Abstract
Neutrophils are the most abundant leukocytes in human peripheral blood, yet their heterogeneity in pregnancy, especially in gestational diabetes mellitus (GDM), remains incompletely understood. Here, we employed InfinityFlow-based surface marker profiling and single-cell RNA sequencing (scRNA-seq) to delineate neutrophil subsets in healthy and GDM pregnancies. We identified a low-density, immature subgroup (CD10⁻CD49d⁺Ig κ⁺) with distinctive morphology and transcriptomic profiles, contrasted against the CD10⁺ segmented mature neutrophils. In healthy pregnancy, these immature low-density neutrophils (LDNs) expanded by mid-gestation, elevating the immature-to-mature (I–M) neutrophil ratio and circulating myeloid progenitor levels throughout gestation. In GDM, this expansion was markedly blunted, with a consistently lower abundance of immature LDNs and progenitors. Flow cytometry and correlation analyses further linked the lower I–M ratio to impaired insulin sensitivity, underscoring a potential immune-metabolic axis in GDM. Collectively, our study provides a framework for neutrophil phenotyping in pregnancy and links disrupted neutrophil equilibrium and pregnancy complications.
Similar content being viewed by others
Introduction
Neutrophils are the most abundant leukocytes in human peripheral blood, accounting for 50–70% of circulating white blood cells1. As first‑line responders to microbial and sterile stimuli, these cells play a vital role in maintaining homeostasis and orchestrating innate immune responses2,3. The long-standing view of neutrophils as a homogeneous population has shifted, with distinct phenotypic and functional subsets now recognized4,5. Among these, low-density neutrophils (LDNs) represent a group of neutrophils with reduced buoyancy that co-segregate with peripheral blood mononuclear cells (PBMCs) during density-gradient separation6. Initially described in patients with systemic lupus erythematosus (SLE) and rheumatoid arthritis7, LDNs have since been linked to solid cancers8,9,10, infections11,12,13, inflammatory disorders14,15,16, and obesity17. However, the precise nature and roles of LDNs remain incompletely understood. LDN populations may comprise immature neutrophil precursors, degranulated mature neutrophils, immunosuppressive polymorphonuclear myeloid-derived suppressor cells (PMN-MDSCs), or a mixture thereof. Notably, pre-analytical variables such as delays in sample processing or the choice of anticoagulant can artifactually inflate apparent LDN frequencies6. These considerations underscore the need for robust marker combinations that define LDNs by intrinsic phenotype rather than density alone.
Pregnancy is a unique immunological state in which the maternal immune system adapts to tolerate the semi-allogeneic fetus. Neutrophils participate in pregnancy by providing both pro-inflammatory (e.g., promoting trophoblast invasion and placental vascular remodeling) and immunoregulatory (e.g., maintaining tolerance at the maternal–fetal interface) functions18,19,20,21. Dysregulation of neutrophil number or function has been associated with pregnancy complications22,23. For example, gestational diabetes mellitus (GDM), a common metabolic disorder of pregnancy, is often accompanied by elevated neutrophil-to-lymphocyte ratios and an expansion of circulating neutrophils24,25. Notably, the emergence of LDNs has been reported in healthy pregnancies26. These cells are thought to support the tolerogenic environment, whereas aberrant LDN profiles have been linked to adverse outcomes. Recent studies suggest that immature or suppressive neutrophils expand in normal pregnancy but are reduced or phenotypically altered in disease states such as GDM, SLE, or miscarriage. For instance, spontaneous abortion has been associated with impaired activation of CD16high LDNs27, while pregnancies complicated by SLE exhibit increased proportions of activated LDNs accompanied by CD62L shedding28. These observations suggest that a deeper resolution of the LDN compartment into biologically meaningful subsets is crucial for understanding the immunometabolic dynamics of pregnancy.
Current knowledge of neutrophil heterogeneity in pregnancy, however, remains incomplete. Conventional markers, including CD66b29,30, CD1531, CD1032, CD11b33, and CD1634, define broad maturation stages but do not fully capture the spectrum of in vivo neutrophil states35. Additional marker combinations that distinguish truly immature cells from band-like or fully segmented neutrophils are needed to clarify their respective roles in maternal–fetal immune tolerance and pregnancy-related disease. In this study, we employed high-dimensional single-cell immunophenotyping (InfinityFlow) and single-cell RNA-seq (scRNA-seq) to delineate neutrophil subpopulations in healthy and GDM pregnancies. We identified a specific combination of surface markers that precisely distinguishes immature LDNs from mature high-density neutrophils (HDNs), and we confirmed that these immature neutrophils are also present at the maternal–fetal interface. Using these markers, we demonstrate that the typical expansion of circulating immature neutrophils during healthy pregnancy is markedly attenuated in GDM. Our findings provide a detailed phenotypic and transcriptomic description of neutrophil heterogeneity during gestation and highlight how dysfunctional neutrophils may contribute to pregnancy complications.
Results
InfinityFlow profiling of circulating neutrophil surface markers during pregnancy
Using density-gradient centrifugation, blood leukocytes can be separated into low-density PBMCs and high-density polymorphonuclear leukocytes (PMNLs, containing high-density neutrophils [HDNs]). In healthy pregnancies (HP), the frequency of Lin−(CD3−CD19−CD56−CD14−)CD45⁺CD15⁺ LDNs within the PBMC fraction was significantly higher than in non-pregnant controls (HC). This expansion was blunted in GDM (Supplementary Fig. 1A). To further investigate neutrophil heterogeneity and resolve the LDN phenotype in pregnancy, we performed a discovery screen with a highly multiplexed flow cytometry platform. Peripheral blood samples were collected from one healthy pregnant woman and one woman with GDM at 38 weeks of gestation. After density-gradient centrifugation, the PBMCs and PMNLs were barcoded with distinct hash antibodies, pooled, and then stained with a set of “backbone” markers (CD3, CD4, LOX-1, CD19, CD56, CD14, and CD15), plus a panel of 364 surface protein markers (Fig. 1A). The InfinityFlow pipeline36,37 was then used to quantify expression and co-expression of these surface proteins across leukocytes, enabling unbiased identification of marker combinations that delineate neutrophil subsets.
A Schematic overview of the InfinityFlow analysis on peripheral blood leukocytes from healthy and GDM pregnancies. Abs, antibodies. B UMAP dimensionality reduction and FlowSOM clustering of leukocytes based on backbone lineage markers, with distinct clusters representing T cells (CD3⁺), B cells (CD19⁺), NK cells (CD56⁺), monocytes (CD14⁺, Mono), and neutrophils (CD15⁺, Neu). C Neutrophils were split into LDNs and HDNs according to hash antibody staining and further analyzed on surface marker expression. D UMAP visualization of CD15+ cells in both healthy and GDM samples, showing LDNs and HDNs distribution based on the expression of 172 selected markers. E Conventional flow cytometry gating and demultiplexing strategy for identifying LDNs and HDNs. F Venn diagram illustrating the overlap of the top differentially expressed surface markers (upregulated or downregulated) in LDNs and HDNs across healthy and GDM pregnancies, derived from InfinityFlow data. G Conventional flow cytometry-based distribution of the top four discriminating markers (CD10, CD49d, CD63, and Ig κ) between LDNs and HDNs in healthy and GDM pregnancies.
We first distinguished major leukocyte lineages using the backbone markers, such as T cells (CD3⁺), B cells (CD19⁺), NK cells (CD56⁺), monocytes (CD14⁺), and neutrophils (CD15⁺) (Fig. 1B and Supplementary Fig. 1B–D). Since CD15+ neutrophils were present in both the high- and low-density populations, we focused on comparing the phenotypes of LDNs versus HDNs. Among the 364 surface markers, 172 were confirmed to be expressed on neutrophils based on isotype controls (Fig. 1C). Uniform manifold approximation and projection (UMAP) visualization using these 172 markers demonstrated that LDNs and HDNs formed distinct clusters in both healthy and GDM pregnancies, indicating clear phenotypic differences between the two neutrophil populations (Fig. 1D).
Differential expression analysis of the InfinityFlow results, validated by conventional flow cytometry, identified nine significantly upregulated (CD111, CD49d, CD63, CD64, CD66b, CD71, β2-microglobulin, Ig κ, and IgG Fc) and six significantly downregulated markers (CD10, CD367, CD85a, TNAP, TRA-2-49, and TRA-2-54) on LDNs compared with HDNs (Fig. 1E, F, Supplementary Fig. 1E and Supplementary Data 2). Notably, CD49d, Ig κ, and CD63 emerged as prominent markers for LDNs, whereas CD10 was the most discriminating marker for HDNs (Fig. 1G and Supplementary Fig. 1F). These findings define a surface-marker signature that can distinguish circulating LDNs from HDNs in pregnancy.
CD10-CD49d+Ig κ+ identify immature LDNs in pregnancy
We next examined expression of the four prominent markers—Ig κ, CD49d, CD63, and CD10—on circulating LDNs and HDNs in a larger cohort of pregnant women (n = 27; Fig. 2A and Supplementary Fig. 2A). Consistent with the InfinityFlow findings, LDNs had higher levels of Ig κ, CD49d, and CD63, whereas HDNs exhibited significantly increased levels of CD10 (Supplementary Fig. 2B). LOX-1, previously identified as an LDN/PMN-MDSC marker in cancer38, was also enriched in pregnancy-associated LDNs (Supplementary Fig. 2C, D). Although these differences were statistically significant, no single marker could completely discriminate LDNs from HDNs (Fig. 2B). However, combinations of these markers effectively distinguished the two neutrophil subsets (Fig. 2C–E and Supplementary Fig. 2E–G). Notably, CD10−CD49d+Ig κ+ neutrophils accounted for 85.7 ± 1.63% of LDNs, while 97.6 ± 0.18% of HDNs were CD10+CD49d−Ig κ− (Fig. 2F), indicating marked discriminating capability.
A Experimental workflow schematic illustrating the labeling and flow cytometry analysis of LDNs and HDNs using the indicated discriminating markers. B Representative histograms (upper) and statistical analysis (lower) of flow cytometric data for CD10, Ig κ, CD49d, and CD63 expression on LDNs and HDNs in pregnancies at 24–28 weeks of gestation (n = 27). Gray dashed lines indicate unstained (Neg) controls. C Flow cytometric gating strategy for demultiplexing HDNs and LDNs (left), with representative plots of CD10, CD49d, and Ig κ expression on LDNs and HDNs (middle); bar graphs (right) showing percentages of cells with different marker combinations in each subset. D Representative ROC curves for marker combinations in distinguishing LDNs from HDNs. E ROC curve analysis evaluating three-marker combinations (CD10, Ig κ, and CD49d) in distinguishing LDNs from HDNs in pregnancies (n = 27); the area under the curve (AUC) was calculated (95% CI: 0.985–0.992). F Proportion of LDNs in the CD10−CD49d+Ig κ+ gate and HDNs in the CD10+CD49d−Ig κ− gate. Error bars: means with SEM. Statistics: unpaired Student’s t test (B). ***P < 0.001.
Leveraging a recent InfinityFlow study in cord blood, we observed that four neutrophil subsets—termed proNeus, preNeus, immature Neus, and mature Neus—could also be resolved in our pregnancy samples, with highly similar marker expression dynamics across these developmental stages (Supplementary Fig. 2H, I). Furthermore, applying the gating strategy proposed by Kwok et al.39(using CD10 vs CD49d in combination with CD11b and CD16) to our flow cytometry data confirmed that pregnancy LDNs are composed almost entirely of immature subsets (CD11b−CD49d⁺ proNeus and CD10−CD11b⁺CD49d⁺ preNeus), with negligible presence of fully mature neutrophils (Supplementary Fig. 2J).
May-Grünwald Giemsa staining of sorted neutrophils confirmed that HDNs uniformly exhibited segmented nuclei. In contrast, the LDN population showed a heterogeneous mix of nuclear shapes (Supplementary Fig. 3A). Within the LDN population, the CD10−CD49d⁺ fraction predominantly exhibited immature nuclear forms (promyelocyte, myelocyte, and metamyelocyte), while CD10⁺ LDNs all had segmented nuclei and thus likely represent mature LDNs (Fig. 3A). The CD10−CD49d⁺ LDN subset expressed low levels of the neutrophil differentiation and activation markers CD11b and CD16 (Fig. 3B). A small subset of eosinophils or basophils also appeared in the CD10⁻ gate but could be excluded using CD49d and Ig κ (Supplementary Fig. 3B, C). We further validated the Ig κ, CD49d, and CD10 marker combination in whole blood, confirming that circulating CD10−CD49d+Ig κ+ neutrophils exhibited immature nuclear morphology, whereas CD10+ neutrophils displayed the typical segmented nuclear morphology (Fig. 3C and Supplementary Fig. 3D, E).
A Manual gating using CD10 and CD49d identifies immature LDNs (CD10−CD49d⁺), with representative micrographs of sorted populations (n = 3). Scale bars, 10 μm. Symbol key: metamyelocytes (○), myelocytes (■) and promyelocytes (★). B Representative histograms and statistical analysis for CD11b, CD62L, and CD16 expression on CD10−CD49d+ LDNs, CD10+ LDNs, and HDNs in pregnancies at 24–28 weeks of gestation (n = 6) and non-pregnant controls (n = 3). Gray dashed lines indicate unstained controls. C Representative flow cytometric analysis identifying immature (CD10−CD49d+Ig κ+) and mature neutrophils (CD10+CD49d−Ig κ−). Sorted populations of immature versus mature neutrophils are shown with corresponding cytospin images illustrating distinct nuclear morphology. Scale bars, 10 μm. D Representative flow cytometric dot plots of CD10−CD49d+Ig κ+ neutrophils in decidua tissues from term placentas. The bar chart shows the percentage of these cells within the CD15+ cells in late-pregnancy peripheral blood (36 weeks, n = 3) and decidua tissues (n = 4). E Representative micrographs (n = 3) of decidual neutrophils, highlighting the kidney-shaped nuclei of CD10−CD49d⁺Ig κ⁺ cells. Dec, decidua. Scale bar: 10 μm. Error bars: means with SEM. Statistics: one-way ANOVA corrected by Tukey’s test (B), unpaired Student’s t test (D). *P < 0.05.
We next asked whether these immature CD10−CD49d+Ig κ+ neutrophils are present at the maternal–fetal interface. To focus on maternal leukocytes and to avoid contamination with fetal cells from villous vasculature, we isolated leukocytes from the decidua parietalis (membrane-associated decidua) rather than the decidua basalis, and quantified CD10−CD49d+Ig κ+ neutrophils. Flow cytometric analysis revealed that these cells accounted for a small but consistent fraction (0.5–2.5%) of CD15+ cells in decidua parietalis of term placentas (Fig. 3D and Supplementary Fig. 3F). These decidual neutrophils predominantly displayed kidney-shaped nuclear morphologies similar to their counterparts in peripheral blood (Fig. 3E). By contrast, the CD10+ neutrophils in the decidua were morphologically mature (i.e., segmented nuclei). Together, these data indicate that the combination of CD10, Ig κ, and CD49d effectively discriminates immature and mature neutrophils in maternal blood and decidual tissues during pregnancy.
scRNA-seq analysis reveals transcriptional profiles of immature neutrophils
To further dissect the heterogeneity of neutrophils in pregnancy, we performed scRNA-seq on FACS-sorted CD10−CD49d+Ig κ+ (presumed immature LDNs) and CD10+ (presumed mature) neutrophils from pregnant blood samples (Fig. 4A). Unsupervised clustering of 43,146 single-cell transcriptomes revealed eight distinct neutrophil clusters (Fig. 4B, C, Supplementary Fig. 4A and Supplementary Data 3). Within the CD10+ (mature) fraction, three clusters of neutrophils were identified, namely Mat-NFKBIA, Mat-CXCR4, and Mat-CXCL1. These mature clusters showed elevated levels of canonical mature markers (B2M, BTG2, CSF3R, CXCL8, FCGR3B, JUNB, NAMPT, NFKBIA, and ZFP36; Fig. 4D). Notably, the Mat-CXCR4 subset appeared to be a senescent-like neutrophil population characterized by high expression of CXCR4, suggesting a role in trafficking back to the bone marrow for clearance40. The Mat-CXCL1 cluster was marked by elevated expression of CXCL1, characteristic of an “explorer” subset capable of initiating neutrophil swarming during infection via the CXCL1-CXCR2 homotypic chemotaxis axis41,42. The Mat-NFKBIA cluster, comprising ~80% of CD10+ neutrophils, encompassed overlapping features of the three mature clusters (Supplementary Fig. 4B).
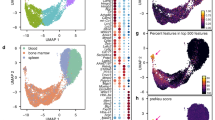
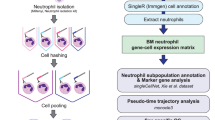
A Schematic of the scRNA-seq workflow. Immature (CD10−CD49d+Ig κ+) and mature (CD10+) neutrophils were sorted from the blood of a healthy pregnant woman and a pregnant woman with GDM at mid-gestation. Each sample was labeled individually with sample tag antibodies, then pooled for library construction and sequencing on the BD Rhapsody scRNA-seq platform. B UMAP plot of 43,146 neutrophils grouped into distinct clusters. Imm, immature, Mat, mature. C UMAP visualization (left) and corresponding cluster proportion (right) for neutrophil populations from healthy (HP) and GDM pregnancies. D Volcano plots displaying differentially expressed genes (left) and regulons (right) in immature versus mature neutrophils. E GSEA of mature neutrophils, highlighting inflammatory response (NES: 2.78, P = 1.63 × 10−18), IL6-JAK/STAT signaling pathway gene set (NES: 2.75, P = 1.18 × 10−14), IFN-α response gene set (NES: 3.01, P = 2.06 × 10−20), IFN-γ response gene set (NES: 3.24, P = 2.44 × 10−33), and TNF-α signaling via NF-κβ (NES: 3.36, P = 2.91 × 10−39). F GSEA of immature neutrophils, revealing enrichment of Myc targets gene sets (M5926 and M5928, NES: −2.56 and −1.97, P = 6.33 × 10−16 and P = 1.07 × 10−4), oxidative phosphorylation (NES: −2.32, P = 2.50 × 10−12), G2M checkpoint (NES: −2.42, P = 5.86 × 10−14), and E2F target (NES: −3.03, P = 3.73 × 10−29). The top portion of the plot shows the running enrichment score as the analysis walks down the ranked list. The bottom portion of the plot shows where the member genes of the gene set appear in the ranked list. NES, normalized enrichment score. G Pseudotime analysis illustrating the progression from the immature-MMP9 cluster to the immature-S100A4 cluster and corresponding changes in MME (CD10), TLR2 (CD282), FCGR2B (CD32), and OLR1 (LOX-1) expression. H The proportion of the “bridging” subset in LDNs (orange) is increased in GDM compared with healthy pregnancies, as assessed by InfinityFlow. I UMAP of total CD15+ cells in a GDM pregnancy (upper) and the corresponding changes in indicated markers along with UMAP-1 (lower). J Zoomed-in views of marker expression (CD10, CD282, CD32, LOX-1) modeled along the transition from LDN (left) to HDN (right). Curves of intensity were fitted using a polynomial model for nonlinear regression.
Pseudobulk differential expression (DE) analysis revealed that these CD10+ mature neutrophils expressed higher levels of pro-inflammatory genes (e.g., IL1A, IL1B, CCL3, and CCL4) and increased activity of immune-related transcription factors (e.g., NFKB2, IRF7, and MAFF) relative to CD10−CD49d+Ig κ+ (immature) neutrophils. Consistently, gene set enrichment analysis (GSEA) showed that the mature neutrophil clusters were enriched for inflammatory pathways (Fig. 4E). By contrast, the CD10−CD49d+Ig κ+ (immature) fraction segregated into five clusters: Imm-DEFA3, Imm-OLR1, Imm-OLFM4, Imm-MMP9, and Imm-S100A4. These immature clusters were enriched for genes associated with antimicrobial defense (e.g., CAMP, LTF, LCN2, MMP8, CEACAM8), proliferation/differentiation (e.g., CDKN2A, ANKEF1), and oxidative stress regulons (e.g., NFE2L1; Fig. 4B–D). The progressive transcription of specific granule (CAMP, LTF, and LCN2) or gelatinase (MMP9) granule genes in these immature neutrophils aligns with known granulopoiesis stages from promyelocytes to band cells43. GSEA of the immature neutrophil signature confirmed enrichment of oxidative phosphorylation, cell cycle, and MYC-related pathways in these cells (Fig. 4F and Supplementary Data 4). Consistently, ex vivo 5-ethynyl-2′-deoxyuridine (EdU) incorporation assays revealed that a fraction of the circulating CD10−CD49d+ LDNs could enter S-phase, demonstrating proliferative capacity (Supplementary Fig. 4C). In addition, we found that CD10−CD49d+ LDNs exhibited a markedly reduced respiratory burst in response to PMA stimulation compared to mature segmented neutrophils (Supplementary Fig. 4D). Moreover, scRNA-seq revealed significant enrichment of ARG1 in the CD10−CD49d+Ig κ+ subset (Supplementary Fig. 4A; immature versus mature neutrophils, Log2FC = −1.93, P = 2.95 × 10−7, Supplementary Data 4), supporting a model in which these circulating immature neutrophils have proliferative potential but limited effector function, consistent with a potentially immunosuppressive phenotype during pregnancy.
Among the immature clusters, Imm-S100A4 occupied a transitional state between the immature and mature neutrophil populations (Fig. 4B). Trajectory analysis indicated that cells progressed from Imm-MMP9 to Imm-S100A4 by upregulating MME (CD10), TLR2 (CD282), and FCGR2B (CD32), while downregulating OLR1 (LOX-1; Fig. 4G, Supplementary Fig. 4E and Supplementary Data 5). Interestingly, we observed that the proportion of such “bridging” population (i.e., the Imm-S100A4 in the scRNA-seq data and the analogous connecting LDNs in the InfinityFlow data) was notably increased in GDM pregnancies compared with healthy pregnancies (Fig. 4H and Supplementary Fig. 4F). InfinityFlow corroborated these transcriptional changes at the protein level, revealing an LDN subpopulation that gradually gained expression of CD10, TLR2, and CD32 while losing LOX-1 along the UMAP-1 axis (Fig. 4I, J). This shift toward a more mature-like phenotype may contribute to immunological alterations observed in GDM, suggesting that dysregulated neutrophil maturation dynamics play a role in pregnancy complications.
Indeed, in immature LDN-like neutrophils (CD10−CD49d⁺Ig κ⁺ subset), GDM samples exhibited increased expression of inflammatory and chemotactic genes (e.g., IL18RAP, CXCL8) and downregulated DNA-replication/cell-cycle programs (e.g., ZCCHC24 and other replication/repair modules). GSEA consistently showed enrichment of cytokine-mediated signaling, inflammatory response, and neutrophil chemotaxis/migration in GDM, whereas healthy pregnancy was enriched for DNA-replication/repair and cell-cycle processes (Supplementary Fig. 5A, B and Supplementary Data 6 and 7). Furthermore, we integrated our single-cell neutrophil dataset with the neutrophil atlas from Montaldo et al.43. A joint neutrophil landscape reveals that neutrophils in pregnancy occupy a transcriptional space distinct from both baseline and G-CSF–mobilized neutrophils (Supplementary Fig. 5C, D). Pregnancy LDNs (CD15⁺CD10−CD49d⁺Ig κ⁺ cells) mapped predominantly to the earliest neutrophil developmental clusters (Clusters 1–3) in the reference, which represent neutrophil-committed progenitors in healthy bone marrow (Clusters 1 and 2) and early immature neutrophils that appear in blood under extreme stimulation (Cluster 3). This suggests that pregnancy mobilizes early neutrophil precursors into circulation in a manner resembling an “extended bone-marrow output”, rather than inducing a full emergency granulopoiesis. In GDM, the neutrophil maturation balance shifted toward intermediate “bridge” clusters (2, 4, and 8) within this continuum (Supplementary Fig. 5E). This compositional skew aligns with the increased “bridging” population we observed in GDM (e.g., the Imm-S100A4 cluster by scRNA-seq and the connecting LDN subset by InfinityFlow).
Attenuated expansion of immature neutrophils in GDM pregnancies
An elevated total neutrophil count in the first trimester of pregnancy has been noted as a risk factor for GDM25. However, the behavior of circulating immature neutrophils in GDM remains poorly understood. While the mature CD10+ neutrophils showed only a modest elevation in GDM, the fraction of immature LDNs (CD10−CD49d⁺Ig κ⁺) declined significantly in GDM pregnancies compared to healthy pregnancies (Fig. 5A, B and Supplementary Fig. 6A–D). Accordingly, the immature (CD10−CD49d+Ig κ+) to mature (CD10+) neutrophil ratio (the I–M ratio) was markedly reduced in GDM (Fig. 5C).
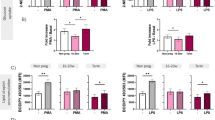
A Whole blood samples from age-matched non-pregnant controls, healthy pregnancies, and GDM pregnancies at 24–28 weeks of gestation were subject to red blood cell lysis and flow cytometry. B Representative dot plots (left) and quantification (right) of immature CD10−CD49d+Ig κ+ and mature CD10+ neutrophils in non-pregnant healthy controls (HC, n = 9), healthy pregnancies (HP, n = 14), and GDM pregnancies (GDM, n = 14). C The ratio of (CD10−CD49d⁺Ig κ⁺) to mature (CD10⁺) neutrophil counts in HP and GDM pregnancies. D Schematic illustration of the workflow for tracking oral glucose tolerance test (OGTT) results in pregnancies where immature neutrophils were previously detected, spanning 10–18 weeks of gestation. EP early pregnancy, MP mid-pregnancy. E Percentage of immature neutrophils (CD10−CD49d+Ig κ+) in CD45+ whole blood cells during early (EP, <14 weeks, n = 40) and mid-pregnancy (MP, ≥14 weeks, n = 35). F Analysis of the temporal trend in the proportion of immature neutrophils from ~11 to ~18 weeks in healthy (n = 60) and GDM (n = 15) pregnancies. G Linear regression of the proportion of immature neutrophils in CD45+ cells relative to gestational weeks of healthy (left, n = 74) and GDM (right, n = 29) cohorts. H Immature-to-mature (I–M) neutrophil ratio at <14, 14–18, and 24–28 weeks in healthy pregnancies (n = 39, 21, 14) and GDM pregnancies (n = 4, 11, 14). Pearson correlations between the I–M ratio and fasting plasma glucose (FPG; I), 1-h (OGTT1h; J), 2-h (OGTT2h; K) post-OGTT (n = 28). L Pearson correlation between the I–M ratio and HOMA-IR (n = 19, 7 not available) at 24–28 weeks of gestation. HOMA-IR homeostasis model assessment of insulin resistance, a dimensionless index. Error bars: means with SEM. Statistics: unpaired Student’s t test (C), Mann–Whitney test (E), one-way ANOVA corrected by Tukey’s test (the upper panel of B), Kruskal–Wallis test followed by Dunn’s test (the lower panel of B), two-way ANOVA followed by Tukey’s test (H). *P < 0.05, **P < 0.01, ***P < 0.001.
To investigate earlier gestational windows, an independent cohort of 75 pregnancies at 10–18 weeks of gestation was examined, with oral glucose tolerance test (OGTT) results obtained at 24–28 weeks (Fig. 5D). Notably, approximately 0.37 ± 0.06% of peripheral blood neutrophils were already CD10−CD49d⁺Ig κ⁺ by 14 weeks (Fig. 5E). In healthy pregnancies, this immature population expanded around 15–16 weeks, whereas women who later developed GDM showed virtually no such increase at that time (Fig. 5F). Combining data from both the first-(10–18 weeks) and second-(24–28 weeks) trimester cohorts confirmed a significantly slower rise in circulating CD10−CD49d+Ig κ+ immature neutrophils in GDM pregnancies compared to healthy pregnancies (Fig. 5G). Additionally, the I–M ratio progressively diverged over gestation between GDM and healthy groups (Fig. 5H and Supplementary Fig. 6E). These findings suggest that an attenuated I–M ratio is a characteristic immunological alteration in GDM pregnancies.
Correlation analyses demonstrated significant negative associations between the I–M ratio and measures of glycemic control, including fasting plasma glucose (FPG), 1-h (OGTT1h), 2-h (OGTT2h) post-OGTT, and homeostasis model assessment of insulin resistance (HOMA-IR; Fig. 5I–L and Supplementary Fig. 6F). Collectively, these results indicate that reduced CD10−CD49d⁺Ig κ⁺ neutrophils in circulation coincide with heightened metabolic disruptions, suggesting a potential immune-metabolic axis in GDM. Thus, an inadequate expansion of immature neutrophils throughout pregnancy may contribute to, or alternatively reflect, the aberrant inflammatory and metabolic states in GDM.
Decreased circulating myeloid progenitor cells associated with GDM
Neutrophil diversity in blood depends partly on the balance between the clearance of aged circulating neutrophils and the release of newly generated neutrophils from the bone marrow. As noted above, our scRNA-seq findings suggest that pregnancy mobilizes bone-marrow early neutrophil precursors into circulation (Supplementary Fig. 5C, D) and GDM pregnancies exhibit fewer circulating immature neutrophils (Fig. 5). These findings prompted us to examine whether pregnancy-associated myelopoiesis is altered in GDM.
We quantified hematopoietic progenitor cells (HPCs) in peripheral blood during the second trimester. Compared to age-matched non-pregnant controls, healthy pregnant women showed a clear elevation of lineage-negative (Lin−)CD45⁺CD34⁺CD38⁺ HPCs in blood (Fig. 6A and Supplementary Fig. 7A), consistent with the concept of enhanced myelopoiesis during normal pregnancy. By contrast, GDM pregnancies displayed a substantially blunted increase in HPCs. The frequencies of circulating common myeloid progenitors (CMPs; CD123+CD45RA−) and granulocyte–monocyte progenitors (GMPs; CD123+CD45RA+) were significantly lower in GDM versus healthy pregnancies (Fig. 6B and Supplementary Fig. 7B). These data support the notion that a robust myelopoietic expansion in healthy pregnancy drives the rise in circulating immature neutrophils. In GDM, this normal pregnancy-associated myelopoiesis appears to be impaired, resulting in a delayed or diminished release of immature neutrophils into the circulation.
A Representative flow plots (left) and quantification (right) of circulating HPCs in age-matched non-pregnant controls (HC, n = 4), healthy pregnancies (HP, n = 15), and GDM pregnancies (GDM, n = 10). B Flow cytometric identification of common myeloid progenitors (CMPs; CD123⁺CD45RA−), granulocyte–monocyte progenitors (GMPs; CD123⁺CD45RA⁺), and megakaryocyte–erythrocyte-biased progenitors (MEPs; CD123−CD45RA−) within CD45⁺ PBMCs. Error bars: means with SEM. Statistics: Kruskal–Wallis test followed by Dunn’s test (A and the right panel of B), Welch’s ANOVA followed by Dunnett’s T3 test (the middle and left panels of B).
Discussion
Neutrophils, traditionally viewed as a homogeneous population, are now recognized to display substantial heterogeneity, especially under stress and pathological conditions3,5,44. Pregnancy represents a unique immunological state that demands finely tuned innate immune adaptations to maintain maternal–fetal tolerance while ensuring immune surveillance22,45. In this study, we leveraged both high-dimensional flow cytometry and scRNA-seq to characterize neutrophil subsets in healthy and GDM pregnancies. Our findings highlight that maintaining an appropriate balance between immature LDNs and mature segmented neutrophils is a feature of normal pregnancy, whereas disruptions in this balance are linked to complications such as GDM.
LDNs, defined by their low buoyancy in density gradients, have been implicated in a range of diseases8,9,11,13,14,46. However, their precise phenotypes and functional roles have remained elusive. Here, we identified a specific combination of surface markers (CD10−CD49d⁺Ig κ⁺) that robustly delineates the immature LDN subset in whole blood during pregnancy. May-Grünwald Giemsa staining confirmed that the CD10−CD49d⁺Ig κ⁺ subset was enriched for promyelocytes, myelocytes, and metamyelocytes, whereas CD10⁺ neutrophils had exclusively segmented nuclei. These observations are consistent with recently defined neutrophil maturation schemes, in which CD49d (α4 integrin) marks neutrophil precursors (proNeu and preNeu), whereas CD10 expression is acquired at the transition to the late immature or band stage. In fact, the neutrophil subsets we identified in mid-gestation pregnancy closely mirror those reported in human cord blood32,35,39, with concordant marker patterns (high CD49d and low CD10 on immature cells). Notably, the CD10−CD49d+ neutrophils in pregnancy were almost uniformly CD16low/CD11blow, distinguishing them from the CD10−CD49d−CD16hi “band” neutrophils typically released during acute infection or G-CSF-driven emergency granulopoiesis39. In our cohorts, such band-like neutrophils were exceedingly rare (~1–2% of circulating neutrophils) in LDNs and resided primarily in the high-density fraction. Thus, the LDN population in healthy pregnancy consists predominantly of early neutrophil precursors rather than the more mature band forms observed in inflammatory settings.
Our scRNA-seq analysis further corroborated the distinction between immature and mature neutrophil populations in pregnancy, revealing divergent transcriptional programs that align with their functional states. Immature CD10−CD49d+ neutrophils exhibited elevated expression of oxidative phosphorylation and cell cycle genes, with lower expression of inflammatory cytokines and immediate early response genes (e.g., IL1B, CCL3, NFKBIA). These patterns suggest that the immature LDNs are a less terminally differentiated population that retains proliferative capacity and is transcriptionally poised for development rather than for effector functions. Functionally, we found that the CD10−CD49d+ LDNs had a markedly attenuated respiratory burst compared to mature neutrophils, indicating limited reactive oxygen species production. Moreover, ARG1 (arginase-1) was significantly upregulated in the immature LDN subset, reminiscent of the PMN-MDSCs that expand during healthy pregnancy to promote fetal tolerance18,21. We also observed high LOX-1 (OLR1), a marker previously identified on suppressive neutrophils in cancer38, on pregnancy LDNs, supporting their immunoregulatory phenotype. Notably, the immature LDNs bound substantial IgG on their surface (κ/λ light chains and Fc), likely due to elevated FcγRI (CD64) expression. We also detected a small but consistent population of these immature neutrophils in the decidua parietalis (maternal–fetal interface), suggesting that they may also participate in local immune regulation and tissue remodeling at the uterus–placenta interface19.
In GDM, the typical mid-gestation expansion of immature CD10−CD49d⁺ LDNs was attenuated, resulting in a lower immature versus mature (I–M) ratio. The remaining immature neutrophils in GDM shifted toward a more mature, activated transcriptomic program, such as upregulating IL1B, IL18RAP, and CXCL8 while downregulating DNA-replication/cell-cycle modules, accompanied by an increased S100A4⁺CD10low “bridge” population. Along this trajectory, loss of LOX-1 and gain of CD10 were suggestive of accelerated maturation and heightened inflammatory readiness. Consistent with this, the I–M ratio inversely correlated with glycemic indices in our cohort. These findings suggest that the neutrophil developmental balance is linked to maternal metabolic status in pregnancy24,25,38,39,43. Our data collectively portray pregnancy LDNs as immature, low-effector neutrophils with potential immunosuppressive roles. This is in stark contrast to the highly reactive, segmented neutrophils that dominate the circulation in acute infections and other pro-inflammatory states43,47,48,49.
Neutrophil diversity can emerge from systemic activation and “emergency granulopoiesis” that accelerates bone marrow neutrophil release50,51. Our data indicate that immature LDNs arise early in healthy pregnancy and expand over gestation. This increase is correlated with elevated levels of circulating myeloid progenitors (CD34+CD38+ CMPs and GMPs), suggesting enhanced myelopoiesis. GDM pregnancies, however, showed a blunted increase in these progenitor populations, consistent with the reduced output of immature neutrophils. Thus, an impaired myelopoietic upregulation in GDM may underlie skewed neutrophil subset balance. It is worth noting that different pregnancy complications may perturb neutrophil dynamics in distinct ways. For example, in preeclampsia and SLE-affected pregnancies, LDN frequencies can be elevated but tend to be skewed toward activated or mature phenotypes (e.g., CD62Llow “aging” neutrophils or pro-inflammatory low-density granulocytes)28,52. In cases of early pregnancy loss, LDN levels are normal, but the CD16high neutrophils display impaired activation27. By contrast, we found that GDM was characterized by a deficiency of immature neutrophils rather than an excess of activated neutrophils. Together, these observations underscore that although altered neutrophil profiles are a common feature of pregnancy complications21,52, the specific nature of the disturbance likely varies by condition. Understanding these nuances will be important for developing targeted interventions.
Beyond pregnancy, circulating immature neutrophils have been described in other stress and disease conditions, including acute infection, sepsis, G-CSF therapy, and cancer50,53,54. For instance, LDNs are predominantly immature (~80–90% CD10⁻) in healthy pregnancy (this study) and in G-CSF–mobilized donors32, whereas in solid tumors they are mostly mature (~10–15% CD10−)10; SLE shows an intermediate pattern (~50% CD10−) with many LDNs being activated. Moreover, unlike in severe infection, pregnancy LDNs contain very few band-form neutrophils53,54. These differences illustrate the plasticity of the neutrophil compartment and highlight the need for standardized nomenclature and phenotyping panels35. Notably, our identification of CD49d (integrin α4) as a key marker for immature neutrophils aligns with recent proposals for neutrophil nomenclature39. CD49d is known to mediate retention of neutrophil precursors in the bone marrow and to facilitate adhesion during intravascular crawling55,56, which may explain its high expression on newly released neutrophils. Meanwhile, the loss of CD10 signifies that these cells have not yet reached full maturation32. The inclusion of an immunoglobulin light-chain marker (Ig κ) in our panel, though unconventional, proved helpful in distinguishing neutrophils from other CD49d+ granulocytes (e.g., eosinophils) and in marking neutrophils with high surface IgG content. The presence of surface-bound Ig on LDNs aligns with their high CD64 expression and may indicate functional differences in how these cells handle immune complexes or interact with antibodies. Future studies should investigate whether Ig-coated immature neutrophils in pregnancy exhibit altered signaling or clearance functions.
In conclusion, our study provides a framework for defining and monitoring neutrophil heterogeneity during human pregnancy. We demonstrate that healthy gestation is characterized by a significant expansion of immature, CD10−CD49d+ neutrophils with immunoregulatory features, whereas GDM is marked by an impairment in this expansion alongside a shift toward pro-inflammatory neutrophil phenotypes. These insights extend our understanding of innate immune adaptation in pregnancy and suggest that the neutrophil developmental balance links maternal immune tolerance with metabolic homeostasis. The marker combination identified here (CD10, CD49d, and Ig-κ binding) provides practical tools for phenotypic tracking of neutrophil subsets and could potentially serve as early indicators of pregnancy complications. By elucidating neutrophil subset dynamics in maternal–fetal health, this work lays the groundwork for further mechanistic studies and points to potential immunomodulatory strategies to improve outcomes in pregnancies complicated by GDM and related conditions.
Methods
Study participants
All blood samples were collected from healthy non-pregnant women (n = 13), healthy pregnant women (n = 99), and pregnant women with GDM (n = 47) at the First Affiliated Hospital of Sun Yat-sen University (Guangzhou, China). According to the clinical guidelines, pregnancies were divided into three different stages: early pregnancy (EP, <14 weeks, n = 75), mid-pregnancy (MP, 14–28 weeks, n = 66), and late pregnancy (LP, >28 weeks, n = 5). All placental samples were collected from healthy pregnant women (n = 4) with normal (physiological) pregnancies who underwent elective cesarean section for clinical indications in the First Affiliated Hospital of Sun Yat-sen University (Guangzhou, China).
The inclusion and exclusion criteria were as follows: all subjects were screened for serum hepatitis B surface antigen (HBsAg), hepatitis C virus (HCV) antibody, hepatitis D virus (HDV) antigen and antibody, HIV antibody, syphilis, and individuals with positive results were excluded from this study. Women with acute infections, fever, severe critical illness, or chronic disease were also excluded from this study. This study was approved by the Clinical Ethics Review Committee of the First Affiliated Hospital of Sun Yat-sen University (No. 766-1[2021]). Written informed consent was obtained from all participants at the time of admission. All ethical regulations relevant to human research participants were followed. Detailed information is provided in Supplementary Table 1 and 2 and Supplementary Data 1.
Cell isolation
Whole blood samples underwent LDN isolation within 45 min from collection to laboratory processing, with transport maintained on ice throughout. All blood samples were collected in BD Vacutainer® K2E (EDTA) tubes (367525; BD) using EDTA as anticoagulant. Peripheral blood mononuclear cells (PBMCs) and polymorphonuclear leukocytes (PMNLs) were isolated using Lymphoprep density-gradient centrifugation. After centrifugation and removal of the plasma, 0.9%NaCl was added to a total volume of 5 mL. The Lymphoprep solution (1858; Serumwerk Bernburg AG) was added to a sterile centrifuge tube, and then the diluted blood was gently layered on top. Following density-gradient centrifugation, PBMCs were collected from the upper layer. Erythrocytes in the high-density layer were removed by dextran (D8270; Solarbio) sedimentation. High-density neutrophils (HDNs) were isolated from the remaining high-density fraction, and residual erythrocytes were lysed using sterile ultrapure water. Finally, the cells were resuspended in PBS and prepared for subsequent experiments.
For whole blood analyses, plasma was removed by centrifugation, and erythrocytes were removed by dextran sedimentation. Erythrocytes were allowed to sediment for 30 min, and the upper phase containing leukocytes was collected. Sterile ultrapure water was used to lyse erythrocytes and cells were resuspended in PBS. Cell viability and cell numbers were determined using a Z Series Coulter Counter (Beckman Coulter).
Placenta samples were obtained from healthy pregnant women with physiological pregnancy following elective cesarean section proposed on clinical grounds in the First Affiliated Hospital of Sun Yat-sen University. Because the decidua basalis lies in intimate apposition to anchoring chorionic villi that contain fetal vasculature, disruption of villous capillaries during dissection or enzymatic digestion can release fetal leukocytes into the preparation. In contrast, the decidua parietalis is distant from the villous tree and shows minimal to no deep invasion by extravillous trophoblast, providing maternal‑specific tissue and a lower risk of fetal carryover. Therefore, we sampled decidua parietalis scraped from the membranes and thoroughly rinsed the tissue before digestion. Decidua parietalis tissues were collected immediately after delivery. In brief, we used a scalpel to carefully scrape off decidua parietalis from chorioamniotic membranes and transferred into PBS57. Fresh decidua tissues were thoroughly washed in PBS and minced into small pieces. Decidual cells were isolated by digesting the tissues with 1 mg/mL collagenase type IV (C5138; Sigma-Aldrich) and 0.01 mg/mL DNase I (D5025; Sigma-Aldrich) in RPMI medium 1640 (31800089; Thermo Fisher Scientific) for 40 min at 37 °C. The suspensions were strained through nylon mesh and treated with RBC lysis buffer (NH4CL2009; TBD). Cell suspensions were filtered and prepared for subsequent experiments.
LEGENDScreen and InfinityFlow pipeline
Blood (10 ml) from a healthy pregnant woman and a pregnant woman with GDM who were matched based on maternal age (28 years old), BMI (28–28.5 kg/m2), method of conception, and gestational weeks (38 weeks) was collected in EDTA-containing tubes (367525; BD). Each donor was screened separately; no cross-donor pooling was performed. LEGENDScreen Human PE Kit (700011; BioLegend), recommendations for LEGENDScreen Plate preparation and staining were performed as indicated by the manufacturer, unless noted otherwise below. Briefly, low‑density PBMCs (approximately 15 million) and high-density PMNLs (approximately 50 million) were isolated by density centrifugation, hashtagged separately, and pooled for flow cytometry analysis, so that each antibody well received cells from both fractions. Initially, the cells were stained with a fixable live/dead dye (423105; BioLegend) for 30 min. Then, cells from different density fractions were stained with different fluorophore-conjugated anti-CD45 before staining with a backbone panel cocktail of antibodies to define the various lineages in the peripheral blood. Cells from different density fractions were washed and mixed for staining with a backbone panel of antibodies for 30 min at 4 °C, including: CD3 (OKT3), CD56 (5.1H11), CD19 (HIB19), LOX-1(15C4), CD34 (581), CD15 (MMA), and CD14 (M5E2). For the LEGENDScreen plates (354 test antibodies + 10 isotype controls; 364 wells), approximately 170,000 cells were aliquoted into individual wells containing specific PE-conjugated marker. After staining for 30 min, plates were washed and fixed before flow cytometry acquisition was performed on Cytoflex S flow cytometer (Beckman Coulter) and analyzed with FlowJo software (TreeStar), and data were subsequently processed through the InfinityFlow pipeline as described elsewhere37. The InfinityFlow pipeline involves regression analysis of the intensities of the PE-bound markers using the intensities of the backbone markers. In detail, we applied the infinity_flow function of this package using a prediction_events_downsampling parameter of 2000 and other default parameters. The output concatenated FCS file was further analyzed using FlowJo software to debarcode the pooled data into four sample groups: CD45+ cells in the low-density fraction of healthy pregnancy (HP_LD_CD45), CD45+ cells in the high-density fraction of healthy pregnancy (HP_HD_CD45), CD45+ cells in the low-density fraction of GDM pregnancy (GDM_LD_CD45), and CD45+ cells in the high-density fraction of GDM pregnancy (GDM_HD_CD45). The separated FCS files of four sample groups were then downsampled to ~20,000 cells per sample and subjected to further analysis in FlowJo. Data preprocessing and FlowSOM clustering were performed in FlowJo. Clusters were manually annotated and gated in FlowJo, and the imputed values for each cluster were exported.
To generate an antibody panel for neutrophils from the low-density and high-density fractions, we inspected the 354 screening antibodies that stained CD15+ cells and focused on the 172 antibodies with detectable signals (the fold change of LDNs or HDNs relative to their respective isotype controls was >1.00 in MFI) and positive proportions exceeding 1%, as determined by 10 isotype controls (Supplementary Data 2). This threshold was chosen because eosinophils and basophils exhibited positive rates of surface markers below 1%.
CD15+ cells in the high-density fraction of healthy and GDM pregnancies were then downsampled to ~3000 cells per sample, and CD15+ cells in the low-density fraction of healthy and GDM pregnancies were then downsampled to ~1000 cells. Sample groups were concatenated, and UMAP was subsequently performed on 172 selected markers expressed on CD15+ cells.
To define up- or downregulated markers in LDNs, we manually gated LDNs and HDNs using single, non-concatenated FACS files from the LEGENDScreen panel and calculated the MFI for each of the 172 neutrophil-expressed markers within each population. Markers were then ranked by the Log2 MFI fold change (LDN versus HDN). We applied a threshold of ∣Log2FC∣ ≥ 0.5 and then took the intersection of these lists across the two independent screens to define the final panel of top markers, as shown in Fig. 1F.
Flow cytometry
The prepared cell suspensions were stained with surface markers. Briefly, single-cell suspension of peripheral blood and decidua was washed and resuspended at 1 × 106 cells/tube. The appropriate antibodies diluted in PBS with 1% FBS and 2 mM EDTA were added to the cells and incubated for 30 min at 4 °C. The strategy for human neutrophil gating from whole blood was CD45+Lin−(CD3, CD19, CD56) CD15+. The strategy for human neutrophil gating from decidua tissues was CD45+Lin−(CD3, CD19, CD56, CD14) CD15+. The strategy for human neutrophil gating from low-density and high-density fractions was CD45+CD15+, in some cases with the additional use of CD193 to exclude contaminating eosinophils. For validating the InfinityFlow results, monoclonal antibodies, including CD10 (HI10a), CD49d (9F10), CD63 (H5C6), and Ig light-chain κ (MHK-49) were used. For the human HSPC analysis, monoclonal antibodies, including lineage cocktail (562722; BD Bioscience), CD45 (J33), CD34 (581), CD38 (HIT2), CD123 (7G3), and CD45RA (HI100) were used. HSPCs gating from PBMCs were identified as CD45+Linlow/−CD34+, GMPs as CD45+Linlow/−CD34+CD38+ CD123+CD45RA+, common myeloid progenitors (CMPs) as CD45+Linlow/−CD34+CD38+ CD123+CD45RA−, and MEPs as CD45+Linlow/−CD34+CD38+CD123−CD45RA−. The gating strategies are provided in Supplementary Figs. 2A, 3F, 6B and 7A in Supplementary Information. Detailed descriptions of the antibodies used for flow cytometry are provided in Supplementary Table 3. Data were acquired on a Cytoflex S flow cytometer (Beckman Coulter) and analyzed with FlowJo software (TreeStar).
Cell sorting
Cells from blood and decidua were stained with surface markers, isolated, and purified by a MoFlo XDP flow cytometer (Beckman Coulter). The strategy for immature LDNs gating from whole blood was CD45+CD15+CD10-CD49d+Ig κ+ and HDNs was CD45+CD15+CD10+Ig κ−CD49d−. The strategy for immature LDNs gating from the PBMC fraction was CD45+CD15+CD10−CD49d+. The strategy for immature neutrophils gating from decidua tissues was CD45+ CD15+CD10−CD49d+Ig κ+. All isolated cells were sorted to a purity of ≥ 95%.
Cytospin
Sorted neutrophil subsets (2 × 104 cells each) were spun onto glass slides using Cytopro 7620 Cytocentrifuge (Wescor, Logan, UT), dried for 10 min, fixed in 4% Paraformaldehyde (158127; Sigma-Aldrich) for 20 min and stained with Giemsa (51811-82-6; Sigma-Aldrich) for 20 min. Images were captured on Nikon Eclipse Ni-E with a 40× and 100× oil immersion objectives, and image brightness was adjusted with Photoshop (Adobe).
EdU incorporation
Sorted neutrophil subsets (CD10−CD49d+ LDNs) or total granulocytes were plated at 106 cells/ml with RPMI + 10% FBS in the absence or in the presence of 10 μM EdU (5-ethynyl-2′-deoxyuridine). After 18 h of culture, cells were collected, washed with PBS + 1% FBS, fixed, permeabilized and incubated with reaction cocktail according to the manufacturer’s instructions (C0071S, Beyotime). Samples were acquired at a Cytoflex S flow cytometer (Beckman Coulter) and analyzed with FlowJo software (TreeStar). A detailed reagent list is reported in Supplementary Table 3.
ROS assay
Freshly isolated PBMCs and PMNLs were washed and resuspended at 1 × 106 cells/ml and left untreated or stimulated with PMA 20 ng/ml and incubated at 37 °C for 30 min. For flow cytometry analysis of ROS content in neutrophils, 1 × 106 PBMCs and PMNLs were stained with fluorescently conjugated antibodies and incubated with 5 μM 2′,7′-Dichlorodihydrofluorescein diacetate (DCFH-DA, Sigma) in the dark for 20 min, acquired at a Cytoflex S flow cytometer (Beckman Coulter), and analyzed with FlowJo software (TreeStar). CD10−CD49d+ LDNs, CD10+ LDNs, and HDNs were identified after gating on Lin−CD15+ cells in the PBMC and PMNL fraction, respectively. A detailed reagent list is reported in Supplementary Table 3.
ROC curve analysis
The intensity scale values of the four most distinctive markers for each cell in LDNs and HDNs were exported using FlowJo. The AUCs for the combination of these markers, assessing their ability to discriminate between LDNs and HDNs, were calculated using SPSS Statistics version 26 (IBM).
Single-cell RNA sequencing
Immature neutrophils (CD45+Lin−CD15+CD10−CD49d+Ig κ+) and mature neutrophils (CD45+Lin−CD15+CD10+) were sorted from peripheral blood of a healthy pregnant woman (25 weeks) and a pregnant woman with GDM (24 weeks). To enable multiplexing, each of the four samples (healthy-immature, healthy-mature, GDM-immature, GDM-mature) was labeled with a unique Sample Tag antibodies in Human Single-Cell Multiplexing Kit (633781; BD Biosciences), following the manufacturer’s instructions. Equal numbers of tagged cells (~12,000 per subset) were then pooled, and the combined cell suspension was loaded into a single cartridge of the BD Rhapsody™ Single-Cell Analysis System. mRNA capture and cDNA synthesis were carried out on-chip following the BD Rhapsody protocol. We constructed both whole transcriptome amplification (WTA) libraries and Sample Tag libraries using the BD Rhapsody WTA workflow. All libraries were sequenced on an Illumina NovaSeq X Plus platform to obtain high-depth 2 × 150 bp reads for transcriptome profiling.
Preprocessing and quality control of scRNA-seq data
Raw Fastq files were processed using the BD Rhapsody Sequence Analysis Pipeline (v2.2.1), aligning to the human reference genome (RhapRef_Human_WTA_2023-02). Subsequent quality control (QC) and filtering were performed in R (v4.3.3, AlmaLinux 8.10) using Seurat (v5.1.0). Cells were retained if they had ≥300 detected genes, <15% mitochondrial RNA, and <1% hemoglobin gene expression. Multiplets or undetermined cells were excluded. For normalization, we applied SCTransform (v0.4.1) under glmGamPoi (v1.14.0) to regress out mitochondrial and ribosomal gene percentages.
Dimensionality reduction and unsupervised clustering analysis
Dimensionality reduction was then executed via Seurat’s RunPCA function, excluding artifactual features such as histone genes, hemoglobin genes, mitochondrial genes, ribosomal genes, and non-coding RNAs. Based on an ElbowPlot, the first 20 principal components were selected for downstream analyses, including uniform manifold approximation and projection (UMAP; RunUMAP) and clustering (FindNeighbors, FindClusters). We examined the resolution parameters with the help of the clustree (v0.5.1) package and altered the resolution to 0.08 in FindClusters.
Reference integration and cell state mapping
To further interpret the identified neutrophil clusters, we leveraged a published single-cell neutrophil atlas as a reference for cell state annotation43. We integrated our neutrophil scRNA-seq data with this atlas using Seurat’s reference mapping workflow. Briefly, the reference dataset was first loaded as a Seurat object with its original cluster labels. We then normalized both reference and query (our dataset) with SCTransform and identified 3000 shared highly variable genes. Using these features, we ran FindTransferAnchors (with reciprocal PCA, 20 dimensions) to find correspondences between the two datasets. We next applied MapQuery, projecting our neutrophil cells into the reference UMAP space and transferring the reference’s cluster labels to each query cell based on gene expression similarity. This procedure effectively mapped each of our cells to a reference-defined neutrophil state.
Single-cell differential gene expression analysis and GSEA
Differentially expressed genes (DEGs) were identified with Seurat’s FindMarkers or FindAllMarkers (Wilcoxon rank-sum or ROC tests) applied to SCT-normalized data. To ensure robust results, we excluded genes prone to technical artifacts (e.g., mitochondria-encoded transcripts, hemoglobin genes, ribosomal genes, immunoglobulin chains, pseudogenes, and unannotated loci) from the testing. Genes detected in >1% of cells and meeting significance (adjusted P < 0.05, Benjamini–Hochberg correction) were considered differentially expressed. GO enrichment analyses were conducted using clusterProfiler (v4.8.1) with org.Hs.eg.db (v3.17.0), while gene set enrichment analysis (GSEA) was performed via fgsea (v1.26.0), using gene lists ranked by fold change. Curated gene sets included the Hallmark collection and Gene Ontology Biological Process (GO:BP) terms from the msigdbr (v7.5.1) resource.
Gene regulatory network inference
Gene regulatory network inference was performed with pySCENIC (v0.12.1) in a Python (v3.9.17) environment. First, a loom file containing the single-cell transcriptomic data (SCT assay) was created using SCopeLoomR (v0.13.0) in R. Then, the transcription factor (TF)–target identification, cis-regulatory motif analysis, and regulon activity scoring were carried out. Specifically, TF–target relationships were inferred using pyscenic grn, motif enrichments were assessed via pyscenic ctx, and regulon activity scores (AUC) were computed using pyscenic aucell. Binarization thresholds for these regulons were subsequently determined with pyscenic. binarization. The resulting regulon activities (both continuous AUC and binarized) were incorporated back into the Seurat object as separate assays.
Trajectory inference
To elucidate potential developmental trajectories of neutrophils and to determine if GDM affects neutrophil maturation dynamics, we performed pseudotime trajectory analysis using slingshot (v2.8.0). Data were converted from a Seurat object to a SingleCellExperiment (v1.22.0) object, and pseudotime plus branching structures were estimated on UMAP embeddings with slingshot. Subsets of neutrophils underwent partial trajectory analysis to explore lineage progression. Differential progression between GDM and HP conditions was assessed using progressionTest and differentiationTest functions in condiments (v1.8.0).
Statistics and reproducibility
Results are expressed as mean ± SEM and mean ± 95% CI. Dataset distribution was evaluated using the Anderson-Darling or the Shapiro-Wilk normality test. For comparing means between two groups, parametric data were analyzed via two-tailed Student’s t-test, whereas the Mann–Whitney U test was utilized for nonparametric datasets. For multiple comparisons, one-way ANOVA followed by Tukey’s multiple comparison test and Welch’s ANOVA Followed by Dunnett’s T3 multiple comparison test (for parametric data) or Kruskal–Wallis test (for nonparametric data) followed by Dunn’s multiple comparison test was applied. For comparative analysis of the immature-to-mature neutrophil ratio, two-way ANOVA followed by Tukey’s post hoc test was used. For correlation analysis, linear regression and nonlinear regression were used to generate the best-fit line for graphical representation, and Pearson’s correlation test was performed to generate P values. P values < 0.05 were considered as statistically significant. All statistical tests were two-sided unless stated otherwise. Detailed descriptions of sample sizes, numbers of replicates, and statistical analyses for each experiment are provided in the corresponding figure legends. All source data underlying the graphs and charts are reported in Supplementary Data 8. Unprocessed microscope images are presented in Supplementary Figs. 8–10.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2021) in the National Genomics Data Center (Nucleic Acids Res 2022), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA-Human: HRA010023), which are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human. All data reported in this paper will be shared by the lead contact upon request.
Code availability
Codes for data analysis and generating specific figures have been deposited to GitHub and can be accessed at https://github.com/ChongEdwardWu/CommsBio_26_Neutrophil_in_Pregnancy58.
References
Ashton, N. Physiology of red and white blood cells. Anaesth. Intensive Care Med. 8, 203–208 (2007).
Herro, R. & Grimes, H. L. The diverse roles of neutrophils from protection to pathogenesis. Nat. Immunol. 25, 2209–2219 (2024).
Kruger, P. et al. Neutrophils: between host defence, immune modulation, and tissue injury. PLOS Pathog. 11, e1004651 (2015).
Ley, K. et al. Neutrophils: New insights and open questions. Sci. Immunol. 3, eaat4579 (2018).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Hassani, M. et al. On the origin of low-density neutrophils. J. Leukoc. Biol. 107, 809–818 (2020).
Hacbarth, E. & Kajdacsy-Balla, A. Low density neutrophils in patients with systemic lupus erythematosus, rheumatoid arthritis, and acute rheumatic fever. Arthritis Rheumatol. 29, 1334–1342 (1986).
Liu, Y. et al. Phenotypic and clinical characterization of low density neutrophils in patients with advanced lung adenocarcinoma. Oncotarget 8, 90969–90978 (2017).
Dumitru, C. A., Moses, K., Trellakis, S., Lang, S. & Brandau, S. Neutrophils and granulocytic myeloid-derived suppressor cells: immunophenotyping, cell biology and clinical relevance in human oncology. Cancer Immunol. Immunother. 61, 1155–1167 (2012).
Vanhaver, C. et al. Immunosuppressive low-density neutrophils in the blood of cancer patients display a mature phenotype. Life Sci. Alliance 7, e202302332 (2024).
Morrissey, S. M. et al. A specific low-density neutrophil population correlates with hypercoagulation and disease severity in hospitalized COVID-19 patients. JCI Insight 6, e148435 (2021).
Rankin, A. N., Hendrix, S. V., Naik, S. K. & Stallings, C. L. Exploring the role of low-density neutrophils during mycobacterium tuberculosis infection. Front. Cell. Infect. Microbiol. 12, 901590 (2022).
Bowers, N. L. et al. Immune suppression by neutrophils in HIV-1 infection: Role of PD-L1/PD-1 pathway. PLOS Pathog. 10, e1003993 (2014).
van den Akker, E. L. T. et al. Ficoll-separated mononuclear cells from sepsis patients are contaminated with granulocytes. Intensive Care Med. 34, 912–916 (2008).
Tay, S. H., Celhar, T. & Fairhurst, A.-M. Low-density neutrophils in systemic lupus erythematosus. Arthritis Rheumatol. 72, 1587–1595 (2020).
Murad, M. et al. Low density neutrophils are increased in patients with behçet’s disease but do not explain differences in neutrophil function. J. Inflamm. 19, 5 (2022).
Sanchez-Pino, M. D. et al. Increased inflammatory low-density neutrophils in severe obesity and effect of bariatric surgery: results from case-control and prospective cohort studies. eBioMedicine 77, 103910 (2022).
Wang, Y. et al. Inhibition of pregnancy-associated granulocytic myeloid-derived suppressor cell expansion and arginase-1 production in preeclampsia. J. Reprod. Immunol. 127, 48–54 (2018).
Amsalem, H. et al. Identification of a novel neutrophil population: proangiogenic granulocytes in second-trimester human decidua. J. Immunol. 193, 3070–3079 (2014).
Ssemaganda, A. et al. Characterization of neutrophil subsets in healthy human pregnancies. PLoS ONE 9, e85696 (2014).
Shi, M. et al. Continuous activation of polymorphonuclear myeloid-derived suppressor cells during pregnancy is critical for fetal development. Cell. Mol. Immunol. 18, 1692–1707 (2021).
Gimeno-Molina, B., Muller, I., Kropf, P. & Sykes, L. The role of neutrophils in pregnancy, term and preterm labour. Life 12, 1512 (2022).
Shardina, K. Y. et al. The role of myeloid-derived suppressor cells in establishing immune tolerance during pregnancy. Cell Tissue Biol. 16, 330–338 (2022).
Pace, N. P. & Vassallo, J. Association between neutrophil-lymphocyte ratio and gestational diabetes—a systematic review and meta-analysis. J. Endocr. Soc. 5, bvab051 (2021).
Sun, T. et al. Elevated first-trimester neutrophil count is closely associated with the development of maternal gestational diabetes mellitus and adverse pregnancy outcomes. Diabetes 69, 1401–1410 (2020).
Köstlin, N. et al. Granulocytic myeloid derived suppressor cells expand in human pregnancy and modulate T-cell responses. Eur. J. Immunol. 44, 2582–2591 (2014).
Ye, H. et al. Dysregulated low-density granulocyte contributes to early spontaneous abortion. Front. Immunol. 14, 1119756 (2023).
Stockfelt, M. et al. Activated low-density granulocytes in peripheral and intervillous blood and neutrophil inflammation in placentas from SLE pregnancies. Lupus Sci. Med. 8, e000463 (2021).
Elghetany, M. T. Surface antigen changes during normal neutrophilic development: a critical review. Blood Cells. Mol. Dis. 28, 260–274 (2002).
Grieshaber-Bouyer, R. & Nigrovic, P. A. Neutrophil heterogeneity as therapeutic opportunity in immune-mediated disease. Front. Immunol. 10, 346 (2019).
Albrechtsen, M. & Kerr, M. A. Characterization of human neutrophil glycoproteins expressing the CD15 differentiation antigen (3-fucosyl-N-acetyllactosamine). Br. J. Haematol. 72, 312–320 (1989).
Marini, O. et al. Mature CD10+ and immature CD10− neutrophils present in G-CSF–treated donors display opposite effects on T cells. Blood 129, 1343–1356 (2017).
Weirich, E. et al. Neutrophil CD11b expression as a diagnostic marker for early-onset neonatal infection. J. Pediatr. 132, 445–451 (1998).
Loken, M., Shah, V., Dattilio, K. & Civin, C. Flow cytometric analysis of human bone marrow: I. Normal erythroid development. Blood 69, 255–263 (1987).
McKenna, E. et al. Neutrophils: need for standardized nomenclature. Front. Immunol. 12, 602963 (2021).
Dutertre, C.-A. et al. Single-cell analysis of human mononuclear phagocytes reveals subset-defining markers and identifies circulating inflammatory dendritic cells. Immunity 51, 573–589.e8 (2019).
Becht, E. et al. High-throughput single-cell quantification of hundreds of proteins using conventional flow cytometry and machine learning. Sci. Adv. 7, eabg0505 (2021).
Condamine, T. et al. Lectin-type oxidized LDL receptor-1 distinguishes population of human polymorphonuclear myeloid-derived suppressor cells in cancer patients. Sci. Immunol. 1, aaf8943 (2016).
Kwok, I. et al. Combinatorial single-cell analyses of granulocyte-monocyte progenitor heterogeneity reveals an early uni-potent neutrophil progenitor. Immunity 53, 303–318.e5 (2020).
De Filippo, K. & Rankin, S. M. CXCR4, the master regulator of neutrophil trafficking in homeostasis and disease. Eur. J. Clin. Investig. 48, e12949 (2018).
Zec, K. et al. Macrophages in the synovial lining niche initiate neutrophil recruitment and articular inflammation. J. Exp. Med. 220, e20220595 (2023).
Lopes, J. P. & Lionakis, M. S. Pathogenesis and virulence of Candida albicans. Virulence 13, 89–121 (2022).
Montaldo, E. et al. Cellular and transcriptional dynamics of human neutrophils at steady state and upon stress. Nat. Immunol. 23, 1470–1483 (2022).
Hedrick, C. C. & Malanchi, I. Neutrophils in cancer: heterogeneous and multifaceted. Nat. Rev. Immunol. 22, 173–187 (2022).
Bert, S., Ward, E. J. & Nadkarni, S. Neutrophils in pregnancy: new insights into innate and adaptive immune regulation. Immunology 164, 665–676 (2021).
Brandau, S. et al. Myeloid-derived suppressor cells in the peripheral blood of cancer patients contain a subset of immature neutrophils with impaired migratory properties. J. Leukoc. Biol. 89, 311–317 (2010).
Hesselink, L. et al. Neutrophil heterogeneity and its role in infectious complications after severe trauma. World J. Emerg. Surg. 14, 24 (2019).
Lokwani, R., Wark, P. A. B., Baines, K. J., Barker, D. & Simpson, J. L. Hypersegmented airway neutrophils and its association with reduced lung function in adults with obstructive airway disease: an exploratory study. BMJ Open 9, e024330 (2019).
Palomino-Segura, M., Sicilia, J., Ballesteros, I. & Hidalgo, A. Strategies of neutrophil diversification. Nat. Immunol. 24, 575–584 (2023).
Lawrence, S. M., Corriden, R. & Nizet, V. The ontogeny of a neutrophil: mechanisms of granulopoiesis and homeostasis. Microbiol. Mol. Biol. Rev. 82, e00057–17 (2018).
Manz, M. G. & Boettcher, S. Emergency granulopoiesis. Nat. Rev. Immunol. 14, 302–314 (2014).
Liu, D. et al. Placenta-derived IL-32β activates neutrophils to promote preeclampsia development. Cell. Mol. Immunol. 18, 979–991 (2021).
Kipnis, E. Neutrophils in sepsis: battle of the bands*. Crit. Care Med. 41, 925 (2013).
Leliefeld, P. H. C. et al. Differential antibacterial control by neutrophil subsets. Blood Adv. 2, 1344–1355 (2018).
Devi, S. et al. Neutrophil mobilization via plerixafor-mediated CXCR4 inhibition arises from lung demargination and blockade of neutrophil homing to the bone marrow. J. Exp. Med. 210, 2321–2336 (2013).
Massena, S. et al. Identification and characterization of VEGF-A-responsive neutrophils expressing CD49d, VEGFR1, and CXCR4 in mice and humans. Blood 126, 2016–2026 (2015).
Xu, Y. et al. Isolation of leukocytes from the human maternal-fetal interface. J. Vis. Exp. 99, e52863 (2015).
Wu, C. ChongEdwardWu/CommsBio_26_Neutrophil_in_Pregnancy: CommsBio neutrophil study—analysis and figure-generation code (v1.0.0). Zenodo, https://doi.org/10.5281/zenodo.17659333 (2025).
Acknowledgements
The authors thank Lingyan Zhu and Yunfeng Chen for helping with FACS sorting and Dr. Renato Ostuni for providing the processed neutrophil scRNA-seq data. This work was supported by project grants from the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2024ZD0532100), the National Natural Science Foundation of China (82401983, 82488101, 32230034, 32322031, and 32571071), the Guangdong Basic and Applied Basic Research Foundation (2023A1515110982), the Guangzhou Basic and Applied Basic Research Foundation (2023A04J0498), the Key-Area Research and Development Program of Guangdong Province (2023B1111020005), and the Science and Technology Planning Project of Guangdong Province (2023B1212060028). Figs. 1A, 2A, 4A and 5A, D were created with BioRender (biorender.com).
Author information
Authors and Affiliations
Contributions
J. Xu, C. Zhu, C. Wu, and H. Chen designed the experiments. J. Xu, Y. Liu, J. Li, and L. Wang performed the experiments. J. Xu, Y. Liu, J. Li, and C. Wu analyzed and interpreted the data. C. Zhu, L. Xie, Y. Liu, L. Ye, Y. Zhang, Z. Wang, and H. Chen provided clinical resources and helpful advice. Z. Wang, L. Zheng, C. Wu, and H. Chen supported and supervised the research project. J. Xu, C. Wu, and H. Chen wrote the manuscript, and all authors contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Dr. Nilanjan Banerjee and Dr. Laura Rodriguez Perez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, J., Zhu, C., Xie, L. et al. Multimodal single-cell protein and RNA profiling unveils dysregulated immature neutrophil dynamics in gestational diabetes mellitus. Commun Biol 9, 316 (2026). https://doi.org/10.1038/s42003-025-09468-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09468-9