Abstract
Numerous computational approaches have been developed to infer cell state transition trajectories from snapshot single-cell data. Most approaches first require projecting high-dimensional data onto a low-dimensional representation; however, this can distort the dynamics of the system. Using epithelial-to-mesenchymal transition (EMT) as a test system, we show that both biology-guided low-dimensional representations and trajectory simulations in high-dimensional state space, not representations obtained with brute force dimensionality-reduction methods, reveal two broad paths of TGF-β induced EMT. The paths arise from the coupling between cell cycle and EMT at either the G1 or G2/M phase, contributing to cell-cycle related EMT heterogeneity. Subsequent multi-plex immunostaining studies confirmed the multiple predicted paths at the protein level. The present study highlights the heterogeneity of EMT paths, emphasizes that caution should be taken when inferring transition dynamics from snapshot single-cell data in two- or three-dimensional representations, and shows that incorporating dynamical information can improve prediction accuracy.
Similar content being viewed by others
Introduction
Throughout the lifetime of every cell, there are critical moments when they encounter intrinsic and extrinsic signals that direct them towards a specific cellular fate. These decisions occur in various contexts, such as embryonic development, wound healing, cellular senescence, and aging. Within a cell, multiple cellular processes function simultaneously; therefore, the decision and execution of a cell fate change often requires coordination between different cellular programs to allocate limited resources and to minimize interference or conflict.
A central, yet long-standing question in cell biology revolves around how the cell cycle couples to and influences other cell fate decisions, such as differentiation, apoptosis, and reprogramming1. The cell cycle is a tightly regulated cellular process, where a proliferative cell cycles through a temporally ordered G1→S→G2→M sequence, with multiple checkpoints deciding on the continuation or halting of the cell cycle to ensure proper cell division.
The cell cycle couples with cell fate in a context-dependent manner. In the case of terminal differentiation, evidence has shown that cells typically withdraw from the cell cycle and enter an arrested state to proceed toward a specific cell fate2. In other cases, cell division is required to allow for cell fate switching, where the period between the end of mitosis and before S-phase entry is a prime window for transcriptional and epigenomic changes to lead to a change in cell fate. The Rb-E2F network, for example, integrates signals from regulators of other cellular processes to decide whether a newly divided cell proceeds through to the G1 phase or exits the cell cycle and enters a quiescent G0 phase1,3. In other cases, the specific cell cycle phase a cell resides in can dictate a cell’s fate2,4,5,6. For example, Gomer and Firtel found that single amoebae differentiated according to its position in the cell cycle, becoming prespore if in the G1 phase and a prestalk cell if in the G2 phase6. The coupling between the cell cycle and cell fate changes has been exploited to modulate cell fate transitions, such as accelerating the reprogramming process and improving the efficacy of cancer treatment2,7. However, the exact mechanisms that govern cell fate decisions, specifically the timing along the cell cycle, remain elusive.
Epithelial-to-mesenchymal transition (EMT) is a cell phenotype transition that has attracted extensive attention in recent years, due to its involvement in many biological processes, including development, wound healing, tissue fibrosis, and cancer metastasis8. During EMT, epithelial cells, characterized by an apical-basal polarity and adherence to a basement membrane and other epithelial cells, lose their epithelial traits and develop mesenchymal traits, characterized by a detachment from the basement membrane, an elongated shape and increased mobility8,9. The onset of EMT is regulated by a core regulatory network involving transcription factors and microRNAs. These regulators are also involved in other cellular processes like proliferation and regulation of the cell cycle10,11,12.
An open question surrounding EMT is whether it proceeds through a single path or multiple paths8. Pseudotime analyses of single-cell RNA-sequencing (scRNA-seq) data of human mammary epithelial (MCF10A) cells treated with transforming growth factor-beta (TGF-β), to induce EMT, concluded that EMT proceeds through a one-dimensional continuum13. Markov transition model analyses on snapshot proteomic data of non-small-cell lung carcinoma cell lines treated with TGF-β also reached a similar conclusion14. On the other hand, Wang et al. performed live-cell imaging studies of TGF-β-induced EMT in a A549, human lung carcinoma epithelial cell line, with endogenous vimentin-RFP labeling in a multi-dimensional composite cell feature space15,16 and found at least two parallel paths that connect the initial epithelial state to the final mesenchymal state.
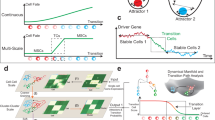
The cell cycle is known to couple with EMT and could therefore be a potential driver for the multiple paths. In the context of human oral squamous cell carcinoma cells, TGF-β treatment induced G1 cell cycle arrest17. In the case of kidney fibrosis and pancreatic ductal adenocarcinoma, a link between cell cycle arrest and partial EMT has been established, where Snai1-induced partial EMT also induces G2/M cell cycle arrest18,19. In other instances, such as during epidermal stem cell differentiation, there is a complete decoupling20. The discrepancy in the reports raises the question of whether the coupling between the cell cycle and EMT is context-dependent. Another factor to consider is the strength of the coupling. In the case of a tight coupling, one would expect cells to arrest from very specific and narrow regions of the cell cycle, such as a checkpoint (Fig. 1a). In contrast, a loose coupling would result in a broad range of cells undergoing EMT in different stages of the cell cycle (Fig. 1b).
Plausible scheme of tight (a) and weak (b) coupling between cell cycle and EMT. Cell cycle checkpoints are indicated by the red lines. c Schematic of the experimental design of scRNA-seq data acquisition. MCF10A cells treated with increasing concentrations of TGF-β over the span of 14 days, pooled together, and then scRNA-seq was performed. d Dynamo transcriptomic vector field of untreated MCF10A cells with stable fixed points. Circled regions are regions used for comparative analyses. e Comparison of the mRNA count between region 1 and region 2. f Comparison of the distribution of cell cycle phases between region 1 and region 2 using cell cycle phases assigned by dynamo. g A nonparametric Mann–Whitney U test was used to compare region 1 and region 2. The top 10 genes per region were selected, and their log2 fold change were plotted.
Addressing these questions is a formidable challenge as it requires unraveling the regulatory mechanisms and transition dynamics of cell phenotypic transition processes, which involve many molecular species that are coupled together. The development of scRNA-seq technology has revolutionized biology research by providing genome-wide snapshots of gene expression profiles in heterogeneous cell populations21,22. Numerous computational approaches have been developed to infer phenotype transition trajectories from snapshot single-cell data. Pseudotime analysis is one class of computational approaches, where sampled single-cell data are ordered based on their progression through a biological process determined by gene expression similarities23. Methods, such as optimal transport, use distributions in the cell state space measured at several time points to infer how cells transition between different cell states24.
Another class of computational methods integrates single-cell expression states with dynamical information for trajectory inference. In a seminal study, La Manno et al. took advantage of the ability to distinguish unspliced and spliced mRNA transcripts from scRNA-seq data, and estimated instantaneous RNA velocity, the speed and direction of change of a high-dimensional vector in the cell expression space25. By taking these instantaneous RNA velocities, estimated using the original splicing-based method or by other improved methods26 as input, Qiu et al. developed dynamo, a machine learning framework that reconstructs a set of multi-dimensional, analytical, and generally nonlinear continuous vector field functions encoding the genome-wide gene regulation relations of the system27. Zhang et al. further developed graph-dynamo for complete dynamical model studies28.
In general, many of the existing dynamics-inference methods are performed on an extremely low-dimensional (e.g., two- or three-dimensional) manifold of the single-cell data23. That is, one implicitly assumes that the very low-dimensional representation faithfully distinguishes different cell states in a biologically meaningful manner. However, this assumption needs to be carefully evaluated, as inappropriate representations may lead to a distortion of the underlying cellular dynamics29,30 and possibly imprecise conclusions, which we will discuss later.
While preliminary analysis in low-dimensional space can aid in visually understanding scRNA-seq data, more emphasis needs to be placed on performing higher-dimensional analysis to corroborate conclusions made from low-dimensional analysis. Similar to molecular dynamics simulations, which use a force field to describe molecular interactions, with dynamo, one can perform deterministic or stochastic simulations with the learned vector fields to generate single-cell trajectories in a high-dimensional (e.g., >20) cell state space to better understand the dynamics within the data.
To better understand EMT dynamics, here we performed vector field analyses using dynamo on scRNA-seq data of MCF10A cells treated with increasing concentrations of TGF-β and analyzed the dynamic cellular paths. To explore the coupling between the cell cycle and EMT, we extended upon the cell cycle framework from the R package Revelio31 to construct a computational pipeline to obtain a cell cycle-informed representation. With this representation, we predicted a loose coupling between the two processes exhibited by broad transition regions, originating from either the G1 or G2/M cell cycle phase. Next, we performed trajectory simulations and transition path analyses to confirm these paths in high-dimensional space. Finally, to validate these findings, we performed iterative indirect immunofluorescence imaging (4i) measurements and confirmed the two predicted EMT paths. Overall, this study advances our knowledge on the paths cells can take during EMT by identifying two broad EMT paths characterized by the coupling between the cell cycle and EMT. In addition, these findings highlight the importance of not only obtaining biologically informed low-dimensional space representations but also performing analyses in high-dimensional space to accurately capture dynamics within scRNA-seq data.
Results
scRNA-seq data of TGF-β induced EMT in MCF10A cells revealed an EMT continuum
We first analyzed the scRNA-seq dataset of human mammary epithelial (MCF10A) cells generated by Panchy et al32. The MCF10A cells were treated with increasing concentrations of TGF-β, an inducer of EMT (0, 12.5, 25, 50, 100, 200, 400, and 800 pM, respectively) for 14 days and then sequenced, encompassing a total of 8983 cells. To confirm EMT progression, we calculated an EMT score for each cell across the entire range of TGF-β concentrations using a Kolmogorov–Smirnov (KS) EMT scoring method, where a negative score indicated a cell is epithelial, while a positive score indicated a cell is mesenchymal33. As the TGF-β concentration increased, the overall KS EMT score increased, indicating that cells overall gained mesenchymal characteristics and lost epithelial characteristics (Fig. S1a), consistent with what was observed by Panchy et al32.
To visualize EMT progression in the gene expression space, we examined the single-cell data with a two-dimensional uniform manifold approximation and projection (UMAP) representation34 (Fig. S1b). With increasing TGF-β concentration, the cells shifted from left to right, suggesting that the first UMAP mode (UMAP1) mainly reflects the TGF-β response, while the second mode (UMAP2) mostly captures TGF-β independent variations. Curiously, the four highest doses (100, 200, 400, 800 pM of TGF-β) settled in the same region on the 2D UMAP (Fig. S1b).
Observing the cells in 3D revealed that the cells with each increasing TGF-β concentration simultaneously moved upward along the UMAP3 axis as well as to the right along UMAP1 (Fig. S1c, d). Amongst the cells treated with the highest TGF-β concentrations, both the KS EMT score and expression of FN1, a mesenchymal gene marker, varied within each TGF-β concentration, reflecting the coexistence of a heterogeneous continuum of mesenchymal states instead of a singular one (Fig. S1e, f), consistent with what has been reported in other studies32,35. Overall, the 3D UMAP provides a clearer representation of EMT progression and reveals how the overall perception can be obscured by the distortion that can occur in 2D representations. This observation resonates with recent debates on the common practice of analyzing single-cell data in a reduced two-dimensional space29,36.
Among the eight TGF-β concentrations, only the cells treated with 12.5 and 25 pM of TGF-β retained a subpopulation of epithelial cells, indicated by the overlap with the untreated cells, which only contain epithelial cells (Fig. S1g–i). The remaining TGF-β concentrations contained little to no epithelial cells as most cells have already undergone EMT and have already moved away from the epithelial region, occupied by the untreated cells. In subsequent analyses on understanding how cells transition from the epithelial to mesenchymal region, we focused on the two lowest TGF-β concentrations (12.5 and 25 pM) in addition to the untreated cells (0 pM) (Fig. 1c). Consistent with the analysis of all TGF-β concentrations, the cells treated with low concentrations of TGF-β were also undergoing EMT (Fig. S2a–f). However, there was less heterogeneity in the end mesenchymal state as most cells end up in a high KS EMT score and high FN1 expression state (Fig. S2b, c). These results suggests that at lower concentrations of TGF-β, cells reach a single mesenchymal state37.
Two-dimensional transcriptomic vector field of untreated MCF10A cells revealed both quiescent and cycling cells but was biologically distortive
To analyze the EMT process, we began with the untreated MCF10A cells. We first applied dynamo to obtain the corresponding RNA vector field (Fig. 1d). The reconstructed vector field identified two stable fixed points, towards which vectors converged, representing stable cell states. To characterize the two stable cell states, we selected cells surrounding each fixed point and performed comparative analyses of mRNA abundance, cell cycle phase distribution and gene expression. Interestingly, cells in region 1 had significantly higher raw mRNA counts than cells in region 2 (with an independent sample two-sided T-test p value 9e−19) (Fig. 1e), indicative of higher total RNA content, although the correlation can be confounded by technical variations. Furthermore, cell cycle phase estimation analysis revealed that region 2 had 17.1 percent more cells in the G1/S phase and 9.7 percent more cells in the M/G1 phase than in region 1 (Fig. 1f). In addition, differential gene expression analysis revealed that region 1 had a positive log fold change for cycling related genes including MKI67, TOP2A, and CENPF (Fig. 1g). Collectively, these results suggest that region 1 corresponds to cells actively proliferating while region 2 may correspond to cells enriched in the quiescent (i.e., G0) state.
Although the 2D UMAP reduced representation of the untreated cells was able to identify two subpopulations of cells, quiescent and proliferating, the representation distorts the cell cycle dynamics. Specifically, the region 1 stable fixed point, identified on the 2D space, is both biologically and mathematically unreasonable, as it represents an actively proliferating cell state. Instead, one would expect to observe circular vector flows to represent the proliferating states. To be more specific, with proliferative cells, one would expect that the single-cell data form a closed geometric manifold with a hollow center along the axis of the closed structure, representing cell cycle progression in the cell state space as shown by Schwabe et al.31.
Two-dimensional transcriptomic vector field revealed two possible EMT transition regions in TGF-β induced EMT of MCF10A cells
A fundamental question in the field of EMT is whether cells undergoing EMT follow a single path or multiple paths8. To investigate this question, we applied the pseudotime method, Monocle 338, to the MCF10A cells treated with 12.5 or 25 pM of TGF-β. This analysis identified a single EMT path (Fig. 2a, b), consistent with previous reports using pseudotime analyses13 and Markovian transition model analyses14 on snapshot single-cell data, but differs from reports from live-cell imaging studies15,16.
Pseudotime analysis of 12.5 (a) and 25 (b) pM TGF-β-treated cells on UMAP space. Vector fields of 12.5 (c) and 25 (d) pM TGF-β treated cells on UMAP space. The red arrows indicate the two EMT paths. TOP2A (e, f) and CCNB1 (g, h) gene expression for both 12.5 and 25 pM TGF-β treated cells on UMAP space, respectively.
To investigate this discrepancy further, we performed vector field analysis using dynamo27. Notably, each vector field demonstrated a consistent pattern of EMT progression, with vectors shifting from left to right through two visually distinct transition paths (Fig. 2c, d), consistent with live-cell imaging results15,16. We noticed that several G2 cell cycle phase related genes, TOP2A and CCNB1, exhibited higher expression in the lower group of cells across the two concentrations of TGF-β-treated cells (Fig. 2e–h). This observation led us to hypothesize that the upper and lower EMT paths observed from the vector fields were due to the cell cycle coupling with EMT at the G1/S and G2/M checkpoints (Fig. 1a, b).
However, the relationship between the cell cycle and EMT is unclear in the current two-dimensional UMAP representation (Box 1). Thus, one needs a representation that can simultaneously reflect cell cycle progression as well as the EMT status of individual cells.
Computational pipeline identified the cell cycle coordinate
To determine whether the cell cycle and EMT couple together, we required a representation that takes cell-cycle information into account, which is not explicitly accounted for in widely used dimensionality reduction methods. Therefore, we developed a four-step pipeline that builds upon the Revelio cell cycle framework from Schwabe et al.31 to generate a biologically meaningful axis to capture cell cycle progression (Fig. 3a).
a The computational pipeline to obtain the cell cycle coordinate. CCNE1 (b) and TOP2A (c) gene expression to confirm cell cycle progression along the x-axis. DEK (d) and PARP1 (e) gene expression confirming quiescence entry into the cell cycle along the y-axis. f The cell cycle-quiescent representation obtained from the untreated MCF10A cells reveals quiescent cell entry into the cell cycle. The cell cycle-EMT representation obtained from the 12.5 (g) and 25 pM (h) TGF-β-treated MCF10A cells. Colors of the data points in (f–h) represent discrete cell cycle stages were assigned using Revelio, which were shown to compare with the cell cycle coordinate.
The pipeline utilizes raw spliced scRNA-seq mRNA counts as input to Revelio31, an R package (Fig. 3a subpanel 1). Revelio assumes that the cell states of proliferative cells approximately reside on a hollow hypercylinder manifold. In step one, Revelio represents the data in a high-dimensional principal component (PC) space and then rotates the coordinate frame to define new axes referred to as dynamical components (DC). The first two DCs, DC1 and DC2, respectively, of the high-dimensional DC space encapsulate most of the cell cycle-dependent variation. In step two, we used an iterative finite temperature string method15,39,40 to obtain a one-dimensional cyclic cell cycle (CC) coordinate on the DC1–DC2 plane (Fig. 3a subpanel 2 and Fig. S3a–f). Next, we identified the cell cycle division point, the split point, where we then unraveled the cell cycle coordinate to form a linear axis with a periodic boundary condition reflecting cell cycle progression (Fig. 3a subpanel 3 and 4). The remaining DCs orthogonal to the DC1–DC2 plane contain variations in the data that are mostly cell cycle independent. Thus, in our subsequent analyses, we employed the CC coordinate and the third dynamical component, DC3, to form a low-dimensional representation describing cell cycle progression and other cellular processes within the data (Fig. 3a subpanel 4). Details are provided in the “Method” section.
The cell cycle representation revealed cycling and quiescent dynamics
We first applied this computational pipeline to the untreated MCF10A cells. Indeed, single-cell data on the DC1-DC2 plane captured the cyclical nature of the cell cycle, which was further reflected in the vector field (Fig. S3g). The obtained CC coordinate accurately captured cell cycle progression, as the gene expression profiles of CCNE1, TOP2A and other cell cycle-related genes showed expected elevated expression at either the G1 or G2/M phase of the CC coordinate (Figs. 3b, c and S4a–c).
In addition, the cycling region previously identified in the UMAP space (region 1, Fig. 1d) was distributed across all cell cycle phases of the cell cycle when mapped to the new CC representation (Fig. S4d, e). The previously identified quiescent region, region 2 (Fig. 1d) ended up concentrated in the upper region of the DC3 axis in the new representation, where further down the DC3 axis marked the transition from quiescence to entry into the cell cycle (Fig. S4d, e). Furthermore, the gene expression of DEK and PARP1, genes identified as universally down-regulated during quiescence41, had reduced expression in the identified quiescent region and higher expression in the actively proliferating region (Fig. 3d, e).
The vector field obtained from the untreated MCF10A cells using the CC-DC3 representation further provided a dynamic view of cell cycle progression, revealing a bifurcation between proliferation and quiescence (Fig. 3f). Above the bifurcation, the vector field flowed upward, indicating cells progressing deeper into quiescence, while below the bifurcation, the downward vectors signal entry into the cell cycle. Such a bifurcation could reflect the quiescence-proliferation transition controlled by a RB-E2F bistable switch, where, after mitosis, daughter cells either enter quiescence or proceed to the G1 phase of the cell cycle42. In conclusion, the CC-DC3 representation correctly captured the quiescence-proliferation dynamics.
The cell cycle-EMT representation revealed coupling between the cell cycle and EMT
Next, we applied our computational pipeline to the 12.5 and 25 pM TGF-β-treated MCF10A cells. Consistent with what was observed with the untreated cell data, the CC coordinate accurately depicted cell cycle progression, corroborated by high expression of CCNE1 and TOP2A in the G1 and G2/M phase, respectively (Fig. S5a–d). Gene expression analysis confirmed that the new y-axis effectively captured EMT progression, with decreasing gene expression levels of CDH1, an epithelial related gene (Fig. S5e, f) and increasing gene expression levels of VIM and FN1, mesenchymal related genes (Fig. S5g–j). That is, as cells moved upward along the EMT axis, they gradually lost their epithelial features and gained mesenchymal features.
Unlike the vector fields in the 2D UMAP representation for 12.5 and 25 pM TGF-β treated cells, the vector field on the CC-EMT representation depicted cells undergoing EMT originating broadly from either the G1 or G2/M phase (Figs. 3g, h and S5k, l). The widespread distribution suggested that the cell cycle and EMT were loosely coupled. If they were tightly coupled, one would expect cells to exit from a narrow region of the cell cycle, likely corresponding to a cell cycle checkpoint.
Overall, our computational pipeline obtained a representation that captured cell cycle progression along a single axis, which enabled the extraction of valuable biological insight, specifically the broad coupling between the cell cycle and EMT.
Trajectory simulations revealed two EMT paths originating from either G1 or G2/M in a multi-dimensional state space
To confirm that the coupling between the cell cycle and EMT at the G1 and G2/M cell cycle phases was not an artifact of dimensionality reduction, we performed trajectory simulations and transition path analyses in a 30-dimensional PCA space, where the vector fields were originally constructed. To obtain trajectories in high-dimensional space, we first identified an epithelial and mesenchymal region from the CC-EMT representation, designating them as potential start and end points, respectively (Fig. 4a). Using the transition matrix, derived from cell neighbor and RNA velocity information25,27, we launched trajectories from the epithelial start region and recorded trajectories that hit the mesenchymal end region, referred to as reactive trajectories.
a Trajectory simulation set-up for 12.5 pM TGF-β treated MCF10A cells with specified start (epithelial) and end (mesenchymal) region. Example trajectories from one round of 12.5 pM (b) and 25 pM (c) TGF-β simulations mapped onto the EMT-cell cycle coordinate representation. Mean trajectories from the d 12.5 pM and e 25 pM TGF-β simulations mapped on to the EMT-cell cycle coordinate representation. Mean trajectories for two clusters in the 3-d DC-EMT space for the 12.5 pM (f) and 25 pM (g) TGF-β simulations. Cell cycle phases were assigned using Revelio.
For both the 12.5 and 25 pM TGF-β-treated cells, we conducted ten rounds of trajectory simulations, generating 100 trajectories per simulation (Fig. 4b, c) and found that they grouped into two distinct clusters (Fig. S6a, b). With these two clusters of trajectories, we used a modified finite temperature string method15,39 to obtain the mean trajectory, similar to that of reaction coordinates in chemical reaction dynamics, per cluster. Projecting this ensemble of mean trajectories to the CC–EMT representation, we observed that the two clusters originated from either the G1 or G2/M phase (Fig. 4d, e). Inspection in 3D further confirmed that the two mean trajectories originated from either the G1 or G2/M phase (Fig. 4f, g).
It is important to note that the number of clusters was user defined, originally set as two, for both the 12.5 and 25 pM TGF-β trajectory simulations. We also tested whether the trajectories could be clustered into three groups. However, for both TGF-β doses the third mean trajectory aligned with either the already established G1 or G2/M trajectories, rather than forming a new path (Fig. S6c–f).
EMT-related genes between the G1 and G2/M paths both exhibited an increase in expression along the EMT axis. However, the G2/M path generally displayed lower expression at the beginning compared to the G1 path (Fig. S7). Additionally, SNAI2, an EMT transcription factor, IL1B, S100A14, C1R, all related to innate immunity, and TNC, which encodes a large extracellular matrix protein, showed differential expression between the two paths (Fig. S7). Further analysis is required to investigate the biological implications of these EMT path differences.
Overall, the trajectory simulation analyses confirmed that MCF10A cells undergoing TGF-β-induced EMT could take one of two paths, transitioning broadly from either the G1 or G2/M region in high-dimensional space.
Multiplex immunostaining confirmed two transition regions in TGF-β treated MCF10A cells
Analyses of scRNA-seq from both 12.5 and 25 pM TGF-β-treated MCF10A cells identified the G1 and G2/M phases as key regions where cells undergo EMT. To test whether these transition regions could be identified on the protein level, we treated MCF10A cells with 0.55 ng/ml (22.5 pM) of TGF-β for 0, 3, and 14 days (Fig. 5a). Cell morphology shifted from the typical round epithelial shape to an elongated, spindle-like form (Fig. 5b). These changes were classical indicators of EMT, confirming that EMT had occurred by 14 days of TGF-β treatment37. Not only did we observe a change in morphology, but we also observed changes in cellular markers. To characterize the changes in cellular markers and morphology, we performed two rounds of iterative indirect immunofluorescence imaging (4i)43 staining (round 1: cyclin A, cyclin B1, vimentin, DAPI, round 2: cyclin D1, DAPI) (Fig. 5a). We selected these proteins to determine the cell cycle phases: cyclin D1 as a marker for the G1 phase, cyclin A and cyclin B1 together as markers for the G2/M phase, and DAPI to determine DNA copy number and vimentin to identify EMT state. An inspection of the vimentin fluorescence intensity revealed that vimentin expression increased from day 0 to day 3, indicating that cells were beginning to undergo EMT (Fig. 5b, c). By day 14, there was a significant rise in vimentin intensity, suggesting that most cells had completed the EMT process (Fig. 5b, c).
a Experimental design: MCF10A cells were treated with TGF-β for 0, 3 and 14 days, followed by two rounds of 4i staining for cell cycle and EMT protein markers. b Representative differential interference contrast (DIC) images (first row) and representative fluorescence images (second row) of vimentin expression in MCF10A cells after 0, 3 and 14 days of TGF-β treatment. Scale bar, 30 μm. c Boxplots showing scaled vimentin expression (0–1) across treatment days. PHATE representation of untreated (d), day 3 (e), and day 14 (f) TGF-β-treated MCF10A cells colored by DAPI intensity. Gray circles indicate untreated reference cells. PHATE-vimentin representation of day 3 (g) and day 14 (h) treated cells, colored by DAPI intensity. PHATE-vimentin representation of day 3 (i) and day 14 (j) treated cells, colored by vimentin. PHATE-vimentin representation of day 3 (k) and day 14 (l) treated cells, with the G1/high vimentin and G2/M-high vimentin regions labeled. The Vim z-axis indicates the relative fluorescence intensity (RFI).
We then learned a manifold representing cell proteomic states across the three time points, with the features we extracted from 4i using PHATE to project the manifold into a 2D space for visualization. The PHATE representation revealed a clear circular cell cycle structure for the untreated cells (Figs. 5d and S8a). Using the untreated cell as a reference, we observed cells in day 3 and 14 in different stages of the cell cycle (Fig. 5e, f).
To examine the coupling between the cell cycle and EMT, we utilized a 3D representation combining the 2D PHATE representation with vimentin intensity as the third axis. For day 3 and day 14, we observed two clusters corresponding to either the G1 or G2/M phase (Fig. 5g, h and S8b–j). Within each cluster, a subset of cells exhibited high vimentin intensity (Fig. 5i, j). The intersection between the G1 or G2/M groups with high vimentin intensity exhibited the coupling we computationally predicted previously between the cell cycle and EMT (Fig. 5k, l). In addition, cells with high vimentin intensity were not confined to a specific region within the G1 or G2/M phase but were broadly distributed, further supporting the loose coupling between the cell cycle and EMT. Interactive 3D plots were provided (Supplementary Data 1–12).
In summary, multiplexed staining and manifold analysis confirmed our computational predictions from analyzing the scRNA-seq data. The proteomic data further corroborated that the cell cycle and EMT loosely couple during TGF-β induced EMT in MCF10A cells, resulting in cells broadly arresting from either the G1 or G2/M phase of the cell.
Discussion
When a proliferating cell undergoes a cell phenotypic transition, it typically needs to temporarily pause or permanently exit the cell cycle to coordinate resource allocation2. Intuitively, one expects that the exit takes place at only a few restricted regions, e.g., cell cycle checkpoints, of the cell cycle, while arrest can take place at various cell cycle stages. The present analysis of single-cell transcriptomic data combined with multiplex 4i protein staining data of MCF10A cells undergoing TGF-β-induced EMT revealed that the arrest points were broadly distributed within the G1 or G2/M phases of the cell cycle.
The observed pattern can be visualized as a wheel and a lever attempting to stop the rotating wheel (Fig. 6). The rotating wheel represents the cell cycle, while the lever represents EMT. With increasing concentrations of TGF-β, more force is applied to the EMT lever, eventually exerting pressure on to the wheel and disrupting cell cycle progression. When the EMT lever comes in contact with the rotating wheel, the cells in this region of the cell cycle halt their progression and transition from the epithelial to the mesenchymal state. This analogy illustrates how TGF-β treatment can induce widespread cell cycle arrest in either the G1 or G2/M phase17,18,19 while also promoting EMT. The difference in EMT programs between the two paths implies a potential mechanism for preferentially activating one path over the other, e.g., the G1 path at normal developmental and physiological conditions as activation of the G2 path could lead to aberrations such as polyploidy. The differential expression along the two paths may also contribute to heterogeneous cellular responses and drug resistance. Further studies are needed to elucidate the implications of the two EMT paths in the context of cell physiology and pharmaceutical intervention. Generally speaking, the observed coupling between cell cycle and EMT underscores the importance of quantitatively understanding the crosstalk between cell phenotypic transitions and the cell cycle when developing biomedical interventions.
Cell cycle and EMT coupling at low (left) and increasing TGF-β concentrations (right).
To perform single-cell data analysis, the standard procedure is to apply a dimensionality reduction method to high-dimensional data to obtain a two-dimensional representation for visualizing such complex data. Many dynamics inference methods, such as trajectory inference, are typically performed on low-dimensional representations of single-cell data, with an implicit assumption that the representation faithfully reflects the cellular processes being studied44. Single-cell omics approaches characterize cell state heterogeneity by quantifying the transcriptomic, proteomic, or epigenomic profiles of individual cells. Thus, a grand challenge is how to identify cellular processes relevant to a specific question of interest. A representation obtained with a brute force dimensionality reduction procedure, which emphasizes the largest contributions to heterogeneity, can conceal or distort the contributions of relevant processes, such as the cell cycle, as shown in this study. Furthermore, most dimensionality reduction methods do not include dynamic information, further worsening the situation. Our computational pipeline, which built upon the Revelio cell cycle framework31 to obtain the cell cycle coordinate, underscores the importance of incorporating biological information when visualizing data in low-dimensional representations.
Examining cellular dynamics in a high-dimensional space, aided by dynamical information such as a transcriptomic vector field, reduces mechanistic ambiguity. With the full dynamical model using RNA velocity information, we performed single-cell trajectory simulations in a high-dimensional state space and confirmed the existence of two distinct trajectories coupling cell cycle and EMT. Note that single-cell trajectories here differ from conventionally use of “trajectory” inference in the single-cell field. The latter typically refers to mean trajectories followed by an ensemble of cells that corresponds to the mean paths discussed in this work. In our case, single-cell trajectories refer to trajectories on a single-cell level.
This study demonstrates how biology-informed representations can reveal mechanistic insights into cellular processes from snapshot single-cell data, which are not transparent otherwise. Future studies can expand along several directions. We followed the Revelio model by assuming that one can approximate the single-cell data manifold to be cylindrical-shaped to describe cell cycle-dependent and -independent variations. While this cylindrical-shape assumption is a valid approximation for cells in homeostasis, as shown in Schwabe et al.31, in the case of multiple cell types, the assumption can be overly simplistic. However, it was a good starting point for our analysis. In addition, we restricted our transition path analyses to a specific final mesenchymal state under several low concentrations of TGF-β. The MCF10A data at higher concentrations of TGF-β suggests the coexistence of different mesenchymal states, thus, more than two types of EMT transitions paths may exist. Moreover, one can extend the trajectory analyses to investigate the coupling between various cellular processes, such as EMT, gain or loss of stemness, and various other cellular fates, such as apoptosis and senescence. Systematic identification of these different transition processes may require trajectory analyses without pre-selected start and end states. In short, improvements in cell state resolution and data sampling, combined with biology-informed in-silico trajectory analyses and experiments, enable systematic characterization and mechanistic insight into how different cellular processes couple to respond to one or multiple environmental and intracellular stimuli.
Methods
Processing of single-cell RNA-seq datasets
The MCF10A scRNA-seq dataset (GSE213753) consists of human mammary epithelial (MCF10A) cells that were treated with increasing concentrations of TGF-β (0, 12.5, 25, 50, 100, 200, 400, and 800 pM) for 14 days and then sequenced. The raw sequencing reads in the form of BAM files were first converted to fastq files using 10x Genomics Cell Ranger bamtofastq version 1.3.0 utility, then Velocyto version 0.17.17 was used to obtain the unspliced and spliced count matrices.
Single-cell RNA-seq analysis
After processing the MCF10A dataset, there was a total of 8983 cells. To obtain a shared UMAP representation for visual comparison between each of the TGF-β doses, all doses were normalized using dynamo version 1.0.0 “recipe_monocle” function where the n_top_genes = 1000, then PCA dimensionality reduction (n_components = 30) was performed, and finally UMAP (n_components = 2) was performed.
To calculate the EMT score, we used a two-sample KS test33, which compares the empirical cumulative distribution function (ECDF) of epithelial and mesenchymal gene sets using SciPy’s “ks_2samp” function. The KS test statistic and p-value for the ECDFEpi ≥ ECDFMes and ECDFEpi ≤ ECDFMes scenarios were calculated. A cell with a negative score is epithelial, while a cell with a positive score is mesenchymal.
To determine which TGF-β doses contained epithelial cells for EMT analysis, we calculated the percentage of area overlap with the untreated data in the UMAP space. This was achieved by configuring an ellipse for each dose. First, the mean coordinate was calculated to obtain the center of the ellipse. Then the eigenvalues from the covariance matrix were calculated to obtain the width and height of the ellipse. The angle of the ellipse was determined using the eigenvectors. To calculate the percentage of area overlap, the difference between the area of the untreated and treated data was taken and then divided by the area of the untreated data. The TGF-β doses with less than 5% area overlap were excluded from analysis.
For downstream analysis, each TGF-β dose was normalized separately “recipe_monocle” function, where the n_top_genes = 2000 to calculate RNA velocity and then the corresponding vector field was obtained using dynamo. For downstream analysis, specifically for the 0, 12.5 and 25 pM TGF-β treated cells, a total of 3787 cells were analyzed.
Assigning cell cycle phase
Two methods were used to determine the cell cycle phase of each cell, dynamo’s “cell_cycle_score” function27 and Revelio’s “getCellCyclePhaseAssignInformation” function31. Both methods follow a similar approach, where each phase is defined by a list of genes. The average gene expression for each phase is calculated, z-normalized per phase and per cell, and the phase with the highest score is the assigned cell cycle phase. The difference between the two methods is that dynamo’s function assigns cells as either M, M/G1, G1/S, S or G2 while Revelio’s function assigns cells as either G1/S, S, G2, G2/M, or M/G1. These assigned discrete cell cycle phase was used to compare with the continuous cell cycle coordinate.
Differential gene expression analysis
Scanpy’s version 1.9.8 “rank_gene_groups” function (method = Wilcoxon), a nonparametric Mann–Whitney U test, was used to identify the top 10 genes that characterize region 1 and 2 (Fig. 1g).
To characterize EMT progression along the UMAP1 axis for both the 12.5 and 25 pM TGF-β-treated cells, starting with an initial list of EMT-related genes8,39,45, we performed Louvain clustering, resolution = 0.4, and obtained two clusters. Using the “rank_genes_groups” function (method = Wilcoxon), we obtained a ranking of the genes most representative of each cluster. With this list of genes, the gene expression of the top ranked genes for the 125 nearest neighbor cells to region a and region b was compared (Fig. S2e).
Constructing the vector field from splicing-based RNA velocities
The Python package, dynamo version 1.0.0, was utilized to obtain the vector fields27. Dynamo builds upon snapshot single-cell data with RNA velocities inferred with various methods. For the datasets used in this study, we inferred the velocities from the unspliced (mRNA with introns) and spliced (mature mRNA without introns) RNA counts. The RNA turnover dynamics for a specific gene i can be described in the following ordinary differential equations25,
where \({u}_{i}\) represents the number of unspliced RNA transcripts for gene i, \({s}_{i}\) represents the number of spliced RNA transcripts of gene i, \(\frac{{{du}}_{i}}{{dt}}\) represents the change rate in unspliced RNA transcripts, \(\frac{{{ds}}_{i}}{{dt}}\) represents the change rate in spliced RNA transcripts, \({\alpha }_{i}\) represents the transcription rate of gene i, \({\beta }_{i}\) represents the splicing rate constant of transcript \({u}_{i}\), and \({\gamma }_{i}\) represents the degradation rate constant of transcript \({s}_{i}\). Assuming a pseudo-steady state approximation with \(\frac{{{du}}_{i}}{{dt}}=0\), the conventional RNA velocity for a gene i can be defined as, \(\frac{{{ds}}_{i}}{{dt}}={u}_{i}-\,\widetilde{{\gamma }_{i}}{s}_{i}\), where \(\widetilde{{\gamma }_{i}}=\,\frac{{\gamma }_{i}}{{\beta }_{i}}\) is a scaled gene-specific degradation rate. RNA velocity can be estimated for each gene in each cell thus, the state change rate of a cell can be represented by a high-dimensional vector of gene velocities. With a further assumption that all genes have the same splicing rate constant \(\beta\), one can use the velocity vector to approximate the transition direction of individual cells in the transcriptomic state space.
Projecting a vector field into a low-dimensional representation
For visualization purposes, the high-dimensional velocities obtained from dynamo are often projected onto a two or three low-dimensional representation. This is commonly achieved by using Pearson, cosine kernels25,26,46,47, or mathematically rigorous GraphVelo method that is based on local linear embedding47 to observe cell fate transitions.
Vector field analysis
Dynamo’s objective is to reconstruct a continuous vector field, i.e., a mechanistic systems biology virtual-cell model, from scRNA-seq gene expression and instantaneous RNA velocity data. The continuous vector field can be described as \( < \frac{d{{\boldsymbol{x}}}}{{dt}} > ={{\boldsymbol{F}}}\left({{\boldsymbol{x}}}\right)\), where \({{\boldsymbol{x}}}\) represents the gene expression state, \( < \frac{d{{\boldsymbol{x}}}}{{dt}} > \) represents the RNA velocity at \({{\boldsymbol{x}}}\) averaged over its k-Nearest Neighbor (kNN) neighborhood, and F represents the learned vector field that maps the gene expression space to the velocity space, reflecting the gene-gene regulations of the system27,39,47.
With the continuous vector field, we can investigate the topology of the vector field through fixed point analysis. There are three types of fixed points: stable, unstable, and saddle points. Stable points are regions where the velocity vectors flow inward from all neighboring cells, such as a stable cell type. Unstable points are regions where the velocity vectors only flow outward, such as a pluripotent cell state, which tends to differentiate quite quickly. Saddle points are regions where some velocity vectors flow inward and some outward, such as a differentiated pluripotent cell, which is relatively stable but also can further differentiate.
Obtaining the one-dimensional cell cycle coordinate
We used a finite temperature string method to define a one-dimensional cell cycle coordinate. The method was originally designed for defining a one-dimensional reaction coordinate that describes the progression of a chemical reaction40 process, and has been adapted to analyze cell state transitions15,39. The finite-temperature string method assumes that all trajectories connecting a start and end region in the state space forms a narrow hyper-tube, and the center line of the hyper-tube is used to define a 1-D reaction coordinate that connects the initial and final regions through an iterative procedure.
While the finite-temperature string algorithm can be applied to find the 1-D path embedded in a multi-dimensional state space, here we restricted it to the DC1–DC2 plane defined by Revelio31. We designed an iterative procedure specifically for identifying the closed cell cycle coordinate (Fig. S3). Since the two DC modes contain most of the cell cycle information, the single cell data points already form a ring-shaped point cloud. We first calculated the center point of the data and generated a circle with a random radius from the center point as our initial guess of the cell cycle path. We then chose two consecutive points along this circle as our starting and ending points. Due to the iterative nature of this procedure, the final path is not sensitive to this choice. Next, we iterated through the following steps,
-
1)
We placed N points, including the starting and ending, called images, along the guessed cell cycle path uniformly with equal arclength distance. The image points divided the whole (2D) state space into Voronoi cells, with each data point assigned to the Voronoi cell with the shortest Euclidean distance. For each Voronoi cell, the centroid of the single cell data was calculated and assigned as a corresponding new image point, except the starting and ending points.
-
2)
With the new image points, we then obtained a new guess of the continuous cell cycle path through cubic spline interpolation, and repeated step 1 until no more single cell data points were reassigned to Voronoi cells.
-
3)
With the path from step 2, we chose an image point, typically far from the initial chosen starting and ending points, to calculate a new radius from the center of the data. With this new radius, a circle was generated as an improved initial guess, and two consecutive points were selected as the starting and ending points. With this improved guess, the above two steps are repeated. One can repeat such re-optimization, but in practice, we found one round was sufficient.
With the final image path from step 3, each cell was assigned a cell cycle coordinate value through interpolation between two nearest image points.
Determining the new cell cycle representation
To obtain the final two-dimensional representation that allowed us to observe how the cell cycle couples with other biological processes, a second axis representing the biological process of interest was needed. The cell by DC matrix output by Revelio showed that the first two dimensions capture a majority of the cell cycle dependent variation therefore, the remaining dimensions must contain the cell cycle independent variation, most likely the biological process being studied. The third DC dimension was designated as the new y-axis to create the new two-dimensional representation for the untreated, 12.5 and 25 pM TGF-β treated cells.
Generating gene expression heatmap and expression plots along the cell cycle coordinate
Since we identified two regions, a quiescent and cycling region, in the untreated MCF10A cells, we selected for only cycling cells to accurately capture the gene expression of cell cycle-related genes along the cell cycle coordinate. To generate the heatmap, the gene expression of several G1 and G2 genes from Schwabe et al.31 were averaged along each point of the cell cycle coordinate. Cells with zero expression were dropped, and each gene was normalized using standard scaling. Similarly, for the expression plots, for each gene, the expression was averaged along each point of the cell cycle coordinate, with zero-expression cells dropped. The smoothed dashed line representing a genes expression pattern used SciPy’s “make_smoothing_spline” function with the regularization parameter set to 1000.
Performing trajectory simulations
The trajectory simulations utilize a transition matrix defined in 30-d PCA space. The transition matrix was obtained using the cosine kernel method25,27. The method uses RNA velocity information to estimate the transition probability (pij) from a cell state i to another neighboring state j in the 30-d state space, \({P}_{{ij}}=\frac{\exp \left[\cos ({{{\bf{v}}}}_{i},{{{\boldsymbol{\delta }}}}_{{ij}})/\sigma \right]}{{\sum }_{m}\exp \left[\cos ({{{\bf{v}}}}_{i},{{{\boldsymbol{\delta }}}}_{{im}})/\sigma \right]}\), with \({{{\bf{v}}}}_{i}\) the RNA velocity vector of cell state i, \({{{\boldsymbol{\delta }}}}_{{ij}}={{{\boldsymbol{x}}}}_{j}-{{{\boldsymbol{x}}}}_{i}\) where \({{{\boldsymbol{x}}}}_{i}\) and \({{{\boldsymbol{x}}}}_{j}\) the expression states of cell state i and j, \(\cos (\cdot ,\cdot )\) denoting the cosine similarity between two input vectors, \(\sigma\) an arbitrary bandwidth parameter (assumed a value of 10 in this study), and the sum is over all kNN neighbors of cell i.
Following chemical rate theory, we performed transition path theory analysis15,39,48 as the following. Aided with the 2D CC-EMT coordinate representation, we chose two threshold values of the EMT coordinate (y), \({\epsilon }_{1} < {\epsilon }_{2}\), d divided the state space into three regions, Epithelial (\({y < \epsilon }_{1}\)), Intermediate (\({\epsilon }_{1}\le y{\le \epsilon }_{2}\)), and Mesenchymal ((\({y > \epsilon }_{2}\)),). We varied the values of \({\epsilon }_{1}\) and \({\epsilon }_{2}\), and found the subsequent analyses were insensitive to the choice, a feature common in transition path theory analysis48. Using the transition matrix, we randomly selected experimentally sampled single-cell data within the Epithelial region as starting points, and launched Gillespie trajectory simulations49 in the full-dimensional state space. Following transition path theory, we termed trajectories starting from the epithelial region and ending at the mesenchymal region without returning to the epithelial region as reactive trajectories. Trajectories that returned to the start region before hitting the end region, termed non-reactive trajectories, were discarded.
Clustering and identifying paths in high dimensional space
For both the 12.5 and 25 pM TGF-β-treated MCF10A cells, ten rounds of 100 reactive trajectories were acquired, totaling 1000 trajectories for each dose. For each trajectory simulation round, the “TimeSeriesKMeans” (n_clusters, metric = softdtw, max_iter = 10, random_state = 0, init = k-means++) function from tslearn version 0.6.3 was applied to cluster the trajectories in the 10-dimensional DC space. The number of clusters, n_clusters, was user defined, but we checked for two and three clusters to ensure that there were only two clusters.
To obtain a mean trajectory for each cluster, we used an adapted finite temperature string method from Wang et al.15,39. The procedure was similar to what was used in obtaining the cell cycle coordinate, but with two major differences. First, the procedure was performed in the leading 10-d DC space. Second, the reactive trajectories also have weights on determining the image points in each Voronoi cells, as detailed in refs. 15,39.
To map the mean trajectories from DC space to other spaces (e.g., CC-EMT) the nearest neighbor cell in DC space was identified and its corresponding coordinate in the target space was used. To compare the mean trajectories obtained in 10-d DC space from each round of trajectory simulations, fastdtw version 0.3.4 was utilized to calculate the Euclidean distance between two trajectories. Pairwise comparisons of all mean trajectories generated a heatmap revealing the number of clusters.
Comparing EMT expression of the two paths
With one set of mean trajectories from one round of 12.5 pM TGF-β trajectory simulations, more points were interpolated using (Fig. S4d) SciPy’s UnivariateSpline function, with a smoothing factor of 0.5, obtaining x coordinates given a set of coordinates along the EMT axis. Having a shared EMT axis allows us to make a one-to-one comparison between the two paths. With these interpolated mean trajectories for the G1 and G2/M paths in CC-EMT space, we identified the corresponding cell states or nearest neighbor in DC space and then converted it into gene expression space using a rotation matrix obtained from Revelio31.
Cell culture
Human mammary epithelial cells (MCF10A; ATCC, CRL-10317) were cultured in DMEM/F-12 medium supplemented with 5% horse serum, 20 ng/mL epidermal growth factor (EGF), 0.01 mg/mL insulin, 500 ng/mL hydrocortisone, and 100 ng/mL cholera toxin. Cells were maintained in a humidified atmosphere of 5% CO₂ at 37 °C and utilized within 10 passages.
For the TGF-β treatment, 1 × 105 MCF10A cells were plated in 35 mm glass-bottom plates (Ibidi, #80137) and cultured for 2–3 days. When cells reached 70–80% confluency, 0.55 ng/mL TGF-β was added to the culture media. The culture media with TGF-β was refreshed every 24 h. TGF-β-treated cells were fixed with 4% PFA after 3 and 14 days, respectively. The fixed cells were then stored at 4 °C for immunofluorescence staining.
4i staining and imaging
Iterative Indirect Immunofluorescence Imaging (4i) staining was performed following the protocol published by Kramer et al.50. The immunofluorescence staining was carried out in two rounds: round 1: cyclin A (1:50, Santa Cruz, sc-271682), cyclin B1 (1:100, R&D, AF6000), vimentin (1:150, Thermofisher, PA5-27231), and DAPI; round 2: cyclin D1 (1:100, Santa Cruz, sc-20044) and DAPI. A Nikon Ti-E2 microscope was used to image under 40X (0.95NA). The following channels were used: DAPI, TRITC, FITC, Cy5. The final image was stitched from 8 × 8 tiles (each 1024 × 1024 pixels) with a stitching overlap of 15% to get one 7117 × 7117-pixel image. All imaging for each channel staining marker was conducted using identical microscope settings and performed on the same day.
Image analysis
For each image, we first subdivided each 7117 × 7117-pixel field into a 4 × 4 grid of 16 tiles as input to the pre-trained Cellpose cyto3 model (3.0.9.dev8+gc03958c)51. Segmentation was performed on each tile using differential interference contrast (DIC) and vimentin channels to generate cell masks. The DAPI channels were used to generate nuclear masks. Nuclear masks required no manual correction due to the high signal-to-noise ratio of DAPI, however, cell contours were manually refined using the DIC channel as reference. Subtle offsets between the two rounds of imaging were corrected by registering nuclear masks with a phase cross-correlation algorithm in the scikit-image package version 0.20.0. We then applied the same shift parameters to the other channels. Channel-specific background intensities were measured in ImageJ and subtracted from every image.
Multiple features were extracted from the images, including the ratio of nuclear mean intensity over cytoplasmic mean intensity for cyclin D1, A, and B1, total nuclear DAPI and total cell vimentin intensity. All features except the total cell vimentin intensity from the three independent experiments were pooled and z-score normalized. Using these features, we applied PHATE version 1.0.11 (k-nearest neighbors = 45) to learn a two-dimensional shared manifold representing cell states across all three time points. Total cell vimentin intensity was then overlaid as the third axis to visualize EMT progression across different days of treatment in a 3D representation.
Clustering was run separately for the three groups: no treatment, 3, and 14 days TGF-β-treated. We used scikit learn’s KMeans clustering with k set to 2 and all other parameters kept at their default settings. The input to KMeans was a one column array containing the DAPI sums for all cells in the given group. After fitting, the two clusters were ordered based on their mean DAPI intensity to maintain a consistent labeling across groups.
Statistics and reproducibility
To reproduce the analyses performed the scripts are available on GitHub at https://github.com/xing-lab-pitt/two-paths.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the raw sequencing data for the MCF10A dataset is available in the NCBI Gene Expression Omnibus under the accession GSE213753. The features extracted from 4i can be found in the supplemental.
Code availability
All code can be found on GitHub https://github.com/xing-lab-pitt/two-paths.
References
Dalton, S. Linking the cell cycle to cell fate decisions. Trends Cell Biol. 25, 592–600 (2015).
Soufi, A. & Dalton, S. Cycling through developmental decisions: how cell cycle dynamics control pluripotency, differentiation and reprogramming. Development 143, 4301–4311 (2016).
Giacinti, C. & Giordano, A. RB and cell cycle progression. Oncogene 25, 5220–5227 (2006).
Nègre, N., Ghysen, A. & Martinez, A.-M. Mitotic G2-arrest is required for neural cell fate determination in Drosophila. Mech. Dev. 120, 253–265 (2003).
Ayeni, J. O. et al. G2 phase arrest prevents bristle progenitor self-renewal and synchronizes cell division with cell fate differentiation. Development 143, 1160–1169 (2016).
Gomer, R. H. & Firtel, R. A. Cell-autonomous determination of cell-type choice in dictyostelium development by cell-cycle phase. Science 237, 758–762 (1987).
Matthews, H. K., Bertoli, C. & de Bruin, R. A. M. Cell cycle control in cancer. Nat. Rev. Mol. Cell Biol. 23, 74–88 (2022).
Yang, J. et al. Guidelines and definitions for research on epithelial–mesenchymal transition. Nat. Rev. Mol. Cell Biol. 21, 341–352 (2020).
Nieto, M. A., Huang, R. Y., Jackson, R. A. & Thiery, J. P. Emt: 2016. Cell 166, 21–45 (2016).
Emadi Baygi, M. et al. Slug/SNAI2 regulates cell proliferation and invasiveness of metastatic prostate cancer cell lines. Tumour Biol. 31, 297–307 (2010).
Mejlvang, J. et al. Direct repression of cyclin D1 by SIP1 attenuates cell cycle progression in cells undergoing an epithelial mesenchymal transition. Mol. Biol. Cell 18, 4615–4624 (2007).
Schuhwerk, H. et al. The EMT transcription factor ZEB1 governs a fitness-promoting but vulnerable DNA replication stress response. Cell Rep. 41, 111819 (2022).
McFaline-Figueroa, J. L. et al. A pooled single-cell genetic screen identifies regulatory checkpoints in the continuum of the epithelial-to-mesenchymal transition. Nat. Genet. 51, 1389–1398 (2019).
Karacosta, L. G. et al. Mapping lung cancer epithelial-mesenchymal transition states and trajectories with single-cell resolution. Nat. Commun. 10, 5587 (2019).
Wang, W., Poe, D., Yang, Y., Hyatt, T. & Xing, J. Epithelial-to-mesenchymal transition proceeds through directional destabilization of multidimensional attractor. eLife 11, e74866 (2022).
Wang, W. et al. Live-cell imaging and analysis reveal cell phenotypic transition dynamics inherently missing in snapshot data. Sci. Adv. 6, eaba9319 (2020).
Takahashi, K. et al. TGF-beta generates a population of cancer cells residing in G1 phase with high motility and metastatic potential via KRTAP2-3. Cell Rep. 40, 111411 (2022).
Lovisa, S. et al. Epithelial-to-mesenchymal transition induces cell cycle arrest and parenchymal damage in renal fibrosis. Nat. Med. 21, 998–1009 (2015).
Qi, R. et al. Snai1-induced partial epithelial–mesenchymal transition orchestrates p53–p21-mediated G2/M arrest in the progression of renal fibrosis via NF-κB-mediated inflammation. Cell Death Dis. 12, 44 (2021).
Cockburn, K. et al. Gradual differentiation uncoupled from cell cycle exit generates heterogeneity in the epidermal stem cell layer. Nat. Cell Biol. 24, 1692–1700 (2022).
Lahnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Method of the year 2013. Nat Methods 11, 1 (2014).
Deconinck, L., Cannoodt, R., Saelens, W., Deplancke, B. & Saeys, Y. Recent advances in trajectory inference from single-cell omics data. Curr. Opin. Syst. Biol. 27, 100344 (2021).
Schiebinger, G. et al. Optimal-transport analysis of single-. Cell Gene Expr. Identifies Dev. Trajectories Reprogramming. Cell 176, 928–943 e22 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Qiu, X. et al. Mapping transcriptomic vector fields of single cells. Cell 185, 690–711 (2022).
Zhang, Y. et al. Graph-dynamo: learning stochastic cellular state transition dynamics from single cell data. Preprint at bioRxiv https://doi.org/10.1101/2023.09.24.559170 (2023).
Chari, T. & Pachter, L. The specious art of single-cell genomics. PLoS Comput. Biol. 19, e1011288 (2023).
Wang, S., Sontag, E. D. & Lauffenburger, D. A. What cannot be seen correctly in 2D visualizations of single-cell ‘omics data? Cell Syst. 14, 723–731 (2023).
Schwabe, D., Formichetti, S., Junker, J. P., Falcke, M. & Rajewsky, N. The transcriptome dynamics of single cells during the cell cycle. Mol. Syst. Biol. 16, e9946 (2020).
Panchy, N., Watanabe, K., Takahashi, M., Willems, A. & Hong, T. Comparative single-cell transcriptomes of dose and time dependent epithelial-mesenchymal spectrums. NAR Genom. Bioinform. 4, lqac072 (2022).
Tan, T. Z. et al. Epithelial-mesenchymal transition spectrum quantification and its efficacy in deciphering survival and drug responses of cancer patients. EMBO Mol. Med. 6, 1279–1293 (2014).
McInnes, L., Healy, J. & Melville, J. Umap: uniform manifold approximation and projection for dimension reduction. Preprint at https://doi.org/10.48550/arXiv.1802.03426 (2018).
Cheng, Y.-C. et al. Reconstruction of single-cell lineage trajectories and identification of diversity in fates during the epithelial-to-mesenchymal transition. Proc. Natl. Acad. Sci. USA 121, e2406842121 (2024).
Lause, J., Berens, P. & Kobak, D. The art of seeing the elephant in the room: 2D embeddings of single-cell data do make sense. PLoS Comput. Biol. 20, e1012403 (2024).
Zhang, J. et al. TGF-β-induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops. Sci. Signal. 7, ra91 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Wang, W., Ni, K., Poe, D. & Xing, J. Transiently increased coordination in gene regulation during cell phenotypic transitions. PRX Life 2, 043009 (2024).
E, W., Ren, W. & Vanden-Eijnden, E. Finite temperature string method for the study of rare events. J. Phys. Chem. B 109, 6688–6693 (2005).
Min, M. & Spencer, S. L. Spontaneously slow-cycling subpopulations of human cells originate from activation of stress-response pathways. PLoS Biol. 17, e3000178 (2019).
Yao, G., Lee, T. J., Mori, S., Nevins, J. R. & You, L. A bistable Rb-E2F switch underlies the restriction point. Nat. Cell Biol. 10, 476–482 (2008).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Taube, J. H. et al. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc. Natl. Acad. Sci. USA 107, 15449–15454 (2010).
Li, T., Shi, J., Wu, Y. & Zhou, P. On the mathematics of RNA velocity I: theoretical. Anal. CSIAM Trans. Appl. Math. 2, 1–55 (2021).
Chen, Y. et al. GraphVelo allows for accurate inference of multimodal velocities and molecular mechanisms for single cells. Nat. Commun. 16, 7831 (2025).
Bolhuis, P. G., Chandler, D., Dellago, C. & Geissler, P. L. Transition path sampling: throwing ropes over rough mountain passes, in the dark. Ann. Rev. Phys. Chem. 53, 291–318 (2002).
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Kramer, B. A., Del Castillo, J. S., Pelkmans, L. & Gut, G. Iterative indirect immunofluorescence imaging (4i) on adherent cells and tissue sections. Bio Protoc. 13, e4712 (2023).
Stringer, C. & Pachitariu, M. Cellpose3: one-click image restoration for improved cellular segmentation. Nat. Methods 22, 592–599 (2025).
Coifman, R. R. & Lafon, S. Diffusion maps. Appl. Comput. Harmon. Anal. 21, 5–30 (2006).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Acknowledgements
The study was supported by NIDDK R01 DK119232 (J.X.), R56 DK (J.X.), NIGMS R01 (J.X.), NSF 2325149 (J.X.), NSF 2205148 (J.X.) and the pilot award from the University of Pittsburgh Hillman Cancer Center (to J.X.). We would like to thank Weikang Wang for the helpful discussions and Kazuhide Watanabe for providing the data.
Author information
Authors and Affiliations
Contributions
Author contributions were determined using the CRediT model: conceptualization: S.H., J.X. Formal analysis: S.H., G.Yu, Z.Z., and K.N. Funding acquisition: J.X. Investigation: S.H., Y.L., G.Yu, Z.Z., and K.N. Methodology: S.H., Y.L., G.Yu, Z.Z., J.Z., Y.Z., and J.X. Project administration: J.X. Supervision: J.X. Visualization: S.H., G.Yu, Z.Z. Writing–original draft: S.H., J.X. Writing–review and editing: S.H., Y.L., G.Yu, Z.Z., A.G., G.Yao, J.X.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Sara Formichetti and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary handling editor: Giulia Bertolin.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, S., Lu, Y., Yu, G. et al. Single cell snapshot analyses under proper representation reveal that epithelial-mesenchymal transition couples at G1 and G2/M. Commun Biol 9, 212 (2026). https://doi.org/10.1038/s42003-025-09487-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09487-6