Abstract
African swine fever (ASF) is a highly contagious viral disease caused by the African swine fever virus (ASFV), which primarily affects pigs. ASFV encodes a variety of proteins that contribute to immune evasion, with the mechanisms of immune escape being diverse, complex, and not yet fully understood. In this study, the MGF 505–3R protein of ASFV was identified as a potential inhibitor of the host’s inflammatory response. We demonstrate that MGF 505–3R suppresses the host antiviral response by promoting the ubiquitin–mediated degradation of MyD88, with the amino acid region 89–277 being essential for this function. Notably, this region directly mediates the interaction with MyD88 and induces its ubiquitination. Furthermore, MGF 505–3R and its derived peptide significantly inhibit the production of type I (IFN–α/β) and type III (IFN–λ) interferons, in addition to impairing NF–κB activation by blocking p65 phosphorylation and nuclear translocation. The MGF 505–3R peptide effectively attenuates the host inflammatory storm, decreasing the expression of cytokines such as TNF–α and IL–1β, and alleviating DSS–induced colitis in male C57BL/6 mice. These findings highlight the dual role of MGF 505–3R in suppressing both inflammatory and interferon pathways, underscoring its potential as a therapeutic candidate for inflammatory diseases and a target for antiviral strategies.
Similar content being viewed by others
Introduction
ASF is an infectious swine disease caused by ASFV. The disease is characterized by rapid onset, high fever, and hemorrhagic lesions1. ASF was first identified in Africa in the early 1920s and began to spread globally in 20072,3. In August 2018, a major outbreak occurred in China, further escalating the challenges of disease prevention and control4. Control of the disease is only possible through strict biosecurity measures in affected areas, which has led to the death or culling of large numbers of pigs, resulting in substantial economic losses5. Due to the high mortality rate of ASF and its impact on the global pork supply chain, researchers are urgently seeking to understand the virus’s pathogenic mechanisms and find more effective control strategies. Investigating how ASFV evades the host immune response, particularly through the inhibition of immune pathways, has become one of the key areas of research.
ASFV belongs to the Asfarviridae family and is a large double-stranded DNA virus6,7. The ASFV genome, classified into 23 genotypes, encodes 150–167 open reading frames (ORFs) that produce over 160 proteins, including 54 structural proteins and more than 100 non-structural proteins8,9,10. These viral proteins exhibit diverse functions and regulate the host immune system through various mechanisms, enabling ASFV to evade immune clearance10. As a result, the immune evasion mechanisms of ASFV are highly complex, leaving significant gaps in current research. Notably, ASFV can evade immune responses by suppressing the host NF–κB signaling pathway, which has become a key focus in ongoing ASFV research.
The NF–κB signaling pathway plays a crucial role in the host immune system, regulating various immune responses, including inflammation, immune cell activation, and cell survival11,12. The activation of this pathway typically occurs in response to pathogen infection or damage signals, promoting the expression of immune factors and enhancing the host’s immune response. However, increasing evidence suggests that ASFV can suppress the NF–κB pathway through different mechanisms, inhibiting inflammation and immune factor production, thus creating an environment conducive to viral replication and survival13,14,15.
The variation in the length of the ASFV genome is primarily attributed to differences in the copy number of multigene families (MGFs). Variations in these regions, including the gain or loss of the p22 gene family, are often associated with genomic differences and contribute to the formation of distinct isolates16. Studies have shown that the deletion of MGF 360/505 significantly reduces the virulence of ASFV, making the gene-deficient vaccine a promising candidate17,18. MGF 505–7R inhibits NF–κB activation by interacting with IKKα in the IKK complex and binds to NLRP3, preventing the formation of inflammasome19. Further studies have demonstrated that MGF 505–7R upregulates the expression of the E3 ubiquitin ligase RNF125 and inhibits Hes5 expression, leading to the degradation of JAK1 and JAK220. Moreover, MGF 360–9L interacts with STAT1 and STAT2, inhibiting IFN–β signaling by degrading both STAT1 and STAT221. MGF 360–12L suppresses IFN–I production by disrupting NF–κB nuclear translocation through competitive inhibition of its interaction with importin α proteins22. These findings suggest that ASFV employs multiple strategies targeting various components of the host immune system, which are critical for immune evasion and persistent infection. However, there is still limited research on how ASFV inhibits NF–κB signaling. Therefore, the precise mechanism of NF–κB inhibition by ASFV remains to be fully elucidated, a key area for the development of effective ASF treatments.
MGF 505–3R is a gene within this family, encoding an 843-bp sequence that produces a 30.91 kDa protein, and is one of the members of the MGF 505 multigene family. While it has been reported that MGF 505–3R degrades TBK1 and inhibits the cGAS–STING pathway23, its potential role in modulating the TLR–MyD88–NF–κB axis and interferon responses remains unclear. This study aimed to screen and identify ASFV proteins involved in the inhibition of host innate immunity, with a focus on MGF 505–3R. We discovered that MGF 505–3R not only downregulated the NF–κB pathway but also broadly suppressed type I and III interferon production. Furthermore, we identified a key functional domain (aa 89–277) responsible for its interaction with and ubiquitin-mediated degradation of MyD88. Notably, a derived peptide (pep3R-1) based on this domain retained potent anti-inflammatory and antiviral inhibitory activities, as validated in both inflammatory and viral infection models. These findings provide novel insights into the multifunctional role of MGF 505–3R in immune evasion and highlight its potential as a candidate for developing peptide-based immunomodulators.
Result
ASFV MGF 505–3R protein significantly inhibits NF–κB signaling activity
To investigate how ASFV evades host innate immunity, we overexpressed MyD88 to activate the TLR–NF–κB signaling pathway and constructed a series of ASFV proteins (Fig. S1) to examine their effects on this pathway. Initially, we found that several viral proteins inhibited the TLR–NF–κB pathway, with MGF 505–3R displaying a notable, dose–dependent effect (Fig. 1A, B). To further explore the role of MGF 505–3R in the NF–κB signaling pathway, the effect of MGF 505–3R on TLR–mediated activation of the NF–κB promoter was assessed using a dual luciferase reporter assay. Activation of the NF–κB promoter by Pam3CSK4 (TLR2), LPS (TLR4), and R848 (TLR7) was evaluated in cells overexpressing ASFV MGF 505–3R. We observed that overexpression of the ASFV MGF 505–3R protein significantly inhibited TLR2/4/7 ligand–induced activation of the NF–κB promoter (Fig. 1C), as well as LPS-mediated activation of the NF–κB promoter in PK–15 cells (Fig. 1D). These findings suggest that MGF 505–3R extensively inhibits TLR ligand-mediated NF–κB promoter activation, likely through its interaction with MyD88 and downstream signaling factors.
A HeLa cells were transfected with pGL4.32–NF–κB–Luc along with either an empty vector, pcDNA3.1–Flag–MyD88, and/or pCMV–myc–MGF 505–3R for 24 h, after which luciferase activity was measured (mean ± S.D., n = 4, *p < 0.05, **p < 0.01). B The experiment was performed as described in (A), but with increasing concentrations of pCMV–myc–MGF 505–3R (mean ± S.D., n = 4, **p < 0.01). C HeLa cells were transfected with pGL4.32–NF–κB–Luc and either an empty vector or pCMV–myc–MGF 505–3R for 24 h, followed by treatment with Pam3CSK4 (100 ng/mL), LPS (100 ng/mL), or R848 (500 ng/mL) for 6 h. Luciferase activity was then measured (mean ± S.D., n = 3, *p < 0.05, **p < 0.01). D The experiment was carried out as described in (A) using PK–15 cells (mean ± S.D., n = 3, ***p < 0.001). E, F Quantitative real-time PCR was performed on RNA extracted from HeLa cells transfected with either an empty vector or pCMV–myc–MGF 505–3R for 24 h, followed by R848 (500 ng/mL) treatment for 6 h (mean ± S.D., n = 3, *p < 0.05, ***p < 0.001). G HeLa cells were transfected with either an empty vector or pCMV–myc–MGF 505–3R for 24 h, followed by LPS (100 ng/mL) treatment for 6 h. At 6 h posttreatment, cells were fixed and stained with DAPI (blue), anti-myc (green) and anti-p65 (red) antibodies. Localization of p65 was visualized by confocal microscopy. Magnification, ×40. H Quantification of p65 nuclear translocation was determined by counting cells from three independent fields (mean ± S.D., n = 3, *p < 0.05).
To further validate the results of the promoter assay, the expression of pro-inflammatory cytokines downstream of NF–κB was measured by qRT-PCR, providing additional evidence of the antagonistic role of the MGF 505–3R protein in innate immunity. Consistent with the results of the NF–κB promoter assay, ASFV MGF 505–3R decreased the transcription levels of IL-6, IL-1β, IFN-γ, and TNF-α in HeLa cells following R848 stimulation (Fig. 1E). Similar suppression was observed for IL-1β, IL-6, IFN-γ, and TNF-α upon LPS stimulation (Fig. S2). Importantly, we found that MGF 505–3R also potently suppressed the expression of type I (IFN-α/β) and type III (IFN-λ) interferons (Fig. 1F), significantly expanding its potential to disrupt the host’s antiviral response.
MGF 505–3R expression blocks the phosphorylation and translocation of p65
The cytoplasmic NF–κB transcription factor subunits p65 and p50 are bound to IκBα in resting cells. Upon activation, p65 is phosphorylated and translocates to the nucleus, where it mediates NF–κB–dependent gene transcription. Following our discovery that MGF 505–3R may act at the level of MyD88 or its downstream signaling, we examined the phosphorylation and nuclear translocation of p65 in these cells. LPS-induced phosphorylation of p65 was significantly suppressed by MGF 505–3R (Fig. S2A). Additionally, we assessed the effect of MGF 505–3R on the nuclear translocation of NF–κB p65. Expression of MGF 505‑3R inhibited LPS‑induced nuclear translocation of p65, as observed by immunofluorescence (Fig. 1G) and quantitative analysis (Fig. 1H). These findings further support the inhibitory role of MGF 505–3R in NF–κB signaling.
MGF 505–3R interacts with MyD88 and degrades MyD88 via the ubiquitin–proteasome
Since MGF 505–3R could extensively suppress NF–κB signaling triggered by various TLR ligands (TLRLs), we hypothesized that its target lies within the signaling pathway downstream of the TLR and upstream of NF–κB. We assessed MyD88 expression and found it to be significantly downregulated after transfection with MyD88 and MGF 505–3R (Fig. 2A). Subsequently, we performed an endogenous MyD88 assay, and the results revealed a similar pattern (Fig. 2B). These findings suggest that MGF 505–3R may inhibit NF–κB activation at the level of MyD88.
A HeLa cells were transfected with pcDNA3.1–Flag–MyD88, empty vector, or pCMV–myc–MGF 505–3R for 24 h. Flag–MyD88 expression was detected by Western blotting, with β-actin used as a loading control. B HeLa cells were transfected with either an empty vector or pCMV–myc–MGF 505–3R for 24 h. Endogenous MyD88 levels were assessed by Western blotting, with GAPDH as a loading control. C HEK 293T cells were transfected with pcDNA3.1–Flag–MyD88, an empty vector, or pCMV–myc–MGF 505–3R for 24 h. Whole cell lysates (WCL) were subjected to immunoprecipitation using anti-Flag or anti-Myc antibodies. The resulting immunoprecipitates were analyzed by Western blotting with the specified antibodies to detect protein–protein interactions. D HeLa cells were transfected with pGL4.32–NF–κB–Luc, along with either an empty vector, pcDNA3.1–Flag–MyD88 or pCMV–myc–MGF 505–3R for 24 h, followed by treatment with inhibitors MG132 (10 μM), BafA1 (10 μM), or 3–MA (25 nM) for 6 h. Luciferase activity was then measured to determine the effect on NF–κB signaling (mean ± S.D., n = 4, **p < 0.01). E, F HEK 293T cells were transfected with pcDNA3.1–Flag–MyD88, empty vector, or pCMV–myc–MGF 505–3R for 24 h. Whole cell lysates (WCL) were immunoprecipitated with anti-Flag, anti-Myc, anti-ubiquitination, anti-k48, or anti-k63 antibodies, and the immunoprecipitates were analyzed by Western blotting using the indicated antibodies to detect protein interactions.
To investigate the mechanism by which MGF 505–3R blocks NF–κB signaling, MGF 505–3R and MyD88 were co-transfected into HEK 293T cells. Co-immunoprecipitation (Co-IP) results confirmed that MGF 505–3R interacted with MyD88, thereby inhibiting NF–κB activation (Figs. 2C and S3A). Furthermore, to examine the mechanism through which MGF 505–3R reduces MyD88 expression, we used protein degradation pathway inhibitors. The ubiquitin–proteasome inhibitor MG132, the autophagy–lysosome inhibitor BafA1, and the autophagy vesicle inhibitor 3–MA were added, and only MG132 restored NF–κB signaling (Fig. 2D and S3B). The lack of effect from BafA1 and 3–MA indicates that the inhibition by MGF 505–3R is independent of autophagy, suggesting that its effect is related to the ubiquitin–proteasome pathway. We further studied the ubiquitination of Flag–MyD88 by immunoprecipitating Flag-tagged proteins, and the results revealed that MGF 505–3R significantly increased the ubiquitination of MyD88 (Fig. 2E). Additionally, MGF 505–3R enhanced K48-linked, but not K63-linked, ubiquitination of MyD88 (Fig. 2F). Thus, our results indicate that MGF 505–3R interacts with MyD88 and promotes its destabilization through K48-linked ubiquitination.
Amino acids 89–277 of MGF 505–3R are the key region for inhibiting NF–κB activity
Through SMART software for bioinformatics analysis, we identified two structural domains within MGF 505–3R and constructed plasmids expressing truncated mutants of MGF 505–3R to pinpoint the specific region responsible for inhibiting NF–κB activation (Fig. 3A, B). The expression of these truncated mutants was verified by Western blotting and immunofluorescence techniques (Fig. 3C). We then assessed the effects of these mutations on NF–κB promoter activation using a MyD88 overexpression assay. Overexpression of MGF 505–3R and MGF 505–3R∆58–83 inhibited MyD88-induced activation of the NF–κB promoter (Fig. 3D). To further investigate the function of this domain, MGF 505–3R∆58–83 and MyD88 were co-transfected into HEK 293T cells. Co-IP results confirmed that MGF 505–3R∆58–83 interacted with MyD88 to exert its inhibitory function (Fig. S5). To delineate the minimal region responsible for MyD88 degradation, a truncated protein encompassing residues 89–277 was expressed. Remarkably, this variant retained the ability to bind MyD88, thereby inducing its ubiquitination and triggering protein degradation (Fig. 3E, F). These findings indicate that the region of MGF 505–3R spanning amino acids 89–277 is essential for suppressing NF–κB activation. We further truncated the 89–277 aa region to identify a more specific set of key sites. However, no clear distinction was observed in the secondary truncation (Fig. S6).
A, B The structural domains of MGF 505–3R were predicted using SMART and SWISS–Model software, revealing a d1myo domain (red) and a conserved MGF 505 family signature domain, DUF249 (green), within the protein. C HEK 293T cells were transfected with truncated versions of MGF 505–3R, MGF 505–3RΔ89–277 or MGF 505–3RΔ58–83, and analyzed by Western blotting and immunofluorescence. D HeLa cells were transfected with pGL4.32–NF–κB–Luc, along with either pcDNA3.1–Flag–MyD88, an empty vector, and/or pCMV–myc–MGF 505–3R, MGF 505–3RΔ89–277, or MGF 505–3RΔ58–83 for 24 h. Luciferase activity was measured to assess the effect on NF–κB signaling. E HEK 293T cells were transfected with truncated versions of MGF 505–3R–89–277, and analyzed by Western blotting and immunofluorescence (mean ± S.D., n = 4, *p < 0.05). F HEK 293T cells were transfected with pcDNA3.1–Flag–MyD88, an empty vector, or pCMV–myc–MGF 505–3R–89–277 for 24 h. Whole cell lysates (WCL) were subjected to immunoprecipitation using anti-Flag, anti-Myc, or anti-ubiquitination antibodies. The resulting immunoprecipitates were analyzed by Western blotting with the specified antibodies to detect protein–protein interactions. G HeLa cells were transfected with pGL4.32–NF–κB–Luc, along with either an empty vector, pcDNA3.1–Flag–MyD88 or pCMV–myc–MGF 505–3R-89-277 for 24 h, followed by treatment with inhibitors MG132 (10 μM), BafA1 (10 μM), or 3–MA (25 nM) for 6 h. Luciferase activity was then measured to determine the effect on NF–κB signaling (mean ± S.D., n = 3, ***p < 0.001).
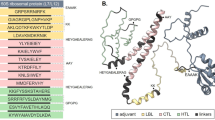
In order to identify potential immunomodulatory bioactive peptides, we performed systematic secondary structure prediction of the full-length MGF 505–3R amino acid sequence using computational tools, including PSIPRED. Our analysis focused on the previously identified functional domain (amino acids 89–277), which is critical for the immunosuppressive activity of MGF 505–3R. Within this region, we prioritized segments that concurrently contain ordered secondary structural elements—specifically, α-helices and β-sheets—as such motifs are likely to confer structural stability and biochemical functionality, improving their potential to mimic or perturb native protein functions in experimental settings. Based on these criteria, we selected two distinct peptide sequences located within this functional region: pep3R–1 (YDLVYKYYDQVKDCHDI) and pep3R–2 (HELNSTCSLKCLFKHAVI). Both peptides are similar in length (17–18 amino acids) and are predicted to exhibit mixed α-helical and β-sheet conformations. The use of two peptides sharing structural features but differing in primary sequence allows for controlled comparative functional assays, helping to rule out nonspecific effects and more accurately attribute any observed bioactivity to sequence-specific properties (Fig. 4A).
A The secondary structure of MGF 505–3R was analyzed using PSIPRED software, and the peptide (pep3R–1) was designed based on the predicted structure. B HeLa cells were transfected with pGL4.32–NF–κB–Luc for 24 h, followed by treatment with LPS (100 ng/mL) and pep3R–1 (40 μM) for 6 h. Luciferase activity was measured to assess NF–κB activation (mean ± S.D., n = 4, **p < 0.01, ***p < 0.001). C Mouse peritoneal macrophages were transfected with empty vector or pCMV–myc–MGF 505–3R for 24 h, then treated with LPS (100 ng/mL) for 6 h. The cells were analyzed by Western blotting to evaluate NF–κB pathway components. HeLa cells were treated with pep3R–1 (40 μM) and various TLR ligands of D LPS (100 ng/mL), E Pam3CSK4 (100 ng/mL), or F R848 (500 ng/mL) for 6 h. The expression of NF–κB–related genes was analyzed by qRT–PCR. G HeLa cells were treated with pep3R–1 (40 μM) and various TLR ligands of R848 (500 ng/mL) for 6 h. The expression of IFN-I/III genes was analyzed by qRT-PCR (mean ± S.D., n = 4, **p < 0.01, ***p < 0.001).
The effect of this peptide on NF–κB promoter activity was assessed through MyD88 overexpression, and the results demonstrated that pep3R–1 significantly inhibited NF–κB promoter activation (Fig. 4B). Additionally, we investigated the phosphorylation of p65 in mouse peritoneal macrophages, and found that LPS-induced p65 phosphorylation was significantly suppressed by pep3R–1 (Fig. 4C). Furthermore, we examined the effect of MGF 505–3R on the mRNA expression of pro-inflammatory cytokines induced by various TLR ligands. The results showed that LPS (Fig. 4D), Pam3CK4 (Fig. 4E), and R848 (Fig. 4F) significantly induced pro-inflammatory cytokine expression, and pep3R–1 markedly inhibited cytokine production in mouse RAW264.7 macrophages. Considering that the full-length protein inhibited the expression of interferon, we assessed whether the pep3R–1 peptide could also inhibit the production of IFN-I/III. Indeed, treatment with pep3R–1 resulted in a significant reduction of IFN-α, IFN-β, and IFN-λ expression levels (Fig. 4G).
MGF 505–3R peptide mitigates the clinical characterization of DSS-induced colitis
The DSS-induced colitis model is widely used to study intestinal inflammatory diseases. To evaluate whether the MGF 505–3R peptide could alleviate colitis by inhibiting the NF–κB signaling pathway, we established a DSS-induced colitis model in mice and administered continuous intraperitoneal injections of MGF 505–3R peptide, pep3R–1.
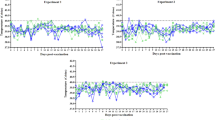
The Disease Activity Index (DAI) is a key indicator for evaluating the severity and progression of colitis. The DAI is based on three critical parameters: body weight change, fecal characteristics, and fecal occult blood in mice. When combined with fecal characteristics (Fig. 5A) and fecal occult blood (Fig. 5B) to calculate the DAI scores, we observed that DAI scores were significantly elevated in the DSS group, while these symptoms were attenuated in the DSS + pep3R–1 group (Fig. 5C). These results suggest that pep3R–1 reduces the DAI score in DSS-induced colitis and delays the progression of clinical symptoms.
A Hemafecia of the mice were observed throughout the experiment. B Fecal occult blood of the mice was assessed during the study. C The DAI score was calculated for each mouse to evaluate the severity of colitis (mean ± S.D., n = 5, *p < 0.05, **p < 0.01, ***p < 0.001). D Macroscopic images of the mouse colon were taken to visually assess inflammation (mean ± S.D., n = 5, *p < 0.05). E Hematoxylin and eosin (HE) staining of colon tissues (magnification ×40, scale bars = 50 μm) to examine histopathological changes. F Histological scores were assigned based on HE-stained sections to assess tissue damage (mean ± S.D., n = 2, *p < 0.05, **p < 0.01). G ELISA detection of the inflammatory factor TNF-α in mouse serum (mean ± S.D., n = 5, *p < 0.05, **p < 0.01). H ELISA measurements of inflammatory cytokine levels, including IL-1β, IL-6, and TNF-α in mouse serum (mean ± S.D., n = 5, *p < 0.05, **p < 0.01). I Measurement of myeloperoxidase (MPO) activity in mouse colon tissue as an indicator of neutrophil infiltration (mean ± S.D., n = 5, **p < 0.01).
In addition to the obvious characteristics observed in the mice, colon length and the expression of inflammatory cytokines are also crucial indicators of colitis. We found that the colon length in the DSS group decreased significantly compared to the control and pep3R–1 groups, indicating substantial colonic inflammation improvement in the pep3R–1 group (Fig. 5D). Pathological analysis revealed that in the DSS-treated mice, large areas of mucosal ulcers were present on the surface of the colon, with extensive epithelial cell loss, complete loss of colonic glands in some mice, and widespread edema in the submucosal layer. Granulocyte infiltration was also observed throughout the intestinal wall. In contrast, the pep3R–1 group showed abundant intestinal glands with no noticeable inflammatory cell infiltration, effectively restoring the epithelial crypt structure and resulting in a significantly reduced histological score (Fig. 5E, F).
The secretion of the inflammatory cytokine TNF-α was detected in the serum of all DSS-treated groups on both the 3rd and 6th days of the experiment. Notably, the levels of TNF-α in the serum of the DSS + pep3R–1 group were lower compared to those in the DSS–only group (Fig. 5G). Furthermore, the levels of inflammatory cytokines IL-6, IL-1β, TNF-α, and MPO in colon tissue were significantly reduced in the DSS + pep3R–1 group compared to the DSS-treated group (Fig. 5G–I). Additionally, we evaluated the anti-inflammatory effects of pep3R–1 in vivo in mice following sub-lethal LPS stimulation. The results demonstrated that pep3R–1 effectively inhibited the LPS-induced inflammatory response and alleviated inflammatory injury in the mice (Fig. S7). These findings suggest that the MGF 505–3R peptide pep3R–1 can inhibit both systemic and localized inflammation in the host, potentially mitigating the inflammatory storm associated with conditions such as DSS-induced colitis.
ASFV MGF 505–3R inhibits the antiviral effect of NF–κB in cells infected with the virus
To further investigate the biological functions of MGF 505–3R, HeLa cells were infected with VSV–EGFP at an MOI of 1.0 for 12 h, followed by harvesting and analysis using fluorescence microscopy and flow cytometry. The results indicated that MGF 505–3R could restore the viral load of VSV in HeLa cells (Fig. 6A), increase the infection rate, and enhance the average fluorescence intensity of individual cells (Fig. 6B), suggesting that MGF 505–3R could inhibit the NF–κB–mediated antiviral response. To assess the inhibitory effect of the 3R peptide on antiviral function, pep3R–1 was selected for further experiments. The findings demonstrated that pep3R–1 also significantly restored the intracellular viral load of VSV–EGFP (Fig. 6D–F). In contrast to pep3R–1, the peptide pep3R–2, which we constructed simultaneously, did not inhibit the antiviral response, although it did exert some inhibitory effects on NF–κB activation and related cytokine production (Figs. S8 and S9). We focused our analysis on the first 20 h post–VSV infection, a period during which viral load and IFN-β levels remained low and unaltered (Fig. S10). At this stage, both MGF 505–3R and peptide pep3R–1 potently inhibited the R848-induced expression of IFN-β (Fig. 6G). This indicates that their mechanism of action directly attenuates cellular antiviral signaling pathways rather than indirectly through modulating VSV replication. Taken together, these results suggest that MGF 505–3R and its peptide pep3R–1 inhibit the antiviral effect of NF–κB in virus-infected cells.
A HeLa cells were transfected with pCMV–Myc–MGF 505–3R (100 ng/mL) for 24 h, stimulated with R848 (5 μg/mL) for 6 h, and subsequently infected with VSV–GFP at an MOI of 1.0 for 12 h. GFP expression indicating VSV replication was visualized using fluorescence microscopy. B, C Flow cytometry analysis of VSV–GFP signals in HeLa cells after the same treatments (mean ± S.D., n = 4, *p < 0.05). D HeLa cells were treated with R848 (5 μg/mL) and pep3R–1 (40 μM) for 6 h, followed by VSV–GFP infection at MOI 1.0 for 12 h. VSV replicative GFP signals were again observed by fluorescence microscopy. E, F Flow cytometry analysis of VSV–GFP signals in HeLa cells after treatment with R848 and pep3R–1 (mean ± S.D., n = 4, *p < 0.05). G HeLa cells were treated with pep3R–1 (40 μM) and various TLR ligands of R848 (500 ng/mL) for 6 h, followed by VSV–GFP infection at MOI 1.0 for 12 h. The expression of IFN-β genes was analyzed by qRT-PCR (mean ± S.D., n = 3, **p < 0.01).
Discussion
ASF has a near 100% fatality rate and can only be controlled through prevention and strict hygiene measures24,25. ASFV is a complex virus, and this complexity has limited our understanding of the functions and precise mechanisms by which ASFV proteins counteract innate immunity26. In this study, we employed a dual-luciferase reporter assay to screen ASFV proteins and identified MGF 505–3R as an innate immune suppressor.
In this study, we focused on the canonical TLR–NF–κB signaling pathway, which can be activated through various mechanisms and plays a critical role in innate immunity as well as antiviral and inflammatory responses27. Several previous studies have demonstrated that ASFV proteins suppress innate immunity via multiple signaling pathways. For instance, ASFV protein F317L enhances IκBα stability by inhibiting IKKβ phosphorylation, thereby blocking NF–κB activation28. Additionally, I329L suppresses TRIF–mediated transcription of downstream cytokines and interferes with IRF3 and NF–κB activation by decreasing TRIF activity29. Our findings demonstrated that MGF 505–3R effectively suppressed NF–κB activation and its downstream inflammatory cytokines, underscoring its role in dampening the host innate immune response. Moreover, MGF 505–3R and its derived peptide pep3R–1 potently inhibited the expression of type I (IFN-α/β) and type III (IFN-λ) interferons, which are critical components of the antiviral defense system. By concurrently disrupting both NF–κB and interferon signaling pathways, MGF 505–3R created a conducive environment for viral replication. These results further substantiate MGF 505–3R as an ASFV–encoded immune evasion protein that plays a critical role in promoting ASFV replication during infection. Our findings suggest that inhibiting NF–κB signaling and inflammatory responses during the early stages of ASFV infection is critical for viral replication. Notably, while early suppression of these pathways facilitates viral proliferation, the late stages of ASFV infection are often characterized by excessive inflammatory responses leading to host mortality30,31,32. This dichotomy underscores a significant distinction between the early and late phases of infection, highlighting the complex interplay between ASFV and the host immune system.
Previous investigations into ASFV proteins have predominantly centered on elucidating their mechanisms through protein–protein interactions. For instance, the ASFV structural protein E120R has been shown to interact with IRF3, thereby inhibiting the production of IFN–β by blocking IRF3 activation33. Similarly, the A528R protein suppresses TLR8-mediated NF–κB signaling by targeting the activation and nuclear translocation of the p65 subunit, effectively attenuating the host’s antiviral and antibacterial responses34. Moreover, the I226R protein has been implicated in the inhibition of the NF–κB signaling pathway through the regulation of NEMO proteins35, while I267L disrupts the interaction between the E3 ubiquitin ligase Riplet and RIG–I, impairing Riplet-mediated K63 polyubiquitination and subsequent RIG–I activation36. These studies collectively highlight the sophisticated strategies employed by ASFV to modulate host innate immune responses via specific protein–protein interactions. In contrast to prior studies, our research reveals that MGF 505–3R not only interacts with MyD88—a pivotal adaptor in the TLR–NF–κB signaling pathway—but also promotes its degradation through K48–linked polyubiquitination, thereby inhibiting NF–κB activation. Although MGF 505–3R inhibited IKK complex-induced NF–κB activation (Fig. S4) in addition to promoting MyD88 degradation, this multifaceted mechanism highlights the sophisticated and layered strategies utilized by ASFV to dysregulate host innate immune signaling. Interestingly, MyD88 plays a crucial role as a common mediator in most TLR signaling pathways37,38,39. However, to our knowledge, few studies have explored the direct interaction of viral proteins with MyD88 in the context of viral immune evasion40,41,42, including its regulation through RNA interactions43. There remains a notable gap in research focused on MyD88-targeted immune modulation by viral proteins.
Furthermore, beyond its suppression of NF–κB signaling, we demonstrated that MGF 505–3R and its derived peptide pep3R–1 potently inhibited the expression of type I (IFN-α/β) and type III (IFN-λ) interferons. This finding significantly expands the immunomodulatory repertoire of MGF 505–3R, suggesting a broader strategy employed by ASFV to dampen the host antiviral response. Notably, a previous study reported that MGF 505–3R targeted the cGAS–STING pathway by promoting TBK1 degradation23. Our work, together with this earlier finding, positions MGF 505–3R as a multifunctional viral protein that concurrently disrupts both the NF–κB and interferon response pathways, thereby creating a more permissive environment for viral replication.
To investigate the molecular mechanism by which MGF 505–3R inhibits NF–κB signaling, we generated truncated mutants of MGF 505–3R and identified the 89–277 amino acid region as the critical domain. Through structure–function analysis, the 89–277 amino acid region of MGF 505–3R was identified as the critical domain responsible for MyD88 binding, ubiquitination, and subsequent degradation. Importantly, a truncated protein encompassing this region (MGF 505–3R–89–277) retained the full capacity to interact with MyD88 and promote its K48-linked polyubiquitination. Notably, the functional core within this domain is conserved across multiple MGF 505 family members (e.g., 1R, 4R, 5R), suggesting a broader, redundant strategy employed by ASFV to dampen host immunity (Fig. S11). This pinpointing of a minimal functional domain not only refines our understanding of the mechanistic basis of MGF 505–3R action but also provides a precise template for the development of targeted inhibitory peptides. Identifying key regions in viral proteins is a common strategy for elucidating their mechanisms. For example, the key site of ASFV E120R was identified through deletion mutants33, and point mutations have been used to study HBV X protein function44. Another approach focuses on recognizing the structural domains of viral and host protein interactions separately, as demonstrated in studies of ASFV pD345L and the IKK complex45. Building on these strategies, we investigated the peptide regions of MGF 505–3R to further explore its mechanism and potential therapeutic applications. For instance, antifungal peptides were developed from CD5 and CD646, and Salmonella TcpS peptides exhibited anti-inflammatory properties47. Our study aims to apply peptide-based insights for both mechanistic understanding and therapeutic advancements. We constructed two peptides based on the critical domain, evaluated their anti-inflammatory functions, and laid the groundwork for future research into anti-inflammatory therapies.
Excessive immune responses lead to inflammation, and persistent activation of Toll-like receptors induces the production of inflammatory cytokines, which can harm the host48,49. Targeting the host’s innate immune response and associated pathways offers promising therapeutic options, as many infectious diseases, autoimmune disorders, and cancers are linked to dysregulated inflammation15,48,49. However, the clinical efficacy and safety concerns of many anti-inflammatory drugs limit their therapeutic value, contributing to substantial healthcare burdens. In contrast, the anti-inflammatory peptide derived from the MGF 505–3R protein of ASFV provides a novel approach to peptide-based therapeutics. This peptide, whose core sequence is notably conserved within a subset of the MGF 505 multigene family, attenuates inflammation by targeting the MyD88–NF–κB pathway and represents a promising candidate for treating inflammatory diseases. Unlike traditional small-molecule drugs, peptide therapies often offer higher specificity and lower toxicity, making them attractive for clinical use. However, challenges such as peptide stability, delivery efficiency, and immunogenicity need to be addressed for practical application. Future research should focus on optimizing the peptide’s structure, improving delivery systems, and evaluating its efficacy in preclinical models. If successful, this peptide could offer a novel treatment for inflammatory diseases and inspire the development of similar immune-modulating peptides derived from viral proteins.
The present study demonstrates that the ASFV protein MGF 505–3R and its derived peptide significantly inhibit the NF–κB signaling pathway and interferon responses, thereby alleviating the host’s antiviral defense and restoring viral replication. This finding aligns with previous studies indicating that ASFV employs various strategies to modulate host immune responses, particularly through the suppression of NF–κB activation coupled with a substantial inhibition of R848-induced IFN-β production, directly linking the immunomodulatory function of MGF 505–3R to a compromised antiviral state in the host. By utilizing the VSV–EGFP system as a surrogate model, we were able to observe that the inhibition of NF–κB by MGF 505–3R and its peptides led to the restoration of viral replication, consistent with the known role of NF–κB in antiviral defense34,50. Although VSV was used as a surrogate model, these results strongly suggest that MGF 505–3R-mediated suppression of NF–κB and interferon signaling is a key strategy that facilitates ASFV replication by weakening the host's intrinsic defense mechanisms.
Our study identified the African swine fever virus (ASFV) protein MGF 505–3R as an immunomodulatory factor that interacts with the host innate immune system to promote immune evasion. We demonstrated that MGF 505–3R suppresses innate immunity by targeting the NF–κB signaling pathway. Specifically, MGF 505–3R interacts with MyD88, modulating its ubiquitination and thereby preventing NF–κB activation. This inhibition reduces inflammatory cytokine production, alleviates inflammation, and suppresses the antiviral effects of NF–κB. Furthermore, structural analysis of MGF 505–3R revealed an inflammation-inhibiting peptide with potential as an anti-inflammatory agent capable of mitigating DSS-induced inflammatory storms. These findings provide novel insights into the immune evasion mechanisms of ASFV and offer new perspectives for the development of anti-inflammatory therapeutics.
Materials and methods
Cells and viruses
The HEK 293T, HeLa, RAW264.7, and PK–15 cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM, Gibco–BRL, USA), supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin (Beyotime Biotech, Shanghai, China). The cells were maintained at 37 °C in a 5% CO₂ incubator.
Peritoneal macrophages were isolated from 6–8-week-old male C57BL/6 mice. After euthanizing the mice, the ventral surface was disinfected with 75% ethanol. To harvest macrophages, 5 mL of sterile, ice–cold phosphate-buffered saline (PBS) was injected into the peritoneal cavity with a 25-gauge needle. The abdomen was gently massaged to dislodge resident cells, and the PBS buffer containing lavaged cells was carefully aspirated. The peritoneal fluid was carefully aspirated with a sterile syringe and transferred to a 15 mL conical tube on ice. The collected fluid was centrifuged at 300 × g for 5 min at 4 °C. The supernatant was discarded, and the cell pellet was resuspended in RPMI 1640 medium supplemented with 10% FBS and 1% penicillin–streptomycin.
The vesicular stomatitis virus–GFP (VSV–GFP) used in this study was generously provided by Dr. Jianzhong Zhu at Yangzhou University.
Construction and transfection of plasmids
PCR primers were designed using Primer Premier 5 software and synthesized by TSingke Biotechnology Co., Ltd. The pcDNA–MGF 505–3R plasmid (GenBank: MN393476.1) was synthesized by Nanjing Genscript Biotechnology Co., Ltd. The MGF 505–3R gene was amplified by PCR and inserted into the EcoR I and Sal I sites of the pCMV–myc vector using the ClonExpress Ultra One Step Cloning Kit. The resulting recombinant plasmid was designated pCMV–myc–MGF 505–3R (Table 1). Truncated mutants of MGF 505–3R were constructed by deleting specific structural domain sequences. The two deletion mutants were named MGF 505–3R∆58–83 and MGF 505–3R∆89–277. The human NF–κB reporter gene and the pcDNA3.1–MyD88 plasmid have been cloned and are preserved in our laboratory.
Plasmid DNA was extracted from recombinant bacteria using the EndoFree Mini Plasmid Kit II (Tiangen, Beijing, China). Purified plasmids were then transfected into HEK 293T, HeLa, or PK–15 cells using Lip2000 Transfection Reagent (Biosharp), following the manufacturer’s instructions. Briefly, purified plasmids were mixed with the transfection reagent in a 1:1 ratio, incubated at room temperature for 20 min, and then slowly added to the cells. The cells were incubated at 37 °C for 24 h to allow for transfection, followed by subsequent experiments.
Design and synthesis of peptides
The design of the peptide pep3R–1 was based on the analysis of the secondary structure of the protein using PredictProtein (https://predictprotein.org/). A portion of the protein exhibiting both α–helix and β–sheet structures was rationally selected for peptide design, yielding two peptides: pep3R–1 (YDLVYKYYDQVKDCHDI) and pep3R–2 (HELNSTCSLKCLFKHAVI). CP was served as a control peptide with the sequence of FQKNHTYIDASRP, while SR4, a known anti-inflammatory peptide, was used as a positive control with the sequence of WLSKSFIKKDWTEYE. To facilitate cellular uptake, a cell-permeating sequence (RQIKIWFQNRRMKWKK) was introduced at the amino-terminal end of each peptide. All peptides were synthesized by Shanghai Sangon Biotechnology Co., Ltd. with a purity of over 95%. The peptides were resuspended in DPBS and stored at −80 °C.
Dual-luciferase reporter assays
The indicated plasmids or empty vectors, along with the reporter plasmids NF–κB–Luc and pcDNA3.1–MyD88, were co-transfected into HEK 293T, HeLa, or PK–15 cells using Lip2000 transfection reagent. After 24 h of transfection, cells were lysed with 5× cell lysis buffer (Vazyme) and shaken at room temperature for 5 min. The cell lysates were then collected by centrifugation to obtain the supernatant. Luciferase activity was measured using the Dual-Luciferase Reporter Assay Kit (Vazyme) according to the manufacturer’s instructions. All experiments were performed in triplicate.
RNA extraction and quantitative PCR (qPCR) assays
Total RNA was extracted from transfected or infected cells using the RNAprep Pure Cell/Bacteria Kit (TaKaRa). For cDNA synthesis, 2 μl of RNA was reverse transcribed using HiScript® III RT SuperMix for qPCR (Vazyme). Quantitative PCR (qPCR) was then performed on the cDNA using ChamQTM Universal SYBR® qPCR Master Mix (Vazyme). Glyceraldehyde–3–phosphate dehydrogenase (GAPDH) was used as a reference gene for internal normalization (Table 1). Relative RNA levels were calculated using the 2–∆∆Ct method. Data from one of the three independent replicates are presented.
Co-immunoprecipitation (Co-IP) and Western blotting (WB)
Plasmids were transfected into HEK 293T cells seeded in 6-well plates (1.0 × 10⁶ cells/well) for 24 h. The cells were then lysed on ice for 10 min using cell lysis buffer for Western and IP (Beyotime). The lysate was centrifuged, and the supernatant was incubated with 20 μl of magnetic bead suspension and treated with the Flag-tag Protein IP Assay Kit with Agarose Gel (Beyotime). The mixture was incubated overnight at 4 °C on a side-swing shaker or rotary mixer. After magnetic separation, the beads were washed with SDS–PAGE loading buffer.
The proteins were electrophoretically separated by SDS–PAGE and transferred to nitrocellulose membranes (NC). The NC membranes were blocked with a blocking solution (5% skimmed milk powder in PBS, 0.1% Tween 20) for 2 h at room temperature (RT). The membranes were then incubated with the primary antibody overnight at 4 °C, followed by incubation with goat anti–mouse/rabbit IgG HRP for 2 h at RT. Protein signals were detected using the ECL chemiluminescence detection kit (Vazyme).
For Co-IP assays, the association of MGF 505–3R or MGF 505–3R–89–277 with Flag–MyD88 was precipitated overnight using magnetic beads coupled with Flag monoclonal antibody. The precipitates were analyzed by WB using mouse anti-Myc monoclonal antibody and rabbit anti-Flag monoclonal antibody.
Anti-viral activity assay
HeLa cells were seeded into 96-well cell culture plates at a density of 1 × 10⁵ cells/mL and incubated at 37 °C in a 5% CO₂ incubator. The cells were transfected with pCMV–myc–MGF 505–3R and incubated for 24 h at 37 °C with 5% CO₂, or for 6 h after the addition of the protein–peptide once the cells had formed a uniform monolayer.
For the viral infection assay, VSV–GFP was diluted in DMEM supplemented with 2% fetal bovine serum (FBS). Virus solution with MOI = 1.0 was added to each well. Control cells (without virus) were included for comparison.
The indicated plasmids or empty vectors, along with pcDNA3.1–MyD88, were co-transfected into HeLa cells using Lip2000 Transfection Reagent according to the manufacturer’s instructions. After a 12 h incubation, cell damage was assessed and documented using fluorescent inverted microscopy or flow cytometry.
Fluorescence microscopy
HeLa cells were seeded on glass coverslips in 24–well plates at a density of 1.5 × 10⁵ cells/well. After 24 h of incubation, cells were transfected with 100 ng of MGF 505–3R and 30 ng of MyD88 plasmids. Following a 24 h transfection period, cells were fixed with 4% paraformaldehyde at room temperature for 30 min and then permeabilized with 0.5% Triton X–100 for 20 min. After permeabilization, the cells were rinsed with PBS, blocked, and then incubated with mouse anti-Myc monoclonal antibody and rabbit anti-p65 polyclonal antibody, followed by corresponding AF488-labeled (for Myc) and AF594-labeled (for p65) secondary antibodies. Images were visualized under a Leica SP8 laser-scanning confocal microscope (LSCM).
DSS-induced colitis in mice
Male C57BL/6 mice (n = 20, aged 6–8 weeks, weight 20–25 g) were purchased from Beijing Vital River Laboratory Animal Technology Co. The mice were housed under controlled conditions in a laboratory with a temperature of 21 °C ± 2 °C and humidity of 50% ± 5%. They were randomly assigned to five groups (n = 4 per group) and allowed to acclimatize for 7 days before the initiation of the study. We have complied with all relevant ethical regulations for animal use.
The control group received water ad libitum daily as a negative control. The DSS group (DSS) was administered 4% DSS daily ad libitum to induce experimental colitis. In the 5–ASA group, mice were treated with 4% DSS ad libitum daily and simultaneously received oral gavage of 100 mg/kg 5–ASA (dissolved in 0.5% CMC–Na). The peptide group consumed 4% DSS ad libitum daily, followed by intraperitoneal injection of 0.1 mL of either control peptide (CP) or pep3R–1 (40 μM) dissolved in DPBS. All treatments were administered for 6 consecutive days according to the described dosing regimen.
During the experimental period, mice were monitored daily for general appearance (e.g., vigor, coat condition). Body weight, fecal consistency, and rectal bleeding were regularly assessed. The Disease Activity Index (DAI) was calculated based on these observations. On day 7, mice were euthanized via intraperitoneal injection of sodium pentobarbital, and serum was collected. The colon was promptly excised, and its length was measured. A 2 cm section of the colorectal tissue was fixed in 4% paraformaldehyde for histological analysis, while another portion was stored at −80°C for further examination. Tissue samples were washed with PBS, weighed, then minced and homogenized in an ice-cold bath. The resulting emulsion was homogenized at 3000 rpm for 10 min. The supernatant was then collected for the assessment of myeloperoxidase (MPO) activity and inflammatory factor concentrations. MPO activity was measured using the MPO Assay Kit, while concentrations of inflammatory cytokines (IL-1β, IL-6, and TNF-α) were quantified using standard ELISA kits.
Evaluation of the anti-inflammatory effects of pep3R–1 peptide in vivo
CP or pep3R–1 was intraperitoneally administered to C57BL/6J mice at a dosage of 10 nmol/g. Following 1 h incubation period post-peptide treatment, the mice were subjected to an intraperitoneal challenge with a sublethal dose of LPS (1 μg/g). At 4 h post-LPS administration, mice were euthanized via intraperitoneal injection of sodium pentobarbital. Spleen tissues and serum samples were harvested from each experimental group. The relative mRNA expression levels of pro-inflammatory cytokines, including TNF-α, IL-6, IFN-γ, and IL-1β, in splenic cells were quantitatively analyzed using reverse transcription qRT-PCR. Concurrently, serum cytotoxicity was evaluated using a commercial LDH Cytotoxicity Assay Kit (Beyotime Biotechnology, China) according to the manufacturer’s protocol.
Statistics and reproducibility
The data were expressed as mean standard deviation (mean SD) and represented the results of two similar experiments. The statistical significance was determined by the GraphPad Prism 8.0 software, and P < 0.05 was considered statistically significant by the paired two-tailed t-test. The symbols “*”, “**”, “***”, and “ns” in the graphs respectively represent P < 0.05, P < 0.01, P < 0.001, and no statistical significance.
Ethical approval
The experimental protocol was approved by the Ethics Committee of the Animal Experiments of Yangzhou University [Approval ID: SYXK (Su) 2022–0044], and conducted according to the guidelines for animal care and ethics.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Uncropped westerns and gels can be found in the Supplementary Information file. Source data can be obtained from Supplementary Data. Additional data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ge, S. et al. Molecular characterization of African swine fever virus, China, 2018. Emerg. Infect. Dis. 24, 2131–2133 (2018).
Eustace Montgomery, R. On a form of swine fever occurring in British east Africa (Kenya Colony). J. Comp. Pathol. 34, 159–191 (1921).
Costard, S. et al. African swine fever: how can global spread be prevented? Philos. Trans. R. Soc. Lond. B Biol. Sci. 364, 2683–2696 (2009).
Zhao, D. et al. 2019. Replication and virulence in pigs of the first African swine fever virus isolated in China. Emerg. Microbes Infect. 8, 438–447 (2019).
Dixon, L. K., Stahl, K., Jori, F., Vial, L. & Pfeiffer, D. U. 2020. African swine fever epidemiology and control. Annu. Rev. Anim. Biosci. 8, 221–246 (2020).
Alonso, C. et al. ICTV virus taxonomy profile: Asfarviridae. J. Gen. Virol. 99, 613–614 (2018).
Chapman, D. A. G., Tcherepanov, V., Upton, C. & Dixon, L. K. Comparison of the genome sequences of non–pathogenic and pathogenic African swine fever virus isolates. J. Gen. Virol. 89, 397–408 (2008).
Aslanyan, L., Avagyan, H. & Karalyan, Z. Whole–genome–based phylogeny of African swine fever virus. Vet. World 13, 2118–2125 (2020).
Goatley, L. C. et al. African swine fever virus NAM P1/95 is a mixture of genotype I and genotype VIII viruses. Microbiol. Resour. Announc. 13, e00067–24 (2024).
Quembo, C. J., Jori, F., Vosloo, W. & Heath, L. Genetic characterization of African swine fever virus isolates from soft ticks at the wildlife/domestic interface in Mozambique and identification of a novel genotype. Transbound. Emerg. Dis. 65, 420–431 (2018).
Abe, T. & Barber, G. N. Cytosolic–DNA–mediated, STING–dependent proinflammatory gene induction necessitates canonical NF–kappaB activation through TBK1. J. Virol. 88, 5328–5341 (2014).
Pietrobon, A. J. et al. Antiviral response induced by Toll–like receptor (TLR) 7/TLR8 activation inhibits human immunodeficiency virus type 1 infection in cord blood macrophages. J. Infect. Dis. 225, 510–519 (2022).
Kawai, T. & Akira, S. TLR signaling. Semin. Immunol. 19, 24–32 (2007).
Kawai, T. & Akira, S. Toll–like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 34, 637–650 (2011).
Kawai, T. & Akira, S. The role of pattern–recognition receptors in innate immunity: update on Toll–like receptors. Nat. Immunol. 11, 373–384 (2010).
Dixon, L. K., Chapman, D. A., Netherton, C. L. & Upton, C. African swine fever virus replication and genomics. Virus Res. 173, 3–14 (2013).
Afonso, C. L. et al. African swine fever virus multigene family 360 and 530 genes affect host interferon response. J. Virol. 78, 1858–1864 (2004).
Zsak, L. et al. African swine fever virus multigene family 360 and 530 genes are novel macrophage host range determinants. J. Virol. 75, 3066–3076 (2001).
Li, J. et al. pMGF505–7R determines pathogenicity of African swine fever virus infection by inhibiting IL–1β and type I IFN production. PLoS Pathog. 17, e1009733 (2021).
Li, D. et al. African swine fever virus protein MGF–505–7R promotes virulence and pathogenesis by inhibiting JAK1– and JAK2–mediated signaling. J. Biol. Chem. 297, 101190 (2021).
Zhang, K. et al. MGF360–9L is a major virulence factor associated with the African swine fever virus by antagonizing the JAK/STAT signaling pathway. mBio 13, e0233021 (2022).
Zhuo, Y. et al. African swine fever virus MGF360–12L inhibits type I interferon production by blocking the interaction of Importin α and NF–κB signaling pathway. Virol. Sin. 36, 176–186 (2021).
Cheng, M. et al. African swine fever virus MGF505–3R inhibits cGAS–STING–mediated IFN–β pathway activation by degrading TBK1. Anim. Dis. 2, 13 (2022).
Sánchez–Vizcaíno, J. M., Mur, L. & Martínez–López, B. African swine fever: an epidemiological update. Transbound. Emerg. Dis. 59, 27–35 (2012).
Arias, M. et al. Approaches and perspectives for development of African swine fever virus vaccines. Vaccines 5, 35 (2017).
Duan, X. et al. Research progress on the proteins involved in African swine fever virus infection and replication. Front. Immunol. 13, 947180 (2022).
Song, K. & Li, S. The role of ubiquitination in NF–kappaB signaling during virus infection. Viruses 13, 145 (2021).
Yang, J. et al. African swine fever virus F317L protein inhibits NF–κB activation to evade host immune response and promote viral replication. mSphere 6, e0065821 (2021).
de Oliveira, V. L. et al. A novel TLR3 inhibitor encoded by African swine fever virus (ASFV). Arch. Virol. 156, 597–609 (2011).
Walczak, M. et al. Blood counts, biochemical parameters, inflammatory, and immune responses in pigs infected experimentally with the African swine fever virus isolate Pol18_28298_O111. Viruses 13, 521 (2021).
Franzoni, G., Pedrera, M. & Sanchez–Cordon, P. J. African swine fever virus infection and cytokine response in vivo: an update. Viruses 15, 233 (2023).
Wang, S. et al. Cytokine storm in domestic pigs induced by infection of virulent African swine fever virus. Front. Vet. Sci. 7, 601641 (2020).
Liu, H. et al. African swine fever virus E120R protein inhibits interferon beta production by interacting with IRF3 to block its activation. J. Virol. 95, e0082421 (2021).
Liu, X. et al. African swine fever virus A528R inhibits TLR8 mediated NF–kappaB activity by targeting p65 activation and nuclear translocation. Viruses 13, 2046 (2021).
Hong, J. et al. I226R protein of African swine fever virus is a suppressor of innate antiviral responses. Viruses 14, 575 (2022).
Ran, Y. et al. African swine fever virus I267L acts as an important virulence factor by inhibiting RNA polymerase III–RIG–I–mediated innate immunity. PLoS Pathog. 18, e1010270 (2022).
O’Neill, L. A., Golenbock, D. & Bowie, A. G. The history of Toll–like receptors – redefining innate immunity. Nat. Rev. Immunol. 13, 453–460 (2013).
Adachi, O. et al. Targeted disruption of the MyD88 gene results in loss of IL–1–and IL–18–mediated function. Immunity 9, 143–150 (1998).
O’Neill, L. A. & Bowie, A. G. The family of five: TIR–domain–containing adaptors in Toll–like receptor signalling. Nat. Rev. Immunol. 7, 353–364 (2007).
Fedosyuk, S., Grishkovskaya, I., de Almeida Ribeiro, E. & Skern, T. Characterization and structure of the vaccinia virus NF–κB antagonist A46. J. Biol. Chem. 289, 3749–3762 (2014).
Feng, N. et al. Enterovirus 71–induced has–miR–21 contributes to evasion of host immune system by targeting MyD88 and IRAK1. Virus Res. 237, 27–36 (2017).
Zhao, Q. et al. Kaposi’s sarcoma–associated herpesvirus–encoded replication and transcription activator impairs innate immunity via ubiquitin–mediated degradation of myeloid differentiation factor 88. J. Virol. 89, 415–427 (2015).
Lingel, A. et al. Kaposi’s sarcoma–associated Herpesvirus reduces cellular myeloid differentiation primary–response gene 88 (MyD88) expression via modulation of its RNA. J. Virol. 90, 180–188 (2016).
Wei, C. et al. The hepatitis B virus X protein disrupts innate immunity by downregulating mitochondrial antiviral signaling protein. J. Immunol. 185, 1158–1168 (2010).
Chen, H. et al. ASFV pD345L protein negatively regulates NF–κB signalling by inhibiting IKK kinase activity. Vet. Res. 53, 32 (2022).
Mourglia et al. In vitro analysis of tandem peptides from Human CD5 and CD6 scavenger receptors as potential anti–cryptococcal agents. J. Fungi 10, 667 (2024).
Xiong, D. et al. Salmonella coiled–coil– and TIR–containing TcpS evades the innate immune system and subdues inflammation. Cell Rep. 28, 804–818.e7 (2019).
Chen, J. Q., Szodoray, P. & Zeher, M. Toll–like receptor pathways in autoimmune diseases. Clin. Rev. Allergy Immunol. 50, 1–17 (2016).
Bahia, M. S., Kaur, M., Silakari, P. & Silakari, O. Interleukin–1 receptor associated kinase inhibitors: potential therapeutic agents for inflammatory– and immune–related disorders. Cell Signal. 27, 1039–1055 (2015).
Wang, T. et al. The African swine fever virus MGF300–4L protein is associated with viral pathogenicity by promoting the autophagic degradation of IKKβ and increasing the stability of IκBα. Emerg. Microbes Infect. 13, 2333381 (2024).
Acknowledgements
This research was supported by the Key research and development program (Modern Agriculture) project of Jiangsu Province (BE2021331, BE2020398), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD). We thank all the editors and reviewers for their valuable comments and suggestions that helped improve the quality of this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.L., D.X., X.J., and Z.P.; methodology: H.L., D.X., L.S., F.W., X.H., and J.H.; software: H.L. and D.X.; formal analysis, H.L. and D.X.; writing—original draft preparation: H.L.; writing—review and editing: D.X., X.J., and Z.P.; visualization: H.L., D.X., and X.K.; supervision: D.X., X.J., Z.P., X.K., D.G., L.S., and C.M.; project administration, X.J. and Z.P.; funding acquisition, X.J. and Z.P. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary handling editors: Dr Suan-Sin Foo and Dr Ophelia Bu.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, H., Sun, L., Wang, F. et al. African swine fever virus–encoded protein MGF 505–3R impairs innate immunity via ubiquitin–mediated degradation of MyD88. Commun Biol 9, 407 (2026). https://doi.org/10.1038/s42003-026-09681-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09681-0