Abstract
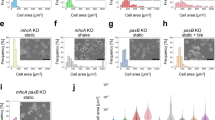
Synchronicity is observed in many biological systems, including development, as seen in vertebrate body axis formation, fly eye development, and development of the social amoeba D. discoideum. Despite its prevalence, quantitative analyses of synchronicity at the single-cell level remain rare. Here we show that synchronicity in D. discoideum is mediated by early cAMP signaling. Using single-cell RNA sequencing (scRNA-seq), we quantify transcriptome similarities between individual cells during development. We show that synchronicity first declines upon starvation but then increases with the onset of cAMP-pulse signaling. Synchronicity remains stable throughout development and differs between prespore and prestalk cells. Genetic perturbations of cAMP production and response reveal that cAMP signaling is essential for establishing and maintaining synchronicity, as its absence leads to highly asynchronous development. These findings highlight the role of cAMP signaling in coordinating transcriptomic and morphological synchronicity, and establish scRNA-seq as a tool for quantitative analysis of developmental synchronicity.
Similar content being viewed by others
Data availability
Raw sequencing reads and processed data have been deposited in the NCBI Gene Expression Omnibus (GEO) under accession number GSE305468. It is available from Zenodo with the DOI 10.5281/zenodo.1848190764. The numerical source data underlying the graphs are available in Supplementary Data 1 (Fig. 2); Supplementary Data 2 (Supplementary Fig. 1); and Supplementary Data 3 (Supplementary Fig. 2a and Fig. 2). All other data and materials are available from the corresponding author on reasonable request.
Code availability
Custom code and reproducible workflows are available at https://github.com/lenatr99/scRNA_dicty, including pipelines for 10x preprocessing, UCE embeddings, UMAP/Leiden clustering, fluorescent-tag assignment, prespore/prestalk classification, synchronicity scoring with bootstrap confidence intervals, and figure generation. The repository includes step-by-step instructions and environment files to enable full reproducibility. It is available from Zenodo with the DOI 10.5281/zenodo.1848180865.
References
Oates, A. C., Morelli, L. G. & Ares, S. Patterning embryos with oscillations: structure, function and dynamics of the vertebrate segmentation clock. Development 139, 625–639 (2012).
Lee, J. D. & Treisman, J. E. In Drosophila Eye Development. Results and Problems in Cell Differentiation (ed K. Moses) Vol. 37 (Springer, 2002).
Spudich, J. A. Dictyostelium discoideum: molecular approaches to cell biology (Academic Press, 1987).
Kessin, R. H. Dictyostelium - Evolution, cell biology, and the development of multicellularity (Cambridge University Press, 2001).
Hall, A. L., Schlein, A. & Condeelis, J. Relationship of pseudopod extension to chemotactic hormone-induced actin polymerization in amoeboid cells. J. Cell Biochem. 37, 285–299 (1988).
Loomis, W. F. Dictyostelium discoideum. A developmental system (Academic Press, 1975).
Kriebel, P. W., Barr, V. A. & Parent, C. A. Adenylyl cyclase localization regulates streaming during chemotaxis. Cell 112, 549–560 (2003).
Sgro, A. E. et al. From intracellular signaling to population oscillations: bridging size- and time-scales in collective behavior. Mol. Syst. Biol. 11, 779 (2015).
Brock, D. A. et al. A Dictystelium mutant with defective aggregate size determination. Development 122, 2569–2578 (1996).
Artemenko, Y., Lampert, T. J. & Devreotes, P. N. Moving towards a paradigm: common mechanisms of chemotactic signaling in Dictyostelium and mammalian leukocytes. Cell. Mol. Life Sci. 71, 3711–3747 (2014).
Antolovic, V., Lenn, T., Miermont, A. & Chubb, J. R. Transition state dynamics during a stochastic fate choice. Development 146. https://doi.org/10.1242/dev.173740 (2019).
Katoh-Kurasawa, M. et al. Transcriptional milestones in Dictyostelium development. Genome Res. 31, 1498–1511 (2021).
Parikh, A. et al. Conserved developmental transcriptomes in evolutionarily divergent species. Genome Biol. 11, R35 (2010).
Rosengarten, R. D. et al. Leaps and lulls in the developmental transcriptome of Dictyostelium discoideum. BMC Genom. 16, 294 (2015).
Van Driessche, N. et al. Global transcriptional responses to cisplatin in Dictyostelium discoideum identify potential drug targets. Proc. Natl. Acad. Sci. USA 104, 15406–15411 (2007).
Van Driessche, N. et al. A transcriptional profile of multicellular development in Dictyostelium discoideum. Development 129, 1543–1552 (2002).
Westbrook, E. R., Lenn, T., Chubb, J. R. & Antolovic, V. Collective signalling drives rapid jumping between cell states. Development 150. https://doi.org/10.1242/dev.201946 (2023).
Kim, J., Heslop-Harrison, P., Postlethwaite, I. & Bates, D. G. Stochastic noise and synchronisation during dictyostelium aggregation make cAMP oscillations robust. PLoS Comput. Biol. 3, e218 (2007).
Laub, M. T. & Loomis, W. F. A molecular network that produces spontaneous oscillations in excitable cells of Dictyostelium. Mol. Biol. Cell 9, 3521–3532 (1998).
Siegert, F. & Weijer, C. Digital image processing of optical density wave propagation in Dictyostelium discoideum and analysis of the effects of caffeine and ammonia. J. Cell Sci. 93, 325–335 (1989).
Brimson, C. A. et al. Collective oscillatory signaling in Dictyostelium discoideum acts as a developmental timer initiated by weak coupling of a noisy pulsatile signal. Dev. Cell 60, 918–933.e914 (2025).
Cai, H. et al. Nucleocytoplasmic shuttling of a GATA transcription factor functions as a development timer. Science 343, 1249531 (2014).
Pitt, G. S. et al. Structurally distinct and stage-specific adenylyl cyclase genes play different roles in Dictyostelium development. Cell 69, 305–315 (1992).
Wang, B. & Kuspa, A. Dictyostelium development in the absence of cAMP. Science 277, 251–254 (1997).
Jermyn, K. A., Duffy, K. T. & Williams, J. G. A new anatomy of the prestalk zone in Dictyostelium. Nature 340, 144–146 (1989).
Sternfeld, J. & David, C. N. Fate and regulation of anterior-like cells in Dictyostelium slugs. Dev. Biol. 93, 111–118 (1982).
Rosen, Y. et al. Universal cell embeddings: a foundation model for cell biology. bioRxiv https://doi.org/10.1101/2023.11.28.568918 (2024).
Eder, M. et al. Systematic mapping of organism-scale gene-regulatory networks in aging using population asynchrony. Cell 187, 3919–3935.e3919 (2024).
Martinez-Jimenez, C. P. et al. Aging increases cell-to-cell transcriptional variability upon immune stimulation. Science 355, 1433–1436 (2017).
Maeda, M. et al. Changing patterns of gene expression in dictyostelium prestalk cell subtypes recognized by in situ hybridization with genes from microarray analyses. Eukaryot Cell 2, 627–637 (2003).
Do, V. H. & Canzar, S. A generalization of t-SNE and UMAP to single-cell multimodal omics. Genome Biol. 22, 130 (2021).
Yang, Y. et al. Dimensionality reduction by UMAP reinforces sample heterogeneity analysis in bulk transcriptomic data. Cell Rep. 36, 109442 (2021).
Salvidge, W. M. et al. Linage priming and cell type proportioning depends on the interplay between stochastic and deterministic factors. eLife 14, RP105512 (2025).
Iranfar, N., Fuller, D. & Loomis, W. F. Transcriptional regulation of post-aggregation genes in Dictyostelium by a feed-forward loop involving GBF and LagC. Dev. Biol. 290, 460–469 (2006).
Corrigan, A. M. & Chubb, J. R. Regulation of transcriptional bursting by a naturally oscillating signal. Curr. Biol. 24, 205–211 (2014).
Chen, G., Shaulsky, G. & Kuspa, A. Tissue-specific G1-phase cell-cycle arrest prior to terminal differentiation in Dictyostelium. Development 131, 2619–2630 (2004).
Shaulsky, G. & Loomis, W. F. Mitochondrial DNA replication but no nuclear DNA replication during development of Dictyostelium. Proc. Natl. Acad. Sci. USA 92, 5660–5663 (1995).
Singer, G., Araki, T. & Weijer, C. J. Oscillatory cAMP cell-cell signalling persists during multicellular Dictyostelium development. Commun. Biol. 2, 139 (2019).
Hashimura, H., Morimoto, Y. V., Yasui, M. & Ueda, M. Collective cell migration of Dictyostelium without cAMP oscillations at multicellular stages. Commun. Biol. 2, 34 (2019).
Hirose, S., Katoh-Kurasawa, M. & Shaulsky, G. Cyclic AMP is dispensable for allorecognition in Dictyostelium cells overexpressing PKA-C. J. Cell Sci. 134. https://doi.org/10.1242/jcs.258777 (2021).
Hirose, S., Chen, G., Kuspa, A. & Shaulsky, G. The polymorphic proteins TgrB1 and TgrC1 function as a ligand-receptor pair in Dictyostelium allorecognition. J. Cell Sci. 130, 4002–4012 (2017).
Kay, R. R. & Thompson, C. R. L. Cross-induction of cell types in Dictyostelium: evidence that DIF-1 is made by prespore cells. Development 128, 4959–4966 (2001).
Thompson, C. R. & Kay, R. R. The role of DIF-1 signaling in Dictyostelium development. Mol. Cell 6, 1509–1514 (2000).
Richardson, D. L., Loomis, W. F. & Kimmel, A. R. Progression of an inductive signal activates sporulation in Dictyostelium discoideum. Development 120, 2891–2900 (1994).
Shaulsky, G., Kuspa, A. & Loomis, W. F. A multidrug resistance transporter/serine protease gene is required for prestalk specialization in Dictyostelium. Genes Dev. 9, 1111–1122 (1995).
Chen, Z. H. & Schaap, P. Secreted cyclic Di-GMP induces stalk cell differentiation in the eukaryote dictyostelium discoideum. J. Bacteriol. 198, 27–31 (2016).
Consalvo, K. M. et al. Extracellular signaling in Dictyostelium. Int. J. Dev. Biol. 63, 395–405 (2019).
Yoshino, R. et al. Regulation of ammonia homeostasis by the ammonium transporter AmtA in Dictyostelium discoideum. Eukaryot Cell 6, 2419–2428 (2007).
Chen, X. et al. Terpene synthase genes in eukaryotes beyond plants and fungi: Occurrence in social amoebae. Proc. Natl. Acad. Sci. USA 113, 12132–12137 (2016).
Chen, X. et al. Diversity and functional evolution of terpene synthases in dictyostelid social amoebae. Sci. Rep. 8, 14361 (2018).
Chen, X. et al. A terpene synthase-cytochrome P450 cluster in Dictyostelium discoideum produces a novel trisnorsesquiterpene. Elife 8. https://doi.org/10.7554/eLife.44352 (2019).
Loomis, W. F. Role of PKA in the timing of developmental events in Dictyostelium cells. Microbiol. Mol. Biol. Rev. 62, 684–694 (1998).
Schaap, P. From environmental sensing to developmental control: cognitive evolution in dictyostelid social amoebas. Philos. Trans. R Soc. Lond. B Biol. Sci. 376, 20190756 (2021).
Warren, J. & Kumar, J. P. Patterning of the Drosophila retina by the morphogenetic furrow. Front. Cell Dev. Biol. 11, 1151348 (2023).
Hirose, S., Santhanam, B., Katoh-Kurosawa, M., Shaulsky, G. & Kuspa, A. Allorecognition, via TgrB1 and TgrC1, mediates the transition from unicellularity to multicellularity in the social amoeba Dictyostelium discoideum. Development 142, 3561–3570 (2015).
Sekine, R., Kawata, T. & Muramoto, T. CRISPR/Cas9 mediated targeting of multiple genes in Dictyostelium. Sci. Rep. 8, 8471 (2018).
Lehmann, P., Katoh-Kurasawa, M., Kundert, P. & Shaulsky, G. Going against the family: perturbation of a greenbeard pathway leads to falsebeard cheating. iScience 27, 111125 (2024).
Kundert, P. et al. A GoldenBraid cloning system for synthetic biology in social amoebae. Nucleic Acids Res. 48, 4139–4146 (2020).
Knecht, D. A., Cohen, S. M., Loomis, W. F. & Lodish, H. F. Developmental regulation of Dictyostelium discoideum actin gene fusions carried on low-copy and high-copy transformation vectors. Mol. Cell Biol. 6, 3973–3983 (1986).
Katoh, M. et al. An orderly retreat: Dedifferentiation is a regulated process. Proc. Natl. Acad. Sci. USA 101, 7005–7010 (2004).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Sour. Softw. 3. https://doi.org/10.21105/joss.00861 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Katoh-Kurasawa, M., Shaulsky, G., Trnovec, L. & Zupan, B. Filtered single cell RNA sequencing AnnData objects for Dictyostelium developmental synchronicity study (AX4, acaA, acaA pkaC) (v1.0.0) [Data set]. Zenodo. https://doi.org/10.5281/zenodo.18481907 (2026).
Trnovec, L. scRNA-seq analysis pipeline for dictyostelium developmental synchronicity (v1.0.0). Zenodo. https://doi.org/10.5281/zenodo.18481808 (2026).
Benabentos, R. et al. Polymorphic members of the lag gene family mediate kin discrimination in Dictyostelium. Curr. Biol. 19, 567–572 (2009).
Acknowledgements
This research was supported by the National Science Foundation (grant number 2319686) through the Integrative Organismal Systems program, as well as by the Slovenian Research and Innovation Agency (grants numbers L2-60154 and P2-0209).
Author information
Authors and Affiliations
Contributions
G.S. conceived the study. G.S., R.C., and M.K.-K. designed the experiments. M.K.-K. performed most of the experiments and data collection, with additional data collected by P.L. and sample processing assistance from Y.L. L.T. led the data analysis and software development, with contributions from M.K.-K. and close guidance from B.Z. G.S. wrote the manuscript. All authors revised and edited the manuscript. G.S., R.C. and B.Z. supervised the project. M.K.-K. and L.T. contributed equally to this work. G.S. and B.Z. jointly oversaw key aspects of the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: David Favero and Rosie Bunton-Stasyshyn. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Katoh-Kurasawa, M., Trnovec, L., Lehmann, P. et al. Early cAMP signaling orchestrates single-cell synchronicity throughout Dictyostelium development. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09806-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09806-5