Abstract
As a major environmental pathogenic factor for various skin diseases, UVB radiation leads to oxidative stress and biomacromolecule damage. Autophagy is a highly conserved catabolic process and serves as one of the main mechanisms to maintain cellular homeostasis. Here, by CRISPR/Cas9-mediated gene deletion, we demonstrate that the essential transcriptional repressor BCL11A is involved in autophagy regulation and participates in the UVB-induced stress response. BCL11A deficiency increases autophagosome formation and enhances the intensity of autophagy flux with or without UVB stress. Mechanistically, ACSS3, rather than autophagy-related genes, is identified as the direct target gene and transcriptionally repressed by BCL11A. Further, BCL11A deficiency reduces DNA damage and ROS to promote survival and inhibit apoptosis under UVB irradiation, which is blocked by pharmacological inhibition of autophagy or BCL11A overexpression. Collectively, BCL11A deficiency promotes autophagy activation to clear ROS and DNA damage, thereby protecting epidermal cells from UVB-induced death.
Similar content being viewed by others
Introduction
Autophagy is an evolutionarily conserved lysosome-dependent process that catabolizes cellular components and recycles materials engulfed by autophagosomes. In various pathological and physiological processes, autophagy is an essential mechanism for maintaining cell viability by removing damaged organelles and materials1. Keratinocytes that possess defective autophagy display a significant reduction in antioxidant capacity and increased oxidative damage2. In other epithelium cells, such as retinal pigment epithelium cells, autophagy attenuates oxidative damage and promotes cell survival by clearing ROS3. Emerging evidence indicates that autophagy also participates in the removal of DNA lesions, which contributes to maintaining cellular viability4, regulating cell radiosensitivity5 and apoptosis6. Ultraviolet B (UVB) irradiation is the most common environmental pathogenic factor for skin, especially the epidermis. DNA damage and ROS production are two major mechanisms by which UVB induces cell damage, eventually leading to various skin diseases. Due to the ability to remove peroxidized substances and damaged nucleotides, autophagy activation is essential to reduce apoptosis, slow down photoaging and effectively suppress inflammation in response to UVB radiation7,8. Moreover, autophagy-deficient keratinocytes are more sensitive to oxidative stress and prone to UVB-induced oncogenic transformation9. Therefore, it is meaningful to further explore the molecular mechanism regulating autophagy under UVB stress.
BCL11A is a key transcriptional suppressor, which inhibits gene expression by recognizing and binding to the TGACCA promoter motif10. It has been well known as the negative transcriptional regulator of the differentiation and development of blood-lineage cells through repressing the expression of hemoglobin genes, such as HBZ and HBG11. This well-established finding has led to the development of the BCL11A gene-editing therapy, which the FDA has approved for the clinical treatment of sickle cell anemia and beta thalassemia10,12. In mammary epithelium, Khaled et al. found that BCL11A was required for epithelial stem cell and lineage-restricted progenitor cell proliferation13. BCL11A deficiency in mice exhibited epidermal permeability barrier defects, along with compromised skin differentiation and altered skin lipid composition14. Skin wound healing is accelerated in mice with epidermal-specific BCL11A KO, as evidenced by the enhanced cell proliferation, epithelialization, angiogenesis, and myofibroblast maturation15. However, the mechanism by which BCL11A participates in epidermal homeostasis under UVB stress remains unclear.
In this study, we discovered that genetic deletion of BCL11A enhances keratinocyte autophagy in a gene dose-dependent manner. Under UVB radiation, BCL11A deficiency promotes autophagosome formation and more intense autophagy flux, which contributes to increased cellular viability as well as reduced DNA damage and ROS production. Mechanically, we identified that the lipid synthesis-related gene, Acyl-CoA Synthetase Short-Chain family member 3 (ACSS3), is the target gene of BCL11A. More importantly, inhibition of autophagy by pharmacological treatment or BCL11A overexpression rescues the protective effect of BCL11A deficiency on HaCaT cells when exposed to UVB radiation. Together, our study reveals that BCL11A deficiency activates autophagy by transcriptionally increasing ACSS3 expression, which promotes cell survival and reduces damage substances under UVB stress. Our work provides mechanistic insights into a potential therapeutic target for relieving epidermic injury and diseases.
Results
BCL11A deficiency increases autophagic vacuoles in HaCaT cells
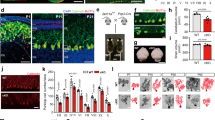
BCL11A knockout (KO) HaCaT cells were generated by CRISPR/Cas9-mediated gene editing to investigate the function of BCL11A in epidermis. Firstly, we designed a gRNA ending in NGG targeting the exon 2 of BCL11A, as shown in Fig. 1a, which was transfected into HaCaT cells by electroporation. Monoclonal heterozygotes (+/−, Het) or homozygotes (−/−, KO) BCL11A KO clones were confirmed by Sanger sequencing and western blot (Fig. 1a, b). More autophagic vacuoles, featured by double-membrane structure and encapsulated contents via transmission electron microscopy (TEM), were identified in both KO and Het cells compared to wild-type (WT) cells, with KO cells exhibiting the highest number (Fig. 1c). To validated the specificity of these autophagic vacuoles, we examined other potential confounding membrane structures, including transport vesicles, multivesicular bodies, and stress granules. No major changes in these structures were observed following BCL11A deletion, besides lipoproteins, which can be distinguished by electron density under TEM (Fig. S1a–h). To investigate whether the loss of BCL11A increased autophagic vacuoles under UVB radiation, WT, KO and Het cells were exposed to UVB radiation. When a mild dose of 30 J m−2 UVB radiation was conducted, all the HaCaT cell lines showed an expected increase of autophagic structures compared with the non-UVB-exposed cells. Further, the number of autophagic vacuoles was highest in KO cells, followed by that in Het cells (Fig. 1d). It indicates that BCL11A deletion, even the haploinsufficiency, is capable to enhance autophagy under UVB radiation. This change was still observed when those cells were subjected to a dose of 100 J m−2 UVB radiation, which caused a higher proportion of cell death (Fig. S1i). Meanwhile, we did not observe significant differences in the structure or contents of autophagic vacuoles among WT, Het and KO cells (Fig. 1c, d). Together, these results suggest that BCL11A deficiency contributes to a gene dose-dependent increase of autophagic vacuoles under UVB radiation or no stress conditions.
a Schematic of the BCL11A gene and the CRISPR/Cas9 guide RNA (sgRNA) target site. b Western Blot and quantitative grayscale analysis of BCL11A haploid knockout and diploid knockout. Grayscale normalized to GAPDH of WT. c, d TEM was employed to examine the morphology and quantify autophagic vacuole numbers in WT, Het and KO cells with or without UVB irradiation. The red arrow indicates the autophagic vacuoles. Scale bar: 1 µm. Data are represented as mean ± SD. (n = 8–9 fields per WT group, n = 11–13 field per HET or KO group). One-way ANOVA, ns no significance, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
BCL11A deficiency enhances UVB-induced autophagic flux
The accumulation of autophagic vacuoles can be caused by either the increased autophagy initiation or blockage of autophagosome degradation. To evaluate whether BCL11A blocks the autophagic flux, WT, Het and KO cells were infected with tandem fluorescent eRFP-EGFP-LC3 lentivirus. We counted the red puncta and the yellow puncta overlapped with RFP and GFP, which indicates autolysosomes and autophagosomes, respectively (Fig. 2a). Consistent with the TEM analysis, KO cells exhibited the highest number of autophagosomes, followed by Het cells, while WT cells showed the lowest levels (Fig. 2b). Under 30 J m−2 UVB exposure, both Het and KO cells displayed significantly higher number of autophagosomes than WT cells and their autophagic flux peaked at 12 h post-irradiation. Bafilomycin A1 (BafA1) was used to block autophagosome degradation, which resulted in the accumulation of autophagosomes and correspondingly decrease of autolysosomes in all the HaCaT cell lines (Fig. 2b, c). KO cells still maintained the highest numbers in autophagosomes, followed by Het and then WT cells, which indicates that BCL11A deficiency promotes rather than impairs autophagic flux. Consistently, compared to WT cells, the expression of LC3І/Ⅱ in Het and KO cells was increased under UVB stress and non-stress conditions, although basal p62 levels in BCL11A-deficient cells were not lower than those in WT cells (Fig. 2d–f). In parallel, BafA1 treatment induced greater accumulation of both p62 and LC3І/Ⅱ in KO cells compared to WT, suggesting a higher autophagy flux in BCL11A-deficient cells (Fig. S2a, b). When exposed to a higher UVB dose of 100 J m−2, BCL11A-deficient cells still significantly enhanced autophagic flux (Fig. S2c–h). To extend these observations in vivo, adeno-associated viral vector (AAV) mediated BCL11A knockdown in the epidermis of mice significantly activated autophagy, as determined by significantly enhanced LC3 and ATG5 expression under UVB exposure (Figs. 2g and S2i). In summary, BCL11A deficiency promotes keratinocyte autophagy flux under UVB stress.
a Representative images of stable eRFP-EGFP-LC3 reporter cells with or without BafA1 treatment. Scale bar: 1 µm. b, c Measurement of autophagosomes and autolysosomes in figure (a), respectively. (n = 12–13 fields per group). d Western Blot analysis of p62 and LC3І/Ⅱ expression in WT, KO and Het cells with or without UVB irradiation. e, f Quantitative grayscale analysis of p62 and LC3I/II protein expression. Grayscale normalized to GAPDH of unexposed WT cells. g The representative immunohistochemical images of LC3 and ATG5 expression in the dorsal skin tissues of BCL11A knockdown mice were obtained after UVB treatment (n = 5). Scale bar: 20 μm. Data are presented as mean ± SD. One-way ANOVA, ns no significance, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 (vs. WT), #P ≤ 0.05, ##P ≤ 0.01, ###P ≤ 0.001 (vs. DMSO).
BCL11A transcriptionally inhibited ACSS3
Transcriptome sequencing was performed to investigate the differences between KO and WT cells and screen the downstream target genes of BCL11A. We observed 195 upregulated genes and 201 downregulated genes, respectively (Fig. 3a). However, functional enrichment analysis of these differentially expressed genes (DEGs) showed no significant GO term directly related to autophagy (Fig. 3b). As a known transcriptional repressor, BCL11A deletion is expected to upregulate its target genes. We picked out autophagy-related genes, including ATG9B, LC3, WIPI1, Beclin1, ATG5, p62, and evaluated their transcript levels using qPCR. None of these genes were upregulated in KO cells compared with WT cells (Fig. 3c, d). Also, the upregulation of BCL11A had no effect on the mRNA level of ATG5 (Fig. S5). Considering that autophagosome formation has been confirmed to depend on the utilization of the existing membrane structures and the synthesis of lipids, especially short-chain fatty acids16,17, we analyzed the upregulated DEGs related to membrane donor and lipid synthesis (Fig. 3e). Among them, ACSS3 was uniquely upregulated at both mRNA and protein levels in BCL11A- deficient cells under both UVB exposed and non-exposed conditions (Fig. 3f, g). ACSS3 is a member of the acyl-CoA synthetase short-chain family and is responsible for producing precursors of lipid synthesis, such as acetyl-CoA and propionate-CoA18. To further explore the mechanism of BCL11A regulating the expression of ACSS3, we analyzed the promoter sequence of ACSS3 and identified a potential BCL11A-binding site (Fig. 3h). Next, chromatin immunoprecipitation (ChIP) was performed to verify the direct binding of BCL11A at the promoter of ACSS3. The results showed that BCL11A was significantly enriched at the promoter of ACSS3 in HaCaT cells with BCL11A overexpression, while the binding was markedly reduced following BCL11A downregulation (Fig. 3i, j). Taken together, these results suggested that BCL11A transcriptionally inhibits the expression of ACSS3.
a Scatter diagram of differentially expressed genes in WT and KO cells with RNA-seq analysis. b GO analysis at biological process level of differentially expressed genes in WT and KO cells. c, d The expression of autophagy-related genes was detected by qPCR. e Differential genes associated with autophagosome formation in WT and KO cells with RNA-seq analysis. f The mRNA levels of ACSS3 were detected by qPCR. g Western Blot and quantitative grayscale analysis of ACSS3 expression. Grayscale normalized to GAPDH of unexposed WT cells. h Analysis of binding site of BCL11A on ACSS3 promoter. i, j ChIP assay detected the enrichment of BCL11A at the promoter of ACSS3 in HaCaT cells with BCL11A overexpression or knockdown. Data are presented as mean ± SD. One-way ANOVA, ns no significance, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
BCL11A deficiency attenuates UVB-induced cell damage via promoting autophagy
To clarify the effect of BCL11A deficiency under UVB-induced stress, keratinocyte damage was evaluated. Under 30 J m−2 UVB radiation, KO and Het cells showed greater proliferative capacity and lower apoptosis rate compared to WT cells, indicating that both BCL11A homozygous and heterozygous deletion confer a protective effect (Fig. 4a–c). Compared to WT cells, BCL11A-deficient cells with higher autophagy flux may be more resistant to UVB radiation. Thus, 3-methyladenine (3-MA) was used to prevent the initiation of autophagy by disrupting autophagosome formation. Since previous experiments confirmed that short-term treatment with 3-MA could not permanently inhibit autophagy in cells, in the cloning formation experiment, we adopted a 24 h treatment with 3-MA (Fig. S4). Inhibition of autophagy significantly suppressed the proliferative capacity and upregulated the apoptosis rate in WT, Het and KO cells upon UVB irradiation (Fig. 4a–c). Notably, the survival rate of BCL11A-deficient cells showed no difference compared to WT and their enhanced colony-forming capacity was abolished upon 3-MA treatment, indicating that HaCaT cells with loss of BCL11A experience a greater dependence on autophagy (Fig. 4a, c). ROS accumulation and DNA damage are the major mechanisms underlying that UVB radiation leads to the decline in cell viability. BCL11A depletion in HaCaT cells decreased ROS accumulation and DNA damage, which was partially reversed by 3-MA treatment (Fig. 4d–f). Consistent with the effects observed at 30 J m−2 UVB, BCL11A deficiency demonstrated protective effects even under a higher-dose UVB exposure (Fig. S3a–f). In vivo, UVB-induced γH2A.X expression was significantly reduced in the mice with epidermal BCL11A knockdown, suggesting that downregulation of BCL11A reduced cell DNA damage (Fig. 4g). In summary, BCL11A deficiency, even heterozygous deletion, is sufficient to cope with UVB-induced ROS and DNA damage through autophagy.
a The survival rate of WT, KO, and Het cells with or without 3-MA assessed by CCK-8 assay at 24 h after 30 J m−2 UVB irradiation. b The flow cytometry quantified the apoptosis rate of WT, KO, and Het cells with or without 3-MA at 24 h 30 J m−2 UVB irradiation. c The representative images of colony formation of WT, KO, and Het cells with or without 3-MA combined with 30 J m−2 UVB irradiation. d, e IF assessed γH2A.X in WT, KO, and Het cells with or without 3-MA after 30 J m−2 UVB irradiation. The average number of γH2A.X was determined manually. Scale bar: 15 µm. f The ROS levels of WT, Het, and KO cells with or without 3-MA after 30 J m−2 UVB irradiation were assessed by fluorescence intensity of DCFH-DA. g The representative immunohistochemical images of γH2A.X in the dorsal skin tissues of BCL11A knockdown mice were obtained after UVB treatment (n = 5). Scale bars: 20 µm. Data are presented as mean ± SD. One-way ANOVA, ns no significance, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 (vs. WT), #P ≤ 0.05, ##P ≤ 0.01, ###P ≤ 0.001 (vs. non-treated with 3-MA).
BCL11A overexpression exacerbates UVB-induced cell damage by inhibiting autophagy
Finally, to further elucidate the role of BCL11A in cell damage and death under UVB stress conditions, we constructed in vivo and in vitro models with BCL11A upregulation. The AAV carrying Bcl11a successfully upregulated BCL11A expression and significantly inhibited autophagy in the epidermis of mouse, as evidenced by the downregulation of LC3 and ATG5 (Fig. 5a, b). In parallel, overexpression of BCL11A in HaCaT cells demonstrated the effective suppression of autophagy activity (Fig. 5c, d). When exposed to 30 J m−2 UVB radiation, the lower proliferation ability and higher apoptosis rate were observed in HaCaT cells with BCL11A overexpression, suggesting that the presence of BCL11A is detrimental to cell survival under UVB radiation (Fig. 5e, f). Additionally, BCL11A overexpression remarkably evaluated ROS accumulation and DNA damage in HaCaT cells both at 30 J m−2 UVB exposure and non-stress conditions (Fig. 5g, h). Consistently, the increase of UVB-induced DNA damage was obtained in the epidermis of mice with BCL11A overexpressed (Fig. 5i). This suggested that autophagy inhibition by BCL11A overexpression exacerbated UVB-induced DNA damage and ROS level. In summary, the presence of BCL11A enhances the sensitivity of epidermal cells to UVB radiation.
a, b Representative immunohistochemistry images show ATG5 and LC3 expression in mice skin with BCL11A overexpressing post-UVB irradiation. Scale: 20 μm. c, d Western blot and quantitative grayscale analysis of BCL11A, LC3І/Ⅱ and ATG5 was conducted in HaCaT cells overexpressing BCL11A. Grayscale normalized to β-action of NC cells. e CCK-8 assay evaluated the viability of HaCaT cells with BCL11A overexpression following UVB exposure. f Flow cytometry measured apoptosis rate of HaCaT cells with BCL11A overexpression after 24 h UVB exposure. g Immunofluorescence assessed γH2A.X in HaCaT cells with BCL11A overexpression and 30 J m−2 UVB exposure (n = 5). Scale: 20 μm. h ROS levels were detected after HaCaT overexpression of BCL11A combined with UVB irradiation. i The representative immunohistochemical images of γH2A.X in the dorsal skin tissues of BCL11A overexpression mice were obtained after UVB treatment (n = 5). Scale bars: 20 µm. Data are presented as mean ± SD. One-way ANOVA, ns no significance, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Discussion
In our study, BCL11A, as a key regulator of skin homeostasis, is identified as a transcriptional factor involved in the regulation of epidermal autophagy. It is evidenced by remarkable increase of autophagosomes and significant enhancement of autophagic flux in BCL11A-deficient cells. Given the established association between autophagy dysregulation and multiple skin pathological conditions, including psoriatic lesions, atopic dermatitis, and skin cancers, this finding is suggested to have important implications19,20. Epidermal autophagy deficiency impaired the resistance of keratinocytes to UV stress in mice2. Hyaluronic acid reduced the facial spots and sagging in human facial skin through autophagy activation21. Furthermore, autophagy upregulation decreased ROS levels, enhanced the preservation of NAD levels and cellular antioxidant ability to maintain cell survival22,23. Defective autophagy activation leads to impaired clearance of DNA double-strand breaks, severely disrupting cellular homeostasis24. However, aberrant autophagy activation or impaired autophagy would be equally detrimental for triggering cell death. MTOR-mediated suppression of autophagy flux inhibits apoptosis in HaCaT cells25. Autophagy inhibition via 3-MA or ATG5 silencing shows a protective function, as a result of the lysosomal damage and reduced cathepsin B levels upon UVA exposure26. In FADD-deficient cells, RIPK1-dependent autophagy was identified as a key contributor to UVB-induced cell death, primarily due to uncontrolled autophagy activation caused by Beclin1 downregulation and its failed interaction with anti-apoptotic proteins27. Here, different pharmacological inhibitors of autophagy were applied to show that BCL11A deficiency promotes a robust autophagy flux via increasing the autophagosome formation and enhances cell survival under UVB radiation, which supports the explanation of protective autophagy.
Notably, the expression of p62 did not show a significant decrease in BCL11A-deficient cells, which appears to be in contrast to eRFP-EGFP-LC3 puncta assays. This may indicate other autophagy receptors involved in BCL11A deficiency-mediated autophagy28. In addition, the autophagy inhibitor did not completely eliminate the protective effect of BCL11A deficiency against UVB stress, suggesting that loss of BCL11A may also confer UVB resistance through alternative mechanisms.
BCL11A is a master regulator of hematopoietic development, controlling B-cell differentiation and the fetal-to-adult hemoglobin switch by suppressing γ-globin10. Recently, the obvious epidermal permeability barrier defects were observed in newborn mice with BCL11A deletion, accompanied by dysregulated lipid metabolism and aberrant epidermal differentiation14. It emphasizes the critical role of BCL11A in skin morphogenesis. When exposed to external stimuli, such as mechanical stress, ablation of BCL11A in murine epidermal keratinocytes promotes rapid closure of excisional wounds both in a cell autonomous manner, likely via accelerating wound re-epithelialization and in a non-cell autonomous manner by enhancing angiogenesis15. It implies that BCL11A may govern mechanisms maintaining homeostasis in mature skin tissue, and its loss reactivates those protective responses when subjected to external stimulations. Here, we show that BCL11A downregulation in mouse epidermis enhances the resistance to UVB radiation. These results underscore the context-dependent role of BCL11A in maintaining epidermal homeostasis. Thus, future investigations are still required to unravel its function and related mechanism and employ the potential target in the prevention and treatment of skin diseases.
Many transcription factors have been verified to regulate autophagy. Most of them directly modulate autophagy-related genes or classical regulators, such as mTOR and PI3K29. Here, BCL11A is identified as a transcription factor regulating autophagy, but does not target the autophagy-related genes, although BCL11A deletion initiates autophagosome formation. Intriguingly, a lipid synthesis-related gene, ACSS3, was identified as the target gene of BCL11A. These align with the previously characterized mechanisms in yeast and mammalian cells that expansion of the phagophore membrane during autophagy is driven by localized de novo phospholipid biosynthesis17,30. Changes in lipid composition of autophagic membranes impair autophagosome completion31. ACSS3 results in the accumulation of lipid droplets and increases the production of succinyl-CoA, involved in lipid synthesis32. An increase in lipoproteins was observed in BCL11A-deficient cells, which may be related to ACSS3 upregulation (Fig. S1d). In this study, the mechanism that how ACSS3 regulates autophagy was not illustrated, but it would be of significant interest for further explored.
Given the critical regulatory role of BCL11A in skin homeostasis, elucidating its transcriptional regulatory mechanism is of great significance. Since transcription factors typically lack well-defined binding pockets, the development of specific small-molecule inhibitors faces significant challenges. To date, no specific small-molecule inhibitor targeting BCL11A has been approved. Based on previous studies in the expression regulation of globin, gene therapies targeting BCL11A have advanced from experimental stages to clinical applications in the treatment of sickle cell disease and β-thalassemia. For instance, the commercial gene therapy product Casgevy was approved in the UK and the US in 202333. Additionally, Yin M’s team has developed BCL11A-targeting nanobodies, offering a potential supplementary approach to gene therapy34. Tenofovir disoproxil fumarate and hydroxyurea have been shown to downregulate BCL11A expression, but the specific targets of its action are not clear35. Certain noncoding RNAs, including miR-486-3p and Lin28B, were also confirmed to inhibit BCL11A expression36. In this study, BCL11A knockdown activates autophagy to alleviate UVB-induced cell damage and promote cell survival. Thus, we suggest that BCL11A knockdown through topical administration of siRNA or other targeted drugs would alleviate the symptoms induced by sun exposure, such as erythema and sunburn, and decrease the risk of sun-exposed skin diseases, including psoriasis and atopic dermatitis. However, it cannot be ignored that the safety of genetic or transcriptional manipulations to suppress BCL11A expression may induce aberrant keratinocyte proliferation. Therefore, extensive in vivo and in vitro studies should be conducted to ensure the safest and most effective therapeutic strategies.
Methods
Cell culture and generation of BCL11A knockout HaCaT cells
The HaCaT cell line was purchased from Shanghai HonSun Biological Technology and maintained with Dulbecco’s modified Eagle medium (Gibco, USA), supplemented with 10% FBS (Every green, China) and 100 µg/mL streptomycin and 100 U/mL penicillin (Gibco, USA) at 37 °C in a humidified 5% CO2 atmosphere. The BCL11A-KO HaCaT cell line was generated using the CRISPR/Cas9 system. A single-guide RNA (sgRNA) targeting the second exon of BCL11A was designed with the following sequence: 5′-TGAACCAGACCACGGCCCGT-TGG-3′ (Cyagen, China).
Animals and UVB irradiation
All animal experiments were approved by the Institutional Animal Care and Use Committee of Southern Medical University (Approval No. SMUL2021194) and complied with the National Institutes of Health Guide for the Care and Use of Laboratory Animals. The male BALB/c mice aged 4–6 weeks were purchased from Guangdong Medical Laboratory Animal Center, randomly assigned to groups, and housed in standard laboratory conditions. After adaptive feeding, the mouse dorsal area (approximately 1 × 2 cm) was depilated and followed by a 24 h acclimation. AAV carrying Bcl11a coding sequence or shRNA targeting Bcl11a (Hanheng, China) was adjusted to a titer of 1.2 × 1012 vg per mL, with a dosage of 50 μL per mouse. The viral suspension was evenly applied to depilated skin, followed by dermal roller puncture to enhance viral transduction. Six weeks post-AAV infection, the dorsal skin was depilation. The next day, mice received intraperitoneal anesthesia and were irradiated with Minimal Erythema Dose, which corresponds to 600 mJ cm−2. Mice were euthanized by cervical dislocation after administering anesthesia at 6 h post-irradiation, and irradiated skin samples were collected. The researchers were blinded during the process of evaluating outcomes.
UVB irradiation of cells
UV irradiation was administered using a Philips lamp at doses of 30 J m−2 and 100 J m−2. Before irradiation, remove the cell culture medium and wash twice with PBS, then add an appropriate volume of PBS according to the culture dish specifications for irradiation. To ensure HaCaT cells in different experiments were subjected to a consistent dose of UVB radiation, the minimal PBS volume required to fully cover the cell layer is calculated and applied. For a 60 mm culture dish, the required PBS volume is calculated to be 1.8 mL based on its base area and a liquid coverage height of approximately 1 mm. For other plate/dish formats, PBS volumes were adjusted proportionally according to the bottom surface area.
Cell transfection
The cells were seeded in 6 cm culture dish and cultured to 50–60% confluence over 24 h. Transfection was performed according to the instructions of Lipofectamine™ 2000 (Invitrogen, USA) transfection reagent, with 2 μg of BCL11A plasmid or empty vector plasmid. After 6 h, the medium was replaced.
eRFP-EGFP-LC3 lentivirus infection and LC3 puncta evaluation
The cells were seeded in a 3.5 cm culture dish at a density of 2 × 105 cells per well and cultured for 24 h. The culture medium was replaced with fresh medium containing the calculated amount of the eRFP-EGFP-LC3 lentivirus (Hanheng, China) (multiplicity of infection = 100, titer = 1 × 108 TU per mL). Polybrene (8 μg per mL) was added to enhance transduction efficiency. The positive cells were selected using 0.5 μg per mL puromycin for at least 3 passages until all cells were stably positive.
Cells infected with eRFP-EGFP-LC3 lentivirus were seeded into 24-well plates and cultured for 24 h. They were preincubated with 100 nM BafA1 for 2 h and then exposed to UVB irradiation. At 6, 12, and 24 h post irradiation, the cells were fixed with 3.7% paraformaldehyde and were imaged on a confocal microscope (Carl Zeiss, Germany). Autophagosomes show up as yellow puncta (the colocalization of red signal and green signal) and autolysosomes as red puncta. Fluorescent puncta were quantified by manually counting ≥10 cells (63× objective).
Colony formation
Cells were seeded in 12-well plates at a density of 500 cells per well and cultured for 24 h. After UVB irradiation, PBS was rapidly removed, and a fresh medium containing 5 mM 3-MA (TargetMol, T1879, China) was supplemented for 24 h. Subsequently, the medium was replaced with fresh culture medium, and the cells were maintained with medium changes every 5 days until colony formation. Cells were fixed with 3.7% paraformaldehyde and then stained with 0.1% crystal violet solution and air-dried for visualization.
CCK-8
Cells were seeded in 96-well plates at a density of 4000 cells per well. Following pretreatment with 5 mM 3-MA (TargetMol T1879, China) for 2 h, cells were exposed to UVB irradiation. At 0 and 24 h after exposure, add 10 µL of CCK-8 (Biosharp, China) per well, and detected its absorbance at 450 nm with a microplate reader.
Apoptosis detection
The cells were seeded in 6 cm culture dishes and cultured for 24 h. Following pretreatment with 5 mM 3-MA (TargetMol T1879, China) for 2 h, the cells were exposed to UVB irradiation. After 24 h, apoptosis was detected by TransDetect Annexin V-FITC/PI apoptosis detection kit (Lianke Biotech Co., China) according to the instructions and quantified by flow cytometer (BD FACSARIA, Grand Island, NY, USA).
ROS detection
Cells were seeded in 96-well plates at 4000 cells per well and cultured for 24 h. Following pretreatment with 5 mM 3-MA (TargetMol, China) for 2 h, the cells were exposed to UVB irradiation. Then, 2,7-dichlorodihydrofluorescein diacetate (Beyotime, Shanghai, China) was used to evaluate cellular ROS levels according to the instructions and quantified via a fluorescence microplate reader.
Western blot
The cells were seeded in 6 cm culture dishes for 24 h, and then treated with UVB irradiation. Following treatment, cells were harvested at 6 and 12 h post-irradiation for protein extraction. Total protein concentrations were determined using a BCA quantification kit (Beyotime, China). Equal amounts of denatured proteins were separated by SDS-PAGE gel, followed by electrophoretic transfer onto 0.45 μm PVDF membranes. Membranes were blocked with 5% skim milk in TBST for 1 h at room temperature, then incubated overnight at 4 °C with primary antibodies. The primary antibodies used were as follows: BCL11A (1:2000, CST 75432, USA), ATG5 (1:2000, CST 12994, USA), Beclin1 (1:2000, CST 3495, USA), LC3A/B (1:2000, CST 12741, USA), SQSTM1/p62 (1:5000, Proteintech 29503-1-AP, China), ACSS3 (1:2000, Proteintech 16204-1-AP, China), and GAPDH (1:5000, Santa 137179, USA). Following TBST washes, membranes were incubated with HRP-conjugated goat anti-mouse IgG(H + L) (1:4000, Beyotome A0216, China) or HRP-conjugated goat anti-rabbit IgG(H + L) (1:4000, Beyotome A0218, China) at room temperature. Protein bands were visualized using enhanced chemiluminescence reagents and captured with a Tanon imaging system (Tanon, China). The quantification was performed using ImageJ software.
Immunofluorescence (IF)
Cells were plated in 24-well plates at a density of 5 × 104 cells per well and cultured for 24 h. Cells were fixed in 3.7% formaldehyde and incubated with a blocking solution for 30 min at room temperature. Cells were probed with the primary antibody γH2A.X (1:1000, CST 9781, USA) overnight at 4 °C. On the subsequent day, cells were washed twice with PBS and incubated with a corresponding secondary antibody for 2 h at room temperature. Subsequently, plates were mounted using ProLong Gold Antifade Mountant with DAPI. Fluorescent images were acquired via confocal microscopy (Zeiss, Germany).
Immunohistochemistry (IHC)
Paraffin-embedded tissue sections were first dewaxed in xylene and rehydrated through an ethanol gradient, followed by antigen retrieval using citrate buffer. After blocking with 5% serum, the sections were sequentially incubated with primary and secondary antibodies. Nuclei were counterstained with hematoxylin before mounting for microscopic observation. The primary antibodies used were as follows: BCL11A (1:2000, CST 75432, USA), LC3A/B (1:2000, CST 12741, USA), ATG5 (1:2000, CST 12994, USA), and γH2A.X (1:1000; CST 9718, USA). Imaging was performed on an inverted microscope (Zeiss, Germany).
Transmission electron microscopy
The cells were seeded in 6-well plates and cultured for 24 h. After 6 h of different doses of UVB irradiation, cells were harvested using 0.25% trypsin-EDTA-free solution (Biosharp, China), washed twice with PBS, and pelleted by centrifugation at 500 × g for 5 min. Cell pellets were fixed overnight at 4 °C in 2.5% glutaraldehyde (Nacalai Tesque, Japan). Ultra-thin sections were observed by TEM. Statistical analysis was performed by manually counting the number of autophagosomes in eight random fields of view.
RNA sequencing and analysis
High-throughput RNA sequencing was performed on WT and KO cell samples utilizing the Illumina HiSeq platform (Genergy Biotechnology, Shanghai). Post-sequencing, alignment outcomes were subjected to screening and identification via DESeq2 software, applying stringent criteria: |log2FC| ≥ 1 and P < 0.05. After differential expression analysis, GO analysis was conducted to ascertain the biological relevance of the identified genes.
Quantitative real-time PCR (qPCR)
Total RNA was extracted from cells using TRIzol reagent (Invitrogen, USA) following the manufacturer’s protocol. cDNA was synthesized with the Evo M-MLV Reverse Transcription Kit (Accurate, China), and target gene expression was quantified by RT-qPCR using SYBR Green Master Mix (TransGen, China). GAPDH served as the internal control, with relative expression calculated via the 2ΔΔCT method. All primer sequences are provided in Supplementary Table 1.
Chromatin immunoprecipitation (ChIP)
ChIP was performed using a commercial kit (Millipore 17-371, Germany), according to the manufacturer’s protocol. Briefly, cells were crosslinked with 1% formaldehyde for 10 min, and chromatin was sonicated to generate DNA fragments ranging from 200 to 1000 bp. The sheared crosslinked chromatins were incubated overnight with 5 μg of anti-BCL11A antibody (ab19487) or IgG antibody. Then, the antibody-antigen-DNA complex was collected by Protein G Agarose. Finally, the purified DNA was quantified by qPCR with SYBR Green Master Mix using ACSS3-specific primers, and the results were normalized to input DNA. All primer sequences are provided in Supplementary Table 1.
Statistics and reproducibility
Statistical analysis was performed using SPSS 25.0 statistical software. Data were expressed as mean ± standard deviation. Student t test was used to compare the data of the two groups, and one-way analysis of variance (ANOVA) was used to compare the data of the three groups and above. P < 0.05 was considered statistically significant. RNA-seq was performed as one independent experiment with three technical replicates, whereas all other experiments included three independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All source data underlying the graphs and charts presented in the main figures are included in Supplementary Data 1. All the full, uncropped blots have been provided in Supplementary Figs. 6–8. The RNA-seq data are deposited in the NCBI Gene https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE306030.
References
Kitada, M. & Koya, D. Autophagy in metabolic disease and ageing. Nat. Rev. Endocrinol. 17, 647–661 (2021).
Song, X. et al. Autophagy deficient keratinocytes display increased DNA damage, senescence and aberrant lipid composition after oxidative stress in vitro and in vivo. Redox Biol. 11, 219–230 (2017).
Wen, C. et al. Gastrodin ameliorates oxidative stress-induced RPE damage by facilitating autophagy and phagocytosis through PPARα-TFEB/CD36 signal pathway. Free Radic. Biol. Med. S0891-5849(24)00616–6 (2024).
Galanopoulou, O. et al. Endonucleosis mediates internalization of cytoplasm into the nucleus. Nat. Commun. 15, 5843 (2024).
Sun, S. et al. Inactivation of ribosomal protein S27-like impairs DNA interstrand cross-link repair by destabilization of FANCD2 and FANCI. Cell Death Dis. 11, 852 (2020).
Xu, L. et al. Suppression of PERK/eIF2α/CHOP pathway enhances oridonin-induced apoptosis by inhibiting autophagy in small-cell lung cancer cells. Biomed. Pharmacother. 175, 116684 (2024).
Chen, R.-J. et al. The roles of autophagy and the inflammasome during environmental stress-triggered skin inflammation. Int. J. Mol. Sci. 17, 2063 (2016).
Wang, Z.-Y. et al. HSP27 protects skin from ultraviolet B -induced photodamage by regulating autophagy and reactive oxygen species production. Front. Cell Dev. Biol. 10, 852244 (2022).
Qiang, L. et al. Autophagy gene ATG7 regulates ultraviolet radiation-induced inflammation and skin tumorigenesis. Autophagy 13, 2086–2103 (2017).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442 (2018).
Sankaran, V. G. et al. Transcriptional silencing of fetal hemoglobin by BCL11A: transcriptional silencing of fetal hemoglobin. Ann. N. Y. Acad. Sci. 1202, 64–68 (2010).
Frangoul, H. et al. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Khaled, W. T. et al. BCL11A is a triple-negative breast cancer gene with critical functions in stem and progenitor cells. Nat. Commun. 6, 5987 (2015).
Li, S. et al. Transcription factor CTIP1/ BCL11A regulates epidermal differentiation and lipid metabolism during skin development. Sci. Rep. 7, 13427 (2017).
Bhattacharya, N. et al. Selective ablation of BCL11A in epidermal keratinocytes alters skin homeostasis and accelerates excisional wound healing in vivo. Cells 11, 2106 (2022).
Xin, Z. et al. Neuroprotective effect of a multistrain probiotic mixture in SOD1G93A mice by reducing SOD1 aggregation and targeting the microbiota-gut-brain axis. Mol. Neurobiol. 61, 10051–10071 (2024).
Schütter, M. et al. Local fatty acid channeling into phospholipid synthesis drives phagophore expansion during autophagy. Cell 180, 135–149.e14 (2020).
Yoshimura, Y. et al. Molecular cloning of rat acss3 and characterization of mammalian propionyl-CoA synthetase in the liver mitochondrial matrix. J. Biochem.161, 279–289 (2017).
Klapan, K. et al. Autophagy and skin diseases. Front. Pharmacol. 13, 844756 (2022).
Wu, X. et al. Exploring the role of autophagy in psoriasis pathogenesis: insights into sustained inflammation and dysfunctional keratinocyte differentiation. Int. Immunopharmacol. 135, 112244 (2024).
Ito, S. et al. 2 kDa hyaluronic acid reduces blemishes and sagging in human facial skin. Clin. Cosmet. Investig. Dermatol. 18, 805–816 (2025).
Kataura, T. et al. Autophagy promotes cell survival by maintaining NAD levels. Dev. Cell 57, 2584–2598.e11 (2022).
Yang, N. et al. A snake cathelicidin enhances transcription factor EB-mediated autophagy and alleviates ROS-induced pyroptosis after ischaemia-reperfusion injury of island skin flaps. Br. J. Pharmacol. 181, 1068–1090 (2024).
Sun, F. et al. Increased DNA damage in full-grown oocytes is correlated with diminished autophagy activation. Nat. Commun. 15, 9463 (2024).
Han, S. et al. Seawater pearl hydrolysate inhibits photoaging via decreasing oxidative stress, autophagy and apoptosis of ultraviolet B-induced human skin keratinocytes. J. Cosmet. Dermatol. 23, 256–270 (2024).
Wu, A. Y.-T. et al. Spatiotemporal roles of AMPK in PARP-1- and autophagy-dependent retinal pigment epithelial cell death caused by UVA. J. Biomed. Sci. 30, 91 (2023).
Antunovic, M. et al. FADD-deficient mouse embryonic fibroblasts undergo RIPK1-dependent apoptosis and autophagy after NB-UVB irradiation. J. Photochem. Photobiol. B 194, 32–45 (2019).
Vargas, J. N. S. et al. The mechanisms and roles of selective autophagy in mammals. Nat. Rev. Mol. Cell Biol. 24, 167–185 (2023).
Kim, Y. et al. The interplay of microRNAs and transcription factors in autophagy regulation in nonalcoholic fatty liver disease. Exp. Mol. Med. 53, 548 (2021).
Rowland, L. A. et al. De novo lipogenesis fuels adipocyte autophagosome and lysosome membrane dynamics. Nat. Commun. 14, 1362 (2023).
Polyansky, A. et al. Phospholipid imbalance impairs autophagosome completion. EMBO J. 41, e110771 (2022).
Wang, L. et al. ACSS3 regulates the metabolic homeostasis of epithelial cells and alleviates pulmonary fibrosis. Biochim. Biophys. Acta Mol. Basis Dis. 1870, 166960 (2024).
Song, X. et al. Gene therapy and gene editing strategies in inherited blood disorders. J. Genet. Genom. 51, 1162–1172 (2024).
Yin, M. et al. Evolution of nanobodies specific for BCL11A. Proc. Natl. Acad. Sci. USA 120, e2218959120 (2023).
Zhang, H. et al. Role of B-cell lymphoma/leukemia 11A in normal and malignant hematopoiesis. Biology 14, 26 (2025).
Saki, N. et al. MicroRNA expression in β-thalassemia and sickle cell disease: a role in the induction of fetal hemoglobin. Cell J. 17, 583–592 (2016).
Acknowledgements
This work was supported by the grants from the National Natural Science Foundation of China (Grant Numbers 82103785 and 82273582) and GuangDong Basic and Applied Basic Research Foundation (2020A1515110543).
Author information
Authors and Affiliations
Contributions
X. D. wrote the manuscript, partly designed research and performed experiments. H. L. performed experiments and collected the data. X. D., Z. F., X. W., and X. N. helped to perform experiments. Z. G., P. L., and Y. D. contributed to animal experiments. Z. D., Y. W., and M. Z. contributed to conceived and designed research, funding acquisition, project administration, supervision, and review. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Shuyu Zhang and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary handling editors: Giulia Bertolin and Manuel Breuer.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, X., Liu, H., Wang, X. et al. BCL11A deficiency protects epidermis from UVB-induced damage through promotion of autophagy. Commun Biol 8, 1417 (2025). https://doi.org/10.1038/s42003-025-08814-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08814-1