Abstract
Guangdong province has diverse local goose breeds with unique characteristics, but there is limited research on their genetic diversity, formation history and breed-specific genes. This study uses whole-genome sequencing to analyze 120 local geese representing four officially recognized breeds and two lines of the folk breed Sanzhou Black goose, as well as five museum goose samples in the region, combined with 56 geese whole genome data from NCBI, with the aim of understanding the genetic diversity and relationships of Guangdong geese, identifying the history of breed formation, and discovering genes linked to various traits in these geese. The results show a considerable degree of genetic diversity in Guangdong local geese. The museum Sanzhou Black goose exhibit multiple ancestral components, and the Magang goose may be derived from the Sanzhou Black goose. A multitude of genes have been identified as related to various functional phenotypes in different Guangdong geese, such as fat deposition (ACER3), feather color (ACTA2), skin pigmentation (GDA), and body growth and development (ARHGAP21), collectively defining the distinctive attributes of indigenous breeds. These results provide a way to study the history of breed formation and a basis for selective breeding or utilization targeting goose breeds in Guangdong.
Similar content being viewed by others
Introduction
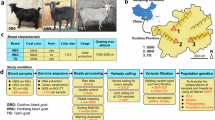
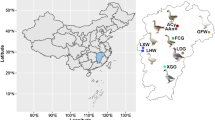
Geese, domesticated in China for over 7000 years and currently one of the most widely used farm animals1, have long played a significant economic role. Both artificial and natural selection have resulted in the development of a wide range of phenotypes and traits in domestic geese. China is home to a considerable number of domestic goose breeds, with 30 native breeds currently identified. Among these, Guangdong province alone is home to five distinct local goose breeds, each exhibiting significant phenotypic differences. Guangdong local goose breeds have rich and diverse phenotypes, including different body size2, plumage colors and fat deposition3. Additionally, as a representative of short-day breeding animal, the Magang goose (MG) is frequently employed to examine the influence of light on reproductive activity in seasonally breeding animals4. Therefore, the study of breed characteristics of various Guangdong local goose breeds is important for genetic improvement of domestic geese and conservation of germplasm resources.
The assessment of genetic diversity, structure, and relationships is crucial for optimizing conservation strategies and sustainable breeding programs. Recent genomic studies have provided insights into Chinese goose breeds. For example, Zheng et al. found that the Lingxian White goose exhibited a lower genetic diversity and a higher inbreeding coefficient than the Guangfeng White goose and the Fengcheng Gray goose using whole genome sequencing5. Jing et al. studied the genetic diversity and genetic structure of domestic geese and found that the genetic diversity of Chinese geese was higher than that of European geese, and verified the accuracy of the classification of Chinese goose germplasm resources6. However, significant gaps remain in our knowledge of Guangdong’s indigenous genetic resources. Notably, current investigations have predominantly focused on selective sweep analysis of mainstream Guangdong breeds2,7. Additionally, there are a number of additional folk breeds in Guangdong, including the Sanzhou Black goose (SZH) and Yangchun white goose (YCB), which warrant further investigation. Consequently, there remains a paucity of systematic exploration of the overall genetic diversity and structure of Guangdong goose breeds.
The temporal dynamics of indigenous breed formation have emerged as a focal subject in domestication studies. The integration of temporal genomic perspectives through historical samples has emerged as a transformative approach for reconstructing domestication histories, particularly in resolving long-standing debates about breed origin. This methodology has successfully elucidated key domestication transitions in chicken, including the discovery of domestication centers8 and the fixation time of domestication gene TSHR9. Similarly, the analysis of historical samples from Bronze Age samples has confirmed that all ancient domestic cattle in northern China belonged to the taurine cattle10. Furthermore, research into ancient DNA has revealed the history of cattle being introduced to the Americas11. Using historical samples, researchers found that contemporary breeds may have been formed in recent centuries12. In the case of the Guangdong goose, the MG was historically attributed to SZH/Yangjiang goose crosses but lacked molecular validation. Therefore, the genomic information of museum geese samples is expected to elucidate the structural changes and breed formation within these populations.
Despite the outstanding germplasm characteristics of the Guangdong local breeds have received great degree of attention, particularly the well-characterized Shitou goose (ST), critical gaps persist in understanding the broader genetic diversity, genetic structure, and genetic relationships of local breeds. In addition, the breed-specific genetic traits in this region remain to be thoroughly examined, which hinders the precise identification and conservation of germplasm resources. In this study, we perform whole-genome sequencing on 120 modern samples and five museum samples. The modern samples including four officially recognized local breeds (ST, Wuzong goose (WZA), MG, YCB), two lines of the folk breed SZH (Maojing (MJ) and Gaoyao (GY)). And the five museum samples representing the one Swan goose (G), two Shitou geese (G-ST), and two Sanzhou Black geese (G-SZH). To comprehensively understand the genetic structure and breed formation history of Guangdong geese, these data were combinedly analyzed with the extant data of 47 modern domesticated geese and nine modern wild Swan goose in public databases. The breed-specific related single nucleotide polymorphisms (SNPs) and Fixation index (FST) analysis were used to determine genomic SNPs and genes related to characteristics of the goose, which will contribute to further genetic and breeding research on goose breeds of Guangdong.
Results
Sequencing and data quality
A total of 24.87–27.27 Gb raw base was obtained for each modern sample, with an average depth between 19.41–21.28× across populations (Supplementary Table S1), providing sufficient data for subsequent analysis. Extant 7 ST, 10 MG, and 10 ZI genome sequencing data (10.10–10.11 Gb raw base for each sample) were obtained from the public database (National Center for Biotechnology Information (NCBI)) (Supplementary Table S1). Additionally, whole genome sequencing data (10.12--17.57 Gb raw base for each sample) for ten Roman goose (RM), ten Hortobágy goose (HT), and nine wild Swan goose (HY) was also downloaded and deployed for joint analysis (Supplementary Table S1). The results of the agarose gel electrophoresis for the museum samples indicated that the DNA extracted from the museum samples was predominantly distributed below 100 bp. In some cases, such as for the G-F (Also known as G) sample, no gel band was detected, suggesting that the DNA concentration was too low for detection (Supplementary Fig. S1). The museum specimens yielded 3.12-–16.39× average sequencing depth for each sample (Supplementary Table S1).
Genomic variation of all modern goose breeds
By combining with the extant dataset in GenBank, a total of 15,071,551 high-quality SNPs were detected among the 176 modern goose genomes. To gain the distribution of SNPs across all modern breeds, the SNP densities were displayed in a 1 Mb window across the entirety of the genome. Among all chromosomes, chromosomes 8, 23, and 24 exhibited the lowest density of SNPs, while chromosomes 25, 26, and 27 exhibited the highest (Supplementary Fig. S2).
Genetic diversity analyses
The genetic diversity analysis of the Guangdong local breed was carried out on the dataset obtained in present study. Individual SNPs were extracted for Guangdong local goose breeds and resulting in 7,936,429 to 9,235,160 SNPs. The lowest number of SNPs was found in the ST, while the highest number of SNPs was found in the YCB (Table 1). The results indicated that the observed heterozygosity (Ho) and the expected heterozygosity (He) for each Guangdong goose breed were found to be 0.321-0.335 and 0.322-0.331, respectively. In addition, the inbreeding coefficient (F) for each breed was found to be 0.012-0.030 based on genome SNPs, while they were 0.259–0.318 based on the runs of homozygosity (ROH). All Guangdong geese exhibited low F, with the MJ population of SZH exhibiting the highest F (0.030) and WZA exhibiting the lowest F (0.012). However, their interbreed genetic distances were low (0.272), indicating a high linkage disequilibrium (LD) distance (R2(0.3) = 712.053). A synthesis of the results of the number of SNPs (NSNP), F, the inbreeding coefficient based on ROH (FROH), and LD revealed that the ST exhibited the lowest genetic diversity, while YCB exhibited the highest genetic diversity (Table 1).
Furthermore, the LD decay of each breed was plotted, and it was observed that the LD distance of each breed gradually increased as R2 decreased (Supplementary Fig. S3). These findings were consistent with those regarding genetic diversity, as ST had the greatest distance at R2 = 0.2, while YCB exhibited the lowest distance. This further demonstrated that the ST was the least genetically diverse of the goose breeds in Guangdong. The YCB, in contrast, exhibited a high level of genetic diversity.
Population structure
To gain further insight into the population structure of the geese in Guangdong, a joint population structure analysis of our dataset with the extant genomic data of European domestic geese (RM and HT) and non-Guangdong geese (ZI) from GenBank was carried out. The results of the NJ tree (NJ-tree) and principal component analysis (PCA) demonstrated that the individual breeds were distinct from one another, with the exception of the MG and SZH (MJ and GY), which exhibited indistinguishable characteristics (Fig. 1). The results of the Ancestral Component Analysis were consistent with those of the NJ-tree and PCA (Fig. 2). When K = 2, domestic geese were divided into two distinct lineages, in which the Chinese geese formed one ancestral component, including SZH (MJ and GY), MG, WZA, YCB, ST, and ZI, and the European geese formed another ancestral component, including HT and RM. It is noteworthy that the ZI component included European domestic geese. When K = 3, which was the optimal K value (cross-validation error = 0.3647), the ST formed a separate lineage. As the value of K increased from four (cross-validation error = 0.3652) to five (cross-validation error = 0.3654), which were close to the optimal K value, the ZI and WZA were gradually divided into independent lineages. The YCB consistently demonstrated a component of the ST pedigree from K = 3 to 5, which is consistent with the distribution observed in the NJ tree. YCB was identified as an independent ancestral component until K = 6. Meanwhile, the two populations of the SZH (MJ and GY) and the MG had consistently exhibited the same ancestral composition, suggesting their close genetic relationship. Therefore, we do not subsequently analyze the SZH (MJ and GY) for breed-specific related SNPs and genes.
A Neighbor-joining tree; B Principal component analysis (PC1 and PC2); C Principal component analysis (PC3 and PC4). Abbreviations: MG: Magang goose, ST: Shitou goose, SZH(MJ): Miaojing line of Sanzhou Black goose, SZH(GY): Gaoyao line of Sanzhou Black goose, YCB: Yangchun White goose, WZA: Wuzong goose, HT: Hortobágy goose, RM: Roman goose, ZI: Zi goose.
Colors represent different ancestral components. Abbreviations: K: The number of ancestral components, MG: Magang goose, ST: Shitou goose, SZH(MJ): Miaojing line of Sanzhou Black goose, SZH(GY): Gaoyao line of Sanzhou Black goose, YCB: Yangchun White goose, WZA: Wuzong goose, HT: Hortobágy goose, RM: Roman goose, ZI: Zi goose.
Genetic relationships between museum and modern samples
To ascertain the changes in genetic structure and the evolutionary trajectory of Guangdong goose throughout the breeding formation in the last several decades, museum samples and modern samples were subjected to a combination of analyses. Considering the presence of wild swan goose in the museum samples, we added nine modern wild swan geese from public databases to the dataset. By retaining SNPs present in both modern and museum samples, we obtained 2,814,728 high-quality SNPs for subsequent museum sample analyses. The NJ tree indicated that when European domestic geese were employed as the root, the historical SZH samples were situated at the root of the WZA, MG, and modern SZH (MJ and GY) branches. The historical ST samples were intermingled with the modern ST branch, while the historical swan goose samples were identified as being at the root of the modern swan geese branch (Supplementary Fig. S4A). The PCA analysis of the museum samples and modern Chinese geese was consistent with the results of the NJ tree, with the historical ST clustered with the modern ST, the historical swan goose clustered with the modern swan goose, and the historical SZH being more closely related to the WZA, MG and modern SZH (MJ and GY) (Supplementary Fig. S4C, D). The population structure analysis yielded identical results to the modern sample analysis, with the different breeds gradually diverging as K increased from 2 to 5 (Supplementary Fig. S4B). Both the historical ST and the swan goose constituted an ancestral component alongside their respective modern samples. Notably, the historical SZH exhibited multiple ancestral components even after the modern groups had diverged into a single ancestral lineage (Supplementary Fig. S4B).
Gene flow
A total of 253 combinations of f3 statistics were evaluated among all modern domesticated goose breeds. The findings revealed statistically significant gene flow between YCB and other Guangdong geese (Z ≤ −3), as well as between HT and RM. Conversely, no significant f3 statistical values were observed for the remaining breeds (Supplementary Table S2). This result aligned with the outcomes of the population structure analysis, suggesting YCB as a mixed breed. The D statistic was found to be comparable to that of f3 statistic. The analysis revealed that significant gene flow occurred between YCB and other local goose breeds (Supplementary Table S3). These findings are supported by the Treemix analysis, which demonstrated the gene flow within the Guangdong goose breeds. When setting the gene flow event to zero, the results of both the maximum likelihood tree and the NJ tree were identical. Conversely, when setting the gene flow event as 1, the maximum likelihood tree indicated a gene flow from the root of MG to YCB. Similarly, when the gene flow event was set to 2, a gene flow from ZI to RM was detected. When gene flow event was 3, we detected the gene flow from ST to ZI (Fig. 3).
A–D population relationships as inferred from the Treemix analyses, with 0–3 possible gene flow event. Abbreviations: MG: Magang goose, ST: Shitou goose, SZH(MJ): Miaojing line of Sanzhou Black goose, SZH(GY): Gaoyao line of Sanzhou Black goose, YCB: Yangchun White goose, WZA: Wuzong goose, HT: Hortobágy goose, RM: Roman goose, ZI: Zi goose.
To assess Swan goose in the museum sample, we added nine modern wild Swan goose samples to the analysis along with the museum sample. The incorporation of the museum samples provided a response to the past status and pedigree composition of the breed. The outgroup f3 statistics showed that the museum swan goose shared a high proportion of its genetic components with modern swan geese, followed by modern ST and other breeds. And the results demonstrated that the historical SZH had the greatest number of genetically shared components with the WZA, followed by the MG, and the SZH (MJ and GY), in accordance with the findings of the phylogenetic tree. The historical ST exhibited the greatest degree of genetic similarity with the modern ST. All museum samples in the present study had the lowest f3 values with European and Zi geese, indicating a lack of significant gene flow with non-Guangdong domestic geese (Supplementary Table S4). The results of the D statistic were found to be consistent with the outgroup f3 statistic (Supplementary Table S5).
Analysis and functional annotation of breed-specific related genes
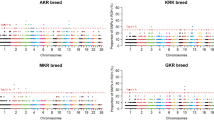
The SNP and genes related to breed-specific traits of Guangdong local geese were identified by detecting breed-specific related SNPs for four officially recognized breeds and ZI. A total of 213,518 to 1,100,759 SNPs were identified for each breed, with ZI exhibiting the most specific SNPs and ST exhibiting the least (Supplementary Table S6). Following filtration using FST, a total of 224-6,568 SNPs were obtained for each breed (Supplementary Table S6 and Fig. 4). A total of 482 SNPs were obtained from YCB, with the majority located on chromosome 15. 224 SNPs were obtained from WZA, with the majority located on chromosome Z. 753 SNP were obtained from MG, with the majority located on chromosomes 5 and Z. 1097 SNP were obtained from ST, with the majority located on chromosomes 1 and 5. Finally, ZI geese obtained 6568 SNPs, with the majority located on chromosomes 1, 2, 3, and Z. It can be observed that all of these regions exhibited high FST values, which indicate that they were subject to strong selection during the formation of each breed. Furthermore, the Z chromosome of different geese exhibited more specific SNPs and higher selection signals.
From the inside to the outside of the circle, the letters a to e stand for the Magang goose, Wuzong goose, Shitou goose, Yangchun White goose, Zi goose. Different colors represent different chromosomes. Abbreviations: A Magang goose, B Wuzong goose, C Shitou goose, D Yangchun White goose, E Zi goose.
The SNPs with the strongest selection signal were then subjected to annotation and yielded 36–720 genes from different goose breeds, including 36 genes from the WZA, 55 genes from the YCB, 96 genes from the MG, 76 genes from the ST, and 720 genes from the ZI (Supplementary Table S6). Subsequently, the gene set for each breed was then submitted to the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis in order to gain further insight into the potential functions of the identified candidate genes (Supplementary Table S7 and Supplementary Table S8). As a result, six significant pathways emerged as significantly enriched by the breed-specific related genes of ST, comprising glutamatergic synapse, cell adhesion molecules and ubiquitin-mediated proteolysis, in which the genes CASR, GLS, CD86, CD99, and CNTN3 were mainly involved. The GO analysis of breed-specific related genes in ST showed significant enrichment in eight GO terms, including L-glutamate transmembrane transport, T cell costimulation and negative regulation of Ras protein signal transduction, among others. A total of 18 significant pathways, including arrhythmogenic right ventricular cardiomyopathy and regulation of actin cytoskeleton were enriched by the breed-specific related genes of YCB. Among these pathways, SGCB, CDH4, EZR, GIT1 and ACTA2 were involved. The GO enrichment analyses of YCB were significantly enriched in four GO terms, including protein binding, collagen binding, extracellular matrix binding, and receptor complex. For MG, 11 pathways were identified as significantly enriched by DPYD, HMGCS1, NOS1, IL11RA, PIGH, GPHN, NFS1, AHCYL2, HRH3, NR3C1, PRLR, NPBWR2, PTGER4, IL11RA, SYCP3, RAD51B and so on, including metabolic pathways, neuroactive ligand-receptor interaction, galactose metabolism and homologous recombination. The GO enrichment analyses of MG enriched in iron-sulfur cluster binding, protein binding, and ATP hydrolysis activity, among others. Furthermore, four pathways, including thermogenesis, cell adhesion molecules, and citrate cycle (TCA cycle), were significantly enriched by the breed-specific related genes of WZA. Among these pathways, GDA, SMARCC1, PTPRT and OGDH were involved. The GO enrichment analyses of WZA were significantly enriched in dephosphorylation. The breed-specific related genes of ZI were found to be significantly enriched to 31 pathways, including the calcium signaling pathway, cell adhesion molecules and oxytocin signaling pathway. Among these pathways, CHRM3, CHRM5, GRM2, GRIN2B, ATP2B2, PDE1C, GNAL, PHKA1, PDGFRB, ERBB4, CACNA1B, PLCB4, VDAC2, MYLK3, TNNC2, RYR3, ADCY1, CACNA2D3, GNAI2, CACNA2D4, ERBB4, GNAL, CACNA2D2, PLCB4 ADCY1, RYR3, MYLK3, RCAN2 were identified as key genes. The GO enrichment analyses of ZI were significantly enriched in 38 GO terms, including protein binding, postsynaptic membrane, intracellular signal transduction, among others.
To further validate the role of varietal breed-specific related genes and SNP loci in different Guangdong breeds, we performed haplotype analyses of the SNP loci linked to these genes. The results showed that there were typical correlations between SNP loci and breeds, such as the loci 6193485 and 215817703 in CM046214.1, associated with ACER3 and AHCYL2, showed significant variation in the MG goose (Fig. 5 and Table 2).
The horizontal axis represents different breeds and the vertical axis represents different gene regions. The Shitou goose genome was used as the reference genome, with 0 (green) representing a pure genotype identical to that of the ST, 1 (black) representing heterozygous, and 2 (red) representing homozygous mutation mutation. The colors at the top represent different breeds. Abbreviations: MG: Magang goose, ST: Shitou goose, YCB: Yangchun White goose, WZA: Wuzong goose.
Discussion
The investigation of the genetic structure and genetic diversity of different breeds in a specific geographic region can facilitate the genetic evaluation, utilization, and conservation of genetic resources. Mitochondrial13 and genome-wide2,14 studies have demonstrated that Chinese geese exhibit high genetic diversity. The present study aimed to analyze the genetic diversity of five local goose breeds in Guangdong. The genetic diversity of the Guangdong local geese was observed to be high, with a considerable number of SNPs, heterozygosity, and low inbreeding coefficients (Table 1), which were in accordance with the findings of a previous genome-wide genetic diversity study on ST and MG14. In comparison to other geese which were considered to have high genetic diversity such as Xingguo Gray goose (F = 0.06 ± 0.04, FROH = 10.99 ± 3.07, R2(0.3)(kb) = 1.45, Ho = 0.28, He = 0.28) and Guangfeng White goose (F = 0.11 ± 0.03, FROH = 10.09 ± 2.62, R2(0.3)(kb) = 1.21, Ho = 0.25, He = 0.26)6, the Guangdong local geese exhibit lower inbreeding coefficients, LD distances, and higher heterozygosity (Table 1). When the observed heterozygosity was less than the expected heterozygosity, inbreeding or selection had occurred in the population15. The observed heterozygosity was higher than or equal to the expected heterozygosity for all Guangdong geese except for SZH(MJ) (Table 1), which suggested potentially targeted artificial selection on the desirable traits during the long-term conservation and utilization. The FROH was considered to be closer to the true inbreeding coefficient16. Contrary to the results of NSNP, He, Ho and F, FROH was higher in Guangdong local goose breeds than in Xingguo Gray goose and Xingguo Gray goose, so the timing of inbreeding occurrence in Guangdong local goose breeds should be further investigated to unravel the reasons for the discrepancies. The observed reduction in genetic diversity of the ST relative to other Guangdong geese may be attributed, at least in part, to its status as the most renowned and widely utilized large-bodied goose in China, which has been subjected to extensive human attention and selective breeding2,17.
Guangdong is home to five different local goose breeds, which play a pivotal role in the regional economy and the conservation of genetic resources associated with local breeds. All the Guangdong domestic geese used in this study exhibit distinct characteristics. The ST, the largest goose breed in the world, has a significantly faster growth rate compared to other local breeds in Guangdong18. The WZA is notable for its more pronounced black feathers than other Guangdong local geese. The MG, a medium-sized breed, is recognized for its early maturity and tendency to fatten easily, making it highly popular in the market. The YCB is the only white-feathered goose breed in Guangdong, valued for both its meat and feather characteristics. The phenotypic diversity and full range of utilization directions of Guangdong local goose breeds allowed us to use them as a model to analyze in depth the genomic differences between different types of local goose breeds and to identify breed-specific related SNPs and functional genes. The Chinese native geese in this study exhibit a similar genetic composition and are clearly distinguished from the European domestic counterparts, which originated from distinct wild geese19,20. Our findings indicate that MG and SZH (MJ and GY) may share a common ancestry (Fig. 1, Fig. 2 and Supplementary Fig. S4), which is consistent with the document record of SZH as the genetic parent of MG21. This may also be the consequence of cross-utilization of breeds during the past few decades. As a direct result, the gander of SZH increased from 2950 g22 in 2008 to 3373 g at present at the 7 weeks of age. In addition, the body weight at 90 days-old of SZH is 4.89 kg, suggesting SZH is larger than the medium sized geese MG23,24. The above results further confirmed the genetic differences between the museum and modern SZH from both MJ and GY. YCB exhibits ancestral components derived from several breeds of geese (Fig. 2), which may have been the source of its white feather, to be included in its pedigree.
Museum samples have been proposed as a valuable resource for the traceability of breeds and the resolution of pedigree composition25. A noteworthy finding is the observation that the G-ST and G can be effectively clustered together in their contemporary modern samples (Supplementary Fig. S4A), indicating that modern samples were subject to early selective breeding and did not exhibit significant group stratification. This suggested that the formation of ST predates the museum sample date of 1980, and showed continuity with modern populations, which was consistent with the documented history of ST formation21. Moreover, the G-SH exhibited multiple ancestral components (Supplementary Fig. S4B), suggesting that the remaining breeds esp. MG may be derived from SZH, a conclusion that aligns with the historical record on these two breeds21. The gene flow between the SZH (MJ and GY), MG and WZA suggests (Supplementary Table S2, S3) that both the MG and WZA in this study were influenced by the SZH (MJ and GY) in their breeding processes. These results supported that MG and WZA introduced the SZH during breeding and subsequently fixed it through selective breeding. Our results confirmed for the first time at the molecular level the history of the formation of the Guangdong domestic goose. In addition, we discovered gene flow between domesticated geese and wild Swan geese as well as domesticated goose breeds19, proving detailed evidence for the complex history of the formation of geese in China.
The Guangdong local geese show displays a remarkable range of phenotypic variation. The ST, for instance, stands as the largest local goose breed in China and globally, with an average weight of over 9 kg26. The MG, on the other hand, belongs to the medium-sized goose category, while the WZA represents the small-sized goose2. Furthermore, there is a discernible distinction in coloration between the various breeds. Consequently, the specific SNP may be associated to the most typical characteristics of the local breeds. Given that ZI goose is known for its high egg production, a range of signaling pathways related to reproduction and neuroendocrinology were identified, such as biological process, the calcium signaling pathway, the oxytocin signaling pathway, the estrogen signaling pathway, and the GnRH signaling pathway, which were found to contain genes that may affect egg-laying traits27. The MAPK signaling pathway, the PI3K-Akt signaling pathway, and other signaling pathways previously validated to be related to egg production in geese28 were also identified. The results of the GO enrichment analysis showed that only a few GO terms were significantly enriched, but all breeds were consistently associated with protein binding terms (Supplementary Table S8). This suggests that the development of their distinct traits is linked to the molecular function of protein binding.
To minimize false positives and he intensity of the screening of breed-specific characterization genes or SNP, the ZI, a representative of the non-Guangdong Goose, was added to the analysis. The annotated SNPs of Guangdong local geese were found to be relate to a variety of traits (Supplementary Table S7). The annotation in ST yielded genes related to body weight, growth, and development, including ARHGAP21, which was identified as a gene associated with the body weight of Yangzhou geese29. Furthermore, the enriched DOCK1 was found to be associated with the growth and development of chickens30, while FBRSL1 was linked to the increase in pectoral muscle weight of ducks31. Additionally, a number of genes related to development were identified, including SSTR418, STT3A32, and TBC1D433, which may be involved in the unique large body trait observed in ST. Multiple genes associated with growth traits have been identified in ST that may be directly related to their larger body weights34 and faster growth rates18. The MG, a local breed with a reputation for its exceptional ability to deposit fat, has been identified as containing several genes that are known to be associated with fat metabolism including ACER335,36 and AHCYL237. Meanwhile, the YCB were found to contain genes associated with feather development (ACTA2)38, body size development (PRDM16)39,40, body weight-related (SGCB)41, and goose beak development (SEPSECS)42. These genes are representative of the characteristics of YCB, a medium to large white-feathered goose breed in Guangdong. Breed-specific related genes further indicate the existence of genes controlling body size in YCB that differ from those found in ST. GDA is believed to be associated with skin pigmentation in Huoyan geese43, and in the context of the present study, it has been found to occur within WZA-specific genes, which may be linked to their ebony skin color of head and foot. In addition, it has been demonstrated that PTPRT is involved in feather growth and maturation in ducks44. This may suggest that PTPRT could be a candidate gene regulating the feather color of WZA. Consequently, genes linked to a multitude of functional phenotypes have been identified in different Guangdong geese, collectively constituting the distinctive characteristics of local breeds. Haplotype analyses further validated that breed-specific related SNPs can reflect the functional traits of the breed (Fig. 5 and Table 2), highlighting significant variations between different breeds. These genes and SNP loci may provide a foundation for subsequent selective breeding or exploitation programs aimed at local goose breeds in Guangdong.
Conclusions
A comprehensive examination of genomic data indicates a considerable degree of genetic diversity in the Guangdong local geese, as evidenced by a substantial number of SNPs, high heterozygosity, and low inbreeding coefficients. The museum SZH exhibited multiple ancestral components, and the modern MG may be derived from the SZH, as it has been well documented. A multitude of genes and SNP loci have been identified that are associated with a variety of functional phenotypes in different Guangdong geese, collectively defining the distinctive characteristics of local breeds. These findings provide a basis for subsequent selective breeding or exploitation programs aimed at local goose breeds in Guangdong.
Methods
Sample collection
120 whole blood samples were collected from four typical breeds, including ST, WZA, MG, YCB and two conservations lines (MJ and GY) of the folk breed SZH in Guangdong province. For each breed/population, 20 whole blood samples were collected from randomly selected individuals of both sexes at ~90 days of age via the sub-wing vein and stored in EDTA-anticoagulated (Ethylenediaminetetraacetic acid anticoagulated) tubes at −80 °C until further analysis. We have complied with all relevant ethical regulations for animal use. All experimental procedures used in this study were approved by the Laboratory Animal Welfare and Animal Experimental Ethical Inspection Board of Foshan University (FOSU202206-28). Skin tissue or phalanx bone were collected from five museum samples made in 1980s, representing one undomesticated Swan goose (G), two Sitou geese (G-ST), and two Sanzhou Black geese (G-SH). In addition, we collected resequencing data from 56 modern samples from public databases (NCBI, http://www.ncbi.nlm.nih.gov/) for joint analysis, including ST45, MG45, ZI45, RM45, HT45 and HY6,19. Detailed sample information has been shown in Supplementary Table S9. Different datasets were applied for the different downstream analyses. In brief, 120 samples of Guangdong local goose breeds collected in this study were used for the genomic diversity analysis of modern Guangdong geese. 47 modern domesticated samples from public database were incorporated in with 120 Guangdong geese for assessing genetic structure and functional gene identification among modern breeds. The nine modern wild Swan geese and the five historical samples were included in the analysis of breed formation history.
Modern sample information
The ST is distinguished by its large size, rapid growth, short feeding period, tolerance of roughage, high feed conversion efficiency and strong adaptability21. Nevertheless, the ST also exhibits comparable deficiencies, including a decline in reproductive performance and inferiority in meat quality3. The MG is a medium-sized goose breed that is renowned for its rapid growth, resilience to roughage, early maturation, and proclivity for rapid fattening. Adult MG exhibit distinctive cranial, cervical, dorsal, and pedal traits, which are often collectively referred to as the ‘four crows’21. The WZA is a small-sized goose species that produces meat of high quality, characterized by its tenderness and flavor, as well as a high proportion of unsaturated fatty acids and low cholesterol3. The breed is characterized by a black beak, black fur, black feet, and a slender head, neck, and bones. Adult WZA are also distinguished by a band of black feathers extending from the base of the bill to the cervical vertebrae21. The YCB is a newly discovered local goose breed in Guangdong in year 2024, which is characterized by medium to large body size, white feathers, and robust disease resistance. YCB is the only white-feathered goose breed in Guangdong. SZH is a small and medium-sized local goose breed. The whole body,y except the chest and abdomen, are black. SZH has fine bones, high meat tenderness, and good meat flavor46.
DNA extraction and genome resequencing
For modern samples, genomic DNA was extracted from blood samples using the Ezup Column Blood Genomic DNA Extraction Kit (Vazyme Biotech Co., Ltd, China) according to the kit instructions. The extracted DNA was subjected to agarose gel electrophoresis to ascertain its integrity, while the DNA concentration and purity were determined using a NanoDrop. All qualified samples were used for library construction and sequenced using the DNBSEQ-T7 platform (PE150). The insert size of the sequencing libraries was the average length ~300 bp for all samples. The average sequencing depth of the samples was greater than 15×.
The DNA experiments conducted on museum specimens were performed in the Ancient DNA Laboratory of Foshan University. In a preliminary step, all equipment was subjected to an initial cleaning procedure involving a 5% (v/v) sodium hypochlorite solution and a subsequent washing with 75% (v/v) alcohol. Following this, all items were exposed to ultraviolet radiation for a duration of one hour. The surface of the museum specimens was meticulously cleaned by gently rubbing it with sandpaper to remove any dust particles. Subsequently, the museum specimens were subjected to a two-stage cleansing process. Initially, they were washed with a 5% (v/v) sodium hypochlorite solution, followed by a rinse with double-distilled water. This was followed by a 30-minute exposure to ultraviolet irradiation. The washed samples were then ground into a fine-grained powder using an automatic quick-grinding machine. Subsequently, a quantity of 100-200 mg of the sample powder was subjected to DNA extraction in accordance with the previously described methodology47. A final volume of 20 μL of DNA was obtained. Agarose gel electrophoresis was used to initially characterize the quality of DNA from museum samples. For each sample, 50 ng of initial DNA was utilized to construct the library with the VAHTS Universal Plus DNA Library Prep Kit for MGI (Vazyme Biotech Co., Ltd., China). In order to monitor contamination, several mock extractions and library preparations were carried out alongside the samples in the same manner. The qualified libraries were then subjected to circularization with the VAHTS Circularization Kit for MGI (Vazyme Biotech Co., Ltd.). Sequencing was performed using the DNBSEQ-T7 platform, with read lengths of 100 bases (PE100). The insert size of the sequencing libraries for the museum samples was more variable, ranging from 100 to 300 bp, due to the degradation of the DNA fragments in these older samples. Each sample yielded 15–68 Gb of raw base data.
Data filtering and SNP genotyping
The raw reads were subjected to filtration using fastp v0.20.048 based on the following criteria: 1) those containing library adapter, 2) those with low-quality bases exceeding 10%, and 3) those with N content exceeding 10%. The clean reads were then aligned to the most recent chromosome-level ST reference genome (GCA_025388735.1)2 using BWA v 0.7.1749 with default parameters. The mapped reads were sorted using the SAMtools software50 and the PCR duplicates were removed using the Picard software (http://broadinstitute.github.io/Picardv2.22.9/). SNPs were identified utilizing the RealignerTargetCreator and IndelRealigner modules in GATK v3.5 software51 and the HaplotypeCaller, CombineGVCFs, and GenotypeGVCFs modules within the GATK v4 software suite52. INDEL in the dataset was removed using the SelectVariants module in GATK 4.0. The SNP filtration was conducted using the VariantFiltration module in GATK, with the following criteria: QD < 5.0, MQ < 40.0, FS > 60, MQRankSum < −12.5, ReadPosRankSum < -8.0, and a minimum of three SNPs in a 10 bp window. The SNP data was filtered using awk software scripts to retain only biallelic in the SNP data. Subsequently, the quality of the SNP data was further assessed utilizing the VCFtools software53 with the following parameter settings: 1) The removal of SNP loci with a SNP deletion rate greater than 90% (--max-missing 0.9); 2) The exclusion of SNPs with a minimum allele frequency less than 0.05 (--maf 0.05). The final SNP dataset was then phased and imputed using Beagle 5.454.
Genetic diversity assessment
To clarify the genetic diversity of different Guangdong local breeds and prevent discrepancies in genetic diversity as a result of differing sequencing depth, only 120 modern geese obtained in this study were employed to estimate the genetic diversity exhibited by the Guangdong local breeds. The parameters in question were NSNP, Ho, He, F, the intraspecific genetic distances (DST), FROH, and LD. The Ho and He were estimated using the VCFtools software (--hardy), while the F was calculated using the VCFtools software (--het). The DST was calculated using the command “plink -distance square 1-ibs”. The FROH was calculated based on a genome size of 1.29 G using the PLINK with the parameter -homozyg -homozyg-snp 30 -homozyg-kb 30. The square of the correlation coefficient (r2) was employed to qualify the degree of linkage between SNP alleles, with this being estimated using PLINK (-r2-ld-window-kb 500-ld-window-r2 0). The LD decay plot was generated using the PopLDdecay software55.
Population structure analyses
To objectively understand the genetic structure and genetic relationship of local domestic geese in Guangdong, we added 47 modern samples of domestic geese, including ZI, HT, RM, ST, and MG downloaded from public databases, to the 120 samples of Guangdong domestic geese obtained in this study for joint analysis. The imputed SNP dataset was filtered using PLINK software56 with the parameter “--indep-pairwise 50 5 0.2” to identify and exclude any linked SNP in the genomic data. The filtered data were then used for subsequent ADMIXTURE analyses. The Neighbor-joining (NJ) tree was employed to estimate the genetic relationships among different breeds, which were calculated using PLINK software with the parameter “--distance-matrix”. Then the NJ tree was constructed by MEGA X software57 and visualized using Figtree v14.4 (http://tree.bio.ed.ac.uk/software/figtree/). PCA was carried out using GCTA software, and the first four components were visualized using R. The ADMIXTURE software58 was used to estimate the ancestral components of each individual. The population structure of Guangdong domestic geese was evaluated by inferring the number of different ancestral components (K).
Gene flow and phylogenetic analyses
To assess the genetic relationship and gene flow between domesticated goose breeds, f3 statistics, D statistics, and Treemix were employed to analyze the possible gene flow of all 167 modern samples. The f3 statistics analysis was performed utilizing qp3pop module within the Admixtools software59. The model employed for f3 statistics is (A, B; C) where C represents the target breed and A and B are the source breeds to test whether C is affected by A and B. All possible combinations of different breeds were tested. When the Z value was found to be significant (Z ≤ -3), it can be concluded that C was affected by A and B. The D-statistics analysis (ABBA-BABA test) was performed using the qpDstat module in Admixtools59. The D-statistics was modeled as (A, B; C, D). When D is less than zero, the ABBA model is obtained, indicating the presence of gene flow between BC or AD, and vice versa. To elucidate the direction of gene flow, the Treemix software60 was employed to calculate the gene flow between different breeds, with the European domestic goose serving as an outgroup and the assuming of 0–3 gene exchange events.
When museum samples and wild Swan geese (HY) were incorporated for analyzing the history of the formation of the Guangdong domestic goose, only SNPs that existed in both modern and museum samples were utilized to perform the same analysis as above. However, in this instance the f3 statistics were replaced by outgroup f3 statistics. Outgroup f3 statistics entail replacing population C in f3 statistics with a population that is not related to A and B, thus enabling evaluation of the gene flow between A and B. In this study, population C was designated as RM.
Breed-specific related genes identification and functional annotation
Due to the uncertainty genetic background of SZH breeds, a comparison was performed on the SNP data from five geese breeds native to Asia. The BCFtools software50 was employed to identify the breed-specific related SNPs that were present in a single breed only, thereby allowing us to gain insight into the unique genetic makeup of each breed. Additionally, selection signature analysis was conducted on the specific SNPs of each breed. The top 1% SNPs were considered to be the most relevant to breed characterization. The SNP loci obtained were then subjected to annotation via snpEff software61. To determine the functions of genes corresponding to breed-specific related SNP loci, the annotated genes were submitted to a customized database (goose database) for KEGG pathway enrichment analysis, and to DAVID62,63 for GO analysis, with a statistical significance threshold of raw P < 0.05 for both analyses.
The functions of the relevant genes were reviewed through the literature, and the SNP loci associated with the breed -specific characteristics were subjected to haplotype analysis using the ST as the reference genome. The haplotypes results were then visualized using OmicShare Tools64.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated in this study have been deposited into the NCBI repository under the accession number PRJNA1286101. The data for 56 other domestic geese, ST, MG, Roman geese, Zi geese, Wild swan geese, and Hortobágy geese are available from NCBI (http://www.ncbi.nlm.nih.gov BioProject IDs: PRJNA72204945, PRJNA76775719, and PRJNA6788156). The uncropped and unedited gel images of Figure S1 is Obtained in the Supplementary Information (Supplementary Fig. S5).
References
Eda, M. et al. Multiple lines of evidence of early goose domestication in a 7,000-y-old rice cultivation village in the lower Yangtze River, China. Proc. Natl. Acad. Sci. USA 119, e2117064119 (2022).
Zhao, Q. Q. et al. Chromosome-level genome assembly of goose provides insight into the adaptation and growth of local goose breeds. Gigascience 12, giad003 (2022).
Chen, G. H. et al. Metabolomic, lipidomic, and proteomic profiles provide insights on meat quality differences between Shitou and Wuzong geese. Food Chem. 438, 137967 (2024).
Liu, J. X. et al. The DNA methylation status of the serotonin metabolic pathway associated with reproductive inactivation induced by long-light exposure in Magang geese. BMC Genom. 24, 355 (2023).
Zheng, S. M. et al. Genomic signatures reveal selection in Lingxian white goose. Poult. Sci. 102, 102269 (2023).
Ouyang, J. et al. Chromosome-level genome and population genomics reveal evolutionary characteristics and conservation status of Chinese indigenous geese. Commun. Biol. 5, 1191 (2022).
Zhang, Y. et al. Morphological, anatomical and histological studies on knob and beak characters of six goose breeds from China. Front. Physiol. 14, 1241216 (2023).
Xiang, H. et al. Early Holocene chicken domestication in northern China. Proc. Natl. Acad. Sci. USA 111, 17564–17569 (2014).
Girdland Flink, L. et al. Establishing the validity of domestication genes using DNA from ancient chickens. Proc. Natl. Acad. Sci. USA 111, 6184–6189 (2014).
Cai, D. W. et al. The origins of Chinese domestic cattle as revealed by ancient DNA analysis. J. Archaeol. Sci. 41, 423–434 (2014).
Delsol, N. et al. Ancient DNA confirms diverse origins of early post-Columbian cattle in the Americas. Sci. Rep. 13, 12444 (2023).
Ostrander, E. A., Wayne, R. K., Freedman, A. H. & Davis, B. W. Demographic history, selection and functional diversity of the canine genome. Nat. Rev. Genet 18, 705–720 (2017).
Zhang, Y. et al. Genetic diversity analysis and breeding of geese based on the mitochondrial ND6 gene. Genes 14, 14081605 (2023).
Gao, G. L. et al. Insights into genetic diversity and phenotypic variations in domestic geese through comprehensive population and pan-genome analysis. J. Anim. Sci. Biotechnol. 14, 150 (2023).
Hoffmann, A. A. et al. An endangered flightless grasshopper with strong genetic structure maintains population genetic variation despite extensive habitat loss. Ecol. Evol. 11, 5364–5380 (2021).
Forutan, M. et al. Inbreeding and runs of homozygosity before and after genomic selection in North American Holstein cattle. BMC Genom. 19, 98 (2018).
Zhang, J. et al. Genome-wide population genetic analysis of commercial, indigenous, game, and wild chickens using 600K SNP microarray data. Front. Genet. 11, 543294 (2020).
Tang, J. et al. Comparative transcriptome analyses of leg muscle during early growth between geese (anser cygnoides) breeds differing in body size characteristics. Genes 14, 14051048 (2023).
Wen, J. H. et al. Origins, timing and introgression of domestic geese revealed by whole genome data. J. Anim. Sci. Biotechnol. 14, 26 (2023).
Shi, X. W., Wang, J. W., Zeng, F. T. & Qiu, X. P. Mitochondrial DNA cleavage patterns distinguish independent origin of Chinese domestic geese and Western domestic geese. Biochem Genet 44, 237–245 (2006).
National Committee of animal genetic resources. National List of Livestock and Poultry Genetic Resources in China. (Ministry of Agriculture and Rural Affairs of the People’s Republic of China, 2022).
Jing, D. L., Pan, X. H. & Lu, X. M. Ecological distribution and breed characteristics of Sanzhou goose (in Chinese). J. Anhui Agri. Sci, 563+566 (2008).
Zhu, C. et al. Effects of dietary supplementation with Bacillus subtilis and bacteriophage on growth performance, intestinal morphology and microbiota structure in 0-90 d MaGang geese. Front Nutr. 12, 1537724 (2025).
Xu, Z. Q. et al. Comparison of carcass performance between Magang goose and Wuzong goose (in Chinese). Guangdong Agric. Sci. 40, 113–116 (2013).
Raxworthy, C. J. & Smith, B. T. Mining museums for historical DNA: advances and challenges in museomics. Trends Ecol. Evol.36, 1049–1060 (2021).
Zhao, Q. et al. Chromosome-level genome assembly of goose provides insight into the adaptation and growth of local goose breeds. GigaScience 12, giad003 (2023).
Gao, G. L. et al. Molecular cloning of the goose GnRH gene and identification of GnRH polymorphisms associated with laying traits. Br. Poult. Sci. 61, 502–507 (2020).
Zhao, H. C. et al. Genome-wide selective signatures mining the candidate genes for egg laying in goose. BMC Genom. 24, 750 (2023).
Zhang, Y. et al. Association analysis of polymorphisms of candidate genes for laying traits in Yangzhou geese. Gene 862, 147249 (2023).
Wu, P. F. et al. Identification of crucial circRNAs in skeletal muscle during chicken embryonic development. BMC Genom. 23, 330 (2022).
Yu, S. M. et al. Resequencing of a Pekin duck breeding population provides insights into the genomic response to short-term artificial selection. Gigascience 12, giad016 (2023).
Qi, H. et al. Analysis of the differentially expressed genes in the combs and testes of Qingyuan partridge roosters at different developmental stages. BMC Genom. 25, 33 (2024).
Espelage, L., Al-Hasani, H. & Chadt, A. RabGAPs in skeletal muscle function and exercise. J. Mol. Endocrinol. 64, R1–R19 (2020).
Zhang, X. et al. DNA Demethylation of myogenic genes may contribute to embryonic leg muscle development differences between wuzong and shitou geese. Int. J. Mol. Sci. 24, 7188 (2023).
Vahedi, S. M., Salek Ardestani, S., Karimi, K. & Banabazi, M. H. Weighted single-step GWAS for body mass index and scans for recent signatures of selection in Yorkshire pigs. J. Hered. 113, 325–335 (2022).
Ibeagha-Awemu, E. M., Peters, S. O., Akwanji, K. A., Imumorin, I. G. & Zhao, X. High density genome wide genotyping-by-sequencing and association identifies common and low frequency SNPs, and novel candidate genes influencing cow milk traits. Sci. Rep. 6, 31109 (2016).
Zhao, L. M. et al. Comparative proteomics reveals genetic mechanisms of body weight in Hu sheep and Dorper sheep. J. Proteom. 267, 104699 (2022).
Li, W. H. et al. Transition from natal downs to juvenile feathers: conserved regulatory switches in Neoaves. Res. Sqr. 15, 4174 (2023).
Wu, Z. et al. Heterogeneity of a dwarf phenotype in Dutch traditional chicken breeds revealed by genomic analyses. Evol. Appl 14, 1095–1108 (2021).
Han, R. L. et al. Novel SNPs in the PRDM16 gene and their associations with performance traits in chickens. Mol. Biol. Rep. 39, 3153–3160 (2012).
Tarsani, E. et al. Discovery and characterization of functional modules associated with body weight in broilers. Sci. Rep. 9, 9125 (2019).
Huang, J. H. et al. Identification of key candidate genes for beak length phenotype by whole-genome resequencing in geese. Front. Vet. Sci. 9, 847481 (2022).
Wen, J. H. et al. Genomic analysis reveals candidate genes underlying sex-linked eyelid coloboma, feather color traits, and climatic adaptation in huoyan geese. Animals 13, ani13233608 (2023).
Ma, S. C. et al. Genome-wide association analysis of the primary feather growth traits of duck: identification of potential Loci for growth regulation. Poult. Sci. 102, 102243 (2023).
Yang, Y. et al. Using comparative genomics to detect mutations regulating plumage variations in graylag (A. anser) and swan geese (A. cygnoides). Gene 834, 146612 (2022).
WEI, J. X. et al. Genetic diversity of Sanzhou black geese and genetic relationship with Guangdong goose breeds (in Chinese). Chin. J. Anim. Sci. 60, 114–119 (2024).
Zhang, X. et al. Ancient mitogenomes reveal the domestication and distribution of cattle during the longshan culture period in North China. Front. Genet. 12, 759827 (2021).
Chen, S. F., Zhou, Y. Q., Chen, Y. R. & Gu, J. FASTP: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
McKenna, A. et al. The Genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. bioRxiv, 201178 (2018).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Browning, B. L., Tian, X., Zhou, Y. & Browning, S. R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet. 108, 1880–1890 (2021).
Zhang, C., Dong, S. S., Xu, J. Y., He, W. M. & Yang, T. L. PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35, 1786–1788 (2019).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19, 1655–1664 (2009).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Fitak, R. R. OptM: estimating the optimal number of migration edges on population trees using Treemix. Biol. Methods Protoc. 6, bpab017 (2021).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–w221 (2022).
Mu, H. et al. OmicShare tools: A zero-code interactive online platform for biological data analysis and visualization. iMeta 3, e228 (2024).
Acknowledgements
This work was supported by the program of survey and identification of livestock and poultry genetic resources in Guangdong Province (20221200340103109), the program of the Construction of DNA fingerprint database for major livestock and poultry breeds (2023-XJS-00-001), Guangdong Rural Science and Technology Commissioner Project (KTP20240679) and Foshan Postdoctoral Research Initiation Grant (BKS209159). We sincerely thank Guangdong Provincial Key Laboratory of Animal Molecular Design and Precision Breeding for providing the experimental platform and high-performance computing platform.
Author information
Authors and Affiliations
Contributions
H.L. and H.X. conceived and designed the study; X.Z., J.W., Z.M., D.H., G.Q., R.Z., R.L., M.H., J.G., F.Y., Q.Z., Q.L., H.X., and H.L. collected the samples and performed the experiments; X.Z. and H.X. analyzed data; X.Z. drafted the manuscript; H.X. and H.L. reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
We have complied with all relevant ethical regulations for animal use. All experimental procedures used in this study were approved by the Laboratory Animal Welfare and Animal Experimental Ethical Inspection Board of Foshan University (FOSU202206-28).
Peer review
Peer review information
Communications Biology thanks Supriya Chhotaray, Suelen Fernandes Padilha and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Aylin Bircan, Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, X., Qiu, G., Huang, D. et al. Genetic assessment and Identification of genes related to characterization of Guangdong local goose breeds based on modern and historical genomes. Commun Biol 8, 1132 (2025). https://doi.org/10.1038/s42003-025-08550-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08550-6