Abstract
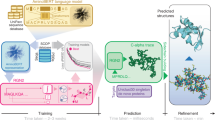
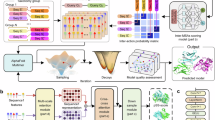
AlphaFold has set a new standard for predicting protein structures from primary sequences; however, it faces challenges with protein complexes across species, engineered proteins, and antigen-antibody interactions, where co-evolutionary signals may be sparse or missing. Herein, we present ProTact, a SE(3)-invariant geometric graph neural network that integrates physics-informed geometric complementarity and trigonometric constraints as inductive biases to enhance protein-protein contact predictions. ProTact is applicable to both experimental and predicted monomer structures and utilizes a modulated key point matching algorithm to approximate accurate docking poses. Experimental evaluations demonstrate that ProTact consistently outperforms state-of-the-art sequence-based and structure-based methods on benchmark datasets, achieving notable relative improvements of 31.63% in average top-10 precision (Precision@10) for CASP 13 and 14 targets and 31.94% for DIPS-Plus datasets on high-quality structures. While performance naturally declines on the more challenging unbound complexes due to large conformational changes, ProTact maintains a competitive edge over baselines. Moreover, when combined with AlphaFold3 as re-scoring functions, ProTact surpasses its default confidence scores, offering over 30.48% improvements in low-MSA contexts. We anticipate that the proposed framework will advance our understanding of protein interactions, functions, and design.
Similar content being viewed by others
Data availability
The datasets used in this study are publicly available: CASP-CAPRI, DB5-Plus, and DIPS-Plus targets were obtained from the DeepInteract repository (https://github.com/BioinfoMachineLearning/DeepInteract), and antibody structures were retrieved from the Structural Antibody Database (SAbDab) (https://opig.stats.ox.ac.uk/webapps/newsabdab/sabdab/). All protein structures were sourced from the RCSB PDB and AlphaFold Protein Structure Database. We used freely available data as described in Methods. The source data behind the graphs in the paper can be found in Supplementary Data 1.
Code availability
The data and code to reproduce the datasets and experiments are available at https://github.com/biomed-AI/ProTact. The specific version of the code used to generate the results presented in this study has been archived in Zenodo55.
References
Altschuh, D., Lesk, A., Bloomer, A. & Klug, A. Correlation of co-ordinated amino acid substitutions with function in viruses related to tobacco mosaic virus. J. Mol. Biol. 193, 693–707 (1987).
Bryant, P., Pozzati, G. & Elofsson, A. Improved prediction of protein-protein interactions using alphafold2. Nat. Commun. 13, 1265 (2022).
Lu, H. et al. Recent advances in the development of protein–protein interactions modulators: mechanisms and clinical trials. Signal Transduct. Target. Ther. 5, 213 (2020).
Arkin, M. R. & Wells, J. A. Small-molecule inhibitors of protein–protein interactions: progressing towards the dream. Nat. Rev. Drug Discov. 3, 301–317 (2004).
Titeca, K., Lemmens, I., Tavernier, J. & Eyckerman, S. Discovering cellular protein-protein interactions: technological strategies and opportunities. Mass Spectrom. Rev. 38, 79–111 (2019).
Jessulat, M. et al. Recent advances in protein–protein interaction prediction: experimental and computational methods. Expert Opin. drug Discov. 6, 921–935 (2011).
Fernández-Recio, J. Prediction of protein binding sites and hot spots. Wiley Interdiscip. Rev. Comput. Mol. Sci. 1, 680–698 (2011).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with alphafold 3. Nature 630, 493–500 (2024).
Krishna, R. et al. Generalized biomolecular modeling and design with rosettafold all-atom. Science 384, 2528 (2024).
Liu, L. et al. Technical report of helixfold3 for biomolecular structure prediction. arXiv preprint arXiv:2408.16975 (2024).
Abanades, B., Georges, G., Bujotzek, A. & Deane, C. M. Ablooper: fast accurate antibody CDR loop structure prediction with accuracy estimation. Bioinformatics 38, 1877–1880 (2022).
Buel, G. R. & Walters, K. J. Can alphafold2 predict the impact of missense mutations on structure? Nat. Struct. Mol. Biol. 29, 1–2 (2022).
Feng, S. et al. Integrated structure prediction of protein–protein docking with experimental restraints using colabdock. Nat. Mach. Intell. 6, 924–935 (2024).
Lu, W. et al. Tankbind: Trigonometry-aware neural networks for drug-protein binding structure prediction. Adv. Neural Inf. Process. Syst. 35, 7236–7249 (2022).
Lu, W. et al. Dynamicbind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model. Nat. Commun. 15, 1071 (2024).
Chelliah, V., Blundell, T. L. & Fernández-Recio, J. Efficient restraints for protein–protein docking by comparison of observed amino acid substitution patterns with those predicted from local environment. J. Mol. Biol. 357, 1669–1682 (2006).
Pierce, B. G. et al. Zdock server: interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Yan, Y., Tao, H., He, J. & Huang, S.-Y. The hdock server for integrated protein–protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Dominguez, C., Boelens, R. & Bonvin, A. M. Haddock: a protein- protein docking approach based on biochemical or biophysical information. J. Am. Chem. Soc. 125, 1731–1737 (2003).
De Vries, S. J., Van Dijk, M. & Bonvin, A. M. The haddock web server for data-driven biomolecular docking. Nat. Protoc. 5, 883–897 (2010).
Stahl, K., Graziadei, A., Dau, T., Brock, O. & Rappsilber, J. Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning. Nat. Biotechnol. 41, 1810–1819 (2023).
Zeng, H. et al. Complexcontact: a web server for inter-protein contact prediction using deep learning. Nucleic Acids Res. 46, 432–437 (2018).
Sanchez-Garcia, R., Sorzano, C. O. S., Carazo, J. M. & Segura, J. Bipspi: a method for the prediction of partner-specific protein–protein interfaces. Bioinformatics 35, 470–477 (2019).
Guo, Z., Liu, J., Skolnick, J. & Cheng, J. Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks. Nat. Commun. 13, 6963 (2022).
Lin, P., Yan, Y. & Huang, S.-Y. Deephomo2. 0: improved protein–protein contact prediction of homodimers by transformer-enhanced deep learning. Brief. Bioinforma. 24, 499 (2023).
Morehead, A., Chen, C., Cheng, J. Geometric transformers for protein interface contact prediction. In: International Conference on Learning Representations (ICLR, 2022). https://openreview.net/forum?id=CS4463zx6Hi
Rao, J. et al. A variational expectation-maximization framework for balanced multi-scale learning of protein and drug interactions. Nat. Commun. 15, 4476 (2024).
Bitbol, A.-F., Dwyer, R. S., Colwell, L. J. & Wingreen, N. S. Inferring interaction partners from protein sequences. Proc. Natl. Acad. Sci. USA 113, 12180–12185 (2016).
Gueudré, T., Baldassi, C., Zamparo, M., Weigt, M. & Pagnani, A. Simultaneous identification of specifically interacting paralogs and interprotein contacts by direct coupling analysis. Proc. Natl. Acad. Sci. USA 113, 12186–12191 (2016).
Szurmant, H. & Weigt, M. Inter-residue, inter-protein and inter-family coevolution: bridging the scales. Curr. Opin. Struct. Biol. 50, 26–32 (2018).
Lawrence, J., Bernal, J. & Witzgall, C. A purely algebraic justification of the kabsch-umeyama algorithm. J. Res. Natl. Inst. Stand. Technol. 124, 1 (2019).
Xu, D. et al. Accurately predicting protein mutational effects via a hierarchical many-body attention network. In: Proc. Annual Conference on Neural Information Processing Systems (NIPS, 2025).
Xie, Z. & Xu, J. Deep graph learning of inter-protein contacts. Bioinformatics 38, 947–953 (2022).
Lin, P., Tao, H., Li, H. & Huang, S.-Y. Protein–protein contact prediction by geometric triangle-aware protein language models. Nat. Mach. Intell. 5, 1275–1284 (2023).
Morehead, A., Chen, C., Sedova, A. & Cheng, J. Dips-plus: the enhanced database of interacting protein structures for interface prediction. Sci. Data 10, 509 (2023).
Lensink, M. F. et al. Blind prediction of homo-and hetero-protein complexes: the casp13-capri experiment. Proteins Struct. Funct. Bioinforma. 87, 1200–1221 (2019).
Lensink, M. F. et al. Prediction of protein assemblies, the next frontier: the casp14-capri experiment. Proteins Struct. Funct. Bioinforma. 89, 1800–1823 (2021).
Si, Y. & Yan, C. Protein language model-embedded geometric graphs power inter-protein contact prediction. Elife 12, 92184 (2024).
He, K., Zhang, X., Ren, S., Sun, J. Deep residual learning for image recognition. In: Proc. IEEE Conference on Computer Vision and Pattern Recognition. 770–778 (CVPR, 2016).
Geiger, M., Smidt, T. E3NN: Euclidean neural networks. arXiv preprint arXiv:2207.09453 (2022).
Tubiana, J., Schneidman-Duhovny, D. & Wolfson, H. J. Scannet: an interpretable geometric deep learning model for structure-based protein binding site prediction. Nat. Methods 19, 730–739 (2022).
Sverrisson, F., Feydy, J., Correia, B.E., Bronstein, M.M. Fast end-to-end learning on protein surfaces. In: Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition. 15272–15281 (CVPR, 2021).
Sorkine-Hornung, O. & Rabinovich, M. Least-squares rigid motion using SVD. Computing 1, 1–5 (2017).
Li, S. et al. Recent advances in predicting protein–protein interactions with the aid of artificial intelligence algorithms. Curr. Opin. Struct. Biol. 73, 102344 (2022).
Kabsch, W. A solution for the best rotation to relate two sets of vectors. Found. Crystallogr. 32, 922–923 (1976).
Liu, S., Liu, C. & Deng, L. Machine learning approaches for protein–protein interaction hot spot prediction: progress and comparative assessment. Molecules 23, 2535 (2018).
Steinegger, M. & Söding, J. MMSEQS2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UNIREF: comprehensive and non-redundant uniprot reference clusters. Bioinformatics 23, 1282–1288 (2007).
Dunbar, J. et al. Sabdab: the structural antibody database. Nucleic Acids Res. 42, 1140–1146 (2014).
Luo, S. et al. Antigen-specific antibody design and optimization with diffusion-based generative models for protein structures. Adv. Neural Inf. Process. Syst. 35, 9754–9767 (2022).
Basu, S. & Wallner, B. Dockq: a quality measure for protein-protein docking models. PloS ONE 11, 0161879 (2016).
Rao, J. biomed-AI/ProTact: ProTact. https://doi.org/10.5281/zenodo.18533525.
Acknowledgements
This study has been supported by the National Natural Science Foundation of China [62041209, 62502553], the Natural Science Foundation of Shanghai [24ZR1440600], the Science and Technology Commission of Shanghai Municipality [24510714300], the China Postdoctoral Science Foundation [2025M771540, GZB20250391], and the Lingang Laboratory Fund [LGL-8888].
Author information
Authors and Affiliations
Contributions
S.Z. and Y.Y. conceived and supervised the project. J.R., D.L., W.L., S.Z., and J.Z. contributed to the algorithm implementation. J.R., D.L., X.Z., W.W., and Q.Y. contributed to the visualization and server implementation. J.R., D.L., Y.R., S.Z., and Y.Y. wrote the manuscript. All authors were involved in the discussion and proofread.
Corresponding authors
Ethics declarations
Competing interests
Y.Y. is an Editorial Board Member for Communications Biology, but was not involved in the editorial review of, nor the decision to publish this article. All the other authors declare no competing interests.
Peer review
Peer review information
: Communications Biology thanks Kundan Sengupta and Zahid Nawaz for their contribution to the peer Communications Biology thanks Gabriele Pozzati and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Laura Rodríguez Pérez.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rao, J., Liu, D., Zhou, X. et al. Accurate protein-protein interactions modeling through physics-informed geometric invariant learning. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09809-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09809-2