Abstract
Gastroesophageal reflux disease (GERD) is a highly prevalent gastrointestinal disorder with complex genetic underpinnings.While genome-wide association studies (GWAS) have identified several GERD-associated loci, traditional GWAS approaches rely on stringent significance thresholds and may miss variants with modest effects that still contribute to disease biology. To enhance the discovery of GERD-associated loci, we developed InsightGWAS, a Transformer-based deep learning model. Using transfer learning, the model was pre-trained on major depressive disorder GWAS data and fine-tuned with GERD GWAS summary statistics. We integrated multi-omics functional annotations, including eQTLs, mQTLs, and epigenomic data, to prioritize candidate variants. Comparative analyses showed that InsightGWAS outperformed logistic regression, XGBoost, and neural networks, achieving superior classification accuracy and reducing false positives. The model replicated known GERD loci and uncovered 209 novel candidate loci, many involved in neurogenic, neuromuscular, and epithelial pathways. Enrichment analyses revealed associations with synaptic transmission, neural development, and cadherin-mediated signaling, suggesting that both nervous system regulation and epithelial integrity contribute to GERD pathophysiology. This study demonstrates the power of deep learning in advancing genetic discovery beyond conventional GWAS. By leveraging transfer learning and multi-omics annotations, InsightGWAS identifies potential disease-asscoated biological pathways underlying GERD, offering promising directions for mechanistic research and potential therapeutic targets.
Similar content being viewed by others
Introduction
Gastroesophageal reflux disease (GERD) is a common multifactorial disorder caused by chronic reflux of gastric acid into the esophagus, with a high prevalence in the population1. It is also the major risk factor for Barrett’s esophagus and esophageal adenocarcinoma2,3. Genetic factors are thought to play a substantial role in GERD, with heritability estimates ranging from 40% to 50%4,5. In recent years, large-scale GWAS meta-analyses have indeed begun to elucidate the genetic architecture of GERD. A meta-analysis of approximately 80,000 GERD cases and 305,000 controls identified 25 independent loci associated with GERD1. Notably, several implicated genes from that study are targets of existing reflux medications or biologically relevant to acid regulation. However, these loci account for only a fraction of GERD heritability, suggesting that additional genetic factors remain undiscovered. Identifying variants requires significantly larger sample sizes, posing challenges that limit the capacity of traditional GWAS to fully unravel the genetic architecture of GERD. Moreover, traditional GWAS approaches rely on stringent significance thresholds and may overlook variants with moderate effects that still contribute to disease biology. Identifying such explanatory variants requires more integrative and flexible analytical frameworks capable of capturing subtle yet biologically meaningful signals.
Epidemiological and genetic studies have revealed a significant relationship between mental disorders and GERD, suggesting shared biological pathways underlying these conditions6. Anxiety and depression, two of the most commonly comorbid mental health conditions with GERD, are reported to exacerbate GERD symptoms through mechanisms such as altered pain perception, increased visceral sensitivity, and dysregulated autonomic nervous system responses7. Genetic correlation analyses further support this association, indicating overlapping genetic factors between GERD and psychiatric disorders6. These findings reveal the complex interplay between neurological and gastrointestinal systems, suggesting the importance of novel computational approaches to studying GERD’s genetic architecture.
Cross-trait analyses have emerged as a powerful approach to uncover shared genetic architecture between GERD and related conditions. These methods leverage genetic correlations and pleiotropic effects to identify loci that influence multiple traits, enhancing discovery beyond what is achievable with traditional GWAS. For GERD, cross-trait meta-analyses with anxiety and depression have revealed loci involved in neuroinflammatory pathways and vagal regulation, shedding light on the shared mechanisms underlying these conditions6. Similarly, analyses integrating GERD, psychiatric conditions and obesity GWAS datasets have identified loci linked to esophageal motility and adiposity, reflecting the metabolic and mechanical interplay contributing to GERD pathophysiology8. By integrating data from related traits, cross-trait studies enhance the detection of novel loci and provide deeper insights into the biological pathways linking GERD with its comorbid conditions.
Beyond traditional GWAS and statistical genetics methods, deep learning has revolutionized genetic studies by offering advanced methods to uncover complex relationships in genomic data9,10,11,12. Transformer-based models are particularly effective in capturing nonlinear relationships and integrating heterogeneous data types, including GWAS summary statistics, functional annotations, and epigenomic data. These models leverage attention mechanisms to prioritize key genetic signals, enabling more accurate identification of disease-associated loci. Additionally, transfer learning, where a model pre-trained on one trait is fine-tuned for another, has proven effective in leveraging shared genetic architecture across traits to enhance discovery.
In this study, we leveraged deep learning to investigate the genetic architecture of GERD. Building on the genetic correlations between GERD and mental disorders, we developed a Transformer-based model to identify GERD-associated loci by leveraging their shared genetic background. This model was pre-trained on major depressive disorder (MDD) GWAS data and refined through transfer learning to enhance sensitivity for GERD-specific genetic variants. By integrating multi-omics functional annotations, including regulatory and epigenomic data, we aim to uncover novel loci and elucidate the biological pathways contributing to GERD pathophysiology.
Results Genetic correlations between GERD and mental disorders
To investigate the shared genetic basis between GERD and several psychiatric disorders, we performed LD score regression (LDSC) using GWAS summary statistics for GERD and six psychiatric conditions (Table 1), including MDD, attention-deficit/hyperactivity disorder (ADHD), anxiety disorders (ANX), bipolar disorder (BIP), autism spectrum disorder (ASD), and schizophrenia (SCZ). The genetic correlations (rg) and P-values are summarized in Supplementary Table 1.
The strongest genetic correlation was observed between GERD and MDD (rg = 0.52, p = 5.08E-134), reflecting a substantial overlap in their genetic architecture. This strong association is consistent with clinical evidence of high comorbidity between GERD and depression and suggests that shared biological mechanisms, such as neuro-immune dysregulation and stress pathways, may contribute to both conditions. In addition to MDD, significant positive genetic correlations were identified with ADHD (rg = 0.53, p = 1.01E-94), ANX (rg = 0.26, p = 1.35E-05), and BIP (rg = 0.11, p = 0.0003), indicating notable shared genetic components with GERD. A modest but significant correlation was also observed with ASD (rg = 0.0821, p = 0.025), suggesting a weaker genetic overlap. In contrast, the genetic correlation with SCZ (rg = 0.0284, p = 0.21) did not reach statistical significance.
Developing the transformer-based deep learning model using MDD GWAS data
The complex genetic architecture of GERD presents substantial challenges for traditional GWAS, particularly due to modest variant effect sizes. To overcome these limitations, we developed InsightGWAS, a Transformer-based neural network, to improve the prediction of variant-phenotype associations by incorporating an expanded set of genetic features and targeted enhancements for greater accuracy. Each genetic variant is treated as a distinct observation, represented by a carefully curated feature set that includes GWAS summary statistics, population-level genetic metrics, and functional annotations relevant to neurological traits. While traditional GWAS methods rely on statistical hypothesis testing and P-value thresholds, InsightGWAS complements these approaches by serving as a post-GWAS prioritization model that uses machine learning to re-rank variants based on their predicted relevance to the trait of interest. This probabilistic approach allows InsightGWAS to recover potentially meaningful loci that may fall below genome-wide significance in conventional analyses, especially in settings with limited sample size or statistical power. As such, InsightGWAS is intended to complement standard GWAS, offering biologically informed variant prioritization rather than replacing formal association testing.
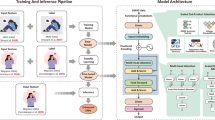
To further address the challenges of defining binary labels for variant associations, we adopted a two-step labeling strategy. Both labels and features were derived from GWAS datasets of MDD, but with different statistical power. Association labels were defined from a large, well-powered MDD GWAS, where variants surpassing the genome-wide significance threshold (P < 5 × 10-8) were labeled as positives and the remainder as negatives. Input features, however, were extracted from a smaller MDD GWAS with lower power, including summary statistics (effect size, P-value, allele frequency, and sample size) as well as multi-omics annotations (eQTLs, chromatin accessibility, conservation scores, etc.). This design allowed the model to learn the genomic patterns underlying true associations from a large dataset while generalizing to weaker signals in smaller cohorts. Here, we specifically selected MDD due to the strong genetic correlation and shared genetic architecture between MDD and GERD, as shown in the genetic correlation analyses. Utilizing MDD data provides well-characterized variant associations, enabling robust model training and enhancing the generalizability of InsightGWAS when applied to GERD and other related traits (Fig. 1).
a The analytical workflow begins with pretraining a baseline model using major depressive disorder GWAS summary statistics, followed by transfer learning with GERD GWAS data. The adapted model is then applied to identify novel GERD-associated variants. b The model architecture is based on a Transformer neural network that integrates GWAS summary statistics with functional genomic annotations. Through red attention-weighted connections, the network captures complex relationships among genetic features to enhance variant prioritization. Blue boxes indicate genomic inputs, and orange boxes represent functional annotations. Organ icons in this figure were obtained from Vecteezy.com and are used under the Free License.
To better understand the contribution of different types of features, we performed an ablation study, systematically removing one feature group at a time and re-evaluating model performance (Supplementary Fig. 1). The results demonstrated that core variant-level features (e.g., chromosome, LD score, frequency) and GWAS summary statistics were the most influential, with the greatest drop in F1 score observed when these groups were removed (ΔF1 = −0.1393 and −0.1109, respectively). In contrast, omitting regulatory features (e.g., chromatin accessibility, TF binding, conservation) led to more modest performance changes, suggesting that these annotations provide complementary, but not primary, predictive power. Regulatory QTLs, including bulk and single-cell eQTLs and mQTLs, showed intermediate importance, supporting their utility in refining association signals.
In summary, InsightGWAS utilizes a Transformer architecture with an attention mechanism to dynamically prioritize key features, enabling the precise identification of variant associations. The model also mitigates data imbalance by focusing on disease-associated variants without the need for undersampling, preserving critical information while minimizing noise. Moreover, its multi-head self-attention mechanism enhances transfer learning by facilitating the integration of insights from large datasets, thereby reducing the need for extensive retraining.
Comparative evaluation of transformer and traditional models
To evaluate the performance of the Transformer-based model (InsightGWAS) against conventional machine learning methods, we compared predicted score distributions using the MDD dataset. Benchmarked approaches included neural networks (DeepGWAS), logistic regression, XGBoost, and ridge regression, all trained on identical input features for consistency (see Methods).
Kernel density estimation (KDE) plots (Fig. 2) revealed that InsightGWAS achieved superior separation between positive and negative classes, with predicted scores tightly clustered near 0 for negatives and near 1 for positives. In contrast, traditional models exhibited more overlap between classes, reflecting weaker discriminative power. The Transformer’s attention mechanism enabled sharper class separation, particularly minimizing false positives, which is critical when prioritizing disease-associated SNPs across millions of loci. In addition, experimental results showed that the Transformer model (InsightGWAS) outperformed traditional approaches in managing data noise and class imbalance, as demonstrated by ROC and DET curve analyses (Supplementary Fig. 2).
Predicted score distributions are shown for Ridge Regression, Transformer (InsightGWAS), Neural Network (DeepGWAS), Logistic Regression, and XGBoost models. KDE plots compare the density of predicted scores for true negative (blue) and true positive (red) labels. Transformer models demonstrate clearer separation between positive and negative predictions compared to conventional methods (Supplementary Data 3).
Overall, our analysis shows InsightGWAS’s advantage in robustly distinguishing true signals from background noise, outperforming prior models under conditions of class imbalance and data complexity (see Supplementary Materials for details).
GERD loci identified by InsightGWAS via transfer learning
To evaluate the performance of the Transformer-based model (InsightGWAS) against conventional approaches, we next applied transfer learning to refine the pre-trained Transformer model for GERD association analysis using the summary statistics of two large-scale GERD GWAS1,13. Following our two-step labeling strategy, features derived from smaller-scale GWAS were paired with labels from larger GERD GWAS datasets, enabling the model to learn from well-powered association signals while remaining sensitive to underpowered variants. To preserve previously learned patterns while adapting to GERD-specific associations, the model was fine-tuned on these datasets with a reduced learning rate of 0.0001. The GERD data were split into training (80%) and validation (20%) sets, achieving a validation accuracy of 98.90% after 100 epochs. Model performance was evaluated across ~7.35 million SNPs, prioritizing precision by applying a stringent decision threshold to minimize false positives. SNPs were classified as significant only when assigned a probability ≥ 99%. This approach resulted in a low false positive rate of ~0.0049% (366/7,350,000), demonstrating the robustness and specificity of the model in detecting GERD-associated loci.
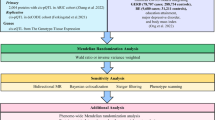
To improve discovery power, we applied enhancements to the largest available GERD GWAS dataset, using summary statistics from An et al1, excluding 23andMe data due to policy restrictions (71,522 cases and 261,079 controls). Independent GWAS signals were identified using PLINK’s clumping procedure, applying an r² threshold of 0.05 to account for LD. Through our InsightGWAS framework, we identified 268 independent signals, which encompassed all 27 genome-wide significant loci reported in the non-23andMe GERD GWAS by An et al1 (Figs. 3,4 and Supplementary Data 1). We next compared our findings with those of Ong et al8, who performed a multi-trait genome-wide association analysis (MTAG) combining GERD with genetically correlated traits, including educational attainment, depression, and body mass index, also based on the summary statistics from An et al. (excluding 23andMe)1. Ong et al8 identified 88 GERD risk loci, including 74 loci not previously reported by An et al1. Notably, our InsightGWAS approach independently predicted 32 of these 74 novel loci, indicating its capacity to prioritize risk loci beyond conventional GWAS significance thresholds and supporting the robustness and reliability of the identified associations.
Venn diagram showing the overlap of indenpendent signals (SNPs after clumping to remove linkage disequilibrium, see Supplementary Data 3 for details.). Coral represents the InsightGWAS-predicted variants, yellow represents variants from conventional GWAS (An et al1.), and purple represents variants identified in the MTAG analysis (Ong et al(8).).
The plot displays −log10(p) association values for GERD from the GWAS by An et al., excluding 23andMe data. Genome-wide significance threshold (p < 5 × 10⁻8) is indicated by the red dashed line. Orange circles represent all variants prioritized by InsightGWAS, illustrating the complete set of SNPs ranked by the model as potentially trait-relevant. Independent association signals were then identified through LD clumping of these prioritized variants. Orange diamonds mark the independent lead loci that correspond to previously reported genome-wide significant regions from An et al1. and Ong et al8. Violet diamonds indicate newly identified independent loci highlighted by InsightGWAS that were not reported in prior GERD studies (Supplementary Data 3).
In addition to replicating the signals reported by An et al1 and Ong et al8, InsightGWAS identified 209 additional loci associated with GERD. Among these, 94 demonstrated nominal replication (P < 0.05) in the Million Veteran Program (MVP) cohort, providing independent support for their significance (Supplementary Data 1)14. Several of these loci converge on pathways regulating gastrointestinal motility, epithelial integrity, and neurohumoral control, all central to GERD pathophysiology. For example, rs6690625 in LEPR influences energy balance, gastric motility, and satiety; rs4791898 in GLP2R contributes to intestinal epithelial maintenance and gastric emptying; and rs73169236 in VIPR2 modulates smooth muscle relaxation. In addition, several identified loci implicate neurogenic and neuromuscular mechanisms in GERD susceptibility. Candidate genes from those loci such as NLGN1 (rs247975), CTNNA3 (rs10733835), SORCS3 (rs7096926), ROBO2 (rs7426688), LAMA2 (rs9492271), and PTN (rs13245564) are involved in neural development, synaptic transmission, sensory processing, and smooth muscle regulation. Together, these loci point to the contribution of altered neural signaling, esophageal sensitivity, and impaired neuromuscular coordination to esophageal motility and lower esophageal sphincter function, providing mechanistic insights into GERD pathogenesis.
Biological Interpretation of GERD-associated loci
To elucidate the functional relevance of the newly identified GERD risk loci, we performed gene and pathway enrichment analyses using the InsightGWAS-predicted loci. Candidate genes were prioritized by MAGMA, selecting those within 500 kb of each locus with P < 0.01 for downstream pathway annotation.
Our pathway enrichment analysis of GERD risk loci revealed progressively deeper biological insights across different analytical approaches (Fig. 5 and Supplementary Data 2). The conventional GWAS (An et al.)1 showed minimal enrichment, with only nominal associations for basic processes like protein binding (GO:0005515). In contrast, MTAG analysis (Ong et al.)8 demonstrated improved detection of neuronal pathways, including synapse organization (GO:0050808) and synaptic structure (GO:0045202). Most strikingly, novel loci from InsightGWAS identified exceptional enrichment for specific neural mechanisms - modulation of chemical synaptic transmission (GO:0050804) and neuron development (GO:0048666) - suggesting potential pathways underlying visceral hypersensitivity. The complete InsightGWAS analysis (all loci) not only confirmed these neural associations but also revealed additional epithelial mechanisms through cadherin binding (GO:0045296) and trans-synaptic signaling (GO:0099537), providing a comprehensive view of GERD’s dual neural-epithelial pathology.
Enrichment results (-log10(P values) and enrichment ratios) for Gene Ontology (GO) terms across four analytical approaches (Supplementary Data 3): conventional GWAS (An et al1.), MTAG analysis (Ong et al8.), novel loci from InsightGWAS (InsightGWAS-new), and all loci from InsightGWAS (InsightGWAS-all).
Discussion
We demonstrate that deep learning methods, particularly transformer-based architectures, provide a powerful complement to conventional GWAS approaches in elucidating the genetic architecture of GERD. By systematically comparing these methods, we show that deep learning uncovers additional genetic associations not detected by traditional or cross-trait analyses. These results suggest that leveraging deep learning will be essential for fully exploiting existing genomic resources and advancing the genetic understanding of GERD.
The genetic overlap between GERD and comorbid conditions such as depression has been increasingly recognized through epidemiological and GWAS studies6,7. Cross-trait analyses have successfully harnessed these correlations to identify novel pleiotropic loci involved in neuroimmune regulation and esophageal function6,8, suggesting their value as a complement to traditional GWAS. Beyond correlation, emerging evidence suggests that the frequent co-occurrence of GERD and depression reflects shared biological mechanisms rather than coincidental comorbidity. Epidemiological studies demonstrate a bidirectional relationship between GERD and depression or anxiety, where each increases the risk of the other15. Genetic and Mendelian randomization analyses further indicate causal links and identify shared genes involved in neuroinflammatory and stress-response pathways16. Mechanistically, several convergent processes have been implicated, including gut–brain axis dysfunction—where altered vagal signaling and microbial composition influence both reflux symptoms and mood regulation—chronic immune–inflammatory activation with cytokine spillover contributing to neuroinflammation, and HPA axis dysregulation that links chronic stress to both esophageal and emotional dysregulation15,17,18. Together, these findings suggest that GERD and psychiatric disorders share overlapping pathophysiological networks that connect peripheral inflammation, neuroendocrine stress, and brain–gut communication. Nonetheless, as both standard and cross-trait approaches face diminishing returns, there is a growing need for innovative computational frameworks capable of unlocking additional insights into GERD’s complex genetic architecture. To address these limitations, we developed InsightGWAS, a Transformer-based deep learning framework designed to advance genetic discovery in GERD and related complex traits. By leveraging transfer learning from large-scale datasets such as MDD and integrating multidimensional genomic annotations—including mQTLs, single-cell eQTLs, and transcription factor binding sites—InsightGWAS efficiently detects both established and novel loci. Its attention-based architecture enables dynamic prioritization of biologically relevant signals across heterogeneous genomic landscapes, overcoming the limitations of previous neural network approaches and offering a robust tool for uncovering the regulatory and functional mechanisms underlying GERD.
By applying InsightGWAS to the GERD GWAS summary statistics by An et al. 1, we replicated many loci reported in subsequent cross-trait studies8 and uncovered additional candidate loci with potential clinical relevance. Among these, 94 demonstrated nominal replication (P < 0.05) in the MVP cohort, providing supportive evidence for their involvement in disease susceptibility. Notably, several genes from these signals are involved in metabolic regulation and esophageal motility, pathways already implicated in GERD pathogenesis. Regional association plots for representative loci are shown in Supplementary Fig. 3. For instance, the LEPR gene (rs6690625) encodes the leptin receptor, linking metabolic and immune pathways. Elevated leptin is associated with GERD and Barrett’s esophagus19 and promotes esophageal inflammation via MIF-driven cytokine induction20. Leptin also correlates with esophageal acid exposure and GERD symptoms21. The GLP2R gene (rs4791898) encodes the glucagon-like peptide-2 receptor, which regulates gastrointestinal motility, gastric relaxation, and energy balance. GLP-2, via GLP2R on enteric neurons and hypothalamic POMC neurons, suppresses feeding, delays gastric emptying, and improves glucose homeostasis22,23. In mouse models, GLP-2 induces gastric fundus relaxation, modulated by nutritional state, and enhances gastric adaptive responses24. Disruption of GLP2R function may accelerate gastric emptying, increase gastric acid exposure, and potentially contribute to GERD susceptibility. The VIPR2 gene (rs73169236) (VIP), which inhibits gastric acid secretion and regulates gut motility. VIP-mediated suppression of acid secretion25,26 may influence esophageal acid exposure and contribute to GERD.
Consistent with the observed genetic correlations between GERD and neuropsychiatric traits, several identified loci indicate neurogenic and neuromuscular mechanisms underlying disease risk. Loci such as rs247975 (NLGN1), rs10733835 (CTNNA3), rs13245564 (PTN), and rs7096926 (SORCS3) map to genes involved in neural development, synaptic signaling, and sensory processing, suggesting that altered neural regulation and esophageal hypersensitivity may contribute to GERD pathogenesis. Additionally, ROBO2, a key regulator of axon guidance and smooth muscle function, modulates vascular cell proliferation via NF-κB signaling27 and harbors mutations in gastrointestinal cancers28, suggesting its involvement in esophageal motility and neuromuscular control. LAMA2, encoding the laminin α2 chain of the basement membrane, shows distinct distribution in esophageal and gastric epithelia29,30, and its dysregulation in Barrett’s esophagus points to a role in epithelial remodeling and barrier maintenance, further contributing to GERD susceptibility.
Our pathway enrichment analyses revealed that while conventional GWAS (An et al.)1 offered limited insights, cross-trait MTAG analyses (Ong et al.)8 uncovered neuronal pathways such as synapse organization, reflecting the shared neurogenetic architecture underlying GERD and neuropsychiatric conditions. Notably, InsightGWAS advanced these findings by showing additional enrichment in neural development, chemical synaptic transmission, and epithelial mechanisms such as cadherin binding, suggesting the dual involvement of neurogenic and epithelial factors in GERD pathogenesis. These results demonstrate that deep learning–based approaches can uncover previously underrecognized biological processes that may contribute to disease susceptibility and progression.
Our study has several limitations. First, due to data availability constraints, the analyses were restricted to European cohorts. Future investigations in non-European populations will be crucial for evaluating the robustness and generalizability of these findings. Second, although we identified novel loci and pathways associated with GERD, their direct functional effects remain unclear. Additional experimental studies will be necessary to elucidate the underlying biological mechanisms and to facilitate clinical translation. Third, residual sample overlap across GWAS datasets may slightly bias genetic correlation estimates; however, LDSC-derived correlations were used only to identify trait pairs rather than as explicit model parameters, so any minor bias is unlikely to affect model performance or overall conclusions.
In conclusion, our study demonstrates the potential of integrating advanced computational frameworks with genomic data to enhance the identification of pathogenic mechanisms in GERD, offering insights that may inform future translational research and therapeutic strategies.
Methods
GWAS data
Summary statistics were obtained from two GERD GWAS datasets. An et al. 1 included 71,522 GERD cases and 261,079 controls from large European cohorts, excluding 23andMe due to policy restrictions. Donertas et al13 analyzed 20,381 cases and 464,217 controls from UK Biobank as part of a broader study on age-related diseases.
For psychiatric traits, data were obtained from the Psychiatric Genomics Consortium (PGC). MDD included two datasets1: a meta-analysis of UK Biobank and PGC cohorts (170,756 cases, 329,443 controls)31 and2 a combined dataset from PGC, deCODE, UK Biobank, and iPSYCH (59,851 cases, 113,154 controls)32. BIP data consisted of 41,917 cases and 371,549 controls33, while ADHD included 38,691 cases and 186,843 controls34. ASD comprised 18,381 cases and 27,969 controls from family-based and iPSYCH cohorts14. Anxiety disorder (ANX) data included 7016 cases and 14,745 controls35, and schizophrenia (SCZ) was based on the largest GWAS meta-analysis to date (76,755 cases, 243,649 controls)36. Detailed sample descriptions are provided in Supplementary Table 1. All GWAS summary statistics used in this study were obtained from previously published studies with appropriate institutional ethics approval and informed consent from participants. For each dataset, all ethical regulations relevant to human research participants were followed, as stated in the original publications.
Genetic correlation analysis
We assessed the genetic correlation (rg) between GERD and psychiatric phenotypes using LDSC, restricting analyses to HapMap3 variants in line with established guidelines37. The estimated range of rg is from −1 to 1, where −1 indicates an absolute negative correlation and 1 an absolute positive correlation. Analyses were conducted with precomputed LD scores from the HapMap3 European reference panel of ~1.2 million common SNPs, derived from the 1000 Genomes Project. LDSC provides approximately unbiased rg estimates even in the presence of sample overlap, as overlap primarily inflates the intercept rather than the regression slope from which rg is derived. In accordance with LDSC best practices, intercepts were left unconstrained to allow for potential sample overlap.
Transformer-based model
The InsightGWAS model utilizes a Transformer encoder architecture to capture complex, nonlinear relationships among genetic features. This design is particularly suited for genomic data, where interactions between elements, such as sQTLs, eQTLs, and positional information, are critical for identifying disease-associated loci. The encoder comprises two layers, each equipped with four multi-head attention mechanisms and a feed-forward network. Input features are embedded and positionally encoded, allowing the model to integrate both interaction patterns and genomic context. The self-attention mechanism dynamically weights input features, focusing on key genetic signals and their dependencies, while multi-head attention captures diverse relationships between features. The encoder’s output is processed through a fully connected layer with sigmoid activation to predict disease associations, eliminating the need for a decoder. To ensure robust training, the model incorporates hidden dimensions (set at 64), layer normalization, and residual connections. The codebase is publicly accessible on GitHub (https://github.com/ziangmeng/MDD-GERD-InsightGWAS), and additional technical details, including architecture specifications, training protocols, and evaluation metrics, are available in the Supplementary Materials.
Input features
The InsightGWAS framework integrates both GWAS-derived statistics and functional annotations at the SNP level. Each input sample corresponds to a single SNP, defined by its chromosomal position and alleles. For each SNP, we first included standard GWAS summary features: effect estimates, standard errors, P-values, allele frequencies, and cohort sample sizes. To account for correlation structure, we added LD metrics derived from European ancestry reference data in the 1000 Genomes Project.
Functional annotations were mapped to SNP genomic coordinates. Regulatory features included tissue- and cell type–specific eQTLs and sQTLs from GTEx and BrainMeta, DNA methylation QTLs, and single-cell eQTLs from brain cell populations (Supplementary Table 2). We also incorporated epigenomic annotations from ENCODE, including chromatin accessibility peaks, transcription factor binding, and histone modification states. These features were aligned to SNP positions and encoded as binary or quantitative predictors depending on the source dataset. To enhance biological relevance and model performance, we further incorporated functional and regulatory annotations, including variants that affect noncoding RNAs (microRNAs, snoRNAs) and transcriptional regulators, as well as trait associations curated from the GWAS Catalog and Open Targets. Evolutionary conservation and predicted functional impact scores (GERP + +, PolyPhen-2, SIFT, CADD) were also integrated, providing measures of deleteriousness.
Together, these SNP-level features enabled the model to jointly assess statistical association, regulatory potential, and functional impact, thereby enhancing the prioritization of loci of biological relevance. Full details of feature preprocessing and model training are provided in the Supplementary Materials.
Comparison with alternative approaches
To benchmark the predictive capabilities of InsightGWAS, we systematically compared it against several established models, including a neural network, XGBoost, logistic regression, and Ridge regression. The neural network, implemented in Keras, followed a multilayer architecture with ReLU activations and a sigmoid output layer, paralleling frameworks such as DeepGWAS10. Training was conducted over 30 epochs with binary cross-entropy loss, incorporating batch normalization to improve convergence and stability. The XGBoost model, a gradient-boosted tree ensemble, was configured in a logistic regression framework with early stopping to prevent overfitting and root mean square log error (RMSLE) as the evaluation metric. Baseline classifiers included logistic regression and Ridge regression (both from scikit-learn), with the latter employing L2 regularization and cross-validation to tune hyperparameters.
Model performance was assessed using a combination of true positive rate (TPR), F1 score, and receiver operating characteristic (ROC) curves, providing a comprehensive evaluation of sensitivity, precision, and classification performance across models. This comparative analysis allowed us to contextualize the added value of the Transformer architecture relative to both traditional statistical and modern machine-learning methods.
Transfer learning
To improve the detection of GERD-associated loci, the Transformer model leveraged a transfer learning framework. The model was initially pre-trained using a MDD dataset, allowing it to capture generalizable patterns of genetic associations across complex traits. This pre-training phase provided a robust initialization for the downstream task. The model was then fine-tuned on GERD-specific data, with the learning rate set at 0.0001. Early stopping was applied during fine-tuning to mitigate overfitting and promote stable convergence. Prior to training, input features from the GERD dataset were standardized, and the data were partitioned into training and validation subsets to enable performance monitoring. Model weights were updated via backpropagation to minimize the loss function, and both validation loss and classification accuracy were tracked to evaluate convergence and optimize generalization.
Statistics and reproducibility
This study utilized publicly available GWAS summary statistics for analysis. For genetic correlation analyses, LDSC was employed to evaluate the relationship between GERD and psychiatric disorders. The statistical significance of the genetic correlations was determined by comparing P-values, with a threshold of P < 0.05 considered significant. The sample sizes considered for the GWAS analysis of GERD included 71,522 cases and 261,079 controls. For psychiatric traits, the datasets ranged from 7016 to 170,756 samples, depending on the disorder. These sample sizes provide robust power for detecting genetic associations, ensuring reliability in the results.
For reproducibility, the study has made all GWAS summary statistics publicly accessible in accordance with relevant data-sharing policies. The data can be accessed through the GWAS Catalog for future validation and follow-up studies.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the GWAS summary statistics analyzed in this study are publicly available. Summary statistics for GERD MDD, BIP, ADHD, autism spectrum disorder (ASD), ANX, and schizophrenia (SCZ) were accessed from published genome-wide association studies in accordance with their respective data usage policies1,13,14,31,32,33,34,35,36.Complete details of the summary statistics, including sample sizes and study references, are provided in Supplementary Table 1.
Code availability
All custom code for the InsightGWAS model, including model architecture and training scripts, is available at https://github.com/ziangmeng/MDD-GERD-InsightGWAS, and the specific version of the code described in this paper has been archived in Zenodo38.
References
An, J., Gharahkhani, P., Law, M. H., Ong, J. S., Han, X. & Olsen, C. M. et al. Gastroesophageal reflux GWAS identifies risk loci that also associate with subsequent severe esophageal diseases. Nat. Commun. 10, 4219 (2019).
Coleman, H. G., Xie, S. H. & Lagergren, J. The epidemiology of esophageal adenocarcinoma. Gastroenterology 154, 390–405 (2018).
Cook, M. B., Corley, D. A., Murray, L. J., Liao, L. M., Kamangar, F., Ye, W. et al. Gastroesophageal reflux in relation to adenocarcinomas of the esophagus: a pooled analysis from the Barrett’s and Esophageal Adenocarcinoma Consortium (BEACON). PLoS One 9, e103508 (2014).
Mohammed, I., Cherkas, L. F., Riley, S. A., Spector, T. D. & Trudgill, N. J. Genetic influences in gastro-oesophageal reflux disease: a twin study. Gut 52, 1085–1089 (2003).
Cameron, A. J., Lagergren, J., Henriksson, C., Nyren, O., Locke, G. R. 3rd & Pedersen, N. L. Gastroesophageal reflux disease in monozygotic and dizygotic twins. Gastroenterology 122, 55–59 (2002).
Ding, H., Jiang, Y., Sun, Q., Song, Y., Dong, S., Xu, Q. et al. Integrating genetics and transcriptomics to characterize shared mechanisms in digestive diseases and psychiatric disorders. Commun. Biol. 8, 47 (2025).
Henning, M., Lindgen, K., Paul, D., Fuchs, C., Niecke, A. & Albus, C. et al. Association between anxiety and reflux symptoms in patients with gastroesophageal reflux disease: a prospective cohort study. Cureus. 16, e73391 (2024).
Ong, J. S., An, J., Han, X., Law, M. H. & Nandakumar, P. andMe Research t, et al. Multitrait genetic association analysis identifies 50 new risk loci for gastro-oesophageal reflux, seven new loci for Barrett’s oesophagus and provides insights into clinical heterogeneity in reflux diagnosis. Gut. 71, 1053–1061 (2022).
Nicholls, H. L., John, C. R., Watson, D. S., Munroe, P. B., Barnes, M. R. & Cabrera, C. P. Reaching the End-Game for GWAS: machine learning approaches for the prioritization of complex disease loci. Front. Genet. 11, 350 (2020).
Li Y., Wen J., Li G., Chen J., Sun Q., Liu W., et al. DeepGWAS: enhance GWAS signals for neuropsychiatric disorders via deep neural network. Res Sq. 2023.
Lakiotaki K., Papadovasilakis Z., Lagani V., Fafalios S., Charonyktakis P., Tsagris M., et al. Automated machine learning for genome wide association studies. Bioinformatics 39, btad545 (2023).
Mieth, B., Rozier, A., Rodriguez, J. A., Hohne, M. M. C., Gornitz, N. & Muller, K. R. DeepCOMBI: explainable artificial intelligence for the analysis and discovery in genome-wide association studies. NAR Genom. Bioinform 3, lqab065 (2021).
Donertas, H. M., Fabian, D. K., Partridge, V. alenzuelaM. F. & Thornton, L. JM. Common genetic associations between age-related diseases. Nat. Aging 1, 400–412 (2021).
Grove, J., Ripke, S., Als, T. D., Mattheisen, M., Walters, R. K., Won, H. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–44 (2019).
Zeng, Y., Cao, S. & Yang, H. The causal role of gastroesophageal reflux disease in anxiety disorders and depression: A bidirectional Mendelian randomization study. Front Psychiatry 14, 1135923 (2023).
Zhan, Z. Q., Huang, Z. M., Zhou, H. B., Xie, Z. X., Chen, Y. Z., Luo, Y. H. et al. Gastroesophageal reflux disease with 6 neurodegenerative and psychiatric disorders: Genetic correlations, causality, and potential molecular mechanisms. J. Psychiatr. Res 172, 244–253 (2024).
He, M., Wang, Q., Yao, D., Li, J. & Bai, G. Association Between Psychosocial Disorders and Gastroesophageal Reflux Disease: A Systematic Review and Meta-analysis. J. Neurogastroenterol. Motil. 28, 212–221 (2022).
Bertollo, A. G., Santos, C. F., Bagatini, M. D. & Ignacio, Z. M. Hypothalamus-pituitary-adrenal and gut-brain axes in biological interaction pathway of the depression. Front Neurosci. 19, 1541075 (2025).
Rubenstein, J. H., Morgenstern, H., McConell, D., Scheiman, J. M., Schoenfeld, P. & Appelman, H. et al. Associations of diabetes mellitus, insulin, leptin, and ghrelin with gastroesophageal reflux and Barrett’s esophagus. Gastroenterology 145, 1237–1244 (2013).
Murata, T., Asanuma, K., Ara, N., Iijima, K., Hatta, W., Hamada, S. et al. Leptin aggravates reflux esophagitis by increasing tissue levels of macrophage migration inhibitory factor in rats. Tohoku J. Exp. Med 245, 45–53 (2018).
Pardak P., Filip R., Wolinski J., Krzaczek M. Associations of obstructive sleep apnea, obestatin, leptin, and ghrelin with gastroesophageal reflux. J. Clin. Med. 10, 5195 (2021).
Guan, X., Shi, X., Li, X., Chang, B., Wang, Y., Li, D. et al. GLP-2 receptor in POMC neurons suppresses feeding behavior and gastric motility. Am. J. Physiol. Endocrinol. Metab. 303, E853–E864 (2012).
Guan, X. The CNS glucagon-like peptide-2 receptor in the control of energy balance and glucose homeostasis. Am. J. Physiol. Regul. Integr. Comp. Physiol. 307, R585–R596 (2014).
Rotondo, A., Amato, A., Baldassano, S., Lentini, L. & Mule, F. Gastric relaxation induced by glucagon-like peptide-2 in mice fed a high-fat diet or fasted. Peptides 32, 1587–1592 (2011).
Wallin, C., Grupcev, G., Emas, S., Theodorsson, E. & Hellstrom, P. M. Release of somatostatin, neurotensin and vasoactive intestinal peptide upon inhibition of gastric acid secretion by duodenal acid and hyperosmolal solutions in the conscious rat. Acta Physiol. Scand. 154, 193–203 (1995).
Nassar, C. F., Abdallah, L. E., Barada, K. A., Atweh, S. F. & Saade, N. E. Effects of intravenous vasoactive intestinal peptide injection on jejunal alanine absorption and gastric acid secretion in rats. Regul. Pept. 55, 261–267 (1995).
Lin, D. S., Zhang, C. Y., Li, L., Ye, G. H., Jiang, L. P. & Jin, Q. Circ_ROBO2/miR-149 axis promotes the proliferation and migration of human aortic smooth muscle cells by activating NF-kappaB signaling. Cytogenet. Genome Res. 161, 414–424 (2021).
Je, E. M., Gwak, M., Oh, H., Choi, M. R., Choi, Y. J., Lee, S. H. et al. Frameshift mutations of axon guidance genes ROBO1 and ROBO2 in gastric and colorectal cancers with microsatellite instability. Pathology 45, 645–50 (2013).
Dave, U., Thursz, M. R., Ebrahim, H. Y., Burke, M. M., Townsend, E. R. & Walker, M. M. Distribution of laminins in the basement membranes of the upper gastrointestinal tract and Barrett’s oesophagus. J. Pathol. 202, 299–304 (2004).
Virtanen, I., Tani, T., Bäck, N., Häppölä, O., Laitinen, L., Kiviluoto, T. et al. Differential expression of laminin chains and their integrin receptors in human gastric mucosa. Am. J. Pathol. 147, 1123–32 (1995).
Howard, D. M., Adams, M. J., Clarke, T. K., Hafferty, J. D., Gibson, J., Shirali, M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–52 (2019).
Wray, N. R., Ripke, S., Mattheisen, M., Trzaskowski, M., Byrne, E. M., Abdellaoui, A. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–81 (2018).
Mullins, N., Forstner, A. J., O’Connell, K. S., Coombes, B., Coleman, J. R. I., Qiao, Z. et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat. Genet. 53, 817–29 (2021).
Demontis, D., Walters, G. B., Athanasiadis, G., Walters, R., Therrien, K., Nielsen, T. T. et al. Genome-wide analyses of ADHD identify 27 risk loci, refine the genetic architecture and implicate several cognitive domains. Nat. Genet. 55, 198–208 (2023).
Otowa, T., Hek, K., Lee, M., Byrne, E. M., Mirza, S. S., Nivard, M. G. et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol. Psychiatry 21, 1391–9 (2016).
Trubetskoy, V., Pardinas, A. F., Qi, T., Panagiotaropoulou, G., Awasthi, S., Bigdeli, T. B. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–8 (2022).
Bulik-Sullivan, B. K., Loh, P. R., Finucane, H. K., Ripke, S., Yang, J., Schizophrenia Working Group of the Psychiatric Genomics, C. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–5 (2015).
Ziangmeng. Ziangmeng/MDD-GERD-InsightGWAS-Model: Frist release of MDD-GERD-InsightGWAS-Model (v1.0) (2025). https://doi.org/10.5281/zenodo.17457573
Acknowledgements
We thank the investigators and participants of the GERD GWAS studies and the corresponding consortia for making their data publicly available. This work was supported by grants from the Special Funds of the Taishan Scholar Project, China (tsqn202211224), the National Natural Science Foundation of China (32270661), and the Excellent Youth Science Fund Project (Overseas) of Shandong, China (2023HWYQ-082). Additional support was provided by the Huzhou Science and Technology Program for Population Health (Key Medical Project) and the Key Research Project of the Joint Science and Technology Plan, co-sponsored by the State Administration of Traditional Chinese Medicine and the Zhejiang Provincial Administration of Traditional Chinese Medicine (GZY-ZJ-KJ-23094).
Author information
Authors and Affiliations
Contributions
Y.W. and Z.M. contributed equally to this work. X.C., N.Z., and Y.W. conceived and supervised the study. Z.M. developed and implemented the deep learning framework. Y.J., Y.W., X.W., H.D., and B.P. contributed to data preprocessing, annotation, and quality control. Y.S., M.G., G.Z., and N.Z. assisted with interpretation of results and literature review. Y.W., X.C., and Z.M. wrote the manuscript with input from all authors. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Zhihao Ding and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Eirini Marouli, Aylin Bircan and George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wei, Y., Meng, Z., Wang, X. et al. Transformer-based InsightGWAS improves GERD genetic discovery via pretraining on GWAS for major depressive disorder. Commun Biol 9, 2 (2026). https://doi.org/10.1038/s42003-025-09177-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09177-3