Abstract
Latin American populations exhibit high genetic and phenotypic diversity shaped by complex admixture histories, yet remain underrepresented in genomic research. Here, we analyze genome-wide data from 432 urban individuals across 13 regions of Peru, including 346 newly genotyped from the Peruvian Genome Project. We reveal fine-scale population structure and demographic patterns shaped by both ancient and recent events. Indigenous American ancestries in urban individuals trace back to ancient north-south interactions consistent with archaeological records, while admixture events occurring within the last 8–10 generations involved sources already admixed between distinct ancestral lineages. Identity-by-descent analyses reveal sustained gene flow in southern Peru, while effective population size trends highlight demographic stability in Lima over the past 25 generations. Sex-biased admixture patterns suggest Indigenous ancestry contribution preferentially mediated by females. These findings provide a comprehensive understanding of Peru’s genetic heritage, enhancing our understanding of human genetic diversity and historical demographic processes in Latin America.
Similar content being viewed by others
Introduction
Less than 0.4% of individuals in genome-wide association studies represent Latin American populations1. This underrepresentation is particularly concerning given the rich cultural and genetic diversity within these populations, which have been shaped by extensive continental admixture over the past 20 generations2,3,4,5,6,7,8 and high population structure derived from their indigenous sources9,10,11. In particular, Peru is a case of remarkable demographic and genetic complexity. Home to approximately 33 million inhabitants and spanning 1.28 million square kilometers—an area comparable to the combined landmasses of the United Kingdom, France, and the Iberian Peninsula—Peru is one of the ancient cradles of civilization in America with a largely Indigenous population, especially in the highlands of the Andes. Notably, Peruvians exhibit considerable Indigenous American genetic ancestry12,13, with the average individual sharing about 50% of their genome with Indigenous groups9,12,14.
The last 500 years of Peruvian history have been part of a process in American social history commonly known as mestizaje, a term scrutinized today from different perspectives, including population genetics15. Initiated during the European invasion, this process involved not only encounters among groups of different ancestries (Indigenous Americans, Europeans, Africans, and later East Asians), but also the imposition of a new social hierarchy whose consequences remain evident today16,17. Yet, despite centuries of mestizaje and profound demographic shifts, Indigenous population structure in Peru has remained deeply influenced by geography10,18. Genetic, archeological, and linguistic evidence suggested that Indigenous Peruvian groups were organized in a latitudinal pattern, with two interaction spheres representing the Northern and Southern mirroring Andean geography18,19. Despite the profound demographic changes brought about by colonialism, present-day Indigenous American groups still exhibit this North-South pattern10,20. However, it remains unknown whether Peruvians now living in urban areas follow the same pattern.
The European invasion and the arrival of enslaved Africans in the 1530 s profoundly shaped Peruvian culture and demography. It transformed its cultural landscape and triggered important socioeconomic changes (e.g., mines and agriculture), resulting in internal migration events21. Following nearly three centuries as a prominent Spanish Viceroyalty, Peru gained independence in 1821, giving rise to a supposedly independent Republic22. During the colonial period, tens of thousands of enslaved Africans were forcibly transported across the Atlantic to Panama and finally shipped along the Pacific coast in Peru to labor in mines and plantations. The Pacific Coast regions supported major economic activities related to sugar cane, cotton, and rice plantations, predominantly reliant on enslaved African labor23, later replaced by East Asian (Chinese and Japanese) workers after the abolition of slavery in 185424,25. In the Amazonian region, during the late 19th and 20th centuries, the rubber industry left profound negative effects on local indigenous communities and favored internal immigration from other Peruvian regions to Amazonian cities26. More broadly, internal migration and rapid urbanization have defined recent Peruvian demography. In 1961, only 50% of Peruvians lived in urban areas, compared to 79.3% today27,28. These voluntary migration flows were later compounded by forced displacements during the internal armed conflict (1980–1992), which drove tens of thousands from southern Andean regions such as Ayacucho, Apurímac, and Puno into Lima’s peripheries29. Nevertheless, despite centuries of admixture dynamics and recent migration, the evolutionary dynamics that shaped the present-day composition of Peruvians in urban areas remain largely unexplored.
In this study, we address two questions regarding the genetic ancestries in Peruvians living in urban areas: (i) How diverse is the genetic composition, and how are their ancestries distributed across Peruvian urban areas? and (ii) how did admixture processes between continental and regional groups in Peru unfold during and after colonial times? To address these questions, we sampled and genotyped 13 urban Peruvian populations as part of the Peruvian Genome Project, including 346 newly genotyped samples and 504 individuals (including 418 Native American and 86 urban individuals) from recent studies9,10 (Fig. 1 and Supplementary Data 1). A notable strength of our study is the inclusion of multiple urban populations, providing insights into ancestry composition, ancestry dynamics, and migration signatures over the past ten generations. This work represents an effort to build resources to address genomic health equity.
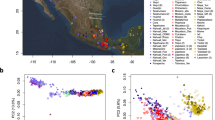
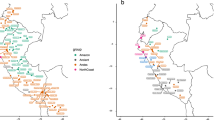
A Genome-wide ancestry proportions inferred by ADMIXTURE K = 4 in urban centers of different dimensions, from Moquegua city (around 96,000 inhabitants) to Trujillo (around 970,000 inhabitants) and Lima, the Peruvian capital (around 10 million inhabitants). Ancestry proportions are described in Supplementary Data 3. B Ancestry-specific PCA for Indigenous American ancestries. Indigenous American references include populations from the Peruvian Pacific Coast, the Andean, and Amazonian regions, represented by gray circles. C SOURCEFIND results for 14 admixed populations show a fine-scale inference of their ancestry proportions (See Supplementary Data 2 and 4). For Indigenous American contributions, each number represents a population: Amazonian (1: Ashaninkas; 2: Awajun; 3: Candoshi; 4: Lamas; 5: Shipibo), Andean (1: Chopccas; 2: Qeros, 3: Uros), and Coast (1: Moche; 2: Tallanes). Indigenous American groups were described in Borda et al. 10. All analyses are based on n = 470 unrelated individuals from PGP and PEL from 1000 Genomes Project (Arequipa = 26, Ayacucho = 35, El Carmen = 37, Cusco = 36, Huaraz = 36, Iquitos = 29, Lambayeque = 12, Lima = 28, PEL = 85, Moquegua = 30, Juliaca = 40, Tacna = 21, Tumbes = 33, Trujillo = 22).
Results
Full details about sampling are described elsewhere30. In this paper, we use urban areas to refer to a broad demographic spectrum, ranging from small cities such as Moquegua (~96,000 inhabitants) to Lima, a metropolis of nearly 10 million inhabitants. As part of the Peruvian Genome Project (PGP), we collected blood from 643 individuals who self-identified as mestizo, representing 13 urban populations across diverse biogeographical regions (Fig. 1). A subset of 86 samples was previously analyzed9. For this study, we generated genome-wide data for 346 individuals using the IlluminaOmni2.5 array (Supplementary Data 1). After QC, and merged with previous PGP data, the final dataset included 432 individuals from urban areas and 2,034,418 SNVs. All procedures were reviewed and approved by the Peruvian Research and Ethics Committee of the Instituto Nacional de Salud (authorization no. OI-003-11 and no. OI-087-13).
Fine-scale structure of indigenous and global ancestries in Peruvian genomes
We merged our dataset with the 1000 Genomes Project high coverage release31, the Human Genome Diversity Project high coverage32 datasets, and recently published Native American data from PGP9,10 (Supplementary Data 2). This later dataset includes Indigenous American populations from the Peruvian Pacific Coast, Andean, and Amazonian regions. Genetic clustering analysis33 shows urban individuals as predominantly three-way admixed with a higher proportion of Native American ancestries (>60% [except for Afro-Peruvians], Fig. 1A, Supplementary Fig. 1, Supplementary Data 3) than European and African ancestries. In general, European and African admixture in urban populations is larger than in Indigenous populations8,9 (Supplementary Fig. 1). Thirteen individuals showed East Asian ancestries (2-18%, Supplementary Fig. 1), likely as a result of the immigration of Chinese and Japanese individuals to Peru during the last 150 years, in part to replace the labor force in Coastal farms after the abolition of slavery in Peru in 185424,25.
Our geographically diverse sampling allows us to identify the heterogeneity of ancestries in urban individuals. ADMIXTURE (Supplementary Fig. 1) and principal component analyses (PCA, Supplementary Fig. 2) revealed a latitudinal differentiation of Indigenous American ancestry proportions. Indigenous ancestry clusters related to the Pacific Coast and Amazonian Natives are predominant in northern urban populations, and Andean components are predominant in southern urban populations.
By coupling local ancestry analyses with PCA34,35 (Ancestry-specific PCA: asPCA) restricted to Indigenous American segments, we showed that most urban groups clustered with Pacific Coast and Andean natives (Fig. 1B). Interestingly, Southern urban groups (i.e., Ayacucho, Arequipa, Moquegua, Tacna, Cusco, and Juliaca) clustered with the Andean Native groups and showed more homogeneity compared with the Northern urban populations and Lima. We tested this homogeneity by performing FST and Identity-by-Descent (IBD) sharing analyses. We showed lower differentiation (lower FST values, Supplementary Fig. 3) and high-IBD sharing between Southern urban populations compared to Northern (Supplementary Fig. 4).
Furthermore, by exploring the haplotype sharing information using the ChromoPainter-SOURCEFIND pipeline4,36, we observed a different composition of Andean ancestries in urban Peruvian populations (Fig. 1C, Supplementary Data 4). Northern populations have two Andean components compared to the South, which includes a third Andean component associated with present-day Aymara speakers’ ancestry (Uros population). Consistent with ADMIXTURE results, Pacific Coast and Amazonian ancestries are more common in urban populations from Northern Peru. SOURCEFIND further indicated that the genetic composition of urban individuals can be explained as a mosaic of different Indigenous American ancestries. As expected, the PEL (Peruvians in Lima, Peru) population from 1000 Genomes Project showed the same ancestry patterns observed in Lima individuals from our dataset. European ancestry is the second most common continental ancestry in urban individuals (Fig. 1A). During colonial times, the primary source of European ancestry was from Spain. However, after independence, the Peruvian government implemented policies that favored the migration of European communities to Peru37. Our asPCA for European ancestries showed admixed Peruvians with higher affinity to a cluster that includes individuals from Spain (IBS) and Italy (TSI) (Supplementary Fig. 5). SOURCEFIND results confirm the predominance of South European ancestry in admixed Peruvians (Fig. 1C, Supplementary Data 4).
There is no precise number of how many enslaved Africans were introduced in Peru throughout the tragic transatlantic slave trade. Still, they concentrated in the coastal region38. Our ADMIXTURE and PCA results (Fig. 1A, Supplementary Fig. 1) show most individuals with genome-wide African ancestries in the coast region, mainly in the North (Trujillo and Tumbes) and South Coast (El Carmen). During the initial decades of the colony, the majority of enslaved Africans originated from West Africa, specifically between the Senegal and Niger rivers38. However, by the late 1580 s, the Congo-Angola region (Central-Southern Africa) emerged as a primary source38. In the asPCA, we observed that African ancestry in Peruvians showed closer similarity to present-day West-Central African populations, although not a complete overlap. Notably, both ADMIXTURE (Supplementary Fig. 1) and SOURCEFIND consistently revealed that, in addition to the predominant West-Central African contribution, there is also a significant genetic affinity with East African (LWK) populations (Fig. 1C, Supplementary Data 4). These groups share greater genetic similarity with Central-Southern African populations, which historically represented one of the principal sources of enslaved Africans brought to the Americas.
It was after Peruvian independence that slavery abolition was achieved39. However, driven by economic interests, an alternative source of inexpensive labor was sought. This led to the influx of East Asian migrants into Peru, with approximately 100,000 Chinese individuals arriving in the 19th century24. The ADMIXTURE results revealed minimal individuals with East Asian-related ancestries (Fig. 1A). Our asPCA analysis of East Asian ancestries indicated that most individuals clustered with the Southern Chinese population, aligning with historical records. Notably, two individuals from Southern Peru (Moquegua) exhibited a higher affinity with Japanese individuals. At the population level, SOURCEFIND detected more East Asian ancestries (~6%) in Lambayeque, a city on the Peruvian Coast (Fig. 1C and Supplementary Data 2).
The signature of admixture shaped present-day admixed peruvians
Our second question investigated how continental and regional groups interacted in Peru during and after colonial times. To address this, we applied two approaches, GLOBETROTTER40 and HierarchyMix41. GLOBETROTTER examines LD decay observed on co-ancestry curves, providing the timing and mode of admixture events and ancestry composition for the hypothetical sources. In contrast, HierarchyMix analyzes the distribution of local ancestry segments. While GLOBETROTTER provides a subcontinental perspective on admixture events, HierarchyMix focuses on the interaction of continental ancestries.
We inferred two major admixture events (See “Methods”, Fig. 2, Supplementary Figs. 6–8, and Supplementary Data 5-6). GLOBETROTTER detected a single admixture event, but the admixed profile of its sources suggests an earlier admixture event (Fig. 2). Complementarily, HierarchyMix inferred admixture events per each pair of continental ancestries. Except for El Carmen, recent events for all Peruvian populations were inferred around 8-12 generations ago, or within the past 215–322.8 years (Fig. 2), assuming a generation time of 26.9 years42. Most populations showed these events as the admixture between a predominantly indigenous American source and a non-indigenous source (Fig. 2, Supplementary Figs. 6–8). Both methods inferred the non-indigenous source as admixed of European and African ancestries, except for Trujillo and El Carmen. HierarchyMix dated these earlier events of African-European admixture around 12-25 generations (Supplementary Figs. 6–8). Moreover, for Trujillo and El Carmen, the earlier admixture event involved Europeans and Indigenous Americans, followed by a recent African contribution (Fig. 2, Supplementary Figs. 6–8).
We inferred the admixture timing and mode in 14 urban Peruvian individuals using GLOBETROTTER. Estimated admixture dates for 14 urban Peruvian populations are shown as black points, with 95% confidence intervals indicated by black lines (See Supplementary Data 5). Bar plots represent the admixture sources inferred by GLOBETROTTER (See Supplementary Data 6), colored by donor populations as in the legend. Populations are ordered roughly from northern to southern Peru. All analyses are based on n = 470 unrelated individuals from PGP and PEL from 1000 Genomes Project (Arequipa = 26, Ayacucho = 35, El Carmen = 37, Cusco = 36, Huaraz = 36, Iquitos = 29, Lambayeque = 12, Lima = 28, PEL = 85, Moquegua = 30, Juliaca = 40, Tacna = 21, Tumbes = 33, Trujillo = 22).
The subcontinental composition of the Indigenous source inferred by GLOBETROTTER aligns with SOURCEFIND results (Fig. 1C), where Andean ancestries dominate in southern groups. This pattern reinforces that the present-day structure in the admixed population mirrors the Native American structure of the last 2000 years18. Furthermore, the genetic profiles of the inferred sources reflect combinations of multiple reference populations. This could indicate either that (i) the sources encompass greater genetic diversity than can be captured by any single Native group, requiring contributions from several populations to explain their composition, or (ii) the sources were themselves already admixed, supporting the presence of older admixture events and more complex interactions among Indigenous groups in the genetic history of Peruvians, as also suggested by Harris et al. 9.
HierarchyMix also inferred admixture events involving East Asian and Indigenous American ancestries. These events were dated several generations before the historical arrival of East Asian migrants in the 19th century, and considering the small contribution of less than 1% in most cases, this signal could be a spurious result from local ancestry inferences. However, our SOURCEFIND results support a significant East Asian contribution to the Peruvian population, with Lambayeque with the highest proportion (~6%). The predominantly Indigenous source for the recent admixture event inferred by GLOBETROTTER showed a small proportion of East Asian ancestries in northern populations (Fig. 2). However, it is unclear if this is confounding and represents the genetic composition of unsampled Indigenous American ancestries.
Reconstructing post-colonial demographic events
The GLOBETROTTER and HierarchyMix results broadly overview major admixture events involving continental ancestries around nine to 12 generations. Considering this, we assumed that DNA shared during the last eight generations could be associated with gene flow among urban groups. Our population structure analyses detected a higher genetic similarity among southern populations (Fig. 1B). Is this higher similarity the result of ancient gene flow during pre-colonial times, recent gene flow during post-colonial times, or both?
To address this question, we investigated patterns of gene flow through IBD segments shorter and longer than 18.75 cM. These two intervals explain the DNA sharing due to the recent common ancestor of two individuals before and after approximately eight generations ago43, respectively (See “Methods”). The short segment interval shows increasing IBD sharing from North to South (Fig. 3A). This mirrors the full IBD sharing patterns (Supplementary Fig. 4), underscoring the influence of ancient gene flow (pre-colonial times) and admixture (colonial times) on increasing the genetic similarity in Southern Peru. Importantly, this pattern has seemingly continued for the last eight generations (Fig. 3B), indicating that recent gene flow follows ancient trends. Furthermore, high IBD sharing between Trujillo and Afrodescendants from El Carmen potentially reflects shared African ancestry. In summary, ancient and recent gene flow have driven the reduced differentiation, or increased connectivity, observed in populations from Southern Peru compared to northern populations.
We organize IBD segments in two intervals corresponding to (A) pre- and colonial gene flow between 14 urban Peruvian populations and B postcolonial and post-admixture events. The first interval includes IBD segments smaller than 18.75 cM. IBD segments greater than 18.75cM represent the DNA sharing of the last eight generations. For visualization purposes, internal sharing was ignored and only values above the median. Populations are ordered roughly from northern to southern Peru. C Effective population size inferred through HapNe for 14 urban populations. Lima (represented by PGP Lima and PEL with green shade with dotted and solid lines) shows continuous population growth. The y-axis is on a log scale. Shading around the lines represents a 95% confidence interval. All analyses are based on n = 470 unrelated individuals from PGP and PEL from 1000 Genomes Project (Arequipa = 26, Ayacucho = 35, El Carmen = 37, Cusco = 36, Huaraz = 36, Iquitos = 29, Lambayeque = 12, Lima = 28, PEL = 85, Moquegua = 30, Juliaca = 40, Tacna = 21, Tumbes = 33, Trujillo = 22).
Furthermore, we leveraged the IBD segment distribution to infer the trajectory of the effective population size (Ne) in recent generations using HapNe44 (See “Methods”). Our analyses revealed a marked population growth in several southern urban groups, including Tacna, Arequipa, and Juliaca, compared to northern groups (Fig. 3C, Supplementary Figs. 9-S10). Notably, both Lima sample and PEL, each representing the capital city, show an even steeper increase in Ne, consistent with recent migration flow into Lima and with the fact that it includes a third part of the country’s total population. In contrast, the El Carmen population, predominantly self-identified as Afro-Peruvian, displays a declining Ne trajectory over the past 25 generations, underscoring a distinct demographic pattern, likely characterized by isolation.
The colonial admixture was characterized by the encounter of continental ancestries, leaving a signature of sex-bias admixture reported in several Latin American populations45,46. Considering our fine-scale sampling, we evaluated sex bias admixture and its regional differences. A comparative analysis of ancestry proportions revealed that, in nine of thirteen populations, European ancestry was significantly higher in autosomes compared to the X chromosome (Wilcoxon test, p < 0.05; Fig. 4, Supplementary Data 3). The complementary pattern was observed for Indigenous American ancestry (Fig. 4 and Supplementary Data 3). The African ancestries showed a similar pattern to that of Europeans in five populations, but with small proportions (Fig. 4 and Supplementary Data 3). These results suggest that, with some exceptions, European admixture in Peruvians was preferentially mediated by males.
Ancestry proportions were inferred using ADMIXTURE. Ancestry proportions are described in Supplementary Data 3. Box plots show the median (center line), interquartile range (box), and data points within 1.5× the interquartile range (whiskers).
Discussion
Urban populations in western South America exhibit high levels of Indigenous American ancestry9,47, in contrast to other South American groups3,4,48,49,50. This study, focusing on urban populations from Peru, builds upon the first phase of the Peruvian Genome Project9,10,51 and incorporates new genome-wide data. Importantly, this cohort is the same as that analyzed in Borda et al.8; however, while the previous study focused on continental-scale migration patterns across Latin America using genotyped and imputed data, the present work provides a detailed, fine-scale characterization of population structure within Peru based exclusively on directly genotyped variants. We replicate previous results and present three key findings about Peruvian genetic composition: (i) the population structure of urban individuals reflects a North-South gradient similar to that observed among Indigenous Peruvians, driven by their elevated Indigenous American ancestry; (ii) Most continental admixture events before eight generations ago were followed by internal migration patterns that align with the North-South continuum; and (iii) the trajectory of the effective population growth in Lima reflects recent rural to urban migration.
Our first contribution builds on efforts to characterize fine-scale ancestries in urban Peruvian populations. The earliest evidence of a latitudinal population structure in Peru came from an autosomal STR study covering 20 of the 24 administrative regions14. Subsequent work using dense genomic data9 and large sample sizes4 expanded on this question but lacked sufficient Native American references to resolve finer-scale ancestry patterns. Here, we not only confirm the latitudinal distribution of Native American ancestries in urban populations but also identify contributions from distinct biogeographical regions—the Pacific Coast, Andes, and Amazon rainforest. This pattern, previously described in Native American communities, is now shown to persist in urban areas, indicating that mestizaje and internal migration did not erase regional genetic structure. Specifically, we found that ancestry related to present-day Aymara speakers is more frequent in southern urban populations, where most Aymara groups are located, while Quechua-related ancestry is present across all urban samples. The widespread distribution of Quechua-related ancestry reflects the long history of interactions across the Andes, shaped by pre-Inca, Inca, and colonial migrations18,52,53
Our second contribution explains the dates and mechanisms through which ancestral source populations interacted, shaping the present-day genetic composition of an urban area population. Here, we identified at least two admixture events. The most recent admixture events were during the last eight to 12 generations involving sources with an admixed genetic profile which suggests previous admixture events. Notably, both methods revealed African-European or Indigenous American-European interactions, but no African-Indigenous American. This pattern is consistent with colonial social hierarchies, in which Spaniards deliberately segregated enslaved Africans and Indigenous peoples, restricting social interactions between them23. The inferred European-related sources in the genetic pool of admixed Peruvians align with historical records, as expected, indicating significant contributions from Southern Europe53. For African ancestries, our results suggest more geographically diverse sources including components associated with West, West-Central, and East African regions. Our GLOBETROTTER analyses indicate that admixture among these components occurred before the last ten generations, with Afrodescendants from El Carmen showing contributions from all three African sources. By the time of independence in 1821, Spaniard invaders had imported over 100,000 enslaved African people to Peru to meet agricultural demands, mainly on the North Coast during the Transatlantic Trafficking of Enslaved Africans52,54. They were brought from ports in the Caribbean region, where European invaders gathered African individuals from diverse ethnic backgrounds to sow confusion and prevent an uprising55. This practice could explain the mixed African ancestries observed in the source population of the most recent admixture event.
For East Asian ancestries, their distribution and diversity align with historical records. We observed these ancestries among individuals in coastal and northern Peru (Fig. 1C). In these regions, there was a huge agricultural development in terms of sugar agriculture. Initially, this development relied on the enslavement of Afrodescendants. However, after the abolition of slavery in 1854, plantation owners sought alternative forms of labor and began importing large numbers of East Asian workers, primarily under indentured contracts. The conditions of these contracts often resembled slavery, as they were exploitative and designed to sustain the profitability of coastal plantations24,56. This migration wave stopped only after international pressure for better life conditions for migrants. These historical processes highlight how economic activities can leave lasting imprints on the genetic diversity of a region. Nevertheless, given the small proportion of ancestry, we have insufficient power to accurately detect the correct admixture timing for this component.
Our third contribution highlights recent demographic events inferred through IBD analyses. Our findings indicate that genetic similarities among urban areas in southern Peru are not only a result of ancient patterns observed in Indigenous Americans from the region but also reflect continuity in migration flows over the last ten generations. This raises the question of how and why people have continued to move in these recurrent patterns. Historically, Native Americans relied on ancestral routes to travel across the Andes. During the invasion and colonial period, the Spanish recognized the importance of these routes, adopting them as the backbone of mobility and economic integration while also establishing new ones57. Over time, extractive and productive activities in Peru—such as silver mining in the Southern Andes, coastal and highland agriculture, and later the Amazonian rubber boom—reshaped migration flows by creating economic hotspots that attracted labor. However, these movements were rarely free. During colonial times, Indigenous populations were compelled to move under systems like the encomienda and the mita, which redirected labor toward mining and other extractive enterprises58. Afrodescendants, brought through the slave trade, were confined largely to plantations, domestic service, and mining, with their movement tightly controlled to safeguard economic interests23. In the Andean region, Indigenous communities remained bound to haciendas through debt peonage and semi-feudal arrangements that restricted their mobility. While in the Amazon, the rubber boom of the late 19th and early 20th centuries further expanded migration into this region, but much of it occurred under exploitative labor regimes resembling slavery. Thus, while economic activities generated new migration patterns across different regions, they also operated within a framework of coercion and restricted movement that disproportionately affected Indigenous and Afrodescendant populations. In recent decades, however, migration has been driven mainly by rural–urban inequalities and political violence, making cities the primary destination, specifically Lima.
Our IBD inference also provide insights about the genetic consequences of the urbanization process in Peruvian groups. Lima, the capital, has long attracted migrants from other regions due to its disproportionate economic development compared to other Peruvian cities. Over the years, thousands have relocated there, making Lima’s population ten times larger than major cities like Trujillo, Arequipa, and Cusco59. This demographic imbalance was not only driven by socioeconomic disparities but by political violence in recent decades60,61,62 and included people from other urban regions but also from Indigenous communities. Importantly, migration from Indigenous communities has also been accompanied by persistent social segmentation in urban areas. Indigenous migrants are often concentrated in low-paid and low-status sectors such as domestic service, street cleaning, gardening, and construction—reflecting persistent inequalities in the integration of rural migrants into city life63. In Peru, these groups are also disproportionately exposed to informal employment, which exacerbates socioeconomic vulnerability64,65. At a regional scale, Peru ranks as the second country in Latin America with the largest absolute number of indigenous people moving to the city in absolute numbers66. Our results are consistent with these historical and contemporary trends, aligning with migration patterns reported until 2015, where regions such as Lima, Arequipa, Tacna, Ucayali, and Tumbes exhibited positive migration rates—congruent with our Ne trajectory analyses. Notably, this trend is even more pronounced in the PEL population, where the Ne trajectory shows a steeper increase.
Our analyses were based on self-identified mestizos, raising the question of what genetics can reveal about this historically and socially constructed category. Colonial societies in the Americas were organized through hierarchical classifications that assigned rights and status according to ancestry (i.e., cultural heritage) and origin. In this context, terms such as mestizo and mestizaje described individuals of mixed European and non-European descent. Today, 60.2% of Peruvians self-identify as mestizo59, yet this identity is fluid. As Paredes67 shows, mestizo status reflects a multidimensional ethno-racial position shaped by phenotype, ancestry, and self-identification. In consequence, mestizo has generally been associated with more socioeconomic advantages than Indigenous identity. In our study, although urban populations on average exhibit substantial non-Indigenous ancestry, they also encompass a wide range of genetic backgrounds, including individuals with nearly full Native American ancestry. This underscores the fluid nature of the mestizo term. We therefore use the terms urban individuals or urban areas to describe our cohort. This framing situates genetic variation within broader processes of migration and urbanization, without treating socially constructed categories as fixed or inherent.
Three limitations can be observed in our study: (a) diversity gaps, poor sampling of the central and Southern Amazonian regions,(b) small sample size to perform demographic analyses, and (c) the power of haplotype-based methods to discriminate ancestries. Our first limitation does not allow us to explore the three clusters identified by Innaconne et al.14. We expect more genomic initiatives to explore the genetic diversity of Indigenous and Admixed groups in central Peru, an important area where the Andean and Amazonian biogeographic regions interact, providing ecosystems with particular selective pressure that shapes their phenotypes. However, our sampling provided interesting clues regarding the diversity and distribution of ancestry components. Our second limitation is our lack of statistical power to correctly identify demographic events related to admixture dynamics and effective population size. For example, our IBD sharing analysis relies on observed IBD distribution, which could be biased due to phasing error, inaccurate IBD detection, or sampling not reflecting population-level patterns. These issues can affect the inference of effective population size; however, our interpretation is based more on the complete picture of the distribution than on particular details. Our third limitation concerns the power of local ancestry inference to discriminate between continental ancestries. This is the case for the inference of segments with East Asian origin, which could be misclassifications of Indigenous American origin and vice versa, generating conflicting results for the inference of admixture between these two ancestral groups. Another example, in our HierarchyMix analyses, is the minimal African ancestry signal combined with small sample size produced spurious inferences of very old admixture events that are unlikely to be reliable. While these limitations restrict the resolution of some of our inferences, they also underscore key directions for future research on the genetic composition of urban populations.
Overall, this work represents another important step in studying genetic history in admixed groups with predominantly Indigenous American ancestry. Several efforts have been performed in Latin America3,10,11,48,68, which, together with our study, represent the effort of local institutions to understand the diversity of their people, people who were underrepresented, but an entire world of genetic and epidemiological histories that we need to understand.
Methods
Ethical approval and informed consent
All study procedures were reviewed and approved by the Peruvian Research and Ethics Committee of the Instituto Nacional de Salud (authorization no. OI-003-11 and OI-087-13). The ethical framework for the collection, stewardship, and analysis of samples was implemented in accordance with international and national standards, including the Declaration of Helsinki. All participants provided written informed consent in the presence of a translator (when needed) and two local witnesses. Participants were explicitly informed of their right to withdraw at any time without explanation. All ethical regulations relevant to human research participants were followed.
Ethics & Inclusion statement
This research was conceived by researchers from the Peruvian National Institute of Health (INS, by its Spanish acronym) and conducted in collaboration with researchers from the University of Maryland and the Universidade Federal de Minas Gerais. INS researchers led the study design and implementation, collected the samples, performed the genotyping, and participated in all stages of data analysis. As noted above, the study was approved by a local ethics review committee. This research does not result in stigmatization, incrimination, discrimination, or any other personal risk to participants. No benefit-sharing measures were discussed, as no biological materials, cultural artefacts, or associated traditional knowledge were transferred outside the country. Finally, local and regional research was integral to the analysis and interpretation of the study findings.
Sampling, genotyping, and quality control
Thirteen populations were sampled in 2010. All details related to sampling were described elsewhere30.
Samples were genotyped using the Illumina Omni2.5 M array in several batches at the Instituto Nacional de Salud (Peruvian National Institute of Health) facilities. We exported raw data to PLINK format using GenomeStudio. We used PLINK1.969 to perform all quality control steps for autosomes and chromosome X. The quality control process at the single-nucleotide variant (SNV) level included: Exclusion of variants with no chromosome information, exclusion of AT/CG genotypes, exclusion of variants with 100% heterozygous, and SNVs out of the Hardy-Weinberg equilibrium (HWE) (threshold 10e-6).
Some batches were genotyped using GRch37. For these batches, liftover to GRch38 was performed using the UCSC liftover tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver) before merging with other GRch38 batches. After merging, we filtered for SNPs and individuals accumulating more than 5% of missing data using PLINK (--geno 0.05 and mind 0.05).
We controlled for relatedness using the KING algorithm70 to identify duplicates and individuals with first- and second-degree relationships. We ran the KING algorithm in the SNPRelate package71 using the snpgdsIBDKING function. To decide which individuals should be removed after relatedness analyses, we used NATORA72. This admixed genotyped data was merged with Peruvian Indigenous American data previously reported9,10 to build the PGP dataset that included 749 individuals (Admixed and Indigenous American Peruvians) and 1,933,621 SNVs.
Phasing
We set the reference allele to the GRCh38 reference allele using the GRCh38_full_analysis_set_plus_decoy_hla.fa file and PLINK. Following this, we phased PGP data using shapeit473 (https://github.com/odelaneau/shapeit4/tree/master) with a reference panel that includes the full 1KGP High coverage31 (n = 3199 individuals) and HGDP data32(n = 827 individuals). To improve phasing quality, we set the mcmc iterations parameters to 10b,1p,1b,1p,1b,1p,1b,1p,10 m. After phasing, we merged Phased PGP with 1KGP and HGDP. Also, we removed multiallelic variants and AT/CG genotypes. Finally, our combined dataset included 4775 individuals, 1,461,568 autosomal SNVs, and 25,418 chromosome X SNVs.
Population structure analyses
We used allele frequency–based methods i.e., principal component analysis [PCA] and genetic clustering to explore population structure in our combined dataset. We chose PCA because it provides a model-free, widely used approach for visualizing major axes of genetic variation and identifying patterns of similarity among our samples and reference populations. Genetic clustering analysis (e.g., ADMIXTURE analyses33) complements PCA by estimating individual ancestry proportions related to the reference population, allowing finer-scale assessment of population structure. In this sense, genetic ancestry was defined based on the proportional similarity between our samples and continental-level reference populations (i.e., African, European, East Asian, and Indigenous American groups). This approach provides a broad-scale framework suitable for characterizing urban populations with complex admixture histories.
Both methods require for unlinked variants. We generated a list of unlinked variants using the SNPRelate snpgdsLDpruning function with a maximum base pair of 1 Mb for the sliding window, a correlation method to calculate LD values, and a threshold of 0.1. With KING relatedness inference and the unlinked SNP list, we ran PCAir74 with a minor allele frequency (MAF) threshold of 5%, kinship threshold of 0.044194, and divergence threshold of -0.044194.
To explore genome-wide ancestry proportions, we ran ADMIXTURE33 from K = 3 to K = 10 in a subset of individuals that includes Full PGP data (admixed and Native Americans), a subset of 1KGP high coverage populations with unrelated individuals (TSI, IBS, CEU, PEL, CLM, MXL, PUR, LWK, MSL, YRI, GWD, JPT, CHB, and CHS), and Native Americans from HGDP (Colombian, Pima, and Maya populations). This subset included unlinked SNVs, and an MAF filter of 5%. We performed ten runs per K value and plotted the run with the highest log-likelihood. To explore differentiation among urban populations, we ran Fst analysis on PLINK using the Weir and Cockerham estimator75.
Identity-by-descent and local ancestry inferences
We removed SNVs with MAF < 0.001 of the full phased data and inferred identity-by-descent (IBD) segments using hap-ibd76 with default parameters. After IBD inference, we ran the merge-ibd-segments software (https://faculty.washington.edu/browning/refined-ibd/merge-ibd-segments.17Jan20.102.jar) to remove gaps between IBD segments. We kept IBD segments greater than 3 cM for subsequent analyses and removed ROH segments.
We performed local ancestry to infer the continental ancestry at the chromosomal level. For this purpose, we ran GNOMIX77 with four continental references based on ADMIXTURE results (Population Structure analyses section). We selected European (n = 404 individuals; Populations: CEU, GBR, IBS, and TSI), African (n = 405 individuals; Populations: YRI, MSL, GWD, and LWK), and East Asian (n = 411 individuals; Populations: CHB, CHS, JPT, and KHV) individuals from 1KGP populations (Supplementary Data 3). Native American reference included 187 individuals from PGP with more than 99% NAT ancestry based on ADMIXTURE K4 results. We ran GNOMIX using the option “best” for model inference parameters. This option uses a random string kernel base + xgboost smoother and is recommended for array data. We also include the rephasing option to correct the phase inference based on local ancestry information.
We ran ancestry-specific PCA (asPCA) to explore the similarity between chromosomal ancestry segments of PGP individuals to specific continental ancestral references using the Browning et al. pipeline35 (http://faculty.washington.edu/sguy/local_ancestry_pipeline/). Before running the asPCA, we convert msp GNOMIX output to viterbi output since the pipeline was designed for rfmix ver1 outputs. The msp output includes ancestry information for loci in each pair of haplotypes. Considering a 4-way admixture, we created an ancestry-specific msp file setting as missing the other three ancestries. To perform asPCA, we kept individuals with a minimum of 30% for European or African ancestry. For East Asian ancestry, we kept individuals with a minimum of 5%.
Admixture dynamics analyses
We explored admixture dynamics in Peruvians by analyzing different evolutionary aspects: (1) admixture timing and mode, (2) recent migration explored by IBD sharing, (3) demographic changes identified by the evolution of effective population size (Ne), and (4) Sex-bias admixture analyses.
First, to analyze the admixture timing and mode, we ran GLOBETROTTER and HierarchyMix to explore admixture timing and mode.
To run GLOBETROTTER, we first ran ChromoPainter-based methods4,36,40 to identify the timing and mode for admixture events. This analysis requires several steps: (i) inference of scaling parameters [recombination and mutation rates], (ii) inference of haplotype sharing (copyvectors and painting samples), (iii) inference of ancestry proportions using haplotype information, and (iv) inference of admixture date and mode. We excluded two Native American populations, Quechuas and Aimaras (Supplementary Data 2), as both carry a substantial proportion of European ancestry. Their inclusion led to a marked reduction in inferred European ancestry compared to ADMIXTURE results. Similarly, we excluded Xibo and Uygur, as they contributed unreliable estimates of Asian ancestry.
Inference of scaling parameters
We ran ChromoPainter version 2 to determine scaling parameters for haplotype sharing inference. For this purpose, we performed a first ChromoPainter run in a subset of chromosomes (1,8,15,22) and a subset of individuals to calculate the scaling parameters.
Inference of haplotype sharing
After calculating the scaling parameters, we set these values in two ChromoPainter runs including all individuals and chromosomes to generate two types of files: copyvectors and painting samples. Our second Chromopainter was to infer copyvectors, we set the parameter -f to select the donors and recipients. We selected all populations as recipients but just Indigenous Americans, European, African, and Asian populations as donors (Supplementary Data 2). Our third Chromopainter run was to infer painting samples, we set the -f mode setting admixed Peruvians as recipients and all other populations as donors.
Ancestry proportions at the subcontinental level
After ChromoPainter runs, we merged copyvectors into one coancestry matrix using the neaverage perl script (https://people.maths.bris.ac.uk/~madjl/finestructure/neaverage.pl.zip) and ran SOURCEFIND4 ver2 (https://github.com/hellenthal-group-UCL/sourcefindV2) to identify ancestry proportions corresponding to donor populations. SOURCEFIND is a Bayesian approach that explores haplotype sharing between donor individuals and recipients to determine ancestry proportions beyond the continental level.
Admixture inference
To determine the date and mode of admixture events, we ran GLOBETROTTER40 (https://github.com/hellenthal-group-UCL/GLOBETROTTER), a software that explores ancestry proportions and linkage disequilibrium decay. We ran GLOBETROTTER in each population and performed 100 bootstrap replications. Crucially, in assessing whether the inferred admixture event in a population signifies a “one-date” or “multiple-dates”, we checked the ancestry curves and the goodness-of-fit (R2) for a single date of admixture (maxR2fit.1date). If the maxR2fit.1date exceeded 99.9%, we interpreted the result as indicative of a one-date admixture, adopting the date and proportions proposed by this mode (criterion confirmed by G. Hellenthal, pers. comm., 2025). However, it’s important to note that this inference pertains to the most recent event and does not preclude the possibility of earlier events.
For HierarchyMix Inference, we use GNOMIX outputs in bed format to run HierarchyMix (https://github.com/Shuhua-Group/HierarchyMix) with default parameters except the number of bootstrap replicates and threshold for ancestry segments. We ran 1000 bootstrap replicates and kept segments greater than 2 cM. No suitable model was identified for four populations (Lima, Lambayeque, Moquegua, and Tacna).
Second, we analyzed IBD sharing among PGP populations to explore recent migration. We kept IBD segments greater than 18.75 cM to explore sharing among populations during the last eight generations that resemble Colonial and post-colonial events. Considering populations X, Y, and Z, we assumed that any IBD segment shared among X and Y populations corresponds to people’s movement among them. However, we cannot rule out that sharing segments between populations could also explain migration from a third source (population Z) to both populations (X and Y) during the same period. To estimate the interpopulation sharing after keeping IBD segments greater than 18.75 cM, we sum the number and the length of IBD segments shared between individuals from different populations. To control for sample sizes, both the total number and the total length were divided by the product of the sample sizes of both populations.
For plotting IBD and GLOBETROTTER results, we modified R codes available at: https://github.com/georgebusby/admixture_in_africa ; and https://github.com/chiarabarbieri/SNPs_HumanOrigins_Recipes
Third, we inferred the distribution of the effective population size during the last 50 generations for PGP data using HapNe44 following the pipeline described here: https://github.com/PalamaraLab/HapNe. Briefly, for each population, each phased chromosome was split into two parts, each divided by the centromere using PLINK. Then, each chromosomal segment was used to infer IBD segments using hap-ibd and then merged with the merge-ibd-segments.jar software. Finally, these outputs were used as inputs for HapNe with default parameters.
Fourth, to interrogate whether a population resulted from sex-bias admixture, we compared admixture proportions from autosomes and chromosome X. For autosomes, we used results from the ADMIXTURE analyses described above. To infer continental ancestry proportions for chromosome X, we ran ADMIXTURE using the --haploid flag. We compared autosomal and chromosome X proportions using the Wilcoxon test.
Statistics and reproducibility
Our full dataset includes 432 individuals from urban populations (Supplementary Data 1); however, most analyses were restricted to unrelated individuals (n = 385) to avoid biases due to familial relatedness. Comparisons between urban populations were conducted on samples ranging from 22 to 40 unrelated individuals per site, reflecting variable sample availability. Most analyses, including PCA, ADMIXTURE, and GLOBETROTTER, were performed across sites to characterize population structure and admixture patterns, while the sex-bias analysis compared autosomal and X chromosome variation within each site. Additionally, heatmaps of pairwise population sharing were used to assess relative relatedness between sites. Continuous variables were compared using a two-sided Wilcoxon rank-sum test with a significance threshold of 0.05. All analyses and visualizations were performed in R (version 4.3) using publicly available software, ensuring transparency and reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data has been deposited at the European Genome-phenome Archive (EGA), which is hosted by the EBI and the CRG, under accession number EGAS00001008264. Further information about EGA can be found on https://ega-archive.org “The European Genome-phenome Archive of human data consented for biomedical research” (https://doi.org/10.1093/nar/gkab1059). All data used for the graphs and charts are included as Supplementary Data and indicated in the Figure caption. For Fig. 1A, the ancestry proportions are observed in Supplementary Data 3. For Fig. 1c, the ancestry proportions are observed in Supplementary Data 4. For Fig. 2, the dates and ancestry proportions are observed in Supplementary Data 5 and 6, respectively. For Fig. 4, ancestry proportions are observed in Supplementary Data 3.
References
Mills, M. C. & Rahal, C. The GWAS diversity monitor tracks diversity by disease in real time. Nat. Genet 52, 242–243 (2020).
Homburger, J. R. et al. Genomic insights into the ancestry and demographic history of South America. PLoS Genet 11, e1005602 (2015).
Kehdy, F. S. G. et al. Origin and dynamics of admixture in Brazilians and its effect on the pattern of deleterious mutations. Proc. Natl. Acad. Sci. USA 112, 8696–8701 (2015).
Chacón-Duque, J.-C. et al. Latin Americans show wide-spread Converso ancestry and imprint of local Native ancestry on physical appearance. Nat. Commun. 9, 5388 (2018).
Ongaro, L. et al. The Genomic Impact of European Colonization of the Americas. Curr. Biol. 29, 3974–3986.e4 (2019).
Gouveia, M. H. et al. Origins, admixture dynamics, and homogenization of the African Gene Pool in the Americas. Mol. Biol. Evolut. 37, 1647–1656 (2020).
Medina-Muñoz, S. G. et al. Demographic modeling of admixed Latin American populations from whole genomes. Am. J. Hum. Genet. 110, 1804–1816 (2023).
Borda, V. et al. Genetics of Latin American diversity project: Insights into population genetics and association studies in admixed groups in the Americas. Cell Genomics 4, 100692 (2024).
Harris, D. N. et al. Evolutionary genomic dynamics of Peruvians before, during, and after the Inca Empire. Proc. Natl. Acad. Sci. USA 115, E6526–E6535 (2018).
Borda, V. et al. The genetic structure and adaptation of Andean highlanders and Amazonians are influenced by the interplay between geography and culture. Proc. Natl. Acad. Sci. USA 117, 32557–32565 (2020).
Sohail, M. et al. Mexican Biobank advances population and medical genomics of diverse ancestries. Nature 622, 775–783 (2023).
Sandoval, J. R. et al. Tracing the genomic ancestry of Peruvians reveals a major legacy of pre-Columbian ancestors. J. Hum. Genet 58, 627–634 (2013).
The 1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Iannacone, G. C. et al. Peruvian genetic structure and their impact in the identification of Andean missing persons: A perspective from Ayacucho. Forensic Sci. Int.: Genet. Suppl. Ser. 3, e291–e292 (2011).
Rodríguez Mega, E. How the mixed-race mestizo myth warped science in Latin America. Nature 600, 374–378 (2021).
de la Cadena, M. ¿son los mestizos híbridos? las políticas conceptuales de las identidades andinas. Univ. Humanist. 61, 51–84 (2006).
Stolcke, V. Los mestizos no nacen sino que se hacen. Avá. Revista de Antropología 14, (2009).
Nakatsuka, N. et al. A paleogenomic reconstruction of the deep population history of the Andes. Cell 181, 1131–1145.e21 (2020).
Urban, M. & Barbieri, C. North and South in the ancient Central Andes: Contextualizing the archaeological record with evidence from linguistics and molecular anthropology. J. Anthropol. Archaeol. 60, 101233 (2020).
Cook, N. D. Patrones de migración indígena en el Virreinato del Perú: mitayos, mingas y forasteros. HISTORICA 13, 125–152 (1989).
Escobar, G. & Beall, C. M. Contemporary Patterns of Migration in the Central Andes. Mt. Res. Dev. 2, 63 (1982).
Anna, T. E. The Peruvian declaration of independence: Freedom by coercion. J. Lat. Am. Stud. 7, 221–248 (1975).
OʹToole, R. S. Bound Lives: Africans, Indians, and the Making of Race in Colonial Peru. (University of Pittsburgh Press, https://doi.org/10.2307/j.ctt5hjpjn 2012).
Gonzales, M. J. Chinese plantation workers and social conflict in Peru in the Late Nineteenth Century. J. Lat. Am. Stud. 21, 385–424 (1989).
Takenaka, A. The Japanese in Peru: History of immigration, settlement, and racialization. Lat. Am. Perspect. 31, 77–98 (2004).
Menton, M. & Cronkleton, P. Migration and Forests in the Peruvian Amazon: A Review. (Center for International Forestry Research (CIFOR), https://doi.org/10.17528/cifor/007305 2019).
INEI. Perú: Perfil Sociodemográfico Informe Nacional. (2018).
Contreras, C. & Cueto, M. Historia Del Perú Contemporáneo. (Instituto de Estudios Peruanos, Peru, 2024).
Coral, I. Desplazamiento por violencia política en el Perú, 1980-1992. In (1994).
Guio, H. et al. The peruvian genome project: expanding the global pool of genome diversity from South America. Front. Genet. 16, 1614021 (2025).
Byrska-Bishop, M. et al. High-coverage whole-genome sequencing of the expanded 1000 Genomes Project cohort including 602 trios. Cell 185, 3426–3440.e19 (2022).
Bergström, A. et al. Insights into human genetic variation and population history from 929 diverse genomes. Science 367, eaay5012 (2020).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Moreno-Estrada, A. et al. Reconstructing the Population Genetic History of the Caribbean. PLoS Genet 9, e1003925 (2013).
Browning, S. R. et al. Local Ancestry Inference in a Large US-Based Hispanic/Latino Study: Hispanic Community Health Study/Study of Latinos (HCHS/SOL). G3 Genes|Genomes|Genet. 6, 1525–1534 (2016).
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of Population Structure using Dense Haplotype Data. PLoS Genet 8, e1002453 (2012).
Bonfiglio, G. Introducción al estudio de la inmigración europea en el Perú. Apuntes: Revista de Ciencias Sociales 93–127 (2015).
Bowser, F. P. The African Slave in Colonial Peru, 1524-1650. (Stanford University Press, Stanford, Calif, 1974).
Sobrevilla Perea, N. The Abolition of Slavery in the South American Republics. Slavery Abolit. 44, 90–108 (2023).
Hellenthal, G. et al. A Genetic Atlas of Human Admixture History. Science 343, 747–751 (2014).
Zhang, S. et al. Reconstructing complex admixture history using a hierarchical model. Brief. Bioinforma. 25, bbad540 (2024).
Wang, R. J., Al-Saffar, S. I., Rogers, J. & Hahn, M. W. Human generation times across the past 250,000 years. Sci. Adv. 9, eabm7047 (2023).
Baharian, S. et al. The Great Migration and African-American Genomic Diversity. PLoS Genet 12, e1006059 (2016).
Fournier, R., Tsangalidou, Z., Reich, D. & Palamara, P. F. Haplotype-based inference of recent effective population size in modern and ancient DNA samples. Nat. Commun. 14, 7945 (2023).
Ongaro, L. et al. Evaluating the impact of sex-biased genetic admixture in the Americas through the analysis of haplotype data. Genes 12, 1580 (2021).
Pena, S. D. J., Santos, F. R. & Tarazona-Santos, E. Genetic admixture in Brazil. Am. J. Med Genet. Pt C. 184, 928–938 (2020).
Nagar, S. D. et al. Genetic ancestry and ethnic identity in Ecuador. Hum. Genet. Genomics Adv. 2, 100050 (2021).
Luisi, P. et al. Fine-scale genomic analyses of admixed individuals reveal unrecognized genetic ancestry components in Argentina. PLoS ONE 15, e0233808 (2020).
Eyheramendy, S., Martinez, F. I., Manevy, F., Vial, C. & Repetto, G. M. Genetic structure characterization of Chileans reflects historical immigration patterns. Nat. Commun. 6, 6472 (2015).
Adhikari, K., Mendoza-Revilla, J., Chacón-Duque, J. C., Fuentes-Guajardo, M. & Ruiz-Linares, A. Admixture in Latin America. Curr. Opin. Genet. Dev. 41, 106–114 (2016).
Alvim, I. et al. The need to diversify genomic studies: Insights from Andean highlanders and Amazonians. Cell 187, 4819–4823 (2024).
Aguirre, C. Breve Historia de La Esclavitud En El Perú: Una Herida Que No Deja de Sangrar. (Fondo Editorial del Congreso del Perú, Lima, 2005).
The Oxford Handbook of Latin American History. (Oxford University Press, Oxford; New York, 2011).
Golash-Boza, T. M. Yo Soy Negro: Blackness in Peru. (University Press of Florida, Gainesville, 2012).
Marcum, A. & Skarbek, D. Why didn’t slaves revolt more often during the Middle Passage? Rationality Soc. 26, 236–262 (2014).
Balan, J. & Liévana, A. Estructuras agrarias y migración interna en una perspectiva histórica: estudios de casos latinoamericanos. Rev. Mex. Sociol. 43, 141–192 (1981).
Marcone, G. The roads of power and the paths of the people: Reconfiguring communities and territories from pre-Hispanic Andean roads to modern Peruvian highways. Journal of Social Archaeology 14696053251348221 https://doi.org/10.1177/14696053251348221 (2025).
Dell, M. The persistent effects of Peru’s mining mita. Econometrica 78, 1863–1903 (2010).
Instituto Nacional de Estadistica e Informatica. Situacion de la Poblacon Peruana 2024: Una mirada de la diversidad étnica. (2024).
Pedersen, D., Tremblay, J., Errázuriz, C. & Gamarra, J. The sequelae of political violence: Assessing trauma, suffering and dislocation in the Peruvian highlands. Soc. Sci. Med. 67, 205–217 (2008).
Loret De Mola, C. et al. The effect of rural-to-urban migration on social capital and common mental disorders: PERU MIGRANT study. Soc. Psychiatry Psychiatr. Epidemiol. 47, 967–973 (2012).
Falaris, E. M. The Determinants of Internal Migration in Peru: An Economic Analysis. Economic Dev. Cultural Change 27, 327–341 (1979).
Oliva Martínez, J. D. Diversidad, resistencia y utopía: los pueblos indígenas de nuestro tiempo. (Tirant Humanidades, Valencia, 2022).
Parodi, J. Ser Obrero Es Algo Relativo: Obreros. Clasismo, Y’Política (Lima: Instituto de Estudios Peruanos, 1986) (1986).
Vincent, S. Pensions, Peasants, and the Informal Economy: Family and Livelihood in Contemporary Peru. Lat. Am. Perspect. 45, 195–211 (2018).
Los pueblos indígenas en América (Abya Yala): desafíos para la igualdad en la diversidad. (Naciones Unidas, CEPAL, Santiago, 2017).
Paredes, C. L. Multidimensional Ethno-racial Status in Contexts of Mestizaje: Ethno-racial Stratification in Contemporary Peru. Socius: Sociological Research for a Dynamic World 4, 1–18 (2018).
Vicuña, L. et al. Novel loci and Mapuche genetic ancestry are associated with pubertal growth traits in Chilean boys. Hum. Genet 140, 1651–1661 (2021).
Purcell, S. et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Leal, T. P. et al. NAToRA, a relatedness-pruning method to minimize the loss of dataset size in genetic and omics analyses. Comput. Struct. Biotechnol. J. 20, 1821–1828 (2022).
Delaneau, O., Zagury, J.-F., Robinson, M. R., Marchini, J. L. & Dermitzakis, E. T. Accurate, scalable and integrative haplotype estimation. Nat. Commun. 10, 5436 (2019).
Conomos, M. P., Miller, M. B. & Thornton, T. A. Robust inference of population structure for ancestry prediction and correction of stratification in the presence of relatedness. Genet. Epidemiol. 39, 276–293 (2015).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358 (1984).
Zhou, Y., Browning, S. R. & Browning, B. L. A fast and simple method for detecting identity-by-descent segments in large-scale data. Am. J. Hum. Genet. 106, 426–437 (2020).
Hilmarsson, H. et al. High Resolution Ancestry Deconvolution for Next Generation Genomic Data. http://biorxiv.org/lookup/doi/10.1101/2021.09.19.460980https://doi.org/10.1101/2021.09.19.460980 (2021).
Acknowledgements
We are deeply grateful to all participants who generously donated blood samples for the Peruvian Genome Project. Their contribution was vital to the success of this research. We thank all those who facilitated the recruitment of participants, including the Direcciones Regionales de Salud from Loreto, Puno, Cusco, La Libertad, Huancavelica, Ica, Piura, Ancash, Arequipa, Ayacucho, Tacna, Ucayali, San Martin, Amazonas; Universidad Andina Nestor Caceres Velasquez Facultad de Ciencias de la Salud, Universidad Nacional Jorge Basadre Grohmann, Universidad Nacional Mayor de San Marcos, Universidad Nacional San Agustín, Universidad Nacional de San Cristóbal de Huamanga, Universidad. Nacional Santiago Antúnez de Mayolo, and Universidad Nacional de Trujillo. This work was supported by the University of Maryland Institute for Health Computing (V.B. and B.A.M.), the Institute for Genome Sciences and Program in Personalized Genomic Medicine at the University of Maryland School of Medicine (V.B., D.V.O., and T.D.O.), and the Instituto Nacional de Salud, Lima, Perú (O.C., C.S., C.P., and H.G.). ET-S and CS-C are supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Apoio à Pesquisa do Estado de Minas Gerais (FAPEMIG), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), and the Brazilian Ministry of Health (Department of Science and Technology-DECIT). M.M. was supported by the CGEn HostSeq/CIHR fellowship (CGE 185054) and the SickKids Restracomp Fellowship.
Author information
Authors and Affiliations
Contributions
The project was conceived by V.B., O.C., C.S., C.P., E.T.S., H.G., and T.D.O. Sampling and Genotyping data were performed by O.C., C.S., C.P., and H.G. Quality control was performed by V.B. Data analysis was performed by V.B., D.V.O., and M. M., C.S.C., I.A., B. A. M., and M.H.G. All authors contributed with data interpretation. V.B., E.T.S., and T.D.O. wrote the manuscript. All authors read the manuscript and contributed with suggestions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Epífania Arango-Isaza and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Borda, V., Caceres, O., Sanchez, C. et al. Unraveling the genetic landscape and admixture dynamics of urban populations across Peru. Commun Biol 9, 410 (2026). https://doi.org/10.1038/s42003-026-09671-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09671-2