tom70p (I) and Cyb2p (J) released from isolated mitochondria in (D) Kgd1p, Tom70p and Cyb2p signals were first normalized to TCE, and then these normalized signals were normalized to WT (+) (For panels E–J, n = 3, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ****p < 0.0001; ns no significance).
Finally, previous study reveals that damaged IMS proteins can be retrotranslocated across the MOM by the TOM complex32. To rule out the possibility that the observed released Kgd1p-GFP reflects export of mistargeted Kgd1p-GFP from IMS, we tested whether osmotic shock of mitochondria, which results in rupture of the MOM and loss of IMS proteins, affects the level of mitochondrial Kgd1p-GFP (Fig. S2). Here, osmotic shock resulted in release of 40% of Cyb2p from mitochondria (Fig. S2D), with no detectable release of Cit1p into the supernatant fraction (Fig. S2E). Thus, the conditions used breach the MOM without affecting MIM integrity. Under these conditions, there is no detectable release of Kgd1p-GFP from osmotically shocked mitochondria (Fig. S2C).
Overall, we detect MAD-linked retrotranslocation of Kgd1p-GFP from the matrix across the MIM and MOM using isolated yeast mitochondria. The observed release is not due to defects in the integrity of mitochondrial outer or inner membranes. Moreover, the GFP tag on Kgd1p does not affect import of Kgd1p-GFP into mitochondria or correct targeting of the protein to the mitochondrial matrix. This retrotranslocation of Kgd1p-GFP across the MOM and MIM (1) is specific for MAD substrates, (2) correlates with MAD-dependent ubiquitination, and (3) is not linked to mitoTAD or protein export from the IMS.
Retrotranslocation of MAD substrates is ATP-dependent and membrane potential-independent
Here, we tested whether retrotranslocation of two MAD substrates, Kgd1p and Pim1p, requires energy and a membrane potential across the MIM (∆Ψ) (Fig. 3). Our previous studies indicate that Pim1p, like Kgd1p, is ubiquitinated in response to exposure of yeast of oxidative stress in mitochondria and that deletion of DOA1 and the associated inhibition of MAD results in an increase in the steady state protein and ubiquitin levels of mitochondrial Pim1p30. Thus, Pim1p is a MAD substrate that localizes to the mitochondrial matrix.
A, D Representative western blots of mitochondrial pellet after releasing (Pellet) and released proteins (Released) from isolated mitochondria from WT (+) and doa1Δ (−) cells in the SUB592 background in the absence or presence of 2 mM ATP and NADH, or 5 µg/ml valinomycin. Release was assessed after incubation of isolated mitochondria for 30 min at 30 °C. The blot was probed with antibodies against GFP for detection of Kgd1p (A) or Pim1p (D), and antibodies against Tom40p and Cyb2p. Total protein load was assessed using TCE. B, E Quantification of Kgd1p (B) and Pim1p (E) in mitochondria in (A) and (D), respectively. C, F Quantification of Kgd1p (C) and Pim1p (F) released from isolated mitochondria in (A) and (D), respectively (n = 4 in (B and C) and n = 3 in (E and F), 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; ns no significance). All signals were first normalized to pellet Cyb2p signals and then normalized to WT (+).
Here, we find that Pim1p-GFP is released from mitochondria under conditions where Tom40p and Cyb2p are quantitatively retained with the organelle, and that the release of Pim1p from mitochondria is increased by deletion of DOA1 (Fig. 3D–F). Moreover, since the GFP-tagged Pim1p that was used is expressed at endogenous levels and quantitatively targeted to mitochondria in living yeast cells (Fig. S3), import of Pim1p-GFP into mitochondria is fully functional under our experimental conditions. Thus, the release of Pim1p-GFP, like Kgd1p-GFP, is due to MAD-linked retrotranslocation of the protein across the MIM and MOM.
To determine whether retrotranslocation of GFP-tagged Kgd1p and Pim1p requires ATP, we added ATP to the assay buffer. Since isolated yeast mitochondria contain some ATP37, addition of ATP supplements mitochondrial ATP levels. Therefore, if retrotranslocation of MAD substrates from the matrix is energy-dependent, then addition of ATP should increase release of GFP-tagged Kgd1p and Pim1p from mitochondria. We found that addition of ATP does not affect the steady state level of either protein in mitochondria (Fig. 3A, B, D, E). However, it does result in a significant increase in the release of GFP-tagged Kgd1p and Pim1p from mitochondria isolated from wild-type and doa1∆ cells (Fig. 3A, C, D, F). Indeed, supplementation with ATP results in a 2-fold and 1.7-fold increase in the release of GFP-tagged Kgd1p and Pim1p from mitochondria isolated from doa1∆ cells compared to WT cells.
To test whether protein retrotranslocation in MAD requires ATP hydrolysis, the non-hydrolyzable ATP analog, ATP-γ-S, was added to the in vitro assay. We found that retrotranslocation of Kgd1p is not affected by supplementation with ATP-γ-S (Fig. S4A, B). Collectively, we obtained evidence that retrotranslocation of MAD substrates from the matrix and across both mitochondrial membranes requires ATP and ATP hydrolysis. Thus, MAD-linked retrotranslocation of damaged, unfolded proteins from the mitochondrial matrix is an active transport event.
Finally, we tested whether this retrotranslocation event is ∆Ψ-dependent by treatment with valinomycin, an ionophore that dissipates ∆Ψ. Treatment of mitochondria isolated from WT or doa1∆ cells with valinomycin has no detectable effect on the levels of GFP-tagged Kgd1p or Pim1p in isolated mitochondria (Fig. 3A, B, D, E). Surprisingly, treatment of mitochondria with valinomycin also has no significant effect on release of either Kgd1p-GFP or Pim1p-GFP (Fig. 3A, C, D, F and S4C) from the mitochondria isolated from WT or doa1∆ cells. Thus, MAD-mediated retrotranslocation of substrates across the MIM and MOM does not require a ∆Ψ across the MIM.
The Tom40p is a channel for retrotranslocation of MAD substrates across the MOM
Here, we used the retrotranslocation assay to determine whether the TOM complex can function as a retrotranslocation channel in the MAD pathway. Since the Tom40p subunit of the TOM complex is essential for yeast cell viability, we studied the effect of deletion of TOM5, which encodes a non-essential subunit of the TOM complex, on retrotranslocation of Kgd1p-GFP from isolated mitochondria. During import of proteins into mitochondria, Tom5p mediates transfer of nuclear-encoded mitochondrial proteins from the Tom70p and Tom20p receptors to the Tom40p pore of the TOM complex38. Tom5p is also required for stabilization of the TOM complex39.
We confirmed that deletion of TOM5 results in destabilization of the TOM complex: there is a significant decrease in mitochondrial Tom40p levels in tom5∆ compared to WT cells (Fig. 4A, C). Importantly, the steady state levels of Kgd1p-GFP and Cyb2p in mitochondria isolated from WT and tom5∆ cells are similar (Fig. 4A, B, D). Thus, deletion of TOM5 destabilizes the TOM complex but has negligible effects on the steady state levels in mitochondria of 2 proteins that undergo TOM complex-dependent import.
A Representative western blots of mitochondrial pellet after releasing (Pellet) and released proteins (Released) from isolated mitochondria from wild-type (WT), doa1Δ, tom5Δ, and doa1Δ tom5Δ cells. The blot was probed with antibodies against GFP for detection of Kgd1p, and antibodies against Tom40p and Cyb2p. Total protein load was assessed using TCE. B–D Quantitation of levels of Kgd1p (B), Tom40p (C) and Cyb2p (D) in mitochondria in (A). E Quantification of mitochondrial proteins released from isolated mitochondria in (A). For panels B–E, signals were first normalized to pellet Cyb2p signals and then normalized to WT. (For panels B–E, n = 4, 1-way ANOVA with Sidak’s multiple comparison test, **p < 0.01; ***p < 0.001; ****p < 0.0001; ns no significance). F, H Representative western blots of mitochondrial pellet after releasing (Pellet) and released proteins (Released) from isolated mitochondria from WT (+) and doa1Δ (−) cells in the presence or absence of 1 mM mPEG. The blot was probed with antibodies against GFP for detection of Kgd1p (F) or Pim1p (H), and against Tom40p and Cyb2p. Total protein load was assessed using TCE. G, I Quantification of released mitochondrial proteins from isolated mitochondria in (F) and (H), respectively (n = 4, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; ns no significance). J, L Representative western blots of mitochondrial pellet after releasing (Pellet) and released proteins (Released) from isolated mitochondria from tom40130/138C (+) and tom40130/138C doa1Δ (−) cells in the presence or absence of 1 mM mPEG. The blot was probed with antibodies against GFP for detection of Kgd1p (J) or Pim1p (L), and against Tom40p and Cyb2p. Total protein load was assessed using TCE. K, M Quantification of released mitochondrial proteins from isolated mitochondria in (J, L) (n = 4, 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ***p < 0.001; ns no significance). In all quantification, signals were first normalized to pellet Cyb2p signals and then normalized to WT without mPEG treatment.
We find that deletion of TOM5 has no significant effect on release of Kgd1p from mitochondria isolated from WT cells. In contrast, we observe a 50% decrease in the release of Kgd1p from tom5∆ doa1∆ cells compared to doa1∆ cells (Fig. 4A, E). Thus, our findings support a role for TOM complex in retrotranslocation of MAD substrates from the mitochondrial matrix under stressed conditions. Since deletion of TOM5 does not completely inhibit TOM complex function, it is possible that TOM complex activity in tom5∆ cells is sufficient to carry out retrotranslocation under basal, non-stressed conditions.
Since deletion of TOM5 can have pleiotrophic effects, we used an alternative approach to study a possible role for the TOM complex as a retrotranslocation channel in the MOM. Here, we tested whether steric inhibition of the lumen of the Tom40p channel affects the retrotranslocation of MAD substrates from mitochondria in vitro using a previously described method32,40. Asn130 and Ser138, two amino acid residues that are in the lumen of the Tom40p channel, were replaced with cysteines using CRISPR. The lumen of the Tom40p channel was then blocked by covalent modification of Cys130 and 138 with maleimide-coupled polyethylene glycol (mPEG) beads. Previous studies revealed that this approach blocks the Tom40p channel and inhibits import of proteins into mitochondria in vitro.
In control studies, we find that retrotranslocation of Kgd1p and Pim1p from mitochondria and accumulation of those MAD substrates in mitochondria is not affected by substitution of 2 residues in the Tom40p channel lumen with cysteines (Figs. 4F–M and S5). In addition, mPEG treatment of mitochondria isolated from yeast expressing WT TOM40 had no detectable effect on (1) the abundance or apparent molecular weight of Tom40p, or (2) retrotranslocation of Kgd1p or Pim1p from mitochondria in vitro (Figs. 4F–I and S5). In contrast, treatment of mitochondria isolated from tom40130/138C cells with mPEG results in an increase in the apparent molecular weight of Tom40p expected by addition of two 5-kD mPEG moieties to the protein (Fig. 4J, L). mPEG treatment has no significant effect on baseline retrotranslocation of Kgd1p or Pim1p from mitochondria isolated from the tom40130/138C mutant. Again, residual TOM complex activity in mPEG-treated tom40130/138C may be sufficient to support retrotranslocation under non-stressed conditions. However, mPEG treatment of mitochondria isolated from tom40130/138C bearing a deletion in DOA1 results in a statistically significant decrease in retrotranslocation of Kgd1p and Pim1p compared to that observed the control (mPEG-treated mitochondria isolated from doa1∆ cells that express WT TOM40, Fig. 4J–M). Thus, we obtained evidence that steric hindrance in the Tom40p channel reduces MAD-linked retrotranslocation of MAD substrates from the mitochondrial matrix under proteostatic stress conditions.
Collectively, our findings indicate that the TOM complex plays a direct role in release of MAD substrates from the matrix to the extramitochondrial space. Since the TOM complex localizes to the MOM where it functions as a protein import channel, our findings support a direct role for the TOM complex as a retrotranslocation channel for transport of unfolded, damaged mitochondrial proteins across the MOM in MAD.
The TOM complex is required for normal MAD function and the cellular response to oxidative stress in vivo
Here, we used two approaches to determine whether TOM complex function as a retrotranslocation channel in MAD is physiologically relevant. First, we studied whether deletion of TOM5 affects interaction of MAD substrates with Cdc48p, the essential subunit of the segregase. Our previous studies revealed that Kgd1p co-immunoprecipitates with Cdc48p and that oxidative stress increases Kgd1p-Cdc48p interactions30. If the TOM complex functions in retrotranslocation of unfolded mitochondrial proteins in MAD is physiologically relevant, then deletion of TOM5 should decrease interaction of Kgd1p with the Cdc48p and the segregase complex in vivo.
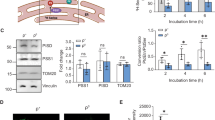
Here, we used co-immunoprecipitation of Myc-tagged Kgd1p with Cdc48p to test the effect of deletion of TOM5 on interaction of a MAD substrate with the segregase complex. Control studies revealed that untagged Kgd1p is not immunoprecipitated with anti-Myc antibody. In addition, we observe co-immunoprecipitation of Cdc48p with Myc-tagged Kgd1p and that this association is reduced by deletion of DOA1. Thus, we confirmed that Kgd1p is associated with the segregase complex and that this association is linked to MAD30 (Fig. 5A, B). Finally, we found that co-immunoprecipitation of Kgd1p-Myc with Cdc48p is reduced by deletion of TOM5 (Fig. 5A, B). These findings support the interpretation that TOM complex functions in retrotranslocation of Kgd1p in MAD and that defects in this process results in reduced interaction of Kgd1p-Myc with Cdc48p.
A Representative western blots of whole cell lysates (Input) and proteins immunoprecipitated with an anti-c-Myc antibody (IP) from whole-cell lysates of non-tagged cells (no tag) or cells expressing Kgd1p-13xMyc in wild-type (WT), doa1Δ, tom5Δ, and doa1Δ tom5Δ cells in the BY4741 background. doa1Δ here represents doa1Δ expressing ubiquitin at similar levels compared to WT. Blots were probed with antibodies against Myc to detect Kgd1p-13Myc and against Cdc48p. Total protein load was assessed using TCE. B Quantification of levels of Cdc48p co-immunoprecipitated with Kgd1p (IP) shown in (A) (n = 3 for each strain, 1-way ANOVA with Dunnett’s multiple comparison test, *p < 0.05). C Growth curves were monitored by measuring optical density at 600 nm (OD600) every 20 min for 60 h in wild-type (WT), doa1Δ, tom5Δ, and doa1Δtom5Δ cells in the presence or absence of 2.5 mM PQ. D Growth rates were calculated from the maximum slope of the growth curve in (C) and shown in SuperPlots, where each color denotes an independent biological replicate, small dots represent individual data points, and triangles represent the mean of the technical replicates within each biological replicate. The p values were calculated from biological replicates. The horizontal bar represents the average of biological replicates and the error bars represent standard error of the mean (SEM) (n = 3, each biological replicate contains >5 technical replicates per strain. Repeated Measurement 2-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001) E The percentage of decreases growth rates in response to PQ (n = 3, each biological replicate contains >5 technical replicates per strain. Repeated Measurement 1-way ANOVA with Sidak’s multiple comparison test, *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001).
Finally, we tested whether deletion of TOM5 affects the oxidative stress response of yeast (Fig. 5C–E). Here, oxidative stress was induced by paraquat (PQ), a redox-active molecule that undergoes ∆Ψ-dependent uptake into mitochondria where it reacts with electrons from the electron transport chain (ETC) to generate superoxides41. PQ treatment reduces yeast growth, as expected. Deletion of DOA1 results in mild growth defects under basal conditions and more severe growth defects produced by PQ exposure. Thus, these studies support a critical role for MAD in the cellular response to PQ-induced oxidative stress30. Next, we tested the effect of PQ treatment on the growth rate of tom5∆, doa1∆ and tom5 doa1∆ yeast cells. We find that there is no significant difference in the growth rate of tom5∆ doa1∆ double mutants compared to tom5∆ single mutants in the presence or absence of PQ treatment (Fig. 5C, D). To test the sensitivity to the growth-inhibiting effects of paraquat, we analyzed the percentage of growth inhibition by PQ. We find that the percentage of inhibition for tom5∆ doa1∆ double mutants is not significantly different compared to those of tom5∆ cells (Fig. 5E). Thus, our data support a role for Tom5p and Doa1p in the cellular response to oxidative stress in yeast and suggest that Tom5p subunit of the TOM complex may play a role in cellular fitness in MAD-mediated cellular response to elevated oxidative stress in mitochondria.
Discussion
Our previous studies revealed that MAD function in mitochondrial protein quality control extends beyond proteins in the MOM and to proteins within the organelle30. Here, we provide the first evidence that MAD substrates can be retrotranslocated from the matrix across both mitochondrial membranes in vitro. Specifically, we reconstituted MAD-dependent retrotranslocation of proteins across the MIM and MOM using isolated mitochondria, and found that this process requires ATP and ATP hydrolysis but does not require mitochondrial membrane potential. Moreover, we obtained evidence for a role for the TOM complex, which mediates protein translocation across the MOM during import of proteins into the organelle, as a retrotranslocation channel in the MOM in the MAD pathway. Thus, our studies provide additional support for the model that proteins within mitochondria are substrates for MAD-dependent mitochondrial quality control and the first mechanistic insight into retrotranslocation of MAD targets from the matrix.
These studies were carried out using two MAD substrates, Kgd1p and Pim1p, that are tagged at their chromosomal loci using GFP. We confirmed that Kgd1p-GFP is faithfully imported into the mitochondrial matrix: it localizes exclusively to mitochondria in living yeast cells and is enriched in mitochondria during subcellular fractionation. Submitochondrial localization studies indicate that Kgd1p-GFP is resistant to protease-mediated degradation of MOM proteins and is therefore not mistargeted to the MOM. Moreover, rupture of the MOM under conditions where the MIM remains intact, does not result in release of Kgd1p-GFP from the organelle. Thus, Kgd1p-GFP is also not mistargeted to the IMS. Since there is no evidence that Kgd1p-GFP is mistargeted to the MOM or IMS, the observed release is not due to mitoTAD or Oma1p function42 in release of mistargeted proteins from the protein translocation machinery in the MOM or MIM. Rather, our findings support the model that Kgd1p-GFP can be retrotranslocated across both the MOM and MIM in vitro.
In addition, we obtained evidence for a role for MAD in retrotranslocation of Kgd1p-GFP from the mitochondrial matrix. We confirmed previous findings30 that Kgd1p-GFP is a MAD substrate: deletion of DOA1 results in increased ubiquitination of mitochondrial Kgd1p-GFP and increased accumulation of Kgd1p-GFP in mitochondria. Importantly, we found that Kgd1p-GFP, but not Cit1p-GFP (an abundant matrix protein that has not been identified as a MAD substrate), undergoes ATP-dependent, ∆Ψ-independent release from mitochondria in vitro. Thus, we obtained evidence that retrotranslocation of Kgd1p-GFP across both the MOM and MIM is specific for MAD substrates.
Consistent with this, we found that deletion of DOA1 results in increased release of Kgd1p-GFP from isolated mitochondria. This finding indicates that Doa1p and Doa1p-mediated targeting of the segregase complex to mitochondria does not have a direct role in retrotranslocation of MAD substrates across mitochondrial membranes. Rather, the increase Kgd1p retrotranslocation observed in mitochondria isolated from doa1∆ compared to WT cells may be due to the increased steady state levels of Kgd1p in mitochondria of doa1∆ cells.
Finally, retrotranslocation of damaged ER proteins from the ER lumen in ERAD is mediated by a Hrd1p43. Our findings support a role for the TOM complex as a protein retrotranslocation tunnel across the MOM in MAD. Specifically, we find that blocking the Tom40p channel using mPEG or destabilization of the TOM complex by deletion of TOM5 inhibits MAD-dependent retrotranslocation in mitochondria isolated from doa1∆ cells. Interestingly, this effect is not observed in mitochondria isolated from yeast that express DOA1. Deletion of DOA1 results (1) in increased ubiquitination and steady state levels of mitochondria Kgd1p-GFP and (2) increased sensitivity to oxidative stress. Therefore, it is possible that residual TOM complex activity in mitochondria isolated from tom5∆ cells or treated with mPEG is sufficient to support basal level retrotranslocation under non-stressed conditions. Nonetheless, our studies support a role for the TOM complex function as the retrotranslocation tunnel in the MOM in MAD.
Since import of proteins into mitochondria requires ∆Ψ, it is surprising that MAD-dependent retrotranslocation from the matrix does not require ∆Ψ. On the other hand, loss of ∆Ψ is a hallmark of damaged mitochondria44 and triggers mitophagy in mammalian systems45. Our finding that retrotranslocation in MAD does not require ∆Ψ indicates that MAD has the capacity to repair damaged mitochondria, including those with no ∆Ψ. However, in contrast to mitophagy, which senses mitochondrial damage by loss of ∆Ψ and is activated by loss of ∆Ψ, MAD senses mitochondrial protein damage but is neither activated nor inactivated by loss of ∆Ψ.
On the other hand, the essential function of the machinery for import of proteins into mitochondria is to ensure that nuclear-encoded mitochondrial proteins, which represent >95% of the proteins within mitochondria, are faithfully imported into the organelle. Therefore, the expectation is that translocation of proteins across the MOM and MIM are irreversible. However, we find that 1) MAD substrates can be retrotranslocated across both mitochondrial membranes, 2) retrotranslocation is dependent upon ATP and ATP hydrolysis and is therefore an active transport process, and 3) an import channel in the MOM is also an export channel in MAD. This raises a fundamental question: how can the TOM complex mediate import and export of proteins from mitochondria without decreasing the efficiency or efficacy of the machinery for import of proteins into mitochondria?
Several observations indicate that retrotranslocation and export of ubiquitinated, damaged mitochondrial proteins in MAD is compatible with import of proteins into the organelle. First, only a small percent of total mitochondrial Kgd1p or Pim1p is retrotranslocated from mitochondria in vitro. Indeed, only 1–2% and 2–7% of the total Kgd1p is released from mitochondria isolated from WT or doa1∆ cells, respectively. Thus, retrotranslocation does not result in large-scale loss of Kgd1p from the mitochondrial matrix. Rather, it presumably only targets damaged mitochondrial proteins. Second, retrotranslocation by the Tom40p channel does not affect the integrity of the organelle. Thus, retrotranslocation of mitochondrial proteins by MAD and the TOM complex does not damage mitochondria or interfere with mitochondrial biogenesis. Third, retrotranslocation across the MIM and MOM is selective: we detect retrotranslocation of two matrix proteins that are known to be oxidation-sensitive and MAD substrates, but not of another abundant matrix protein, Cit1p (Fig. 1).
Finally, studies from our group and others indicate that MAD is critical for the cellular response to oxidative stress21,23,30. However, since there are proteases in mitochondria that reduce the toxicity of unfolded proteins and disrupt protein complexes with folding errors17, it is not clear what function MAD provides in mitochondrial protein quality control for intramitochondrial proteins beyond that provided by mitochondrial proteases. MAD may function downstream of mitochondrial proteases: it may target large proteolytic fragments of misfolded mitochondrial proteins or proteins released by protease-mediated degradation of misfolded mitochondrial protein complexes for degradation by the proteasome. In addition, we find that Pim1p, the Lon protease of the mitochondrial matrix, is a MAD substrate that undergoes retrotranslocation across the MOM and MIM. Consistent with this, DOA1 genetically interacts with PIM1, the Lon protease gene23. Thus, MAD may also function in proteostatic control of proteins that are not substrates for mitochondrial proteases, including the proteases themselves.
Overall, our studies support a model for MAD function in protein quality control of intramitochondrial proteins. According to this model, damaged and unfolded proteins in the mitochondrial matrix are retrotranslocated across the MIM. The TOM complex then mediates retrotranslocation of unfolded matrix proteins across the MOM. Retrotranslocation of MAD substrates from mitochondria is dependent upon ATP and ATP hydrolysis but does not require a membrane potential across the MIM. Finally, the segregase complex mediates transfer of retrotranslocated proteins from the surface of the organelle to the proteasome for degradation.
While our studies provide the first mechanistic understanding of retrotranslocation of proteins in the MAD pathway, they also raise fundamental questions. What protein(s) mediates retrotranslocation across the MIM? Does retrotranslocation occur at contact sites between the MOM and MIM? How are unfolded proteins maintained in an unfolded, retrotranslocation-competent state prior to membrane retrotranslocation? What is the signal for retrotranslocation across the MIM and MOM? How are different mitochondrial proteostasis processes coordinated and how do they contribute to overall quality control in the organelle? These and other fundamental questions regarding the MAD pathway are the topic of ongoing and future studies.
Methods
Yeast growth conditions
All S. cerevisiae strains were derived either from the wild-type strain BY4741 or from SUB59235. All strains are listed in Table 1. Cells were grown in glucose-based rich media (yeast extract/peptone/dextrose, YPD) for growth rate measurement, grown in synthetic complete medium without uracil (SC-Ura) to mid-logarithmic phase (optical density OD600 = 0.1–0.3) for the measurement of mitochondrial redox state using mito-roGFP1, and grown in SC medium for all other experiments. All cells were grown at 30 °C with 200 rpm shaking.
Yeast strain construction
To generate deletion strains, the target genes were replaced with cassettes containing the selectable marker HphMX6 using homologous recombination. Cassettes containing HphMX6 flanked by 40 bp of homology to regions immediately upstream and downstream of the target gene were amplified from plasmids pCY3090-0246 (Addgene plasmid # 36231). The PCR fragments were transformed into cells using the lithium acetate method. Transformants containing selectable markers were selected on YPD containing 50 µg/ml Hygromycin B (Invitrogen, 10687010).
tom40130/138C strains containing Asn-to-Cys and Ser-to-Cys alterations at amino acid 130 and 138 from mutations of AA-to-TG at nucleotide 388/389 and C-to-G at nucleotide 41340,47 were generated in SUB592 background using the CRISPR/Cas9 tool box from Dr. Tom Ellis’ lab48. This tool box contains a gRNA expression plasmid (pWS082, Addgene plasmid # 90516) and a CRISPR/Cas9 expression plasmid with a Hygromycin selectable marker (pWS175, Addgene plasmid # 90962). gRNA sequences were designed as a 20-mer guide sequence immediately preceding a PAM sequence (5′-NGG-3′) near the mutation sites, and then generated using two annealed 26 bp oligonucleotides containing the GACT overhang at the 5′ and GTTT at the 3′ for Esp3I Golden Gate assembly into pWS082. An additional TT was included to complete the HDV ribozyme sequence:
5′ GACTTTNNNNNNNNNNNNNNNNNNNN 3′
3′ AANNNNNNNNNNNNNNNNNNNNCAAA 5′
These oligonucleotides were hybridized and ligated into the gRNA expression vector pWS082 using Golden Gate assembly. Next, pWS082 containing sgRNA and pWS175 were linearized by digestion with EcoRV and Esp3I, respectively, and the 1000 bp-fragment from pWS082 containing sgRNA and the backbone (10000 bp-fragment) from pWS175 were gel purified. Donor oligonucleotides containing the 3 mutations of tom40 130/138 C, the PAM sequence mutation that eliminates the PAM sequence but does not change amino acid sequence, and guide sequence flanked by 40 bp of homology to regions immediately upstream and downstream of the guide sequence were generated and hybridized. Yeast cells were transformed with linearized pWS175, sgRNA containing fragment from pWS082 and donor DNA, and the transformants were selected on YPD plates containing 50 µg/ml Hygromycin B. Positive transformants were confirmed by sequencing.
Growth rate analysis
Yeast cells were grown to mid-log phase in YPD (OD600 = 0.3–0.5) and then diluted to OD600 = 0.07. 10 μL of these cultures were added to 200 μL YPD or YPD containing 2.5 mM paraquat (PQ) in a well of a 96-well flat-bottom plate (Corning, Corning, NY). The optical density of the culture (OD600) was measured every 20 min for 60 h using a plate reader (Tecan Infinite M200, Research Triangle Park, NC). Each strain was plated in sextuplicate. The growth curves were plotted from the average OD600 over time, and maximum growth rate (slope) calculated as the greatest change in OD600 over a 240-min interval in 60 h, using linear regression in Magellan software.
Imaging
Mito-roGFP1 imaging was performed as described previously with minor modification49,50. Cells were transformed with a plasmid bearing the mito-roGFP1 sequence using the lithium acetate method. Cells containing mito-roGFP1 plasmids were grown in SC-Ura medium in the presence or absence of 2.5 mM PQ for 8 h to mid-log phase. Images were acquired on an Axioskop 2 microscope with a 100x/1.4 NA Plan-Apochromat objective (Zeiss, Thornwood, NY) and an Orca ER cooled charge-coupled device (CCD) camera (Hamamatsu Photonics, Hamamatsu City, Japan) using excitation by an LED light source (CoolLED pE-4000, Andover, UK) at 365 and 470 nm for the oxidized and reduced form of roGFP, respectively. All channels were acquired with a modified GFP filter (Zeiss filter 46 HE without excitation filter, dichroic FT 515, emission 535/30). Images were collected through the entire cell depth with 21 z-sections at 0.3-µm intervals and were deconvolved using a constrained iterative restoration algorithm with the following parameters: 507 nm excitation wavelength, 60 iterations, 100% confidence limit (Volocity 5.5, Quorum Technologies). After subtracting background and thresholding, the reduced/oxidized ratio was calculated by dividing the intensity of the reduced channel (λex = 470 nm, λem = 525 nm) by the intensity of the oxidized channel (λex = 365 nm, λem = 525 nm) in Volocity software.
To detect the localization of Kgd1p or Pim1p, cells with Kgd1p-GFP or Pim1p-GFP were incubated in SC with 100 nM MitoTracker Red CM-H2Xros (ThermoFisherScientific, M7513) for 30 min. MitoTracker Red-stained cells were washed with SC for 3 times and imaged as previously described using excitation using a 561 nm LED for MitoTracker Red and 470 nm LED for GFP with a dual eGFP/mCherry cube (59222, Chroma, Bellows Falls, VT). Images were deconvolved using a constrained iterative restoration algorithm with calculated PSFs assuming 507 nm and 610 nm emission for GFP and MitoTracker Red, respectively, using 60 iterations and a 100% confidence criterion for termination.
Isolation of mitochondria
Mitochondria were isolated as described previously51,52. 500 mL cells were grown in 2-L flasks to mid-log phase in SC or SC containing 2.5 mM PQ for 8 h at 30 °C with shaking at 200 rpm. Cells were collected by centrifugation at 1500 × g for 5 min at 4 °C and washed with water once. Supernatants were removed and the weight of the “wet” cell pellet was measured. Cells were then incubated in 20 mL Tris-DTT buffer (0.1 M Tris-SO4, pH 9.4 and 10 mM DTT) ( ~ 5 ml/g yeast wet weight) for 15 min at 30 °C with shaking, washed once with 20 mL SP buffer (1.2 M Sorbitol and 20 mM KPi, pH 7.4), and incubated with 20 mL SP buffer containing Zymolyase 20 T (Seikagaku Corporation, Tokyo, Japan) (7.5 mg/g yeast wet weight) at 30 °C for 40 min with shaking to remove cell walls. Spheroplasts were collected by centrifugation at 4500 × g at 4 °C for 5 min, washed once with 20 mL ice-cold SEH buffer (0.6 M Sorbitol, 20 mM HEPES-KOH, pH 7.4, 2 mM MgCl2) containing a protease inhibitor cocktail (PI-1: 0.5-mg/ml Pepstatin A, 0.5 µg/ml Chymostatin, 0.5 µg/ml Antipain, 0.5 µg/ml Leupeptin, and 0.5 µg/ml Aprotinin; PI-2: 10 µM Benzamidine–HCl and 1 µg/ml 1,10-Phenanthroline; 1 mM PMSF), and then homogenized using 15 forceful strokes of a pre-chilled glass/glass Dounce homogenizer (Wheaton Science Products, Millville, NJ). The homogenate was subjected to low-speed centrifugation (1500 × g) for 5 min, and the supernatant obtained was subjected to high-speed centrifugation (12,000 × g) for 10 min at 4 °C. The resulting pellet was resuspended in 1 mL ice-cold SEH buffer containing protease inhibitor cocktails. To further remove debris, 1 ml of mitochondria-enriched fraction was subjected to 2 rounds of low-speed centrifugation (700 × g for 5 min and 1500 × g for 5 min) at 4 °C using a benchtop microcentrifuge. The supernatant obtained was centrifuged at 12,000 × g for 10 min at 4 °C, and the mitochondria-contained pellet was resuspended in 200 µL ice-cold SEH buffer. Mitochondrial protein concentration was measured by BCA protein assay (Pierce, 23225). To remove mitochondrial outer membrane proteins, isolated mitochondria were incubated in the presence or absence of proteinase K (10 µg/mL) for 30 min at 0°C. After inactivation of proteinase K by addition of protease inhibitors, mitochondria were separated from the proteinase K-containing solution. To generate osmotic stress, mitochondria were incubated with hypotonic buffer (20 mM HEPES-KOH, pH 7.4, 2 mM MgCl2) for 20 min at 0 °C, and then separated from the hypotonic buffer. The proteins in the supernatants were precipitated with 10% TCA for 1 h at −20 °C and washed with acetone at −20 °C.
Mitochondrial protein releasing assay
Released mitochondrial proteins were collected as previously described with some modification32. In the time-course releasing assay in Fig. 1A, D, isolated mitochondria were incubated in the assay buffer (0.6 M Sorbitol, 20 mM HEPES-KOH, pH 7.4, 2 mM MgCl2, 5 mM Methionine, 10 mM KPi pH 7.4) and a protease inhibitor cocktail at 30 °C for 10, 30, and 60 min. In other experiments, isolated mitochondria were incubated in the assay buffer containing 2 mM ATP, 2 mM NADH and a protease inhibitor cocktail at 30 °C for 30 min. To treat isolated mitochondria with non-hydrolyzed ATP, 2 mM ATP-γ-S was added into assay buffer. To inhibit mitochondrial membrane potential, isolated mitochondria were treated with 5 µg/ml valinomycin in assay buffer. To modify cysteine residues in Tom40p channels of mitochondria, isolated mitochondria were treated with 1 mM mPEG-Mal (Sigma-Aldrich, 63187) in assay buffer containing 2 mM ATP, 2 mM NADH and a protease inhibitor cocktail at 30 °C for 30 min. Mitochondrial pellets were then collected by centrifuging at 12,000 × g for 5 min at 4 °C. Supernatant proteins were precipitated with 10% TCA for 1 h at −20 °C and washed with acetone at −20 °C. Mitochondrial pellet and TCA-precipitated supernatant proteins were then resuspended in 2xSDS sample buffer (20% Glycerol, 120 mM Tris/HCl pH 6.8, 4% SDS, 180 mM DTT, 0.02% bromophenol blue) and boiled for 5 min.
Pull-down of His-tagged proteins
400 µg of mitochondrial proteins were lysed in 500 ml Cell Lysis Buffer (CLB, 50 mM Tris pH 8.0, 300 mM NaCl, 0.1% Tween 20, protease inhibitor cocktail, 10 mM NEM) containing 250 µl glass beads with vortex for 4 min at 4 °C. The lysates were separated from beads and debris by centrifuging at 12,000 × g and incubated with 50 ml CLB-washed Ni2 + -NTA magnetic beads (HisPur Ni-NTA Magnetic Beads, Thermo Scientific, Grand Island, NY) for 1 h at 4 °C with rotate. Beads with His-tagged proteins were separated from other proteins by placing in the Magnetic Separation Rack for 1 min, and were washed with 500 ml Washing Buffer (50 mM Tris pH 8.0, 300 mM NaCl, 0.1% Tween 20, 10 mM imidazole, protease inhibitor cocktail) for 3 times. Proteins were eluted by incubating with 60 ml of Elution Buffer (50 mM Tris pH 8.0, 300 mM NaCl, 0.1% Tween 20, 300 mM imidazole, protease inhibitor cocktail) for 5 min with rotation. Protein samples were added 4xSDS Sample buffer and subjected to polyacrylamide gel electrophoresis and western blots.
Immunoprecipitation
Cells were lysed in 500 µl Cell Lysis Buffer (50 mM Tris pH 8.0, 150 mM NaCl, 10 mM EDTA, 1% Triton X-100, 10% glycerol, protease inhibitor cocktail, 10 mM NEM) containing 250 µl glass beads with vortex for 5 min at 4 °C. The lysates were separated from beads and debris by centrifuging at 12,000 × g for 5 min at 4 °C and incubated with 10 µg anti-Myc antibodies for 16 h at 4 °C with low-speed rotation. The lysates with antibodies were incubated with Pierce Protein A Magnetic Beads that were washed with Bead Washing Buffer (1x TBS, 0.1% Tween-20) for 1 h at 4 °C. Beads with antibody-bound proteins were separated from other proteins by placing in the Magnetic Separation Rack for 1 min, and were washed with 500 µl ice-cold Washing Buffer (50 mM Tris pH 8.0, 150 mM NaCl, 1% Triton X-100, 10% glycerol, protease inhibitor cocktail) for 3 times. Proteins were eluted by incubating with 60 µl of 1xSDS Sample Buffer (10% Glycerol, 60 mM Tris/HCl pH 6.8, 2% SDS, 90 mM DTT, 0.01% bromophenol blue) containing protease inhibitor cocktail and subjected to polyacrylamide gel electrophoresis and western blots.
Western blot analysis
Western blot analysis was performed using standard procedures on PVDF membranes (Immobilon-FL; EMD Millipore, Billerica, MA). Equal amounts of protein lysates were loaded onto a SDS-PAGE gel containing 0.5% trichloroethanol (TCE) for protein detection. Prior to transfer, the gel was exposed to UV light (300 nm) for 2.5 min to activate protein-crosslinking activity of TCE53. TCE-crosslinked proteins were detected by exposure of gels to 300 nm illumination for 4 s using a ChemiDoc™ MP Imaging System (Bio-Rad, Hercules, CA). Total proteins crosslinked by TCE were used as loading controls. The gel was then transferred to a PVDF membrane (Immobilon-FL; EMD Millipore, Billerica, MA). After transfer, the PVDF membrane was blocked with 5% skim milk for 1 h, incubated with 5% skim milk containing primary antibodies for 16 h, and secondary antibodies for 1 h. Proteins in blots were detected using Luminata Forte Western HRP substrate (MilliporeSigma, Burlington, MA) and the ChemiDoc™ MP Imaging System (Bio-Rad, Hercules, CA). The primary antibodies used in these studies were mouse monoclonal antibodies against GFP (#11814460001, Roche) and Myc (Clone 9E10), and rabbit polyclonal antibody against Cdc48p (#62-303, As One International Inc), hexokinase (LS‑C59302, LSBio, MA) Cyb2p, Cit1p, Tom70p and Tom40p (415 T, a gift from Dr. Carla Koehler).
Statistics and Reproducibility
For multiple group comparisons, p values were determined by a 1-way ANOVA with Dunnett’s or Sidak’s test. Bar graphs show the mean and SEM. For SuperPlots54, each color denotes an independent biological replicate. Small dots represent individual data points, and triangles represent the mean of the technical replicates within each biological replicate. p values were determined by a Repeated Measurement 1-way ANOVA with Sidak’s test. All statistical analysis was conducted using GraphPad Prism9 (GraphPad Software). For all tests, p values are classified as follows: *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All source data used to produce the graphs in this work are provided in Supplementary Data. Uncropped blots/gels are provided in Supplementary Fig. 6. Further data and materials of this study are available from the corresponding author upon reasonable request.
References
Spinelli, J. B. & Haigis, M. C. The multifaceted contributions of mitochondria to cellular metabolism. Nat. Cell Biol. 20, 745–754 (2018).
Sharma, A., Smith, H. J., Yao, P. & Mair, W. B. Causal roles of mitochondrial dynamics in longevity and healthy aging. EMBO Rep. 20, e48395 (2019).
Chacinska, A., Koehler, C. M., Milenkovic, D., Lithgow, T. & Pfanner, N. Importing mitochondrial proteins: machineries and mechanisms. Cell 138, 628–644 (2009).
Guo, R., Zong, S., Wu, M., Gu, J. & Yang, M. Architecture of human mitochondrial respiratory megacomplex I2III2IV2. Cell 170, 1247–1257.e1212 (2017).
Kawamata, H. & Manfredi, G. Proteinopathies and OXPHOS dysfunction in neurodegenerative diseases. J. Cell Biol. 216, 3917–3929 (2017).
Ng, M. Y. W., Wai, T. & Simonsen, A. Quality control of the mitochondrion. Dev. Cell 56, 881–905 (2021).
Oh, E., Akopian, D. & Rape, M. Principles of ubiquitin-dependent signaling. Annu. Rev. Cell Dev. Biol. 34, 137–162 (2018).
Ravanelli, S., den Brave, F. & Hoppe, T. Mitochondrial quality control governed by ubiquitin. Front Cell Dev. Biol. 8, 270 (2020).
Ling, Q. et al. Ubiquitin-dependent chloroplast-associated protein degradation in plants. Science 363, https://doi.org/10.1126/science.aav4467 (2019).
Schmidt, O. et al. Endosome and Golgi-associated degradation (EGAD) of membrane proteins regulates sphingolipid metabolism. EMBO J. 38, e101433 (2019).
Ling, Q. et al. The chloroplast-associated protein degradation pathway controls chromoplast development and fruit ripening in tomato. Nat. Plants 7, 655–666 (2021).
Taylor, E. B. & Rutter, J. Mitochondrial quality control by the ubiquitin-proteasome system. Biochem Soc. Trans. 39, 1509–1513 (2011).
Brodsky, J. L. Cleaning up: ER-associated degradation to the rescue. Cell 151, 1163–1167 (2012).
Christianson, J. C. & Ye, Y. Cleaning up in the endoplasmic reticulum: ubiquitin in charge. Nat. Struct. Mol. Biol. 21, 325–335 (2014).
Izawa, T., Park, S. H., Zhao, L., Hartl, F. U. & Neupert, W. Cytosolic protein Vms1 links ribosome quality control to mitochondrial and cellular homeostasis. Cell 171, 890–903 e818 (2017).
Martensson, C. U. et al. Mitochondrial protein translocation-associated degradation. Nature 569, 679–683 (2019).
Song, J., Herrmann, J. M. & Becker, T. Quality control of the mitochondrial proteome. Nat. Rev. Mol. Cell Biol. 22, 54–70 (2021).
Weidberg, H. & Amon, A. MitoCPR-A surveillance pathway that protects mitochondria in response to protein import stress. Science 360, https://doi.org/10.1126/science.aan4146 (2018).
Shang, F. & Taylor, A. Ubiquitin-proteasome pathway and cellular responses to oxidative stress. Free Radic. Biol. Med. 51, 5–16 (2011).
Ye, Y., Tang, W. K., Zhang, T. & Xia, D. A mighty “Protein Extractor” of the cell: structure and function of the p97/CDC48 ATPase. Front Mol. Biosci. 4, 39 (2017).
Heo, J. M. et al. A stress-responsive system for mitochondrial protein degradation. Mol. Cell 40, 465–480 (2010).
Tanaka, A. et al. Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J. Cell Biol. 191, 1367–1380 (2010).
Wu, X., Li, L. & Jiang, H. Doa1 targets ubiquitinated substrates for mitochondria-associated degradation. J. Cell Biol. 213, 49–63 (2016).
Xu, S., Peng, G., Wang, Y., Fang, S. & Karbowski, M. The AAA-ATPase p97 is essential for outer mitochondrial membrane protein turnover. Mol. Biol. Cell 22, 291–300 (2011).
Azuma, Y. et al. Identification of ter94, Drosophila VCP, as a strong modulator of motor neuron degeneration induced by knockdown of Caz, Drosophila FUS. Hum. Mol. Genet. 23, 3467–3480 (2014).
Qiu, L. et al. Structure and function of the PLAA/Ufd3-p97/Cdc48 complex. J. Biol. Chem. 285, 365–372 (2010).
Deshwal, S., Fiedler, K. U. & Langer, T. Mitochondrial proteases: multifaceted regulators of mitochondrial plasticity. Annu. Rev. Biochem. 89, 501–528 (2020).
Nishimura, K., Kato, Y. & Sakamoto, W. Essentials of proteolytic machineries in chloroplasts. Mol. Plant 10, 4–19 (2017).
Ruggiano, A., Foresti, O. & Carvalho, P. Quality control: ER-associated degradation: protein quality control and beyond. J. Cell Biol. 204, 869–879 (2014).
Liao, P. C., Wolken, D. M. A., Serrano, E., Srivastava, P. & Pon, L. A. Mitochondria-associated degradation pathway (MAD) function beyond the outer membrane. Cell Rep. 32, 107902 (2020).
Li, J. et al. The CDC48 complex mediates ubiquitin-dependent degradation of intra-chloroplast proteins in plants. Cell Rep. 39, 110664 (2022).
Bragoszewski, P. et al. Retro-translocation of mitochondrial intermembrane space proteins. Proc. Natl. Acad. Sci. USA. 112, 7713–7718 (2015).
Bragoszewski, P., Gornicka, A., Sztolsztener, M. E. & Chacinska, A. The ubiquitin-proteasome system regulates mitochondrial intermembrane space proteins. Mol. Cell Biol. 33, 2136–2148 (2013).
Johnson, E. S., Ma, P. C., Ota, I. M. & Varshavsky, A. A proteolytic pathway that recognizes ubiquitin as a degradation signal. J. Biol. Chem. 270, 17442–17456 (1995).
Spence, J. et al. Cell cycle-regulated modification of the ribosome by a variant multiubiquitin chain. Cell 102, 67–76 (2000).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21, 921–926 (2003).
Wachter, C., Schatz, G. & Glick, B. S. Protein import into mitochondria: the requirement for external ATP is precursor-specific whereas intramitochondrial ATP is universally needed for translocation into the matrix. Mol. Biol. Cell 5, 465–474 (1994).
Dietmeier, K. et al. Tom5 functionally links mitochondrial preprotein receptors to the general import pore. Nature 388, 195–200 (1997).
Schmitt, S. et al. Role of Tom5 in maintaining the structural stability of the TOM complex of mitochondria. J. Biol. Chem. 280, 14499–14506 (2005).
Gornicka, A. et al. A discrete pathway for the transfer of intermembrane space proteins across the outer membrane of mitochondria. Mol. Biol. Cell 25, 3999–4009 (2014).
Cocheme, H. M. & Murphy, M. P. Complex I is the major site of mitochondrial superoxide production by paraquat. J. Biol. Chem. 283, 1786–1798 (2008).
Krakowczyk, M. et al. OMA1 protease eliminates arrested protein import intermediates upon mitochondrial depolarization. J. Cell Biol. 223, https://doi.org/10.1083/jcb.202306051 (2024).
Baldridge, R. D. & Rapoport, T. A. Autoubiquitination of the Hrd1 ligase triggers protein retrotranslocation in ERAD. Cell 166, 394–407 (2016).
Zorova, L. D. et al. Mitochondrial membrane potential. Anal. Biochem 552, 50–59 (2018).
Quinn, P. M. J., Moreira, P. I., Ambrosio, A. F. & Alves, C. H. PINK1/PARKIN signalling in neurodegeneration and neuroinflammation. Acta Neuropathol. Commun. 8, 189 (2020).
Young, C. L., Raden, D. L., Caplan, J. L., Czymmek, K. J. & Robinson, A. S. Cassette series designed for live-cell imaging of proteins and high-resolution techniques in yeast. Yeast 29, 119–136 (2012).
Qiu, J. et al. Coupling of mitochondrial import and export translocases by receptor-mediated supercomplex formation. Cell 154, 596–608 (2013).
Shaw, W. M. et al. Engineering a model cell for rational tuning of GPCR signaling. Cell 177, 782–796.e727 (2019).
Liao, P. C., Yang, E. J. & Pon, L. A. Live-cell imaging of mitochondrial redox state in yeast cells. STAR Protoc. 1, 100160 (2020).
Vevea, J. D., Alessi Wolken, D. M., Swayne, T. C., White, A. B. & Pon, L. A. Ratiometric biosensors that measure mitochondrial redox state and ATP in living yeast cells. J. Vis. Exp. 22, 50633 (2013).
Liao, P. C., Bergamini, C., Fato, R., Pon, L. A. & Pallotti, F. Isolation of mitochondria from cells and tissues. Methods Cell Biol. 155, 3–31 (2020).
Liao, P. C., Boldogh, I. R., Siegmund, S. E., Freyberg, Z. & Pon, L. A. Isolation of mitochondria from Saccharomyces cerevisiae using magnetic bead affinity purification. PLoS One. 13, e0196632 (2018).
Ladner, C. L., Yang, J., Turner, R. J. & Edwards, R. A. Visible fluorescent detection of proteins in polyacrylamide gels without staining. Anal. Biochem. 326, 13–20 (2004).
Lord, S. J., Velle, K. B., Mullins, R. D. & Fritz-Laylin, L. K. SuperPlots: communicating reproducibility and variability in cell biology. J. Cell Biol. 219, https://doi.org/10.1083/jcb.202001064 (2020).
Acknowledgements
We thank members of the Pon laboratory for technical assistance and valuable discussion, Dr. Steven P. Gygi (Harvard Medical School, Boston) for providing the SUB592 strain, and Dr. Carla Koehler (University of California, Los Angeles) for providing Tom40p antibody. This work was supported by awards from the National Institutes of Health (NIH) (GM122589 and AG051047) to L.P., and the National Science and Technology Council in Taiwan (NSTC 111-2311-B-007-013-MY3) and the Yushan Fellow Program by the Ministry of Education (MOE), Taiwan (MOE-111-YSFAG-0002-001-P1) to P.L.; We thank Dr. Theresa Swayne in the Confocal and Specialized Microscopy Shared Resource the Herbert Irving Comprehensive Cancer Center at Columbia University Medical Center for valuable discussion. The Confocal and Specialized Microscopy Shared Resource is supported in part by an award from the NIH/NCI (5 P30 CA13696).
Author information
Authors and Affiliations
Contributions
Conceptualization: P.L. and L.A.P.; validation: P.L., C.A.T., T.L., C.H., K.F., and I.B.; formal analysis: P.L., C.A.T., T.L., C.H., K.F., and I.B.; investigation: P.L., C.A.T., T.L., C.H., K.F., and I.B.; writing-original draft: P.L.; writing – review & editing: P.L. and L.A.P.; supervision: P.L. and L.A.P.; project administration: P.L. and L.A.P.; funding acquisition: P.L. and L.A.P.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Felix Boos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Dr Giulia Bertolin and Dr Ophelia Bu.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liao, PC., Lin, TY., Tsang, C.A. et al. Tom40 functions as a channel for protein retrotranslocation in the mitochondria-associated degradation (MAD) pathway. Commun Biol 8, 1122 (2025). https://doi.org/10.1038/s42003-025-08549-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08549-z