Abstract
Zika virus (ZIKV) infection is an international concern of public health emergency and has been associated with severe neurodevelopmental abnormalities. ZIKV-encoded envelope protein was virulent to neural cells, but its role in the neurodifferentiation process is not well known. Thus, we used mouse embryonic stem cells (mESCs) to mimic early neural differentiation in vitro and comprehensively studied the role of ZIKV envelope protein in the early directional differentiation of the neural lineage. We found that the ZIKV envelope protein does not affect the pluripotency of mESCs, but is a strong blocker of early neural differentiation. In both monolayer and suspension culture neural differentiation models, abnormalities arise from neural lineage commitment to early neuronal differentiation. During differentiation, envelope protein downregulated a large number of neurodevelopmentally relevant genes, therefore inhibiting multiple key biological processes, including neuroactive ligand-receptor interactions, neuron, synaptic processes, and dendritic spine development. ZIKV envelope protein disrupts early directed differentiation within neural lineages, providing key evidence for ZIKV-mediated neurodevelopmental abnormalities such as fetal microcephaly.
Similar content being viewed by others
Introduction
Zika virus (ZIKV), an enveloped neurotropic Flavivirus, was isolated from a feverish rhesus monkey in Uganda in 1947 and initially spread only in Africa1. Subsequently, it spread from the African continent to Asia and the Pacific Islands region, eventually reaching the Americas2,3. ZIKV can attack the central nervous system (CNS) and peripheral nervous system (PNS)4,5, leading to severe neurological disorders and congenital malformations, resulting in microcephaly of fetuses and babies, called congenital Zika syndrome (CZS)6.
The genome of ZIKV is a single-stranded positive-sense RNA of approximately 10.8 kb in length. It contains one open reading frame (ORF) flanked by 5’- and 3’-untranslated regions (UTRs). The ORF encodes a polyprotein that is processed into three structural proteins (capsid protein, C; precursor membrane protein, prM; and envelope protein, Env), as well as seven nonstructural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5)7,8. Structural proteins are responsible for the assembly and invasion of viral particles into host cells, while non-structural proteins regulate viral genome RNA replication or mediate the host immune response9. Interaction of Env protein with cell surface attachment receptors facilitates the entry of viral particles into the host cells9.
N-glycosylation is one of the most important post-translational modifications in proteins, occurring at a highly conserved tripeptide sequence: Asn-X-Ser/Thr. The process involves the covalent attachment of an oligosaccharide chain (glycan) to the amide nitrogen atom of a specific amino acid (asparagine, Asn, N) in the protein10. During the early ZIKV epidemic, African and several other historical strains lacked glycosylation motifs in the Env protein11. However, contemporary epidemic strains developed glycosylation sites during passage11. The N154 glycosylation site is typically involved in the binding to cell surface receptors (such as DC-SIGN and DC-SIGNR), facilitating viral entry into the host. Viruses lacking N154 glycosylation exhibit reduced virulence, with decreased viral loads in the brain and blood12. The N-linked glycosylation at position 154 of Env is considered to be a key determinant of ZIKV virulence and neuroinvasiveness.
ZIKV has been shown to infect various types of cells and tissues such as fetal neurons, neuroblastoma cells, mesenchymal stem cells, and primary trophoblast cells13,14,15. It can also directly infect monolayers, 3D neurospheres, and human induced pluripotent stem cell (iPSC) in vitro16,17. Neurogenesis is a key process by which neural stem cells (NSCs) or neural progenitor cells (NPCs) differentiate into neurons, and defects in neurogenesis lead to developmental neurological disorders, including microcephaly18. ZIKV targets NPC and down-regulates microcephaly-related genes such as microcephaly protein 1 (MCPH1), cancer susceptibility candidate 5 (CASC5), or aberrant spiny microcephaly-associated protein (ASPM), resulting in the malfunction of NPC and its derivatives19. ZIKV causes brain developmental disorders and affects the development of NSCs in mice model19.
Embryonic stem cells (ESCs) are a type of highly undifferentiated cell isolated from early embryos. Under specific conditions, ESCs maintain pluripotency and differentiate into specific cell types. ESCs can sequentially differentiate into NSCs and NPCs in vitro, and further differentiate into neurons. These neurons subsequently mature, forming typical structures such as axons and dendrites20. Mouse embryonic stem cells (mESCs) primarily maintain pluripotency in vitro through the 2i (PD0325901, CHIR99021)/LIF pathway21. PD0325901 inhibits the FGF-MAPK pathway, while CHIR99021 suppresses the GSK pathway22. Mouse leukemia inhibitory factor (LIF) activates STAT3 and bone morphogenetic protein (BMP) to induce differentiation-inhibiting proteins23. N2B27 differentiation is a well-established method for inducing mESCs to differentiate into a monolayer neural lineage in vitro, supporting the long-term survival of neurons in the central nervous system. This method is simple to operate and reproducible. Using serum-free adhesion differentiation, the addition of N2, B27, and other growth factors induces cells to differentiate into NPCs and neurons24. KSR differentiation is through Serum Replacement Knockout (Knockout™ SR, KSR) medium to direct mESCs toward a telencephalon precursor. KSR medium can induce mESCs to differentiate into neural spheres composed of NPCs and neurons25.
ZIKV Env protein can cause direct toxicity to neurons by over activating poly adenosine diphosphate-ribose polymerase 1 (PARP1)26. ZIKV Env protein also alters cellular properties by inducing pro-neural genes and modulating microRNA circuitry, thereby inducing premature differentiation of fetal neural stem cells and impeding neuronal differentiation27. Previous studies have linked ZIKV Env protein to neurodevelopmental defects observed in congenital Zika syndrome. Our study established a method to induce Env expression in undifferentiated mouse embryonic stem cells and traced its effects throughout the continuum from neural lineage commitment to early neuronal differentiation. This offers a unique perspective on how early Env exposure disrupts neural lineage specialization from the stem cell stage. We found that Env/N154A barely affects the pluripotency, proliferation, and cell cycle of mESCs, but impairs their ability to spontaneously form embryoid bodies (EBs) and differentiate into three embryonic layers. Differentiation into the neural lineage revealed that Env targets multiple genes and specific biological processes, blocking the ability of mESCs to differentiate into early neurons.
Results
Env of ZIKV has little effect on the pluripotency, proliferation, and cell cycle, but significantly inhibits the free differentiation of mESCs
The mESC 46C expresses GFP under control of the Sox1 promoter (Sox1-GFP mESCs)28. To investigate the biological functions of ZIKV Env protein from neural lineage commitment to early neuronal differentiation in vitro, we constructed 46C embryonic stem cell line 46C-Env expressing flag-tagged Env (Fig. 1A–E) by using Lentiviral vector-mediated gene transduction. The empty Lentiviral vector-transduced 46C was designated as wild-type cell line (WT). The expression of flag-tagged Env was confirmed at the protein level and mRNA level by Western blotting and RT-qPCR, respectively, in either 293T cells (Fig. 1B, C) or mESCs (Fig. 1D, E). To determine whether mutation of the key glycosylation site of Env can cause distinct consequence, a mESCs 46C-N154A was made expressing Env containing the N-to-A mutation of a glycosylation site at the amino acid position 154 (Fig. 1A)29. The stable expression of flag-tagged Env-N154A was confirmed in parallel with the non-mutated Env (Fig. 1B–E).
A Diagram showing ZIKV Env protein and mutation at N154A. B, C RT-qPCR and Western blot results of Env expression in HEK293T cells 48 h after transfection. D, E RT-qPCR and Western blot results of Env expression in mESCs. F Representative immunofluorescence images of the expression of pluripotent markers NANOG and OCT4 in 46C-WT and -Env/N154A. Scale bar, 50 μm. G RT-qPCR of representative pluripotent gene expression in 46C-WT and -Env/N154A. H Cell cycle analysis of 46C-WT and -Env/N154A. I CCK8 proliferation assay at 24 h, 48 h, 72 h, 96 h, respectively. J Morphological images of EB differentiation of 46C-WT and -Env/N154A at D4, D8 and D18. Scale bar, 100 μm or 200 μm, respectively. K, L Expression of three-layer markers measured by RT-qPCR. The relative expression levels were quantitated using ImageJ software. The error bars represent the mean ± SD, and the significance level was calculated by Student’s t test (two-tailed, equal variance) (ns, not statistically significant, ∗, p < 0.05; ∗∗, p < 0.01; and ∗∗∗, p < 0.001) (n ≥ 3).
The effects of Env or N154A expression on the pluripotency of mESCs were assessed by analyzing the expression of pluripotency hall marker genes using immunofluorescence staining (Fig. 1F) or RT-qPCR (Fig. 1G and Supplementary Data). The expression of Nanog and Oct4 was not affected by either Env or N154A (Fig. 1F, G). The transcripts of Sox2 and Klf4 were not affected by Env (Fig. 1G). N154A only slightly reduced the transcript of Sox2 (Fig. 1G). Taken together, expression of Env or N154A barely affects the pluripotency of the mESCs, suggesting that the mESCs remain the ability to differentiate in multiple directions.
ZIKV infection induces mitotic abnormalities in human neural progenitor cells, and Env induces G2/M cell cycle arrest in neuroendocrine PC12 cells30,31. We then examined whether Env affects the cell cycle and growth rate of mESC and found that Env or N154A did not significantly change the cell cycle compared to WT (Fig. 1H), although the early growth rate of mESCs was slightly reduced by Env (Fig. 1I and Supplementary Data).
The inherent property of pluripotent stem cells is the ability to form embryoid bodies (EBs), which recapitulate early embryogenesis32. As ZIKV can enter the developing fetal brain via the placenta33, the free differentiation into EBs in vitro was tested, and we found that the EBs were significantly smaller and unable to form cavities on day 18 in the presence of either Env or N154A (Fig. 1J). Next, the ability of mESCs to differentiate freely toward three embryonic layers was measured by detecting the three-layer markers (Fig. 1K, L). Interestingly, almost all three kinds of germline genes (Pax6 and Nestin in the ectoderm; Gata4 and Sox17 in the endoderm, and Gsc and T in the mesoderm) were significantly down-regulated by Env at day 4 or day 8 (Fig. 1K, L and Supplementary Data). In contrast, N154A expression exerts different effects on the three-layer genes. N154A expression significantly affected Nestin expression at D8. The mesodermal gene Gsc showed mild downregulation at D4, while the T was significantly upregulated at both D4 and D8. For endodermal genes Gata4 and sox17, both were significantly downregulated at D4. However, the regulatory pattern of Gata4 gradually shifted from downregulation to upregulation between D4 and D8 (Fig. 1K, L). These results indicate that Env protein expression suppresses the ability of mESCs to differentiate into the three germ layers in vitro, whereas N154A expression exerts varying degrees of influence on this process.
Env of ZIKV inhibits neurogenesis of mESCs in N2B27 medium
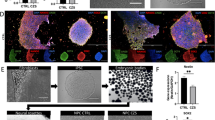
The effects of Env on the process from neural lineage commitment to early neuronal differentiation were investigated by subjecting mESCs to monolayer differentiation in N2B27 medium (Fig. 2A)34. The differentiation on days 3-8 (D3-8) was recorded and the sox1-GFP (neural stem cell marker) fluorescence of 46C-WT mESCs gradually increased over the time course (Fig. 2B). On D6, WT appeared as a large and distinct rosette organized in clusters of neural precursors (neural rosettes). In contrast, 46C-Env showed a few dispersed neural rosettes. On D8, a large number of neurons were observed in WT rather than 46C-Env (Fig. 2B). Meanwhile, the fluorescence intensity in 46C-Env was always lower than that of WT (Fig. 2B), suggesting that Env may have affected neural differentiation. We analyzed the microscopy images of 46c-WT and 46C-Env/N154A, counting clearly defined rosettes formed in at least three independent experiments (Supplementary Fig. S1A in Supplementary Information). Quantitative results show that compared to 46C-WT, 46C-Env exhibits a significant reduction (50%) in neural rosettes. 46C-N154A demonstrates a 70% reduction in neural rosettes (Supplementary Fig. S1A). Flow cytometry (FCM) analysis showed that the number of cells expressing sox1-GFP was significantly lower in 46C-Env than in WT from day 3 to 5, suggesting that Env significantly inhibits neural stem cell formation during the early stage of neural differentiation (Fig. 2C and Supplementary Data).
A Schematic representation of monolayer differentiation of mESCs. B Morphological images of 46C-WT and -Env/N154A during differentiation between day 3-and day 8 in N2B27 medium. Scale bar, 100 μm or 50 μm, respectively. C Flow cytometry analysis to determine the percentage of Sox1-GFP positive cells in differentiated 46C-WT and -Env/N154A cells at D3-8. D Expression of neural marker genes detected by RT-qPCR. E Expression of Pax6, β-III-Tubulin, and GAPDH detected in 10 μg of total protein by Western blot at D7. The relative expression levels were quantitated using ImageJ software. The error bars represent the mean ± SD. The statistical significance level was calculated by one-way analysis of variance (ANOVA). The comparison group is Env vs. WT, N154A vs. WT (ns, not statistically significant, ∗, p < 0.05; ∗∗, p < 0.01; and ∗∗∗, p < 0.001) (n ≥ 3).
Next, the neural stem cell markers (Sox1 and Nestin), neural progenitor cell markers (Pax6 and Neurod1), neuronal marker (β-III-Tubulin), and mature neuronal marker (early dendritic marker) (Map2) were analyzed by RT-qPCR (Fig. 2D and Supplementary Data). The transcripts of those genes were significantly lower in 46C-Env than WT. Western blotting further confirmed that the proteins of neural stem cell marker (Pax6) and neuronal marker (β-III-Tubulin) were significantly reduced on day 7 of differentiation (Fig. 2E and Supplementary Data). To further confirm this effect, we performed immunofluorescence staining for β-III Tubulin on D10 of differentiation. The results revealed a significant reduction in the number of neurons using Env-expressing cells, confirming that early neuronal generation was severely suppressed (Supplementary Fig. S1B in Supplementary Information). Therefore, the expression of Env blocks neural lineage determination and the formation of neural progenitor cells, while simultaneously inhibiting the formation of early neurons.
Compared to 46C-Env, 46C-N154A exhibits a more pronounced inhibitory effect on the process from neural lineage commitment to early neuronal differentiation. Microscopic observations revealed that 46C-N154A exhibits fewer neural rosettes and neurons compared to 46C-Env (Fig. 2B). FCM analysis revealed a significantly lower proportion of Sox1-GFP expressing cells than that of WT and 46C-Env (Fig. 2C), indicating that 46C-N154A may cause more severe effects in the early stage of neural differentiation. Moreover, the transcripts of Sox1, Pax6, Nestin, Neurod1 β-III-Tubulin, and Map2 were significantly downregulated by 46C-N154A when compared with 46C-Env (Fig. 2D). This indicates that 46C-N154A exerts a more pronounced inhibitory effect on early neural differentiation. However, Western blot analysis of Pax6 and β-III Tubulin at the protein level revealed that 46C-Env and 46C-N154A exerted comparable inhibitory effects. Immunofluorescence analysis revealed no significant difference in the expression of the early neuronal marker β-III-tubulin between 46C-Env and 46C-N154A (Supplementary Fig. S1B). Nevertheless, both the expression of Env and N154A exhibited significant inhibitory effects compared to WT in N2B27 medium.
Env of ZIKV inhibits the conversion of mESCs into neuronal cells in suspension culture in KSR medium
To further validate the inhibitory effect of ZIKV Env on early neural lineage differentiation, we induced neural differentiation of mESCs towards the telencephalon in KSR (Knockout Serum Replacement) medium (Fig. 3A)25,35. The morphology and fluorescence intensity of neurospheres at D3-7 were recorded. The fluorescence intensity of Sox1-GFP gradually increased with the time in WT, but the fluorescence of 46C-Env was consistently weaker than that of WT (Fig. 3B). In addition, the morphology of 46C-Env was less smooth than WT (Fig. 3B). FCM analysis of Sox1-GFP further confirmed that 46C-Env inhibited neural differentiation (Fig. 3C and Supplementary Data). Moreover, RT-qPCR analysis showed a significant reduction in the expression of neural stem cell, neural progenitor cell, early neuron, and mature neuron markers in 46C-Env cells (Fig. 3D; Supplementary Data). Pax6 and β-III Tubulin were also downregulated by Env during KSR neural differentiation as evidenced at D7 (Fig. 3E; Supplementary Data). Immunofluorescence analysis of differentiated D10 cells revealed that β-III Tubulin expression was suppressed (Supplementary Fig. S2 in Supplementary Information).
A Schematic representation of three-dimensional neural differentiation of mESCs in KSR medium. B Representative cell morphology of neurospheres derived from 46C-WT and -Env/N154A cell lines at D3-7. Scale bar, 100 μm or 50 μm, respectively. C Flow cytometry analysis of the KSR differentiation process at D3-7. D Expression of neural marker genes detected by RT-qPCR at D4 and D7. E Expression of Pax6, β-III-Tubulin, and GAPDH detected in 10 μg of total protein by Western blot at D7. The relative expression levels were quantitated using ImageJ software. The error bars represent the mean ± SD. The statistical significance level was calculated by one-way analysis of variance (ANOVA). The comparison group is Env vs. WT, N154A vs. WT (ns, not statistically significant, ∗, p < 0.05; ∗∗, p < 0.01; and ∗∗∗, p < 0.001) (n ≥ 3).
Compared to 46C-Env, 46C-N154A has weaker Sox1-GFP fluorescence (Fig. 3B) and the lower percentage of fluorescence positive cells (Fig. 3C), indicating that 46C-N154A may have more severe effects in early neural differentiation. 46C-N154A exhibits stronger suppression of transcripts for Sox1 and Pax6 (Fig. 3D), consistent with the N2B27 differentiation results (Fig. 2D). Conversely, the inhibitory effect of N154A expression diminishes for Nestin, Neurod1 β-III-Tubulin, and Map2 at day D4/7 in KSR medium. Western blotting and immunofluorescence results indicate that 46C-Env/N154A exhibits the similar inhibitory effect compared with WT (Fig. 3E and Supplementary Fig. S2).
Collectively, these findings indicate that Env expression induces the same inhibitory effect in KSR medium as in N2B27 medium, while N154A expression produces a stronger inhibitory effect than Env expression. This inhibition persists throughout the entire process from neural lineage commitment to early neuronal differentiation.
Env of ZIKV inhibits neurodevelopment-related pathways under monolayer neural differentiation
To further investigate the changes in biological processes during N2B27 differentiation, RNA-Seq was performed on D0 and D7 post-differentiation and the results were shown in Fig.436,37. The volcano plot showed 358 differentially expressed genes (DEGs) in 46C-Env compared to WT after 7 days differentiation, in which 190 genes were up-regulated and 168 genes were down-regulated. In particular, the genes with significant down-regulation changes were identified at the top of the volcano plot, suggesting that they may represent the potential key factors involved in physiological responses (Fig. 4A)38. To comprehensively analyze transcriptional perturbations in the ZIKV Env-expressing cells, we employed three complementary bioinformatics approaches: Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis to map differentially expressed genes to specific signaling and metabolic pathways; Gene Ontology (GO) analysis to characterize the macro-level functional attributes of differentially expressed genes; and Gene Set Enrichment Analysis (GSEA) to detect biologically coherent processes exhibiting coordinated changes across the entire gene expression landscape, thereby avoiding the omission of significant signals due to threshold settings.
A Volcano plots show the expression of DEGs in 46C-Env compared with 46C-WT on day 7 after N2B27 differentiation. Up- and down-regulated genes are plotted in red and blue, respectively. B, C KEGG pathway analysis was performed on up- and down-regulated DEGs. D, E GO analysis of the top 10 GO terms for up- and down-regulated DEGs. F GSEA analysis of DEGs enriched in “Alanine Aspartate and Glutamate Metabolism” and “Spinal Cord Association Neuron Differentiation”. G Heatmap showing expression of D0 and D7 DEGs in 46C-Env compared with 46C-WT (n = 3). Each color block in the figure represents the computational result for a single gene in a single sample. The color scale represents expression values standardized by row Z-score.
KEGG pathway analyses were performed on DEGs to identify potential regulatory pathways in the process of neural differentiation affected by Env. KEGG classification showed that “Viral myocarditis”, “p53 signaling pathway” and “Calcium signaling pathway” were enriched in up-regulated pathways (Fig. 4B). Pathways related to amino acid metabolism, neurons and synaptic, including “Alanine, aspartate and glutamate metabolism”, “Synaptic vesicle cycle”, and “Neuroactive ligand-receptor interaction” were enriched for down-regulated genes (Fig. 4C)38.
GO functional annotation to reveal the potential biological role of ZIKV Env in neural differentiation. The top 10 highly enriched GO terms include biological process (BP), cellular component (CC), and molecular function (MF), among which, up-regulated genes in “G protein-coupled receptor activity” and “Transmembrane signaling receptor activity” were enriched (Fig. 4D). For down-regulated genes, the BP clustering terms were mainly enriched with “neurotransmitter loading into synaptic vesicle”, “neurotransmitter biosynthetic process”, “regulation of synapse assembly”, and “positive regulation of dendritic spine morphogenesis”. CC clustering terms are mainly rich in “synaptonemal complex”, “synaptonemal structure”, “condensed nuclear chromosome”, and “nuclear chromosome”, these terms appear to be associated with prophase I of the first meiotic division. MF categories included “Thyroid hormone binding”, “D-aspartate oxidase activity”, and “neuropeptide receptor activity” (Fig. 4E)38. These results suggest that down-regulated genes in 46C-Env are mainly associated with neurotransmitters, meiosis and the synaptonemal complex, and amino acid metabolism.
GSEA was used to further identify Env-affected biological processes in neural differentiation. In 46C-Env cells, most genes were highly enriched in “Alanine Aspartate and Glutamate Metabolism” and “Spinal Cord Association Neuron Differentiation”, both of which were downregulated (Fig. 4F). The heatmap analysis showed that neural development-related genes were activated, and DEGs were enriched in down-regulated genes related to neurotransmitters, synapsis, and amino acid metabolism at D7 differentiation (Fig. 4G)38.
We determined the transcript level of representative DEGs by RT-qPCR (Supplementary Fig. S3A in Supplementary Information), and the results were consistent with RNA-seq. We also validated the expression of Kruppel-like factor 2 (Klf2), developmental pluripotency associated 2 (Dppa2), and Calcium-binding protein 2 (Calb2) by Western blotting (Supplementary Fig. S3B in Supplementary Information). Klf2 participates in regulating NF-κB-mediated immune responses, protects against neural injury, and promotes axonal regeneration39,40. In neonatal rats, Klf2 expression suppresses neuronal apoptosis and alleviates nerve injury41. Klf2 may play a key role in both neurotrauma and axonal growth. We detected Klf2 expression in mESCs expressing Env/N154A after seven days of differentiation in N2B27 medium and found that Klf2 was activated at both the mRNA and protein levels (Supplementary Fig. S3B). Calb2 binds to excess intracellular calcium ions, buffering calcium concentrations and protecting neurons from excitotoxicity42,43. Dysregulation of Calb2 may lead to neuronal damage. In cells expressing Env, Calb2 is significantly inhibited, and N154A expression exerts a more pronounced inhibitory effect on Calb2 (Supplementary Fig. S3B). Our data support the notion that Env expression leads to suppression of neurodevelopment-related genes during early directional differentiation in the neural lineage.
Env of ZIKV inhibits neuronal development and synapse assembly in KSR suspension medium
The transcriptome of 46C-WT and 46C-Env cells was also analyzed during the KSR neural differentiation assay and the results were shown in Fig.537,44. RNA-Seq showed that 902 up-regulated and 403 down-regulated genes were identified at day 4 post-differentiation at D4 (Fig. 5A)45.
A Volcano plots show the expression of DEGs in 46C-Env compared with 46C-WT on day 4 after KSR differentiation. Up- and down-regulated genes are plotted in red and blue, respectively. B, C KEGG pathway analysis was performed on up- and down-regulated DEGs at D4. D, E GO analysis of the top 10 GO terms for up- and down-regulated DEGs at D4. F GSEA analysis of DEGs enriched in “Central Nervous System Neuron Differentiation” and “Synapse Assembly”. G Heatmap showing the expression of D4 DEGs in 46C-Env compared with 46C-WT (n = 3). Each color block in the figure represents the computational result for a single gene in a single sample. The color scale represents expression values standardized by row Z-score.
KEGG pathway analysis showed that “Calcium signaling pathway” and “Efferocytosis” were enriched in the up-regulated pathways on day 4 after differentiation (Fig. 5B). However, ‘Neuroactive ligand-receptor interaction’ and “Axon guidance” were enriched in the down-regulated pathway (Fig. 5C)45. GO analysis showed that for the up-regulated genes, “translation initiation factor activity”, “signaling receptor activity”, and “G protein-coupled receptor activity” were highly enriched (Fig. 5D). For down-regulated genes, the annotated BP, CC, and MF categories were similar to that of the N2B27 differentiation (Fig. 4E). The BP category includes “neuron differentiation” and “nervous system development”. The CC category includes “synaptonemal complex” and “synaptonemal structure”. The MF category includes “DNA-binding transcription factor activity” (Fig. 5E)45. GSEA showed that “Central Nervous System Neuron Differentiation” and “Synapse Assembly” were enriched in the down-regulated pathway (Fig. 5F). Heatmap analysis showed the enrichment of DEGs in down-regulated genes associated with neuronal differentiation and nervous system development after 4 days of differentiation (Fig. 5G)45.
We further validated the expression of representative DEGs by performing RT-qPCR (Supplementary Fig. S4A in Supplementary Information). Wnt7a, a member of the Wnt family, plays an important role in embryonic development and cell fate regulation, and specifically regulates CNS angiogenesis46,47. Grik2 (Glutamate Receptor Ionotropic, Kainate 2), a receptor for the neurotransmitter glutamate, belonging to the kainate receptor subtype48. Kainate receptors are present on both presynaptic and postsynaptic membranes, mediating excitatory synaptic transmission and regulating neurotransmitter release49. Robo2 (Roundabout Guidance Receptor 2) is a repulsive receptor that mediates repulsive signals for axon guidance and cell migration50. We further validated the expression of Cdkn1c and L1td1 (upregulated DEGs), Wnt7a, Grik2, and Robo2 (downregulated DEGs) at the protein level by Western blotting (Supplementary Fig. S4B in Supplementary Information), with results consistent with those obtained by RT-qPCR. These data further indicate that ZIKV envelope protein damages neural development in KSR medium by suppressing neurodevelopment-related genes, downregulating synaptic assembly and neuronal development, while simultaneously activating calcium signaling pathways and G protein-coupled receptors.
Discussion
The ability of ZIKV to invade and persist in the central nervous system has attracted worldwide attention. ZIKV Env is involved in key steps such as fusion of virus-cell membranes, assembly and release of viral particles, and has important functions in virus-induced neuropathogenesis30,51,52. In this study, we revealed that ZIKV Env may contribute to ZIKV-induced neuropathogenesis by inhibiting dendritic spine development, synaptic assembly, and synapsis, while activating G protein-coupled receptors and calcium signaling pathways as shown in this mESCs-based neural differentiation models (Fig. 6).
Normal neural differentiation of mESCs (Top panel) and abnormal differentiation of mESCs expressing ZIKV Env under N2B27 or KSR conditions (Bottom panel).
We expressed ZIKV Env protein stably in mouse embryonic stem cells, overcoming the limitations imposed by viral replication interference on our research. Using two in vitro methods for directed neural differentiation, i.e., N2B27 (monolayer culture) and KSR (suspension culture). We systematically investigated the role of Env from neural lineage commitment to early neuronal differentiation at both two-dimensional and three-dimensional levels. In previous studies, Env was found to cause cell growth inhibition and cycle arrest30,31. However, our study found that stem cells expressing Env retained the ability to differentiate in multiple directions and barely affected cell cycle and proliferation (Fig. 1).
Since apoptosis is triggered during ZIKV infection53, we also examined the apoptosis rate under both differentiation modalities and found that Env and N154A down-regulated apoptosis under N2B27 differentiation (Supplementary Fig. S5A, B in Supplementary Information). We detected activated caspase-3 (cleaved caspase-3) at D7 on N2B27 differentiation (Supplementary Fig. S5C in Supplementary Information). No significant difference was observed in the level of activated caspase-3 (cleaved caspase-3). Simultaneously, Caspase-9 expression was reduced by Env/N154A, while p53 remained unchanged and the Bax/Bcl-2 ratio was slightly increased (Supplementary Fig. S5C). Based on these findings, we hypothesize that the envelope protein inhibits apoptosis by mainly suppressing the Caspase-9-mediated intrinsic pathway. In KSR medium, N154A also significantly protected the cells from apoptosis at D7 differentiation (Supplementary Fig. S5D, E in Supplementary Information), which is consistent with the phenomenon observed during N2B27 differentiation. In addition, multiple cellular inflammatory factors have been detected in the brains of newborns infected with ZIKV, suggesting that ZIKV infection may induce other forms of cell death54. Viruses have evolved immune evasion mechanisms that activate or suppress pyroptosis55. ZIKV was shown to induce brain atrophy by triggering pyroptosis in neural progenitor cells through caspase-1 and gasdermin D (GSDMD) activation56 or induce placental cell pyroptosis by activating gasdermin E (GSDME) both in vivo and in vitro57. We examined pyroptosis in two differentiation modes and found that Env had negligible effects on pyroptosis. However, N154A induced pyroptosis pathway in both N2B27 and KSR media by activating caspase-1, NLRP3, and GSDMD (Supplementary Fig. S6A, B in Supplementary Information). This suggests that the stronger differentiation arrest caused by N154A may be causally associated with the pyroptosis pathway.
mESCs have the ability to differentiate into three embryonic layers (endoderm, mesoderm, and ectoderm) that give rise to all organs58,59. The nervous system is formed by the proliferation and invagination of ectodermal cells59. We showed that Env expression inhibits embryoid bodies and downregulates ectodermal hall markers. N154A expression produced similar results to Env expression in that it significantly inhibited embryoid body formation during EB differentiation. However, N154A expression produced varying degrees of impact on the three-layer markers. ZIKV downregulates cerebellar-related genes19, inhibits human neural stem cell growth, attacks neural progenitor cells60, and impairs neuronal development, leading to impaired neuronal homeostasis61,62. Env expression may serve as a major player as in N2B27 and KSR differentiations as Env expression inhibited the expression of neural stem cells and neuronal marker genes such as sox1, β-III Tubulin, blocked the production of neural rosettes and nerve fibers, and retarded cellular development (Figs. 1–3). Both modes of differentiation confirm that Env expression affects the differentiation of mESCs towards their neural fates. Flow cytometry results showed that Env-expressing cells caught up with WT in terms of Sox1-GFP positive cell numbers at Days 6/7. However, N154A-expressing cells gradually caught up with WT at D8 in N2B27 medium but exhibited a significant downward trend at D8 in KSR medium. This suggests that Env expression may exert differential effects on the two distinct differentiation modes.
N-glycosylation is thought to be required for ZIKV infection, and glycosylation ablation causes mild virulence12,63. Our research has found that N154A exhibits a stronger inhibitory effect than wild-type Env, from neural lineage commitment to early neuronal differentiation. In previous studies, cells transfected with the envelope protein in neuronal cells exhibited neurotoxicity, but the absence of glycosylation sites (N154A) significantly enhanced this toxicity. In neural stem cells, only N154A induced toxicity, the wild-type Env protein did not exhibit toxicity. Glycosylated envelope proteins facilitate efficient viral particle assembly and release. Simultaneously, the Env protein may protects the host cells it infects (neurons and NSCs) through a specific mechanism. Non-glycosylated Env is inefficiently assembled into viral particles, resulting in reduced viral load. Large quantities of unassembled Env protein are abnormally released into the extracellular space, damaging nearby uninfected cells and thereby causing more severe toxicity26. Additionally, we observed that wild-type Env does not induce significant pyroptosis, indicating that the inhibition of neuronal differentiation by Env may not be associated with pyroptosis. Conversely, the N154A mutant induced robust pyroptosis, which provides a mechanistic explanation for the more severe phenotype observed with N154A. This is reminiscent of the studies showing that in newborn mice, glycosylation deficiency can induce inflammatory and necrotic immune responses, thereby enhancing neurotoxicity64. We found that the ZIKV Env protein establishes a protected intracellular environment for viral replication by inhibiting caspase-9-mediated endogenous apoptosis (Supplementary Fig. S5 in Supplementary Information). Non-glycosylated Env protein due to misfolding or abnormal intracellular aggregation, specifically activate the pyroptosis pathway (Supplementary Fig. S6 in Supplementary Information). The potent pyroptotic signal they trigger overrides the anti-apoptotic protective effect, driving cells toward inflammatory lytic death and resulting in enhanced neurotoxicity.
RNA-seq analysis of Env-expressing cells during monolayer neural differentiation at D7 showed that Env reduced the expression levels of neurodevelopment-related genes, including the X-linked lymphocyte regulatory (Xlr) gene family, which are associated with the regulation of dendritic branching, the number and morphology of dendritic spines65, and are important regulators of neuronal function in the mouse brain66. Slc17a6, also known as VGLUT2 (Vesicular Glutamate Transporter 2), plays a key role in the central nervous system. It is primarily responsible for loading glutamate into synaptic vesicles, thereby serving as the primary excitatory neurotransmitter in synaptic transmission67. Kcnc2, the 3.2 protein encoded by KCNC2 (Potassium Voltage-Gated Channel Subfamily C Member 2), is mainly expressed in cortical and hippocampal fast-spiking GABAergic interneurons68. The genes down-regulated by Env were mainly enriched for biological processes related to neurodevelopment. For instance, “neurotransmitter metabolic process”, in which acetylcholine is the main neurotransmitter in the autonomic ganglia and influences synaptic transmission in the peripheral nervous tissue of the central nervous system69. “positive regulation of dendritic spine morphogenesis”, the dendritic spine is the main site of synapse formation between neurons and is associated with memory formation and storage70. “Neurotransmitter loading into synaptic vesicle”, after neurotransmitters are synthesized within the presynaptic neuron, they are loaded into synaptic vesicles (SVs) for storage, awaiting a stimulus signal from the neuron for release and transmission. Information transfer between neurons occurs through the cytosolic action of neurotransmitter-filled synaptic vesicles71. ZIKV infects mature neuronal cells in adult brain tissue and successfully replicates. ZIKV targets brain regions associated with memory function (frontal cortex and hippocampus) in the central nervous system, causing synaptic and memory impairment in immunocompetent mice72. ZIKV also induces significant activation of TNF-α in mouse brains, accompanied by microglial proliferation. Neutralizing TNF-α attenuates microglial proliferation and reduces complement system protein C3, ultimately reversing synaptic damage and memory deficits in mice72. Our results indicate that ZIKV Env disrupts neurodevelopmental processes including synaptic vesicle cycle and regulation of synapse assembly (Figs. 4, 5 and 6), providing mechanistic information for the previous observations72.
Other Env-modulated processes include “Alanine, aspartate and glutamate metabolism” and “D-Amino acid metabolism”. Amino acids are important markers of functional subtypes of cortical neurons and play an important role in development. Synaptic transmission involved in the encoding of sensory information uses amino acid neurotransmitters, glutamate, gamma-aminobutyric acid (GABA) and glycine, as the main neurochemicals73. D-Amino acids are physiological regulators of NPC homeostasis in the developing brain74. ZIKV NS4A downregulates glutamine transporters, altering glutamatergic networks and leading to NPC depletion in zebrafish brains75. Our findings reveal that the Env protein can also influence neural development by regulating amino acid metabolism. Interestingly, we also identified biological pathways associated with the synapsis, such as “synaptonemal structure” and “synaptonemal complex”. Normal proliferation and differentiation of neural progenitor cells depend on precise cell cycle regulation and genomic integrity. ZIKV infection induces DNA damage in neural progenitor cells, leading to unexpected activation of cyclin and triggering mitotic catastrophe76. Abnormal expression of the ‘synapsis complex’ may cause NPC cells to enter an abnormal cell cycle arrest state, leading to the abnormal activation of NPCs. The Env protein not only specifically inhibits the early neural differentiation program but also disrupts cellular genomic expression and cycle regulation, causing cells to lose their ability to differentiate normally.
Similar to two-dimensional differentiation N2B27 cultures, in three-dimensional differentiation KSR cultures, RNA-seq analyses showed that Env significantly reduced the expression of neurodevelopmentally relevant genes on D4. Pax6, the transcription factor, is a determinant of the neuronal differentiation fate of stem cells and ensures the success of the neurogenesis process77. Nnat (neuronal element) is translated peripherally and externally in neuronal dendrites and is associated with synaptic plasticity, which is involved in brain development78. Cxcr4, a G protein-coupled receptor that activates a variety of intracellular signaling pathways79. Interaction of viral envelope proteins with CXCR4 receptors can cause neurotoxicity79,80. Down-regulated genes were significantly enriched in “Neuroactive ligand-receptor interaction”, “Axon guidance”, “neuron differentiation”, “nervous system development”, “Regulation of synapse assembly”, “Neurotransmitter receptor activity”, “Positive regulation of dendritic spine development”, and “synaptonemal complex” (Fig. 5). ZIKV infection causes neurobehavioral and motor deficits in mice81. Neonatal mice infected with ZIKV exhibit delayed startle reflex latency and enhanced pre-pulse inhibition (PPI). Adult mice demonstrate reduced startle reflex amplitude and PPI impairment. Gender differences in startle reflex responses are also observed following adult ZIKV infection81. Our study reveals that the Env protein may influence neural circuit formation by downregulating Axon guidance and synaptic assembly, which may lead to abnormalities in sensorimotor circuits (Figs. 4, 5, and 6). The more severe behavioral deficits observed in neonatal mice compared to adult mice, further indicate ZIKV-induced neurological damage exhibits developmental stage dependency81. ZIKV infection leads to neurodevelopmental abnormalities, and our results further demonstrate that ZIKV Env protein contributes to such abnormalities.
We analyzed the pathways that were co-enriched in DEGs under both N2B27 and KSR differentiation modalities, and found that the co-enriched upregulated pathways and biological functions mainly include “Calcium signaling pathway” and “G-protein-coupled receptor” (Supplementary Fig. S7A–C in Supplementary Information). “Neuroactive ligand-receptor interaction”, “Positive regulation of dendritic spine morphogenesis”, and “Regulation of post synapse organization” were co-down-regulated (Supplementary Fig. S7D–F in Supplementary Information). Here, we propose a hypothesis of “stage-specific” disruption by ZIKV Env affecting neurodifferentiation. The study speculates that Env may suppress neural progenitor cell fate determination (e.g., Sox1 and Pax6) at early stages and interfere with neuronal development (e.g., β-III Tubulin and Map2) at later stages, targeting neurodevelopment-related pathways, dynamically impacting the entire process of neural development. Additionally, it was unexpectedly discovered that Env activates calcium signaling and GPCR pathways, potentially exacerbating neurological dysfunction through abnormal excitability.
ZIKV infection can cause severe neurodevelopmental defects, and ZIKV encodes structural and nonstructural proteins that may also act individually or in combination to cause neurotoxicity. A single amino acid mutation (serine to aspartic acid at position 139) in the pre-membrane protein prM82 and a single amino acid substitution (lysine to arginine substitution at position 101) in coat protein C83 result in exacerbated neurotoxicity. NS2A causes microcephaly by destabilizing the adhesion junction complex (AJ)84. NS2B-NS3 cleave Septin-2 and cause defective cytoplasmic division, impairing the cell cycle and leading to neural progenitor cytotoxicity85. NS4A-NS4B synergistically inhibits the Akt-mTOR pathway, leading to defective neurogenesis and induction of autophagy86. NS5 protein interacts with host proteins at the base of primary cilia in neural progenitor cells to promote ciliopathy and premature differentiation of neural progenitor cells87. ZIKV Env is also neurotoxic. Env alters neural stem cell properties by inducing immature differentiation of pre-neuronal genes in human foetal neural stem cells (fNSCs) and modulating microRNA circuitry27. Envelope proteins of other viruses are also neurotoxic. For example, human immunodeficiency virus 1 (HIV-1) surface glycoprotein (gp120) activates c-Jun N-terminal kinase (JNK) and p42 extracellularly regulated kinase (ERK) to induce apoptosis in neurons and microglia88. The human endogenous retrovirus (HERV) envelope protein is regulated by the TAR (trans-activation reactive) DNA-binding protein 43 (TDP43), which may cause neurodegeneration89. Our research indicates that Env expression alone disrupts differentiation programs through non-cytotoxic pathways, suggesting the potential pathogenic risk of non-infectious viral proteins. Env vaccines or antibody therapies require assessment of neurotoxicity risks to avoid side effects similar to those of HIV gp120.
In conclusion, our study reveals that ZIKV Env significantly inhibits the developmental process from neural lineage commitment to early neuronal differentiation stages independently of viral replication, rather than through conventionally recognized pathways such as apoptosis or proliferation arrest, offering a novel perspective on the teratogenic mechanisms of ZIKV. In this model, Env inhibits neuroactive ligand-receptor interaction, neuron, synaptic processes, and dendritic spine development, but activates Calcium signaling pathway and G protein-coupled receptor signaling pathway, constituting the molecular mechanisms of ZIKV-induced neurodevelopmental defects.
Material and methods
Cells
Mouse embryonic stem cell line (mESC) 46 C (sox1-gfp) is a kind gift from Professor Mingze Yao (Shanxi University) and was incubated at 37 °C in a cell culture incubator containing 5% CO2 and cultured on 0.2% gelatin-coated plates in Dulbecco’s modified Eagle’s high sugar medium (DMEM, Gibco). The medium was supplemented with 20% fetal bovine serum (FBS, SOFRA), 1 mM non-essential amino acids (Gibco), 1 × GlutaMAX (Gibco), 50 units/mL penicillin and 50 μg/mL streptomycin (Solarbio, China), 0.1 mM β-mercaptoethanol (Gibco), 1000 U/mL leukemia inhibitory factor (LIF, Sino Biological Inc.), and 2i inhibitors (3 μM CHIR99021 and 1 μM PD0325901). The culture medium was renewed daily, and the cells were passaged at a density of 70% with trypsin containing 0.25% EDTA (Solarbio, China) at a rate of 1/20-1/50. HEK293T cells were maintained in DMEM supplemented with 10% fetal bovine serum. All cell lines used in this study were tested for mycoplasma by PCR every fortnight and were free of mycoplasma contamination.
Plasmid construction and generation of 46C-Env/N154A expressing cell lines
The plasmid pEGFP-C3-ZIKV-Env contains Env gene of ZIKV isolate ZIKV/H. sapiens/Brazil/Natal/2015 (NCBI ID: NC_035889.1)18. The Env gene was released by EcoRI and BamHI and then inserted into puromycin-resistant pSIN-Flag-ENNSPC vector to produce pSIN-Flag-ZIKV-Env, in which Env gene was under control of EF-1-alpha promoter and bovine growth hormone (bGH) polyadenylation signal. This plasmid was mutated by overlapping PCR to produce pSIN-FLIG-ZIKV-N154A, which encodes a mutant Env containing a point mutation at the amino acid position 154 to eliminate the N-glycosylation site.
Each of pSIN-Flag-ENNSPC, pSIN-Flag-ZIKV-Env, and pSIN-Flag-ZIKV-N154A was co-transfected with psPAX2 and pMD2G into 293T cells to package recombinant lentivirus as described previously90. The mESC 46 C was transduced with the resultant pseudo-lentiviruses in the presence of 2 μL polybrene for 48 h and then screened in the medium containing 1 μg/mL puromycin (Sty551, Beyotime, China) to establish mESCs stably expressing only Flag tag (designated as 46C-WT and used as negative control throughout the experiments), N-terminal Flag-tagged ZIKV Env or Env-N154A (designated as 46C-Env or 46C-N154A, respectively). The established stable cell lines were characterized by Western blotting or RT-qPCR.
RNA isolation and RT‑qPCR
Total RNAs were isolated using RNAiso-plus (Catalog No. 9109, Takara, Japan). One microgram total RNA as template, cDNA synthesis was performed with the HiScript®III RT SuperMix Kit (Catalog No. RR036A, Takara, Japan). Three-replicate PCR amplifications were conducted on the CFX Connect system (Bio-Rad) using the SYBR Green qPCR Master Mix Kit (Me5 Biotechnology Co., Ltd.). RT-qPCR reactions were performed under the following conditions: initial denaturation at 95 °C for 30 s, followed by 39 cycles (95 °C for 10 s, 60 °C for 10 s, 72 °C for 30 s). β-actin served as the internal reference gene, and relative expression levels of target genes were calculated using the ΔΔCT method. Primer sequences are detailed in Supplementary Information (Supplementary Table S1).
Immunofluorescence staining
Cells were washed three times with PBS for 5 min each time, fixed with 4% paraformaldehyde for 15 min at 4 °C, permeabilized with 0.3% TritonX-100 for 10 min, and blocked with 10% goat serum for 1 h at room temperature. Cells were then incubated overnight at 4 °C with a primary antibody in the presence of 0.1% TritonX-100, 1% goat serum, 1% BSA. After three washes of 5 min each, cells were stained with secondary antibody for 2 h at room temperature in the dark. Subsequently, cells were stained with 4′,6-diamidino-2-phenylindole (DAPI) (1 μg/mL, 10 min), visualized with a confocal laser scanning microscope (Zeiss LSM 710). Primary antibodies used in immunofluorescence staining included anti-OCT4-Mouse-Monoclonal (1:200, Santa Cruz Biotechnology, USA, Cat No. sc-5279), anti-NANOG-Rabbit-Polyclonal (1:500, Novus Biologicals, USA, Cat No. NB100-58842), and anti-β-III tubulin-Mouse-Monoclonal (1:200, ABclonal, China, Cat No. A18132), Alexa Fluor 594-conjugated anti-Mouse IgG (H + L) (1 : 1000, Proteintech, Cat No. RGAM004), and Alexa Fluor 594-conjugated anti-Rabbit IgG (H + L) (1 : 1000, Proteintech, Cat No. RGAR004).
Cell cycle analysis
The mESCs were cultured to 70% density in 6-well plates, digested into single cells with 0.25% trypsin-EDTA (Solarbio, China), and then washed with Dulbecco’s Phosphate Buffered Saline (DPBS). Approximately 5 × 105 cells were fixed in 70% ethanol at 4 °C for 24 h, washed twice with DPBS (Hyclone), stained with 500 μl of PI staining reagent (containing 20 × propidium iodide, 50 × RNase), and incubated at 37 °C for 30 min in the dark. The cell cycle was measured using a flow cytometer (BECKMAN COULTER) and the raw data were analyzed using FlowJo 10.8.1 and GraphPad Prism.
Cell proliferation assay
2000 mESCs cells/well were seeded in 96-well culture plates. At 24, 48, 72 and 96 h of incubation, 10 μL of CCK8 (CCK8, Beyotime, China) was added to 100 μL of culture medium at 37 °C for 2 h in the dark. Absorbance at 450 nm was recorded using a Synergy H1MD plate reader (BioTek).
Formation of embryoid bodies (EBs)
A suspension drop (1000 cells per drop) is covered with a 150 mm dish for culturing the cells, and PBS is added to the dish to allow formation of EBs in differentiation medium. the medium is free of LIF and 2i inhibitors. After 4 days, the EBs are transferred to a 100 mm dish for culture in differentiation medium. Cell morphology was observed under a fluorescence microscope (Zeiss Axio Imager). Total RNA was then extracted and the expression of the three germline marker genes was detected by RT-qPCR. All experiments were independently repeated at least for three times.
N2B27 monolayer neural differentiation
Prior to monolayer neural differentiation induction, mESC were cultured for three generations in N2B27 medium with 2i and LIF inhibitors (N2B27 medium supplemented with 1 × sodium pyruvate, 1000 U/mL LIF, 3 μM CHIR99021 and 1 μM PD0325901) to exclude FBS. mESCs were cultured in 12-well plates coated with 0.4% gelatin overnight, and then N2B27 Neural Differentiation Medium was added. N2B27 Neural Differentiation Medium contained 50% DMEM/F12 (Sevenbio, Beijing, China), 0.5% N2 (Gibco), 50% Neurobasal (Gibco), 1% B27 (Gibco), 1 mM NEAA (Gibco), 1 × GlutaMAX (Gibco), and 0.1 mM β-mercaptoethanol (Gibco). The medium was replaced freshly daily. The RNA and protein samples were prepared at the indicated time points for further analysis using RT-qPCR or Western blotting.
KSR differentiation
Two hundred fifty thousand cells of mESCs were cultured in 6 cm diameter plates in KSR differentiation medium GMEM (Gibco) containing 7% Knockout Serum Replacement (Gibco), 1 mM NEAA, 1 × GlutaMAX, 1 mM sodium pyruvate and 0.1 mM β-mercaptoethanol (Gibco). The medium was changed every two days. The RNA and protein samples were prepared at the indicated time points for further analysis using RT-qPCR or Western blotting.
Flow cytometry
Cells were digested into single cells with 0.25% trypsin-EDTA, resuspended in PBS, and then the SOX1-GFP+ cells were sorted and analyzed using flow cytometry (BD LSRForessaTMX-20). Data were analyzed using FlowJo 10.8.1 software. The gating strategy is as follows: First, delineating the target cell population (P1) in the FSC-A/SSC-A scatter plot. Subsequently, screening single cells (P2) from P1 in the FSC-A/FSC-H scatter plot to exclude adherent cells. The proportion of GFP-positive cells was determined as follows: on the basis of histogram of fluorescence intensity in the FITC channel (detecting GFP) from untransfected GFP-negative control cells, a threshold is set (typically placing >99% of negative cells to the left of the threshold). Subsequently, the percentage of cells with fluorescence intensity exceeding this threshold is calculated within the P2 gate of the experimental group.
Western blotting
Cells are lysed in 1 × Laemmli buffer and then denatured at 100 °C for 10 min. Ten micrograms of total protein (whole cell lysates) were separated in 10–15% SDS-PAGE, transferred to polyvinylidene difluoride (PVDF) membranes, blocked with 5% skimmed milk in PBS. The membranes were incubated with primary antibodies overnight at 4 °C in PBS containing 1% skimmed milk, washed for three times using TBST, incubated with HRP-conjugated anti-mouse or anti-rabbit secondary antibodies for 1 h at room temperature. The protein bands were visualized using the enhanced chemiluminescent (ECL) system (Perkin Elmer) and then analyzed using ChemoStar 6.0 Imaging Software (INTAS).
Primary antibodies used in Western blotting include anti-DYKDDDDK-Tag-Mouse-Monoclonal (1:2000, Proteintech, Cat No. 66008-4-Ig), anti-PAX6-Rabbit-Polyclonal (1:1000, Proteintech, Cat No. 12323–1-AP), anti-β-III-Tubulin-Mouse-Monoclonal (1:2000, Abclonal, Cat No. A18132), anti-Cleaved Caspase3-Rabbit-Polyclonal (1:1000, Proteintech, Cat No. 25128-1-AP), anti- Caspase 9/P35-Rabbit-Polyclonal (1:300, Proteintech, Cat No. 10380-1-AP), anti-Phospho-P53(Ser15)-Rabbit-Polyclonal (1:2000, Proteintech, Cat No. 28961-1-AP), anti-Bax-Mouse-Polyclonal (1:10000, Proteintech, Cat No. 60283-2-Ig), anti-Bcl2-Rabbit-Polyclonal (1:500, Proteintech, Cat No. 26593-1-AP), anti-NLRP3-Rabbit-Polyclonal (1:1000, abcam, Cat No. ab4207), anti-GSDMD-Rabbit-Monoclonal (1:1000, abcam, Cat No. ab209845), anti-Caspase1-Rabbit-Monoclonal (1:1000, abcam, Cat No. ab207802), anti-KLF2-Rabbit-Polyclonal (1:200, Signalway Antibody, Cat No. 46591), anti-DPPA2-Rabbit-Polyclonal (1:1000, Signalway Antibody, Cat No. 62249), anti-Calretinin-Rabbit-Polyclonal (1:500, EpiZyme, Cat No. P011307), anti-p57 Kip2-Rabbit-Monoclonal (1:500, Zenbio, Cat No. R380868), anti-L1TD1-Rabbit-Polyclonal (1:1000, Signalway Antibody, Cat No. 62248), anti-WNT7A-Rabbit-Polyclonal (1:1000, Signalway Antibody, Cat No. #38653-1), anti-GRIK2-Rabbit-Monoclonal (1:500, Zenbio, Cat No. R389225), anti-ROBO2-Rabbit-Polyclonal (1:1000, Proteintech, Cat No. 21635-1-AP), anti-β-actin-Rabbit-Polyclonal (1:2000, Proteintech, Cat No: 20536-1-AP), anti-GAPDH-Mouse-Polyclonal (1:5000, EpiZyme, Cat No. LF205). The secondary antibodies include horseradish peroxidase (HRP)-conjugated Goat anti-Rabbit IgG (H + L) (1:2000, Abclonal, Cat No. A5014) or HRP-conjugated Goat anti-Mouse IgG (H + L) (1:5000, Proteintech, Cat No. SA0000-1-1). Uncropped Western blots were shown in the Supplementary Information (Uncropped Western blots).
Apoptosis
Cells were treated with trypsin and then transferred to clean 1.5 ml microtubes. The collected cells were stained with APC and 7-AAD staining solution according to the manufacturer’s instructions (Annexin V-APC/7-AAD Apoptosis Kit, Elabscience, Cat. No. AK17929). Cells were selected based on FSC and SSC and the gating of cell population was determined by healthy control cells. Ten thousand cells per sample were analyzed with a flow cytometer (BECKMAN COULTER). Double-negative were healthy cells, Annexin V-APC single-positive cells were early apoptotic cells, AnnexinV-APC and 7-AAD double-positive cells were necrotic or late apoptotic cells, and 7-AAD single-positive cells were nude nucleated cells. Data were analyzed by CytExpert software program.
RNA sequencing
RNA-seq was performed by Shanghai Applied Protein Technology (Co., Ltd., Shanghai, China). RNA from each sample was extracted using TRIzol. RNA concentration was determined, RNA integrity was assessed, and then the cDNA libraries were prepared and sequenced on the MGISEQ-T7 platform. Volcano and heatmaps were used to show genes with statistically significant differential expression. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were used to find the differentially enriched pathways and biological processes. In addition, we used Gene Set Enrichment Analysis (GSEA) for gene enrichment analysis. Bioinformatic analysis was performed using the OECloud tools at https://cloud.oebiotech.com and https://www.bioinformatics.com.cn, online platform for data analysis and visualization91,92.
Statistical analyses
All data are statistical results of at least three independent experiments and are expressed as mean ± SD. Statistical analysis was performed using GraphPad Prism 9 software (GraphPad Software, La Jolla, CA, USA) for one-way analysis of variance (ANOVA). In all experiments, ns represents statistical non-significance.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are openly available in figshare at https://doi.org/10.6084/m9.figshare.2881490938; https://doi.org/10.6084/m9.figshare.2881490645; https://doi.org/10.6084/m9.figshare.2892983944; https://doi.org/10.6084/m9.figshare.2892987836; and https://doi.org/10.6084/m9.figshare.3108316037. The uncropped and unprocessed original images for all Western blots shown in this study are seen in the Supplementary Information file (Uncropped Western Blots). The recombinant plasmid generated in this study is derived from fully sequenced and publicly available components (backbone: Addgene #342963; insert: GenBank accession number: NC_035889.1). The physical plasmid is available from the corresponding author upon request under a standard Material Transfer Agreement (MTA).
References
Dick, G. W., Kitchen, S. F. & Haddow, A. J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520 (1952).
Lanciotti, R. S. et al. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 14, 1232–1239 (2008).
Faria, N. R. et al. Zika virus in the Americas: early epidemiological and genetic findings. Science 352, 345–349 (2016).
Aid, M. et al. Zika virus persistence in the central nervous system and lymph nodes of rhesus monkeys. Cell 169, 610–620.e614 (2017).
Christian, K. M., Song, H. & Ming, G. L. Pathophysiology and mechanisms of Zika virus infection in the nervous system. Annu. Rev. Neurosci. 42, 249–269 (2019).
Freitas, D. A. et al. Congenital Zika syndrome: a systematic review. PloS ONE 15, e0242367 (2020).
Hasan, S. S., Sevvana, M., Kuhn, R. J. & Rossmann, M. G. Structural biology of Zika virus and other flaviviruses. Nat. Struct. Mol. Biol. 25, 13–20 (2018).
Shi, Y. & Gao, G. F. Structural Biology of the Zika Virus. Trends Biochem. Sci. 42, 443–456 (2017).
Guo, M., Hui, L., Nie, Y., Tefsen, B. & Wu, Y. ZIKV viral proteins and their roles in virus-host interactions. Sci. China Life Sci. 64, 709–719 (2021).
Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta Mol. Cell Res. 1833, 2430–2437 (2013).
May, M. & Relich, R. F. A comprehensive systems biology approach to studying Zika virus. PloS ONE 11, e0161355 (2016).
Carbaugh, D. L., Baric, R. S. & Lazear, H. M. Envelope protein glycosylation mediates Zika virus pathogenesis. J. Virol. 93, https://doi.org/10.1128/jvi.00113-19 (2019).
Jorgacevski, J. et al. ZIKV strains differentially affect survival of human fetal astrocytes versus neurons and traffic of ZIKV-laden endocytotic compartments. Sci. Rep. 9, 8069 (2019).
Ramos da Silva, S., Cheng, F., Huang, I. C., Jung, J. U. & Gao, S. J. Efficiencies and kinetics of infection in different cell types/lines by African and Asian strains of Zika virus. J. Med. Virol. 91, 179–189 (2019).
Gavegnano, C. et al. Jak inhibitors modulate production of replication-competent Zika virus in human Hofbauer, trophoblasts, and neuroblastoma cells. Pathog. Immun. 2, 199–218 (2017).
Ming, G. L., Tang, H. & Song, H. Advances in Zika virus research: stem cell models, challenges, and opportunities. Cell Stem Cell 19, 690–702 (2016).
Garcez, P. P. et al. Zika virus impairs growth in human neurospheres and brain organoids. Science 352, 816–818 (2016).
Mlakar, J. et al. Zika virus associated with microcephaly. N. Engl. J. Med. 374, 951–958 (2016).
Li, C. et al. Zika virus disrupts neural progenitor development and leads to microcephaly in mice. Cell Stem Cell 19, 120–126 (2016).
Vieira, M. S. et al. Neural stem cell differentiation into mature neurons: mechanisms of regulation and biotechnological applications. Biotechnol. Adv. 36, 1946–1970 (2018).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Zhang, F., Pang, C., Zhu, H. & Chen, Y. Timely stimulation of early embryo promotes the acquisition of pluripotency. Cytom. Part A J. Int. Soc. Anal. Cytol. 101, 682–691 (2022).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Wongpaiboonwattana, W. & Stavridis, M. P. Neural differentiation of mouse embryonic stem cells in serum-free monolayer culture. J. Vis. Exp. e52823, https://doi.org/10.3791/52823 (2015).
Watanabe, K. et al. Directed differentiation of telencephalic precursors from embryonic stem cells. Nat. Neurosci. 8, 288–296 (2005).
Steiner, J. P. et al. Neurotoxic properties of the Zika virus envelope protein. Exp. Neurol. 367, 114469 (2023).
Bhagat, R. et al. Zika virus E protein alters the properties of human fetal neural stem cells by modulating microRNA circuitry. Cell Death Differ. 25, 1837–1854 (2018).
Aubert, J. et al. Screening for mammalian neural genes via fluorescence-activated cell sorter purification of neural precursors from Sox1-gfp knock-in mice. Proc. Natl. Acad. Sci. USA Suppl 1. 100, 11836–11841 (2003).
Cheng, M. L. et al. Pathogenicity and structural basis of Zika variants with glycan loop deletions in the envelope protein. J. Virol. 96, e0087922 (2022).
Liu, J. et al. Zika virus envelope protein induces G2/M cell cycle arrest and apoptosis via an intrinsic cell death signaling pathway in neuroendocrine PC12 cells. Int. J. Biol. Sci. 14, 1099–1108 (2018).
Souza, B. S. et al. Zika virus infection induces mitosis abnormalities and apoptotic cell death of human neural progenitor cells. Sci. Rep. 6, 39775 (2016).
Brickman, J. M. & Serup, P. Properties of embryoid bodies. Wiley Interdiscip. Rev. Dev. Biol. 6, https://doi.org/10.1002/wdev.259 (2017).
Calvet, G. et al. Detection and sequencing of Zika virus from amniotic fluid of fetuses with microcephaly in Brazil: a case study. Lancet Infect. Dis. 16, 653–660 (2016).
Ying, Q. L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat. Biotechnol. 21, 183–186 (2003).
Furukawa, Y. et al. Effects of knockout serum replacement on differentiation of mouse-induced pluripotent stem cells into odontoblasts. Bull. Tokyo Dent. Coll. 63, 75–83 (2022).
Xing, L. & Ma, Z.-H. N2B27 neural differentiation of mouse embryonic stem cells expressing ZlKV envelope protein. Figshare, https://doi.org/10.6084/m9.figshare.28929878 (2026).
Xing, L. Expression of ZlKV envelope protein in mouse embryonic stem cells. Figshare, https://doi.org/10.6084/m9.figshare.31083160 (2026).
Xing, L. ZIKV_ENV_N2B27_Dataset.xlsx, figshare, V2, https://doi.org/10.6084/m9.figshare.28814909 (2026).
Jha, P. & Das, H. KLF2 in Regulation of NF-κB-mediated immune cell function and inflammation. Int. J. Mol. Sci. 18, https://doi.org/10.3390/ijms18112383 (2017).
Wu, F. & Li, C. KLF2 up-regulates IRF4/HDAC7 to protect neonatal rats from hypoxic-ischemic brain damage. Cell Death Discov. 8, 41 (2022).
Wang, Q. et al. Klf2-Vav1-Rac1 axis promotes axon regeneration after peripheral nerve injury. Exp. Neurol. 343, 113788 (2021).
Schwaller, B. The use of transgenic mouse models to reveal the functions of Ca2+ buffer proteins in excitable cells. Biochim. Biophys. Acta 1820, 1294–1303 (2012).
Maksimova, M. A. et al. Interneuron functional diversity in the mouse accessory olfactory bulb. eNeuro 6, https://doi.org/10.1523/eneuro.0058-19.2019 (2019).
Ma, Z. H. & Xing, L. KSR neural differentiation of mouse embryonic stem cells expressing ZIKV envelope protein, figshare, https://doi.org/10.6084/m9.figshare.28929839 (2026).
Xing, L. ZIKV_ENV_KSR_Dataset.xlsx, figshare, V2, https://doi.org/10.6084/m9.figshare.28814906 (2026).
Liebner, S. et al. Wnt/beta-catenin signaling controls development of the blood-brain barrier. J. Cell Biol. 183, 409–417 (2008).
Daneman, R. et al. Wnt/beta-catenin signaling is required for CNS, but not non-CNS, angiogenesis. Proc. Natl. Acad. Sci. USA 106, 641–646 (2009).
Stolz, J. R. et al. Clustered mutations in the GRIK2 kainate receptor subunit gene underlie diverse neurodevelopmental disorders. Am. J. Hum. Genet. 108, 1692–1709 (2021).
Nomura, T. et al. A pathogenic missense mutation in kainate receptors elevates dendritic excitability and synaptic integration through dysregulation of SK channels. J. Neurosci. 43, 7913–7928 (2023).
López-Bendito, G. et al. Robo1 and Robo2 cooperate to control the guidance of major axonal tracts in the mammalian forebrain. J. Neurosci. 27, 3395–3407 (2007).
Zhao, F. et al. Extracellular vesicles from Zika virus-infected cells display viral E protein that binds ZIKV-neutralizing antibodies to prevent infection enhancement. Embo J. 42, e112096 (2023).
Dai, L. et al. Structures of the Zika virus envelope protein and its complex with a flavivirus broadly protective antibody. Cell Host Microbe 19, 696–704 (2016).
Turpin, J. et al. Apoptosis during ZIKA virus infection: too soon or too late? Int. J. Mol. Sci. 23, https://doi.org/10.3390/ijms23031287 (2022).
de Sousa, J. R. et al. In situ inflammasome activation results in severe damage to the central nervous system in fatal Zika virus microcephaly cases. Cytokine 111, 255–264 (2018).
Zhang, Y. et al. Pyroptosis, a double-edged sword during pathogen infection: a review. Cell Death Discov. 11, 289 (2025).
He, Z. et al. Neural progenitor cell pyroptosis contributes to Zika virus-induced brain atrophy and represents a therapeutic target. Proc. Natl. Acad. Sci. USA 117, 23869–23878 (2020).
Zhao, Z. et al. Zika virus causes placental pyroptosis and associated adverse fetal outcomes by activating GSDME. eLife 11, https://doi.org/10.7554/eLife.73792 (2022).
Hadas, R. et al. Temporal BMP4 effects on mouse embryonic and extraembryonic development. Nature 634, 652–661 (2024).
Nakanoh, S. Exploring early extraembryonic cells of epiblast origin: questions on human amniotic ectoderm and extraembryonic mesoderm. Dev. Biol. 524, 80–86 (2025).
Tang, H. et al. Zika virus infects human cortical neural progenitors and attenuates their growth. Cell Stem Cell 18, 587–590 (2016).
Nielsen-Saines, K. et al. Delayed childhood neurodevelopment and neurosensory alterations in the second year of life in a prospective cohort of ZIKV-exposed children. Nat. Med. 25, 1213–1217 (2019).
Stokes, C. et al. The human neural cell atlas of Zika virus infection in developing brain tissue. Cell reports. Medicine 6, 102189 (2025).
Annamalai, A. S. et al. Zika virus encoding nonglycosylated envelope protein is attenuated and defective in neuroinvasion. J. Virol. 91, https://doi.org/10.1128/jvi.01348-17 (2017).
Guo, Y. et al. The ablation of envelope protein glycosylation enhances the neurovirulence of ZIKV and cell apoptosis in newborn mice. J. Immunol. Res. 2021, 5317662 (2021).
Di Segni, M. et al. Xlr4 as a new candidate gene underlying vulnerability to cocaine effects. Neuropharmacology 168, 108019 (2020).
Sideromenos, S. et al. The metabolic regulator USF-1 is involved in the control of affective behaviour in mice. Transl. Psychiatry 12, 497 (2022).
Wu, J. Y. et al. Regulation of states of consciousness by supramammillary nucleus glutamatergic neurones during sevoflurane anaesthesia in mice. Br. J. Anaesth. https://doi.org/10.1016/j.bja.2024.10.023 (2024).
Li, L. et al. Investigation of novel de novo KCNC2 variants causing severe developmental and early-onset epileptic encephalopathy. Seizure 101, 218–224 (2022).
Konishi, S., Tsunoo, A. & Otsuka, M. Enkephalins presynaptically inhibit cholinergic transmission in sympathetic ganglia. Nature 282, 515–516 (1979).
Aguilar-Hernandez, L. et al. Memory and dendritic spines loss, and dynamic dendritic spines changes are age-dependent in the rat. J. Chem. Neuroanat. 110, 101858 (2020).
Gasnier, B. The loading of neurotransmitters into synaptic vesicles. Biochimie 82, 327–337 (2000).
Figueiredo, C. P. et al. Zika virus replicates in adult human brain tissue and impairs synapses and memory in mice. Nat. Commun. 10, 3890 (2019).
Kalloniatis, M. & Tomisich, G. Amino acid neurochemistry of the vertebrate retina. Prog. Retin Eye Res. 18, 811–866 (1999).
Semenza, E. R. et al. D-cysteine is an endogenous regulator of neural progenitor cell dynamics in the mammalian brain. Proc. Natl. Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2110610118 (2021).
Sow, A. A., Jamadagni, P., Scaturro, P., Patten, S. A. & Chatel-Chaix, L. A zebrafish-based in vivo model of Zika virus infection unveils alterations of the glutamatergic neuronal development and NS4A as a key viral determinant of neuropathogenesis. PLoS Pathog. 20, e1012756 (2024).
Rychlowska, M., Agyapong, A., Weinfeld, M. & Schang, L. M. Zika virus induces mitotic catastrophe in human neural progenitors by triggering unscheduled mitotic entry in the presence of DNA damage while functionally depleting nuclear PNKP. J. Virol. 96, e0033322 (2022).
Manuel, M. N., Mi, D., Mason, J. O. & Price, D. J. Regulation of cerebral cortical neurogenesis by the Pax6 transcription factor. Front. Cell Neurosci. 9, 70 (2015).
Joseph, R. M. Neuronatin gene: Imprinted and misfolded: Studies in Lafora disease, diabetes and cancer may implicate NNAT-aggregates as a common downstream participant in neuronal loss. Genomics 103, 183–188 (2014).
Lee, C. et al. Macrophage activation through CCR5- and CXCR4-mediated gp120-elicited signaling pathways. J. Leukoc. Biol. 74, 676–682 (2003).
Williams, K. S., Seawell, J. A., Zhuravleva, V., Pierre, K. & Meeker, R. B. Cooperative interactions between neurotrophin receptors and CXCR4 regulate macrophage phenotype and susceptibility to activation by HIV. J. Neurovirol. 30, 406–422 (2024).
Souza, I. N. O. et al. Different outcomes of neonatal and adult Zika virus infection on startle reflex and prepulse inhibition in mice. Behav. Brain Res. 451, 114519 (2023).
Yuan, L. et al. A single mutation in the prM protein of Zika virus contributes to fetal microcephaly. Science 358, 933–936 (2017).
Song, G. Y. et al. A single amino acid substitution in the capsid protein of Zika virus contributes to a neurovirulent phenotype. Nat. Commun. 14, 6832 (2023).
Yoon, K. J. et al. Zika-virus-encoded NS2A disrupts mammalian cortical neurogenesis by degrading adherens junction proteins. Cell Stem Cell 21, 349–358.e346 (2017).
Li, H. et al. Zika virus protease cleavage of host protein septin-2 mediates mitotic defects in neural progenitors. Neuron 101, 1089–1098.e1084 (2019).
Liang, Q. et al. Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 19, 663–671 (2016).
Saade, M. et al. Multimerization of Zika virus-NS5 causes ciliopathy and forces premature neurogenesis. Cell Stem Cell 27, 920–936.e928 (2020).
Lannuzel, A. et al. Human immunodeficiency virus type 1 and its coat protein gp120 induce apoptosis and activate JNK and ERK mitogen-activated protein kinases in human neurons. Ann. Neurol. 42, 847–856 (1997).
Li, W. et al. Human endogenous retrovirus-K contributes to motor neuron disease. Sci. Transl. Med. 7, 307ra153 (2015).
Liu, J. et al. Construction and identification of recombinant HEK293T cell lines expressing non-structural protein 1 of Zika virus. Int. J. Med. Sci. 14, 1072–1079 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE 18, e0294236 (2023).
Acknowledgements
The authors sincerely thank Mr. Tinglin Ren for his technical guidance and critical comments on cell culture, neural differentiation techniques, and flow cytometer operation. The authors also appreciate the staff at Shanghai Applied Protein Technology Co., Ltd. for their technical support for transcriptomic analysis. This work was supported by the Program of Graduate Innovation Research of Shanxi Province, China (2024KY119) and the Programme of Introducing Talents of Discipline to Universities (D21004).
Author information
Authors and Affiliations
Contributions
Zi-Hui-Ma: Conceptualization, investigation, analysis, visualization, writing–original draft, writing–editing. Yan Wang: investigation, analysis. Nahla Ahmed Hassaan: Conceptualization, investigation, analysis. Xia-Nan Chu: Resources. Pei-Hua Wang: Resources. Changxin Wu: Resources. Li Xing: Conceptualization, supervision, project administration, funding acquisition, writing–editing. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Raiane Oliveira Ferreira, Newton G. Castro, Isis de Oliveira Souza and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary handling editors: Shani Stern and Benjamin Bessieres. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, ZH., Wang, Y., Hassaan, N.A. et al. ZIKV envelope protein is a strong blocker of early directional differentiation in the neural lineage. Commun Biol 9, 395 (2026). https://doi.org/10.1038/s42003-026-09672-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09672-1