Abstract
Bacterial carboxyl-terminal processing proteases (CTPs) are essential for protein quality control, signal transduction, and cell adaptation. Typically, activation of CTPs involves substrate binding to the PDZ domain or an adaptor protein. Recent studies of CTPs from various bacterial species have indicated structural diversity in CTP oligomerisation. However, the activation mechanisms and rationale for these different oligomeric forms are not well characterised or understood. Here, we present biophysical analyses of CtpA from Helicobacter pylori, which assembles into a trimer-of-dimer (hexameric) configuration. Hydrogen–deuterium exchange mass spectrometry shows that CtpA transitions between resting and active states independently of substrate binding. Cryo-electron microscopy and crystal structural analysis of CtpA further reveal that only one subunit per dimer is active at a time, driven by asymmetric conformational changes. This asymmetric activation supports a cooperative mechanism in which hexameric subunits synchronise to regulate proteolytic activity. Coordinated inter-subunit interactions and concerted movements of the PDZ domain and motile loop generate three self-compartmentalised catalytic units that enable processive substrate degradation. We also identify intra- and intermolecular interactions that stabilise functional states, allowing adaptor-independent activation. These findings open a new avenue towards understanding the key elements of oligomeric assembly in protease activation.
Similar content being viewed by others
Introduction
Carboxyl (C)-terminal processing proteases (CTPs) are ATP-independent serine proteases that mediate proteolytic cleavage at the C-termini of substrate proteins. These proteases occur in all domains of life and are crucial for protein quality control and signal transduction1,2,3. In bacteria, CTPs regulate key physiological processes such as cell wall remodelling, stress responses and cell cycle progression4,5,6,7,8,9. Although the protein sequence of a CTP can vary from approximately 400–700 amino acid residues in length, sequence homology is used to classify bacterial CTPs into three subfamilies: CTP-1, CTP-B and CTP-310. Studies of representative structures in each subfamily, such as Prc in Escherichia coli, CtpB in Bacillus subtilis (BsCtpB) and CtpA in Pseudomonas aeruginosa (PaCtpA)9,11,12, have revealed diverse activation mechanisms.
One major function of bacterial CTPs is the modulation of peptidoglycan (PG) metabolism. In E. coli, Prc works with the adaptor NlpI to degrade hydrolases, such as MepS, thus ensuring proper remodelling of the PG layer during growth and division4,11. In P. aeruginosa, a parallel system involving PaCtpA and lipoprotein LbcA targets hydrolases such as MepM and PA1199, underscoring a conserved strategy in bacterial envelope regulation5. BsCtpA additionally contributes to DNA damage recovery in B. subtilis by degrading the checkpoint inhibitor YneA, thus facilitating cell division post-repair8. CTP-null strains in Brucella suis, Burkholderia mallei and Staphylococcus aureus all exhibit reduced virulence13,14,15, highlighting the importance of these proteases in bacterial pathogenesis and their potential as therapeutic targets. Despite these insights, our understanding of CTP function is incomplete due to the limited number of substrates identified.
A typical bacterial CTP consists of an N-terminal domain, a PDZ domain, a protease domain (further subdivided into cap and core subdomains) where the catalytic triad resides, and a C-terminal domain (Fig. 1a)9,11,12. Despite the highly flexible orientation of the substrate-binding PDZ domain, the PDZ-protease unit is the most structurally conserved, whereas the N- and C-terminal domains participating in intermolecular interactions that generate different CTP oligomeric forms are variable9,16,17. For example, BsCtpB, PaCtpA, and Acinetobacter baumannii S41 peptidase dimerise by packing their N-terminal helices into a four-helix bundle9,12,16. The PaCtpA dimer further trimerises via the trans-interacting loop and long helix (C-helix) located at the C-terminal of each subunit to form a hexamer12,16. Prc remains monomeric, and its N- and C-terminal β-strands associate to create a bowl-shaped structure17, with the PDZ domain acting as a regulatory switch9,17,18. In the resting state, this domain docks over the protease tunnel, thus misaligning the catalytic residues, while in the active state, it moves outward and realigns the active site for cleavage (Fig. 1b). Prc requires its PDZ domain for activation11,17, whereas the PDZ domain of BsCtpB is autoinhibitory, such that its deletion yields a protease with constitutive activity9.
a The domain organisation of CtpA in H. pylori. The residue range of each domain and the positions of the catalytic triad are marked. b The resting CTP (left) and active CTP (right) are depicted with the different domains marked.
Recent studies have observed that membrane adaptor proteins may also regulate CTP activity. The crystal structure of Prc in complex with the adaptor NlpI and substrate MepS shows that NlpI facilitates Prc activity by bringing MepS and Prc together11,19. By contrast, the adaptor LbcA not only acts as a medium between PaCtpA and its substrates but also regulates PaCtpA activity by stabilising its hexamer12,16. Taken together, these observations suggest that bacterial CTPs have evolved to use different molecular designs for enzyme activation and regulation. Understanding how these structural differences influence CTP behaviour is essential for gaining deeper insights into their functional roles.
In this work, we combine protein crystallography, cryo-electron microscopy (cryo-EM) and hydrogen–deuterium exchange mass spectrometry (HDX-MS) to study the structural basis of Helicobacter pylori CtpA (HpCtpA), a member of the CTP-3 subfamily. H. pylori is the primary pathogen colonising human stomachs and its presence strongly correlates with the incidence of gastric cancer20. CtpA is one of the least well-characterised of the 20 proteases encoded in the H. pylori genome21. Its importance and pathological relevance are indicated by its high sequence conservation and detection of the ctpA gene in a majority of clinical isolates22. Here, we present the proteolytic mechanism of HpCtpA with respect to the putative substrate HP1076, which acts as a co-chaperone of the flagellin export chaperone FliS23 and is involved in bacterial motility24. Our structural and dynamic data demonstrate that HpCtpA is a conformationally dynamic protease that undergoes structural transitions for intrinsic auto-activation and exhibits a processive nature in trimer-of-dimer (i.e., hexameric) assembly. These findings exemplify a new mechanism of CTP self-assembly for effective proteolysis.
Results and discussion
Structures of H. pylori CtpA
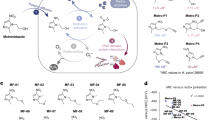
H. pylori CtpA was first identified in a pull-down assay conducted to find interacting partners of HP1076 (Supplementary Fig. 1a). Compared with the control experiments using GST-FliS and GST as bait (lanes 4 and 5), two additional bands were observed in the GST-FliS-HP1076 pull-down product (lane 6). These two bands were subjected to mass spectrometry analysis, which identified them as CtpA and cleaved HP1076, with the C-terminal 21 amino acid residues undetected. Given that CtpA functions as a C-terminal processing protease, we next studied the proteolytic activity of CtpA towards HP1076. To ensure the intracellular expression of recombinant CtpA using an E. coli expression system, we deleted a 20-amino-acid sequence from the N-terminus, which was predicted to be a signal peptide, from the construct. However, the resulting protein product was unstable (Supplementary Fig. 1b). Thus, multiple N-terminal truncated CtpA mutants were screened. Among these, as the most stable construct retaining the longest N-terminal end, CtpAΔN46 (residues 47-454) was selected (hereafter referred to as CtpA or HpCtpA) and used for subsequent investigations in this study. From the results of the CtpA activity assays, a cleaved product of HP1076 was detected over time only in the presence of wild-type CtpA (Supplementary Fig. 1c). The inactive mutant CtpAS300A (substitution of serine at residue 300 with alanine)9,11,12, which was set up in parallel produced no cleaved band, indicating that the serine at residue 300 is critical for CtpA’s cleavage activity. The direct interaction of CtpA with HP1076 was further confirmed using MST, which showed a Kd of 15.4 μM (Supplementary Fig. 1d). Moreover, both CtpA and HP1076 were detected in the membrane fraction; while CtpA is membrane-bound, HP1076 appears to be predominantly membrane-associated, although it is also localised in the cytosol (Supplementary Fig. 1e). These results showed that HP1076 is a putative substrate of CtpA.
Analytic ultracentrifugation (AUC) revealed that CtpA is a hexamer in solution (Supplementary Fig. 1f) and presents as a triangular ring structure as revealed by transmission electron microscopy (TEM) (Supplementary Fig. 1g). AUC also identified a minor species with a molecular weight of 474 kDa, which may represent residual protein impurities or aggregates of CtpA formed during ultracentrifugation. However, TEM did not reveal any higher-order oligomeric assemblies. We further obtained the crystal structure of CtpA at a resolution of 3.4 Å (Table 1). CtpA has a hexameric ring or trimer-of-dimer structure (Supplementary Fig. 2a), resembling that of PaCtpA. An N-terminal dimerisation domain (NTD) and C-terminal dimerisation domain (CTD) flanking the protease core are responsible for hexamerization. Regarding the PDZ domains, three with clear electron densities are compact and located at one face of the hexamer. The remaining three PDZ domains, with weak densities, are located on the opposite face and shifted away from the main body. Of these, two were bound to substrate peptides presumably co-purified from E. coli. Based on the orientations of the PDZ domains and the presence of substrate peptides, the CtpA crystal structure was determined to contain three subunits in the resting state, two subunits in the active state and one subunit in an intermediate state (Supplementary Fig. 2b–e). The α3 belonging to the protease cap subdomain is flexible in the intermediate subunit structure. In each subunit, about 30–40 residues upstream of the C-helix remain unassigned.
To reduce the structural flexibility induced by substrate proteolysis, a triple mutant CtpAS300A/K325A/Q329A (CtpATM) with a disabled catalytic triad9,11,12 was generated. A structural model of CtpATM was built using a cryo-EM map with a resolution of 3.13 Å (Supplementary Fig. 3 and 4, Table 2). Each face of the CtpATM hexamer comprises three subunits in the same state, either active or resting (Fig. 2a). For the active face side of the hexamer, a substrate peptide modelled with poly-alanine was assigned next to the shifted PDZ domain (Fig. 2b, c). A structural comparison of the resting and active subunits revealed that, along with the shift in the PDZ domain, the hinge region (residues 104–107 and 189–192) connecting the PDZ domain to the cap and core subdomains is transformed from loops to a β-sheet (Fig. 2c). Furthermore, the active subunits were found to exhibit other conformational changes, including movement of the NTD and cap subdomain as described in BsCtpB and PaCtpA9,12,16. Residues 376–392 are modelled between the core subdomain and the CTD (Fig. 2c). This segment and residues 393–422, which are predicted to be disordered, constitute the motile loop (ML), so named for its flexible position within the active and resting subunits. Specifically, ML either contacts the PDZ domain or inserts into the cleft between an active and a resting subunit. As described in later sections, this ML region helps to regulate CtpA activity.
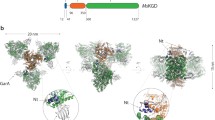
a Cryo-EM density map of the CtpATM hexamer. The resting and active subunits are indicated by cold and warm colours, respectively. The substrates are coloured in yellow. b Structure of CtpATM in a cartoon presentation. The subunits are coloured as in (a). The substrates are depicted in molecular surface mode. c Enlarged view showing the differential conformations relative to the protease core domain in the resting state (left) and active state (right). The domains are coloured as depicted in Fig. 1a, and the substrate is coloured in brown with a transparent molecular surface. The motile loops are displayed as thick ribbons.
Structural dynamics of CtpA
The co-existence of resting and non-resting states revealed in the CtpA crystal structure and CtpATM cryo-EM structure suggests that the protease switches between these two states by swinging the PDZ domain inwards and outwards. We used HDX-MS to investigate the intrinsic dynamics of CtpA. Similar studies of CTPs have not been documented previously. The CtpATM construct was denatured with urea, refolded and purified to remove the endogenously bound substrate from E. coli, yielding apo CtpATM. In total, 91.1% of the CtpA sequence was covered by the MS (Supplementary Fig. 5a and Supplementary Table 1). For each subunit within the CtpA hexamer, limited deuterium uptake was observed near the active site (residues 304–320) and the C-terminal dimerisation interface (residues 430–438) even after 2 h of labelling, indicating the rigidity of these structures (Supplementary Figs. 5b and 6). Conversely, the surface solvent-exposed area showed significant deuterium uptake within a short period, in line with the structural model. Bimodal distribution patterns were observed, especially for the peptides covering α3 in the protease cap subdomain and the hinge connecting the PDZ domain (residues 90–113) and part of the ML (residues 393–422) (Fig. 3a, b). This bimodal distribution pattern may indicate the presence of multiple conformations25. It is plausible that CtpA subunits exist in equilibrium between the active and resting states in solution, independently of substrate binding. We speculate that the resting form is dominant because it has a larger interface area between the PDZ domain and the main body of CtpA than that of the active form (777.4 vs 363.5 Å2). Next, to monitor the molecular dynamics of CtpA during substrate binding, the apo CtpATM sample was incubated with an excess of HP1076 to shift the equilibrium of the hexamer subunits to the active conformation to the greatest extent possible. Complexing CtpA with HP1076 showed a protective effect of deuterium uptake at multiple sites, including α2 of the NTD (residues 73–82), part of the PDZ domain (residues 105–142), part of the protease core subdomain (residues 238–256) and the ML contacting region containing residues 345–364 (Fig. 3c and Supplementary Fig. 5c, d). The regions exhibiting decreases in deuterium uptake are likely to be related to the interfaces that come in contact with the substrate HP1076 and/or the structural changes induced upon HP1076 binding. These include the tunnel surrounded by the protease cap and core subdomains, the PDZ domain, the hinge and the region in the activated protease core subdomain near the ML. The decreases in deuterium uptake also suggest that, apart from the PDZ domain, which reaches outwards, the remainder of active CtpA has a more compact and rigid structure that provides additional protection from deuteriation of the NTD and cap subdomains. The bimodal spectral distribution of α3 and the ML region was also found in the CtpA-HP1076 sample (Supplementary Fig. 7), indicating that when CtpA was saturated by the substrate, the structure still contained a combination of subunits with different conformations. This finding is consistent with the crystal and cryo-EM structures, in which half of the subunits in one hexamer were found to exist in the resting state. These results suggest a potential structural constraint that prevents all subunits in the hexamer from shifting to the active state simultaneously.
a HDX-MS stacked spectral plots of peptides covering residues 90–113 and 393–422 from apo CtpATM. The HDX intervals were 0, 0.5, 1, 10, 60, and 120 min. The bimodal distribution (green and blue peaks) indicates the simultaneous presence of multiple states. b The peptides in (a) are mapped in the resting state (blue; upper panel) and active state (red; lower panel) of the CtpA structure. c Docking of HP1076 to the CtpA subunit bound to the substrate peptide. The summed HDX difference of 10- and 120 min labelling is mapped on the CtpA structure. Unidentified peptides in HDX-MS are coloured in black. Enlarged view shows the contact interface between the C-terminal tail of HP1076 and CtpA. The sequence of the peptide with the largest HDX difference is labelled. Kinetic plots of peptides with a significant ΔHDX are shown at the bottom, with error bars representing the standard deviation (n = 3 independent experiments).
Asymmetric conformational changes and assembly of the self-compartmentalisation unit in CtpA
The crystal structure of CtpA and the cryo-EM structure of CtpATM both reveal three resting subunits and three non-resting subunits located at opposite faces of the hexamer, and this subunit arrangement led us to explore the regulatory mechanism of CtpA activation. We attempted to visualise the dynamic conformational changes in wild-type CtpA using cryo-EM. Heterogeneous refinement resulted in three maps corresponding to CtpA in different conformations: (I) three resting subunits at one face and three non-resting subunits at the other face (map resolution: 3.43 Å); (II) two resting subunits and one non-resting subunit at one face, and one resting subunit and two non-resting subunits at the other face (map resolution: 3.59 Å); and (III) six resting subunits (map resolution: 3.49 Å) (Fig. 4a). The differential arrangement of the resting and non-resting subunits within a CtpA hexamer implies that at least one subunit within a CtpA dimer mediated by the NTD (N-dimer) is in the resting state. On this basis, a maximum of three subunits within a CtpA hexamer would be active. This conclusion is consistent with the HpCtpA structures revealed in this study and the currently available PaCtpA structures12,16.
a Cryo-EM density maps of three particle populations of wild-type CtpA identified via heterogeneous refinement. Resting and non-resting subunits are labelled with cold and warm colours, respectively. b Surface view of an N-terminal dimer showing the asymmetric conformation. The dynamic unit is depicted transparently with secondary structures. Static units 1 and 2 are coloured in blue, and the PDZ domains are coloured in yellow. Each static unit moves to either side as indicated by the purple arrows. The enlarged view shows hydrogen bonds within the dynamic unit. The two involved subunits are shown as semi-transparent ribbon models, coloured green and purple, respectively. c A self-compartmentalising working unit presented in surface view. The active (Sa) and resting (Sr) subunits are coloured in red and blue, respectively. The PDZ domains are indicated by dotted lines. The motile loops provided by the C-dimer partners are depicted as ribbons and indicated as arrowheads. These two C-dimer partners are presented transparently. The substrate is coloured in yellow. The enlarged window illustrates the ML1 binding interface. Residues involved in hydrogen bond and salt bridge formation are marked. Steric hinderance from the resting PDZ domain that repels ML2 is marked by a red arrow.
This phenomenon of at least one resting state per N-dimer may be related to the mechanism of CtpA subunit activation. A detailed structural analysis revealed that extensive hydrogen bonds and salt bridges are formed between the two NTDs and protease cap subdomains (excluding α3) of each subunit in a single N-dimer. This group of interactions form a rigid and isolated module (referred to as the dynamic unit) (Fig. 4b, Supplementary Fig. 8 and Supplementary Table 2). At the two distal ends of the N-dimer, the protease core subdomain and the CTD constitute another rigid module referred to as static units 1 and 2 (Fig. 4b). Within a single N-dimer, constraints from the C-terminal dimerisation interfaces maintain the ring structure. Therefore, the positions of static units 1 and 2 are largely fixed, leaving the dynamic unit to shift towards either static unit 1 or 2. This shift is coupled with the outward movement of the corresponding PDZ domain in the active subunit. Consequently, an asymmetric conformational change happens within one N-dimer in which only one subunit within the N-dimer can be activated, while the other subunit necessarily remains at rest (Fig. 4b). Following this transformation, the protease tunnel is exposed in the activated subunit for substrate binding. Although BsCtpB shares an N-terminal dimerisation pattern with HpCtpA and PaCtpA, its C-terminal dimerisation is facilitated by a four-helix domain positioned between the protease domain and the C-helix, rather than the long C-helix used in HpCtpA and PaCtpA (Supplementary Fig. 9). This difference in dimerisation interface architecture not only results in a transition of the oligomeric state from hexameric to dimeric but also enables both subunits within BsCtpB to be independently active. Therefore, we speculate that the C-dimerisation interface of BsCtpB does not generate structural constraints. These findings imply that switching of the PDZ domain can be regulated by other structural modules.
The asymmetric conformational change observed in each N-dimer would reduce the catalytic power of CtpA by half. We recognised this necessary change when we examined the structural details of the N-dimers and their respective C-dimers (i.e., the CtpA dimer mediated by the CTD) in both CtpATM and CtpA in conformations I and II. We found that each N-dimer and the two MLs (ML1 and ML2), each from a C-dimer partner, assemble to form a self-compartmentalising protease unit (Fig. 4c). Regarding the active subunit (Sa), its narrow protease tunnel could be accessed via either face of the hexamer. However, substrate entrance at one face is blocked by the PDZ domain of the resting subunit (Sr) and ML1 from the C-dimer partner of Sa. Specifically, ML1 fills the gap between the protease core subdomain of Sa and the PDZ domain of Sr. The occupancy of ML1 in this region is achieved by a shift of the PDZ domain in Sa, which leads to exposure of the NTD and protease core subdomain. This exposure enables extensive hydrogen bonding and salt bridge formation between ML1 and the surrounding region (Fig. 4c, Supplementary Fig. 8 and Supplementary Table 3). In contrast, the PDZ domain of Sr occupies the interface on the NTD and the protease core subdomain of Sr, thus repelling ML2 from the C-dimer partner of Sr. ML2 moves with the shifted PDZ domain of Sa and partially seals the bottom of the self-contained unit (Fig. 4c). Therefore, the concerted swapping of MLs with subunit activation acts as a gatekeeper to ensure one-way substrate entry in Sa. While the specific interaction between ML2 and the PDZ domains of Sa cannot be defined due to the low local resolution, Glu-382 in ML1 is linked to His-85 of the protease core in Sa via hydrogen bonds (Fig. 4c). CtpA activity was found to be largely reduced in the H85A and E382A mutants and totally abolished in the E382R mutant (Supplementary Fig. 10). In PaCtpA, the equivalent residue His-84 functions as the acid residue of the catalytic triad16. Our results further demonstrate the importance of the MLs for full proteolytic function. Both His-85 and Glu-382 are highly conserved among CtpAs but not CtpBs; these residues are likely to have evolved in a pairwise manner for activity regulation (Supplementary Fig. 8).
ATP-independent processive proteolysis of HpCtpA is mediated by Arg-162
A self-compartmentalising protease can undergo processive proteolysis, during which the substrate is continuously processed, without releasing the intermediate, until digestion is completed26,27. The processivity of CtpA towards HP1076 was assayed using reversed-phase high-performance liquid chromatography (HPLC) (Supplementary Fig. 11). The HPLC profiles of the product peaks varied for the non-processive protease trypsin. However, proteolysis by CtpA yielded a decrease in the substrate peak and an increase in the product peaks over time. These results indicate that CtpA undergoes processive digestion. In previous research on the ATP-independent processive protease MtaLonC, an electrostatic ratchet mechanism was proposed in which a protonated Glu attracts the nascent C-terminus of the substrate, which is translocated along the proteolytic groove in each cleavage cycle28. In CtpA, the basic residue Arg-162 at the ligand binding site in the PDZ domain can be readily protonated under the experimental condition of pH 7.5. Therefore, we speculate that Arg-162 acts to attract substrates (Supplementary Fig. 12). The resting PDZ domain and ML1 shield the substrate to prevent its escape in the opposite direction towards the activated PDZ domain and facilitate translocation of the nascent intermediate in the self-compartmentalised unit. Consistent with these observations, HDX-MS revealed that the deuterium exchange of the ML was protected after being bound with HP1076 (Supplementary Fig. 5c).
Self-compartmentalisation and its associated processivity comprise a strategy utilised by proteases to control proteolysis, thus preventing proteins from unintentional degradation. The target substrate must be recognised, unfolded and translocated through a narrow pore to reach the catalytic sites29. This process is often ATP-driven, as exemplified by ClpX in ClpXP protease and Lon protease28. In H. pylori, four ATP-dependent self-compartmentalising proteases are present in the genome: HslVU, ClpXP, Lon and FtsH30,31. These proteases share similar features, namely self-compartmentalisation via protomer oligomerisation into barrel-shaped structures31. Here, trimerisation of the dimer in CtpA generates three self-compartmentalisation units, each with an asymmetric conformation. Substrate translocation is presumed to be driven by the protonated Arg-162. These properties allow CtpA to efficiently control its activity in an ATP-free periplasmic environment, at a cost of half of the CtpA molecules.
Substrate recognition and conformation maintenance of CtpA
The conserved residue Arg-168 in BsCtpB is crucial for substrate sensing and active conformation maintenance9. The equivalent residue Arg-162 in CtpA is salt-bridged with Glu-96 in α3 of the protease cap subdomain, where it stabilises the PDZ domain in the Sa subunits (Fig. 5a). Mutants R162A and R162E in CtpA exhibited a loss of enzyme activity, although the effect of R162E was greater (Supplementary Fig. 10). The correct orientation of Arg-162 is likely to be related to the cation–π interaction with a highly conserved Phe-105 in the hinge of the PDZ domain. An electron density connecting the C-terminus of the substrate and the side chain of Phe-105 is observed in CtpATM, highlighting the involvement of Phe-105 in substrate binding (Fig. 5a). In line with these observations, the F105R mutant exhibited drastically impaired activity, whereas the F105Y mutant retained proteolytic activity (Supplementary Fig. 10). The 3.7 Å crystal structure of the F105R mutant revealed a hexamer with all protomers in the resting state, further suggesting that Phe-105 is essential for maintaining the active conformation (Fig. 5b and Table 1). Additionally, HDX-MS analysis of the CtpATM/F105R mutant revealed monomodal spectral patterns for peptides spanning residues 90–113 and 394–422 (Supplementary Figs. 13 and 14). Comparison with HDX-MS data from the CtpATM mutant further confirmed that the F105R mutation impaired the conformational transition from the resting to the active state.
a The substrate binding site in an active subunit. The enlarged view shows how the stabilisation of the active state is mediated by the salt bridge between Glu-96 and Arg-162 and the cation–π interaction between Phe-105 and Arg-162 (dotted box). Processive proteolysis of the C-terminus of the substrate is driven by the pronated Arg-162. The cryo-EM density near the C-terminus of the substrate peptide is shown. b The crystal structure of the CtpAF105R mutant, displayed in surface view. The six resting subunits are labelled in cold colours, and one of the subunits is transparent. c The crossing angles of the two α-helices in the C-terminal dimerisation interface of CtpA from P. aeruginosa (left) and H. pylori (right).
Although HpCtpA and PaCtpA are structurally very similar, the former is autoactivated in vitro. In PaCtpA, LbcA must bind to the NTD to activate the protease and enable trimer-of-dimer formation through the CTD12,16. Structural comparisons of the two CtpA proteases revealed various insights, leading us to propose that the activation mechanism is species-specific. Although the CTD domains are structurally conserved, the crossing angle of helices α7 in the interface of the PaCtpA apo form is about 12° greater than that in the interface of HpCtpA (Fig. 5c). This results in a smaller interface area (319 Å2) in the PaCtpA C-dimer than in HpCtpA (436 Å2). The interaction between LbcA and the NTD domain allows a particular PDZ domain to shift in the N-dimers and the ML to move inward to promote C-dimer formation. All cap subdomains in the crystal structure of the PaCtpA hexamer are in the resting state12, further suggesting that PaCtpA does not intrinsically possess an active-resting state equilibrium as seen in HpCtpA. In contrast, the binding interface at the CTD in HpCtpA is sufficient to maintain the hexameric form even if all subunits are in the resting state.
In conclusion, the structural and biophysical characterisations of the self-compartmentalised HpCtpA indicate its molecular dynamics, where only one subunit can be active per N-dimer. The concerted movement of the PDZ domain and the MLs from the C-dimer partners during the conversion between resting and active states only allows the unfolded C-terminus of a substrate to enter the active subunit and be processed in a processive manner. This work also explains the role of conserved Arg-162 and Phe-105 (Supplementary Fig. 8) in interacting with the C-terminus of a substrate in the activation process and opens a new perspective to understand the structural basis of C-terminal proteases. It remains unclear why HpCtpA acts so differently from PaCtpA, and thus, additional biological functional studies are needed.
Methods
H. pylori growth conditions
H. pylori G27 strain (kindly provided by Professor Karen Ottemann, Department of Microbiology and Environmental Toxicology, University of California) was cultured at 37 °C on Columbia blood agar with 5% defibrinated horse blood under microaerobic conditions (5% CO2, 4% O2, and 91% N2) produced by AnaeroGen gas packs (Oxoid). Brucella broth containing 10% (v/v) foetal bovine serum (BB10) was used for liquid H. pylori culture.
Pull-down assays
The pull-down assays were performed as described previously23. Briefly, purified GST, GST-FliS and GST-FliS-HP1076 were immobilised to glutathione Sepharose (Cytiva), followed by incubating with the cell lysate of H. pylori G27 cells in the buffer containing 20 mM HEPES (pH 7.5), 137 mM NaCl, 27 mM KCl, 5% glycerol, 0.1% Tween-20 and 10 mM DTT for 3 h at 4 °C. After washing three times, the resin beads were mixed with SDS-PAGE sample loading buffer and analysed with SDS-PAGE.
CtpA expression and purification
Fragments of DNA encoding different CtpA truncations were amplified from the cell lysate of H. pylori G27 strain and subcloned into the pAC28m vector to add an N-terminal His6 tag. The plasmids expressing different mutants were constructed using site-directed mutagenesis based on corresponding wild-type protein expression plasmids. The primers used in this study are listed in Table 3. Proteins were expressed in E. coli BL21(DE3) after induction using 0.25 mM isopropyl-β-D-thiogalactopyranoside (IPTG) at 20 °C for 20 hours. Cells were sonicated in lysis buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl and 60 mM imidazole and then centrifuged at 20,000 × g for 1 h at 4 °C to collect the supernatant. After filtering with 0.22 μm pore size filters, the supernatant was mixed and incubated with Ni-NTA agarose (Cytiva) for 1 h at 4 °C, followed by washing using the lysis buffer. Proteins were eluted with elution buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl and 300 mM imidazole. The eluant was further purified with a mono STM 4.6/100 PE column (Cytiva) in 20 mM Tris-HCl (pH 7.5), and a 150 mM to 1 M NaCl gradient and followed by gel filtration with a HiLoad® 16/600 Superdex® 200 PG column (Cytiva) in the buffer containing 20 mM Tris-HCl (pH 7.5) and 150 mM NaCl. For the purification of apo CtpATM, 8 M urea was added in the lysis buffer before sonication and used all through the Ni-NTA purification. After eluted with 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 60 mM imidazole and 8 M urea, the eluant was dialysed to remove urea, followed by ordinary purification steps described above.
Activity assay of CtpA on HP1076 and immunoblot detection of HP1076
Recombinant HP1076 protein was mixed with either wild-type CtpA or CtpA mutants at a molar ratio of 10:1 in a reaction buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, and 1 mM DTT. The mixtures were incubated at room temperature for various time intervals to assess time-dependent cleavage. Reactions were terminated by the addition of SDS-PAGE loading buffer and subsequently boiled for 5 min. Protein samples were separated by SDS-PAGE and transferred to PVDF membranes. Immunoblotting was performed using a rabbit anti-HP1076 antibody to detect both full-length HP1076 and its cleaved products. Donkey anti-rabbit IgG-HRP (Santa Cruz, Cat# sc-2313) was used as the secondary antibody.
Cell fractionation
An overnight liquid culture of H. pylori was centrifuged with 6000 × g for 10 min at 4 °C to harvest the cells. The cell pellet was resuspended in 10 mM Tris-HCl (pH 8.0) to an OD600 of 10, followed by sonication to break the cells. The cell lysate was centrifuged with 6000 g for 10 min at 4 °C to remove the cell debris and unbroken cells. The supernatant was further ultracentrifuged with 100,000 × g for 1 h at 4 °C. The supernatant containing cytosolic and periplasmic fractions and the pellet containing membrane fraction were collected for the subsequent immunoblotting analysis. Mouse anti-CtpA and rabbit anti-HP1076 antibodies were used to detect CtpA and HP1076, respectively. Mouse anti-OMP (Santa Cruz, Cat# sc-57779) was used to detect both OMP and HSP. Goat anti-mouse IgG-HRP (Santa Cruz, Cat# sc-2005) and donkey anti-rabbit IgG-HRP (Santa Cruz, Cat# sc-2313) were used as secondary antibodies to detect the corresponding primary antibodies.
Microscale thermophoresis (MST)
HP1076 was fluorescently labelled with the Monolith NT Protein Labelling Kit RED-NHS (Nano Temper Technologies). CtpAS300A was serially diluted to concentrations ranging from 165 μM to 5.04 nM, followed by mixed with 45 nM of labelled HP1076 in a 1:1 volume ratio. The assays were performed in a monolith NT.115 instrument (Nano Temper Technologies) with 70% excitation power and 40% MST power at 20 °C. The data was analysed by the MO. Affinity Analysis software.
Protein crystallisation and structure determination
The crystallisation experiments were set on sitting-drop vapour-diffusion crystallisation plates. The crystals of wild-type CtpA were grown in the buffer containing 100 mM bis-Tris-HCl (pH 6.5), 100 mM magnesium chloride, 25% PEG 3350 at 16 °C. The crystals of CtpAF105R mutant were grown in the condition A8 of Morpheus® crystallisation screen (Molecular Dimensions). The X-ray diffraction data was collected with TPS 05 A at the National Synchrotron Radiation Research Centre, Taiwan. The datasets were processed using HKL200032 or iMosflm33. The crystals of wild-type CtpA were in the space group P1211, and the CtpAF105R crystals were in the space group H3. Molecular replacement was used to solve the structures with Phaser34 in the Phenix suite35. Pruned and truncated structure models of Photosystem II D1 C-terminal processing protease (PDB ID: 1FC6) and CtpB from B. subtilis (PDB ID: 4C2E) were used together as search models to perform molecular replacement for wild-type CtpA, with six chains in the asymmetric unit. The resting protomer of wild-type CtpA was used as a search model for CtpAF105R mutant, with four chains in the asymmetric unit. Structure building, refinement and subsequent rebuilding were performed using COOT36 and Phenix. The protein coordinates have been deposited to Protein Data Bank (PDB ID: 9JR1 and 9KU0). The structural images were prepared using USCF ChimeraX37 and PyMol.
Hydrogen deuterium exchange mass spectrometry (HDX-MS)
CtpATM in complex with HP1076 was prepared by mixing apo CtpATM with HP1076 and incubated overnight at 4 °C. Stock solutions of 10 μM CtpATM either in apo form or in complex with HP1076, or CtpATM/F105R were prepared by diluting them in protein buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl. To perform HDX experiments, 3 μl of the stock solutions were added into 47 μl of HDX buffer (20 mM Tris-HCl, 150 mM NaCl in D2O, pD 7.5) at room temperature. Control experiments without HDX were performed by diluting protein samples in the corresponding deuterium-free protein buffer. After different incubation time periods (0.5, 1, 10, 60, 120 min), each HDX was quenched by adding 50 μl of pre-cooled quenching buffer (20 mM Tris-HCl (pH 1.7), 150 mM NaCl) to reach a final pH of 2.5. Immediately after quenching, 50 μl of the sample was injected into a Acquity M-Class UPLC system linked with a HDX Manager (Waters, USA) at 0 °C. The samples were online digested with an Enzymate BEH-Pepsin column (Waters, 300 Å, 5 μm, 2.1 × 30 mm) at 25 °C and subsequently desalted with a BEH C18 Trap Column (Waters, 130 Å, 5 μm, 300 × 50 mm) for a total of 3 min at 100 μl/min. The peptides were eluted through a BEH C18 column (Waters, 130 Å, 1.7 μm, 2.1 × 100 mm) using a linear gradient from 95% solvent A (H₂O with 0.1% formic acid) and 5% solvent B (acetonitrile with 0.1% formic acid) to 5% solvent A and 95% solvent B over 12 minutes at a flow rate of 100 µL/min. Mass spectra were acquired with an m/z range of 100–3000 using the Waters SYNAPT G2-Si Mass Spectrometry in MSE mode. Three replicates were performed for each CtpA sample with different incubation time.
HDX data analysis
The mass spectra raw data were processed by ProteinLynx Global Server to generate the peptide lists. The following filtering parameters were set in DynamX 3.0 to remove low-quality signal peaks: Minimum intensity: 10,000; Maximum sequence length: 30; Minimum products per amino acid: 0.3; Maximum MH+ Error (ppm): 5; Minimum PLGS score: 7. The quality of resulting signal peaks after filtering was further inspected manually. The average standard deviation of the deuterium uptake was below 0.1 Da. Difference in deuterium uptake with 95% confidence interval (CI) was considered as a significant change for differential HDX. Uptake figures were produced by Microsoft Excel and relative deuterium uptake was mapped onto the crystal model of proteins using PyMol. HX-Express338 was used to perform binomial fitting and bimodal deconvolution of the spectra.
Cryo-EM sample preparation and data collection
To prepare the cryo-EM grids, 4 μl of protein sample at a concentration of 2 mg ml-1 was applied to the glow-discharged holey carbon grids (Quantifoil R1.2/1.3). After a wait time of 5 s and a blotting time of 3 s, the grids were plunge frozen into liquid ethane using Vitrobot Mark IV (Thermo Fisher) operated at 4 °C and 100% humidity. Data acquisition was performed using a Titan Krios microscope (Thermo Fisher) operated at 300 kV and equipped with a GIF Quantum energy filter and a Gatan K3 direct electron detector. Automatic image collection was carried out using SerialEM39 with a slit width of 20 eV on the energy filter in the super-resolution counting mode at a magnification of ×130,000 for CtpATM and ×105,000 for wild-type CtpA, yielding calibrated pixel sizes of 0.92 Å and 0.83 Å, respectively. The defocus range was set from -1.5 to -2.5 μm. Each movie stack was dose-fractionated to 32 frames with a total dose of ~50 e Å-2.
Cryo-EM data processing and map calculation
The movies collected were motion-corrected with MotionCor240, followed by contrast transfer function estimation (CTF) with CTFFIND41. Image processing was performed using Relion42 and cryoSPARC software43. Particles were auto-picked and processed with 2D classification using Relion. Noise and junk particles were removed by multiple rounds of 2D classification. Selected 2D classes were used for automated particle picking by Gautomatch (http://www.mrc-lmb.cam.ac.uk/kzhang/). After removing junk particles by multiple rounds of 2D classification, ab initio reconstruction and heterogeneous refinement were performed to identify the best classes of particles. Non-uniform refinement was further applied to generate the final cryo-EM density maps.
For wild-type CtpA, 1,583,803 particles were picked by template picking and further extracted and imported into cryoSPARC. The particles were subjected to 2D classification with a mask diameter of 226 Å. 561,739 particles were left to perform ab initio reconstruction. After heterogeneous refinement in cryoSPARC, 155,703 particles were selected and subjected to nonuniform refinement with C3 symmetry followed by local refinement, yielding a map for CtpA conformation I with an overall resolution of 3.4 Å (EMD-62575); 133,262 particles were selected and refined to yield a map for CtpA conformation II with an overall resolution of 3.6 Å (EMD-62576); 144,375 particles were selected and refined to yield a map for CtpA conformation III with an overall resolution of 3.5 Å (EMD-62553).
For CtpATM, 1,449,887 particles were picked by template picking and further extracted and imported into cryoSPARC. The particles were subjected to 2D classification with a mask diameter of 226 Å. 381,365 particles were left to perform ab initio reconstruction. After heterogeneous refinement in cryoSPARC, 151,917 particles were selected and subjected to nonuniform refinement with C3 symmetry followed by local refinement, yielding a map with an overall resolution of 3.1 Å (EMD-62573).
Model building and coordinate refinement
An initial predicted model was generated with DeepTracer44,45,46 after submitting the cryo-EM map of CtpATM and its sequence file in fasta format. The crystal structure of wild-type CtpA was superimposed onto the initial model and modified to fit the density in COOT. The missing structures were manually built in COOT. Cycles of model building and refinement were performed using COOT and real_space_refine in Phenix. The geometry of the model was refined with the ISOLDE47 plugin in ChimeraX. The final refined model was validated using MolProbity48. The protein coordinates have been deposited to Protein Data Bank (PDB ID: 9KU3, 9KUB, 9KUC and 9KSP) The hydrogen bonds and contact interfaces were analysed with the PDBePISA server49. The structural images were prepared using USCF ChimeraX and PyMol.
Absorbance-based analytical ultracentrifugation
Sedimentation velocity studies of CtpA were performed using a Beckman proteomeLab XL-I analytical ultracentrifuge equipped with an An-60 Ti rotor at 16 °C. Double-sector centrifuge cells were loaded with 380 μl of the sample and 400 μl of the reference buffer. Data from sedimentation at 28,000 rpm were collected at 4 min intervals for 12 h in a continuous mode. Sedimentation velocity data were fitted to a c(s) continuous size distribution model using SEDFIT50 to determine the sedimentation coefficients.
Processive proteolysis monitored by reverse-phase HPLC
HP1076 (50 μg) was mixed with CtpA (5 μg) in the buffer containing 20 mM Tris-HCl (pH 7.5) and 150 mM NaCl and incubated for 0, 15, 30, 60, 90 min at room temperature. The reaction was quenched by adding the same volume of 7.4 M guanidine hydrochloride. For the negative control, HP1076 (50 μg) was mixed with trypsin (3 μg) and incubated for 0, 15, 30, 60, 90 min at 37 °C, followed by quenching of the reaction by adding the same volume of 7.4 M guanidine hydrochloride. Immediately after quenching, 5 μl of the sample was injected onto an XBridge BEH C18 column (Waters, 130 Å, 5 μm, 4.6 × 250 mm) at a flow rate of 1.2 ml/min with a 1260 Infinity II LC System equipped with a 1290 Infinity II Diode Array Detector (Agilent). A 0 to 100% acetonitrile in 0.1% trifluoroacetic acid gradient in 30 min was performed to elute the degradation products at a flow rate of 1.2 ml/min. The peaks were detected at 210 nm wavelength.
Statistics and reproducibility
For HDX-MS analysis, mass differences greater than 0.19 Da were considered significant. Statistical comparisons between CtpATM with and without HP1076 binding were performed using Student’s t-test, with significance defined as p < 0.05. Each dataset was generated from more than three independent experiments. All other experiments were also independently repeated at least three times to ensure reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM maps and atomic coordinates of CtpA and CtpATM have been deposited in the Electron Microscopy Data Bank (https://www.ebi.ac.uk/pdbe/emdb/) under accession number EMD-62576, EMD-62553, EMD-62573, and EMD-62575, and in the Protein Data Bank (https://www.rcsb.org) under accession number 9KUC, 9KSP, 9KU3, and 9KUB, respectively. The crystal structure models of CtpA and CtpAF105R have been deposited in the Protein Data Bank under accession number 9JR1 and 9KU0, respectively. The HDX-MS data have been deposited to the ProteomeXchange Consortium via the PRIDE51 partner repository with the dataset identifier PXD068881. The numerical source data underlying the graphs presented in this study are available in Supplementary Data 1 and 2. Uncropped and unedited blot and gel images are provided in Supplementary Fig. 15. All other data are available from the corresponding author upon reasonable request.
References
Keiler, K. C., Patrick, R. H. W. & Sauer, R. T. Role of a peptide tagging system in degradation of proteins synthesized from damaged messenger RNA. Science 271, 990–993 (1996).
Otero-Asman, J. R. et al. The Prc and CtpA proteases modulate cell-surface signaling activity and virulence in Pseudomonas aeruginosa. iScience 26, 107216 (2023).
Che, Y. et al. C-terminal processing of reaction center protein D1 is essential for the function and assembly of photosystem II in Arabidopsis. Proc. Natl. Acad. Sci. USA 110, 16247–16252 (2013).
Singh, S. K., Parveen, S., SaiSree, L. & Reddy, M. Regulated proteolysis of a cross-link–specific peptidoglycan hydrolase contributes to bacterial morphogenesis. Proc. Natl. Acad. Sci. USA 112, 10956–10961 (2015).
Srivastava, D. et al. A proteolytic complex targets multiple cell wall hydrolases in Pseudomonas aeruginosa. mBio. 9, e00972–18 (2018).
Saoud, J., Mani, T. & Faucher, S. P. The tail-specific protease is important for Legionella pneumophila to survive thermal stress in water and inside amoebae. Appl. Environ. Microbiol. 87, e02975–20 (2021).
Roy, R., You, R., Lin, M. & Lin, N. Mutation of the carboxy-terminal processing protease in Acinetobacter baumannii affects motility, leads to loss of membrane integrity, and reduces virulence. Pathogens 9, 322 (2020).
Burby, P. E., Simmons, Z. W., Schroeder, J. W. & Simmons, L. A. Discovery of a dual protease mechanism that promotes DNA damage checkpoint recovery. PLoS Genet. 14, e1007512 (2018).
Mastny, M. et al. CtpB assembles a gated protease tunnel regulating cell-cell signaling during spore formation in Bacillus subtilis. Cell 155, 647–658 (2013).
Rawlings, N. D., Barrett, A. J. & Bateman, A. MEROPS: the peptidase database. Nucleic Acids Res. 38, D227–D233 (2010).
Su, M. et al. Structural basis of adaptor-mediated protein degradation by the tail-specific PDZ-protease Prc. Nat. Commun. 8, 1516 (2017).
Hsu, H., Wang, M., Kovach, A., Darwin, A. J. & Li, H. Pseudomonas aeruginosa C-terminal processing protease CtpA assembles into a hexameric structure that requires activation by a spiral-shaped lipoprotein-binding partner. mBio. 13, e0368021 (2022).
Bandara, A. B., Sriranganathan, N., Schurig, G. G. & Boyle, S. M. Carboxyl-terminal protease regulates Brucella suis morphology in culture and persistence in macrophages and mice. J. Bacteriol. 187, 5767–5775 (2005).
Bandara, A. B. et al. A disruption of ctpA encoding carboxy-terminal protease attenuates Burkholderia mallei and induces partial protection in CD1 mice. Microb. Pathog. 45, 207–216 (2008).
Bojer, M. S. et al. SosA inhibits cell division in Staphylococcus aureus in response to DNA damage. Mol. Microbiol. 112, 1116–1130 (2019).
Hsu, H., Wang, M., Kovach, A., Darwin, A. J. & Li, H. P. aeruginosa CtpA protease adopts a novel activation mechanism to initiate the proteolytic process. EMBO J. 43, 1634–1652 (2024).
Chueh, C. et al. Structural basis for the differential regulatory roles of the PDZ domain in C-terminal processing proteases. mBio. 10, e01129–19 (2019).
Sommerfield, A. G. & Darwin, A. J. Bacterial Carboxyl-terminal processing proteases play critical roles in the cell envelope and beyond. J. Bacteriol. 204, e0062821 (2022).
Wang, S. et al. Structural basis for recruitment of peptidoglycan endopeptidase MepS by lipoprotein NlpI. Nat. Commun. 15, 5461 (2024).
Polk, D. B. & Peek, R. M. Helicobacter pylori: gastric cancer and beyond. Nat. Rev. Cancer 10, 403–414 (2010).
Tomb, J. F. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539–547 (1997).
Gharibi, S. et al. Relationship between histopathological status of the Helicobacter pylori infected patients and proteases of H. pylori in isolates carrying diverse virulence genotypes. Microb. Pathogenesis 110, 100–106 (2017).
Lam, W. W. L. et al. Molecular interaction of flagellar export chaperone FliS and cochaperone HP1076 in Helicobacter pylori. FASEB J. 24, 4020–4032 (2010).
Douillard, F. P., Ryan, K. A., Hinds, J. & O’Toole, P. W. Effect of FliK mutation on the transcriptional activity of the σ54 sigma factor RpoN in Helicobacter pylori. Microbiology 155, 1901–1911 (2009).
Na, S., Lee, J., Joo, J. W. J., Lee, K. & Paek, E. deMix: decoding deuterated distributions from heterogeneous protein states via HDX-MS. Sci. Rep. 9, 3176 (2019).
Thompson, M. W., Singh, S. K. & Maurizi, M. R. Processive degradation of proteins by the ATP-dependent Clp protease from Escherichia coli. Requirement for the multiple array of active sites in ClpP but not ATP hydrolysis. J. Biol. Chem. 269, 18209–18215 (1994).
Butler, S. M., Festa, R. A., Pearce, M. J. & Darwin, K. H. Self-compartmentalized bacterial proteases and pathogenesis. Mol. Microbiol. 60, 553–562 (2006).
Li, S. et al. Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex. Sci. Adv. 7, eabj9537 (2021).
Baumeister, W., Walz, J., Zühl, F. & Seemüller, E. The proteasome: paradigm of a self-compartmentalizing protease. Cell 92, 367–380 (1998).
Kim, D. Y. & Kim, K. K. Crystal structure of ClpX molecular chaperone from Helicobacter pylori. J. Biol. Chem. 278, 50664–50670 (2003).
De Mot, R., Nagy, I., Walz, J. & Baumeister, W. Proteasomes and other self-compartmentalizing proteases in prokaryotes. Trends Microbiol 7, 88–92 (1999).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Cryst. D. 67, 271–281 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl Crystallogr 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Guttman, M., Weis, D. D., Engen, J. R. & Lee, K. K. Analysis of overlapped and noisy hydrogen/deuterium exchange mass spectra. J. Am. Soc. Mass Spectrom. 24, 1906–1912 (2013).
Mastronarde, D. N. SerialEM: a program for automated tilt series acquisition on tecnai microscopes using prediction of specimen position. Microsc. Microanal. 9, 1182–1183 (2003).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Nakamura, A. et al. Fast and automated protein-DNA/RNA macromolecular complex modeling from cryo-EM maps. Brief. Bioinform. 24, bbac632 (2023).
Si, D. et al. DeepTracer web service for fast and accurate de novo protein complex structure prediction from Cryo-EM. Algorithms and Methods in Structural Bioinformatics, edited by Haspel, N., Jagodzinski, F. & Molloy, K., pp. 101–114. (Springer, 2022).
Pfab, J., Phan, N. M. & Si, D. DeepTracer for fast de novo cryo-EM protein structure modeling and special studies on CoV-related complexes. Proc. Natl. Acad. Sci. USA 118, e2017525118 (2021).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D. Struct. Biol. 74, 519–530 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgements
This research was funded by the Hong Kong Research Grants Council (General Research Fund 14117622 and Collaborative Research Fund C4012-16E) and National Natural Science Foundation of China (NSFC, No. 31900046 and 82372269 to H.Z., No. 82172465 to D.W.). We extend our gratitude to the staff at beamlines BL15A and TPS05A of the National Synchrotron Radiation Research Centre, Taiwan, China. The authors thank the Cryo-EM Centre of Southern University of Science and Technology for data collection and HPC-Service Station.
Author information
Authors and Affiliations
Contributions
Conceptualization, K.S. and S.W.N.A.; methodology, K.S., P.K.S., S.P.K.T., H.Z., and S.W.N.A.; investigation, K.S., L.Y., C.Y.M., P.K.S., S.P.K.T., and H.Z.; writing – original draft, K.S.; writing – review and editing, K.S., P.K.S., S.P.K.T, H.Z., and S.W.N.A.; funding acquisition, S.W.N.A.; resources, K.F.L., D.W., H.Z., and S.W.N.A.; supervision, S.W.N.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Daniel Bonsor and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Janesh Kumar and Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, K., Yan, L., Mok, C.Y. et al. Structures of Helicobacter pylori C-terminal protease CtpA reveal a new mode of self-contained proteolytic processing. Commun Biol 8, 1828 (2025). https://doi.org/10.1038/s42003-025-09175-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09175-5