Abstracts
Paliurus hemsleyanus Rehd., a deciduous shrub or small tree endemic to China, is valued for its hardiness, economic and ornamental importance, and widespread used as a rootstock for Chinese jujube (Ziziphus jujuba Mill.). Despite its ecological and economic significance, genomic resources for this genus remain limited. Here, we assemble a haplotype-resolved, telomere-to-telomere (T2T), gap-free genome of P. hemsleyanus (2n = 24), representing the genus Paliurus. The genome comprises two haplotypes of 306.65 Mb and 306.21 Mb, with contig N50 values of 24.91 Mb and 24.94 Mb, respectively. Each haplotype encodes over 29,000 protein-coding genes, with all centromeres and telomeres fully predicted. Allele-specific expression analysis reveals a positive correlation between gene expression divergence and sequence variation, indicating functional differentiation between haplotypes. Comparative genomic analysis shows relatively stable genome evolution within Rhamnaceae, with all examined extant species containing 12 chromosomes. Disease resistance (NLR) genes exhibit a root-preferred expression pattern, and allelic copies expressed more strongly than non-allelic ones between haplotypes. Ascorbic acid (AsA) metabolic genes show leaf-preferred expression; and moreover MDHAR genes exhibit Rhamnaceae-specific tandem duplications, suggesting lineage-specific adaptive evolution. This high-quality genome provides an essential resources for evolutionary studies, functional genomics, breeding, and the conservation of Rhamnaceae species.
Similar content being viewed by others
Introduction
Paliurus hemsleyanus Rehd., commonly known as the Chinese Coin Tree (referred to as cointree in this study), is an important deciduous shrub or small tree species of the cosmopolitan plant genus Paliurus of Rhamnaceae family1. This tree is endemic to China’s montane forests, found at elevations up to 1600 m across north to south provinces. Noted for its ecological resilience, rated hardy to USDA Zone 6 and RHS H5, it thrives on dry slopes and in nutrient-poor soils. Cointree offers both economic values, such as containing bioactive triterpenes in root2, and ecological services, such as soil stabilization1. In addition, it serves as a preferred rootstock for Chinese jujube (Ziziphus jujuba Mill.), one of the five most important economic forest tree species, where grafted scions exhibit enhanced resistance to phytoplasma induced witches’ broom disease, the world-wide most destructive disease in jujube production3,4. Despite its ecological and economic significance, cointree lacks high-quality genomic resources, underscoring the need for a comprehensive genome assembly to advance understanding of its genome biology and evolutionary history.
Nucleotide-binding leucine-rich repeat proteins (NLRs) serve as central guardians of plant immunity, orchestrating defense responses through their modular domain architecture5. These resistance (R) proteins typically adopt a tripartite structure: an N-terminal signaling domain (TIR, CC, or RPW8), a central NB-ARC domain that binds ATP/GTP to regulate activation, and a C-terminal LRR region that dynamically evolves to recognize pathogen effector. This structural plasticity enables NLRs to function as molecular switches, directly sensing pathogen invasion via LRR-mediated effector interactions and triggering immune signaling cascades through conserved NB-ARC and N-terminal domains6. Genomic surveys across angiosperms reveal remarkable NLR diversity, with inventories ranging from ~300 in diploids like Vitis davidii7 to over 2,000 in polyploid wheat (Triticum aestivum)8, highlighting their adaptive expansion under external stress. Functional studies further demonstrate that NLR expression dynamics critically shape disease outcomes. For instance, the Yr87/Lr85 NLR from Aegilops species confers dual resistance to wheat stripe rust (Puccinia striiformis) and leaf rust (P. triticina)9 while atypical NLR paralogs (Rpp6907-7/Rpp6907-4) in soybean (Glycine max) provide broad-spectrum rust immunity10. These examples underscore NLRs’ dual role as both pathogen-specific sentinels and versatile regulators of plant immunity11. Despite cointree’s renowned stress tolerance, particularly to drought and phytoplasma disease, its NLR repertoire remains uncharacterized, a critical gap this study addresses through T2T genome analysis.
Ascorbic acid (AsA, vitamin C), a ubiquitous non-enzymatic antioxidant in plants, is synthesized through four metabolic pathways: the L-galactose, D-galacturonic acid, inositol, and L-gulose pathways. Among these, the L-galactose and D-galacturonic acid pathways dominate in most plant species12. Beyond its biosynthetic role, AsA plays a crucial function in maintaining redox homeostasis by scavenging reactive oxygen species (ROS) through tightly coordinated degradation and regeneration cycles. During H₂O₂ detoxification, AsA is oxidized to monodehydroascorbate (MDHA) by ascorbate peroxidase (APX). MDHA is either directly reduced back to AsA by monodehydroascorbate reductase (MDHAR) or undergoes disproportionation to form AsA and dehydroascorbate (DHA). DHA is subsequently reduced to AsA by dehydroascorbate reductase (DHAR) using reduced glutathione (GSH) as an electron donor, while glutathione reductase (GR) regenerates GSH from its oxidized form (GSSG). These sequential reactions comprise the AsA–GSH cycle, which maintains antioxidant homeostasis and sustains electron flow from NADPH to H₂O₂. Through this cycle, plants enhance their tolerance to both biotic and abiotic stresses13. For example, in Actinidia chinensis, the transcription factor AcePosF21 interacts with the R2R3-MYB protein AceMYB102 to activate GDP-L-galactose phosphorylase 3 (AceGGP3), thereby reducing ROS accumulation under cold stress14. Similarly, exogenous AsA application in tomato enhances salt tolerance by stabilizing chlorophyll biosynthesis and improving stomatal function15. In cointree, tissue-specific coordination of AsA metabolism, particularly the oxidization and regeneration pathway, likely underpins its ability to sustain physiological activity under adverse conditions.
Haplotype-resolved telomere-to-telomere (T2T) genomes have revolutionized the study of complex genomic regions by enabling precise annotation of centromeres, telomeres, and allelic variation. However, the genomes of cointree and the genus Paliurus remain unexplored, limiting insights into NLR clusters, AsA pathway genes, and haplotype-specific allelic variation. Here, we integrated PacBio HiFi, Oxford Nanopore (ONT) ultralong reads, and Hi-C data to generate a haplotype-resolved T2T genome for cointree and for the genus Paliurus. We assembled all 24 chromosomes (12 per haplotype) with fully annotated repeat sequences and protein-coding genes. Allele-specific expression analysis revealed a positive correlation between expression levels and allelic divergence. Comparative analyses showed a relatively stable genomic structure in Rhamnaceae compared with Rosaceae. NLR genes exhibited root-preferred expression, with bi-allelic genes expressed significantly more than single-allele genes. Tissue-specific analysis of AsA metabolism revealed higher expression and AsA content in leaf, alongside a lineage-specific tandem expansion of MDHAR genes in Rhamnaceae. This high-quality haplotype-resolved genome provides a valuable resource for investigating Rhamnaceae genome biology and evolution, supporting future comparative, functional, and applied studies.
Results
Haplotype-resolved T2T genome assembly and validation
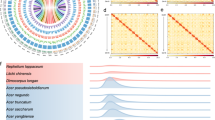
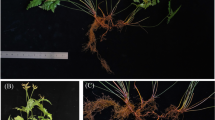
Mature leaves of cointree were used for whole-genome DNA sequencing, while RNA was extracted from root, stem, and leaf for transcriptome sequencing (Fig. 1A). Genome size was estimated at ~299 Mb with ~0.74% heterozygosity, based on 43.06 Gb (71×) of clean paired-end reads from the MGISEQ-2000 platform (Supplementary Table 1 and Fig. 1B). For assembly, we used 12.5 Gb (41.7×) of PacBio HiFi CCS reads, 53.78 Gb (180×) of ultra-long Oxford Nanopore (ONT) reads, and 60.5 Gb (201×) of Hi-C data. The initial haplotype-resolved assembly yielded 100 and 87 contigs for HapA and HapB, with contig N50 values of 23.86 Mb and 23.65 Mb, respectively (Supplementary Table 2). After removing redundant and low-quality contigs, 15 and 14 high-quality contigs remained for HapA and HapB (Supplementary Table 2 and Supplementary Data 1). Hi-C interaction data enabled the clustering and anchoring of contigs into 12 chromosomes per haplotype, with 68.79% of Hi-C reads identified as valid (Supplementary Table 3). Five remaining gaps were closed based on HiFi read mapping (Supplementary Table 4 and Supplementary Fig. 1). The final HapA and HapB assemblies measured 306.66 Mb and 306.21 Mb, with contig N50s of 24.91 Mb and 24.94 Mb, respectively (Fig. 1C and Table 1).
A Morphology of the cointree: the whole tree (photographed in the greenhouse of Hebei Agricultural University) (upper left), together with its branch and leaves (upper right) and root (bottom left); as well as a fruit photograph collected outdoors (bottom right). Root, stem, and leaf tissues from this tree were used for RNA-seq sequencing, with three biological replicates per tissue in this study. Scale bars are shown at the bottom right of each photograph. B K-mer frequency distribution for genome size estimation and heterozygosity assessment. C Circular genome overview. From outer to inner tracks: (1) chromosome ideogram (Mb scale), where A and B indicate HapA and HapB, respectively, followed by chromosome number; (2) gene density; (3) repeat density; (4) SNP/Indel density; (5) structural variantion distribution; (6) GC content; and (7) inter-chromosomal synteny between haplotypes. D Hi-C contact map, with darker colors indicating higher interaction frequency. ChrA and ChrB denote the chromosomes of the HapA and HapB genomes, respectively.
Assembly quality was evaluated in three steps. First, Hi-C data confirmed structural accuracy: 59.48 Gb (98.31%) of valid interactions were used to construct Hi-C contact maps on both haplotypes, showing strong diagonal signals consistent with chromosomal proximity (Fig. 1D and Supplementary Fig. 2). Second, BUSCO analysis revealed 98.90% completeness in both haplotypes, and mapping the HiFi reads back to the two genomes showed both 99.74% mapping rates, further supporting the high completeness of the assemblies (Table 1 and Supplementary Table 5). The LTR Assembly Index (LAI), a metric for assessing assembly continuity and completeness, reached 18.55 and 18.62 for HapA and HapB, respectively, indicating high assembly quality (Table 1). Finally, Merqury assessment showed QV scores of 51.44 (HapA) and 51.51 (HapB), with base error rates below 0.0001, indicating high assembly accuracy (Supplementary Table 6).
Repeat and protein-coding gene annotation
We identified 145.88 Mb (47.57%) and 145.04 Mb (47.37%) of repetitive sequences in HapA and HapB, respectively, with transposable elements (TEs) as the major components, accounting for 91.61% and 91.45%. Class I TEs constituted over 50% in both haplotypes, among which Gypsy and Copia were the most abundant, representing 12.44% and 6.76% in HapA, and 12.52% and 6.75% in HapB, respectively. The proportion of intact LTRs reached 3.47% in HapA and 3.38% in HapB (Table 1 and Supplementary Data 2). All telomere sequences were identified, ranging from 689 bp to 11,609 bp in HapA and from 1,872 bp to 12,660 bp in HapB, with average lengths of 7650 bp and 7601 bp, respectively (Supplementary Data 3). Centromeric regions were predicted on all chromosomes using QuartTET16, ranging from 101,414 bp to 2,254,431 bp in HapA and 109,227 bp to 2,149,307 bp in HapB, with average lengths of 875,550 bp and 764,555 bp, respectively. These regions were mainly composed of long terminal repeat retrotransposons (LTR-RTs) (Supplementary Table 7). Alignment of jujube ChIP-seq data17 to HapA did not reveal distinct peak signals, whereas mapping of jujube centromeric retrotransposons (CRJs) showed enrichment across all chromosomes, despite suboptimal alignment. In addition, a small number of tandem repeats (TRs) were identified by QuartTET on chromosomes 2, 3, 8, and 11, and their positions overlapped with CRJ enrichment sites (Supplementary Fig. 3).
Combining the de novo prediction, protein homology search, and transcripts-based evidence from RNA-seq (Supplementary Table 1), we finally identified a total of 29,165 and 29,238 protein-coding genes in HapA and HapB genomes, of which 93.73% and 93.60% have been functionally annotated by either eggNOG-mapper or Interproscan databases, respectively, with 17,402 and 17,437 genes having GO annotations and 12,114 genes and 12,150 genes having KEGG annotations (Table 1). Using the RNA-seq data from three tissues, root, stem and leaf, 19,283 (66.12%) genes and 19,201 (65.67%) genes expressed in at least one tissue for HapA and HapB, respectively.
We divided the expression values (FPKM) of each gene into five levels according to published studies18,19: very high (FPKM > = 1000), high (FPKM 100–1000), medium (FPKM 10–100), low (FPKM 1–10), and very low (FPKM 0–1). In leaves, the proportions of expressed genes in these five intervals for HapA were 0.38%, 5.90%, 30.93%, 15.96%, and 46.82%, respectively; while for HapB they were comparable, at 0.31%, 5.08%, 30.17%, 16.97%, and 47.46%. Overall, expression patterns across the three tissues were similar: genes with very high expression accounted for the smallest fraction (less than 0.5%), and genes with very low expression accounted for the largest fraction (over 45%), while approximately 60% of the remaining genes showed medium expression levels (Supplementary Table 8 and Supplementary Fig. 4).
Variation and allele-specific expression (ASE) between two haplotypes
Collinearity analysis between HapA and HapB revealed a perfect genomic synteny (Fig. 2A), however contained substantial variations. In detail, A total of 1,295,745 single nucleotide polymorphisms (SNPs) and 348,631 insertions/deletions (Indels) were identified, with chromosome 4 exhibiting the highest number of variations (Supplementary Table 9). Additionally, 13,763 structural variants (SVs) were detected, including 6883 insertions, 6825 deletions, 28 translocations, 16 inversions, and 11 duplications. The majority of these variants (58.78%) ranged from 50 to 100 bp in length, with a sharp decline in frequency observed for variants between 100 and 250 bp (Supplementary Fig. 5). Collectively, these variants accounted for approximately 3.88% of the total genome size and affected 66.66% (19,442/29,165) of protein-coding genes by introducing sequence differences in their coding regions.
A Whole-genome collinearity comparison between HapA and HapB. B Sequence identity distribution of allelic gene pairs between the two haplotypes. Numbers in brackets indicate the proportion of sequences falling within each identity interval. C Distribution of nonsynonymous mutation counts across allelic gene pairs. D Allele-specific expression between HapA and HapB; colors represent different log₂ fold change (log2FC) values between haplotypes. E Pearson’s correlation between allelic sequence identity and expression divergence. The x-axis shows allelic gene identity, and the y-axis shows allelic expression divergence (absolute logFC). Linear regression fitting was applied, and the p-value was calculated from the Student’s t-test of the correlation coefficient’s significance (n = 829). F–H Box-plot distribution of SNPs, Indels, and SVs in coding regions, 2 kb upstream regions, and introns, along with Student’s t-test statistical comparisons between DEGs (n = 829) and non-DEGs (n = 27,036). I Schematic diagram showing the 285 bp insertion in the first exon of the ASS1 gene on HapB chromosome. The figure was initially generated in R using the “gggenes” and “ggplot2” packages, and subsequently refined proportionally in Adobe Illustrator. J Expression profile of PhAASS1 in leaf, root, and stem tissues.
We identified 27,865 allelic gene pairs between the two haplotypes, which exhibited a high average sequence identity of 98.57% (Fig. 2B and Supplementary Data 4). To further characterize allele-specific variations, we analyzed nonsynonymous substitutions within coding regions and found that 48.63% (13,550/27,865) of the allelic gene pairs harbored such mutations (Fig. 2C and Supplementary Data 4), and among these, the majority (approximately 80%) contained fewer than three nonsynonymous changes per gene pair. To assess whether allelic sequence divergence resulted in differential gene expression, we conducted differential expression analysis using the RNA-seq data from leaves. A total of 2.98% (829/27,865) of allelic gene pairs displayed significant expression imbalance between the haplotypes (Fig. 2D and Supplementary Data 5). Gene Ontology (GO) enrichment analysis revealed that these ASE genes were primarily involved in defense responses and stress-related processes. Moreover, KEGG pathway analysis indicated significant enrichment in pathways such as ribosome function and plant–pathogen interaction (Supplementary Fig. 6).
For the 829 ASEs, the sequence identity of the allelic pairs was significantly negative correlated with their expression divergence evaluated (logFC) (r: −0.152, p value: 0.0002), indicating that greater allelic sequence divergence is associated with greater expression divergence (Fig. 2E and Supplementary Data 5). In addition, the average number of SNPs, Indels, and SVs was all significantly higher in ASEs compared with non-ASEs. Within ASEs, the average numbers of these three types of variants in the 2 Kb upstream region (largely encompassing the promoter) and introns were more than an order of magnitude higher than in coding regions (Fig. 2F–H and Supplementary Data 6).
For allelic pairs with extensive variations (249 out of 829; defined as carrying structural variants or more than the average number of SNPs or small indels) in the 2 kb upstream region, GO enrichment highlighted processes such as arginine biosynthesis, metabolism of cell wall macromolecules, aminoglycan and chitin metabolism, and catabolic processes related to aminoglycan and chitin. KEGG enrichment revealed pathways including arginine biosynthesis, biosynthesis of amino acids (particularly arginine), and carbon fixation (Supplementary Fig. 7 and Supplementary Data 5).
Detailed analysis of these 249 ASEs identified one notable allelic pair encoding argininosuccinate synthase: PhAASS1, functional in the HapA genome, and a nonfunctional allele in HapB (pseudo-PhBASS1) disrupted by a 285 bp insertion in the first exon (Fig. 2I and Supplementary Fig. 8a). This insertion was supported by HiFi reads (Supplementary Fig. 8b). PhAASS1 was expressed at relatively low levels in leaf, but was nearly absent in root and stem (Fig. 2J). In addition, two other tandem copies of this gene family were present in both haplotypes and showed moderate expression, particularly in leaves (Supplementary Fig. 8c).
Evolutionary insights from comparative genomics of cointree and relatives
To perform comparative genomic analysis with cointree, we included Vitis vinifera (Vvi) and Glycine max (Gma) as outgroups, along with 16 additional Rosales species with publicly available genome sequences (Supplementary Data 7). These comprised three Rhamnaceae species (Ziziphus jujuba [Zju], Sageretia thea [Sth], and Rhamnella rubrinervis [Rru]); six Rosaceae species (Malus domestica [Mdo], Pyrus communis [Pco], Prunus persica [Ppe], Rosa chinensis [Rch], Argentina anserina [Aan], and Fragaria vesca [Fve]); two Elaeagnaceae species (Elaeagnus mollis [Emo] and Hippophae salicifolia [Hsa]); one Ulmaceae species (Ulmus minor [Umi]); one Urticaceae species (Urtica dioica [Udi]); one Moraceae species (Morus notabilis [Mno]); and two Cannabaceae species (Humulus lupulus [Hlu] and Cannabis sativa [Csa]). Including cointree (Paliurus hemsleyanus [Phe]) (HapA used here), a total of 19 species were analyzed.
Based on the protein-coding genes of the 19 species, a total of 6940 gene families were found to be shared among them. Among the 14,766 gene families which contained 95.4% genes identified in cointree, 410 were species-specific (Fig. 3A and Supplementary Table 10). The four Rhamnaceae species exhibited highly similar distributions of unassigned genes, unique gene families, single-copy, and multi-copy gene families, whereas the other Rosales showed more variable patterns, with gene family numbers generally positively correlated with genome size (Fig. 3B and Supplementary Data 8). A maximum-likelihood phylogenetic tree constructed from 114 single-copy orthologous genes revealed that cointree is most closely related to jujube, with their divergence estimated at 5–17 million years ago (Mya). Gene family expansion and contraction analysis identified 447 expanded and 833 contracted gene families in cointree, derived from 71 and 697 ancestral expansion and contraction events of Rosales, respectively, across the 19 species (Fig. 3C).
A Petal diagram showing the shared and unique gene families among cointree and five other plant species. B Numbers of unassigned genes and gene families across 19 species. C Phylogenetic tree of the 19 species constructed using 114 single-copy orthologous genes, with Vvi and Gma as outgroups. Divergence times in million years (Mya) are shown in red. Pie charts indicate gene family expansions (purple) and contractions (blue). The published ancestral eudicot karyotype (AEK), together with our inferred ancestral karyotypes of Rhamnaceae (RAK) and Rosaceae (ARK), are displayed at the corresponding nodes. Karyotypes of each of the 19 species are shown next to species names. The red star marks the γ whole-genome triplication event, and the blue star indicates recent species-specific whole-genome duplications in Mdo and Pco. D Ks and 4DTv distributions of paralogous gene pairs within seven representative species, highlighting whole-genome duplication events (ss: species-specific). E Ks and 4DTv distributions of orthologous gene pairs between cointree (Phe) and each of the other six representative species, reflecting speciation events.
Based on the Phylogenetic tree, we inferred the ancestral karyotypes of Rosaceae and Rhamnaceae, the two largest families within Rosales in terms of sequenced species, and reconstructed the karyotype evolution of Rosales using the seven published ancestral eudicot karyotype (AEK) chromosomes (Fig. 3C and Supplementary Fig. 9)20. The results showed that the six Rosaceae species were derived from eight ancestral chromosomes (ARK, Ancestral Rosaceae Karyotype). Within this family, Rch, Fve, and Aan each possess seven chromosomes, whereas Mdo and Pco have 17 chromosomes, indicating a recent whole-genome duplication (WGD) accompanied by chromosomal fissions and/or fusions after their divergence. By contrast, Ppe retained eight chromosomes and did not experience this WGD event, a pattern also supported by collinearity dot plots. For Rhamnaceae, the four analyzed species were derived from 12 ancestral chromosomes (RAK, Ancestral Rhamnaceae Karyotype), with all extant species maintaining 12 chromosomes (Supplementary Figs. 10 and 11). Some chromosomes were entirely derived from single AEK chromosome, such as chromosome 1 of Rru and Sth, and chromosome 12 of Phe, all from AEK5, as well as chromosome 10 of Rru, chromosome 3 of Phe, and chromosome 3 of Zju, all from AEK2. This suggests that, compared with Rosaceae, Rhamnaceae experienced fewer chromosomal fusion events and did not undergo large scale lineage-specific duplications.
We further investigated gene-based collinearity and calculated the synonymous substitution rate (Ks) and the four-fold synonymous third-codon transversion (4DTv) among paralogs and orthologs in studied 17 Rosales species. All exhibited the shared ancient whole-genome triplication event (γ event), with Ks and 4DTv peaks ranging from 1.36 to 1.71 and from 0.47 to 0.59, respectively (Fig. 3D, Supplementary Figs. 12 and 13, and Supplementary Table 11). In contrast, recent species-specific peaks varied among species. Within Rosaceae, Mdo (apple) and Pco (pear) exhibited pronounced whole-genome duplication (WGD) signals, as did the two Elaeagnaceae species, Emo and Hsa, all of which displayed elevated peaks relative to other taxa (Fig. 3D, Supplementary Fig. 12). Peak values were nearly identical within each family (Ks = 0.180 and 0.184, 4DTv = 0.063 and 0.063 for Mdo and Pco; Ks = 0.376 and 0.348, 4DTv = 0.129 and 0.121 for Hsa and Emo), suggesting a family-specific duplication pattern. Moreover, the results suggest that the lineage-specific WGD possibly occurred earlier in Elaeagnaceae than in Rosaceae (Supplementary Table 11). The γ-triplication event is also evident in the intra-species dot plot, showing a three-copy pattern. In addition, a species-specific WGD is observed in Mdo, which, together with the γ-event, results in a six-copy pattern (Supplementary Fig. 13).
This species-specific peaks were absent or weak in all other Rosales. Within Rhamnaceae, all four species retained minor peaks, while among other Rosaceae members, only Rch exhibited a small peak (Fig. 3D). This minor peak is not observable in the dot plot, unlike the γ-event or the species-specific duplication in Mdo. We further examined the relationship between Ks peak height and genome size, and found that the γ-event peak height was significantly negatively correlated with genome size (p = 0.01, Student’s t-test), whereas species-specific peaks showed a significant positive correlation (p = 0.004, Student’s t-test) (Supplementary Fig. 14). Beyond paralogous comparisons, analysis of orthologous gene pairs using Ks and 4DTv revealed the closest relationship between Zju (jujube) and Phe (cointree), with Ks = 0.074 and 4DTv = 0.027, indicating a more recent divergence compared with other Rhamnaceae species, including Rru (Ks = 0.364, 4DTv = 0.137) and Sth (Ks = 0.348, 4DTv = 0.129) (Fig. 3E and Supplementary Table 11). Furthermore, the γ-event is also evident in the dot plot of inter-species comparisons (Supplementary Fig. 13).
The profiles and tissue expression of NLR genes in cointree
One of the most notable features of cointree is its stress resilience, which may be associated with its abundant repertoire of NLR genes. We identified a total of 853 NLRs, including 427 in HapA and 426 in HapB, classified into four subfamilies: 142/131 Coiled-Coil NB-ARC Leucine-Rich Repeats (CNLs), 183/191 NB-ARC Leucine-Rich Repeats (NLs), 96/98 Toll/Interleukin-1 Receptor NB-ARC Leucine-Rich Repeats (TNLs), and 6/6 Resistance to Powdery Mildew 8 NB-ARC Leucine-Rich Repeat (RNLs) in HapA/HapB, respectively (Supplementary Data 9). These genes were broadly distributed across both haplotypes, often forming clusters generated by tandem duplications (Supplementary Fig. 15 and Supplementary Data 9). Phylogenetic analysis of conserved domains revealed clear differentiation among CNL, NL, and TNL subfamilies, while RNLs were interspersed within the NL clade. Several genes were ambiguously nested between NL and CNL clades, consistent with UMAP clustering, which indicated closer sequence similarity between NLs and CNLs than between NLs and TNLs (Fig. 4A and Supplementary Fig. 16).
A UMAP clustering of NLR genes based on sequence similarity, showing relationships among CNL, TNL, NL, and RNL subfamilies. B Distribution of NLRs in both haplotypes across different FPKM intervals for three tissues. C Ternary plot of expression levels for 853 NLRs across all transcriptome samples. Each dot represents a gene triad with R, S, and L coordinates. Genes near vertices are dominant, those along edges are suppressed, and grey dots near the center indicate balanced triads. D Venn diagram of highly expressed NLR genes (FPKM > 5.46) across three tissues. E Ratio of TE insertions in the four NLR types across two haplotypes. F Boxplot of total TE numbers in each NLR type with Student’s t-test statistical comparisons. The sample size used to derive statistics for each NLR type are indicated below each plot: CNL (n = 146), NL (n = 183), RNL (n = 6), and TNL (n = 96) in HapA; and CNL (n = 131), NL (n = 191), RNL (n = 6), and TNL (n = 98) in HapB. G Boxplot showing FPKM distributions of allelic versus non-allelic genes within the CNL (n = 240 vs. n = 36), NL (n = 290 vs. n = 87), and TNL (n = 170 vs. n = 25) families, with Student’s t-test statistical comparisons. Non-allelic genes were not present in the RNL family and are therefore excluded. H Venn diagram of NLRs exhibiting ASE across three tissues. I Collinearity analysis showing a ~ 246 kb deletion on chromosome 9 of HapA, with associated simple repeats (SRs) and transposable elements (TEs). J Ultra-long ONT read alignments confirming the ~250 kb deletion, supported by approximately half the read depth. K Distribution of NLR gene types across four Rhamnaceae species. L Maximum-likelihood phylogenetic tree of RNLs from four Rhamnaceae species.
Transcriptome profiling revealed that the 853 NLRs were predominantly expressed in roots. Within the moderate expression range (FPKM 10–100), root tissue showed higher expression levels compared to leaf and stem (Fig. 4B and Supplementary Data 10), a pattern further supported by ternary plot analysis of all NLRs (Fig. 4C). Overall, 209 NLR genes exhibited expression above the global mean (FPKM = 5.46, calculated across nine samples: three tissues × three replicates) (Supplementary Data 9). To assess tissue-specific expression, we averaged FPKM values across three replicates per tissue and identified NLRs exceeding the global mean in any tissue, confirming the root-enriched expression of highly expressed NLRs (Fig. 4D). Furthermore, only one NLR, RNL1, was consistently expressed across all tissues (FPKM > 100), with exceptionally high expression in roots (average FPKM 893 in HapA and 700 in HapB), more than fivefold higher than in leaf and stem, providing a striking example of root-dominant expression (Supplementary Data 9).
We next examined the association between transposable elements (TEs) and NLR subfamilies, focusing on the upstream 2 kb, gene body, and downstream 2 kb regions of each gene. Unlike the genome-wide pattern dominated by Gypsy elements (Supplementary Data 2), Copia elements were the predominant TE type across all NLR subfamilies in both HapA and HapB (Fig. 4E and Supplementary Data 11). Among the four subfamilies, TNLs harbored significantly more TEs than the others, particularly compared with CNLs (Fig. 4F and Supplementary Data 11). Consistent with this pattern, CNL expression appeared largely unaffected by TE abundance across the three tissues (Supplementary Fig. 17), whereas TNL expression showed a clear negative correlation with TE abundance in stem and leaf, with higher TE density associated with reduced expression (Supplementary Fig. 17a, b). Interestingly, this correlation was absent in root (Supplementary Fig. 17c), suggesting that root-specific regulatory mechanisms may buffer against TE-mediated repression and thereby sustain the dominant activity of NLRs in this tissue.
Comparison of allelic versus non-allelic NLRs revealed significantly higher expression in allelic pairs across CNLs, NLs, and TNLs (Fig. 4G, Supplementary Data 9 and Supplementary Data 12). ASE analysis further supported root as the dominant tissue for NLR activity, with the largest number of ASEs detected in this organ (Fig. 4H). A striking structural example was a~246 kb deletion on chromosome 9 of HapB (22.52–22.77 Mb in HapA), which eliminated 13 tandem NLRs (10 NLs [NL162–NL171] and 3 CNLs [CNL139–CNL141]) (Fig. 4I, J). None of these genes was expressed, providing a clear case of reduced expression in non-allelic NLRs. We further examined the distribution of simple repeats and transposable elements within and outside this region but found no obvious differences (Fig. 4I), suggesting that other mechanisms may underlie the transcriptional silencing of these non-allelic NLRs.
Finally, interspecific comparison of NLR repertoires in Zju, Rru, and Sth revealed marked variation, with 566, 346, and 68 NLRs, respectively (Fig. 4K, Supplementary Data 13 and 14). The extremely low number in Sth may reflect its smaller genome size (198 Mb; Supplementary Data 7). Interestingly, despite its reduced repertoire, Sth retained a comparable number of RNLs to cointree, whereas only a single RNL was detected in both Zju and Rru (Fig. 4L). This unusual pattern suggests a distinct evolutionary trajectory of RNLs in Rhamnaceae and highlights them as promising targets for future functional studies.
Expression and evolution of the AsA metabolic pathway
We identified 91 genes involved in AsA metabolism in the cointree genome, including 48 in HapA and 43 in HapB. Of these, 28 HapA and 24 HapB genes were assigned to biosynthesis pathways, 12 and 11 to oxidation, and 8 and 8 to regeneration (Fig. 5A and Supplementary Data 15). Among the four known AsA biosynthesis pathways, the L-galactose and D-galacturonic acid pathways predominated across root, stem, and leaf, with all key enzymes identified and expressed. Biosynthetic genes showed significantly higher mean expression in stem and leaf compared with root (Fig. 5B and Supplementary Data 16), consistent with AsA content, which was highest in leaf (Fig. 5C and Supplementary Data 17).
A Schematic of the AsA metabolic pathway with a heatmap of gene expression in root, stem, and leaf (left to right). Key enzymes include GDP-L-galactose phosphorylase (GGP), L-galactose-1-phosphate phosphatase (GPP), L-galactose dehydrogenase (GDH), L-galactono-1,4-lactone dehydrogenase (GLDH), D-galacturonic acid reductase (GalUR), aldonolactonase (Alase), dehydroascorbate reductase (DHAR), monodehydroascorbate reductase (MDHAR), L-ascorbate oxidase (AO), and L-ascorbate peroxidase (APX). Expression values represent the mean of three biological replicates, log2-transformed and normalized to z-scores for heatmap visualization. Genes are grouped into four principal AsA biosynthetic pathways. B Aggregate expression levels of AsA pathway genes (n = 91) across root, stem, and leaf. C AsA content in root, stem, and leaf based on three biological replicates. Error bars represent the standard deviation (SD) of three biologically independent replicates (n = 3). For B and C, significance was assessed using Student’s t-test; pairs without labeled p-values are not significantly different. D Heatmap of pairwise log2FC comparisons of FPKM values among tissues for MDHAR and DHAR genes. Log2FC values are shown in each grid. Rows were clustered using Pearson correlation distance, and columns were clustered using Ward’s criterion. E Maximum-likelihood phylogenetic tree of MDHAR genes from 20 representative species. Six major clades are labeled; genes from different species are color-coded, with cointree genes highlighted by a red dotted line. F Collinearity plot of tandemly duplicated MDHAR genes in Rhamnaceae compared with other Rosales species and two high–AsA species (citrus and kiwifruit). MDHAR genes are highlighted in orange, and red curves link the key gene (MDHARs3) from cointree tandem clusters to its orthologs in other Rosales species.
In the L-galactose pathway, GME and GPP showed >3-fold higher expression in stem and leaf compared to root. In the D-galacturonic acid pathway, only GaLUR13 displayed root-specific expression among the 14 GaLUR genes. In the oxidation pathway, AO1 and AO4 showed lowest expression in root, with AO4 absent from root. Among the seven APX genes, APX2, APX3, and APX7 displayed strong tissue specificity, with high expression in aerial tissues but little or none in root. In the regeneration pathway, two DHAR and six MDHAR genes were identified. DHAR1 was leaf-preferred, while DHAR2 showed ~fivefold higher expression overall, especially in stem. Among MDHARs, MDHAR3 was the most highly expressed across all tissues, followed by MDHAR1. By contrast, MDHAR2 was root-preferred, and three other members (s1–s3) showed very low expression (Fig. 5A, D and Supplementary Data 15). Overall, only GaLUR13 and MDHAR2 displayed root-enriched expression, consistent with the relatively low AsA accumulation in root. Notably, although stem exhibited gene expression comparable to leaf, its AsA content was unexpectedly lower, even below root (though not significantly), suggesting additional regulation or metabolic mechanisms.
Comparative analysis revealed lineage-specific expansion of the MDHAR family in Rosales. Phylogenetic analysis of MDHAR genes from 20 species (Supplementary Data 18), including high–AsA species (jujube, chestnut rose, kiwifruit, pepper, orange), grouped MDHARs into three conserved orthologous clades (Clades 1–3). In Rosales, additional lineage-specific expansions were detected, forming three separate clades: a Rhamnaceae-specific tandem expansion (Clade 4), a Rosaceae-specific expansion (Clade 6), and an expansion shared across other Rosales species (Clade 5). The Rhamnaceae-specific cluster was most pronounced, with jujube showing the largest expansion (13 genes), consistent with our previous reports3. In contrast, high–AsA species outside Rosales expanded only within conserved clades. For example, kiwifruit expanded across all three conserved clades, chestnut rose expanded in Clade 2, and pepper and orange showed no expansion (Fig. 5E). Collinearity analysis confirmed that MDHAR tandem clusters are unique to Rhamnaceae, with the key cluster member showing clear orthologous relationships in corresponding collinear regions of other Rosales species (red curve in Fig. 5F). In contrast, only weak synteny was observed in more distant lineages such as citrus and kiwifruit, with no evidence of MDHAR tandem cluster expansion (Fig. 5F).
Discussion
Cointree is valued for its distinctive sombrero-shaped winged fruits (samaras), its adaptability in cultivation, and its economic and ornamental importance1,2,4,21. Yet, despite these attributes, genomic resources for this species have been lacking. Here, we present the fully haplotype-resolved T2T and gapless genome assembly for cointree, a representative for the genus Paliurus within Rhamnaceae. This high-quality genome establishes a robust foundation for exploring genome evolution, deciphering genes linked to stress tolerance, and advancing breeding and conservation strategies.
T2T genomes have enabled precise characterization of centromeric architecture in plants22. Our recent study in jujube revealed a centromere architecture dominated by LTR-RTs17. In cointree, bioinformatic predictions indicated centromeres of 100 kb–2 Mb, with TRs, common in species such as Arabidopsis23 and soybean24, restricted to chromosomes 2, 3, 8, and 11 (Supplementary Fig. 3). Co-localization of CRJs with TR-enriched regions suggests a TE-dominated centromere organization in cointree. These findings suggest a TE-dominated centromere organization similar to jujube, a hypothesis that requires confirmation by ChIP-seq with a cointree-specific CENH3 antibody.
Haplotype-resolved assembly enabled the ASE analysis, which can reveal regulatory variation underlying trait differences in plants25. In cointree, 829 significant ASEs were enriched for defense- and stress-response functions (Supplementary Fig. 6). This is consistent with ASE-driven immune modulation reported in other plants26,27. Allelic expression divergence positively correlated with sequence variation, especially in promoter regions. Notably, 249 ASEs with extensive variation were enriched in the arginine biosynthesis pathway (Supplementary Fig. 7), previously linked to nitrogen-deficiency tolerance via photosynthesis and amino acid metabolism regulation28. A clear example is an allelic pair of argininosuccinate synthase genes: PhAASS1 (functional in HapA) and a nonfunctional HapB allele disrupted by a 285 bp insertion. This case highlights how structural variation drives ASE, underscoring the potential role of allelic asymmetry in metabolic flexibility.
Comparative analyses show that Rhamnaceae genomes are relatively conserved within Rosales. All four examined Rhamnaceae species share a haploid chromosome number of 12 and modest genome sizes (~285 Mb on average; Supplementary Data 7), contrasting sharply with the more dynamic genome evolution observed in Rosaceae, for example, apple (~742 Mb), which experienced a lineage-specific whole-genome duplication followed by extensive chromosomal rearrangements29. The Ks and 4DTv analyses revealed minor species-specific duplication peaks across Rhamnaceae, consistent with our previous findings in jujube3, and suggesting the occurrence of small-scale duplications that may contribute incrementally to genome expansion. These findings contrast with the large-scale genome expansions driven by whole-genome duplications in other lineages30, reflecting distinct evolutionary trajectories within Rosales and underscoring the need for broader comparative genomic analyses to clarify the mechanisms and evolutionary significance of these patterns.
Plant NLR genes are central to innate immunity, enabling rapid recognition and response to diverse pathogens6. Our transcriptome analyses revealed pronounced root-preferential expression of NLRs in cointree, particularly among moderately to highly expressed genes. This root-biased pattern aligns with a cross-lineage meta-analysis showing root-enriched NLR expression in most non-Brassicaceae species31. As roots are the primary interface with soilborne pathogens and abiotic stressors, elevated root NLR expression reflects an adaptive strategy for robust belowground immune surveillance. Root-specific NLR clusters, such as those in tomato, evolved for nematode defense32, further support this organ-specific functional specialization. Additionally, the higher expression of allelic versus non-allelic NLR genes suggests a regulatory advantage of retaining both haplotypes. A notable case is the ~250 kb deletion in HapB, which resulted in complete loss of expression for the corresponding NLRs in HapA. This mirrors findings in pear, where haplotype-specific genes exhibited significantly lower expression than bi-allelic genes33. Such allelic diversity likely enhances transcriptional robustness and functional plasticity, enabling more effective immune responses under variable environmental conditions. Structural variation, including deletions and promoter divergence, emerges as a key driver of differential expression and a potential force in the evolutionary maintenance of NLR gene pairs33,34.
AsA is a central antioxidant that enhances plant tolerance to multiple stresses, including salinity, drought, and heat, by scavenging reactive oxygen species (ROS) and maintaining redox homeostasis35. In cointree, AsA accumulation was highest in leaf, consistent with the elevated expression of biosynthetic and regeneration genes in aerial tissues. This pattern reflects the high demand for ROS detoxification in photosynthetically active tissues36. By contrast, stem exhibited comparable expression levels to leaf but unexpectedly lower AsA content, suggesting that additional regulatory layers, such as substrate availability, enzyme activity, or transport, may contribute to tissue-specific AsA accumulation, as reported in kiwifruit and tomato37,38. The evolutionary history of MDHAR genes provides further insights. We observed Rhamnaceae-specific tandem expansion, with the largest expansion in jujube and cointree. Given that MDHAR plays a key role in AsA regeneration from its oxidized forms, such expansion may contribute to maintaining AsA pools under stress conditions. Interestingly, other high–AsA species outside Rosales (e.g., citrus, kiwifruit) did not show MDHAR expansions, indicating that different lineages may have evolved distinct strategies for sustaining high AsA levels. For instance, kiwifruit has been reported to increase AsA through enhanced biosynthesis39. Together, our findings highlight both conserved and lineage-specific mechanisms shaping AsA metabolism. In cointree and related Rhamnaceae, tandem duplication of MDHAR may represent a unique evolutionary route to reinforce AsA recycling capacity, complementing biosynthetic pathways. Future functional studies, such as enzyme activity assays and stress-induction experiments, will be essential to test the adaptive significance of these expansions.
In conclusion, this study presents a haplotype-resolved telomere-to-telomere genome assembly of Paliurus hemsleyanus, providing a high-quality reference for the genus Paliurus and the family Rhamnaceae. Our analyses reveal a relatively stable genome structure within Rhamnaceae, root-preferred expression of NLR genes, and a positive correlation between allele-specific expression and allelic divergence. Tissue-specific profiling of AsA metabolism highlights leaf as the primary site for AsA accumulation, and a lineage-specific tandem expansion of MDHAR genes occurred in Rhamnaceae. These findings advance our understanding of genomic organization, gene expression, and evolutionary patterns in cointree, offering a valuable resource for future comparative genomics, functional studies, and breeding efforts.
Methods
Sample preparation and sequencing
Botanical samples of cointree were preserved at the Chinese Jujube Research Centre, Hebei Agricultural University. Genomic DNA was extracted from young leaf tissues using a modified cetyltrimethylammonium bromide (CTAB) method40, in which the selective precipitation and CsCl gradient steps were omitted to reduce cost and reagent toxicity compared with the original protocol. Genome sequencing was conducted using three platforms: PacBio HiFi, Oxford Nanopore Technologies (ONT), and MGISEQ-2000. For PacBio HiFi sequencing, high-molecular-weight DNA was sheared into 15–18 kb fragments using a g-TUBE (Covaris, MA, USA), and SMRTbell libraries were prepared with the SMRTbell Express Template Preparation Kit 2.0 (PacBio, CA, USA) according to the manufacturer’s protocol. For ONT sequencing, libraries were prepared and sequenced on a PromethION platform (Oxford Nanopore Technologies, UK). Reads with Q scores greater than 7 were retained after base calling and quality filtering of the raw FASTQ files. For short-read sequencing (NGS), paired-end libraries with insert sizes of 200–400 bp were constructed and sequenced on the MGISEQ-2000 platform following standard procedures. In Hi-C sequencing, approximately 2 g of fresh leaves were used for library construction. The procedure involved formaldehyde fixation, chromatin cross-linking, nuclei isolation, restriction enzyme digestion, DNA ligation, end-repair, purification, and quantification41. The resulting library was sequenced on the MGISEQ-2000 platform, and quality control was performed following the same procedure as for paired-end sequencing.
RNA sequencing and data analysis
For RNA-seq, root, stem, and leaf were harvested separately from the same cointree individual with three biological replications. RNA for each sample was collected using a plant RNA isolation kit (Tiangen Biotechnology Co.). Libraries were prepared using the TruSeq RNA Library Preparation Kit (Illumina, USA) following the manufacturer’s instructions. Brief procedures include mRNA purification using oligo poly-T probes, cDNA synthesis, adaptor ligation, size selection and purification, PCR, PCR product purification, and library quality evaluation. Finally, the library was sequenced on an Illumina Novaseq platform to obtain 150 bp paired-end reads.
The raw paired-end RNA-seq reads were first performed for quality control using fastp42. Then the clean reads were mapped to each of the two haplotypes of the cointree genome using STAR (v2.7.10) with default parameters43. The result BAM files for each haplotype were used as the input to the RSEM44 to calculate gene expression levels in fragments per kilobase of exon per million mapped reads (FPKM). DEGs were identified with DESeq245, using thresholds of |log₂ fold change| > 1 and p < 0.05.
T2T genome assembly and evaluation
Genome size was estimated from clean NGS reads using Jellyfish (v2.3.1)46 and GenomeScope (V2.0)47. Haplotype-resolved genome assembly was performed using Hifiasm (v0.19.8-r603) with parameters “--ul-cut 20000 -D10 --hom-cov 31” based on PacBio HiFi, ONT, and Hi-C data48. Redundant overlaps in contigs were removed using Purge_Dups (v1.2.5)49, and the resulting de-redundant contigs of HapA and HapB were together used to construct chromosome-level assemblies with YaHS (v1.2)50 using Hi-C data. Initial gap filling was performed with quartTET (v1.2.5)16, and remaining gaps were manually closed using the HiFi reads. Assembly quality and completeness were evaluated using BUSCO (v5.4.4) with the embryophyta_odb1051 and Merqury (V1.3)52. LTR-RTs were annotated using GenomeTools (v1.6.5)53 and LTR_FINDER_parallel (V1.3)54, and the LAI was calculated with LTR_retriever (v3.0.2)55 to assess genomic continuity.
Genome annotation
Interspersed repetitive elements were identified using both ab initio prediction and homology-based methods. RepeatModeler56 was first employed to construct a species-specific repeat library, which was then combined with the Repbase database (http://www.girinst.org/repeatbase) as input to RepeatMasker57 for genome-wide repeat annotation.
Protein-coding gene prediction was performed using an integrative approach combining homology-based, transcriptome-based, and ab initio methods. Homologous protein sequences from Malus domestica, Arabidopsis thaliana, Populus przewalskii, Prunus persica, Prunus armeniaca, and Pyrus pyrifolia were aligned to the genome using GeMoMa58, and transcripts were assembled with StringTie43, and then transcripts were assembled using StringTie59. These assemblies were input into PASA for gene model prediction60. For ab initio prediction, the StringTie-assembled transcripts were used to train Augustus61, which generated a gene model optimized for cointree. The outputs from all approaches were integrated using EVidenceModeler (EVM)60. Gene models located exclusively within duplicated regions were removed based on whole-genome alignment, and transposon-related genes were filtered using TransposonPSI62.
Identification of telomere and centromere sequences
Telomeric sequences were identified by scanning the genome for the typical plant telomeric repeat (5′-CCCCTAAA/TTTAGGG-3′) using the Seqkit software63. Centromeres for both haplotypes were detected using quartTET with default parameters.
Identification of inter-haplotype variation
The two haplotypes were aligned using Nucmer from the MUMmer suite (v4)64 with the parameters --maxmatch -c 100 -l 100. SNPs and Indels were identified using the delta-filter and show-coords subroutines. Structural variants (SVs), including inversions, translocations, and duplications, were detected by aligning the haplotypes with Minimap265, and the resulting alignment was analyzed using SyRi (v1.6.1)66.
Gene family clustering and phylogenetic tree construction
Single-copy orthologous genes among cointree and 18 representative angiosperms were identified using OrthoFinder (v2.5.5)67 with default parameters. Phylogenetic trees were constructed based on these genes using IQ-TREE2 (v2.2.0)68. Divergence times were estimated with MCMCTREE (v4.9)69, calibrated using fossil-based divergence times between Glycine and Vitis (109-124 Mya), as well as between Gylcine and Pyrus (94-104 Mya), from TimeTree (http://www.timetree.org/). MCMCTREE was run for 5,000,000 iterations with a sampling frequency of 30. Gene family expansion and contraction were inferred using CAFE5 (v1.1)70 based on the inferred phylogeny.
Ancestral karyotype reconstruction
The overall pipeline is shown in Supplementary Fig. 9. Briefly, the wgdi pipeline71 was used to infer ancestral karyotypes for Rosaceae (six species) and Rhamnaceae (four species) families. Pairwise comparisons of genome protein sequences were first conducted using BLAST, and the results were used to generate collinearity dot plots72 with the -d parameter in wgdi. After removing redundant comparisons, two representative pairs in Rosaceae (Mdo vs. Aan and Ppe vs. Rch) and one in Rhamnaceae (Phe vs. Sth) were retained for analysis. From these comparisons, chromosome-level collinear blocks identified from the dot plots were regarded as ancestral chromosomes, denoted as RAK (ancestral Rhamnaceae karyotype) and ARK (ancestral Rosaceae karyotype) (Supplementary Fig. 10). Protein sequences of RAK and ARK were then reconstructed using the parameters -icl, -bi, -c, -km, and -k. To further refine RAK and ARK, their sequences were aligned with the four Rhamnaceae species and six Rosaceae species, respectively, using the -k parameter, which produced the final karyotypes of RAK and ARK (Supplementary Fig. 11). Finally, the seven published Ancestral Eudicot Karyotype (AEK) sequences20 were used as a reference to determine the karyotype diagrams of RAK, ARK and the extant species.
Collinearity analysis and calculation of Ks and 4DTv
Collinearity analysis of both paralogous and orthologous genes was performed using McSCan73. Homologous CDS sequences from the MCScan results were aligned using ParaAT (v2.0)74, and Ks were estimated with KaKs_Calculator (v2.0)75, using the YN model. The 4DTv was calculated using the Perl script Calculate_4DTV_correction.pl3,76,77.
Identification of NLR genes and AsA genes
NLR genes in each species were identified using NLR-Annotator (v2)78, followed by InterProScan to confirm the presence of NB-ARC (PF00931), TIR (PF01582), or RPW8 (PF05659) domains. Conserved domain sequences were aligned with MAFFT, and UMAP-based dimensionality reduction was performed using the R packages seqinr, umap, and Rtsne. The ternary plot in Fig. 4C is referred the study of oat genome79. AsA pathway, genes were identified through functional annotation and sequence alignment, with Pfam and SMART used to confirm conserved structural domains.
Statistics and reproducibility
Gene expression comparisons between haplotypes (Fig. 4G) and between different tissues (Fig. 5B) were performed using Student’s t-test. KEGG and GO enrichment analyses were performed using hypergeometric tests to identify significantly enriched pathways and GO terms among the DEGs relative to the whole-genome background. Ks calculations with KaKs_Calculator utilized Fisher’s exact test to assess statistical significance.
Whole-genome sequencing was conducted using leaf tissue without biological replicates. For RNA-seq, root, stem, and leaf tissues were each sampled with three biological replicates, resulting in a total of nine samples. Biological replicates were defined at the sampling stage; for example, different parts of the root from a single individual tree were independently sampled three times for sequencing.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data from this project, including the genome assembly, and raw sequencing reads, have been deposited in the National Genomics Data Center (NGDC) under project number PRJCA038194. The biological sample accession numbers for root, stem, and leaf are SAMC5008325, SAMC5008326, and SAMC4946747, respectively. In addition, the haplotype-resolved genome assemblies have been deposited in the NCBI database under project numbers PRJNA1338467 (HapA) and PRJNA1338466 (HapB), with all raw sequencing reads available under PRJNA1338467. The haplotype-resolved genome assemblies, GFF3 annotation files, protein and coding sequence datasets for both haplotypes, as well as the ARK and RAK sequences and their corresponding protein and coding sequences for all species analyzed in this study, are also available in the Figshare repository80. The gene information for 829 differentially expressed allelic genes, along with 249 genes showing extensive variation (defined as carrying structural variants or more than the average number of SNPs or small indels), is provided in Supplementary Data 5. Gene IDs, names, and expression levels (FPKM) from three biological replicates of root, stem, and leaf tissues for both NLR and AsA metabolic genes are available in Supplementary Data 9, 14, 15, and 18. Raw data file for Figs. 2B–E, F–H, 3B, 4B, E–G, K, and 5C, and Supplementary Fig. 17 were provided in Supplementary Data 4, 5, 6, 8, 10, 11, 12, 13, 16, 17, and 9, respectively. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The codes used in this study are available on GitHub at https://github.com/Zhang-Shufeng/cointree-genome, have also been deposited in the Zenodo repository81.
References
Johnson, O. ‘Paliurus hemsleyanus’ from the website Trees and Shrubs Online. https://www.treesandshrubsonline.org/articles/paliurus/paliurus-hemsleyanus/ (2024).
Lee, S.-S., Shy, S.-N. & Liu, K. C. Triterpenes from Paliurus hemsleyanus. Phytochemistry 46, 549–554 (1997).
Yang, M. et al. Insights into the evolution and spatial chromosome architecture of jujube from an updated gapless genome assembly. Plant Commun. 4, 100662 (2023).
Xu, Y. et al. Identification of high tolerance to jujube witches’ broom in Indian jujube (Ziziphus mauritiana Lam.) And mining differentially expressed genes related to the tolerance through transcriptome analysis. Plants 12, 2082 (2023).
Wu, T., Al-Mamun, H. A., Edwards, D., Batley, J. & Dolatabadian, A. Genome-wide identification and prediction of disease resistance genes in Hirschfeldia incana. Agric. Commun. 2, 100049 (2024).
Shepherd, S., Yuen, E. L. H., Carella, P. & Bozkurt, T. O. The wheels of destruction: plant NLR immune receptors are mobile and structurally dynamic disease resistance proteins. Curr. Opin. Plant Biol. 74, 102372 (2023).
Luo, Y. et al. Phased T2T genome assemblies facilitate the mining of disease-resistance genes in Vitis davidii. Horticult. Res. 12, uhae306 (2025).
Gu, L., Si, W., Zhao, L., Yang, S. & Zhang, X. Dynamic evolution of NBS-LRR genes in bread wheat and its progenitors. Mol. Genet. Genom. 290, 727–738 (2015).
Sharma, D. et al. A single NLR gene confers resistance to leaf and stripe rust in wheat. Nat. Commun. 15, 9925 (2024).
Hao, Q. et al. A pair of atypical NLR-encoding genes confers Asian soybean rust resistance in soybean. Nat. Commun. 15, 3310 (2024).
von Dahlen, J. K., Schulz, K., Nicolai, J. & Rose, L. E. Global expression patterns of R-genes in tomato and potato. Front. Plant Sci. 14, 1216795 (2023).
Quinones, C. O., Gesto-Borroto, R., Wilson, R. V., Hernandez-Madrigal, S. V. & Lorence, A. Alternative pathways leading to ascorbate biosynthesis in plants: lessons from the last 25 years. J. Exp. Bot. 75, 2644–2663 (2024).
Xiao, M. et al. The multiple roles of ascorbate in the abiotic stress response of plants: antioxidant, cofactor, and regulator. Front. Plant Sci. 12, 598173 (2021).
Liu, X., Bulley, S. M., Varkonyi-Gasic, E., Zhong, C. & Li, D. Kiwifruit bZIP transcription factor AcePosF21 elicits ascorbic acid biosynthesis during cold stress. Plant Physiol. 192, 982–999 (2023).
Chen, X. et al. Ascorbic acid mitigates salt stress in tomato seedlings by enhancing chlorophyll synthesis pathways. Agronomy 14, 1810 (2024).
Lin, Y. et al. quarTeT: a telomere-to-telomere toolkit for gap-free genome assembly and centromeric repeat identification. Hortic. Res. 10, uhad127 (2023).
Lin, D. et al. Structural composition and evolution of jujube centromere reveal a dominant role for LTR Retrotransposons. Hortic. Res. 12, uhaf244 (2025).
Nowicka, A. et al. The transcriptional landscape of the developmental switch from regular pollen maturation towards microspore-derived plant regeneration in barley. Crop J. 12, 1064–1080 (2024).
Yang, X., Wang, X., Yao, J. & Duan, D. Dynamic chromatin aaccessibility underlies generation-biased gene expression in the brown alga Ssaccharina japonica. Plant Cell Environ. 9 (2025).
Wang, Z. et al. A high-quality Buxus austro-yunnanensis (Buxales) genome provides new insights into karyotype evolution in early eudicots. BMC Biol. 20, 216 (2022).
Wu, Z., Raven, P. H. & Hong, D. Flora of China. Sci. Press 12, 1–534 (2007).
Naish, M. & Henderson, I. R. The structure, function, and evolution of plant centromeres. Genome Res. 34, 161–178 (2024).
Naish, M. et al. The genetic and epigenetic landscape of the Arabidopsis centromeres. Science 374, eabi7489 (2021).
Liu, Y. et al. Pan-centromere reveals widespread centromere repositioning of soybean genomes. Proc. Natl. Acad. Sci. USA 120, e2310177120 (2023).
Guo, M. et al. Genome-wide allele-specific expression analysis using Massively Parallel Signature Sequencing (MPSS) reveals cis- and trans-effects on gene expression in maize hybrid meristem tissue. Plant Mol. Biol. 66, 551–563 (2008).
Zhang, X. et al. Haplotype-resolved genome assembly provides insights into evolutionary history of the tea plant Camellia sinensis. Nat. Genet. 53, 1250–1259 (2021).
Yang, H. et al. A haplotype-resolved gap-free genome assembly provides novel insight into monoterpenoid diversification in Mentha suaveolens ‘Variegata. Hortic. Res. 11, uhae022 (2024).
Chen, Q. et al. Arginine increases tolerance to nitrogen deficiency in malus hupehensis via alterations in photosynthetic capacity and amino acids metabolism. Front. Plant Sci. 12, 772086 (2021).
Jung, S. et al. Whole genome comparisons of Fragaria, Prunus and Malus reveal different modes of evolution between Rosaceous subfamilies. BMC Genom. 13, 129 (2012).
Ren, R. et al. Widespread whole genome duplications contribute to genome complexity and species diversity in angiosperms. Mol. plant 11, 414–428 (2018).
Munch, D. et al. The brassicaceae family displays divergent, shoot-skewed NLR resistance gene expression. Plant Physiol. 176, 1598–1609 (2018).
Lüdke, D. et al. A root-specific NLR network mediates immune signaling of resistance genes against plant parasitic nematodes. Plant Cell 37, koaf145 (2025).
Sun, M. et al. Haplotype-resolved, gap-free genome assemblies provide insights into the divergence between Asian and European pears. Nat. Genet. 57, 2040–2051 (2025).
Yu, H. et al. Haplotype-resolved genome assembly provides insights into the genetic basis of green peach aphid resistance in peach. Curr. Biol. 35, 2614–2629.e5 (2025).
Anjum, N. A., Umar, S. & Chan, M.-T. Ascorbate-Glutathione Pathway And Stress Tolerance In Plants (Springer Science & Business Media, 2010).
Foyer, C. H. & Noctor, G. Ascorbate and glutathione: the heart of the redox hub. Plant Physiol. 155, 2–18 (2011).
Bulley, S. et al. Enhancing ascorbate in fruits and tubers through over-expression of the L-galactose pathway gene GDP-L-galactose phosphorylase. Plant Biotechnol. J. 10, 390–397 (2012).
Mellidou, I. & Kanellis, A. K. Genetic control of ascorbic acid biosynthesis and recycling in horticultural crops. Front. Chem. 5, 50 (2017).
Liu, X., Wu, R., Bulley, S. M., Zhong, C. & Li, D. Kiwifruit MYBS1-like and GBF3 transcription factors influence l-ascorbic acid biosynthesis by activating transcription of GDP-L-galactose phosphorylase 3. N. Phytol. 234, 1782–1800 (2022).
Allen, G. C., Flores-Vergara, M. A., Krasynanski, S., Kumar, S. & Thompson, W. F. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat. Protoc. 1, 2320–2325 (2006).
Belton, J.-M. et al. Hi–C: a comprehensive technique to capture the conformation of genomes. Methods 58, 268–276 (2012).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 1–16 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Zhou, C., McCarthy, S. A. & Durbin, R. YaHS: yet another Hi-C scaffolding tool. Bioinformatics 39, btac808 (2023).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Gremme, G., Steinbiss, S. & Kurtz, S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE/ACM Trans. Comput Biol. Bioinform. 10, 645–656 (2013).
Ou, S. & Jiang, N. LTR_FINDER_parallel: parallelization of LTR_FINDER enabling rapid identification of long terminal repeat retrotransposons. Mob. DNA 10, 48 (2019).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Plant Physiol. 176, 1410–1422 (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Chen, N. Using Repeat Masker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinforma. 5, 1–4.10. 14 (2004).
Keilwagen, J., Hartung, F. & Grau, J. GeMoMa: homology-based gene prediction utilizing intron position conservation and RNA-seq data. Methods Mol. Biol. 1962, 161–177 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using evidencemodeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, W465–W467 (2005).
Haas, B. TransposonPSI: an application of PSI-blast to mine (Retro-)Transposon ORF homologies. https://transposonpsi.sourceforge.net (2010).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Marcais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Mendes, F. K., Vanderpool, D., Fulton, B. & Hahn, M. W. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics 36, 5516–5518 (2020).
Sun, P. et al. WGDI: a user-friendly toolkit for evolutionary analyses of whole-genome duplications and ancestral karyotypes. Mol. Plant 15, 1841–1851 (2022).
Tang, H. & Lyons, E. Unleashing the genome of Brassica rapa. Front. Plant Sci. 3, 172 (2012).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49–e49 (2012).
Zhang, Z. et al. ParaAT: a parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 419, 779–781 (2012).
Wang, D., Zhang, Y., Zhang, Z., Zhu, J. & Yu, J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom. Proteom. Bioinforma. 8, 77–80 (2010).
Hasegawa, M., Kishino, H. & Yano, T. -a Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 22, 160–174 (1985).
Shen, Y. et al. Chromosome-level and haplotype-resolved genome provides insight into the tetraploid hybrid origin of patchouli. Nat. Commun. 13, 3511 (2022).
Steuernagel, B. et al. The NLR-annotator tool enables annotation of the intracellular immune receptor repertoire. Plant Physiol. 183, 468–482 (2020).
Kamal, N. et al. The mosaic oat genome gives insights into a uniquely healthy cereal crop. Nature 606, 113–119 (2022).
Shufeng, Z. et al. https://doi.org/10.6084/m9.figshare.30444779 (2025).
Shufeng, Z. et al. https://doi.org/10.5281/zenodo.17509296 (2025).
Acknowledgements
This work was supported by the Projects of the General Program of the National Natural Science Foundation of China (Grant No. 32171817); the Special Research Projects for the New Talent of Hebei Agricultural University, Hebei Province, China (Grant No. YJ2020025).
Author information
Authors and Affiliations
Contributions
M.Y. and M.L. conceived and designed the project. S.Z., Y.Y., and J.Z. assisted with sample collection. S.Z., J.G., D.L., M.Y., and J.S. performed genome assembly and gene annotation. Y.Lan, Y.Y. and Z.Z. performed transcriptome analyses. J.L., Y.L., Y.Z., D.L., and Z.Z. assisted with bioinformatics analyses and data visualization. S.Z., M.Y., and J.G. wrote the manuscript. M.Y. revised the manuscript. All authors have reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Tao Shi, Yongzhi Yang, Huilong Du and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Tobias Goris and David Favero. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, S., Guo, J., Lin, D. et al. Haplotype-resolved t2t genome of paliurus hemsleyanus provides insights into rhamnaceae evolution and genome biology. Commun Biol 8, 1744 (2025). https://doi.org/10.1038/s42003-025-09283-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09283-2