Abstract
Acinetobacter baumannii is recognized as the paradigm of multidrug-resistant superbug, topping the WHO priority list of critical human pathogens. Interestingly, it senses and responds to blue light, which modulates global aspects of its physiology, including pathogenicity. We hypothesized that light could serve as a signal to synchronize the bacterial physiology to the host’s behavior and/or to the environment. At environmental temperatures, light regulation is mainly governed by the photoreceptor BlsA. In this work, we identified the existence of daily rhythms in blsA promoter activity displaying a robust response to light, as well as endogenous circadian rhythms in A. baumannii. In fact, we show that blsA promoter activity can be synchronized to 24-hour blue light-dark cycles, which immediately resynchronizes after a phase shift. Upon release to constant darkness, bacterial populations present free-running oscillations with a period close to 24 hours. Furthermore, our data indicate that BlsA is involved in synchronization to light-dark cycles. In fact, under constant darkness without previous entrainment, A. baumannii is rhythmic, but acrophases are not clusterized, behaving as the blsA mutant under light-dark cycles. Our work contributes to the developing field of circadian clocks in bacterial pathogens, which could impact in the microorganisms’ lifestyle and pathogenicity.
Similar content being viewed by others
Introduction
Circadian rhythms, observed across all domains of life, enable organisms to anticipate and prepare for daily changes in environmental conditions resulting from the periodic movement of the Earth. Among bacteria, circadian rhythms have been traditionally studied in cyanobacteria, which are photosynthetic microorganisms. In fact, genes responsible for the core clock of cyanobacteria have been identified, including kaiA, kaiB, and kaiC, along with other crucial components like the response regulator RpaA and two regulatory histidine kinases, CikA and SasA1. Recently, it has been shown that non-photosynthetic soil microorganisms, such as Bacillus subtilis, exhibit a light-dependent circadian system that behaves in a complex fashion, similar to the circadian systems of multicellular eukaryotes2,3. Moreover, it was shown that a clinical isolate of the gut bacterium Klebsiella aerogenes exhibited melatonin and temperature-dependent rhythms in the expression of the motility gene motA4,5. Although the relevance of these circadian rhythms in pathogenesis has not been explored, these findings suggest that the circadian clock of K. aerogenes may entrain to host cues in vivo. Finally, sustained temporal oscillations in lipid droplets have been reported in Pseudomonas aeruginosa synchronized by temperature cycles6.
Although light-dependent circadian rhythms have not been reported in bacterial pathogens, it is well established that important pathogens such as Acinetobacter baumannii, Pseudomonas aeruginosa, Brucella abortus, and Staphylococcus aureus sense and respond to light, showing a global photic modulation of bacterial physiology and, most interestingly, of determinants of pathogenicity7,8,9,10,11. A large body of evidence demonstrates that light signal transduction and physiological responses are mainly governed by blue light through the FAD (BLUF)-type photoreceptor BlsA at environmental temperatures in A. baumannii7,8,12. In fact, blue light modulates iron uptake, tolerance, and susceptibility to antibiotics, desiccation tolerance, competition with other microorganisms, antioxidant enzyme levels, metabolism, biofilm formation, and quorum sensing7,8,13,14,15,16,17. BlsA is a global regulator able to bind and antagonize the functioning of different transcriptional regulators such as Fur, the iron metabolism repressor, and AcoN, the acetoin catabolism repressor, in a light-dependent manner15,18. We have also shown that A. baumannii responds to light modulating virulence in a human keratinocyte epithelial infection model7 as well as quorum sensing16 at 37 °C, through a BlsA-independent mechanism7,16. Moreover, the two-component system BfmRS has been shown to transduce light signaling, as it is involved in photoregulation of motility and desiccation tolerance in A. baumannii strain V15, while evidence strongly indicates that BfmR phosphorylation levels are also modulated by light17. Finally, BfmRS antagonizes blsA transcription17, and has been shown to physically interact with BlsA19.
A. baumannii presents an alarming propensity to develop multi-drug resistance, leading to serious outbreaks in the hospital setting20,21,22, while there are also increasing reports of community-acquired A. baumannii infections across the globe23. Well-described manifestations of A. baumannii include pneumonia, septicemia, bloodstream and urinary tract infections, wound infections or osteomyelitis, endocarditis, and meningitis23. Infections associated with multi-drug resistant (MDR) A. baumannii are linked to higher rates of morbidity and mortality20,21,23. Accordingly, carbapenem-resistant Acinetobacter recently topped the WHO priority list of bacteria that require research and development of novel therapeutic strategies22.
A. baumannii’s lifestyle and pathophysiology represent the paradigm of a resist-persist pathogen, highlighting the importance of its environmental tenacity and physiological state at the onset of infection in its success as pathogen23. In fact, the pathogen’s ability to survive, persist, and thrive in clinical settings is directly linked to acquisition and selection of multi-drug resistance leading to hospital outbreaks24,25. The direct transmission of the microorganism from the environment to the immunocompromised patient tissues through medical devices, which bypass natural host barriers (such as endotracheal tubes), facilitates infection and constitutes one of the most widespread mechanisms leading to disease in the clinical setting. Lifestyle strategies supporting A. baumannii persistence in the environment include the formation of biofilms24,25,26,27,28, a condition that is more favored at environmental temperatures of 23-28 °C rather than at 37 °C for many strains, and entrance into the viable but non-culturable (VBNC) state29. In fact, systems specifically adaptive for its environmental lifestyle have been described, such as the light-sensing and response system, BlsA, which has been shown to be operative at temperatures up to 25 °C8,30.
The bacterial physiological state at the onset of infection is determinant in infection outcome and host fate23,31,32. In fact, a key determinant in patient survival is the early initiation of effective antimicrobial therapy, whose success depends on the bacterial antibiotic susceptibility and virulence status at the onset of infection23.
In this work, we studied the existence of circadian rhythms in the critical pathogen A. baumannii, which could modulate the bacterial physiological state at the onset of infection during the environmental, pre-infectious bacterial life. Since blue light has been shown to extensively modulate A. baumannii physiology, and is an environmental clue entraining circadian rhythms in most of the experimental models examined so far, we studied the effect of blue light on circadian synchronization and entrainment. The existence of such rhythms could modulate bacterial persistence in the environment, antibiotic susceptibility, and pathogenicity, impacting virulence and opening new paths for treatment of infections produced by these pathogens.
Results
Daily and free-running rhythms of blsA promoter activity in A. baumannii
We have shown that blue light exerts a global modulation of A. baumannii’s physiology at moderate temperatures such as 23 °C through the blsA photoreceptor7,8,12,13,15,16,17. To study the existence of rhythms under light-dark cycles and free-running (FR) under constant darkness conditions, we evaluated blsA promoter activity using a luciferase reporter transcriptional fusion. For this, A. baumannii V15 pLPV1Z-pblsA::luc clone 1 cultures were grown without shaking in lysogeny broth (LB)33 under 12 h blue light (bL)-12 h dark (D) 12bL:12D photoperiod for 4 days and then released to DD for 2.5 more days, at 23 °C. Cultures formed robust wall biofilms under these conditions (Fig. S1). This strain expresses a luciferase reporter under the control of the blsA promoter from the pLPV1Z plasmid34. Results shown in Fig. 1A represent all traces, i.e., the total of wells and average + SD, registered from a total of 3 biological replicates. All traces and all rhythmic (i.e., still cycling under constant environmental conditions) traces corresponding to each biological replicate are shown in Fig. S2A–F.
A Luminescence reporter activity of A. baumannii V15 pLPV1Z-pblsA::luc clone 1. The cultures were incubated in bLD cycles for 4 days and then released to DD. The graph includes all traces of three independent experiments, n = 141. Black/blue bars indicate dark/light periods, respectively. Luminescence signals are shown as mean ± SD in blue (line and shadow, respectively), and the individual wells are represented in grey lines. B Average endogenous period of rhythmic populations of A. baumannii V15 pLPV1Z-pblsA::luc clone 1 (26 ± 2.4 h, n = 124). Data are shown as mean ± SD. C Average amplitude of the luminescence rhythm of rhythmic populations (n = 124) of A. baumannii V15 pLPV1Z-pblsA::luc clone 1. Data are shown as mean ± SD. P-value was calculated using Wilcoxon signed-rank test (****p < 0.0001). D Rayleigh plots showing the phase of the bioluminescent peak under cyclic conditions (bLD, black dots) and the bioluminescent peak of the first day of release to FR (DD, white dots) for the rhythmic population (bLD: ZT22.9 ± 0.8 h, R = 0.98 and DD: CT14.6 ± 1.5 h, R = 0.92; n = 124). Lines represent the average peak phase of pLPV1Z-pblsA::luc activity (mean vectors for the circular distributions) of each group. The length of the vector represents the strength of the phase clustering while the angle of the vector represents the mean phase. Individual data points are plotted outside the circle. The central circle represents the threshold for p = 0.05. E–G Individual luminescence plots for the rhythmic populations of A. baumannii V15 pLPV1Z-pblsA::luc clone 1 for biological replica 1, 2 and 3, respectively. Bacteria were grown at a constant temperature of 23 °C. B–D analysis included three biological replicates. n = total number of samples; 1 sample = 1 well of a 96-well plate. n rhythmic = number of samples exhibiting circadian rhythmicity under free-running conditions (see Methods for definition).
We first analyzed whether blue light could act as a zeitgeber in A. baumannii. The pblsA::luc reporter showed that blsA promoter activity responds to bLD cycles establishing a robust diurnal rhythm with a clear phase relationship with the zeitgeber, in which activity increases during the dark phase and decreases during the light phase (Fig. 1A and Fig. S2A–F). The higher or lower maximal values of luminiscence tend to anticipate zeitgeber transitions, suggesting that the bacteria are not only passively responding to external stimuli (Fig. 1A and Fig. S2A–F). Interestingly, a free running-rhythm (FR) in blsA promoter activity was maintained when the cultures were released to DD, showing a calculated period of 26 ± 2.4 h (rhythmic N = 124 populations, total N = 141, 87.9% rhythmic) (Fig. 1B and Supplementary Table 1), indicating the existence of a genuine circadian component. The amplitude of the oscillation in promoter activity was significantly greater under cyclic conditions than in DD (bLD: 0.014 ± 0.003 DNLU vs DD: 0.004 ± 0.002 DNLU, n = 124 populations rhythmic; Wilcoxon signed-rank test, ****p < 0.0001) (Fig. 1A and C). Under bLD conditions, the bioluminescence peaks occurred at zeitgeber time (ZT) 22.9 ± 0.8 h, which shifted to circadian time (CT) 14.6 ± 1.5 h under DD (Fig. 1D). A broad transition peak appeared on the first day under constant DD (Fig. 1A), suggesting that a zeitgeber masking component is present, despite an entrainment component cannot be discarded. Acrophases (times of bioluminescent peaks) determined on the broad transition peak of the first day under DD may not represent a reliable indicator for defining phase, since it reflects a population parameter very sensitive to small individual variations. Thus, the blue light zeitgeber is able to synchronize blsA promoter rhythms with a robust and clear phase relationship, while the bacterial populations still remain rhythmic under constant conditions, indicating the presence of an endogenous circadian oscillator. Overall, the data indicate that these important bacterial pathogens possess circadian rhythms with a clear response to light. Interestingly, promoter activity behaved similarly among the independent populations (i.e., different wells) in DD, increasing and decreasing following the same pattern, suggesting that the populations’ rhythms might be coordinated by some internal clue or remain as a post-effect of light exposure. Figure 1E–G shows luminescence data retrieved from individual rhythmic wells representative of the three independent experiments performed. Figure S3 shows similar experiments as above but performed using strains A. baumannii V15 pLPV1Z-pblsA::luc clones 2–4, recovered from independent transformation events. The data for these clones is consistent with that shown for clone 1.
Oscillations in blsA expression levels
To further validate the data obtained above, we studied whether promoter activity determined by luciferase activity correlated with blsA transcript levels. For this, A. baumannii V15 strain cultures were placed in 24 well-plates, incubated under 12bL:12D photoperiod and then released to constant darkness at 23 °C, in conditions similar as described above.
Samples were recovered from individual wells at selected timepoints corresponding to the end of the dark and light phases of the last day under LD photoperiod, as well as timepoints compatible with peaks and valleys observed under DD in Fig. 1 and Fig. S2. Total RNA was extracted, and blsA transcript levels were estimated by Reverse Transcription followed by Quantitative PCR (RT-qPCR), using blsA-specific primers and rpoB and recA as normalizers13. Our data indicate significant differences in blsA RNA transcript levels at the indicated timepoints at each day, both in LD and in DD-released cultures determined by RT-qPCR (Fig. 2), validating the oscillations found using the luciferase reporters at the RNA level.
blsA transcript levels were estimated by RT-qPCR in A. baumannii V15 wild-type cultured under 12bL:12D photoperiod cycles, and then released to constant darkness at 23 °C. The data shown are the mean ± SD of normalized relative quantities (NRQ)49 in three biological replicates measured in samples prepared from bacteria grown in LB broth. Significant differences determined by ANOVA followed by Tukey’s multiple comparison test (p < 0.05) are indicated by different letters. Gray and light blue colors correspond to dark and blue light incubations, respectively.
A. baumannii instantaneously resynchronizes to changes in the blue light zeitgeber
To further study synchronization and determine whether the rhythms could be reset by the zeitgeber, we applied a classical phase-shift approach in which bacterial A. baumannii V15 pLPV1Z-pblsA::luc clone 1 populations were first entrained to bLD cycles for 3 d (bLD1) and then were subjected to a phase shift caused by a 6-h night extension (lights onset were shifted from 7:00 AM to 1:00 PM), and incubated for 2 more days under the new scheme (bLD2). Bacterial populations showed a robust response to the zeitgeber, presenting an instantaneous resynchronization to the new bLD2 schedule (Fig. 3A and Fig. S4A). This fast response seems indicative of a masking phenomenon. It is interesting to note that the amplitude of the peaks remained conserved with respect to the previous bLD1 days before night lengthening (Fig. 3B), indicating that blsA promoter activity is not affected by culture aging or nutrient deprivation. From this, it follows that the reduction in amplitude observed when cultures are released to DD (Fig. 1(A, D) and S2D) is a characteristic of the endogenous rhythms and not an artifact resulting from those conditions. Finally, when cultures were released to DD after resynchronization, endogenous circadian rhythms were observed, while the phase appeared not conserved with respect to the new bLD2 phase (peak during bLD2: ZT2 22 ± 0.1 h; peak during DD: CT 5.9 ± 1.4 h) (Fig. 3C), again indicating the contribution of a masking component.
A Average A. baumannii V15 pLPV1Z-pblsA::luc clone 1 population luminescence rhythms after a 6-h phase shift in the 3rd LD day before constant darkness. The arrow indicates the time of the phase shift. The graph includes all traces of one independent experiment, n = 44. Black/blue bars indicate dark/light periods. Luminescence signals are shown as mean ± SD in blue (line and shadow, respectively), and the individual wells are represented in grey lines. B Average amplitude of the luminescence rhythm of rhythmic populations (n = 38). Data are shown as mean ± SD. Two-sample paired t-test, ****p < 0.0001. C Rayleigh plots showing the phase of the bioluminescent peak under cyclic conditions (bLD 1, black dots: 23.2 ± 0.2 h, R = 0.99; bLD 2, blue dots: 22 ± 0.1 h, R = 0.99; n = 38) and the bioluminescent peak on the first day of release to FR (DD, white dots: 5.9 ± 1.4 h, R = 0.93; n = 38) for the rhythmic population. Rayleigh test, P < 0.001. D–F Similar experiment as in (A-C) with the modification that the 6-h phase shift was applied at the last LD day. D The graph includes all traces of one independent experiment, which were all rhythmic, n = 11. The arrow indicates the time of the phase shift. Black/blue bars indicate dark/light periods. Luminescence signals are shown as mean ± SD in blue (line and shadow, respectively), and the individual wells are represented in grey lines. E Average amplitude of the luminescence rhythm of rhythmic populations (n = 11). Data are shown as mean ± SD. Two-sample paired t-test, ****p < 0.0001. F Rayleigh plots showing the phase of the bioluminescent peak under cyclic conditions (bLD 1, black dots: 21.8 ± 0.4 h, R = 0.99; bLD 2, blue dots: 22.5 ± 0.5 h, R = 0.99; n = 11) and the bioluminescent peak on the first day of release to FR (DD, white dots: 2.3 ± 0.5 h, R = 0.99; n = 11) for the rhythmic population. Rayleigh test, P < 0.001. Bacteria were grown at a constant temperature of 23 °C. G, H Individual luminescence plots for the rhythmic populations of A. baumannii V15 pLPV1Z-pblsA::luc clone 1 from the two resynchronization experiments, respectively. Bacteria were grown at a constant temperature of 23 °C. n = total number of samples; 1 sample = 1 well of a 96-well plate. n rhythmic = number of samples exhibiting circadian rhythmicity under free-running conditions (see Methods for definition).
To further study transition events observed in the first day under constant darkness (DD1, Fig. 1A) and determine whether entrainment or masking events are taking place, we performed a similar experiment as above, but with the modification that the 6-h night extension was applied to the last day under LD before application of constant darkness. Here again, a rapid response to the zeitgeber was observed, while endogenous circadian rhythms in DD presented a relative phase conservation (peak during bLD2: ZT2 22.5 ± 0.5 h; peak during DD: CT 2.3 ± 0.5 h) with respect to the new bLD2 phase (Fig. 3D, F), suggesting an entrainment component. Overall, our data show that a) there is a strong response to the blue light zeitgeber in blsA promoter activity, where masking and entrainment appear as contributing components leading to instantaneous resynchronization, and b) there are coordinated endogenous circadian rhythms under constant darkness.
BlsA is involved in synchronization to blue light in A. baumannii
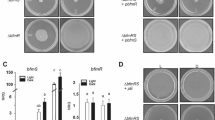
We further evaluated the behavior of another A. baumannii strain, ATCC 17978, under similar conditions as those described above. Our results show that 17978 harboring plasmid pLPV1Z-pblsA::luc presented a robust response to the light zeitgeber during the bLD photoperiod similarly to the V15 pLPV1Z-pblsA::luc strain, with a nocturnal increase in promoter activity (Fig. 4A, all traces of all replicates; Fig. S5, all traces and rhythmic population of individual biological replicates). Moreover, the presence of an endogenous rhythm was observed in DD (Figs. 4A and S5), which exhibited characteristics of canonical light entrainment (81.3% samples synchronized, 64% entrained) (Supplementary Table 1). It should be noted that cultures formed wall and pellicle biofilms, assessed qualitatively by visual inspection as well as quantitatively (Fig. S1). On the other hand, mutation in the blsA gene led to a significant decrease in circadian amplitude under bLD conditions, while the endogenous rhythm was maintained in DD (52-57.1% of the synchronized samples were rhythmic both in wt and blsA mutant, respectively; Chi square test p = 0,4654) (Fig. 4A, B, Fig. S5 and Supplementary Table 1). Most interestingly, synchronization to the blue light zeitgeber was severely compromised in ΔblsA pLPV1Z-pblsA::luc populations (only 46.6% samples synchronized; Chi square test p < 0.0001); while entrainment presented non-statistically significant differences with respect to the wild type (37.5% entrained, Chi square test p < 0.1) (Figs. 4A, B; S5 and Supplementary Table 1). The acrophase dispersion in rhythmic populations was higher in the bLD for 17978 ΔblsA pLPV1Z-pblsA::luc strain (Watson-Wheeler test for homogeneity of angles, p = 0.0312, *p < 0.05) (Fig. 4C). We did not observe any significant differences in the circadian period in ΔblsA pLPV1Z-pblsA::luc (24.02 ± 1.6 h, n = 16 mutant rhythmic, n total = 60, vs. 24.3 ± 1.4 h, n = 25 control rhythmic, n total = 59) (Fig. 4D and Supplementary Table 1). Thus, the overall results indicate that BlsA is involved in synchronization to the zeitgeber but does not affect the endogenous rhythm component.
A Reporter activity of A. baumannii 17978 pLPV1Z-pblsA::luc clon B populations. The graphs shows all traces of two independent experiments (A. baumannii 17978 pLPV1Z-pblsA::luc clone B, n = 59; 17978 ΔblsA pLPV1Z-pblsA::luc clone B, n = 60). The cultures were incubated in bLD cycles for 4 days and then released to DD. Black/blue bars indicate dark/light periods. Luminescence signals are shown as mean ± SD in blue (line and shadow, respectively), and the individual wells are represented in grey lines. B Average amplitude of the luminescence rhythm of rhythmic populations of A. baumannii 17978 pLPV1Z-pblsA::luc clone B (n = 25) and 17978 ΔblsA pLPV1Z-pblsA::luc (n = 16). Data are shown as mean ± SD. Two-way ANOVA followed by Tukey’s multiple comparisons test. ****p < 0.0001; ns non-significant. C Rayleigh plots showing the phase of the bioluminescent peak under cyclic conditions (bLD, black dots) and the bioluminescent peak on the first day of release to FR (DD, white dots) for the rhythmic population: A. baumannii 17978 pLPV1Z-pblsA::luc clone B (bLD: ZT21.6 ± 0.8 h, R = 0.97 and DD: CT19.3 ± 2.9 h, R = 0.69; n = 25) and 17978 ΔblsA pLPV1Z-pblsA::luc (bLD: ZT22.4 ± 2 h, R = 0.84 and DD: CT16.8 ± 3.9 h, R = 0.48; n = 16). Lines represent the average peak phase of pLPV1Z-pblsA::luc activity (mean vectors for the circular distributions) of each group. The length of the vector represents the strength of the phase clustering, while the angle of the vector represents the mean phase. Individual data points are plotted outside the circle. The central circle represents the threshold for p = 0.05. D Average endogenous period of rhythmic populations of A. baumannii 17978 pLPV1Z-pblsA::luc clone B (24.3 ± 1.4 h, n = 25) and 17978 ΔblsA pLPV1Z-pblsA::luc (24.02 ± 1.6 h, n = 16). Data are shown as mean ± SD. Wilcoxon rank sum test, ns non-significant. E, F Individual luminescence plots for the rhythmic populations of A. baumannii 17978 pLPV1Z-pblsA::luc clone B (E) and 17978 ΔblsA pLPV1Z-pblsA::luc (F). Bacteria were grown at a constant temperature of 23 °C.
A. baumannii is rhythmic under constant dark conditions
To further explore endogenous circadian rhythms in A. baumannii, we next decided to study the blsA promoter activity under constant darkness conditions from the beginning of the experiment and for 7.5 days. The presence of rhythmic oscillations was observed in populations of both V15 pLPV1Z-pblsA::luc clone 1 and 17978 pLPV1Z-pblsA::luc clone B (51% and 44% rhythmic, respectively), with a calculated period of 23.9 ± 4,5 and 24.7 ± 4, respectively (Fig. 5A, B, E-F), indicating the existence of endogenous circadian rhythms under constant dark conditions. Single wells of rhythmic samples are shown in Fig. 5D and H. Interestingly, acrophases were not clusterized but distributed along different hours of the day (Fig. 5C and G). Consistent results were observed for biological replicates, shown in Fig. S6. This indicates that in the absence of previous zeitgeber exposure, the independent populations, i.e., the different wells, loose coordination and behave quite independently, suggesting that light is the signal synchronizing their behavior (DD in Figs. 1A, 4A). Also interesting is the fact that 17978 pLPV1Z-pblsA::luc clone B under DD behaves similarly as 17978 ΔblsA pLPV1Z-pblsA::luc in LD. This makes sense since this strain is compromised in synchronization to the zeitgeber, consistent with being a photoreceptor mutant deficient in light sensing, without significant effects under DD.
Luminescence reporter activity of A. baumannii V15 pLPV1Z-pblsA::luc clone 1 (A–D) and 17978 pLPV1Z-pblsA::luc clon B (E–H) populations under DD for 7.5 days. The graph in A shows all traces of A. baumannii V15 pLPV1Z-pblsA::luc clone 1, n = 47, while only rhythmic samples are shown in (B), n = 24. Baseline-detrended, normalized luminescence is shown. Average endogenous period (FRP) of rhythmic populations of A. baumannii V15 pblsA::luc clone 1 (23.9 ± 4.5 h, n = 24), and the time window for period calculation is indicated in the top right of panel (B). A representative experiment from two biological replicates is shown. Individual wells are represented in colored lines. C Rayleigh plots showing the phase of the bioluminescent peak under constant darkness DD conditions for the rhythmic population (DD, black dots: 21.9 ± 4.9 h, n = 24; R = 0.1, for the rhythmic population). Lines represent the average peak phase of pLPV1Z-pblsA::luc activity (mean vectors for the circular distributions) of each group. The length of the vector represents the strength of the phase clustering, while the angle of the vector represents the mean phase. Individual data points are plotted outside the circle. The central circle represents the threshold for p = 0.05. (D) Individual luminescence plots for the rhythmic populations of A. baumannii V15 pLPV1Z-pblsA::luc clone 1. The graph in E shows all traces of 17978 pLPV1Z-pblsA::luc clon B, n = 36, while only rhythmic samples are shown in (F), n = 19. Baseline-detrended, normalized luminescence is shown. Average endogenous period (FRP) of rhythmic populations of A. baumannii 17978 pLPV1Z-pblsA::luc clone B (24.67 ± 4 h, n = 19) and the time window for period calculation are indicated in the top right of B. A representative experiment (biological replica 1) from two biological replicates is shown. Individual wells are represented in colored lines. G Rayleigh plots showing the phase of the bioluminescent peak under constant darkness DD conditions for the rhythmic population (DD, black dots: 12.6 ± 5 h, n = 16; R = 0.1, for the rhythmic population). Lines represent the average peak phase of pLPV1Z-pblsA::luc activity (mean vectors for the circular distributions) of each group. The length of the vector represents the strength of the phase clustering, while the angle of the vector represents the mean phase. Individual data points are plotted outside the circle. The central circle represents the threshold for p = 0.05. (H) Individual luminescence plots for the rhythmic populations of A. baumannii 17978 pLPV1Z-pblsA::luc clon B. The parameters were determined using BioDare2. Bacteria were grown at a constant temperature of 23 °C. n = total number of samples; 1 sample = 1 well of a 96-well plate. n rhythmic = number of samples exhibiting circadian rhythmicity under free-running conditions (see Methods for definition).
Discussion
It has recently been ascertained that many non-phototrophic pathogens are capable of sensing and responding to light via specialized molecular systems such as photoreceptors7,8,9,10. In fact, we have extensively shown that the critical human pathogen A. baumannii presents a global response to light, including modulation of pathogenicity and virulence7,8,12,13,14,16,17,18,30,35,36,37,38,39. This prompted us to determine whether this response to light is actually part of a more specialized mechanism, such as a light-dependent circadian pacemaker. Given that A. baumannii’s environmental life plays a key role in its pathophysiology, and that the bacterial physiological state at the onset of infection, which could be defined by environmental circadian rhythms, determines the outcome of infection and host fate23, we decided to perform our studies at environmental temperatures that mimic the initial, pre-infectious bacterial state.
In this work, we provide evidence indicating that blsA promoter activity shows a robust response to bLD cycles leading to daily rhythms, which anticipate zeitgeber changes, suggesting an endogenous oscillation. In fact, endogenous free-running rhythms were observed under constant darkness. Daily rhythms are very important as they show variations in bacterial physiology due to changing environmental conditions throughout the day, while the existence of endogenous rhythms suggests that this pathogen has evolved mechanisms to anticipate periodic variations in these conditions.
Interestingly, our data strongly suggest the contribution of both masking and entrainment components in the response to light of circadian rhythms in the V15 strain. Masking is an important evolutionary mechanism that allows an organism to respond to changes in exogenous stimuli (e.g., light-dark cycle, social cues, temperature, food, drugs), thereby enabling the organism to act immediately and appropriately31. In addition, masking can complement and integrate with entrainment if the cues align with arousal and the circadian system40.
The circadian response to light depends on the timing of light exposure, as well as its intensity, duration, wavelength, and prior light exposure history. For example, the threshold for human entrainment requires relatively bright light; on the contrary, the circadian system of mice is exquisitely sensitive to light41: a 12 L:12D photocycle using light intensities as low as 0.01–0.1 lux will photoentrain murine rhythms. The robust response observed in the bLD cycle indicates that the minimum light timing and intensity (see methods) information necessary has been reached in our conditions. Blue light sensing is an integral part of circadian rhythms in all the experimental models examined so far. It should be noted that we are using very low light intensity (8 μE m−2 s−1), more than 4 times lower than that reported in photoentrained circadian rhythms evidenced in B. subtilis2. Thus, a priori, we would not expect the blue light intensity used in our setup to be too high to lead to masking effects prevailing over entrainment. Also interesting is that in DD (after LD conditions), the whole bacterial population behaves similarly, i.e., blsA promoter activity increases and decreases following the same pattern in different wells, suggesting that the populations might be coordinated by some internal clue or remain in phase as a post-effect of light exposure. Indeed, as non-photosynthetic bacteria constitute poorly characterized organisms in terms of circadian rhythmicity, we can now conclude that synchronization by light is a conserved mechanism. In fact, bacteria introduce a new dimension in the understanding of circadian rhythms, and it would thus not be surprising that non-canonical features are discovered regarding circadian rhythms in these microorganisms. It is also likely that different strains present differential characteristics regarding synchronization and entrainment by light, as well as endogenous circadian components, most probably because of differential lifestyle, antibiotic susceptibility profile, pathogenicity, etc.
In this work, we characterized blsA as it is the photoreceptor governing photoregulation at environmental temperatures in A. baumannii, showing that it is involved in synchronization to the zeitgeber during light-dark cycles. BlsA production is under feedback control, as the promoter firing is compromised in the blsA mutant, involving other components such as the BfmRS system, which has previously been shown to modulate blsA expression17 and physically interact with BlsA19. Further work studying the involvement of BfmRS in clock entrainment, as well as rhythmicity in other genes and phenotypic responses, will contribute to gain full understanding of circadian rhythms in this critical pathogen. Interestingly, under constant darkness in the absence of entrainment, A. baumannii wild-type strains behaved similarly to the blsA mutant under light-dark cycles, consistent with ΔblsA being a photoreceptor mutant deficient in light sensing. In this context, it would be interesting to also study circadian rhythms at the host’s physiological temperature, which is beyond the scope of this work. Moreover, the effect of temperature entrainment, which could occur after infection of the host, is also a possibility to be explored.
A. baumannii’s rhythm was detected in conditions of biofilm formation, as occurred with B. subtilis2, which is known to be a bacterial social behavior regulated by quorum sensing16. In this context, another interesting aspect to further explore is whether communication between the bacteria, these apparent “independent individuals” that constitute the bacterial population, is taking place in circadian rhythm generation. Several questions arise here such as whether the bacteria interact through quorum sensing, a phenomenon known to be modulated by light in A. baumannii biofilms16,18; or how is the circadian mechanism transmitted to new generations in dividing short-lived bacteria, considering that the circadian clock is thought to be dependent on state variables, i.e., substances that reflect time, whose concentrations might be disrupted by the cell division process. In this sense, the possibility that P. aeruginosa quorum sensing and light sensing integration to control virulence and biofilm formation is under circadian regulation has been proposed before42. Moreover, in cyanobacteria a memory effect has been identified, which spans over the cell cycle interval (i.e., the physiological state of a bacterium is sustained for several cell divisions)1.
This work contributes to establishing that bacterial pathogens are subjected to circadian regulation, opening a new full area of research that will likely lead to reinterpretation of previous data on the pathogen’s behavior. The observation of daily and circadian rhythms in different strains, given the high genetic heterogeneity present within the A. baumannii species due to genome plasticity, plasmid content, etc, is an additional and positive validation. Since A. baumannii is phylogenetically related to K. aerogenes, but distant from B. subtilis and Synechococcus sp. (Fig. S7), microorganisms in which the existence of circadian clocks has been demonstrated1,2,3,4,5, our work contributes to broadening the evolutionary spectrum of circadian rhythms in bacteria. We expect this work will inspire future efforts to investigate whether bacterial pathogens’ circadian rhythms modulate their behavior in the environment, and most importantly, whether they can synchronize their behavior to the host’s circadian rhythm and immune response to optimize infection, which was demonstrated in other pathogens. Indeed, it has been recently shown that internal timekeeping mechanisms in the malaria parasite synchronize with the host’s circadian rhythm. An example of this is the synchronization of the rupture of red blood cells with the completion of the parasite’s asexual cell cycle43.
Changes in bacterial antibiotic susceptibility, infection outcomes or persistence in the environment influenced by the time of the day could introduce modifications in treatment schedules to optimize medical interventions and prevention of critical infections, offering new opportunities for the development of targeted therapeutic strategies to combat infectious diseases.
In summary, our work contributes to the developing field of circadian clocks in bacterial human pathogens, indicating the existence of daily as well as endogenous circadian rhythms in a critical pathogen, which could impact the microorganisms’ lifestyle and its pathogenicity.
Methods
Media
Lysogeny Broth (LB)33 composition was triptein 10 g/L (Britania), NaCl 5 g/L (Ciccarelli), and yeast extract 5 g/L (Britania). When required agar 1.5 g/L (Britania) was included.
Strains and constructions
The strains used in this study are reported in Table S1. The promoter region of the blsA gene was amplified using primers blsA_EcoRI_FW (5′- GAATTCagtattacaaattgaacgtgt -3′) and blsA_BamHI_REV (5′- GGATCCaagacttccgtgaaatataaa -3′). Primers were purchased at Integrated DNA Technologies, USA. High-fidelity polymerase chain reaction products were digested with EcoRI and BamHI enzymes (Promega) and cloned into the corresponding sites of pLPV1Z harboring the promoterless luxABCDE genes34. The correct construction was verified by sequencing, and pLPV1Z-PblsA::luc was subsequently introduced into A. baumannii V15 strain by transformation.
Light settings
Samples were exposed to blue light emitted by nine-LED (light-emitting diode) arrays with an intensity of 6 to 10 µmol photons/m2/s and peak emission centered at 462 nm 8. Light intensity was measured using a radiometer/photometer (Flame-T, OceanOptics). Temperature was set at 23 °C, and fluctuations in the incubator were less than 0.5 °C.
Zeitgeber (i.e., “time giver” or entraining agent) time 0 or ZT0 (9:00 am) indicates the time at which lights were turned on. Circadian Time (CT) refers to a specific time in the free-running conditions (constant darkness, DD, and constant temperature of 23 °C). Photo and thermal conditions were controlled with an I-291PF incubator (INGELAB, Argentina), and temperature was monitored using DS1921H-F5 iButton Thermochrons (Maxim Integrated, USA).
Luminescence assays
For all assays, A. baumannii cells were cultured in white 96-well plates (Greiner) under static conditions in LB broth (250 µl for well) from an initial OD660 of 0,05. Plates were sealed with a transparent optical film (ThermalSeal RT2RRTM, EXCEL Scientific) to avoid evaporation and contamination, and the seal over each well was perforated twice to avoid condensation and allow oxygen exchange. Cultures were exposed to light-darkness cycles (12 h blue light (bL) and 12 h dark (D)) for 4 days, after which the cultures were released to constant darkness (free-running condition). All experiments were carried out in temperature-controlled incubators (I-291PF incubator, INGELAB, Argentina). The temperature was kept constant at 23 °C. Temperature fluctuations in the incubator due to lights being on or off were less than 0.5 °C. The luminescence of each well was integrated for 10 s every 30 min (Berthold Centro LB 960 microplate luminometer, Berthold Technologies). Microwin 2000 software version 4.43 (Mikrotek-441 Laborsysteme) was programmed to leave the plate outside the luminometer after each recording to expose A. baumannii cells to the environmental cues. It should be noted that blue light is not expected to affect luciferase activity measurement, since blue light application is spatially and temporally separated from luciferase emission detection. In fact, bioluminescence was measured inside the luminometer equipment, while the plate was ejected between readings, for exposure to blue light.
For phase-shift assays, cells were entrained for 3 days under a bLD cycle and then were subjected to a phase shift caused by a 6-h night extension. After 1 (or 3) more days under regular bLD, cells were released into constant darkness conditions.
For free-running (FR) assays, cells were exposed for 10 days to constant darkness conditions.
In all cases, 1 sample = 1 well of a 96-well plate.
The presence of biofilms was assessed qualitatively, by eye, as a pellicle forming at the air-liquid interface in the well. Also, pellicles and wall biofilms were quantified as described in the Biofilms assays item below.
Analysis of FRP following 4 days of entrainment with bLD cycles
In all cases, the first 24 to 36 hours of recording were removed due to accumulation of the luciferase enzyme. Raw data were analyzed using CircaLuc v0.7, a Shiny application developed in our laboratory for the analysis of periodic luminescence data, freely available at https://ispiousas.shinyapps.io/circaluc/44,45. CircaLuc linearly detrends and smooths luminescence signals (with adjustable parameters), estimates the circadian period using the Lomb-Scargle (LS) periodogram (via the Lomb R package)46, with an oversampling of 30. The acrophase (time at peak) and amplitude of each signal were estimated using the Cosinor method by fitting a cosine waveform to the data using a non-linear least squares regression implemented through the NLS function of base R, with R² used to evaluate the goodness of fit. The data is shown as mean ± SD of luminescence. Periods were considered as circadian when their duration was between 18 and 33 hours. Background signals from the bacteria transformed with the empty luc-plasmid or LB broth alone were at least 10 folds of magnitude lower than signals retrieved from the strains expressing luc directed from the blsA promoter.
Under bLD conditions, any luminescent signal of a sample with a period of 24 h and an R2 adjustment ≥ 0.5 was considered to be “synchronized”. In the case of DD conditions, any luminescent signal with a period range between 18 and 33 h and an R2 adjustment ≥ 0.5, which had previously synchronized to the bLD cycle, was considered to be “rhythmic”. We considered “entrained” samples to be those rhythmic samples whose acrophases in FR conditions had a difference of less than 3 h with respect to that of the entrainment conditions. To avoid confusion, the following nomenclature is used in this work: “percentage of synchronized” means the number of synchronized samples over the total number of samples tested; “percentage of rhythmic under DD” means the number of rhythmic samples over the total number of samples tested; and “percentage of entrained” represents the number of entrained samples over the number of rhythmic samples.
Analysis of FRP in the absence of entrainment
For data shown in Fig. 5 and Fig. S6, data analysis of the period was performed using Fast Fourier Transform, nonlinear least squares function of the BioDare2 suite47, only for samples that passed the Jonckheere-Terpstra-Kendall (JTK) cycle test for rhythmicity (P < 0.01) on nonnormalized, baseline detrended data. The P values were corrected using the Benjamini-Hochberg procedure for multiple testing. Periods were considered circadian when their duration was between 18 and 34 hours.
Plotting data
Graphs were generated using either GraphPad Prism software version 8.0.1 or the ggplot2 package in R version 4.4.348. Final figures were generated using Biorender (https://app.biorender.com/).
Bacterial cultures for RNA extraction
A. baumannii V15 cells were cultured in 24-well microplates under static conditions in LB broth at 23 °C from an initial OD660 of 0,05. The bacteria were incubated for 4 days under 12 L/12D photoperiod and then released to constant darkness. 2 ml samples were retrieved at the selected timepoints indicated in the text and figures. The samples were centrifuged, and the pellets were saved at -80 °C until further use.
RNA extraction
RNA was extracted following procedures described in Muller et al., 201713. Briefly, pelleted cells were mixed with 0.5 ml of lysis buffer (0.1 M sodium acetate, 10 mM EDTA, 1% SDS) in a boiling-water bath. Cell lysates were extracted three times at 60 °C with one volume of phenol, which was adjusted to pH 4.8 with 50 mM sodium acetate, and then once with chloroform at room temperature. The RNA precipitated overnight at -20 °C with 2.5 volumes of ethanol was collected by centrifugation, washed with 70% ethanol, and dissolved in diethyl pyrocarbonate (DEPC)-treated deionized water at 65 °C. RNA quality and quantity were evaluated by gel electrophoresis and by determination of the relationship of absorbance at 260 nm to absorbance at 280 nm.
Gene expression analysis via RT-qPCR
Total RNA (0.5 µg) was treated with RQ1 RNase-free DNase (Promega). First-strand cDNA synthesis was performed using random priming and Moloney Murine Leukemia Virus Reverse Transcriptase (MMLV; Invitrogen). PCR was performed in an AriaMx thermal cycler (Agilent Technologies, Santa Clara, CA, USA) using SYBR Green I to monitor double-stranded DNA synthesis. Normalized relative quantities (NRQ) of blsA were obtained using the qBase method49 with recA and rpoB as reference genes for normalization across samples. The qPCR reactions were performed the primers V15blsA.rtF 5’CCTGTGTTATGCCAGCCAACGAA3’ and V15blsA.rtR 5’ TCAACGACCTCTTGTTCGCCTTCT3’ for blsA; rpoBrt.F 5’CAGAAGTCACGCGAAGTTGAAGGT3’ and rpoBrt.R 5’AACAGCACGCTCAACACGAACT3’ for rpoB; and recAF.rt 5’TACAGAAAGCTGGTGCATGG3’ and recAR.rt 5’TGCACCATTTGTGCCTGTAG3’ for recA15. The stability of the normalizers across all samples was determined by calculating the coefficient of variation (CV) of their NRQs. Both reference genes showed CVs below 25% (0.237 for rpoB and 0.249 for recA), which is within the accepted range for stably expressed reference genes. Melting curve analyses at the end of the process and ‘no template controls’ were performed to ensure product-specific amplification without primer-dimer artifacts.
Biofilm assays
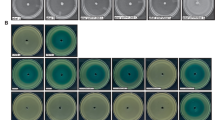
For biofilm assays, a dilution in fresh LB broth to OD660 = 0.05 of an overnight shaking culture grown at 37 °C of each strain of interest medium was inoculated into 96 96-well plate. The cultures were then incubated for 4 days statically in 12bLD:12D photocycle (LD4), and a 5th day in DD (DD1) at 23 °C. Wall biofilms, i.e., biofilms that form in the liquid-plastic interphase, were visualized and quantified by crystal violet staining (Fig. S1 A and B), as described in Mussi et al., 20108. Briefly, the supernatant of each well at each condition (n = 5) was aspirated and rinsed thoroughly with distilled water. Then, a 0.1% crystal violet solution was added and incubated for 1 hour, after which the wells were rinsed again thoroughly with distilled water. The biofilms attached to the tube walls were visualized and photographed, and then solubilized using an 80:20 ethanol–acetone solution. The OD580/OD660 ratio was used to normalize the amount of biofilm formed to the total cell content of each sample tested to avoid variations due to differences in bacterial growth under different experimental conditions. Pellicles that formed on the culture surfaces at the air-liquid interphase were detected by visual inspection. Also, to estimate pellicles on the liquid surfaces, the culture medium was removed gently with a Pasteur pipette without losing pellicles. The cells that remained in the tubes were resuspended in 1 ml of sterile phosphate-buffered saline (PBS) solution, and the cell density was determined by measurement of OD660. The amount of biofilm formed by each sample was normalized to its total biomass, which was determined by measuring the OD660 of the whole culture in each well (Fig. S1 C)8.
Statistics and reproducibility
All statistical analyses were carried out using Prism 8 software version 8.4.3. Normality was checked using the Shapiro-Wilk test. When data were normally distributed, means between two groups of values were compared using the two-sample Student’s t-test or paired t-test, and two-way ANOVA was used to compare the means between more than two groups. When the assumption of normality was not met, Wilcoxon rank sum test, Wilcoxon signed-rank test or the Watson-Wheeler test for homogeneity of angles were employed. Chi-square test was performed to compare proportions. Results were considered significant at 5% significance level. The exact sample sizes (“n” values) as well as the number of biologically independent experiments are indicated in the figure legends. Luminometry experiments involved sample sizes (n) ranging from 11 to 60, and 2 to 3 biologically independent experiments. RT-qPCR, biofilm formation, and growth experiments involved sample sizes (n) ranging from 3 to 5, and 3 biologically independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Code availability
The CircaLuc code is available on Zenodo at link https://doi.org/10.5281/zenodo.16740196.
References
Swan, J. A., Golden, S. S., LiWang, A. & Partch, C. L. Structure, function, and mechanism of the core circadian clock in cyanobacteria. J. Biol. Chem. 293, 5026–5034 (2018).
Eelderink-Chen, Z. et al. A circadian clock in a nonphotosynthetic prokaryote. Sci. Adv. 7, https://doi.org/10.1126/sciadv.abe2086 (2021).
Sartor, F. et al. The circadian clock of the bacterium B. subtilis evokes properties of complex, multicellular circadian systems. Sci. Adv. 9, eadh1308 (2023).
Paulose, J. K., Cassone, C. V., Graniczkowska, K. B. & Cassone, V. M. Entrainment of the Circadian Clock of the Enteric Bacterium Klebsiella aerogenes by Temperature Cycles. iScience 19, 1202–1213 (2019).
Paulose, J. K., Wright, J. M., Patel, A. G. & Cassone, V. M. Human gut bacteria are sensitive to melatonin and express endogenous circadian rhythmicity. PloS one 11, e0146643 (2016).
Wagner, P. M. et al. Rhythms in lipid droplet content driven by a metabolic oscillator are conserved throughout evolution. Cell Mol. Life Sci. 81, 348 (2024).
Tuttobene, M. R. et al. Light modulates important pathogenic determinants and virulence in ESKAPE Pathogens Acinetobacter baumannii, Pseudomonas aeruginosa, and Staphylococcus aureus. J. Bacteriol. 203, https://doi.org/10.1128/JB.00566-20 (2021).
Mussi, M. A. et al. The opportunistic human pathogen Acinetobacter baumannii senses and responds to light. J. Bacteriol. 192, 6336–6345 (2010).
Mukherjee, S., Jemielita, M., Stergioula, V., Tikhonov, M. & Bassler, B. L. Photosensing and quorum sensing are integrated to control Pseudomonas aeruginosa collective behaviors. PLoS Biol. 17, e3000579 (2019).
Swartz, T. E. et al. Blue-light-activated histidine kinases: two-component sensors in bacteria. Science 317, 1090–1093 (2007).
Rinaldi, J. et al. The beta-scaffold of the LOV domain of the Brucella light-activated histidine kinase is a key element for signal transduction. J. Mol. Biol. 420, 112–127 (2012).
Abatedaga, I. et al. Integration of Temperature and Blue-Light Sensing in Acinetobacter baumannii Through the BlsA Sensor. Photochem. Photobiol. 93, 805–814 (2017).
Muller, G. L. et al. Light Modulates Metabolic Pathways and Other Novel Physiological Traits in the Human Pathogen Acinetobacter baumannii. J. Bacteriol. 199, https://doi.org/10.1128/JB.00011-17 (2017).
Ramirez, M. S. et al. White and blue light induce reduction in susceptibility to minocycline and tigecycline in Acinetobacter spp. and other bacteria of clinical importance. J. Med. Microbiol. 64, 525–537 (2015).
Tuttobene, M. R., Cribb, P. & Mussi, M. A. BlsA integrates light and temperature signals into iron metabolism through Fur in the human pathogen Acinetobacter baumannii. Sci. Rep. 8, 7728 (2018).
Tuttobene, M. R. et al. Blue light directly modulates the quorum network in the human pathogen Acinetobacter baumannii. Sci. Rep. 11, 13375 (2021).
Perez Mora, B. et al. BfmRS encodes a regulatory system involved in light signal transduction modulating motility and desiccation tolerance in the human pathogen Acinetobacter baumannii. Sci. Rep. 13, 175 (2023).
Tuttobene, M. R. et al. Quorum and light signals modulate acetoin/butanediol catabolism in Acinetobacter spp. Front. Microbiol. 10, 1376 (2019).
Chitrakar, I. et al. Structural basis for the regulation of biofilm formation and iron uptake in A. baumannii by the Blue-Light-Using Photoreceptor, BlsA. ACS Infect. Dis. 6, 2592–2603 (2020).
Spellberg, B. & Bonomo, R. A. The deadly impact of extreme drug resistance in Acinetobacter baumannii. Crit. care Med. 42, 1289–1291 (2014).
Strachan, C. R. & Davies, J. The Whys and Wherefores of Antibiotic Resistance. Cold Spring Harbor Perspectives in Medicine 7, https://doi.org/10.1101/cshperspect.a025171 (2017).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Wong, D. et al. Clinical and pathophysiological overview of acinetobacter infections: a century of challenges. Clin. Microbiol. Rev. 30, 409–447 (2017).
Mea, H. J., Yong, P. V. C. & Wong, E. H. An overview of Acinetobacter baumannii pathogenesis: Motility, adherence and biofilm formation. Microbiol. Res 247, 126722 (2021).
Gaddy, J. A., Tomaras, A. P. & Actis, L. A. The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and in the interaction of this pathogen with eukaryotic cells. Infect. Immun. 77, 3150–3160 (2009).
McQueary, C. N. & Actis, L. A. Acinetobacter baumannii biofilms: variations among strains and correlations with other cell properties. J. Microbiol. 49, 243–250 (2011).
Tomaras, A. P., Dorsey, C. W., Edelmann, R. E. & Actis, L. A. Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology 149, 3473–3484 (2003).
Tomaras, A. P., Flagler, M. J., Dorsey, C. W., Gaddy, J. A. & Actis, L. A. Characterization of a two-component regulatory system from Acinetobacter baumannii that controls biofilm formation and cellular morphology. Microbiology 154, 3398–3409 (2008).
Konig, P. et al. The VBNC state: a fundamental survival strategy of Acinetobacter baumannii. mBio 14, e0213923 (2023).
Golic, A. E. et al. BlsA Is A Low To Moderate Temperature Blue Light Photoreceptor In The Human Pathogen Acinetobacter baumannii. Front. Microbiol. 10, 1925 (2019).
John, T. J. & Kompithra, R. Z. Eco-epidemiology triad to explain infectious diseases. Indian J. Med Res 158, 107–112 (2023).
Chen, W. Host-pathogen interactions in Acinetobacter baumannii infection: recent advances and future challenges. Future Microbiol. 15, 841–845 (2020).
Bertani, G. Lysogeny at mid-twentieth century: P1, P2, and other experimental systems. J. Bacteriol. 186, 595–600 (2004).
Lucidi, M. et al. New Shuttle Vectors for Real-Time Gene Expression Analysis in Multidrug-Resistant Acinetobacter Species: In Vitro and In Vivo Responses to Environmental Stressors. Appl. Environ. Microbiol. 85, https://doi.org/10.1128/AEM.01334-19 (2019).
Arana, N. et al. Light regulation in critical human pathogens of clinical relevance such as Acinetobacter baumannii, Staphylococcus aureus and Pseudomonas aeruginosa. Photochem. Photobiol. Sci. https://doi.org/10.1007/s43630-023-00437-x (2023).
Abatedaga, I. et al. Characterization of BLUF-photoreceptors present in Acinetobacter nosocomialis. PloS one 17, e0254291 (2022).
Ramirez, M. S., Muller, G. L., Perez, J. F., Golic, A. E. & Mussi, M. A. More than just light: clinical relevance of light perception in the nosocomial pathogen acinetobacter Baumannii and other members of the genus Acinetobacter. Photochem. Photobiol. 91, 1291–1301 (2015).
Golic, A. et al. Staring at the cold sun: blue light regulation is distributed within the genus Acinetobacter. PloS one 8, e55059 (2013).
Pezza, A. et al. Through the eyes of a pathogen: light perception and signal transduction in Acinetobacter baumannii. Photochem. Photobiol. Sci. 18, 2363–2373 (2019).
Gall, A. J. & Shuboni-Mulligan, D. D. Keep your mask on: the benefits of masking for behavior and the contributions of aging and disease on dysfunctional masking pathways. Front Neurosci. 16, 911153 (2022).
Foster, R. G., Hughes, S. & Peirson, S. N. Circadian photoentrainment in mice and humans. Biology 9, https://doi.org/10.3390/biology9070180 (2020).
Sartor, F. & Kovacs, A. T. Rhythmic spatial self-organization of bacterial colonies. mBio 13, e0170322 (2022).
Rijo-Ferreira, F. et al. The malaria parasite has an intrinsic clock. Science 368, 746–753 (2020).
Lamberti, M. L. et al. Clock gene homologs lin-42 and kin-20 regulate circadian rhythms in C. elegans. Sci. Rep. 14, 12936 (2024).
Spiousas, I. gnacio spiousas/CircaLuc. Zenodo https://doi.org/10.5281/zenodo.16740196 (2025).
Van Dongen, H. P., Olofsen, E., VanHartevelt, J. H. & Kruyt, E. W. A procedure of multiple period searching in unequally spaced time-series with the Lomb-Scargle method. Biol. Rhythm Res. 30, 149–177 (1999).
Zielinski, T., Moore, A. M., Troup, E., Halliday, K. J. & Millar, A. J. Strengths and limitations of period estimation methods for circadian data. PloS one 9, e96462 (2014).
Wickham, H. ggplot2. (Springer, 2016).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 8, R19 (2007).
Acknowledgements
This work was supported by grants from the Agencia Nacional de Promoción Científica y Tecnológica to MAM (PICT 2019-01484), Secretaría de Ciencia, Tecnología e Innovación (Provincia de Santa Fe) to MAM (PEIC I + D 2023-255), and DG (PICT 2021-1051). MAM, RER, BPM, and DG are career investigators of CONICET, while VP and NA are fellows from the same institution. We thank Dr. Ignacio Spiousas (Escuela de Educación, Universidad de San Andrés) for his kind assistance using CircaLuc and data analyses. We also thank Dr. Adrián E. Granada (Universitat Medizin, Berlin, Germany) for his kind assistance using PyBoat and data analyses.
Author information
Authors and Affiliations
Contributions
Conceptualization: V.P., B.E.P.M., M.L.M., D.A.G. and M.A.M. Formal analysis: R.E.R., V.P., B.E.P.M., M.L.M., N.A., M.L.L., J.I.F., M.B.A., D.A.G. and M.A.M. Funding acquisition: M.A.M and D.A.G. Investigation: V.P., B.E.P.M., M.L.M., M.L.L., J.I.F., M.B.A., G.D.V., M.F., D.A.G. and M.A.M. Methodology: R.E.R., V.P., B.E.P.M., M.L.M., N.A., D.A.G. and M.A.M. Project administration: M.A.M. Visualization: V.P., M.L.M., J.I.F., M.B.A. and M.A.M. (Writing—original draft: M.A.M and D.A.G. Writing—review & editing: V.P., B.E.P.M., M.L.M., J.I.F., M.B.A., G.D.V., M.F., D.A.G. and M.A.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Ranjana Pathania and David Favero.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Permingeat, V., Perez Mora, B.E., Migliori, M.L. et al. The critical human pathogen Acinetobacter baumannii exhibits light-regulated circadian rhythms. Commun Biol 8, 1385 (2025). https://doi.org/10.1038/s42003-025-08732-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08732-2