Abstract
Cryo-imaging in cellular biology provides the means to visualize the cellular interior at close-to-native conditions. A cornerstone in the field has been cryo-correlative light and electron microscopy (cryo-CLEM), with cryo-visible light fluorescent microscopy (cryo-VLFM) providing the specificity by tagging macromolecules or structures and cryo-electron tomography (cryo-ET) for the structural details at molecular level. The large resolution gap between these techniques, however, is limiting this correlative workflow as cryo-ET targets are often smaller than the resolution limit of cryo-VLFM. Here we introduce cryo-soft X-ray tomography (cryo-SXT) as an intermediate step that can compensate for the partial view caused by the lost cellular material due to FIB-milling and limited resolution of cryo-VLFM by providing invaluable cellular context information in 3D to the cryo-ET dataset within an integrated workflow. This work shows that X-ray and electron imaging are not mutually exclusive, creating opportunities for further correlative imaging strategies.
Similar content being viewed by others
Introduction
Correlative multimodal imaging is a powerful tool for the study of complex systems, such as those found in biology1,2,3,4,5,6. Correlating data obtained on the same sample by different modalities gives complementary information, significantly promoting a more comprehensive interpretation of the studied cellular process. Major challenges for correlative multimodal imaging have been the compatibility of sample preparation and support, the drastic increase in complexity as each step adds damage risk that can hinder subsequent steps, and the need to keep the workflow within a radiation dose limit, depending on the required resolution.
In cellular biology, correlative light and electron microscopy (CLEM) has been performed for many years at room temperature (RT)7. Recently, with the emergence of direct electron detectors and advanced image processing algorithms, cryo-ET has gained momentum, and thus cryo-CLEM has set off in many laboratories8,9. It is now possible to generate 3D cryo-ET datasets at molecular resolution in near-native conditions. However, to generate high resolution (i.e., sub-nanometer) data, sample thickness (<200 nm) and radiation dose (<150 e/A2 ~ 550 MGy10,11) need to be controlled12. Thus, for most unicellular or complex biological samples thinning is required, nowadays mostly done by cryo-focused Ion Beam (FIB) milling13. Cryo-VLFM is employed for targeting sites of interest, although inefficiently due to the resolution limitation and insufficient spatial accuracy for precise localization of the fluorescently labelled structures of interest, especially in the axial dimension. Cryo-VLFM resolution is restricted by the lack of dedicated high-numerical aperture cryo-immersion objectives, which limits currently the achievable resolution to 350 nm laterally and to more than 1 µm in the axial direction. When aiming for small (<200 nm) structures, cryo-lamellae are often milled at the wrong depth, missing the target partially or completely.
Here we introduce cryo-SXT14,15 as an intermediate step, which can provide precious information regarding the 3D cellular context of up to 10 µm thick samples down to 30 nm resolution (at full dose)16, otherwise lost during cryo-FIB milling. With both lateral and axial resolution being higher than any existing cryo-VLFM modality, cryo-SXT provides the accuracy needed to optimize lamellae milling17. In cryo-SXT at the so-called “water window” energy region (284–543 eV), oxygen-based structures absorb little, while carbon-based structures absorb strongly the incoming photons. This provides a natural contrast of the cellular ultrastructure without additional staining. Moreover, intact cells can be imaged as a whole, without the need for sectioning. Cryo-SXT has been used in a correlative mode with cryo-VFLM (correlated light and X-ray microscopy, cryo-CLXM)4,18,19,20,21,22,23,24, and more recently in multimodal experimental setups also alongside cryo-CLEM5,25 or others3. Utilizing the three modalities together on the same cell within an integrated workflow (cryo correlative light, X-ray, and electron microscopy: cryo-CLXEM), however, has to our knowledge never been reported yet.
Results
Theoretical proof of the feasibility of the proposed workflow
Sample preparation is highly compatible for the three modalities as they use the same support and cryo-fixation approach. However, as radiation damage is induced by both X-rays26,27 and electrons28,29, their combination requires special care and evaluation of the resolution-dose tradeoff for imaging the sample. In cryo-soft X-ray imaging at Mistral (ALBA)30, a photodiode is used to measure the photon flux reaching the sample at a given energy, and the percentage of photons absorbed is determined by the difference between the transmitted signal with and without the sample. In EM, what is commonly referred to as the electron dose describes the electron fluence - the number of electrons per unit surface to which the sample is exposed (expressed in e⁻/Ų). This value can be translated into an equivalent radiation dose, via a voltage-dependent conversion factor (see “Methods” section) to be represented in MGy11. While there are various types of interactions31,32, soft X-rays mainly interact through photoelectric absorption, although a small portion is elastically scattered (Rayleigh Scattering), while electrons mainly interact through inelastic (Compton) scattering. In both cases, the absorbed dose depends on the composition and thickness of the sample, among other parameters related to the incoming beam (see “Methods” section). Despite the different interactions, the visible radiation effects are similar11, and a previous exposure can hasten the onset thereof. Hence, it can be assumed that a sample has a certain “dose-budget” that can be spent before visible damage to the ultrastructure becomes evident, depending on the desired resolution, and well before sublimation in the form of bubbling due to radiolysis of water molecules occurs33. Notably, the lack of visible damage does not imply its absence; molecular bonds will be broken at much earlier timepoints34,35. In our calculations, considering a goal of 5 nm resolution or better with cryo-ET36, we determined a total dose for cryo-SXT of a few tens of MGy27,37, corresponding approximately to 10–15% of the maximum dose-budget for vitrified ice sublimation. This was then considered for the subsequent cryo-ET acquisition (see “Methods” section).
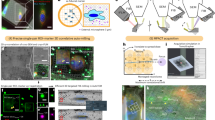
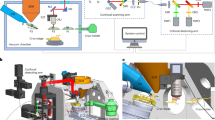
To briefly summarize the experimental procedure (Fig. 1), adherent cells were grown on gold EM grids and plunge-frozen in liquid ethane. Fluorescence labeling of the nucleus, mitochondria, and acidic compartments was performed shortly before plunging. Vitrified samples were then imaged in a cryo-confocal microscope to, firstly, make a 2D map of the grid to assess the quality in terms of grid damage, ice quality, and cell density, and secondly, to identify and image suitable cells. Samples were then sent to the cryo-SXT Mistral beamline (ALBA Synchrotron, Spain) and loaded into the microscope. With their in-line visible light microscope (VLM) a 2D map is acquired and correlated with the confocal map to identify previously imaged cells. A single cryo-SXT tomogram per cell was acquired of up to 10 cells per grid, keeping tight control on the dose, reducing the photon flux when required. It is important to note that balance must be found between exposure and resulting contrast (i.e., SNR) to avoid losing the contextual information provided by cryo-SXT. Post-acquisition, samples were shipped back to Institut Pasteur and loaded into the cryo-FIB scanning electron microscope (cryo-FIB-SEM). Imaged cells were identified, and ~300 nm thin lamellae were cut, aiming for the area previously imaged by cryo-SXT. The samples were then loaded into a cryo-transmission electron microscope (cryo-TEM), and tilt series of features of interest were acquired (see “Methods” section and Supplementary Fig. 1 for more details).
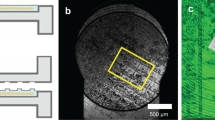
Cells are seeded on the grids and plunge frozen in liquid ethane (E, a). The samples are imaged by cryo-confocal microscopy (b), cells of interest are identified and z-stacks are acquired. Samples are then shipped to the synchrotron to be loaded in dedicated sample holders and cryo-SXT unidirectional tilt series are acquired (c). Samples are then loaded into a cryo-FIB-SEM, in which a focused ion beam (FIB) is used to trim away from the previously imaged cells of interest (d). Using a cryo-TEM, bidirectional tilt series are acquired in the prepared lamellae (e). Finally, all the data are reconstructed and correlated before further analysis (f).
Targeted resolution is still achievable in both modalities despite the reduced dose during acquistion
Figure 2 shows the full correlation from the 3 modalities in the same area from one of the imaged grids (see Supplementary Fig. 2 for additional examples from two other grids). The fluorescent 1 µm microspheres can easily be identified in the cryo-confocal and cryo-SXT datasets. Using this information, the area of the cryo-SXT tomogram (in X and Y) could be localized within the cryo-FIB-SEM. At the cryo-TEM, medium magnification maps at 8700x (corresponding to an effective pixelsize of 14.77 Å) of the lamellae were acquired, and sites for the acquisition of tilt series were chosen based on distinct features, like the two shown examples: mitochondria and multivesicular bodies, which can be easily identified in the cryo-SXT tomograms, were also labelled by fluorescent markers. Precise correlations were performed afterwards. An estimate of the achieved spatial resolution (in cryo-SXT commonly referred to as half-pitch resolution) was made by Fourier shell correlation even/odd (cut-off at 0.2536) and were found to be ~55 and ~70 nm for the cryo-SXT volumes at the calculated total doses of ~100 and ~76 MGy (with 25 and 40 nm Fresnel Zone Plates lenses, respectively), and ~5 nm for the cryo-ET volume (see Supplementary Fig. 3). In addition to the tilt series, we acquired still-images to measure the allowed cumulative dose on the target. To this aim we took a series of 70 images at 0° tilt, resulting in a total (exposure) dose of up to ~200 e/A2 (~740 MGy) on the sample (see Supplementary Fig. 4), which did not show any apparent damage to visible membrane structures.
A shows the Z-projection of the confocal stack acquired at the cryo-Stellaris 8 confocal microscope. Blue – Hoechst, Green – Mitotracker Green, Red – Fluorescent Microspheres (1 µm), Yellow – Lysotracker deep red. B shows the 2D cryo-SXT mosaic of the same grid square. The yellow square in A and B are shown overlayed in (C). The red square corresponds to the cryo-SXT tomogram area of which a single slice is shown in (D) (see Supplementary Video 1). The pink square corresponds to the lamella position, with the arrow indicating the direction (E). The green squares highlight the area of the cryo-ET acquisitions. In D’ and D’’ single slices from the reconstructed cryo-SXT tomogram, showing a multivesicular body (see Supplementary Video 2) and part of a mitochondrion, respectively, are shown, which can also be observed by cryo-ET in (E’, E’’). Scalebars: A–C: 10 µm; D: 2 µm; D’, D’’: 1 µm; E: 1 µm; E’, E’’: 200 nm.
Discussion
Herein, we have developed an integrated cryo-CLXEM workflow demonstrating that acquisition of cryo-electron tomograms for the visualization of eukaryotic cells is possible after cryo-confocal and cryo-SXT imaging. Although the throughput remains limited in our study—with approximately 15% of the cryo-SXT imaged samples having more than 50% overlap with the prepared FIB-lamellae, we believe that the potential of this cryo-correlative multiscale workflow is evident. Furthermore, it can be adapted to other X-ray-based cryo-correlative pipelines, aiming at imaging vitrified cells, as in the current work, we show that X-ray and electron-based imaging are not mutually exclusive if the dose is kept under control.
The estimated resolutions are in an acceptable range considering the reduced dose and SNR used in this proof-of-concept study. For studying rare events that are inherently difficult to locate, cryo-CLXEM will allow precise targeting, in addition to securing the cellular context for cryo-ET data interpretation. Further investigations using sub-tomogram averaging would need to be performed on known macromolecules (e.g., in situ ribosomes) to evaluate the ultimate resolution achievable. Note that one of the major bottlenecks is the targeting at the cryo-FIB-SEM system, which directly correlates to the throughput. The small field of view (FoV) contained few to no correlative markers, which, in addition, were mostly indistinguishable from the accumulated ice-crystal contamination from the FIB-angle point of view. With future technological developments, we expect this to improve. Meanwhile, intermediate solutions, like using more distinguishable correlative fiducial markers and/or using correlation software like 3DCT38 could enhance the workflow. Moreover, we suggest the development of software tools to allow calculating in advance the total dose a specific sample could stand for a given spatial resolution, following what other scientific communities have done, e.g., the protein crystallography field39. As a last consideration, accessibility to high resolution X-ray techniques at synchrotron-based facilities are open to users free of charge through public calls40. Additionally, laboratory X-ray microscopes are now commercially available, which should allow a larger community to take advantage of these techniques.
Overall, the addition of cryo-SXT provides a wealth of complementary information to classical cryo-ET datasets especially in the case of vitrified cells. The larger FoV and penetration depth, which allow visualizing the local cellular environment, can be crucial for the interpretation of structures within the lamellae. Furthermore, cryo-SXT has been widely used to describe the effect of drugs3,21,41,42 or the reorganization of organelles4,25,43,44, among others, and with our proposed workflow, the affected areas can now be targeted for high-resolution data acquisition, potentially beyond the here reported 5 nm. Additionally, with scientists experiencing the need to go to more complex models, like tissue or organoids, cryo X-ray imaging can provide a real benefit over cryo-VLM when it comes to targeting areas of interest45,46,47,48. The developments of multiscale CLXEM workflows covering macromolecular to cellular dimensions and cellular to tissular dimensions are now at reach.
Methods
Sample preparation
Vero E6 cells (Vero 76, clone E6, Vero E6, ATCC® CRL-1586TM, simian kidney cell line) were obtained from ATCC (USA) and were maintained in DMEM GlutaMax medium supplemented with 10% FBS and 1% Pen/Strep. 150.000 cells were seeded in a 35 mm ibidi culture dish, on 200 Mesh 1/4 holey carbon gold EM-grids from Quantifoil. Hoechst nuclear dye, Mitotracker green, and Lysotracker deep-red (all from Thermo Fischer Scientific) were added shortly before plunge-freezing, considering the manufacturers' recommended concentration and incubation time, washed with PBS, and returned to complete medium until freezing. The grids were frozen using the Leica EMGP plunge-freezer, keeping the humidity in the chamber above 70% and adding 3 µl of 1 µm fluorescent microsphere (red FluoSpheres™, Invitrogen) to the grids before blotting. Grids were blotted for 5 s, plunged into liquid ethane kept at −180 °C, clipped into the specialized Thermo Fischer autogrids for cryo-FIB-milling, and stored in liquid nitrogen until further use.
Cryo-confocal imaging
Samples were loaded into the cryo-Stellaris 8 confocal system (Leica Microsystems) equipped with a HC PL APO 50x/0.90 cryo objective, a white light laser (WLL), and three HyD detectors. A first 2D widefield map was created of the entire grid to assess the quality of the grid in terms of damage, number of cells in suitable positions, and quality of the ice. Then, cryo-confocal stacks were acquired of up to 10 cells of interest per grid, with the pinhole set to 1 Au, the zoom set to 1.7x to have the FOV be one grid square only, and the z-step was determined using the system-optimization feature.
Cryo-SXT imaging
Samples were shipped to Mistral Beamline at the Alba Synchrotron in Spain using a dry shipper (CX100). There they were loaded into their dedicated dual tilt sample holders49 and inserted into the cryo-SXT microscope30. The magnification was chosen to give an effective pixel size of 13 nm for the 40 nm Fresnel Zone Plate (FZP) and 11 nm for the 25 nm FZP. First, the photodiode was inserted to measure the real flux at the sample position. Then, the sample was loaded, and a brightfield and fluorescence map were acquired using the in-built visible light microscope. Then, using the 2D fluorescence map from the cryo-confocal system, the cells that were previously imaged were identified. Only one X-ray tomogram per cell was acquired, as the X-ray radiation exposure extends beyond the FOV. The dose on the sample was controlled by reducing the gap of the entry and exit slit of the monochromator to about half of the usual dose used to obtain 30 nm resolution16, or even lower. X-ray images were acquired every 1 degree, up to ±65 degrees, if possible, for a total of up to 131 images. The samples were then unloaded and shipped back to Institut Pasteur, Paris, France.
Cryo-lamellae preparation
For the cryo-Focused Ion Beam (FIB) milling, samples were loaded into the Thermo Fischer Aquilos 2 system. Using the Maps software, the acquired SEM overview map of the grid was aligned with the previously acquired visible light microscopy maps to identify the areas imaged by cryo-confocal and cryo-SXT. Using the 2D-X-ray mosaic images, the exact area within the grid square was identified and marked. Before starting the milling process an organometallic platinum layer was deposited on the grids for 75 s using the gas injection system. The Auto-TEM software was used for automatic lamellae cutting, from 1 nA for initial rough milling, going down gradually until 30 pA for the final fine polishing step, aiming for a final thickness of 300 nm. After completion, the lamellae were inspected manually and polished further if any curtaining was visible.
Cryo-EM
Samples were loaded into the 300 kV cryo-TEM Titan Krios, equipped with a Falcon 4i direct electron detector (Thermo Fisher Scientific) and a Selectris X energy filter (Thermo Fisher Scientific), using the autoloader system. Datasets were collected using SerialEM (Version 4.1.4)50 with the energy filter at 10 eV (zero loss), objective aperture of 100 \({{\upmu }}{{\rm{m}}}\) and a pixel size of 3.104 Å. The exposure time was chosen to have a maximum of 110 e/ Å2 for the full tilt series. The tilt series was acquired utilizing a dose-symmetric scheme, acquiring an image every 3°, from 70° to −50°, starting at 10°, for a total of up to 41 images at defocus set to -2 \({{\rm{\mu }}}{{\rm{m}}}\).
Data processing
Cryo-ET frames were aligned using MotionCor 2 within the Scipion package51. Both cryo-SXT and cryo-ET tilt series were aligned with AreTomo52 and reconstructed using either the SIRT algorithm of Tomo3D53 for the cryo-SXT datasets and using the Filtered Back Projection with the SIRT-like filter of IMOD54 for the cryo-ET datasets. Initial correlation of the data was done with scNodes55, and for a finer correlation we utilized the ecCLEM plugin56 in ICY57.
Resolution estimation
The resolution was estimated by Fourier shell correlation (FCS) of the split tilt series, as reported previously (Carrascosa et al.36). For the cryo-SXT datasets, as no frames were collected, the tilt series were divided into even and odd tilts. For the cryo-EM datasets, the frames were divided. AreTomo was used for the alignment, and the datasets were reconstructed using the WBP algorithm with the thickness limited to include only cellular features. For the calculation, the resolution FSO protocol of XMIPP3 was used within the Scipion environment, and the Global FSC output was used.
Doses estimation
Table 1 where:
-
e- = electron
-
1 eV = 1,6 \({{\cdot }}\) 10−19 J = energy of 1 e- under a potential difference of 1 V
-
px = pixel size
-
ph = photon
Table 1 Parameters definition X-rays radiation-dose formula:
$$D\left[{G}_{y}\right]=\frac{{{{\rm{\mu}}}}_{l}\left[{{\rm{\mu}}}{m}^{-1}\right]\cdot N\left[\frac{ph}{{{\rm{\mu}}}{m}^{2}s}\right]\cdot E\left[J\right]\cdot\,{t}_{\exp}^{{tot}}\left[s\right]}{{{{\uprho}}}\left[\frac{{kg}}{{{\rm{\mu}}}{m}^{3}}\right]}$$where:
D = radiation dose measured in Grey = \({{{\rm{G}}}}_{{{\rm{y}}}}\left[{{\rm{J}}}/{{\rm{kg}}}\right],{{\rm{which}}}\) is the amount of energy deposited through inelastic scattering per unit mass of specimen, measured in Joule per kilogram
-
\({{\rm{\rho }}}\) is the density of the material, where ρ ~ 0.94 g/cm3 is the average value for amorphous ice
-
\({{{\rm{\mu }}}}_{l}\) is the linear absorption coefficient, where µl ~ 0.1 µm−1 is the average value for amorphous ice
-
\({{\rm{N}}}\) = Flux/unit area = (number of photons/s)/unit area
-
E [J] = energy of the X-rays source measured in Joule
-
ttotexp = total exposure time
Electron radiation-dose formula:
Conversion of e-/Å2 to \({{MG}}_{y}\): [\({{MG}}_{y}\)] = CF \({{\cdot }}\) [e-/Å2] Table 2
Parameters for cryo-soft X-rays tomography half-dose acquisition at Mistral Beamline:
-
Source energy: E = 520 eV = 8.33 × 10−17 J
-
Fresnel zone plates (FZP) = 40 nm
-
Pixel size (px) = 13 nm
-
Flux (total flux at 520 eV at sample position with ES = 30 µm and XS = 15 µm) = 2.08 \({{\cdot }}\) 1010 ph/s (measured with photodiode)
-
XS (for acquisitions) = 8 µm
→ Flux (for acquisitions) = 53% of Flux (at sample position) = 1.11 \({{\cdot }}\) 1010 ph/s
-
Field of View (FoV) = 13 µm × 13 µm = 169 µm2
-
Total n° projections = 131
-
texp = 1 s (Exposure time per projection)
-
µl ~ 0.1 µm−1
-
ρ ~ 0.94 g/cm3 = 0.94 \({{\cdot }}\) 10−15 kg/ µm3
Calculation of Fluence F = N \({{\cdot }}\) t [ph/µm2] (equivalent to Exposure Dose Ed [é/Å2] in EM):
Before the sample:
Flat field: f b = 39,000 cts/s (average value on all tilt series projections)
Fluence: F b = 6.7 \({{\cdot }}\) 107 ph/s µm2 (fluence reaching the sample per FoV per projection)
Total Fluence: F bTOT = 131 \({{\cdot }}\) F b = 0.86 × 1010 ph/µm2 per tilt series
Total Dose: D bTOT [\({{{\rm{MG}}}}_{{{\rm{y}}}}\)]= µl/ρ \({{\cdot }}\) FbTOT \({{\cdot }}\) E \({{\cdot }}\) texp = 76.20 \({{{\bf{MG}}}}_{{{\bf{y}}}}\)
After the sample:
Flat field: f a = 17540 cts/s (average value on all tilt series projections)
Fluence: F a = 3.7 \({{\cdot }}\) 107 ph/µm2 (fluence reaching the sample per FoV per projection)
Total Fluence: F aTOT = 131 \({{\cdot }}\) F a = 0.48 \({{\cdot }}\) 1010 ph/µm2 per tilt series
-
→ 55% adsorbed fluence, supposing linear response of the FZP.
-
→ total adsorbed dose = 0.55 \({{\cdot }}\) DTOT [\({{{\rm{MG}}}}_{{{\rm{y}}}}\)]= 41.92 \({{{\bf{MG}}}}_{{{\bf{y}}}}\)
Parameters for full-dose cryo-electron tomography acquisition on the Titan Krios:
-
Source energy: E = 300 keV
-
Total n° projections = 41
-
Pixel size (px) = 3.104 Å
-
texp = 1.77 s (Exposure time per projection)
-
N (Dose rate) = 25.58 e−/px2/s
-
F (Fluence, Exposure dose) = 4.78 e−/Å2
-
DTOT (Total Radiation Dose per tilt series) = (F \({{\cdot }}\) 41)/texp = 110.48 e−/Å2
Calculation of Fluence/Exposure Dose F[é/A2] for EM (equivalent to Fluence F[ph/µm2] for X-rays):
Before the sample:
Fluence: F b = 4.78178 e-/ Å2 (fluence reaching the sample per FoV per projection)
Total Fluence: F bTOT = 41 \({{\cdot }}\) F b/texp = 110 e−/Å2 per tilt series
Total Dose: F bTOT (e−/ Å2) \({{\cdot }}\) 3.7 = 407 \({{{\bf{MG}}}}_{{{\bf{y}}}}\)
After the sample:
Fluence: F a = 0.89 e-/ Å2 (fluence reaching the sample per FoV per projection)
Total Fluence: F aTOT = 41 \({{\cdot }}\) F a/texp = 20.61 e-/Å2 per tilt series
Total Dose: F aTOT (e-/ Å2) \({{\cdot }}\) 3.7 = 76.26 \({{{\bf{MG}}}}_{{{\bf{y}}}}\)
-
→ 81.3% adsorbed fluence
-
→ Total adsorbed dose = 0.813 × DbTOT [\({{MG}}_{y}\)] = 330.74 \({{{\rm{MG}}}}_{{{\rm{y}}}}\)
At 300 kV: [\({{{\rm{MG}}}}_{{{\rm{y}}}}\)] = CF \({{\cdot }}\) [e-/Å2], with CF = 3.71, as biological material is mostly composed of H20. Table 3
Final consideration. To achieve resolutions better than 3 Å, exposure should be kept below 30 e-/Å2 35, or 111 \({{{\rm{MG}}}}_{{{\rm{y}}}}\). DbTOT has to be kept to minimum to allow high resolution reconstructions and structure determination afterwards, keeping in mind that for a good cryo-SXT tomogram, 40-60% absorption provides good S/N ratio, without using denoising algorithms. Absorption depends highly on composition and thickness and photon flux will be different for each cryo-SXT source.
References
De Boer, P., Hoogenboom, J. P. & Giepmans, B. N. G. Correlated light and electron microscopy: ultrastructure lights up! Nat. Methods 12, 503–513 (2015).
van Rijnsoever, C., Oorschot, V. & Klumperman, J. Correlative light-electron microscopy (CLEM) combining live-cell imaging and immunolabeling of ultrathin cryosections. Nat. Methods 5, 973–980 (2008).
Conesa, J. J. et al. Unambiguous intracellular localization and quantification of a potent iridium anticancer compound by correlative 3D Cryo X-ray imaging. Angew. Chem. Int. Ed. 59, 1270–1278 (2020).
Duke, E. M. H. et al. Imaging endosomes and autophagosomes in whole mammalian cells using correlative cryo-fluorescence and cryo-soft X-ray microscopy (cryo-CLXM). Ultramicroscopy 143, 77–87 (2014).
Mendonça, L. et al. Correlative multi-scale cryo-imaging unveils SARS-CoV-2 assembly and egress. Nat. Commun. 12, 4629 (2021).
Jamme, F. et al. Synchrotron multimodal imaging in a whole cell reveals lipid droplet core organization. J. Synchrotron Radiat. 27, 772–778 (2020).
Kukulski, W. et al. Correlated fluorescence and 3D electron microscopy with high sensitivity and spatial precision. J. Cell Biol. 192, 111–119 (2011).
Lučić, V. et al. Multiscale imaging of neurons grown in culture: from light microscopy to cryo-electron tomography. J. Struct. Biol. 160, 146–156 (2007).
Sartori, A. et al. Correlative microscopy: bridging the gap between fluorescence light microscopy and cryo-electron tomography. J. Struct. Biol. 160, 135–145 (2007).
Henderson, R. Cryo-protection of protein crystals against radiation damage in electron and X-ray diffraction. Proc. R. Soc. Lond. B Biol. Sci. 241, 6–8 (1990).
Baker, L. A. & Rubinstein, J. L. Radiation damage in electron cryomicroscopy. Methods Enzymol. 481, 371–388 (2010).
Hoffmann, P. C. et al. Structures of the eukaryotic ribosome and its translational states in situ. Nat. Commun. 13, 7435 (2022).
Wagner, F. R. et al. Preparing samples from whole cells using focused-ion-beam milling for cryo-electron tomography. Nat. Protoc. 15, 2041–2070 (2020).
Schneider, G. et al. Three-dimensional cellular ultrastructure resolved by X-ray microscopy. Nat. Methods 7, 985–987 (2010).
Parkinson, D. Y., McDermott, G., Etkin, L. D., Le Gros, M. A. & Larabell, C. A. Quantitative 3-D imaging of eukaryotic cells using soft X-ray tomography. J. Struct. Biol. 162, 380–386 (2008).
Otón, J. et al. Characterization of transfer function, resolution and depth of field of a soft X-ray microscope applied to tomography enhancement by Wiener deconvolution. Biomed. Opt. Express 7, 5092 (2016).
Schneider, G. Cryo X-ray microscopy with high spatial resolution in amplitude and phase contrast. Ultramicroscopy 75, 85–104 (1998).
Carzaniga, R., Domart, M. C., Duke, E. & Collinson, L. M. Correlative cryo-fluorescence and cryo-soft X-ray tomography of adherent cells at European synchrotrons. Methods Cell Biol. 124, 151–178 (2014).
Spink, M. C. et al. Correlation of cryo soft X-ray tomography with Cryo fluorescence microscopy to characterise cellular organelles at beamline B24, Diamond Light Source. Microsc. Microanalysis 124, 151–178 (2018).
Okolo, C. A. et al. Sample preparation strategies for efficient correlation of 3D SIM and soft X-ray tomography data at cryogenic temperatures. Nat. Protoc. 16, 2851–2885 (2021).
Groen, J. et al. Correlative 3D cryo X-ray imaging reveals intracellular location and effect of designed antifibrotic protein-nanomaterial hybrids. Chem. Sci. 12, 15090–15103 (2021).
Hagen, C. et al. Multimodal nanoparticles as alignment and correlation markers in fluorescence/soft X-ray cryo-microscopy/tomography of nucleoplasmic reticulum and apoptosis in mammalian cells. Ultramicroscopy 146, 46–54 (2014).
Cinquin, B. P. et al. Putting molecules in their place. J. Cell Biochem 115, 209–216 (2014).
Hagen, C. et al. Correlative VIS-fluorescence and soft X-ray cryo-microscopy/tomography of adherent cells. J. Struct. Biol. 177, 193–201 (2012).
Castro, V., Pérez-Berna, A. J., Calvo, G., Pereiro, E. & Gastaminza, P. Three-dimensional remodeling of SARS-CoV-2-infected cells revealed by cryogenic Soft X-ray tomography. ACS Nano 17, 22708–22721 (2023).
Guo, L. et al. The study of radiation damage of yeast cells in Cryo-soft X-ray tomography. in Selected Papers of the Chinese Society for Optical Engineering Conferences held October and November 2016 (eds. Lv, Y., Le, J., Chen, H., Wang, J. & Shao, J.) 102551Q (2017). https://doi.org/10.1117/12.2268715.
Howells, M. R. et al. An assessment of the resolution limitation due to radiation-damage in X-ray diffraction microscopy. J. Electron Spectros Relat. Phenom. 170, 4–12 (2009).
Egerton, R. F. Dose measurement in the TEM and STEM. Ultramicroscopy 229, 113363 (2021).
Karuppasamy, M., Karimi Nejadasl, F., Vulovic, M., Koster, A. J. & Ravelli, R. B. G. Radiation damage in single-particle cryo-electron microscopy: Effects of dose and dose rate. J. Synchrotron Radiat. 18, 398–412 (2011).
Sorrentino, A. et al. MISTRAL: a transmission soft X-ray microscopy beamline for cryo nano-tomography of biological samples and magnetic domains imaging. J. Synchrotron Radiat. 22, 1112–1117 (2015).
Henke, B. L., Gullikson, E. M. & Davis, J. C. X-ray interactions: photoabsorption, scattering, transmission, and reflection at E = 50-30, 000 eV, Z = 1-92. At. Data Nucl. Data Tables 54, 181–342 (1993).
Reimer, L. & Kohl, H. Transmission Electron Microscopy Physics of Image Formation (Springer, 2008).
Leapman, R. D. & Sun, S. Cryo-electron energy loss spectroscopy: observations on vitrified hydrated specimens and radiation damage. Ultramicroscopy 59, 71–79 (1995).
Henderson, R. The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules. Q Rev. Biophys. 28, 171–193 (1995).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. Elife 4, e06980 (2015).
Carrascosa, J. L. et al. Cryo-X-ray tomography of vaccinia virus membranes and inner compartments. J. Struct. Biol. 168, 234–239 (2009).
Atakisi, H., Conger, L., Moreau, D. W. & Thorne, R. E. Resolution and dose dependence of radiation damage in biomolecular systems. IUCrJ 6, 1040–1053 (2019).
Arnold, J. et al. Site-specific cryo-focused ion beam sample preparation guided by 3D correlative microscopy. Biophys. J. 110, 860–869 (2016).
Paithankar, K. S. & Garman, E. F. Know your dose: RADDOSE. Acta Crystallogr D. Biol. Crystallogr 66, 381–388 (2010).
Lightsources.org. https://lightsources.org/.
Kapishnikov, S. et al. Mode of action of quinoline antimalarial drugs in red blood cells infected by Plasmodium falciparum revealed in vivo. Proc. Natl. Acad. Sci. USA 116, 22946–22952 (2019).
Perez-Berna, A. J. et al. Monitoring reversion of hepatitis C virus-induced cellular alterations by direct-acting antivirals using cryo soft X-ray tomography and infrared microscopy. Acta Crystallogr. D Struct. Biol. 77, 1365–1377 (2021).
Pérez-Berná, A. J. et al. Structural changes in cells imaged by soft X-ray cryo-tomography during hepatitis C virus infection. ACS Nano 10, 6597–6611 (2016).
Chen, J. H. et al. Soft X-ray Tomography Reveals HSV-1-Induced Remodeling of Human B Cells. Viruses 14, 2651 (2022).
Zhang, Y. et al. Structure-function mapping of olfactory bulb circuits with synchrotron X-ray nanotomography. Preprint at https://doi.org/10.1101/2025.04.24.650439 (2025).
Kuan, A. T. et al. Dense neuronal reconstruction through X-ray holographic nano-tomography. Nat. Neurosci. 23, 1637–1643 (2020).
Shahmoradian, S. H. et al. Three-dimensional imaging of biological tissue by cryo x-ray ptychography. Sci. Rep. 7, 6291 (2017).
Bosch, C. et al. Functional and multiscale 3D structural investigation of brain tissue through correlative in vivo physiology, synchrotron microtomography and volume electron microscopy. Nat. Commun. 13, 2923 (2022).
Valcarcel, R. et al. New holder for dual-axis cryo soft X-ray tomography of cells at the Mistral beamline. In Proc. Mechanical Engineering Design of Synchrotron Radiation Equipment and Instrumentation (eds. Schaa, R. W., Tavakoli, V.K. & Tilmont, M.) (2018).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
de la Rosa-Trevín, J. M. et al. Scipion: a software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
Zheng, S. et al. AreTomo: an integrated software package for automated marker-free, motion-corrected cryo-electron tomographic alignment and reconstruction. J. Struct. Biol. X 6, 100068 (2022).
Agulleiro, J. I. & Fernandez, J. J. Fast tomographic reconstruction on multicore computers. Bioinformatics 27, 582–583 (2011).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Last, M. G. F., Voortman, L. M. & Sharp, T. H. scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy. Nat. Methods 20, 1445–1446 (2023).
Paul-Gilloteaux, P. et al. eC-CLEM: flexible multidimensional registration software for correlative microscopies. Nat. Methods 14, 102–103 (2017).
de Chaumont, F. et al. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Acknowledgements
Cryo-SXT data were collected at Mistral beamline at Alba Synchrotron during experimental sessions 2023087704. This work was supported by grants from the Programme Fédérateur de Recherche 6 (PFR6) SARS-CoV-2 and COVID-19. We acknowledge the cryo-ET expertise and assistance of the Institut Pasteur’s NanoImaging Core facility, created and supported by a PIA grant (EquipEx CACSICE: ANR-11-EQPX-0008). The DIHP unit receives is part of the LabEx IBEID and from the LabEx MI. Furthermore, JE acknowledges funding by the ANR PRC RabReprogram.
Author information
Authors and Affiliations
Contributions
J.G. performed all the experimental steps and wrote the manuscript. A.S.R., E.P., and J.G. conceived the study. A.S.R. supervised the study. E.P. collected the cryo-SXT data at Mistral (ALBA). A.S.R. and E.P. provided the dose calculations and scientific discussions. J.E. provided edits to the paper and financial support. A.G. and S.K. provided scientific discussions. A.B. provided financial support, and M.V. provided facility support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: Sergey Kapishnikov is part of SiriusXT, a company developing a cryo-SXT laboratory system. The other authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Hideki Shigematsu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Janesh Kumar and Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Groen, J., Gazi, A., Kapishnikov, S. et al. Cryo-CLXEM introduces cryo-SXT to bridge the resolution gap in cryo-CLEM. Commun Biol 8, 1677 (2025). https://doi.org/10.1038/s42003-025-09072-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09072-x