Abstract
Zoonotic pathogens are increasingly emerging in previously low-risk regions with the accelerating process of globalization. Understanding the genomic evolution and spatial dissemination of these pathogens is essential for the rapid response and targeted control of future outbreaks. Brucellosis, caused by Brucella melitensis, is one of the global neglected zoonotic diseases now expanding beyond its traditional endemic zones. Here, we present the comprehensive genomic analysis of B. melitensis isolates from an emerging hotspot of human brucellosis in Southwest China. A total of 103 isolates were collected from 10 cities across Yunnan in 2019-2022. Phylogenetic analyses of the 850 global genomes revealed the coexistence of a dominant lineage (Lineage 1.1) and a novel lineage (Lineage 1.3) in non-epidemic regions in Southwest China. Importantly, we identified lineage-specific patterns of genomic differentiation, indicating the great variability of evolutionary dynamics among the co-circulating lineages. Our results showed higher rates of gene gain and loss events for Lineage 1.1, and a decrease in gene similarity along the gradient of spatial distance for Lineage 1.3. Additionally, spatiotemporal phylogeographic inference indicated that Lineage 1.1 has undergone recent interprovincial spread within China, while Lineage 1.3 remains geographically restricted to Yunnan. Our findings highlight the distinct evolutionary and transmission dynamics of B. melitensis in a non-traditional epidemic region. We provide critical insights into the value of genome-informed lineage-specific surveillance and targeted control strategies in addressing the emerging threat of neglected zoonotic diseases in the context of rapid ecological and social change.
Similar content being viewed by others
Introduction
Zoonotic diseases continue to pose challenges to human health and socioeconomic development. Emerging evidence is increasingly highlighting the surge of many neglected zoonotic diseases and their under-recognized global threats1. Brucellosis, caused by Gram-negative coccobacilli of the genus Brucella, remains one of the leading neglected zoonotic diseases2,3. The substantial disease burden of human brucellosis has been identified in regions with extensive livestock farming and insufficient public health infrastructure4. Assessing transmission dynamics is fundamental for alleviating disease burden in these high-risk regions.
The transmission dynamics of brucellosis may vary considerably across regions. This is largely attributable to the multifaceted drivers and interconnected mechanisms of disease dynamics. Of the many Brucella species, human infections by Brucella melitensis found in goats and sheep are widely recorded worldwide, accounting for up to 90% of all the reported cases5. Human infections occur primarily through occupational exposure, putting individuals such as abattoir workers, veterinarians, laboratory technicians, farmers, and livestock producers at increased risk6. Additionally, human infections are often linked to international travel or the consumption of imported foods. This suggests that the accelerating process of globalization has the potential to exacerbate the global transmission of the disease7.
Brucellosis has re-emerged as a public health threat in traditional epidemic areas in the Northern provinces of China since the mid-1990s8. Despite various control programs, the prevalence of human brucellosis has significantly increased in recent years9,10. Importantly, the spatial expansion of human brucellosis has demonstrated a shift from pastoral areas in Northern provinces to the rural and urban regions across the country11. The annual average growth rate of brucellosis incidence in southern provinces reached 31.5% in 2015–2021, which is markedly higher than that observed in the north12. Consistent with this, a marked increase in human brucellosis infections has been recorded in several low-risk Southern provinces such as Yunnan, Guangdong, and Guangxi10,13. This upsurge of human infections among traditionally non-endemic areas suggests latent brucellosis risks and underscores the need for region-specific surveillance and control strategies.
Addressing the threat of neglected zoonotic diseases requires a holistic approach, integrating epidemiological and genetic features of the human and animal isolates in the One Health framework14,15. However, the lack of surveillance may have led to a substantial under-reporting of human infections. Relatedly, the limited sampling effort may have complicated the inference of the spatiotemporal diffusions of the disease. It is noted that none of the existing B. melitensis genomes originating from China provided in the public repositories were isolated from Yunnan province (Supplementary Data 1). Given the emerging challenge to public health and socioeconomic sustainability, uncovering the genomic and epidemiological characteristics of Brucella strains circulating in Yunnan is essential for understanding transmission dynamics.
Brucellosis control efforts in China have historically prioritized the high-prevalence northern pastoral regions, yet the disease’s southward expansion poses unique epidemiological challenges that demand tailored surveillance approaches. Among southern provinces, Yunnan documented the highest human infections, with an approximately 90% surge in human incidence (from 0.78 to 1.48 cases per 100,000 population) between 2020 and 202112. This epidemiological shift occurs against a backdrop of severe genomic surveillance gaps where southern regions remain dramatically underrepresented. Our study addresses this disparity by establishing the genomic landscape for B. melitensis in Yunnan, capturing lineage-specific adaptation patterns and transmission networks.
The aim of this study was to investigate the spatiotemporal diffusion of brucellosis in non-traditional epidemic regions in China. To this end, we sampled and sequenced a total of 103 B. melitensis strains from 10 different cities in Yunnan Province. By leveraging these genomic data, we investigated genomic differentiation and spatial structuring of brucellosis on the global and finer spatial scales. Integrating the genomic, phylogenetic, and epidemiological analyses illustrated how evolutionary trajectory and spatial dynamics amplified brucellosis threats to humans. The insights gained from this study will inform targeted interventions of the disease in non-traditional epidemic regions, with broader implications for the global mitigation of neglected zoonotic diseases.
Methods
Strain isolation
B. melitensis isolates were collected from confirmed human cases in 2019–2022 across 10 cities in Yunnan Province, Southwest China (Supplementary Data 2). The sampling effort was aligned with the spatiotemporal diffusion of human infections, covering a broad range of cities in Yunnan Province. Specifically, 21 isolates were collected in 2019 and 2020, followed by 31 isolates in 2021 and 34 in 2022. Our sampling involved 27 isolates from Kunming, 16 each from Honghe and Qujing, with the remaining strains collected from Dali, Yuxi, Chuxiong, Zhaotong, Baoshan, Lijiang and Wenshan. For each clinical isolate, the sampling date and the residential address of the infected patient were recorded. Temporal and geographic distances between isolates were subsequently evaluated to further characterize the spatiotemporal pattern of genomic differentiation (see “Model inference of genomic differentiation”).
Genome sequencing
For whole-genome sequencing, we extracted DNA using the High Pure PCR Template Preparation Kit. Libraries were subsequently prepared and 2 × 150 bp paired-end sequencing was carried out on the Illumina NextSeq platform. The raw sequencing reads were processed through the following steps. Initially, read quality was assessed by FastQC v0.12.1 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc). Adaptors and low-quality bases were trimmed using fastp v0.23.416. Genome assemblies were generated using a de novo approach with Unicycler v0.5.0.1217. The quality of the resulting assemblies was subsequently assessed using QUAST v5.0.218. All assemblies met the following quality control thresholds: number of contigs <300, N50 > 50,000, and average coverage >30×.
Pan-genome analyses and openness estimation
To construct the pan-genome, we collected 747 publicly available B. melitensis genomes from the National Center for Biotechnology Information’s (NCBI) Genome Dataset (https://www.ncbi.nlm.nih.gov/datasets/genome/?taxon=29459) as of August 30, 2024, and integrated them with the B. melitensis genomes sequenced in this study. Of these, we identified 118 genomes that originated from China; notably, none were isolated from Yunnan Province. Our isolates provide the genomic characterization of B. melitensis in Yunnan Province, filling a critical surveillance gap for Southwest China.
Pan-genome analyses are used to study the genetic diversity of B. melitensis isolates. We started with gene prediction of the assembled genomes using Prokka v1.14.619, and subsequently performed core-genome identification and gene family clustering using Panaroo v1.5.220. To facilitate comparison with previous studies, we additionally ran the same workflow using Roary v3.13.021. Genes present in more than 95% of all genomes were defined as core genes, while the others were classified as accessory genes. Of the accessory genes, those found in 15–95% of genomes were categorized as soft genes, and those present in fewer than 15% of genomes were rare genes22,23. Functional annotation of genes was performed based on the amino acid sequences using eggNOG v.5.024.
In addition, the openness of the pan-genome was evaluated by quantifying the number of novel genes identified as the sample size increased. For each evolutionary lineage, we computed the pan-genome openness using a presence/absence matrix generated by Panaroo21. In this matrix, rows correspond to genomes and columns to genes, with binary entries (0/1) indicating the absence or presence of a gene. First, genomes belonging to the targeted lineage were selected from the full dataset. Then, for a series of increasing sample sizes n (e.g., n = 5, 10, …, N, where N is the total number of genomes for the lineage), we randomly sampled n genomes and calculated two quantities: the core-genome size, defined as the number of genes present in all n genomes, and the pan-genome size, defined as the number of genes present in at least one of the n genomes. This random sampling was repeated (100 replicates) for each n, and the median pan-genome size P(n) was recorded.
The openness of the pan-genome was assessed by calculating the number of new genes discovered when the sample size increases by one, defined as
A regression analysis was then performed on the log-transformed data, fitting the model
where b quantifies the rate at which new genes are acquired with the addition of genomes. The slope of the regression line b < 1 indicates that as more genomes are analyzed, the number of new genes decreases slowly, suggesting the pan-genome is open25. Conversely, if the slope b > 1, the number of new genes reaches a plateau as more genomes are added, indicating that the pan-genome is closed.
Assessment of genome similarity
We assessed the genomic similarity between isolates using the average nucleotide identity (ANI) and gene content similarity (GCS). ANI is used to evaluate overall nucleotide sequence similarity between genome pairs. ANI was computed based on pairwise comparisons of genome sequences using the R package ape v5.826. GCS was calculated as the proportion of shared genes relative to the total number of unique genes between two genomes, providing a measure of gene content variation (GCV)27. Each genome was represented as a binary vector based on a presence–absence matrix, with 1 indicating gene presence and 0 indicating absence. Pairwise GCS between genomes was subsequently estimated using the Jaccard similarity coefficient28,29.
Phylogeny and lineage profiling
Phylogenetic analysis included isolates obtained from NCBI Genome Dataset and those sequenced in our study. For SNP detection, B. melitensis 16 M (ASM712v1) was selected as the reference. Each strain’s sequence was then mapped to B. melitensis 16 M using Snippy 4.6.0 (https://github.com/tseemann/snippy). Recombinant regions were identified and removed using Gubbins 2.2.030, generating the dataset for the subsequent multiple sequence alignments and SNP detection. Using this data, a phylogenetic tree was constructed using maximum likelihood analysis in RAxML v8.2.1331, applying the GTRGAMMA model of rate heterogeneity and optimizing substitution rates. Based on tree topology, clustering was conducted using the Bayesian Analysis of Population Structure32. Specifically, initial clustering was performed with multiple parameters to achieve high-resolution clusters (Supplementary Fig. 1). Clusters with low support were subsequently merged using the bootstrap method, resulting in all lineages with bootstrap values greater than 80 (Supplementary Fig. 2). The major clusters and branches were labeled following a structured hierarchical naming system. While multiple studies have reconstructed B. melitensis phylogenies, there still lacks a unified lineage nomenclature system. Current classifications employ inconsistent tesminology—using geographic labels, numeric codes, or study-specific terms—creating substantial challenges for cross-study comparison33,34,35. We defined major lineages in the numerical order. Lineage 1 was further divided into Lineage 1.1, 1.2, and 1.3. Similarly, Lineages 2 and 3 were divided into sub-lineages. To evaluate the geographic distribution of lineage diversity, samples with associated geographic metadata (n = 811) were grouped by area. Lineage diversity was quantified using Shannon entropy calculated from the lineage distribution in each area.
Time-scaled phylogeny
We performed phylogeographical analyses to characterize the spatiotemporal transmission of B. melitensis strains in Asia. The phylogeographic distribution of taxa was inferred using SNP-based whole-genome alignments of B. melitensis strains in BEAST v1.10.436. More specifically, B. melitensis strains with the known isolation dates were used to calibrate the molecular clock and date phylogenetic tree. The optimal evolutionary model was selected by evaluating 88 candidate substitution models using Jmodeltest2 v2.1.1037. Three independent runs of the model were carried out using a strict clock, with a chain of 200,000,000 generations and a recording rate of every 1000 generations. The convergence of the MCMC topology and parameters was estimated in the Tracer v1.6 program. All effective sample size values were >20038.
Spatial prevalence and gene content
To assess the association between geographic spread and GCV, spatial prevalence was defined as the number of counties in which each lineage was detected per year39. For each isolate, total gene content was defined as the number of genes present, while soft gene content and rare gene content were defined as the proportion of genes classified as soft or rare, respectively. These gene content metrics were then aggregated by lineage and year to obtain their annual means. Linear regression models were used to evaluate the lineage-specific relationships between spatial prevalence and the annual mean of gene content. To further explore functional correlates of spatial prevalence27, pathway-level gene content was defined as the proportion of pathway-associated genes present in each genome, based on GO pathway annotations derived from eggNOG results. Linear regressions were then performed to assess associations between pathway-level gene content and spatial prevalence across lineages. Statistical significance of enrichment was tested using a permutation test with Benjamini-Hochberg correction (adjusted p < 0.05).
Estimation of gene gain and loss rates across evolutionary time
To assess the association between gene gain/ loss events and evolutionary history, we applied a modified version of the Panstripe40 method. Gene presence–absence matrices were derived from Panaroo annotations. Ancestral gene content states were reconstructed using a parsimony-based algorithm, allowing quantification of the expected number of gene gain (0\(\to\)1) and loss (1\(\to\)0) events along each internal branch. These expectations were aggregated across all genes within each category (i.e., the pan, soft, and rare genes) to yield branch-level summaries of gene turnover. Evolutionary time along each internal branch was extracted from a time-scaled phylogeny reconstructed using BEAST, and the branch depth (distance from root to parent node) was also recorded to account for phylogenetic structure and potential biases at deeper nodes.
We then constructed generalized linear models with a quasi-Poisson distribution to account for overdispersion in event counts. Two separate models were fitted for gene gain and gene loss events, using evolutionary time as the main explanatory variable. Branch depth was included as a covariate to adjust for potential confounding due to phylogenetic depth. Lineage identity was modeled as an interaction term with evolutionary time to allow for lineage-specific effects. The models were established using the following form:
where \({Y}_{{ij}}\) is the expected number of gene gain or loss events for branch \(i\) in lineage \(j\), \({T}_{{ij}}\) represents evolutionary time, \({D}_{{ij}}\) is branch depth, and \({L}_{j}\) is an indicator for lineage. The interaction term \({T}_{{ij}}\times {L}_{j}\) allows the rate of change in gene events over time to vary by lineage.
Separate models were fitted for each gene category (i.e., the pan, soft, and rare genes). Model coefficients were used to estimate the lineage-specific marginal effects of evolutionary time on gene gain and loss. Statistical significance was assessed using confidence intervals derived from the estimated standard errors of the marginal effects. This framework allowed us to quantify and compare the tempo of genome evolution across distinct gene categories and lineages.
Model inference of genomic differentiation with spatial and climatic factors
To evaluate the association between genomic differentiation and both geospatial and environmental factors, we first obtained monthly climatic data, i.e., mean temperature, dewpoint temperature, surface pressure, and total precipitation, provided by the fifth-generation reanalysis (ERA5)41 from the European Center for Medium-Range Weather Forecasts. For each B. melitensis isolate, we extracted local climate conditions using its geographic coordinates and sampling time. Annual averages of these climate factors were further calculated to minimize the influence of seasonal fluctuations. With these datasets, we subsequently evaluated the pairwise differences of climate conditions between isolates.
We established generalized additive models (GAMs) to investigate how intra-lineage genomic differentiation is associated with geospatial and climatic factors. The two genomic similarity metrics, i.e., ANI and GCS, were used as the response variables. Pairwise differences in climate conditions, together with geographic distance and sampling time interval, were included as potential factors in the GAMs. The GAMs were fitted separately for each lineage. The models were specified as:
where Genomic similarity represents either ANI or GCS between isolate pairs; s(geo_distance, k = 5) and 297 s(time_interval, k = 5) denote smooth functions capturing potential nonlinear 298 relationships between genomic differentiation and geographic or temporal distance; and Δpressure, Δdewpoint_temp, Δtemp and Δprecipitation represent pairwise differences in surface pressure, dewpoint temperature, mean temperature, and total precipitation, respectively.
Models were fitted using the mgcv package (version 1.9-1)42 in R v4.1.2. Diagnostic evaluations were conducted, including assessment of residual distributions and goodness-of-fit metrics (e.g., adjusted R2, explained deviance), to ensure model adequacy and validity. The significance of smooth and linear predictors was tested via approximate F tests, with corresponding p values reported.
Phylogeographical analyses
Based on the time-scaled phylogeny, we investigated the spatial structure of B. melitensis strains at both global and finer geographic scales. To this end, we extracted the time to the most recent common ancestor (tMRCA) for all isolate pairs. For the investigation of global spatial structure, pairs were categorized into four tMRCA groups (0–10 years, 10–50 years, 50–100 years, and >100 years). Geographical relationship of the isolate pairs was classified as local (within province), regional (within country), national (between countries), or continental (between continents). To evaluate the finer-scale structuring of Lineage 1.1 and 1.3 within China, isolate pairs were stratified by tMRCA and binned into geographic distance intervals ranging from <100 km to >3000 km.
We further evaluated the relative risks (RRs) of transmission across spatial scales43,44,45. RRs were calculated by comparing the probability that a given isolate pair was observed within each geographic category, relative to a reference category (regional or specified distance bands). Confidence intervals for RRs were estimated using bootstrapping.
Global transmission dynamics of Lineage 1
To investigate global transmission dynamics of B. melitensis Lineage 1, we calculated the probability (P) that a pair of isolates, sampled within a specified temporal interval, shared their most recent common ancestor (MRCA) within defined time intervals: 0–10 years, 10–50 years, 50–100 years, and >100 years. Spatial categories were defined as local (within the same province), regional (within the same country but different provinces), national (between different countries), and continental (between different continents).
The probability (P) was calculated using the following formula:
Relative risk (RR) was then computed by comparing probabilities across categories using the following formula:
The reference category was set to regional transmission. Confidence intervals for the RR estimates were obtained through bootstrapping (1000 iterations), sampling pairs of isolates with replacement and recalculating RR each iteration. The 95% confidence intervals were determined from the 2.5 and 97.5 percentiles of the bootstrapped distribution.
Local transmission dynamics of Lineage 1
For finer-scale transmission dynamics within China (specifically Lineages 1.1 and 1.3), pairs of isolates were grouped based on geographic distance intervals: <100 km, 100–200 km, 200–500 km, 500–1000 km (reference), 1000–2000 km, and >3000 km. We calculated probabilities (P) similarly, defined as:
Since the maximum geographic distance among samples within Yunnan Province was 646 km, we defined the 500–1000 km interval as the reference category, delineating provincial from extra-provincial transmissions. RR for each distance interval compared to the reference (500–1000 km) was calculated by:
Confidence intervals for RRs were also calculated via bootstrapping with 1000 iterations, resampling isolate pairs and recalculating probabilities and relative risks for each distance interval.
Incorporation of livestock trade data into transmission risk models
To evaluate the hypothesis that interprovincial livestock trade networks significantly contributed to the dissemination of Brucella in Yunnan Province, we extended our epidemiological models to integrate province-level livestock production and trade data. This approach allows us to move beyond a static, geographic view of risk to a dynamic one that accounts for the pathways through which infection is likely to be introduced into new regions.
Human brucellosis incidence data, based on onset date from January 2004 to December 2020, were obtained from the National Notifiable Disease Surveillance System via the Chinese Public Health Science Data Center. As a proxy for the movement of live animals—direct data for which is scarce—we utilized China’s interprovincial physical food trade dataset46. This dataset provides estimates of pairwise trade flows of cattle and sheep meat between provinces. We reasoned that the trade of meat products reflects the underlying movement of live animals from production areas to markets and slaughterhouses, which is a known critical risk factor for Brucella transmission. For each province and year, we aggregated these data to calculate the total inflow and outflow of cattle and sheep meat.
We employed GAMs using the bam function in the R package mgcv to model the log-transformed incidence of human brucellosis in province \({j}\), denoted as \(\log \left({{Inc}}_{j}\right)\). The baseline model was specified as:
where \({P}_{j}\) represents cattle and sheep production, s(\({\text{Province}}_{j}\), bs = “re”) and s(Year, bs = “re”) denote province- and year-specific random effects to account for unmeasured spatial and temporal heterogeneity, and s(Month, bs = “cr”) is a cyclic regression spline capturing seasonal variation.
To quantify the additional explanatory power of trade, we constructed an extended model:
where \({I}_{j}\) denotes the total inflow of cattle and sheep meat into province \(j\), and \({O}_{j}\) denotes the corresponding total outflow. This model specification is crucial as it allows the functional relationship between trade flows and disease incidence to vary non-linearly and uniquely for each province (via the by argument), capturing the heterogeneous role of trade across diverse epidemiological contexts within China. Both models assumed Gaussian-distributed errors on the log incidence scale. Model performance was compared using the Akaike information criterion and explained deviance.
Statistical analysis of occupational data
To assess the association between B. melitensis lineages and the occupation of infected individuals, we categorized cases into two groups, i.e., farmer and non-Farmer. Due to the sample size in some categories, a Fisher’s exact test was used to compare the occupational distributions between Lineage 1.1 and Lineage 1.3. The p-value of less than 0.05 was considered statistically significant.
Statistics and reproducibility
All statistical analyses were conducted in R (v4.1.2) unless otherwise stated. Analyses of pan-genome openness, gene gain and loss dynamics, and lineage-specific genomic differentiation were implemented using Roary (v3.13.0), Panaroo (v1.5.2), and custom R scripts. For evaluating the association between genomic differentiation and spatioclimatic factors, GAMs were fitted using the mgcv package (v1.9-1). Geographic distance and sampling interval were modeled as smooth terms, and differences in climatic variables (mean temperature, dewpoint temperature, surface pressure, and total precipitation) were modeled as linear predictors. Significance of predictors was assessed by approximate F tests (p < 0.05). Lineage-specific gene gain and loss rates were estimated using quasi-Poisson generalized linear models, with evolutionary time as the fixed effect and lineage as the interaction term. Confidence intervals of model coefficients were used to assess statistical significance. For categorical data (e.g., occupational distributions between lineages), Fisher’s exact tests were conducted, and p < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Global phylogeny reveals a novel B. melitensis lineage
A total of 103 B. melitensis isolates were collected from confirmed human brucellosis cases in Yunnan Province, Southwest China, from 2019 to 2022. Together with 747 publicly available B. melitensis genomes, we established a global dataset to investigate the phylogenetic and genomic structure of the pathogen. The whole-genome SNP phylogenetic analysis identified three lineages and eight sub-lineages of the 850 B. melitensis isolates (Fig. 1A). Spatially explicit distribution of the isolates suggests that these lineages were concentrated in the Eurasian regions (Fig. 1B, Supplementary Fig. 3A). Isolates in Lineage 1 were primarily distributed in Asia (90.26%), while the majority of the isolates from Lineage 3 (95.38%) were predominantly from European countries. Although Lineage 2 was associated with diverse origins, approximately half of its isolates (53.66%) originated from African countries (Supplementary Fig. 3B). Additionally, estimates of Shannon entropy indicated that lineage composition was highly diverse across geographic areas (Fig. 1C). Although no overall correlation between lineage diversity and sample size (p = 0.719, Supplementary Fig. 3C), the higher diversity in North America and Western Europe may reflect multiple independent introductions.
A Whole-genome SNP-based phylogenetic tree of the global B. melitensis isolates, comprising 747 publicly available genomes and 103 isolates sequenced from Yunnan Province, China. The tree was plotted using Grafen’s transformation, showing topological relationships. The isolates in major lineages and sub-lineages are distinguished by colors. Outer rings indicate the geographic region (continent and area) and date of sampling. Isolates in this study were marked. B Geographic distribution of B. melitensis isolates included in this study. Countries are shaded according to the total number of isolates collected. The circles represent the proportion of lineages in each area, with circle size proportional to the total number of isolates. Colors of the circles indicate different lineages. C Relationship between sample size and Shannon entropy of lineage distribution across geographic areas. Areas with above-average diversity are shown in yellow, and others in gray. D Power-law regression analysis based on the association between the number of genomes and new genes on the log scale, with shaded areas representing the 95% confidence intervals. E Relationship between pairwise evolutionary time and gene family copy number variation. The median estimates of the regression (50th percentile) and the 95% confidence interval were marked.
Finer-scale spatial distribution of the isolates also revealed sub-lineage-specific spatial clusters (Supplementary Fig. 3D). Our results showed that East Asian isolates were predominantly clustered in Lineage 1.1, while those from West Asia were concentrated in Lineage 1.2. Moreover, Lineage 2.2 exclusively comprised isolates from African countries, while isolates from Lineage 2.3 were predominantly from Central America and Africa. Additionally, isolates in Lineage 3.1 were mainly found in Southern and parts of Western Europe, while those in Lineage 3.2 were distributed across diverse regions in Europe and North Africa. This finding suggests that geographic isolation may have played a major role in shaping the spatial distribution of B. melitensis lineages.
Investigation using pan-genome presence–absence variation (PAV) revealed lineage-specific clustering of accessory genome, indicating the robust genomic differentiation among the lineages (Supplementary Fig. 4). We subsequently performed power-law regression analysis to examine the pan-genome openness within each lineage. We identified that all three lineages were characterized by closed pangenomes (Fig. 1D). To further investigate evolutionary processes, we recovered gene family copy number information from the Panaroo presence/absence matrix and observed increasing differences between sample pairs over evolutionary time (Fig. 1E). Further assessment of the contribution of homologous recombination to genetic variation indicated that recombination rates estimated by Gubbins were consistently low across all lineages (Supplementary Fig. 5). Such low recombination rates support the hypothesis that B. melitensis evolves largely through clonal expansion rather than frequent horizontal gene transfer.
Note that the Yunnan isolates cover a broad range of endemic areas within the province (Fig. 2A, B). In view of the consistency between the distribution of sequenced isolates and that of the reported human cases (Supplementary Note 1, Supplementary Fig. 6), the isolates are highly representative of B. melitensis across the province. We found that these isolates belonged to Lineage 1 (Fig. 2C), while further diverging into three sub-lineages: Lineage 1.1 (n = 79), Lineage 1.2 (n = 5), and a novel Lineage 1.3 (n = 19). It is noted that Lineage 1.3 formed a distinct monophyletic clade, which is unique to Yunnan. Temporal dynamics showed that Lineage 1.1 remained the dominant lineage over the years; Lineage 1.3 accounted for ~50% of isolates in 2019 but gradually declined to ~20% in 2020, and less than 15% in 2021–2022 (Fig. 2D). Further analysis revealed that the number of Lineage 1.1 isolates increased over the study period, while Lineage 1.2 and 1.3 showed no clear temporal trend (Supplementary Fig. 7). This indicates that the shift in sub-lineage proportions was driven primarily by the expansion of Lineage 1.1.
A Monthly incidence of human brucellosis and the number of B. melitensis isolates sequenced in this study in Yunnan from 2019 to 2022. The inset shows a significant positive correlation between the number of sequenced isolates and the total number of cases, and thus the minimal sampling bias. B Geographic distribution of the sequenced Yunnan B. melitensis isolates. Lineage of isolates and density of human cases were distinguished by colors. C Phylogenetic tree of the Yunnan B. melitensis isolates. Sub-lineages of the isolates were distinguished by color. The tree was plotted using Grafen’s transformation, showing topological relationships. D The annual variation of the relative proportions of B. melitensis lineages from 2019 to 2022.
We further investigated the epidemiological characteristics of cases, identifying a striking divergence of the patient occupation between Lineage 1.1 and 1.3. Although the sample size for Lineage 1.3 is limited, its cases were almost exclusively found in farmers, whereas Lineage 1.1 infected individuals across a broader range of occupations, including farmers, herders, office staff, students, and retirees. A Fisher’s exact test confirmed that this occupational distribution was significantly different between the two lineages (p = 0.0477).
Spatial prevalence correlates with GCV across lineages
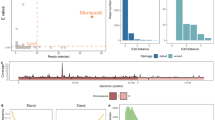
We evaluated the relationship between gene content composition and the annual number of detected counties for each sub-lineage to investigate how genomic features may vary along geographic expansion. We found that the total gene content showed no significant association with geographic expansion (Fig. 3A). In contrast, accessory gene content changed with geographic expansion and showed comparable trends across Lineage 1.1, 1.2, and 1.3. Specifically, higher spatial prevalence was significantly associated with a reduction in soft gene content but an increase in rare gene content (Fig. 3B, C). We further validated this genomic-geographic relationship by examining the rates of gene gain and loss across internal branches over evolutionary history. We revealed a significant increase in the rate of gene gain and loss events per year of evolutionary time of Lineage 1.1, suggesting highly dynamic pan-genomic patterns (Fig. 3D). Consistent with the genome level dynamics, further analyses showed that in Lineage 1.1, evolutionary time was associated with the rates of both gene gain and loss events for soft genes, but only with the rate of gene gain for rare genes (Fig. 3E). In contrast, we identified no evidence of a significant relationship between gene gain/loss rates and evolutionary time for Lineage 1.2 (Supplementary Fig. 8) or Lineage 1.3 (Fig. 3D, E). These results suggest that the gene content of Lineage 1.1 may have become increasingly individualized over time and that the accumulation of rare genes may have contributed to its environmental adaptability.
Lineage-specific relationship between the number of counties and the A total, B soft, and C rare gene content. Error bars indicate the 95% confidence intervals of gene content for each lineage in the sampling year. Shaded areas represent 95% confidence intervals of the linear regression across all lineages. D Estimated gene gain and loss rates (expected event counts per unit evolutionary time) at the pan-genome level, stratified by lineage. Error bars indicate 95% confidence intervals. E Gene gain and loss rates for soft and rare genes in Lineage 1.1 and Lineage 1.3. Error bars indicate 95% confidence intervals. F Enriched GO pathways whose gene content is significantly associated with spatial prevalence. Each node represents a pathway; edges denote Jaccard similarity based on shared genes. Node size indicates effect size, and color reflects the direction and magnitude of association. Modules were identified based on shared gene content.
Our findings from the pathway-level analysis supported the association between functional gene content and spatial prevalence. Notably, several stress response pathways, tRNA-related pathways, and ion-binding-associated pathways increase significantly with the number of detected counties (Fig. 3F, Supplementary Table 1). These enrichments suggest that genes involved in environmental sensing and response mechanisms may facilitate adaptation to varied local conditions, supporting spatial expansion of specific lineages through ecological flexibility.
Distinct geographic and climatic constraints of Brucella lineages
In view of the differences in pan-genome dynamics and adaptive signatures between Lineage 1.1 and 1.3, we further sought to investigate how genomic variation is associated with geographic distance, sampling time interval and climatic factors. We observed a significant decline of core-genome ANI and GCS with increasing geographic distance for Lineage 1.3 (Fig. 4A, B). This distance–decay relationship revealed that isolates collected from geographically distant regions share fewer genes and exhibit greater nucleotide divergence, with an average reduction of 0.6% in ANI and ~2.1% in GCS per 100 km. However, we detected no evidence of significant associations between sampling dates and genomic variation for both lineages (Fig. 4A, B). Importantly, environmental variables, including annual temperature, precipitation, surface pressure, and dewpoint temperature, may have had a limited effect on genomic dynamics (Fig. 4C, D). These results indicate that spatial isolation may have dominated the genomic diversification of Lineage 1.3 by restricting geographic dispersal but promoting localized evolution of the strains.
A, B Relationships between pairwise genomic similarity (ANI and gene content similarity) and geographic distance and time interval of sampling across isolates. Lines represent smoothed GAM fits, with shaded areas indicating 95% confidence intervals. C, D Estimated effects of climate factor differences on ANI and gene content similarity stratified by lineage. Each point represents the effect size, and error bars indicate a 95% confidence interval. Asterisks indicate statistically significant associations (p < 0.05).
In contrast, the genomic similarity among isolates in Lineage 1.1 varied minimally across geographic distances and the sampling time interval, indicating frequent gene flow or a more recent common ancestry of the isolates across Yunnan (Fig. 4A, B). However, we detected significant correlations between climatic differences and genomic similarity for strains within Lineage 1.1. Specifically, pairwise differences in annual mean temperature were significantly associated with ANI divergence, while differences in total annual precipitation were significantly associated with both ANI and GCS (Fig. 4C, D). These findings suggest that climatic factors may have contributed to genomic differentiation within Lineage 1.1. In contrast, the same analysis for Lineage 1.2 revealed no significant associations between the tested factors and either ANI or GCS (Supplementary Table 2).
Localized dissemination of Lineage 1.3 in Yunnan
Inference of the global dynamics of Lineage 1 identified the great variation of B. melitensis transmission risk across spatiotemporal scales. The recent transmissions (MRCA < 10 years) were exclusively restricted to provincial and national boundaries, with a markedly elevated relative risk of local transmissions (RR = 28.17, 95% CI: 12.6–81.0) as compared to those on broader spatial scales. Consistent with this, local transmissions remained dominant (RR = 3.41, 95% CI: 2.44–4.86) in the intermediate term (10–50 years), though larger-scale connections emerged with relatively lower risks (national: RR = 0.08, 95% CI: 0.03–0.15; continental: RR = 0.004, 95% CI: 0.002–0.011). Note that local and long-distance transmissions collectively shaped the spatial structure of global B. melitensis around 50–100 years ago. Relative risks of local transmission declined (RR = 0.51, 95% CI: 0.27–0.90), while those of the continental transmission increased (RR = 0.38, 95% CI: 0.09–0.83) as compared to the recent and intermediate transmissions. At MRCA times >100 years, transmission across national boundaries became dominant, with the highest relative risk (RR = 3.78, 95% CI: 2.25–7.88) across spatial scales. These results demonstrate that the evolution of Lineage 1 is characterized by strong spatial structure over time, with long-distance transmissions over the early evolutionary stages but progressively concentrated in local areas in recent years (Fig. 5A).
A Relative risk (RR) of spatial transmission for Lineage 1 isolates, stratified by the estimated time to the most recent common ancestor (tMRCA). Transmission events were grouped into four evolutionary intervals (<10 years, 10–50 years, 50–100 years, and >100 years), and assessed across four geographic scales (local, regional, national, and continental). RRs >1 indicate elevated transmission risk compared to the reference group. B Spatial patterns of Lineage 1.1 and Lineage 1.3 across China, with pairwise links colored by tMRCA category. Insets highlighted transmissions within Yunnan Province. C Distance-stratified RRs of spatial transmission within Lineage 1.1 (left) and Lineage 1.3 (right). Solid lines indicate point estimates of RR, with dashed lines representing 95% confidence intervals. Shaded areas denote different distance bins.
Given the elevated risks of the recent transmissions on the local and regional scales, we make a lineage-stratified inference of B. melitensis dynamics in China. By comparing the tMRCA for samples within Yunnan Province and those from other provinces, we observed distinct spatial dissemination patterns between Lineage 1.1 and Lineage 1.3 (Fig. 5B). We further differentiated the risks of transmission within and outside the province (Fig. 5C). Lineage 1.1 showed elevated relative risks of short-distance transmissions within the province, with estimated RRs of 12.8 (95% CI: 4.7–37.31) for <100 km and 9.4 (3.32–27.07) for the 100–200 km range. The longer geographic distance resulted in a marked increase in the relative risk, with an estimate of 53.6 (95% CI: 21.85–142.6) for transmission events spanning more than 3000 km. These findings suggested that Lineage 1.1 may have undergone substantial geographic expansion across broader regions within China. Despite the limited sample size, we also observed that the dissemination pattern of Lineage 1.2 resembled that of Lineage 1.1 (Supplementary Fig. 9). In contrast, Lineage 1.3 showed limited evidence of national-scale dissemination. Short-distance transmissions within 100 km were associated with the highest RR (19.16, 95% CI: 12.50–29.94); however, increasing spatial distance was linked to a substantial reduction in RRs. Such a decrease in risk with increasing geographic distance suggests primarily localized transmission and limited geographic expansion of the novel lineage. The alternative measures of geographic distance have a marginal impact on the findings (Supplementary Figs. 10 and 11).
To address potential inflation of interprovincial transmission risk estimates due to uneven sampling across provinces, we further incorporated epidemiological data to evaluate the impact of livestock trade on the spread of Brucella. Models incorporating trade data demonstrated a better fit to the observed epidemic dynamics in Yunnan Province (Supplementary Fig. 12A). Higher livestock trade volumes were associated with increased transmission risk (Supplementary Fig. 12B), with the contribution of livestock trade to the Yunnan epidemic peaking ~2015 (Supplementary Fig. 12C). In addition, case occupation patterns differed substantially between lineages. Lineage 1.3 infections were predominantly among farmers and herders, while Lineage 1.1 infections occurred across a broader occupational spectrum (farmers, herders, office staff, students, and retirees) (Supplementary Fig. 12D). The proportion of patients from non-agricultural occupations also peaked in 2015 (Supplementary Fig. 12E, F). These epidemiological findings collectively suggest an increasing risk of interprovincial Brucella dissemination.
Discussion
Brucellosis remains a persistent public health challenge worldwide, particularly in regions where veterinary infrastructure and disease surveillance are often under-resourced47. By analyzing B. melitensis genomes in Southwest China, we provide the in-depth genomic and epidemiological characterization of B. melitensis in the non-traditional epidemic region. Our findings elucidate how distinct evolutionary trajectories and spatial transmission dynamics of B. melitensis lineages have contributed to the emergence and dissemination of brucellosis in this region, providing insights into the epidemiology of this neglected zoonosis.
We revealed the coexistence of a widely distributed lineage and a previously unreported, geographically restricted lineage in a non-traditional epidemic region in China. It is worth noting that Lineage 1.1 and 1.3 exemplified divergent patterns of genomic differentiation, highlighting the variability in evolutionary dynamics among co-circulating lineages. The GCV of Lineage 1.1 was significantly associated with both evolutionary time and spatial prevalence, together with an accumulation of accessory genes, particularly rare genes, along the lineage’s phylogeny. Functional enrichment analyses showed that these genes are often involved in stress response, transport functions, and membrane-related processes, which are typically implicated in ecological adaptation48. Consistently, the observed increase in rare gene content associated with the geographic expansion of Lineage 1.1 provides further evidence for its potential in adaptive evolution. Therefore, the evolutionary success of Lineage 1.1 is suggestive of the adaptive acquisition of functions that may have contributed to the enhanced ecological flexibility of the lineage. In contrast, genomic differentiation in Lineage 1.3 may have been predominantly shaped by geographic isolation. The declined genomic similarity with spatial separation indicated that physical distance and local isolation are key drivers of genomic differentiation for this novel lineage. Additionally, we found no significant associations between genomic variation and either sampling time or climatic factors. This reinforces that spatial isolation, rather than environmental selection, may have dominated the evolution of Lineage 1.3. Although the pan-genome of this lineage appears relatively stable, the significant intra-lineage diversity observed across collection sites implies ongoing micro-evolutionary processes within a geographically constrained context. Such findings align with broader theories of isolation-by-distance in bacterial populations49.
It is important to note that, unlike established systems for pathogens like Mycobacterium tuberculosis, the current absence of a unified whole-genome sequencing-based typing framework for Brucella species hinders comparability across studies due to disparate nomenclature schemes. To robustly define a new sub-lineage despite this limitation, we applied multiple lines of evidence to substantiate the designation of Lineage 1.3 as distinct. Phylogenetically, Lineage 1.3 forms a monophyletic clade with a significantly earlier tMRCA than Lineages 1.1 and 1.2, indicating early divergence. Ecologically, it exhibits distinct geographic and environmental associations. Epidemiologically, and despite the limited sample size, Lineage 1.3 cases were predominantly restricted to farmers, suggesting a pattern consistent with localized spillover. In contrast, Lineage 1.1 infections occurred across a diverse range of occupations—consistent with introduction through broader transmission networks like interprovincial livestock trade. Complementary epidemiological modeling, which supports trade-mediated introduction as a key driver of the human case surge in Yunnan, further corroborates the external origin of Lineage 1.1.
Our findings of spatial structuring indicate the unique patterns of the transmission dynamics of the two lineages. Notably, adaptive expansion may have facilitated the broad dissemination of Lineage 1.1, while geographic confinement and isolation may have shaped the local persistence of Lineage 1.3. Phylogeographic reconstructions revealed that Lineage 1.1 has undergone extensive interprovincial dissemination, particularly over intermediate evolutionary timescales. Elevated risks of transmission were observed across both short and long distances, indicating potential for widespread distribution beyond local foci. However, the transmission probability of Lineage 1.3 declined with geographic distance, suggesting confinement to intra-provincial spread. Such that recent brucellosis cases in Yunnan may result from local persistence of an endemic lineage as well as independent introductions of more adaptive strains. These findings highlight that the risks of both cryptic endemic persistence and silent introduction events are amplified in regions with intermittent surveillance and limited public health resources. Therefore, the implementation of lineage-specific surveillance strategies, with Lineage 1.3 requiring focused local monitoring and Lineage 1.1 demanding broader, cross-regional containment measures.
The zoonotic nature of brucellosis underscores that human infections primarily originated from livestock exposure, either through direct contact or consumption of contaminated products. While human-to-human transmission is rare, occupational exposure to goat and/or sheep remains the dominant risk factor. Our analyses revealed striking lineage-specific epidemiological patterns (Supplementary Fig. 12): lineage 1.3 cases were almost exclusively farmers, suggesting localized agricultural spillover; while Lineage 1.1 infections were recorded among a broader range of occupations (farmers, herders, office staff, students, and retirees), implying complex transmission routes, possibly through livestock trade or food transportation networks. These occupational disparities may drive the divergent evolutionary trajectories and spatial dynamics between lineages. However, the lack of systematic livestock sampling limited the direct reconstruction of the transmission networks. Implementing a One Health framework—integrating human, animal, and environmental surveillance—is critical to verify these hypothesized transmission pathways and elucidate the evolutionary trajectories of co-circulating lineages across extended timescales.
It is recognized that the closed pan-genome structure of B. melitensis is not consistent with previous studies50. This discrepancy may be partially attributable to the methodological differences among the studies51. For example, we employed Panaroo to merge gene families based on conserved genomic contexts20, thereby reducing annotation artifacts that may inflate accessory gene counts. To further explore this discrepancy, we examined gene family copy numbers and found increasing variation over evolutionary time. This suggests that previous analyses may have overestimated genome openness by treating copy number variation as gene presence/absence. While gene families remain largely conserved, divergence in copy number contributes to intra-lineage genomic diversity and may confound accessory genome estimates. These insights underscore the importance of selecting appropriate analytical tools in microbial genomic studies.
Our study contributes to the understanding of the establishment of neglected zoonotic diseases in new ecological contexts. Although brucellosis has not historically been regarded as a major concern in Southwest China, the ongoing globalization of trade and shifts in agricultural practices may further accelerate such dynamics, enabling diseases like brucellosis to expand into areas previously considered low risk52. Although Lineage 1.2 was sporadically detected in Yunnan and other provinces, our analysis was constrained by a limited sample size. In contrast, the discovery of the distinct Lineage 1.3 in Yunnan highlights critical gaps in current genomic surveillance, particularly in understudied and low-incidence regions.
In conclusion, our study illustrates the complex interplay of local adaptation, global connectivity, and pathogen evolution that collectively drives the emergence and dissemination of B. melitensis lineages in Southwest China. By integrating genomic, phylogenetic, and epidemiological approaches, we provide an evidence base for more nuanced interventions to mitigate brucellosis risks. The findings underscore the vigilant monitoring and proactive management of neglected zoonotic diseases in non-traditional epidemic and currently low-risk regions.
Ethics statement
This study complies with all relevant ethical regulations. The approval was obtained from the Medical Ethics Committee of Yunnan Institute of Endemic Disease Control and Prevention (NO. ICDC-2025014). Informed consent was obtained from all human participants prior to sample collection.
Data availability
The raw sequence data reported in this paper have been deposited in the GSA database (China National Center for Bioinformation) under GSA CRA030806. The source data for the main and Supplementary Figs. are available in FigShare public repository54 (https://doi.org/10.6084/m9.figshare.30555551.v1). Publicly Brucella melitensis genomes were obtained from the NCBI Genome Dataset, and accession numbers for all genomes are provided in Supplementary Data 1. Climatic data were derived from the fifth-generation reanalysis41 and accessed via Google Earth Engine. Food trade data were obtained from China’s interprovincial physical food trade dataset46. All other data are available from the corresponding author on reasonable request.
Code availability
The codes used to perform the analysis are available in the GitHub repository (https://github.com/xiyanglee/brucellosis-wgs-yunnan) and Zenodo public repository53 (https://doi.org/10.5281/zenodo.17260137).
References
Halliday, J. E. B. et al. Endemic zoonoses in the tropics: a public health problem hiding in plain sight. Vet. Rec. 176, 220–225 (2015).
Pappas, G., Papadimitriou, P., Akritidis, N., Christou, L. & Tsianos, E. V. The new global map of human brucellosis. Lancet Infect. Dis. 6, 91–99 (2006).
Gwida, M. et al. Brucellosis - regionally emerging zoonotic disease? Croat. Med. J. 51, 289–295 (2010).
World Health Organization and others. The control of neglected zoonotic diseases: a route to poverty alleviation: report of a joint WHO. (2006).
Corbel, M. J. Brucellosis in Humans and Animals. (World Health Organization, 2006).
Vonesch, N. et al. Emerging zoonotic viral infections of occupational health importance. Pathog. Dis. 77, ftz018 (2019).
Saker, L. et al. Globalization and infectious diseases: a review of the linkages. (2004).
Chen, S. et al. Increasing threat of brucellosis to low-risk persons in urban settings, China. Emerg. Infect. Dis. 20, 126–130 (2014).
Ji-qiu, W. National Brucellosis intervention pilot county survey on the economic losses. Chin. J. Control Endem. Dis. 13, 929040 (2008).
Zhong, Z. et al. Human brucellosis in the People’s Republic of China during 2005-2010. Int. J. Infect. Dis. 17, e289–e292 (2013).
Sun, G.-Q. et al. Transmission dynamics of brucellosis: mathematical modelling and applications in China. Comput. Struct. Biotechnol. J. 18, 3843–3860 (2020).
Yang, H. et al. Epidemic characteristics, high-risk areas and space-time clusters of human Brucellosis - China, 2020–2021. China CDC Wkly 5, 17–22 (2023).
Li, Z. et al. Epidemiological characteristics of brucellosis in Yunnan Province from 2008 to 2018 and prediction of grey GM (1, 1) model. Chin. J. Endem. 40, 225–229 (2021).
Organization, W. H. & others. Taking a multisectoral one health approach: a tripartite guide to addressing zoonotic diseases in countries. (Food & Agriculture Org., 2019).
Erkyihun, G. A. & Alemayehu, M. B. One Health approach for the control of zoonotic diseases. Zoonoses 2, 963 (2022).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, 1–22 (2017).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Matthews, C. A., Watson-Haigh, N. S., Burton, R. A. & Sheppard, A. E. A gentle introduction to pangenomics. Brief. Bioinform. 25, bbae588 (2024).
Tantoso, E. et al. To kill or to be killed: pangenome analysis of Escherichia coli strains reveals a tailocin specific for pandemic ST131. BMC Biol. 20, 146 (2022).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Sterzi, L. et al. Genetic barriers more than environmental associations explain Serratia marcescens population structure. Commun. Biol. 7, 468 (2024).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Domingo-Sananes, M. R. & McInerney, J. O. Mechanisms that shape microbial pangenomes. Trends Microbiol 29, 493–503 (2021).
Tu, Q. & Lin, L. Gene content dissimilarity for subclassification of highly similar microbial strains. BMC Genomics 17, 647 (2016).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15–e15 (2015).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinform. 30, 1312–1313 (2014).
Tonkin-Hill, G., Lees, J. A., Bentley, S. D., Frost, S. D. W. & Corander, J. Fast hierarchical Bayesian analysis of population structure. Nucleic Acids Res 47, 5539–5549 (2019).
Abdel-Glil, M. Y. et al. Core genome multilocus sequence typing scheme for improved characterization and epidemiological surveillance of pathogenic brucella. J. Clin. Microbiol. 60, e0031122 (2022).
Pisarenko, S. V. et al. Global evolution and phylogeography of Brucella melitensis strains. BMC Genomics 19, 353 (2018).
Rabinowitz, P. et al. Whole genome sequence analysis of Brucella melitensis phylogeny and virulence factors. Microbiol. Res. 12, 698–710 (2021).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 4, vey016 (2018).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772–772 (2012).
Drummond, A. J., Ho, S. Y. W., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLOS Biol. 4, e88 (2006).
Liao, J. et al. Nationwide genomic atlas of soil-dwelling Listeria reveals effects of selection and population ecology on pangenome evolution. Nat. Microbiol. 6, 1021–1030 (2021).
Tonkin-Hill, G. et al. Robust analysis of prokaryotic pangenome gene gain and loss rates with Panstripe. Genome Res. 33, 129–140 (2023).
Copernicus Climate Change Service. ERA5 monthly averaged data on single levels from 1940 to present. Copernicus Climate Change Service (C3S) Climate Data Store (CDS) https://doi.org/10.24381/CDS.F17050D7 (2019).
Wood, S. N. Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. J. R. Stat. Soc. 73, 3–36 (2011).
Salje, H. et al. Dengue diversity across spatial and temporal scales: Local structure and the effect of host population size. Science 355, 1302–1306 (2017).
Moura, A. et al. Emergence and global spread of Listeria monocytogenes main clinical clonal complex. Sci. Adv. 7, eabj9805 (2021).
Belman, S. et al. Geographical migration and fitness dynamics of Streptococcus pneumoniae. Nature 631, 386–392 (2024).
Luo, L. et al. A dataset of interprovincial food trade flows in China. Sci. Data 12, 943 (2025).
Godfroid, J. Brucellosis in livestock and wildlife: zoonotic diseases without pandemic potential in need of innovative one health approaches. Arch. Public Health Arch. Belg. Sante Publique 75, 34 (2017).
Kornspan, D., Zahavi, T. & Salmon-Divon, M. The acidic stress response of the intracellular pathogen brucella melitensis: new insights from a comparative, genome-wide transcriptome analysis. Genes 11, 1016 (2020).
Long, G. S. et al. A 14th century CE Brucella melitensis genome and the recent expansion of the Western Mediterranean clade. PLOS Pathog. 19, e1011538 (2023).
Yang, Z. et al. Comparative genomic analysis provides insights into the genetic diversity and pathogenicity of the genus Brucella. Front. Microbiol. 15, 1389859 (2024).
Manzano-Morales, S., Liu, Y., González-Bodí, S., Huerta-Cepas, J. & Iranzo, J. Comparison of gene clustering criteria reveals intrinsic uncertainty in pangenome analyses. Genome Biol. 24, 250 (2023).
Rohr, J. R. et al. Emerging human infectious diseases and the links to global food production. Nat. Sustain. 2, 445–456 (2019).
Li, X. iyang Source code ‘Genom. Spat. heterogeneity Brucella melitensis underscores latent brucellosis risks non-Endem. zones Southwest China’ Creators https://doi.org/10.5281/zenodo.17260137 (2025).
Xiyang Li. Source data for ‘Genomic and spatial heterogeneity of Brucella melitensis underscores latent brucellosis risks in non-endemic zones in Southwest China’. https://doi.org/10.6084/m9.figshare.30555551.v1 (2025).
Acknowledgements
This study was supported by grants from the Seeds funding of Nanjing Medical University (NMUR20220001). The funder of the study had no role in the study design, data collection, data analysis, data interpretation, or writing of the manuscript.
Author information
Authors and Affiliations
Contributions
R. Li and Z. Gao designed the research; F. Yang and B. Yu contributed materials; X. Li, Z. Lou, W. Zhao, and X. Lu performed data analyses; D. Lin, H. Zhao, J. Lu, and X. Ding wrote the paper; X. Yang, Q. Yang, and M. Shi planned the projects and participated in paper preparation. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Kevin Daly, with co-reviewer Louis L’Hôte, and Junjie Yue for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, F., Li, X., Yu, B. et al. Genomic and spatial heterogeneity of Brucella melitensis underscores latent brucellosis risks in non-endemic zones in Southwest China. Commun Biol 8, 1824 (2025). https://doi.org/10.1038/s42003-025-09218-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09218-x