Abstract
Excessive activation of the NLRP3 inflammasome drives the pathogenesis of diverse inflammatory diseases. However, the clinical application of NLRP3 inflammasome inhibitors remains a significant challenge. Here, we screen a natural product library of 126 compounds and identify Nimbolide (NIM), a triterpenoid from Azadirachta indica, as a potent suppressor of IL-1β secretion. Cellular studies reveal that NIM dose-dependently suppresses NLRP3 inflammasome activation, thereby the blocking Caspase-1 cleavage, IL-1β release, and pyroptosis in macrophages. Importantly, NIM exhibits high selectivity for NLRP3 inflammasome, showing no significant inhibition of non-NLRP3 inflammasomes. Mechanistically, NIM exerts dual effects by suppressing both NF-κB-dependent priming and NLRP3 inflammasome assembly. Molecular investigations reveal that NIM directly targets the Lys565 within the NLRP3 NACHT domain, thereby hindering inflammasome assembly. Using male C57BL/6 and Nlrp3-knockout mice, we demonstrate that NIM administration effectively alleviates inflammation and pathological damage in models of LPS-induced acute respiratory distress syndrome (ARDS) and DSS-induced ulcerative colitis. Collectively, our findings highlight NIM as a natural inhibitor that targets both the priming and assembly phases of NLRP3 inflammasome activation, offering a dual-modulatory strategy for treating NLRP3-driven inflammatory disorder.

Similar content being viewed by others
Introduction
Inflammation serves as a pivotal driver of pathogenic processes in many human diseases, for which there are no disease-modifying drugs1. The NLRP3 inflammasome, a key protein molecular machine of the innate immune system, represents a multi-protein complex critical for inflammatory responses2. Composed of the sensor protein NLRP3, the adaptor protein ASC, and the effector protein pro-Caspase-1, the NLRP3 inflammasome assembles in response to diverse stimuli, including damage-associated molecular patterns (DAMPs, like uric acid) and pathogen-associated molecular patterns (PAMPs, like nigericin)3,4,5. Active Caspase-1 then drives maturation of IL-1β and IL-18, while also initiating pyroptosis, a lytic form of programmed cell death characterized by plasma membrane rupture and release of pro-inflammatory DAMPs6. Dysregulated NLRP3 inflammasome activity has been implicated in a spectrum of human diseases, from autoimmune disorders like rheumatoid arthritis, and autoinflammatory disease like cryopyrin-associated periodic syndromes, to metabolic conditions such as obesity-related cardiomyopathy7 and type 2 diabetes8, as well as neurodegenerative disorders including Alzheimer’s disease9. The capacity of NLRP3 to integrate diverse pathological cues positions it as a central hub in inflammation-driven pathogenesis10. Thus, intervening in the NLRP3 inflammasome will emerge as a pivotal approach for the treatment of inflammatory diseases.
In recent decades, pharmacological inhibition of the NLRP3 inflammasome has emerged as a promising therapeutic strategy for inflammatory diseases, garnering increasing attention due to its critical role in bridging innate immunity and pathological inflammation11, such as diabetes12, osteoarthritis13, and Alzheimer’s disease14. The sulfonylurea inhibitor MCC950 represents a landmark inhibitor that potently blocks NLRP3 by targeting the Walker B motif in the NACHT domain15,16. However, its clinical progression was limited by unfavorable toxicokinetic properties and potential hepatotoxicity6. Similarly, while other direct inhibitors such as CY-09 and OLT1177 have shown promise, challenges regarding off-target effects and bioavailability remain17,18. However, no such drugs have yet been approved by the FDA for clinical use. Thus, there is an urgent need to identify novel molecular scaffolds with distinct mechanisms of action, like ZAP-180013, to overcome these barriers19.
Natural products and their structural analogues have historically made a major contribution to pharmacotherapy for diseases20. Nimbolide (NIM), a triterpenoid derived from the leaves and flowers of Azadirachta indica, has demonstrated efficacy in treating cancer and inflammatory diseases in recent years. Zhang et al. exhibited the protective effect of NIM against obesity in rats via alteration of Nrf2/HO-1 pathway21. Gu and his colleagues found that NIM against N-methyl-N-nitrosourea induced gastric cancer via alteration of apoptosis and NF-κB signaling pathway22. NIM was also characterized as a potential candidate for targeting the toll-like receptors pathway in rheumatoid arthritis23. Although NIM exhibits broad therapeutic activities, no reports have described its anti-inflammatory effects on the NLRP3 inflammasome.
In this study, NIM was found to potently inhibit the excessive activation of NLRP3 inflammasome, thereby blocking pyroptosis in macrophages. We observed that NIM targets NLRP3 Lys565, blocking the interaction between NLRP3 and ASC, thereby impeding the assembly of the inflammasome. In animal models, NIM significantly alleviated Lipopolysaccharides (LPS)-induced acute respiratory distress syndrome (ARDS) and mitigated DSS-induced acute ulcerative colitis, with its therapeutic effect being abolished in NLRP3 knockout (Nlrp3KO) mice. Collectively, this study identifies NIM as a potent natural agent for ameliorating inflammatory diseases through targeting the NLRP3 inflammasome.
Results
Nimbolide inhibits NLRP3 inflammasome activation and pyroptosis in macrophages
With the goal of reducing the release of pro-inflammatory cytokines, we employed a screening system to identify the natural products with highly effective anti-inflammatory effect (Fig. 1A). We found that Nimbolide (over 80%) is the natural agent with the most potent inhibitory activity against IL-1β release among 126 candidates (Fig. 1B, listed in Supplementary Table 1). Subsequently, the optimal administration concentration of NIM was determined via CCK-8 assay. We then selected an administration concentration no higher than 5 μM to ensure cell viability remained above 85% (Fig. 1C). Subsequently, we systematically evaluated the efficiency of NIM to inhibit excessive activation of the NLRP3 inflammasome. The results showed that NIM effectively suppressed adenosine triphosphate (ATP)-stimulated Caspase-1 release and IL-1β secretion in bone marrow-derived macrophages (BMDMs) under 1, 2, and 5 μM concentrations, exhibiting potency comparable to that of the standard inhibitor MCC950 (Fig. 1D, quantification in Supplementary Fig. 1A; Fig. 1E). While the release of IL-6 remained unaffected by NIM treatment (Fig. 1F). Next, we investigated the effect of NIM on NLRP3 inflammasome-mediated pyroptosis. GSDMD, the effector protein of pyroptosis, forms pores in the cell membrane after Caspase-1-mediated cleavage, leading to cellular perforation and content leakage24. By detecting the formation of GSDMD cleavage products (p32), we found that NIM effectively suppressed the cleavage of GSDMD, comparably to the positive drug (Fig. 1G, quantification in Supplementary Fig. 1B). Concurrently, NIM reduced both the number of dead cells as measured by propidium iodide (PI) staining (Fig. 1H, I) and the release of lactate dehydrogenase (LDH) into the supernatant (Fig. 1J). Collectively, we demonstrate that NIM potently inhibits the activation of the NLRP3 inflammasome and the subsequent NLRP3-mediated pyroptosis.
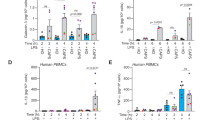
A Heatmap representing the screening of 126 natural products (10 μM) for IL-1β inhibition in culture supernatant of BMDMs. The arrow indicates NIM. The heatmap represents the mean values from these independent experiments. B Structure of NIM. C Cell viability of BMDMs treated with increasing concentrations of NIM for 24 h, assessed by CCK-8 assay. D Immunoblot analysis of cleaved Caspase-1 (p20) and IL-1β in culture supernatants (SN) and proteins in whole-cell lysates (Lys) of BMDMs. Cells were primed with LPS (500 ng/mL) for 3 h, treated with NIM (1, 2, or 5 μM) or the inhibitor MCC950 (5 μM) for 30 min, and stimulated with ATP (2.5 mM) for 30 min. E, F Levels of IL-1β (E) and IL-6 (F) in culture supernatants from BMDMs treated as in D, measured by ELISA. G Immunoblot analysis of full-length gasdermin D (GSDMD, p53) and the cleaved N-terminal fragment (p32) in lysates of BMDMs treated as in D. H Representative fluorescence images of BMDMs stained with Propidium Iodide (PI, red) and Hoechst (blue). Scale bar: 100 μm. The cells were stimulated with LPS (500 ng/mL) for 3 h, treated with NIM (5 μM) or MCC950 (5 μM) as previously described, then were challenged with ATP (2.5 mM) for 1 h. I Quantification of the percentage of PI-positive cells from (H). J Lactate dehydrogenase (LDH) release in supernatants of BMDMs. The cells were stimulated with LPS (500 ng/mL) for 3 h, treated with NIM (1, 2, and 5 μM) or MCC950 (5 μM) as previously described, then were challenged with ATP (2.5 mM) for 1 h. Data are representative of three biologically independent experiments and presented as the mean ± SEM. Statistical significance was determined by one-way ANOVA followed by Tukey’s multiple comparisons test (E, F, I, J). *P < 0.05, **P < 0.01, ***P < 0.001, significantly different; ns, not significant.
NIM is a specific inhibitor of the NLRP3 inflammasome
Next, we investigated NIM’s sensitivity to different stimuli activated NLRP3 in BMDMs. Extracellular ATP induces potassium efflux by directly activating the P2X7 receptor on the membrane, while nigericin acts as a potassium ionophore, promoting potassium exchange across mitochondrial membranes25. Thus, we explored the regulatory effect of NIM on nigericin-induced excessive activation of the NLRP3 inflammasome. Results showed that NIM similarly inhibited Caspase-1 release and IL-1β secretion to a comparable extent as the positive control (Fig. 2A, quantification in Supplementary Fig. 2A; Fig. 2B). We also tested alum potassium sulfate dodecahydrate (Alum) crystals, which activate NLRP3 by rupturing lysosomes and releasing cathepsin B26. Consistent with previous findings, NIM exhibited similar efficacy against this stimulus (Fig. 2C, quantification in Supplementary Fig. 2B; Fig. 2D). Furthermore, NIM effectively inhibited inflammasome activation triggered by imiquimod, which is known to activate potassium efflux-independent NLRP3 inflammasome19 (Supplementary Fig. 2C, D). We also evaluated the efficacy of NIM in human THP-1 cells and found that it attenuated the release of inflammatory cytokines induced by LPS plus nigericin (Fig. 2E, quantification in Supplementary Fig. 2E; Fig. 2F). Based on these results, we speculated that NIM exerts robust inhibitory effects on NLRP3 inflammasome activation induced by diverse stimuli. To determine whether NIM targets inflammasomes composed of different sensors, we challenged BMDMs with double-stranded DNA (dsDNA) and bacterial flagellin, agonists for the AIM2 inflammasome and NLRC4 inflammasome, respectively. NIM did not attenuate inflammation mediated by either the AIM2 (Supplementary Fig. 3A, B) or NLRC4 inflammasome (Supplementary Fig. 3C, D). Collectively, these findings establish NIM as a potent and specific inhibitor uniquely targeting the NLRP3 inflammasome.
A Immunoblot analysis of cleaved Caspase-1 (p20) and IL-1β in culture supernatants (SN) and proteins in whole-cell lysates (Lys) of BMDMs. Cells were primed with LPS (500 ng/mL) for 3 h, treated with NIM (1, 2, or 5 μM) or the inhibitor MCC950 (5 μM) for 30 min, and stimulated with Nigericin (10 μM) for 30 min. B Levels of IL-1β in SN from BMDMs treated as in A, measured by ELISA. C Immunoblot analysis of p20 and IL-1β in culture SN and proteins in Lys of BMDMs. Cells were primed with LPS (500 ng/mL) for 3 h, treated with NIM (1, 2, or 5 μM) or the inhibitor MCC950 (5 μM) for 30 min, and stimulated with Alum (300 μg/mL) for 2 h. D Levels of IL-1β in culture supernatants from BMDMs treated as in C, measured by ELISA. E Immunoblot analysis of Caspase-1 (p10) and IL-1β in culture SN and proteins in Lys of THP-1 cells. Cells were treated with NIM (1, 2, or 5 μM) or the inhibitor MCC950 (5 μM) for 30 min, challenged with LPS (500 ng/mL) for 3 h, and stimulated with Nigericin (10 μM) for 30 min. F Levels of human IL-1β in SN from BMDMs treated as in E, measured by ELISA. Data are representative of three biologically independent experiments and presented as the mean ± SEM. Statistical significance was determined by one-way ANOVA followed by Tukey’s multiple comparisons test (B, D, F). *P < 0.05, **P < 0.01, ***P < 0.001, significantly different; ns, not significant.
NIM suppressed assembly of NLRP3 inflammasome by interrupting NLRP3–ASC interaction
Previous reports have indicated that NIM exerts anti-inflammation effects by regulating the TLR4/NF-κB signaling pathway23. We therefore investigated whether NIM mediates its anti-NLRP3 inflammasome activity through NF-κB-related priming step. Based on LPS + ATP stimulation, we employed two administration protocols: long-term pretreatment with NIM before LPS exposure (to assess NF-κB signaling) and short-term post-treatment with NIM after LPS priming (to evaluate NLRP3 inflammasome activity). As shown in Fig. 3A (quantification in Supplementary Fig. 4A) and 3B, both protocols effectively inhibited the secretion of mature IL-1β and Caspase-1. However, immunoblot analysis revealed that short-term treatment did not reduce the protein levels of NLRP3 or pro-IL-1β (Fig. 3A, quantification in Supplementary Fig. 4B). We further evaluated the transcriptional and signaling events. RT-qPCR analysis demonstrated that the mRNA expression levels of Nlrp3 and Il1b remained unchanged following short-term treatment (Supplementary Fig. 4C, D). Furthermore, we examined the upstream signaling pathways essential for priming. Immunoblot analysis showed that short-term NIM treatment did not inhibit the LPS-induced phosphorylation of NF-κB (p65) or ERK1/2 (Supplementary Fig. 4E, F). Consistent with these findings, short-term treatment failed to suppress the secretion of TNFα, a cytokine dependent on NF-κB signaling but independent of NLRP3 activation (Supplementary Fig. 4G). Collectively, these data provide evidence that short-term NIM treatment acts downstream of the transcriptional priming step. In contrast, long-term pretreatment potently suppressed p65 and ERK1/2 phosphorylation (Supplementary Fig. 4H, I), reduced NLRP3 and pro-IL-1β protein levels (Fig. 3A, quantification in Supplementary Fig. 4B), and inhibited TNFα secretion (Supplementary Fig. 4G). Notably, while both treatments significantly reduced IL-1β secretion, long-term pretreatment exhibited a stronger inhibitory effect than short-term post-treatment. This enhanced efficacy is likely attributable to the dual inhibition of both the NF-κB-dependent priming step and the subsequent inflammasome activation. In contrast, short-term treatment inhibited only the activation step without affecting the transcription and translation of inflammatory precursors. Building on prior findings, our results show that NIM can exert dual inhibitory effects on NF-κB-related priming and the activation step of the NLRP3 inflammasome through distinct administration protocols.
A Immunoblot analysis of cleaved Caspase-1 (p20) and IL-1β in culture supernatants (SN) and proteins in whole-cell lysates (Lys) of BMDMs. Cells were subjected to two treatment protocols: short-term (primed with LPS for 3 h, then treated with NIM for 30 min before ATP stimulation) or long-term (treated with NIM for 0.5 h, then stimulated with LPS for 3 h before ATP stimulation). B Levels of IL-1β in SN from BMDMs treated as in (A), measured by ELISA. C Intracellular potassium concentration in LPS-primed BMDMs treated with NIM and stimulated with ATP. D–F Representative confocal microscopy images of BMDMs stimulated with LPS + ATP in the presence or absence of NIM. Cells were stained with Fluo-4 (D), MitoSOX (E), or MitoTracker (F). Scale bars: 10 μm. G Co-immunoprecipitation (Co-IP) analysis of the NLRP3–NEK7 interaction. Lysates from LPS + ATP-stimulated BMDMs treated with NIM were immunoprecipitated with an anti-NEK7 antibody. H Co-IP analysis of the NLRP3–ASC interaction. Lysates were immunoprecipitated with an anti-ASC antibody. I Immunoblot analysis of ASC oligomerization in the insoluble fraction (pellet) of BMDMs lysates. Data are representative of three biologically independent experiments and presented as the mean ± SEM. Statistical significance was determined by one-way ANOVA followed by Tukey’s multiple comparisons test (B, C). *P < 0.05, **P < 0.01, ***P < 0.001, significantly different; ns not significant.
Next, we sought to determine the mechanism by which NIM inhibits the NLRP3 inflammasome. As potassium efflux is a critical upstream event in NLRP3 inflammasome activation, we examined the effect of NIM on LPS + ATP-induced potassium efflux. Results showed that NIM at various concentrations had no impact on intracellular potassium levels, indicating that NIM does not inhibit the NLRP3 inflammasome by regulating potassium efflux (Fig. 3C). Given that cytosolic calcium mobilization has been linked to potassium dynamics, we used the Fluo-4 fluorescent probe to measure calcium changes and found no effect of NIM on calcium flux (Fig. 3D). Additionally, mitochondrial damage often accompanies NLRP3 inflammasome activation27, yet NIM had no significant impact on mtROS (Fig. 3E) or mitochondrial injury production (Fig. 3F). Collectively, these data indicate that NIM does not target key upstream events in NLRP3 inflammasome activation.
Subsequently, we hypothesized that NIM might regulate inflammasome assembly. While the interaction between NEK7 and NLRP3 is a critical step driving inflammasome formation after potassium efflux28, NIM did not alter their complex formation (Fig. 3G, quantification in Supplementary Fig. 5A). We therefore investigated other steps in the assembly process. Co-immunoprecipitation assays revealed that NIM significantly suppressed the interaction between NLRP3 and ASC (Fig. 3H, quantification in Supplementary Fig. 5B). Further cross-link analysis of downstream events showed that NIM inhibited ASC oligomerization, a key process in inflammasome assembly (Fig. 3I, quantification in Supplementary Fig. 5C). To directly visualize the inhibition of inflammasome assembly, we assessed ASC speck formation using confocal microscopy. Consistent with the cross-link analysis, NIM treatment significantly reduced the percentage of cells forming ASC specks upon LPS + ATP challenge, to an extent comparable to MCC950 (Supplementary Fig. 5D, E). These findings demonstrate that NIM disrupts the interaction between NLRP3 and ASC, thereby blocking the assembly of the NLRP3 inflammasome.
NIM directly targets NLRP3’s Lys565 site in the NLRP3 NACHT domain to inhibit oligomerization
To clarify the certain target of NIM, we performed the drug affinity responsive target stability (DARTS) assay which relies on differences in pronase-mediated proteolytic degradation. As shown in Fig. 4A (quantification in Supplementary Fig. 6A), NIM treatment protected NLRP3 from the pronase-mediated degradation in BMDMs, whereas it offered no protection for NEK7, ASC, or internal control. Similar results were observed in HEK293T cells transfected with overexpression plasmid (Fig. 4B, quantification in Supplementary Fig. 6B). These results indicated that the NIM directly binds to NLRP3 protein. NLRP3 comprises an N-terminal pyrin domain (PYD); a central NACHT domain; and a C-terminal LRR domain29. PYD is the effector domain for inflammasome formation; NACHT, a central ATPase domain, mediates protein oligomerization upon activation; and LRR domain acts as the signal sensor30. To determine the certain domain to which NIM binds, we constructed plasmids encoding individual each domain and performed DARTS assay in HEK293T. As shown in Fig. 4C (quantification in Supplementary Fig. 6C), NIM treatment specifically protected NACHT domain, indicating that NIM targets NACHT domain of NLRP3. Surface plasma resonance (SPR) assay with recombined human partial NLRP3 protein was performed and we found that NIM directly bound to NLRP3 with high affinity (Kd = 5.2 μM, Fig. 4D). Based on the function of NACHT domain in self-association, we investigated the effect of NIM on NLRP3 oligomerization using semi-denaturing detergent agarose gel electrophoresis (SDD–AGE). Interestingly, the oligomerization was blocked by NIM treatment (Fig. 4E, quantification in Supplementary Fig. 6D). These results demonstrate that NIM directly binds to the NACHT domain of NLRP3, thereby blocking its oligomerization. Next, to elucidating the binding mode, we performed molecular docking simulation. The results showed that NIM forms critical hydrogen bonds with lysine at position 565 (Lys565) in the NACHT domain of NLRP3, stabilizing NIM within the NLRP3 binding pocket (Fig. 4F). Subsequently, we constructed NLRP3 K565A mutant and performed a DARTS assay. Unlike the wild-type protein, the K565A mutant was not protected from pronase-mediated degradation by NIM (Fig. 4G, quantification in Supplementary Fig. 6E), demonstrating that the interaction is strictly dependent on the K565 residue. Consistently, Co-immunoprecipitation (co-IP) assays revealed that NIM failed to break the recruitment of ASC to the K565A mutant (Supplementary Fig. 7A), confirming that the K565 residue is critical for the inhibitory action of NIM. To further examinate the functional targeting of the NACHT domain, we examined the efficacy of NIM against specific auto-active NLRP3 disease variants with distinct activation mechanisms31. We reconstituted the inflammasome in HEK293T cells expressing D21H (PYD domain), F304C (NACHT domain), or E690K (LRR domain) mutants. As shown in Supplementary Fig. 7B–D, NIM effectively inhibited the E690K mutant. However, it failed to inhibit D21H and F304C, likely due to their distinct activation mechanisms. Recent studies have identified K565E as a strongly active mutant that drives inflammasome activation31. Crucially, Nimbolide failed to inhibit the hyperactivity of K565E, confirms that K565 is required for the inhibitory action of NIM (Supplementary Fig. 7E). These data corroborate our findings that NIM specifically targets the NACHT domain to prevent oligomerization. Given the NACHT domain’s ATPase activity, we subsequently investigated the effect of NIM on this activity. The results showed that NIM did not inhibited NLRP3’s ATPase activity (Supplementary Fig. 7F). Collectively, these findings demonstrate that NIM directly targets the Lys565 site within NACHT domain, inhibiting the oligomerization of NLRP3, thereby blocking the assembly of the NLRP3 inflammasome.
A DARTS assay performed in BMDMs lysates. Lysates were incubated with NIM or vehicle followed by pronase digestion. B DARTS assay in HEK293T cells transfected with plasmids encoding Flag-NLRP3, Flag-NEK7, or Flag-ASC. C Domain mapping using DARTS in HEK293T cells expressing Flag-tagged LRR, NACHT, or PYD domains of NLRP3. D SPR analysis of the binding affinity between NIM and recombinant human NLRP3 protein. E Analysis of NLRP3 oligomerization by SDD-AGE and total NLRP3 expression by SDS-PAGE in LPS + ATP-stimulated BMDMs treated with NIM. F Molecular docking simulation of NIM (orange, stick) binding to the NLRP3 NACHT domain (cartoon, PDB ID: 6NPY). The enlarged view illustrates the predicted hydrogen bond (yellow dashed line) between NIM and the Lys565 residue (purple sticks). G DARTS assay in HEK293T cells transfected with K565A mutant. Data are representative of three biologically independent experiments.
NIM alleviated LPS-induced ARDS in mice model via suppressing NLRP3 inflammasome
Numerous studies have demonstrated the critical involvement of the NLRP3 inflammasome in sepsis and its complications, with significantly elevated NLRP3 protein levels observed in the blood and immune cells of patients with ARDS32. We therefore evaluated the therapeutic efficacy of NIM in a murine model of ARDS. After three days of pre-treatment with NIM injections, septic ARDS was induced via intraperitoneal LPS administration, and tissues were harvested 12 h later (Fig. 5A). Notably, the administration of NIM at this dose was well-tolerated, with no significant body weight loss or hepatotoxicity observed during the treatment period (Supplementary Fig. 8A–D). Pathological analysis revealed that NIM treatment in wild-type mice significantly ameliorated lung tissue pathology, including reduced cellular infiltration and alveolar collapse (Fig. 5B, C). Protein concentration analysis of bronchoalveolar lavage fluid (BALF) showed that NIM attenuated LPS-induced increases in BALF protein content, indicating improved blood-air barrier integrity (Fig. 5D). The lung dry-to-wet weight ratio, a marker of pulmonary edema, was also improved by NIM (Fig. 5E). Cell counting confirmed reduced cellular infiltration in lung tissues (Fig. 5F), and immunohistochemical staining revealed decreased accumulation of F4/80+ macrophages (Fig. 5G) and Ly6G+ neutrophils (Fig. 5H) after NIM treatment, demonstrating the suppression of the inflammatory. To assess NIM’s regulation of the NLRP3 inflammasome during septic ARDS, we measured IL-1β levels in lung tissues and serum, finding significant suppression by NIM (Fig. 5I, J), which proved that NIM alleviates septic ARDS primarily through NLRP3 inflammasome inhibition.
A Schematic diagram of the experimental protocol. 10–week–old Wild-type (WT) and Nlrp3-deficient (Nlrp3KO) mice were pretreated with NIM (10 mg/kg, i.p.) or vehicle for three consecutive days, followed by an intraperitoneal injection of LPS (10 mg/kg) to induce sepsis-associated ARDS. Tissues were collected 12 h post-challenge. Panel (A) was created with BioRender.com. B Representative H&E staining of lung tissue sections. Scale bars: 100 μm. C Lung injury scores based on histological analysis of B. The legend in (C) applies to (D–F, I, J). D–F Assessment of pulmonary barrier function and inflammation, measured by total protein concentration in BALF (D), lung wet/dry weight ratio (E), and total cell counts in BALF (F). G, H Representative immunohistochemical staining of lung tissues for F4/80 (G) and Ly6G (H). Scale bars: 100 μm. I, J Concentrations of IL-1β in serum (I) and lung tissue (J) measured by ELISA. Data are presented as mean ± SEM (n = 6 biologically independent mice per group). Statistical significance was determined by one-way ANOVA followed by Tukey’s multiple comparisons test (C–F, I, J). *P < 0.05, **P < 0.01, ***P < 0.001, significantly different; ns not significant.
To further validate the broad therapeutic potential of NIM against direct pulmonary damage, we employed an LPS intratracheal instillation model (Supplementary Fig. 9A). Histological analysis showed that pretreatment with NIM or the positive control MCC950 significantly alleviated lung tissue damage (Supplementary Fig. 9B, C). Crucially, NIM preserved the blood-air barrier and suppressed inflammation, as evidenced by reductions of protein concentration in BALF, lung wet/dry ratio, and inflammatory cell infiltration (Supplementary Fig. 9D–F). Moreover, NIM effectively suppressed the secretion of IL-1β and prevented the activity of Caspase-1 in lung tissues, with efficacy comparable to MCC950. (Supplementary Fig. 9G, H).
Validation experiments in Nlrp3KO mice showed that genetic deletion of NLRP3 attenuated ARDS progression, and NIM provided no additional benefit in these mice (Fig. 5A–J), indicating that its therapeutic effects are dependent on NLRP3 inhibition. Collectively, these findings establish that NIM mitigates ARDS by inhibiting the NLRP3 inflammasome.
NIM ameliorates NLRP3-mediated ulcerative colitis in an NLRP3-dependent manner
In the pathogenesis of ulcerative colitis, the NLRP3 inflammasome also plays a critical mediating role33,34. We established a mouse model of ulcerative colitis by administering 2.5% DSS in drinking water. After initial body weight measurement and recording, mice were provided with 2.5% DSS in their drinking water for six consecutive days, followed by regular drinking water until day 10 (Fig. 6A). Throughout this period, daily body weights were recorded, and mice received two different concentrations of NIM via oral gavage (Fig. 6A). Calculation of weight change percentage showed that DSS administration caused about 25% body weight loss in wild-type mice, whereas NIM treatment mitigated this decline to less than 20% (Fig. 6B). Assessment of the disease activity index (DAI) revealed that NIM significantly attenuated disease progression, improving parameters including weight loss, stool consistency, and intestinal bleeding (Fig. 6C). On day 10, colon tissues were harvested, and measurement of colon length demonstrated that ulcerative colitis-induced shortening was markedly reversed by NIM treatment (Fig. 6D). Pathological analysis further showed that NIM reduced inflammatory cell infiltration and intestinal mucosal damage in colon tissues (Fig. 6E, F). Detection of NLRP3-mediated IL-1β secretion confirmed that NIM effectively decreased IL-1β levels in colon tissues (Fig. 6G). Consistent with prior findings, experiments in Nlrp3KO mice showed that NIM did not enhance the disease-improving effects of genetic NLRP3 deletion, as the protective benefits of NIM were abrogated in these models (Fig. 6A–G). Collectively, these results demonstrate that NIM significantly alleviates NLRP3-mediated ulcerative colitis.
A Schematic diagram of the experimental protocol. 10-week-old male WT and Nlrp3KO mice were administered 2.5% DSS in drinking water for 6 days, followed by regular water for 4 days. NIM (2.5 or 5 mg/kg) or vehicle was administered daily via oral gavage. Panel A was created with BioRender.com. B, C Disease progression monitored daily, shown as the percentage of body weight change (B) and Disease Activity Index (DAI) (C) over the 10-day period. D Colon lengths were measured after harvested on Day 10. The legend in (D) applies to panels (F, G). E Representative H&E staining of colon tissue sections. Scale bar: 100 μm. F Histological injury scores based on (E). G Levels of IL-1β in colon tissue lysates measured by ELISA. Data are presented as mean ± SEM (n = 6 biologically independent mice per group). Statistical significance was determined by one-way ANOVA followed by Tukey’s multiple comparisons test (B–D, F, G). *P < 0.05, **P < 0.01, ***P < 0.001, significantly different; ns, not significant.
Discussion
In this study, we reported the identification of NIM as a NLRP3-targeted inhibitor, with therapeutic potential for treating ARDS and ulcerative colitis. Our findings demonstrated that NIM exhibited broad and specific inhibitory activity against the NLRP3 inflammasome activation in macrophages, significantly reducing the secretion of IL-1β and Caspase-1. Mechanistically, NIM potently inhibited the interaction between NLRP3 and ASC during the assembly of the inflammasome, reducing ASC oligomerization and thereby alleviating pyroptosis. At the molecular level, we identified NLRP3 as the direct target of NIM. NIM formed a stable hydrogen bond with Lys565 in the NACHT domain of NLRP3, which disrupts NLRP3 oligomerization. In animal models of human diseases, NIM effectively mitigated the progression of ARDS and ulcerative colitis, effects that were completely abrogated when NLRP3 was genetically eliminated. Collectively, our study establishes NIM as a promising NLRP3-targeted inhibitor for inflammatory diseases and provides a novel molecular scaffold for developing next-generation NLRP3 inflammasome therapeutics.
Natural products serve as critical sources for drug design and development, offering both pharmacological efficacy and biological safety. Many of which have already entered clinical use, particularly in the field of critical care medicine, such as ambroxol and atropine35. Our laboratory has previously discovered numerous natural-source drugs with substantial potential for clinical application. 18β-glycyrrhetinic acid was identified as a potent suppressor of hepatic Angiotensinogen to suppressed sepsis induced myocardial dysfunction (SIMD)36. Emodin, a compound isolated from rhubarb, could be utilized as a potential Sirt3 modulator to treat SIMD37. During the drug screening process, we also identified potential candidate compounds, such as Clinodiside A and Eurycomanone, which inhibited IL-1β with an efficiency greater than 70% but less than 80%. While Eurycomanone is known for its anti-tumor and pro-apoptotic effects38,39, the pharmacological actions of Clinodiside A remain unexplored. Detailed activity testing, mechanistic investigation, and drug development targeting these two compounds constitute potential directions for further studies. In this study, we report NIM as a targeted inhibitor of NLRP3 for the treatment of ARDS and ulcerative colitis. Unlike other well-known natural or synthetic NLRP3 inhibitors, NIM as a structurally complex triterpenoid provides novel insights into the structural understanding40. Structurally, while well-known agents like MCC950 rely on the sulfonylurea, NIM possesses a complex triterpenoid scaffold. This unique chemical architecture provides novel structural insights and allows NIM to target K565 residue within the NLRP3 NACHT domain, a new binding mode distinct from previously characterized inhibitors. Mechanistically, NIM blocked the oligomerization of NLRP3 rather than inhibiting ATPase activity. This structural and mechanistic difference, combined with its dual ability to suppress both NF-κB priming and inflammasome activation, positions the triterpenoid NIM as a promising lead candidate for ameliorating NLRP3-driven diseases. Thus, further structural optimization of NIM from a medicinal chemistry perspective represents a critical step for the clinical translation of NIM and its derivatives.
The therapeutic efficacy of NIM in preclinical models of ARDS and ulcerative colitis underscores the clinical relevance of targeting NLRP3. In ARDS, NIM attenuated lung injury by suppressing the activation of NLRP3 inflammasome, while in ulcerative colitis, it restored epithelial integrity and inhibited IL-1β production. A unique pharmacological feature identified in this study is NIM’s dual-modulatory activity, which directly disrupts the post-translational assembly of the NLRP3 inflammasome during short-term treatment and suppresses the NF-κB-dependent transcriptional priming step during long-term treatment. This profile offers a distinct advantage for managing acute inflammatory storms. Broad pharmacological inhibition of NF-κB has raised concerns regarding off-target systemic immunosuppression and impaired host defense. However, NIM does not only rely on upstream blockade. By binding to the NLRP3 NACHT domain, NIM ensures that the terminal effect of inflammation is suppressed, providing potent therapeutic efficacy without relying on the abolish of upstream immune signaling. This allows for potent therapeutic efficacy via precise downstream inhibition, instead of the reliance on broad upstream immune suppression. Thus, drug development based on NIM could provide novel therapeutic modalities for emergency and acute-care settings in emergency departments and ICUs, where urgent anti-inflammatory intervention is critical, particularly for diseases like septic ARDS and ulcerative colitis that require rapid-acting emergency treatments41.
Our findings elucidated the pharmacological mechanism by which NIM acts as an NLRP3 inhibitor, suppressing NLRP3 oligomerization, reducing excessive activation of the NLRP3 inflammasome, and alleviating inflammation. A structural comparison with positive inhibitors highlights the unique value of NIM. For instance, MCC950, a potent and specific sulfonylurea-based NLRP3 inhibitor, functions by stabilizing the inactive conformation of the NLRP3 NACHT domain, thereby preventing ATP hydrolysis16. NIM is similar to Oridonin, another diterpenoid that forms a covalent bond with Cysteine 279 on the NACHT domain42. They both exhibit an ATPase-independent mechanism of action distinct from that of MCC950. The identification of NIM expands the chemical space of natural NLRP3 inhibitors, offering a novel scaffold for drug design. While our data demonstrates the high selectivity of NIM, we must consider potential off-target effects before clinical application. However, the fact that natural products possess multiple targets in vivo necessitates more extensive proteomic analysis to rule out specific interactions43. This is particularly important given that other small molecule inhibitors have faced setbacks in clinical trials due to unforeseen liver toxicity or poor pharmacokinetics44. Despite these advantages, the clinical translation for triterpenoids like NIM is often failed by poor bioavailability and rapid metabolic clearance45. These data in the drug development process need further investigation. While our study confirms pharmacodynamic efficacy in acute models, the physicochemical properties of NIM require optimization. Addressing these pharmacokinetic liabilities, alongside absorption, distribution, metabolism, and excretion (ADME) profiling, remains the primary challenge for transforming this natural product into a viable therapeutic candidate. Additionally, crystal structure for direct interactions between natural products and NLRP3 protein are currently unavailable. Such structural biology insights will serve as the most valuable basis for guiding medicinal chemistry optimization of natural product-derived inhibitors in the future.
In this study, we report that NIM acts as a potent NLRP3 inhibitor, effectively suppressing Caspase-1 release and IL-1β secretion induced by excessive NLRP3 inflammasome activation in both human and murine macrophages, thereby inhibiting pyroptosis. Mechanistically, NIM directly targets Lys565 within NACHT domain of NLRP3, disrupts the interaction between NLRP3 and ASC, attenuates ASC oligomerization, and blocks inflammasome assembly. In vivo, we demonstrate that NIM administration potently inhibits NLRP3-mediated ulcerative colitis and ARDS, with therapeutic efficacy abrogated in Nlrp3-deficient mice. Collectively, our findings establish NIM as a promising candidate for development and clinical application as an NLRP3 inflammasome inhibitor.
Methods
Reagents
LPS (#L2880), ATP (#A3377), and Alum (#237086) were from Sigma (St. Louis, MO, USA). Nigericin (#N849347) was purchased from Macklin (Shanghai, China). Nimbolide (CAS No. 25990-37-8, Cat No. #T16324) was bought from Targetmol (Shanghai, China). Poly(dA:dT) (#tlrl-patn) and flagellin (#tlrl-flic) were supplied by InvivoGen (Toulouse, France). MitoSOX (#M36008) was obtained from Thermo Fisher Scientific (Carlsbad, CA, USA). MitoTracker (#C1049B), Protein A/G agarose (#P2012), and N-(methoxycarbonyl methyl)-6-methoxyquinolinium bromide (MQAE, #S1082) were obtained by Beyotime (Shanghai, China). Anti-NEK7 (#ab133514), anti-GSDMD (#ab209845), and anti-human Caspase-1 (#ab179515) antibodies were purchased from Abcam (Cambridge, UK). Anti-ASC (#67824S), anti-ERK1/2 (#9102S), anti-phospho-ERK1/2 (#9101S), and anti-phospho-NF-κB p65 (#3033S) were from Cell Signaling Technology (Danvers, MA, USA). Anti-NF-κB p65 (#T55034) were purchased from Abmart (Shanghai, China). Anti-NLRP3 (#AG-20B-0014) and anti-mouse Caspase-1 (#AG 20B-0042) antibodies were from Adipogen (San Diego, CA, USA). Anti-mouse IL-1β (#AF-401-NA) antibody was obtained from R&D Systems (Minneapolis, MN, USA). Anti-human IL-1β (#AF5103) antibody was purchased from Affinity Biosciences (Changzhou, China). Anti-Flag (#20543-1-AP), anti-HA (#66006-2-Ig), and anti-β-Actin (#66009-1-Ig) antibodies were bought from Proteintech (Rosemont, IL, USA).
Animal models
All experimental animal protocols were approved by the Hangzhou Medical College Animal Policy and Welfare Committee. We have complied with all relevant ethical regulations for animal use. Mice were housed in a specific pathogen-free (SPF) environment with strictly controlled housing conditions, including regulated temperature (23 ± 2 °C), humidity (55 ± 10%), and light cycle (lights on from 7:00 AM to 7:00 PM). Investigators were blinded to group allocation during data analysis. No animals were excluded from analysis.
For ARDS models, male C57BL/6 or Nlrp3KO mice (8–10 weeks old) were randomized into three groups (n = 6 biologically independent mice per group): normal, LPS, and LPS + NIM groups. For the treatment groups, NIM (10 mg/kg) or vehicle was administered intraperitoneally 3 consecutive days prior to LPS challenge prior to the LPS challenge. To induce systemic inflammation-associated lung injury, mice received an intraperitoneal injection of LPS (10 mg/kg). Control mice received vehicle alone. Mice were euthanized by carbon dioxide inhalation at 12 h post-LPS challenge. To induce direct pulmonary injury, mice were anesthetized by isoflurane, and the trachea was exposed. LPS (5 mg/kg) dissolved in 50 μL sterile saline was instilled intratracheally. Control mice underwent the same procedure with an equal volume of sterile saline. Mice were euthanized by carbon dioxide inhalation at 6 hours post-LPS challenge. At the experimental endpoints, BALF was collected to quantify total cell numbers, protein concentrations, and cytokine levels via enzyme-linked immunosorbent assay (ELISA). Lung tissues were harvested and used for the wet/dry weight ratio, histological examination, and cytokines analysis. For histological examination, lung tissues were sectioned at 5 μm thickness. Sections were stained with hematoxylin and eosin (H&E) to evaluate the degree of lung injury using light microscopy. The severity of injury was judged according to the following criteria: no injury = 0; injury to 25% of the field = 1; injury to 50% of the field = 2; injury to 75% of the field = 3; and diffuse injury = 4. Lung injury score was calculated as the sum of scores from different views of the lung tissue section of each mouse.
For ulcerative colitis models, male C57BL/6 or Nlrp3KO mice (8–10 weeks old) were randomized into four groups (n = 6 biologically independent mice per group): normal, DSS (2.5% w/v dextran sodium sulfate in drinking water for 6 days), DSS + NIM (2.5 and 5 mg/kg, oral gavage daily from day 1–10). Body weight, diarrhea, and rectal bleeding were monitored daily to calculate DAI. On day 10, mice were euthanized by carbon dioxide inhalation, and colons were harvested to measure length and processed for histology to score epithelial damage, crypt loss, and inflammatory cell infiltration. Colonic tissue lysates were analyzed for IL-1β via ELISA.
Cell culture and inflammasome activation
BMDMs were isolated from the femoral bone marrow of C57BL/6 mice aged 8 to 10 weeks. The cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM; #C11995500BT, Thermo Fisher Scientific), which was supplemented with 10% fetal bovine serum (FBS), 20% L929 mouse fibroblast-conditioned supernatant, 100 U/mL penicillin, and 100 μg/mL streptomycin. After a culture period of 7 days, the BMDMs were harvested and subsequently utilized for further experimental procedures. L929 mouse fibroblasts (catalog #GNM28), HEK-293T cells (catalog #GNHu17), and THP-1 cells (catalog #TCHu57) were procured from the Shanghai Institute of Biochemistry and Cell Biology. L929 mouse fibroblasts and HEK-293T cells were maintained in DMEM supplemented with 10% FBS and penicillin-streptomycin, while THP-1 cells were cultured in RPMI-1640 medium (#C11875500BT, Thermo Fisher Scientific) supplemented with 10% FBS and penicillin-streptomycin.
For inflammasome activation assays, BMDMs (1 × 106 cells/well) were seeded in culture plates and cultured overnight. For NLRP3 inflammasome activation, cells were primed with LPS (500 ng/mL) for 3 h. Following priming, the cells were incubated with NIM at different concentrations or MCC950 for 30 min. Subsequently, cells were stimulated with ATP (2.5 mM), Nigericin (10 μM), Imiquimod (10 μg/mL) for 30 min, or Alum (300 μg/mL) for 2 h. For NLRC4 and AIM2 inflammasome activation, cells were primed with LPS (500 ng/mL) for 3 h and treated with NIM for 30 min. Cells were then stimulated by transfecting flagellin (0.1 μg/mL) or poly(dA:dT) (2 μg/mL) using Lipofectamine 2000 for 2 h. Supernatants and cell lysates were collected immediately after stimulation for ELISA and immunoblotting analysis, respectively.
CCK-8 assay
BMDMs (5 × 104 cells/well) were seeded in 96-well plate and cultured overnight. The culture medium was replaced with medium containing gradient of concentrations of NIM for 24 h. The cell survival rate after NIM treated were detected using CCK-8 kit (#CA1210; Solarbio, Beijing, China) referring to the user manual from the producer.
LDH assay
BMDMs (1 × 105 cells/well) were stimulated with 500 ng/mL LPS for 3 h and treated with varying doses of NIM. Then BMDMs were stimulated with 2.5 mM ATP for 1 h. The LDH level was determined using an LDH releasing assay (#C0016, Beyotime) according to the manufacturer’s instructions.
PI staining
BMDMs (1 × 105 cells/well) were plated in a fluorescent petri dish and cultured overnight. The cells were stimulated with LPS (500 ng/mL) for 3 h, treated with NIM or MCC950, then were challenged with ATP (2.5 mM) for 1 h. The cell death level was determined using a Cell Death Detection Kit (#C1062M, Beyotime) according to the manufacturer’s instructions. After that, the cells were observed using a fluorescence microscope.
Immunoblotting and co-immunoprecipitation assay
Protein samples were extracted from cells or tissues using lysis buffer supplemented with protease and phosphatase inhibitors. The protein concentration was determined using Bradford assay. Equal amounts of protein were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) on 8–12% gels and then transferred onto polyvinylidene fluoride (PVDF) membranes. The membranes were blocked with 5% bovine serum albumin (BSA) in Tris-buffered saline containing 0.05% Tween-20 (TBST) for 1 h at room temperature. Subsequently, the membranes were incubated overnight at 4 °C with primary antibodies diluted in blocking buffer (1:1000). After washing three times with TBST, the membranes were incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature. Protein bands were visualized using an enhanced chemiluminescence detection system, and images were captured using a chemiluminescence imaging system. The relative band intensities were quantified using ImageJ software (NIH, Bethesda, MD, USA) and normalized to the loading control.
To analyze the oligomerization of NLRP3, SDD-AGE was performed. Briefly, BMDMs were lysed in non-denaturing lysis buffer (0.5% Triton X-100, 50 mM Tris-HCl pH 7.4, 150 mM NaCl, 10% glycerol, and 0.1% SDS) supplemented with protease inhibitors. Lysates were resuspended in sample buffer (0.5× TBE, 0.1% SDS, 10% glycerol, and 0.025% Bromophenol Blue) and incubated at room temperature for 5 min. Samples were resolved on a vertical 1.5% agarose gel in TBE buffer containing 0.1% SDS at 80 V for 60 min at 4 °C. Proteins were transferred onto PVDF membranes for immunoblotting with anti-NLRP3 antibody.
In the co-immunoprecipitation assay, cell lysates were incubated with decoy antibodies overnight at 4 °C, and then were immunoprecipitated with protein G agarose beads at 4 °C for 4 h. After that, the immunoprecipitation samples were subjected to immunoblotting to detect the coprecipitated proteins. Total lysates were also subjected to Western blot analysis as an input control.
ELISA
BMDMs and THP-1 cells (1 × 106 cells/well) were stimulated and treated with varying doses of NIM. After incubation, the medium was collected and used for ELISA. Lung and colon tissues were lysed in tissue lysis buffer, followed by protein quantification. According to the manufacturer’s instructions, the level of mouse IL-6 (#88-7064-88, Thermo Fisher Scientific), IL-1β (#88-7013-77, Thermo Fisher Scientific), TNF-α (#88-7324-76, Thermo Fisher Scientific) or human IL-1β (#88-7064-86, Thermo Fisher Scientific) in supernatants or lysis were measured using ELISA kits.
ASC oligomerization assay
BMDMs (2 × 106 cells/well) have undergone inflammasome stimulation and drug treatment, they are lysed using lysis buffer containing 0.5% Triton X-100 and protease inhibitors. After lysis, the supernatant and precipitate are separated by centrifugation. The precipitation is resuspended in a solution of 2 mM suberic acid bis (3-sulfo-N-hydroxysuccinimide ester sodium salt (BS3, CAS No. 127634-19-9, Cat No. #S855494; Macklin, Shanghai, China), and incubated at room temperature for 30 min for protein cross-linking. After the cross-linking reaction is completed, the precipitate is collected and subjected to subsequent analysis by Western blotting.
Potassium concentration determination
BMDMs (2 × 106 cells/well) were stimulated and treated with different doses of NIM. Following three-time washes with 0.9% normal saline, the cells were lysed using ultrapure water for 20 min at 37 °C. After sample collection and centrifugation, the potassium concentrations in the supernatants were measured using a Cobas-c-311 Automatic Biochemistry Analyzer (Roche, Germany).
Fluorescence staining
BMDMs (1 × 105 cells/well) were plated in a fluorescent petri dish and cultured overnight. The cells were then stimulated and treated with varying concentrations of NIM as described earlier. ATP and either 40 nM MitoTracker, 5 μM Fluo-4 or 2.5 µM MitoSOX were added together. Then the cells were washed three times with PBS and fixed with 4% paraformaldehyde. Finally, the cells were counterstained with DAPI and observed using a fluorescence microscope.
DARTS analysis
Cells were lysed with lysis buffer and then aliquoted into tubes after protein quantification. The cell lysates were incubated with NIM or DMSO at 4 °C overnight, followed by pronase (#10165921001, Roche; 50 ng/μg protein) at room temperature for different time: protein ≥ 70 kDa, 30 s; 40 kDa ≤ protein < 70 kDa, 60 s; 10 kDa ≤ protein < 40 kDa, 120 s; protein < 10 kDa, 180 s. The reactions were immediately terminated by addition of SDS loading buffer and boiling at 100 °C. The samples were analyzed by immunoblotting.
SPR assay
The binding kinetics between the analyte and ligand were analyzed using Biacore X100 (Cytiva, Uppsala, Sweden) instrument. Recombinant Human NLRP3 protein was coupled to a CM5 chip (#BR100399, Cytiva). For kinetic analysis, different doses of NIM were diluted in HBS-EP+ buffer (#BR100826, Cytiva) and injected over the protein-immobilized surface at various concentrations (0.24–62.5 µM). The association and dissociation phases were monitored in real-time, and the sensor gram data were collected. The binding kinetics were calculated using Biacore evaluation software.
Molecular docking
The human NLRP3 crystal structure (PDB ID: 6NPY) was retrieved from the Protein Data Bank. AutoDock Tools were applied to prepare and parametrize the receptor protein and ligands. The docking grid documents were generated by AutoGrid of sitemap, and AutoDockFR 1.0 was used for docking simulation. The optimal pose was selected to analyze interaction. Finally, the protein-ligand interaction figure was generated by PyMOL.
Immunohistochemistry analysis
The lung tissues from mice were cut into 4 μm sections. After incubating 3% hydrogen peroxide at room temperature for 20 minutes to inactivate endogenous peroxidases, the tissues were incubated overnight with primary antibodies at 4 °C. The tissue sections were washed three times with TBST and then incubated with an HRP-IgG secondary antibody at 37 °C for 15 min. After DAB staining using DAB Substrate Kit (#DA1010, Solarbio), all slides were stained with hematoxylin and then observed using a microscope.
RNA Extraction and RT-qPCR
Total RNA was extracted from BMDMs using the RNA Extraction Kit (#AG21024; Accurate Biotech, Changsha, China) according to the manufacturer’s instructions. The concentration and purity of the isolated RNA were determined using a NanoDrop 2000 spectrophotometer (ThermoFisher Scientific). Subsequently, 1 μg of total RNA was reverse-transcribed into cDNA using the PrimeScript RT reagent Kit (#RR037A; Takara, Japan). Quantitative real-time PCR was performed using SYBR premix Ex Taq (#639676, Takara) on CFX Connect system (Bio-Rad). Primer sequences were included in Supplementary Table 2. The relative mRNA expression levels of target genes were calculated using the 2-ΔΔCT method and the data were normalized relative to the expression levels of the control group.
Reconstituted NLRP3 inflammasome in HEK293T cells
HEK293T cells were transfected with plasmids encoding NLRP3 inflammasome components or NLRP3 mutants. HEK293T cells (2 × 106 cells/well) were seeded in culture plates and cultured overnight. The cells were co-transfected with plasmids expressing human NLRP3 inflammasome components, including WT GFP-NLRP3 or mutated GFP-NLRP3 (400 ng), His-ASC (100 ng), HA-Caspase-1 (400 ng), and FLAG-IL-1β (200 ng) using Lipofectamine 2000. The effect of NIM treatment on activation of reconstituted NLRP3 inflammasome was examined 24 h after transfection.
Statistics and Reproducibility
All statistical analyses were performed using Prism 8 software (GraphPad Prism, San Diego, CA, USA). For comparison between two groups, data were analyzed by the unpaired two-tailed Student’s t-test. For more than two groups, data were evaluated by one-way ANOVA, followed by Tukey’s multiple comparisons test. P < 0.05 was considered significant. Details on the number of biological replicates and the sample sizes are provided in the figure legends. Randomization was used for animal grouping, and investigators were blinded to group allocation during data analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Coll, R. C. & Schroder, K. Inflammasome components as new therapeutic targets in inflammatory disease. Nat. Rev. Immunol. 25, 22–41 (2025).
Fu, J. & Wu, H. Structural Mechanisms of NLRP3 Inflammasome Assembly and Activation. Annu Rev. Immunol. 41, 301–316 (2023).
Huang, Y., Xu, W. & Zhou, R. NLRP3 inflammasome activation and cell death. Cell Mol. Immunol. 18, 2114–2127 (2021).
Yang, Y., Wang, H., Kouadir, M., Song, H. & Shi, F. Recent advances in the mechanisms of NLRP3 inflammasome activation and its inhibitors. Cell Death Dis. 10, 128 (2019).
Seok, J. K., Kang, H. C., Cho, Y. Y., Lee, H. S. & Lee, J. Y. Therapeutic regulation of the NLRP3 inflammasome in chronic inflammatory diseases. Arch. Pharm. Res. 44, 16–35 (2021).
Mangan, M. S. J. et al. Targeting the NLRP3 inflammasome in inflammatory diseases. Nat. Rev. Drug Discov. 17, 588–606 (2018).
Soták, M., Clark, M., Suur, B. E. & Börgeson, E. Inflammation and resolution in obesity. Nat. Rev. Endocrinol. 21, 45–61 (2025).
Deng, Y. et al. Obesity Enables NLRP3 Activation and Induces Myocardial Fibrosis via Hyperacetylation of HADHa. Diabetes 72, 1597–1608 (2023).
McManus, R. M. et al. NLRP3-mediated glutaminolysis controls microglial phagocytosis to promote Alzheimer’s disease progression. Immunity 58, 326–343 (2025).
Wang, L. & Hauenstein, A. V. The NLRP3 inflammasome: Mechanism of action, role in disease and therapies. Mol. Asp. Med 76, 100889 (2020).
Yan, L. -j et al. Hypocrellin A from an ethnic medicinal fungus protects against NLRP3-driven gout in mice by suppressing inflammasome activation. Acta Pharmacologica Sin. 46, 1016–1029 (2025).
Ward, R. et al. NLRP3 inflammasome inhibition with MCC950 improves diabetes-mediated cognitive impairment and vasoneuronal remodeling after ischemia. Pharm. Res. 142, 237–250 (2019).
Jia, L. et al. Ginkgolide C inhibits ROS-mediated activation of NLRP3 inflammasome in chondrocytes to ameliorate osteoarthritis. J. Ethnopharmacol. 325, 117887 (2024).
Shan, X. et al. Kai-Xin-San ameliorates Alzheimer’s disease-related neuropathology and cognitive impairment in APP/PS1 mice via the mitochondrial autophagy-NLRP3 inflammasome pathway. J. Ethnopharmacol. 329, 118145 (2024).
Coll, R. C. et al. A small-molecule inhibitor of the NLRP3 inflammasome for the treatment of inflammatory diseases. Nat. Med. 21, 248–255 (2015).
Tapia-Abellán, A. et al. MCC950 closes the active conformation of NLRP3 to an inactive state. Nat. Chem. Biol. 15, 560–564 (2019).
Jiang, H. et al. Identification of a selective and direct NLRP3 inhibitor to treat inflammatory disorders. J. Exp. Med 214, 3219–3238 (2017).
Marchetti, C. et al. OLT1177, a β-sulfonyl nitrile compound, safe in humans, inhibits the NLRP3 inflammasome and reverses the metabolic cost of inflammation. Proc. Natl. Acad. Sci. USA 115, E1530–E1539 (2018).
Kim, W. et al. A potent NLRP3 inhibitor effective against both MCC950-sensitive and -resistant inflammation. Cell Chem. Biol. 32, 1125–1139 (2025).
Atanasov, A. G., Zotchev, S. B., Dirsch, V. M. & Supuran, C. T. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 20, 200–216 (2021).
Zhang, L., Li, Y., Sun, D. & Bai, F. Protective Effect of Nimbolide against High Fat Diet-induced Obesity in Rats via Nrf2/HO-1 Pathway. J. Oleo Sci. 71, 709–720 (2022).
Gu, Y., Liu, B., Xia, X., Luo, C. & Ren, Y. Chemoprotective effect of nimbolide against N-methyl-N-nitrosourea induced gastric cancer via alteration of apoptosis and NF-κB signaling pathway. Acta Cir. Bras. 40, e402125 (2025).
Israr, M. et al. Nimbolide attenuates complete Freund’s adjuvant induced arthritis through expression regulation of toll-like receptors signaling pathway. Phytother. Res 37, 903–912 (2023).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Di Virgilio, F., Dal Ben, D., Sarti, A. C., Giuliani, A. L. & Falzoni, S. The P2X7 Receptor in Infection and Inflammation. Immunity 47, 15–31 (2017).
Hornung, V. et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat. Immunol. 9, 847–856 (2008).
Nakahira, K. et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 12, 222–230 (2011).
He, Y., Zeng, M. Y., Yang, D., Motro, B. & Núñez, G. NEK7 is an essential mediator of NLRP3 activation downstream of potassium efflux. Nature 530, 354–357 (2016).
Hochheiser, I. V. et al. Structure of the NLRP3 decamer bound to the cytokine release inhibitor CRID3. Nature 604, 184–189 (2022).
Xiao, L., Magupalli, V. G. & Wu, H. Cryo-EM structures of the active NLRP3 inflammasome disc. Nature 613, 595–600 (2023).
Feng, S. et al. Mechanisms of NLRP3 activation and inhibition elucidated by functional analysis of disease-associated variants. Nat. Immunol. 26, 511–523 (2025).
Gu, L. et al. NLRP3 promotes inflammatory signaling and IL-1β cleavage in acute lung injury caused by cell wall extract of Lactobacillus casei. Commun. Biol. 8, 20 (2025).
Xue, J. C. et al. Natural products modulate NLRP3 in ulcerative colitis. Front Pharm. 14, 1265825 (2023).
Zhou, J., Yang, Q., Wei, W., Huo, J. & Wang, W. Codonopsis pilosula polysaccharide alleviates ulcerative colitis by modulating gut microbiota and SCFA/GPR/NLRP3 pathway. J. Ethnopharmacol. 337, 118928 (2025).
Wang, Z. et al. Polypharmacology of ambroxol in the treatment of COVID-19. Biosci. Rep. 43, https://doi.org/10.1042/bsr20221927 (2023).
Shi, M. et al. Identification of 18β-glycyrrhetinic acid as an AGT inhibitor against LPS-induced myocardial dysfunction via high throughput screening. Biochem Pharm. 223, 116127 (2024).
Xu, Y. et al. Sirt3 is a novel target to treat sepsis induced myocardial dysfunction by acetylated modulation of critical enzymes within cardiac tricarboxylic acid cycle. Pharm. Res 159, 104887 (2020).
Xu, W. et al. Eurycomanone inhibits osteosarcoma growth and metastasis by suppressing GRP78 expression. J. Ethnopharmacol. 335, 118709 (2024).
Okba, M. M., Ezzat, M. I., Shehabeldine, A. M. & Ezzat, S. M. Eurycomanol and eurycomanone as potent inducers for cell-cycle arrest and apoptosis in small and large human lung cancer cell lines. Nat. Prod. Res. 37, 1856–1862 (2023).
Bosch, J. PPI inhibitor and stabilizer development in human diseases. Drug Discov. Today Technol. 24, 3–9 (2017).
Villar, J. et al. Dexamethasone treatment for the acute respiratory distress syndrome: a multicentre, randomised controlled trial. Lancet Respir. Med 8, 267–276 (2020).
He, H. et al. Oridonin is a covalent NLRP3 inhibitor with strong anti-inflammasome activity. Nat. Commun. 9, 2550 (2018).
Chen, X. et al. Target identification of natural medicine with chemical proteomics approach: probe synthesis, target fishing and protein identification. Signal Transduct. Target Ther. 5, 72 (2020).
Zahid, A., Li, B., Kombe, A. J. K., Jin, T. & Tao, J. Pharmacological Inhibitors of the NLRP3 Inflammasome. Front Immunol. 10, 2538 (2019).
Sharmila, A. et al. Nanoformulated Terpenoids in Cancer: A Review of Therapeutic Applications, Mechanisms, and Challenges. Cancers (Basel) 17, 3013 (2025).
Acknowledgements
This study was supported by the National Key Research and Development Program of China (2021YFC2501800 to Z.C.Z.), the National Natural Science Foundation of China (82272182, 82472177 to Z.C.Z.), the Zhejiang Provincial Natural Science Foundation of China (LHDMD22H020001 to Z.C.Z.), the Key Research and Development Program of Zhejiang Province (2024C3186 to Y.A.X.), and Major Project of National-Zhejiang Provincial Administration of Traditional Chinese Medicine (GZY-ZJ-KJ-24030 to Y.A.X.).
Author information
Authors and Affiliations
Contributions
H.X., Y.X., and Z.Z. conceptualized the study and designed the experiments. H.X., Y.L., and W.L. performed the experiments and methodology. H.X., Y.L., W.L., and S.S. curated and analyzed the data. H.X. wrote the original draft. Y.X. and Z.Z. reviewed and edited the manuscript. Y.X. and Z.Z. provided resources and supervision. Y.X. and Z.Z. acquired funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Radhwan Al-Zidan, Shouya Feng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Dr. Si Ming Man and Dr. Ophelia Bu. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, H., Lin, Y., Luo, W. et al. Nimbolide ameliorates ARDS and ulcerative colitis by disrupting NLRP3 inflammasome activation. Commun Biol 9, 246 (2026). https://doi.org/10.1038/s42003-026-09524-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09524-y