Abstract
The transcription factor p53 is exquisitely sensitive and selective to a broad variety of cellular environments. Several studies have reported that oxidative stress weakens the p53-DNA binding affinity for certain promoters depending on the oxidation mechanism. Despite this body of work, the precise mechanisms by which the physiologically relevant DNA-p53 tetramer complex senses cellular stresses caused by H2O2 are still unknown. Here, we employed native mass spectrometry (MS) and ion mobility (IM)-MS coupled to chemical labelling and H2O2-induced oxidation to examine the mechanism of redox regulation of the p53-p21 complex. Our approach has found that two reactive cysteines in p53 protect against H2O2-induced oxidation by forming reversible sulfenates.

Similar content being viewed by others
Introduction
The p53 protein is a transcription factor that regulates the expression of dozens of target genes related to cell cycle checkpoints, DNA repair, and apoptosis1. The human p53 protein contains 393 amino acids (aa) with four functional domains. The disordered N-terminal region (aa 1–42) is a transcriptional activation domain2,3. The C-terminal region contains an oligomerization domain (aa 323–356) and a regulatory domain (aa 360–393). In the central part of p53 is the sequence-specific DNA-binding domain (DBD, aa 91–289)2,3.
Biophysical studies have shown that the isolated p53-DBD can recapitulate properties of the full-length p534,5,6. The DBD not only interacts with specific promoter response elements (RE), but also with many proteins and cofactors7,8,9,10. Approximately 50% of human tumours carry a mutation in TP53, of which 90% are missense mutations in the DBD-encoding sequence11. A consequence of these mutations is the impairment of the structure or the p53-DNA binding activity and/or specificity4. The function of p53 is strictly regulated in the cell by a variety of cellular and molecular mechanisms, such as post-translational modifications or long-range interactions between p53 domains12,13,14,15,16.
Hydrogen peroxide, the most important reactive oxygen species (ROS), has been shown to be capable of triggering both canonical DNA damage response (DDR) and oxidative signaling pathways, resulting in the transcriptional activation of p53 targets17,18,19,20,21,22. Early in vitro and in vivo studies have already shown that p53 binds to promoter sites in reducing conditions, but not in oxidizing conditions23,24,25. Uberti et al. concluded that, following H2O2 insult, oligodendroglia-like cells express the p21 protein in a p53-independent manner20. In H2O2-treated IMR-90 cells, Chen et al. found that p53 and p21 are induced by oxidative signaling rather than DDR pathways26. Barton group has reported that DNA-mediated charge transport, induced by ultraviolet radiation, promotes the oxidative dissociation of p53 from the DNA repair-associated Gadd45 RE through an electron-based mechanism involving the oxidation of a cysteine residue (Cys277 or Cys275) and disulfide formation22. Similarly, Buzek et al. explored p53’s DNA binding in ultraviolet radiation-exposed WS-1 cells, showing its dissociation from the Gadd45 sequence but remaining bound to the cell cycle-regulating p21 RE. Site-directed mutagenesis experiments revealed that Cys277 was responsible for gene-selective transcription18.
Well established bottom-up mass spectrometry (MS)-based proteomics plays a major role deciphering protein composition, abundances, and post-translational modifications27. In the bottom-up MS approach, protein sequences are easily identified and quantified after enzymatic digestion into amenable smaller peptides, whose sequences are commonly determined by liquid-chromatography coupled to tandem mass spectrometry (LC-MS/MS)28. A body of work has shown that Cys oxidation in p53 itself may stabilize and activate p53 in vitro and in vivo. Using a bottom-up MS and a differential alkylation approach, it was discovered that Cys182 and Cys277 in the endogenous p53 of MCF7 and HCA2 cells were responsive to the thiol oxidant diamide29. Similarly, Barton and co-workers identified Cys277 and Cys141 as key residues involved in the oxidative dissociation of the p53-DNA complex22, while Fersht and co-workers reported that Cys141 and Cys124 were the most reactive towards electrophiles30. Further studies have demonstrated that p53 can form intermolecular disulfides with 14-3-3θ and 53BP1 proteins involving Cys27731.
While bottom-up MS has been invaluable in identifying peptide-level modifications, it does not address proteins as intact entities. Proteins in biological systems exist as intact proteoforms, representing the diverse forms in which a protein product from a single gene can be found32. Today’s bottom-up MS technologies, which identify and quantify peptides derived from proteins rather than intact protein themselves, fail to provide protein-level information because the peptide-to-proteoform connectivity is lost33. In contrast to conventional MS-based proteomics, native MS intends to analyze intact proteins and their non-covalent complexes preserving the native-like state34. If combined with tandem mass spectrometry, this approach called native top-down is a powerful tool for higher-order structural characterization of protein complexes35,36,37,38,39. Upon such and many other successes, the past three decades have witnessed remarkable progress in native MS and established a new phase in structural biology40,41,42,43,44,45.
Consistently to previous reports, Langridge-Smith and co-workers using a top-down MS approach identified Cys182 and Cys277 as the most reactive Cys residues in DNA-free p53-DBD towards N-ethylmaleimide (NEM)46. Notwithstanding, the examination of the Cys reactivity against H2O2 in the DNA-p53 tetramer complex at the protein level remains unclear and is one of the aims of this study. For heterogeneous systems, measuring the masses of native complexes may not be sufficient to unambiguously identify structural information. In a seminal work by Clemmer and Jarrold, the capabilities of ion mobility-mass spectrometry (IM-MS) were demonstrated by revealing that a single charge state can present several conformations47. Since then, IM-MS has been employed to study protein systems and characterize their conformation and dynamics48,49,50,51,52. IM-MS is especially useful for analyzing heterogeneous samples and requires less sample preparation than other high-resolution techniques such as X-ray crystallography, NMR, and cryo-electron microscopy. Additionally, many systems that are not amenable to analysis using mentioned structural techniques can be interrogated by IM-MS. Thus, IM-MS has become an established method for investigating biomolecules53,54,55. IM-MS provides information about charge state distribution (CSD) and collision cross section (CCS). Proteins with larger CCS and/or extended conformations undergo a greater number of collisions with the buffer gas, leading to longer drift times or reduced mobility56. However, the resolution of the IM device may not be sufficient to distinguish closely related protein conformations. To aid in this purpose, gas-phase collisional activation (CA) can be used to probe subtle structural differences between similar conformations and study protein ion stability and dynamics57. For example, recording IM-MS experiments with increasing CA can be used to study unfolding mechanisms and estimate lab-frame unfolding energies58,59,60,61,62. These experiments, commonly referred to as collision-induced unfolding (CIU), have provided a practical approach with multiple applications such as uncovering ligand-based stabilization in TTR proteins63, characterization of biotherapeutic antibodies64 or metalloproteins62,65. A quantitative framework to determine activation energy of dissociation/unfolding has been proposed, providing a link between CIU and thermodynamic parameters66.
These conflicting results highlight the need for further experiments to clarify how the tumor transcription factor p53, in complex with DNA, responds to cellular stress induced by H2O2. To this end, we used hybrid MS approaches, including IM-MS, high-resolution native MS, bottom-up MS, and top-down MS. Following Cysteine alkylation, native MS was used to profile reactive Cys residues, and bottom-up/top-down MS identified them in both DNA-free and DNA-bound p53 states. We then monitored the H2O2 oxidation reaction on the DNA-p53 tetramer using native MS, and IM-MS was used to characterize the conformational space and monitor H2O2-induced structural changes. Dimedone, a probe that reacts with sulfenic acids, was then used to validate the H2O2 oxidation mechanism. Combining all this information, we showed how p53 can self-regulate and protect itself against H2O2 fluctuations using its Cys residues.
Results
Mapping the conformational landscape of p53 on DNA binding
The native mass spectrum of p53-DBD, referred to here as “p53” or “WTp53”, presents a bimodal CSD, suggesting the coexistence of at least two conformational families: one compact and globular, and the other more extended. In the absence of DNA, p53 is seen as a free monomer with one Zn(II) bound (Fig. 1A and Table S1). Previous studies have demonstrated that full-length p53 is capable of forming tetrameric complexes in the absence of DNA, as observed with an ultrastable quadruple mutant of p53 or full-length WT p5367,68,69. However, the inherent instability of full-length p53 makes it challenging to conduct experiments. Since isolated p53-DBD can recapitulate many physical properties of full-length p53, and most of the hot spot mutations are located in the DBD, it has become the preferred choice in structural studies4,5,6.
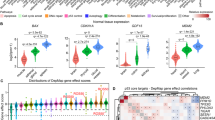
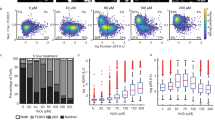
Native mass spectra of WTp53 in the absence (A) and incubated with 0.25 equivalents of p21 RE (E). Travelling wave (TW) ion mobility (IM)-derived collision cross sections (CCS) of quadrupole-selected charge states of WTp53 and (WTp53)4p21 (B, F). The CCS values were calculated from three replicates, and the error bars plot along the CCS axis. Collision-induced unfolding (CIU) heat maps and CCS profiles for WTp539+ (C, D) and (WTp53)4p2120+ (G, H). The PDBs used to illustrate monomer and tetramer complexes are 1OCJ and 3TS8, respectively.
The charge state (z) region from 8 ≤ z ≤ 10 is assigned to a compact conformation that spans only three charge states, while the higher charge states (ranging 11 ≤ z ≤ 20), accounting for 2–5% of relative intensities, are assigned to forms presenting more disordered conformations. The Rayleigh limit (ZR) permits us to estimate the theoretical maximum number of positive charges that a globular protein can accommodate70, and it was found here to be 12. The Rayleigh limit also demarks the transition between compact and extended conformations (Fig. S1A).
To examine the conformers of p53 in more detail, we employed traveling wave ion mobility (TWIM)-MS. This allowed us to acquire arrival time distributions for each charge state, which were converted to TWCCSN2 using a TWIMS calibration procedure. Extended conformations or ions with larger CCS undergo a greater number of collisions with the buffer gas and, as a result, have a longer drift time and reduced mobility than those that are more compact. The capability of IM-MS to separate conformational states has brought a powerful addition to the structural and dynamic interrogation of heterogeneous protein systems, which are not readily analyzed by other structural techniques71,72,73,74,75. TWIM-derived CCS distributions confirm that the four charge states with 8 ≤ z ≤ 11 populate a similar compact conformation, with TWCCSN2 values spanning ~130 Å2 (Figs. 1B, S1 and Table S2). Comparing the experimental TWCCSN2 values with the theoretical CCS (PDB: 1OCJ, 2792 Å2) calculated using the Projection Approximation using Rough Circular Shapes (PARCS) algorithm76 shows a slightly gas-phase collapse of the native-like p53 conformations, in agreement with previous data68,77. For ions with z > 11, multiple unfolded conformations are observed, and their size increases up to ~4000 Å2 for z = 18, which is consistent with previous measurements9.
Gas-phase activation of protein ions via CA can probe subtle structural differences between similar conformations, estimate ion stabilities and dynamics, and characterize unfolding mechanisms78. IM-MS under different CA conditions, recorded for mass-selected WTp539+ ions, reveals a unfolding mechanism via a transition from the native compact α conformation with TWCCSN2 ~ 2173 Å2 to an unfolded γ conformer with TWCCSN2 ~ 2700 Å2, passing through a β intermediate conformation (Fig. 1C, D).
In the presence of DNA (p21 RE), WTp53 forms a (Zn-p53)4DNA tetramer complex appearing at 40–50% relative intensity, most likely via (Zn-p53)2DNA dimer assembly, as this form is also observed (Fig. 1E). Monomeric disordered species are now present at levels lower than 1% of the relative intensity, suggesting that the remaining unbound p53 species are in a compact conformation. The conformational properties of the tetramer were examined by analyzing TWIM-derived CCS distributions of individual charge states, which showed a well-conserved single conformer with TWCCSN2 ~ 5700 Å2 in all cases (Fig. 1F and Table S2). A close agreement was found between theoretical (PDB: 3TS8, 5644 Å2) and experimental CCS values for (Zn-p53)4DNA tetramer complex.
To compare the conformational diversity between monomeric and tetrameric species, we calculated the full width half maximum (∆TWCCSN2) (Table S2). The p53 monomer exhibits ∆TWCCSN2 ranging from 26 to 36 Å2, while the (p53)4DNA complex displays ∆TWCCSN2 values in the 67–74 Å2 range (Table S2). To understand the conformational changes of monomeric p53 upon DNA binding, we normalized the ∆TWCCSN2 to a single monomeric specie. When the ∆TWCCSN2 values of the (p53)4DNA complex are normalized to a single monomeric specie (∆TWCCSN2/n° subunits), they lead to ∆TWCCSN2 values of 16–19 Å2. As these values are much lower than the ∆TWCCSN2 values for p53 monomer (ranging from 26 to 36 Å2), we conclude that the monomer adopts a more compact conformation when bound to DNA. These results align with previous molecular dynamics simulations, which have reported a DNA-induced conformational change in p53, leading to a less solvent-accessible and compact conformation10. Specifically, these simulations identified that conformational changes in the L1 loop at the DNA-binding interface are coupled to shifts in S6-S7 loop. This regulatory mechanism of p53 upon DNA binding serves to protect p53 from interactions with partners that could potentially inhibit its transcription-dependent functions. Gas-phase CA of WT(p53)4DNA20+ ions reveals a complex multi-step unfolding mechanism (Fig. 1G, H), in contrast to the relatively simply unfolding process seen at the monomer level (Fig. 1C). We observe that the tetrameric DNA-p53 complex populate partially unfolded intermediates that are stable on the millisecond timescale. Atomic force microscopy studies have also revealed that binding to DNA alters the mechanical unfolding pathway, primarily due to the formation of hydrogen bonds with DNA, which leads to structural stabilization79.
To further support the idea that native MS and IM-MS can reproduce the structural features of p53, we performed identical experiments on one of the most common hot-spot mutations in p53, namely, R248Q80. The crystal structure of p53-DBD in complex with DNA reveals that R248 interacts with the minor groove of DNA, and therefore, R248Q is referred to as a DNA-contact mutant3. Our results show that R248Q exhibits a similar bimodal CSD in the native mass spectrum and a two-step mechanism with larger unfolding energies compared to the WT (Fig. S2). Furthermore, incubation of R248Qp53 with different stoichiometries of two RE, p21 and mdm2, does not progress to the formation of the tetrameric DNA-p53 complex (Fig. S3). Altogether, this data, in agreement with previous results, demonstrates that the mutation disrupts p53-DNA binding properties and confirms that native MS can accurately reproduce not only the specific stoichiometry of the (Zn-p53)4DNA complex, but also retains the structural features observed in solution.
Profiling reactive Cys residues in the (Zn-p53)4DNA complex
Cellular levels of p53 are tightly controlled by ubiquitination and proteasomal degradation81, and under cellular oxidative stress induced by ROS, p53 can be activated to regulate transcription of genes involved in cell cycle as p21 or DNA repair as Gadd45, among others17,18,19,20,21,22. As cysteines are a preferred target for redox-linked regulatory modifications82, we profiled reactive Cys residues with native MS and chemical labelling with NEM. The p53 monomer contains ten Cys residues, from which three (Cys176, Cys238, and Cys242) together with His179 coordinate a structural Zn(II)83. Focusing on the isolated WTp53 first, two/three Cys react fast upon NEM incubation without alteration of the Zn(II) binding properties (Fig. S4A). As the reaction proceeds, up to five Cys residues are modified, forming Zn-p53(NEM)5 complex (Fig. S4B, C), suggesting that from the remaining five Cys residues, three coordinate Zn(II) and two are less accessible to the solvent. Then, we observed a cooperative modification mechanism whereby Zn(II) dissociates, and all of Cys residues were NEM-labelled leading to apo-p53(NEM)10 (Fig. S4D). In agreement with the structural role of Zn(II), the protein presents in higher charge states and more extended forms at this stage. Conflicting results have been published regarding the identification of reactive Cys in p5321,22,29,46. Fersht and co-workers performed Cys alkylation on p53-DBD and full-length p53 in the absence of DNA and determined by bottom-up MS that Cys141 and Cys124 were the most reactive30. Langridge-Smith and co-workers used NEM to label reactive Cys on DNA-free p53-DBD, and top-down MS identified Cys182 and Cys277 as NEM-labelled46. Barton and co-workers, using ultraviolet radiation, determined that Cys141 may be involved in the DNA-mediated oxidation DNA(Gadd45)-p53 complex21. In a later study, they reported that Cys277 is required for the oxidative dissociation of the p53-DNA complex and that mutation of Cys275 lowers p53’s affinity for DNA22. Taken together, these reports show how intricate is to attempt to unravel the mechanistic details of p53 oxidation at the molecular level.
Native mass spectrometry was used to examine the reactivity of the Cys residues in the (Zn-p53)4DNA complex formed upon DNA(p21) incubation (Fig. 2). As in the case of the free monomer, two Cys residues reacted fast upon NEM addition, and the p53NEM2 was able to form a DNA-p53 tetramer complex (Fig. 2A, B). CIU experiments for WTp53NEM29+ revealed a CCS increase of ~150 Å2, a sharp unfolding transition with absence of β intermediate, and increased gas-phase stability (from 280 to 308 eV) than for WTp539+ ions (Fig. 2C). Although these gas-phase stabilities are specific to the instrument and experimental conditions used66, it allowed us to estimate relative stabilities between ions of the same charge state.
Native mass spectrum of WTp53 incubated with 0.25 equivalents of p21 RE and 0.5 mM NEM recorded after 0, 5, 15, and 25 min of reaction (A). Plot of individual signal intensities as a function of the reaction time (B). Collision-induced unfolding (CIU) heat maps for quadrupole-selected p53NEM29+ (C). Schematic representation of reaction mechanism (D). Purple and yellow spheres represent Zn(II) ions and S atoms from Cys residues, respectively. The orange moiety indicates a NEM modification.
Such a shift towards larger CCS values was also observed for the WT(p53NEM2)4DNA19+/20+ ions compared with WT(p53)4DNA19+/20+ (Fig. S5). This may suggest that NEM binds to surface-exposed Cys residues, resulting in an increased size and stability of the protein complex. Interestingly, binding the third NEM moiety to p53 almost completely disrupts p53-DNA binding, while Zn(II) remains bound to p53 (Fig. 2A). Further alkylation triggers Zn(II) dissociation and protein unfolding. From these results, we may conclude that two reactive Cys residues in p53 are partially involved in DNA binding, as evidenced by a 10–20% decrease in the relative intensity of the complex, and the third one completely abrogates p53-DNA complex formation. (Fig. 2D).
In order to localize the Cys residues harboring NEM, we performed native top-down MS experiments in which the intact protein ions were quadrupole-selected and subjected to collision-induced dissociation (CID) (Fig. 3A). We observe a CID spectrum mostly dominated by N-terminal b-fragment ions, where Cys124, Cys135, Cys141 residues were NEM-free (Fig. 3B). We found two characteristic y-fragment ions that localize Cys277-NEM labeled residue (inset Fig. 3A). Calculating the electrostatic potential shows that identified Cys277-NEM labeled lies in the positive electrostatic potential DNA-binding interface region (Fig. 3C). While electrostatic potential values provide valuable insights, other factors such as steric hindrance (bulkiness around the nucleophilic atom), solvent, charge and protein stability effects also influence nucleophilicity84. The Cys277 residue is located in the loop S10-H2 and shows higher solvent-accessible surface area (SASA) than median (Fig. 3D).
Native top-down mass spectrum of quadrupole-selected p53NEM210+ ions obtained after incubation of the WTp53 with 0.25 equivalents of p21 RE and 0.5 mM NEM for 5 min (A). Fragment location map from native top-down MS where the Cys277 modified by NEM is indicated by yellow (B). Electrostatic potential calculated for PDB:1OCJ (C). Solvent-accessible surface area (SASA) values for the Cys residues in p53 (D). The dashed line indicates the median SASA value. The MS/MS spectra supporting the annotation of NEM modification on Cys277 (E) and Cys182 (F). The Cys residues in p53 are coloured in yellow, and Cys NEM-labeled residues identified are marked in yellow in the peptide.
Considering previous data18,22,29,30,31,85, we can conclude that Cys277, a residue in close contact with DNA, is a reactive residue in DNA-bound p53 states although not critical for p53 function per se. Indeed, the results show that Cys277 alkylation did not lead to a complete abrogation of the tetrameric p53-DNA complex. Our data agree with in vitro cell studies that observed that the p53 C277S mutant was still transcriptionally activated18.
Using top-down MS, we did not localize the second NEM modification. However, considering that Cys124, Cys135, Cys141 residues were NEM-free and Cys176, Cys238, and Cys242 binds Zn(II), either Cys182, Cys229 or Cys275 was labeled by NEM (Fig. 3B). By employing a bottom-up MS approach, in which the protein is digested into peptides, and their masses are then measured, we found that Cys182 and Cys277 were labelled by NEM (Fig. 3E, F). The reactivity of Cys182 towards NEM can be rationalize by its high SASA values, allowing it to easily react with NEM (Fig. 3D). Our results are in agreement with Langridge-Smith and co-workers, that also identified Cys182 and Cys277 as the most reactive Cys residues in DNA-free p53-DBD46.
In an effort to reactivate p53 mutants, Cys alkylation has gained attention, and several small electrophiles have been developed30,85. Therefore, we tested the capabilities of NEM to reactivate R248Qp53, a mutant with impaired DNA binding properties. As in the case of WT-p53, two Cys were alkylated by NEM, and the fragments obtained by native top-down localize NEM in Cys277. Upon Cys residue alkylation, the protein did not form a complex with DNA, leading us to conclude that alkylation of the Cys277 residue impair p53-DNA binding properties rather than reactivate them (Fig. S6).
Discerning the protective role of reactive Cys against oxidation
Once we characterized the Cys residues reactivity in the DNA-p53 complex, we aimed to answer the following question: how does (Zn-p53)4DNA(p21) tetramer respond to an H2O2 insult? As abovementioned, several reports tried to address this issue, leading to ambiguous results mainly because it is not clear if the p21 protein is expressed in a p53-independent or dependent manner. Among ROS, H2O2 is an important intracellular messenger86. The moderate reactivity and high activation energies of the reaction with Cys residues results in a free H2O2 diffusion in the cell. As a consequence, it may reach long distances and target less surface-exposed proteins. In a first attempt, we incubated DNA-free p53 at increasing H2O2 concentrations and recorded a native MS spectrum after 5 min of reaction. Using a far beyond physiological H2O2/p53 molar ratios, H2O2 oxidized two thiolate anions to the sulfenate form (Cys-SO−, ∆m = 16 Da each one), which further reacted with excess H2O2 to yield two irreversible sulfinate species (Cys-SO2−, ∆m = 32 Da each one) (Fig. S7). Using an excess of ~200 molar equivalents of H2O2 to p53, which aligns with relative concentrations found in vivo87, sulfenic acid (Cys-SOH) formation was accompanied by the formation of disulfides, and any sulfinate species were detected (Fig. 4). Interestingly, the CSD shifted towards larger z and closely resembled that obtained for metal-free p53, albeit with Zn(II) remaining bound to p53 (Fig. 4A–C). This suggests a solution-phase conformational change altering SASA is induced by disulfide formation. Indeed, a second population with lower IM-derived CCS values was obtained for (apo-p53)2S-S9+ ions (where S-S stands for disulfide) than for Zn-p53, indicating that disulfide formation yields to more-compact and ordered conformations (Fig. 4D, E). As we were able to work under controlled oxidation conditions, we investigated the mechanism of oxidation by high-resolution native MS, which enabled real-time monitoring of the reaction (Fig. 4F, G). According to our results, the oxidation mechanism of Zn-p53 by H2O2 leads to the formation of bis-disulfides (apo-p53)2S-S and proceeds by a stepwise mechanism (Figs. 4F and S8). Accurate mass measurements allowed us to conclude that the formation of bis-disulfides starts by the attack of two cysteine residues on two H2O2 molecules to yield two sulfenic acids ((Zn-p53)2Cys-OH), which reacts with Zn(II)-binding cysteine residues with concomitant Zn(II) dissociation and disulfide formation ((apo-p53)2S-S) (Fig. S8).
Native mass spectrum of WTp53 (A) incubated with 1 mM EDTA (B) or 2 mM H2O2 (C). IM-MS monitored the H2O2 oxidation of WTp53 over time (D). Travelling wave (TW) ion mobility (IM)-derived collision cross sections (CCS) of quadrupole-selected 9+ ions of Zn-p53 and (apo-p53)2S-S (E) Calculated relative abundances from the mass spectrometry data in (F) and globally fitted to a two consecutive reactions A → B → C (G). Individual steps in the H2O2 oxidation mechanism of WTp53 (H). Purple and yellow spheres represent Zn(II) ions and S atoms from Cys residues, respectively.
The formation of the disulfide product induces a conformational change, as evidenced by IM-MS. The observation of the (Zn-p53)2Cys-OH intermediate is in good agreement with our NEM results that determined the presence of two reactive Cys residues in p53. The calculated relative abundances from the mass spectrometry data were globally fitted to two consecutive reactions A → B → C, where A is Zn-p53, B is (Zn-p53)2Cys-OH, and C is (apo-p53)2S-S (Fig. 4). The observed first-order rate constants (kobs) were 0.09 s−1 and 0.08 s−1 at 523 K. Although qualitatively satisfactory, the data may not be directly compared with other experimental solvated data. We have based it on a single charge state, and we are then assuming this is equally representative of the protein in solution and that the ionization properties of the modified species are comparable during the calculation of relative abundances, and we ignore other factors (e.g., overlapping of isotopic distributions or estimation of S/N ratio), all of which are a potential source of errors that can alter the kobs estimation. Nevertheless, it can be clearly seen that the concentration of Zn-p53 decreases exponentially while (Zn-p53)2Cys-OH increases reaching a maximum value, and the product (apo-p53)2S-S increased continuously (Fig. 4). As the reaction proceed, (apo-p53)2S-S cooperatively reacts with H2O2 to yield (apo-p53)5S-S via a sulfenic acid intermediate (apo-p53)2S-S,2Cys-OH (Figs. S8 and 4G). To further confirm the formation of disulfides, aliquots of the reaction mixture were taken at various time points, and free Cys labelled with NEM (Fig. S9A–C). In the absence of H2O2, ten Cys residues were modified by NEM, indicating that all the Cys residues were in their reduced form (Cys-SH) (Fig. S9D). Upon H2O2 reaction, the species with 4, 6, and 8 NEM appeared, indicating that 6, 4, and 2 Cys residues were oxidized (Fig. S9E). Increasing H2O2 concentration resulted in the increase of apo-p53(NEM)6, suggesting a preference formation of a bis-disulfide product, as hypothesized in a previous report88.
Finally, we examine the H2O2 oxidation of the (Zn-p53)4DNA(p21) tetramer. Time-resolved measurements were used to monitor the relative intensities of the species involved in the reaction (Fig. 5). As in the prior experiment, two sulfenic acids are rapidly formed in both the DNA-free ((Zn-p53)2Cys-OH) and DNA-bound ((Zn-p53)2Cys-OH)4DNA(p21) p53 states (Fig. 5A–C). Moreover, the detection of signals with ∆m = 64 Da only in the DNA-bound complexes suggests either Cys oxidation in p53 or oxidative DNA damage, likely producing oxidized guanines (Fig. 5A). Guanine has the lowest oxidation potential compared to the other DNA bases and is therefore, the most readily oxidized base89. Indeed, biologically relevant 8-oxo guanine patterns have been mapped in human cells treated with physiological concentrations of H2O2, resembling two mutational signatures found in cancer89. Monitoring the mass distribution of the DNA-p53 complexes and monomeric p53 formation over time allowed us to calculate relative abundances and propose a plausible oxidation mechanism (Fig. 5C). The formation of ((Zn-p53)2Cys-OH)4DNA(p21) is followed by an oxidative dissociation, where the sulfenic acids in p53 react with Zn(II)-binding cysteine residues, resulting in concomitant Zn(II) dissociation and disulfide formation, which leads to dissociation from a promoter sequence for the p21 gene (Fig. 5C). The (apo-p53)2S-S species are eventually further oxidized by H2O2, yielding (apo-p53)5S-S. We next examined whether these mechanisms also occur in the presence of zinc metallothionein-2 (MT), a H2O2 scavenger90. The role of MT in redox processes is evidenced by the fact that its production increases in response to various oxidative stressors91,92. Additionally, cells that overexpress MT exhibit enhanced protection against oxidative stress93,94,95. Using 200:2:1 molar equivalents of H2O2:Zn7MT2:p53, we observed the rapid dissociation of six out of seven Zn(II) ions from MT2 and the formation of two sulfenic acids in Zn-p53 (Fig. S10A). Within 10 min of reaction with H2O2, all 20 cysteines of MT2 are oxidized, leading to the concomitant dissociation of all seven zinc ions (Fig. S10B). The sulfenic acids in p53 react as above with the formation of (apo-p53)5S-S. These results reflect the role of the two reactive Cys residues in protecting p53 against H2O2.
Native mass spectrum of WTp53 pre-incubated with 0.25 equivalents of p21 RE incubated with 2 mM H2O2 was monitored over time (A). Peaks were assigned for monomeric p5314+ species and for (p53)4p2119+. Calculated relative abundances from the time-resolved mass spectrometry data (B). Individual steps in the H2O2 oxidation mechanism of DNA-p53 tetramer complex (C). Initially, H2O2 oxidizes the ((Zn-p53))4DNA(p21) complex, forming Cys sulfenic acids (Cys-OH). Under physiological conditions, sulfenic acids in p53 react with non Zn(II)-binding cysteine residues, forming reversible disulfides that can be reduced by the buffer redox agents. Under oxidative conditions, when PRDX/TRX becomes oxidized, the complex undergoes oxidative dissociation. Here, sulfenic acids in p53 react with Zn(II)-binding cysteine residues, resulting in Zn(II) dissociation and disulfide formation, ultimately leading to dissociation from the promotor sequence. Zn(II), S atoms from Cys residues, and O atoms from H2O2 are represented by purple, yellow and red spheres, respectively.
To probe the existence of two sulfenic acids as intermediates in the H2O2 oxidation reaction and validate the reaction mechanism, we used dimedone, a compound that selectively traps sulfenic acids90. Direct injection of the reaction mixture resulted in unresolved charge state distributions, potentially leading to inaccurate mass assignments. For improved resolution and mass accuracy, the reaction mixture was separated and analyzed by a top-down liquid chromatography-mass spectrometry (LC-MS). The LC-MS chromatogram revealed the presence of two distinct populations: one composed of p53 molecules that did not react with H2O2, and the other comprising molecules that were oxidized and reacted with dimedone (Fig. 6A, B). Deconvolution of the mass spectrum and mass simulation revealed that two Cys residues reacted with dimedone, confirming that (Zn-p53)2Cys-OH is a main intermediate in the H2O2 oxidation reaction (Fig. 6C, D). To localize these dimedone modifications, we employed a top-down and bottom-up MS approach. Intact MS/MS measurements did not lead to any informative fragment that can map the Cys modifications (Fig. S11). However, by using a bottom-up MS approach we were able to annotate peptides in which Cys141 and Cys182 were modified by dimedone (Fig. 6E, F). Peptides covering the Cys141 and Cys182 sequence were manually checked and showed nearly complete sequence coverage, validating the mapped modifications (Fig. 6E, F). In contrast to the nucleophilic addition reaction with NEM, Cys141, not Cys277, was the reactive residue towards H2O2. Redox-sensitive Cys residues are often deprotonated at physiological pH to undergo sulfenation90. In fact, the energy barrier for the oxidation of a thiol by H2O2 (~50 kcal∙mol−1) is two times higher than of a thiolate (~28 kcal∙mol−1)91. Calculations of the pKa values for the Cys residues using PropKa analysis revealed that Cys182 (pKa 7.55), but not Cys141 (pKa 11), is an acidic cysteine residue. Considering the combined evidence from mass spectrometry and computational data, we can hypothesize that, owing its high SASA value (Fig. 3D) and acidic character, Cys182 in its thiolate form initiates the H2O2 oxidation reaction. The protein conformational changes induced by this reaction alter the protein microenvironment, lowering the Cys141 pKa and triggering the oxidation cascade.
Base peak chromatogram (BPC) from the intact LC-MS analysis of WTp53 reaction with H2O2 and incubated with dimedone (30 μM, 2 mM, and 5 mM concentrations, respectively) (A). Mass spectrum for the scans corresponding to the modified protein shown in the BPC (B). Deconvoluted spectrum and simulated isotopic distributions for the three species annotated (C). Schematic representation of the workflow used to trap sulfenic acids by dimedone (D). MS/MS spectra for the TCPVQLWVDSTPPPGTR140–156 peptide (E) and CSDSDGLAPPQHLIR182–196 peptide (F) that were identified with Cys141 and Cys182 dimedone-labeled, respectively.
Discussion
Different mechanisms are involved in the stabilization of p53 and the transcriptional activation of target genes responsible for DNA damage repair, cell cycle arrest, or apoptosis, by both redox signaling and DNA damage. Hydrogen peroxide, the most important ROS in redox signaling, has been shown capable of triggering both canonical DDR pathways and, through upstream ATM and stress-activated protein kinases JNK and p38MAPK pathways, leading to the phosphorylation and stabilization of p5317. A body of work has studied the effect of H2O2 on p53 showing that different mechanisms are involved in p53 stabilization and activation of target genes involved in DNA damage repair, cell cycle arrest, or apoptosis by redox signaling and DNA damage. For instance, early in vitro and in vivo reports have shown that p53 binds to promoter sites under reducing conditions but not under oxidizing conditions23,24,25. More recently, Barton and co-workers determined that p53 is likely irreversible oxidized when HeLa cells are incubated with millimolar H2O2 concentrations, although its transcriptional activity was not assessed21. In this sense, Uberti et al. showed that H2O2 induces a p53 nuclear translocation in oligodendroglia-like (OLN 93) cells and that they constitutively express p21 in a p53-independent manner20. However, it is not clear whether the observed activation of p53 by H2O2 is mediated through DDR, redox signaling, or both. Chen et al. addressed this question and found that H2O2-treated IMR-90 cells induced p53 and p21, while the levels of oxoG in DNA remained stable, indicating that DDR was not responsible for p53 and p21 activation26. Barton and co-workers, using electrophoretic mobility shift assays, determined that DNA-mediated charge transport (CT) can sequence-selectively promote the oxidative dissociation of p53 bound to the Gadd45 sequence, while p53 remains unaltered on the p21 promoter21. Using a bottom-up proteomics approach, they concluded that Cys141 forms a disulfide bond with an unlocalized Cys residue, promoting p53 dissociation from DNA. In a later work22, the authors employed a more sensitive mass spectrometry assay based on multiple reaction monitoring, which elucidated that Cys277 or Cys275, rather than Cys141, are involved in disulfide formation and p53 dissociation from DNA. Consistent with their findings, Buzek et al. using a bio-oligo pull-down DNA binding assay, investigated the DNA binding activity of p53 on ultraviolet-radiated WS-1 human fibroblast cells18. They found that p53 dissociates from the Gadd45 sequence but not from the p21 response element (RE), and that Cys277 is critical for the differential affinity of p53 to distinct REs.
Besides signaling pathways upstream of p53 or controlled by redox signaling, Cys oxidation by H2O2 in p53 itself has also been suggested to stabilize and activate p53. For instance, APR-249 (an alkylating reagent tested in a phase I/IIa clinical trial) reactivates mutant p53 by targeting Cys124 and Cys27785. Many other small molecules have been developed to reactivate mutant p53 proteins by covalently binding to cysteines, such as CP-31398, HO-3867,KSS-9, MIRA-1, PK11007, and STIMA-1. Several attempts have been made to identify the redox-active Cys residues in p53. Held et al. used a differential alkylation approach with the alkylating reagent NEM and found that Cys182 and Cys277 in endogenous p53 of MCF7 and HCA2 cells were sensitive to the thiol oxidant diamide29. However, the specific type of reversible cysteine oxidation remained to be determined.
Consistently, Scotcher et al. identified that Cys182 and Cys277 in monomeric p53 were reactive to the alkylating reagent NEM in vitro46. The metal binding status of p53 has also been linked to the sensitivity for Cys oxidation. Wu and Momand showed that at least one Cys residue per p53 molecule was highly sensitive to oxidation when MCF7 cells were treated with pyrrolidine dithiocarbamate (PDTC), a metal chelator96. PDTC treatment correlated with the oxidation state of Cys residues on p53 in vivo97. This body of work clearly indicates that Cys oxidation by H2O2 in p53 itself may play a role in p53 stabilization and activation in vivo. Despite being a subject of study for more than 25 years, yet it is not fully understood how Cys oxidation in p53 regulates p53 stability and function. This is partly due to the milieu of components involved, which hinders the isolation and study of Cys oxidation by H2O2 from DDR or redox signaling. This complexity is our rationale for conducting the study in vitro. When coupled with high-resolution native MS, we are able to resolve a chemical reaction that likely could not be done by other means due to the heterogeneity of the protein and reaction mixture.
A major concern and possible limitation of our conclusions may relate to the amount of H2O2 used in the experiments. Physiological H2O2 concentration is estimated to be buffered in the low nanomolar range, typically between 1 and 100 nM98. However, these estimations are based on quantifying extracellular H2O2, followed by calculating intracellular H2O2 concentration using a theoretical 100-fold extracellular-to-intracellular peroxide gradient99. Recent estimations, utilizing genetically encoded H2O2 probes like HyPer-399, have reported even higher gradient concentrations, up to 650-fold100. However, it has been noted that there are intracellular spaces with even zero H2O2 concentration, which would make the gradient tend toward infinity101. Overall, rough estimations of the average basal H2O2 level and its fluctuations suggest different intracellular H2O2 concentrations, ranging from the nano to even picomolar range102,103. Under oxidative stress, however, H2O2 levels can increase up to 10,000 nM98. Our in vitro experiments used H2O2 concentrations much higher than physiological levels in absolute terms. Additionally, we employed a p53 concentration a million times higher than the reported physiological range (0.5–2 ng/ml)104. Considering that H2O2 is estimated to be between 10–100 nM and p53 between 0.005 and 0.04 nM, there is an excess of 250 to 20,000 molar equivalents of H2O2 to p53. In relative terms, we employed an excess of ~200 molar equivalents of H2O2 to p53, which aligns with relative concentrations found in vivo. Furthermore, our in vitro experiments appear to capture two different oxidation mechanisms. Using a far beyond physiological H2O2/p53 molar ratios, we observed that Cys oxidation rapidly forms two irreversible sulfinate species. However, using a seemingly reasonable H2O2/p53 molar ratios, the formed sulfenic acids do not react with an excess of H2O2 to form sulfinate species but instead react with free thiols to form disulfides—an oxidation product that can be reversed by the action of the redox buffers present in the cell, such as TRX or GRX.
There is a large body of work showing that oxidation of zinc finger transcription factors occurs both in vitro and in vivo. These include heat shock protein 33105,106, anti-sigma factor RsrA, protein kinase C and replication protein A107. In the above examples, H2O2 sensing occurs through the oxidation of the zinc-binding Cys residues. The zinc finger NEMO required for NF-kB activation is also target for H2O2, likely through Cys179 oxidation.108. H2O2 has also been demonstrated to oxidize cysteinyl thiols, inducing the formation of sulfenic acids and disulfide bonds in various protein targets such as arylamine N-acetyl transferase-1, indoleamine 2,3-dioxigenase, phospholipase A1, SUMO E1 UBA2 and PTP1B109 as well as the bacterial transcription factor OxyR110.
Another major question to consider is whether or not p53 can be oxidized in the nucleus in the presence of peroxidases, which contain catalytic Cys residues that react much faster than Cys residues in p53 with H2O2. Shi and Dansen excellently reviewed two PRDX models, the flood-gate model and the PRDX relay model111. Briefly, the PRDX model provides an explanation of how intrinsically unreactive protein thiols can be oxidized in response to low H2O2 concentration, as observed in proteomics studies. In this model, redox signaling starts at very low H2O2 concentration, oxidizing PRDX. Redox signaling gradually changes to damage signaling when PRDX becomes overoxidized and H2O2 is able to escape their scavenging function. Even though the intrinsic reactivity of the Cys residues in p53 is much lower compared to Cys residues in PRDX and TRX, as evidenced above, there is a large body of evidence that p53 itself is redox-sensitive in vivo.
The mechanism by which H2O2 regulates the stability and activity of the DNA-p53 complex through Cys residues in p53 itself remains unknown and was a motivation for our study. By applying ion mobility-mass spectrometry, we first characterized the gas-phase conformational landscape and structural properties of the p53 monomer, and DNA-p53 tetramer. Then, using NEM, we found that p53 contains two reactive Cys residues in both DNA-free and DNA-bound states. Using top-down and bottom-up MS approaches, we identified these residues as Cys277 and Cys182. Next, we monitored the H2O2 oxidation reaction on the DNA-p53 tetramer in real-time using native MS. Under physiologically relevant H2O2/p53 molar ratios, we observed that two cysteine residues attacked H2O2 molecules to yield two sulfenic acid intermediates per monomer. These intermediates later reacted with Zn(II)-binding cysteine residues, resulting in concomitant Zn(II) dissociation and disulfide formation, ultimately leading to the loss of transcriptional activity. To validate the proposed H2O2 oxidation mechanism, we used dimedone, a reagent that selectively reacts with sulfenic acids. We indeed observed the formation of two sulfenic acids, which were later localized to be formed at Cys141 and Cys182. Due to the high SASA value and acidic character of Cys182, we hypothesized that the mechanism starts with Cys182 attacking H2O2, accompanied by a conformational change that lowers the pKa of Cys141 and triggers the oxidation cascade. Despite losing its transcriptional activity when oxidized, the apoprotein can be restored to its functional state under normal cellular conditions through the action of redox (including TRX, GRX) and metal buffers (such as metallothioneins, ZnT, and Zip zinc transporters). Initially, the disulfides can be reversed to thiols/thiolates by the action of TRX or GRX. Considering the cellular free Zn(II) concentration ranging from 10−11 to 10−9 M112, and the metal binding properties of p53 (Kd ~ 10−15 M)113, it is anticipated that reduced p53 will undergo remetallation. Our work evidences how p53 can self-regulate and defend itself against H2O2 fluctuations through its cysteine residues and reconciles the previously published data.
Methods
Expression and purification of WTp53, R248Qp53 and metallothionein
The expression vector pET15B encoding the human p53-DBD (residues 91-312) was a gift from Cheryl Arrowsmith (Addgene plasmid # 24866).114. Metallothionein-2 was overexpressed in E. coli cells (expression vector deposited in Addgene, plasmid ID 105693). The protein was overexpressed and purified in bacterial system as described in the Supporting information.
Nanoelectrospray Ionization
All MS and IM-MS experiments were conducted using nanoelectrospray and samples prepared at 5–10 µM in 200 mM ammonium acetate (pH 6.8) and desalted using micro Bio-Spin 6 columns (Bio-Rad) prior to any experiment. The samples were then ionized from a borosilicate glass capillary (O.D. 1.2 mm, I.D. 0.9 mm, World Precision Instruments, Stevenage, UK) produced in-house using a Flaming/Brown P-1000 micropipette puller (Sutter Instrument Co., Novato, CA, USA). Ions were produced by applying a positive potential of 0.9–1.4 kV via a platinum wire (Goodfellow).
High resolution native MS and top-down MS
MS experiments were performed on a Q-Exactive UHMR Orbitrap mass spectrometer (Thermo-Fisher Scientific, Bremen, Germany) equipped with nESI source. The spray voltage were in the 1–1.4 kV range, source temperature was set at 250 °C, and the S-lens RF was set at 100. No in-source trapping was applied. The ions were transported to the HCD cell with an injection energy of 10 eV where they were cool down by doubling the N2 gas pressure in the HCD cell (trapping gas pressure 2) compared to the standard value. Full MS scan data were acquired with the noise threshold parameter set to 3.64 at a set resolution of 12,000 and 200,000 at m/z 400. The native mass spectra was processed using Xcalibur 4.1 and in house Python 3.5 scripts.
Native top-down experiments were performed in a Synapt XS HDMS by quadrupole-selecting particular ions and increasing the trap collision energy. Data was analyzed using MassLynx v4.2 (Waters Corp., UK) and masses assigned using ProSight Lite115. Electrostatic surface was calculated by USCF Chimera116 using the PDB:1OCJ. Propka 2.0 was used to calculate pKa values117.
Bottom-up MS
To perform bottom-up MS experiments, N-ethylmaleimide- or dimedone-labelled p53 proteins were buffer exchanged to 100 mM ammonium bicarbonate. Trypsin (Sigma-Aldrich) was added at 1:1 (w/w), followed by the addition of 0.1% RapiGest SF (Waters Corp., UK) and overnight digestion at 37 °C. Samples were acidified to 0.5% formic acid (FA) to quench digestion, centrifuged, and desalted on Stage Tips eluting the peptides with 80:20 ACN:H2O 0.1% FA. After removing ACN by speed-vac, the peptides were resuspended in 0.1% FA. Samples were analyzed via LC-MS using a Waters Acquity UPLC mclass system coupled to a Synapt XS HDMS operated in a positive ion and resolution mode. Peptides were first trapped on a Waters Acquity BEH C18 1.7 µm VANGUARD column and then separated on a Waters Acquity UPLC BEH C18 1.7 µm, 1.0 · 100 mm. Mobile phase A consisted of 0.1% FA (v/v) in MilliQ water, while mobile phase B consisted of 0.1% FA (v/v) in ACN. The LC gradient was supplied at 40 µl·min-1 over 22 min gradient (5–35%B). Mass spectrometry analysis was performed using MSe data independent acquisition, which used a collision energy ramp from 20 to 45 V in a mass range of 50–200 m/z for the high energy scans. Raw LC-MSe data files were processed using ProteinLynx Global Server 3.0.3 (Waters Corp., UK) and searched against human p53 fasta protein sequence (UniProt P04637). The search was performed using 10 ppm as a precursor mass tolerance and a fragment mass tolerance of 5 ppm with three minimum fragment ion matches per peptide. Trypsin was set as the protease with a maximum of two missed cleavage allowed. Cysteine N-ethylmaleimide, cystine, and Cysteine dimedone were set as variable modifications. Leu-Enkephalin (556.27 m/z) was used as a lock mass. Peak lists were exported as text files and analyzed using custom scripts in Python 3.5.
Ion Mobility-Mass Spectrometry
IM-MS experiments were carried out on a Synapt XS HDMS (Waters Corporation, Manchester, UK). The ion source was operated under gentle conditions to prevent ion activation (source temperature 30 °C, cone voltage 10 V, source offset 1). All of the experiments were carried out in sensitivity mode to maximize ion transmission at the expense of peak resolution. CIU experiments were performed by recording ion arrival time distributions under different trap collision energies in the 0–60 V range of a quadrupole-selected ions. Wave velocity and wave heights were set up at 300 ms−1 and 20 V, respectively. The helium cell and nitrogen traveling wave were operated at 200 and 75 ml min−1. The trap DC bias was set up at 35 V to minimize ion activation. The mass spectra were calibrated using 2 μg μl−1 NaI (1:1 water:isopropanol). Activation energies were reported as laboratory frame energy (Elab), accounting for the charge state of the mass-selected ion and are specific to Synapt XS HDMS and experimental conditions used66. Arrival time distributions were converted to TWCCSN2 using a TWIMS calibration procedure118. Briefly, serum albumin (Bovine), concanavalin a (jack bean) and cytochrome C (equine heart) purchased from Sigma-Aldrich were dissolved in 200 AmAc and diluted to a 10 µM protein concentration, and the ATD recorded using identical parameters. Measurements were done in triplicate in different days to account for any source of variation, and data averaged. The literature CCSN2 values for the standards were obtained from A. P. France et al.118. CCS calibration was performed using IMSCal19 (Waters Corp., UK)119. Data were analyzed by means Masslynx v4.2 (Waters Corp., UK), ORIGAMI120, CIUSuite 257, and custom Python 3.5 scripts.
DNA oligonucleotides
All the oligonucleotides for DNA binding studies were synthesized by Sigma-Aldrich. The oligonucleotides encoding for the p21 recognization element were 5′ F-GAACATGTCCCAACATGTTG-3′ and 5′ R-CAACATGTTGGGACATGTTC-3′. Olignucleotides encoding for the mdm2 recgonition element were 5′F-GGGCTGGTCAAGTTGGGACACGTCCGGCGT3′ and 5′R-ACGCCGGACGTGTCCCAACTTGACCAGCCC-3′. Annealing was performed by heating the oligonucleotides at 95 °C for 10 min, and gradually cooling to 25 °C. The concentration of double stranded DNA used for the experiments was based on that all DNA was annealed, which to our purposes is accurate enough.
Cysteine alkylation experiments
To profile reactive cysteine residues, 10 µM of either DNA-free (WTp53 and R248Qp53) or DNA-bound (WTp53) was incubated with 0.5–1 mM NEM at 25 °C. Aliquouts were withdrawn at 5, 15, 30, and 60 min and native mass spectrum or IM-MS was recorded.
Oxidation experiments
Stock solutions of H2O2 were freshly prepared in miliQ H2O A set of reactions involving different experimental approaches were performed: (i) Isolated p53 monomer 10 μM (200 mM ammonium acetate) was incubated with 2 and 20 mM H2O2 for 5 min in ice. Samples were desalted using micro Bio-Spin 6 columns (Bio-Rad), loaded to in-house prepared needles and analyze using an Orbitrap UHMR mass spectrometer with a resolution set up at 200,000 @ m/z 400; (ii) Time course oxidation experiments of the isolated p53 monomer were performed by incubating 10 μM p53 (200 mM ammonium acetate) with 2 mM H2O2. The sample was loaded into the needle and the reaction was monitored using Orbitrap UHMR mass spectrometer operating at resolution 200,000 @ m/z 400, or Synapt XS HDMS.; (iii) A control experiment that shows the CSD shift was performed by incubating 10 μM p53 with 1 mM EDTA for 30 min at 25 °C. The sample was desalted using micro Bio-Spin 6 columns (Bio-Rad) and analyzed by native MS and IM-MS using Orbitrap UHMR mass spectrometer and Synapt XS HDMS.; (iv) 10 μM p53 was incubated with 0.25 molar equivalents of p21 for 10 min. Then, 2 mM H2O2 was added and the sample loaded in the needle. The reaction was monitored online by using an Orbitrap UHMR mass spectrometer, operating at resolution 12,000 @ m/z 400.; (v) 10 μM p53 was incubated with 0.25 molar equivalents of p21 for 10 min. Then, 20 μM of Zn7MT2 was added and incubated for 10 min. After this, 2 mM H2O2 was added and the sample loaded in the needle. The reaction was monitored online by using an Orbitrap UHMR mass spectrometer, operating at resolution 12,000 @ m/z 400.
Alkylation and oxidation experiments
These experiments attempt to probe the formation of disulfides by labeling free Cys residues. In a first experiment, p53 monomer was incubated with 5 mM NEM (15 min, 25 °C, dark), the NEM excess removed by C18-Ziptip and analyzed under denaturing MS conditions using a Synapt XS HDMS. A second experiment p53 was first incubated with 2 or 5 mM H2O2 (5 min, 25 °C) and then 5 mM NEM was added and incubated for 15 min in 25 °C, dark. Sample later purified by C18-Ziptip and directly analyzed.
Sulfenic acid trapping by top down LC-MS dimedone experiments
To probe the existence of sulfenic acids as intermediates during the oxidation reaction, 30 μM of p53 was incubated with 0.5–10 mM H2O2 in the presence of 5 mM dimedone (Sigma-Aldrich). Aliquots were taken after 15, 60, 120, and 300 min, and the reaction quenched by fivefold dilution and acidification (50:50 ACN:H2O 0.1% FA). Samples were analyzed via LC-MS using a Waters Acquity UPLC mclass system coupled to a Synapt XS HDMS operated in a positive ion and resolution mode. A 5 µl of sample was injected and proteins then separated on a Waters Acquity UPLC BEH C4 1.7 µm, 2.1 · 50 mm. Mobile phase A consisted of 0.1% FA (v/v) in MilliQ water, while mobile phase B consisted of 0.1% FA (v/v) in ACN. The LC gradient was supplied at 10 µl·min-1 over 20 min gradient (15–90%B). Full MS data was acquired in a range of 500–3000 m/z. Raw LC-MS data files were processed using Masslynx v4.2 (Waters Corp., UK) and Python 3.5 scripts.
Kinetic analysis
The m/z spectrum was divided into two distinct populations: unfolded conformation ions (z > 11) and ions forming a native compact structure (z = 8–10). The choice of charge state 14 for the monomeric species was based on several considerations: first, the signals in the m/z region corresponding to 14+ were baseline resolved, allowing us to directly use the intensity for kinetic analysis. In contrast, the native charge states (8–10) exhibited an asymmetric baseline, and the peaks were not completely desolvated. Considering these factors, relying on their intensity could introduce errors. Additionally, ionization and transmission effects may vary to a further extent for different intermediates in native-like ions. To validate our approach, we performed similar kinetic analyzes with different ions from the unfolded population. We opted not to use deconvolution spectra, as they would incorporate the abovementioned source of errors. Regarding the ions corresponding to the p53-DNA complex, we chose the most intense ion 19+ since they exhibited similar transmission and desolvation effects. Thus, the intensities from the single charge state of the mass spectrum acquired in the experiment ii (Fig. 4F, G) were normalized to 0–1 on every scan and fitted with Eqs. (1–3), that correspond to two consecutive first-order reactions:
where [A]0 is the initial concentration of the reactant, k1 and k2 are the reaction constants and t is reaction time.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry data have been deposited to Figshare repository (10.6084/m9.figshare.25352884).
References
Levine, A. J., Momand, J. & Finlay, C. A. The p53 tumour suppressor gene. Nature 351, 453–456 (1991).
Joerger, A. C. & Fersht, A. R. Structural biology of the tumor suppressor p53. Annu. Rev. Biochem. 77, 557–p582 (2008).
Cho, Y., Gorina, S., Jeffrey, P. D. & Pavletich, N. P. Crystal structure of a p53 tumor suppressor-DNA complex: understanding tumorigenic mutations. Science 265, 346–355 (1994).
Blanden, A. R. et al. Zinc shapes the folding landscape of p53 and establishes a pathway for reactivating structurally diverse cancer mutants. eLife 9, e61487 (2020).
Friedler, A., Veprintsev, D. B., Hansson, L. O. & Fersht, A. R. Kinetic instability of p53 core domain mutants. J. Biol. Chem. 278, 24108–24112 (2003).
Pradhan, M. R. et al. Simulations of mutant p53 DNA binding domains reveal a novel druggable pocket. Nucleic Acids Res. 47, 1637–1652 (2019).
Butler, J. S. & Loh, S. N. Zn2+-dependent misfolding of the p53 DNA binding domain. Biochemistry 46, 2630–2639 (2007).
Han, C. W., Lee, H. N., Jeong, M. S., Park, S. Y. & Jang, S. B. Structural basis of the p53 DNA binding domain and PUMA complex. Biochem. Biophys. Res. Commun. 548, 39–46 (2021).
Jurneczko, E. et al. Probing the conformational diversity of cancer-associated mutations in p53 with ion-mobility mass spectrometry. Angew. Chem. 52, 4370–4374 (2013).
Lambrughi, M. et al. DNA-binding protects p53 from interactions with cofactors involved in transcription-independent functions. Nucleic Acids Res. 44, 9096–9109 (2016).
Baugh, E. H., Ke, H., Levine, A. J., Bonneau, R. A. & Chan, C. S. Why are there hotspot mutations in the TP53 gene in human cancers? Cell Death Differ. 25, 154–160 (2018).
Krois, A. S., Dyson, H. J. & Wright, P. E. Long-range regulations of p53 DNA binding by its intrinsically disordered N-terminal transactivation domain. Proc. Natl. Acad. Sci. USA. 115, E11302–E11310 (2018).
Veprintsev, D. B. et al. Core domain interactions in full-length p53 in solution. Proc. Natl. Acad. Sci. USA. 103, 2115–2119 (2006).
Bode, A. M. & Dong, Z. Post-translational modification of p53 in tumorigenesis. Nat. Rev. Cancer 4, 793–805 (2004).
Friedel, L. & Loewer, A. The guardian’s choice: how p53 enables context-specific decision-making in individual cells. FEBS Lett. 289, 40–52 (2021).
Levine, A. J. p53, the cellular gatekeeper for growth and division. Cell 80, 323–331 (1997).
Shi, T., van Soest, D. M. K., Polderman, P. E., Burgering, B. M. T. & Dansen, T. B. DNA damage and oxidant stress activate p53 through differential upstream signaling pathways. Free Radic. Biol. Med. 172, 298–311 (2021).
Buzek, J., Latonen, L., Kurki, S., Peltonen, K. & Laiho, M. Redox state of tumor suppressor p53 regulates its sequence-specific DNA binding in DNA-damaged cells by cysteine 277. Nucleic Acids Res. 30, 2340–2348 (2002).
Chin, P. L., Momand, J. & Pfeifer, G. P. In vivo evidence for binding of p53 to consensus binding sites in the p21 and GADD45 genes in response to ionizing radiation. Oncogene 15, 87–99 (1997).
Uberti, D. et al. Hydrogen peroxide induces nuclear translocation of p53 and apoptosis in cells of oligodendroglia origin. Mol. Brain Res. 65, 167–175 (1995).
Augustyn, K. E., Merino, E. J. & Barton, J. K. A role for DNA-mediated charge transport in regulating p53: oxidation of the DNA-bound protein from a distance. Proc. Natl. Acad. Sci. USA. 104, 18907–18912 (2007).
Schaefer, K. N. et al. Oxidation of p53 through DNA charge transport involves a network of disulfides within the DNA-binding domain. Biochemistry 54, 932–941 (2015).
Hainaut, P. & Redox, J. Milner modulation of p53 conformation and sequence-specific DNA binding in vitro. Cancer Res. 53, 4469–4473 (1993).
Rainwater, R., Parks, D., Anderson, M. E., Tegtmeyer, P. & Mann, K. Role of cysteine residues in regulation of p53 function. Mol. Cell. Biol. 15, 3892–3903 (1995).
Parks, D., Bolinger, R. & Mann, K. Redox state regulates binding of p53 to sequence-specific DNA, but not to non-specific or mismatched DNA. Nucleic Acids Res. 25, 1289–1295 (1997).
Chen, Q. M., Liu, J. & Merrett, J. B. Apoptosis or senescence-like growth arrest: influence of cell-cycle position, p53, p21 and bax in H2O2 response of normal human fibroblasts. Biochem. J. 347, 543–551 (2000).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Altelaar, A. F. M., Munoz, J. & Heck, A. J. R. Next-generation proteomics: towards an integrative view of proteome dynamics. Nat. Rev. Genet. 14, 35–48 (2013).
Held, J. M. et al. Targeted quantitation of site-specific cysteine oxidation in endogenous proteins using a differential alkylation and multiple reaction monitoring mass spectrometry approach. Mol. Cell. Proteom. 9, 1400–1410 (2010).
Kaar, J. L. et al. Stabilization of mutant p53 via alkylation of cysteines and effect on DNA binding. Protein Sci. 19, 2267–2278 (2010).
Shi, T. et al. p53 forms redox-dependent protein-protein interactions through cysteine 277. Antioxidants 10, 1578 (2021).
Smith, L. M. & Kelleher, N. L. Consortium for top down proteomics. Proteoform: a single term describing protein complexity. Nat. Methods 10, 186–187 (2013).
Tamara, S., den Boer, M. A. & Heck, A. J. R. High-resolution native mass spectrometry. Chem. Rev. 122, 7269–7326 (2022).
Heck, A. J. R. Native mass spectrometry: a bridge between interactomics and structural biology. Nat. Methods 5, 927–933 (2008).
Lermyte, F., Tsybin, Y. O., O’Connor, P. B. & Loo, J. A. Top or middle? Up or down? Toward a standard lexicon for protein top-down and allied mass spectrometry approaches. J. Am. Soc. Mass Spectrom. 30, 1149–1157 (2019).
Gault, J. et al. Combining native and ´omics´ mass spectrometry to identify endogenous ligands bound to membrane proteins. Nat. Methods 17, 505–508 (2020).
Lantz, C. et al. Native top-down mass spectrometry with collisionally activated dissociation yields higher-order structure information for protein complexes. J. Am. Chem. Soc. 144, 21826–21830 (2022).
Wu, D. & Robinson, C. V. Native top-down mass spectrometry reveals a role for interfacial glycans on therapeutic cytokine and hormone assemblies. Angew. Chem. 61, e202213170 (2022).
Kelleher, N. L. et al. Top down versus bottom up protein characterization by tandem high-resolution mass spectrometry. J. Am. Chem. Soc. 121, 806–812 (1999).
Katta, V. & Chait, B. T. Observation of the heme-globin complex in native myoglobin by electrospray-ionization mass spectrometry. J. Am. Chem. Soc. 113, 8234–8535 (1991).
Rostom, A. A. & Robinson, C. V. Detection of the intact GroEL chaperonin assembly by mass spectrometry. J. Am. Chem. Soc. 121, 4718–4719 (1999).
Tran, J. C. et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature 480, 254–258 (2011).
Vimer, S., Ben-Nissan, G. & Sharon, M. Direct characterization of overproduced proteins by native mass spectrometry. Nat. Protoc. 15, 236–265 (2020).
Rose, R. J., Damoc, E., Denisov, E., Makarov, A. & Heck, A. J. R. High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies. Nat. Methods 9, 1084–1086 (2012).
Chorev, D. S. et al. Protein assemblies ejected directly from native membranes yield complexes for mass spectrometry. Science 362, 829–834 (2018).
Scotcher, J. et al. Identification of two reactive cysteine residues in the tumor suppressor protein p53 using top-down FTICR mass spectrometry. J. Am. Soc. Mass Spectrom. 22, 888–897 (2011).
Clemmer, D. E., Hudgins, R. R. & Jarrold, M. F. Naked protein conformations: cytochrome c in the gas phase. J. Am. Chem. Soc. 117, 10141–10142 (1995).
Bargeen, S. M. et al. Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction. Nat. Commun. 13, 4377 (2022).
Michaelevski, I., Eisenstein, M. & Sharon, M. Gas-phase compaction and unfolding of protein structures. Anal. Chem. 82, 9484–9491 (2010).
Wang, S. C. et al. Ion mobility mass spectrometry of two tetrameric membrane protein complexes reveals compact structures and differences in stability and packing. J. Am. Chem. Soc. 132, 15468–15470 (2010).
Young, L. M. et al. Screening and classifying small-molecule inhibitors of amyloid formation using ion mobility spectrometry-mass spectrometry. Nat. Chem. 7, 73–81 (2015).
Wang, H. et al. Native mass spectrometry, ion mobility, electron-capture dissociation, and modeling provide structural information for gas-phase apolipoprotein e oligomers. J. Am. Soc. Mass Spectrom. 30, 876–885 (2019).
Hofmann, J., Hahm, H. S., Seeberger, P. H. & Pagel, K. Identification of carbohydrate anomers using ion mobility-mass spectrometry. Nature 526, 241–244 (2015).
Abi-Ghanem, J., Rabin, C., Porrini, M., Rosu, F. & Gabelica, V. Compaction of RNA hairpins and their kissing complexes in native electrospray mass spectrometry. J. Am. Soc. Mass Spectrom. 31, 2035–2043 (2020).
Van Dyck, J. F. et al. Sizing up DNA nanostructure assembly with native mass spectrometry and ion mobility. Nat. Commun. 13, 3610 (2022).
Gabelica, V. & Marklund, E. Fundamentals of ion mobility spectrometry. Curr. Opin. Chem. Biol. 42, 51–59 (2018).
Polasky, D. A., Dixit, S. M., Fantin, S. M., & Ruotolo, B. T. CIUSuite 2: next-generation software for the analysis of gas-phase protein unfolding data. Anal. Chem. 91, 3147–3155 (2019).
Al-jabity, A. et al. Combined chemical modification and collision induced unfolding using native ion mobility-mass spectrometry provides insights into protein gas-phase structure. Chem. Eur. J. 27, 13783 (2021).
Van Schaick, G. et al. Online collision-induced unfolding of therapeutic monoclonal antibody glyco-variants through direct hyphenation of cation Exchange chromatography with native ion mobility-mass spectrometry. Anal. Chem. 95, 3932–3939 (2023).
Deslignière, E. et al. Combination of IM-based approaches to unravel the coexistence of two conformers on a therapeutic multispecific mAb. Anal. Chem. 94, 7981–7989 (2022).
Zheng, X., Kurulugama, R. T., Laganowsky, A. & Russell, D. H. Collision-induced unfolding studies of proteins and protein complexes using drift tube ion mobility-mass spectrometer. Anal. Chem. 92, 7218–7225 (2020).
Peris-Díaz, M. D., Barkhanskiy, A., Liggett, E., Barran, P. & Krężel, A. Ion mobility mass spectrometry and molecular dynamics simulations unravel the conformational stability of zinc metallothionein-2 species. Chem. Commun. 59, 4471–4474 (2023).
Hyung, S. J., Robinson, C. V. & Ruotolo, B. T. Gas-Phase unfolding and disassembly reveals stability differences in ligand-bound multiprotein complexes. Chem. Biol. 16, 382–390 (2009).
Tian, Y. & Ruotolo, B. T. The growing role of structural mass spectrometry in the discovery and development of therapeutic antibodies. Analyst 143, 2459–2468 (2018).
Dong, S., Shirzadeh, M., Fan, L., Laganowsky, A. & Russell, D. H. Ag+ ion binding to human metallothionein-2A is cooperative and domain specific. Anal. Chem. 92, 8923–8932 (2020).
Donor, M. T., Shepherd, S. O. & Prell, J. S. Rapid determination of activation energies for gas-phase protein unfolding and dissociation in a Q-IM-ToF mass spectrometer. J. Am. Soc. Mass. Spectrom. 31, 602–610 (2020).
Arlt, C. et al. An integrated mass spectrometry based approach to probe the structure of the full-length wild-type tetrameric p53 tumor suppressor. Angew. Chem. Int. Ed. 128, 1–6 (2016).
Pagel, K., Natan, E., Hall, Z., Fersht, A. R. & Robinson, C. V. Intrinsically disordered p53 and its complexes populate compact conformations in the gas phase. Angew. Chem. Int. Ed. 52, 361–365 (2012).
Tidow, H. et al. Quaternary structures of tumor suppressor p53 and a specific p53-DNA complex. Proc. Natl. Acad. Sci. USA. 104, 12324–12329 (2001).
Beveridge, R. et al. A mass spectrometry-based framework to define the extent of disorder in proteins. Anal. Chem. 86, 10979–10991 (2014).
Beveridge, R., Migas, L. G., Kriwacki, R. W. & Barran, P. E. Ion mobility mass spectrometry measures the conformational landscape of p27 and its domains and how this is modulated upon interaction with cdk2/cyclin A. Angew. Chem. Int. Ed. 58, 3114–3118 (2019).
Eldrid, C. et al. Gas phase stability of protein ions in a cyclic ion mobility spectrometry traveling wave device. Anal. Chem. 91, 7554–7561 (2019).
Uetrecht, C. et al. Ion mobility mass spectrometry of proteins and protein assemblies. Chem. Soc. Rev. 39, 1633–1655 (2010).
Norgate, E. L. et al. Cold denaturation of proteins in the absence of solvent: implications for protein storage. Angew. Chem. 61, e202115047 (2022).
May, J. C. & Russell, D. H. A mass-selective variable-temperature drift tube ion mobility-mass spectrometer for temperature dependent ion mobility studies. J. Am. Soc. Mass Spectrom. 22, 1134–1145 (2011).
Turzo, S. B. A. et al. Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction. Nat. Commun. 13, 4377 (2022).
Faull, P. A. et al. Utilising ion mobility-mass spectrometry to interrogate macromolecules: Factor H complement control protein modulates 10-15 and 19-20 and the DNA-binding core domain of tumour suppressor p53. Int. J. Mass Spectrom. 298, 99–110 (2010).
Dixit, S. M., Polasky, D. A. & Ruotolo, B. T. Collision induced unfolding of isolated proteins in the gas phase: past, present, and future. Curr. Opin. Chem. Biol. 42, 93–100 (2018).
Taniguchi, Y. & Kawakami, M. Variation in the mechanical unfolding pathway of p53DBD induced by interaction with p53 N-terminal region or DNA. PLoS ONE 7, e49003 (2012).
Wong, K.-B. et al. Hot-spot mutants of p53 core domain evince characteristic local structural changes. Proc. Natl. Acad. Sci. USA. 96, 8438–8442 (1999).
Sparks, A. et al. The degradation of p53 and its major E3 ligase mdm2 is differentially dependent on the proteasomal ubiquitin receptor S5a. Oncogene 33, 4685–4696 (2014).
Paulsen, C. E. & Carroll, K. S. Cysteine-mediated redox signalling: chemistry, biology, and tools for discovery. Chem. Rev. 113, 4633–4679 (2013).
Loh, S. N. The missing zinc: p53 misfolding and cancer. Metallomics 2, 442–449 (2010).
Ferrer-Sueta, G. et al. Factors affecting protein thiol reactivity and specificity in peroxide reduction. Chem. Res. Toxicol. 24, 434–450 (2011).
Zhang, Q., Bykov, V. J. N., Wiman, K. G. & Zawacka-Pankau, J. APR-246 reactivates mutant p53 by targeting cysteines 124 and 277. Cell Death Dis. 9, 439 (2018).
Gough, D. R. & Cotter, T. G. Hydrogen peroxide: a Jekyll and Hyde signaling molecule. Cell Death Dis. 2, e213 (2011).
Sies, H. & Jones, D. P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell. Biol. 21, 363–383 (2020).
Scotcher, J. et al. Redox regulation of tumour suppressor protein p53: identification of the sites of hydrogen peroxide oxidation and glutathionylation. Chem. Sci. 4, 1257–1269 (2013).
Jin, S.-G., Meng, Y., Johnson, J., Szabó, P. E. & Pfeifer, G. P. Concordance of hydrogen peroxide-induced 8-oxo-guanine patterns with two cancer mutation signatures of upper GI tract tumors. Sci. Adv. 8, eabn3815 (2022).
Maret, W. & Thiolate, B. L. Vallee ligands in metallothionein confer redox activity on zinc clusters. Proc. Natl. Acad. Sci. USA. 95, 3478–3482 (1998).
Ghorbel, I. et al. Expression of metallothioneins I and II related to oxidative stress in the liver of aluminium-treated rats. Arch. Physiol. Biochem. 122, 214–222 (2016).
Zhang, B. et al. Activity of metal-responsive transcription factor 1 by toxic heavy metals and H 2 O 2 in vitro is modulated by metallothionein. Mol. Cell. Biol. 23, 8471–8485 (2003).
Rodríguez-Menéndez, S. et al. The zinc-metallothionein redox system reduces oxidative stress in retinal pigment epithelial cells. Nutrients 10, 1874 (2018).
Cai, L., Klein, J. B. & Kang, Y. J. Metallothionein inhibits peroxynitrite-induced DNA and lipoprotein damage. J. Biol. Chem. 275, 38957–38960 (2000).
Schwarz, M. A. et al. Metallothionein protects against the cytotoxic and DNA-damaging effects of nitric oxide. Proc. Natl. Acad. Sci. USA. 92, 4452–4456 (1995).
Wu, H. H., Thomas, J. A. & Momand, J. p53 protein oxidation in cultured cells in response to pyrrolidine dithiocarbamate: a novel method for relating the amount of p53 oxidation in vivo to the regulation of p53-responsive genes. Biochem. J. 351, 87–93 (2000).
Wu, H. H. & Momand, J. Pyrrolidine dithiocarbamate prevents p53 activation and promotes p53 cysteine residue oxidation. J. Biol. Chem. 273, 18898–18905 (1998).
Stone, J. R. & Yang, S. Hydrogen peroxide: a signaling messenger. Antioxid. Redox Signal. 8, 243–270 (2006).
Bilan, D. S. et al. HyPer-3: a genetically encoded H2O2 probe with improved performance for ratiometric and fluorescence lifetime imaging. ACS Chem. Biol. 8, 535–542 (2013).
Huang, B. K. & Sikes, H. D. Quantifying intracellular hydrogen peroxide perturbations in terms of concentration. Redox Biol. 2, 955–962 (2014).
Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: oxidative eustress. Redox Biol. 11, 613–619 (2017).
Lyublinskaya, O. & Antunes, F. Measuring intracelular concentration of hydrogen peroxide with the use of genetically encoded H2O2 biosensor HyPer. Redox Biol. 24, 101200 (2019).
Ezerina, D., Morgan, B. & Dick, T. P. Imaging dynamic redox processes with genetically encoded probes. J. Mol. Cell Cardiol. 73, 43–49 (2014).
Song, K.-S. et al. Detection and quantification of Tp53 and p53-anti-p53 autoantibody immune complex: promising biomarkers in early stage lung cancer diagnosis. Biosensors 12, 127 (2022).
Winter, J., Ilbert, M., Graf, P. C. F., Ozcelik, D. & Jakob, U. Bleach activates a redox-regulated chaperone by oxidative protein unfolding. Cell 135, 691–701 (2008).
Ilbert, M. et al. The redox-switch domain of Hsp33 functions as dual stress sensor. Nat. Struct. Mol. Biol. 14, 556–563 (2007).
Ilbert, M., Graf, P. C. F. & Jakob, U. Zinc center as redox switch-new function for an old motif. Antioxid. Redox Signal. 8, 835–846 (2006).
Morgan, M. J. & Liu, Z.-G. Crosstalk of reactive oxygen species and NF-kB signaling. Cell Res. 21, 103–115 (2011).
Barrett, W. C. et al. Roles of superoxide radical anion in signal transduction mediated by reversible regulation of protein-tyrosine phosphatase 1B. J. Biol. Chem. 274, 34543–34546 (1999).
Zheng, M., Aslund, F. & Storz, G. Activation of the OxyR transcription factor by reversible disulfide bond formation. Science 279, 1718–1721 (1998).
Shi, T. & Dansen, T. B. Reactive oxygen species induced p53 activation: DNA damage, redox signaling, or both? Antioxid. Redox Signal. 33, 839–859 (2020).
Krężel, A. & Maret, W. Zinc-buffering capacity of a eukaryotic cell at physiological pZn. J. Biol. Inorg. Chem. 11, 1049–1062 (2006).
Blanden, A. R. et al. Zinc shapes the folding landscape of p53 and esbalishes a pathway for reactivating structurally diverse cancer mutants. eLife 9, e61487 (2020).
Ayed, A. et al. Latent and active p53 are identical in conformation. Nat. Struc. Biol. 8, 756–760 (2001).
Fellers, R. T. et al. ProSight Lite: graphical software to analyze top-down mass spectrometry data. Proteomics 15, 1235–1238 (2014).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Bas, D. C., Rogers, D. M. & Jensen, J. H. Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins 73, 765–783 (2008).
France, A. P., Migas, L. G., Sinclair, E., Bellina, B. & Barran, P. E. Using collision cross section distributions to assess the distribution of collision cross section values. Anal. Chem. 92, 4340–4348 (2020).
Richardson, K., Langridge, D., Dixit, S. M. & Ruotolo, B. T. An improved calibration approach for traveling wave ion mobility spectromeytry: robust, high-precision collision cross sections. Anal. Chem. 93, 3542–3550 (2021).
Migas, L. G., France, A. P., Bellina, B. & Barran, P. E. ORIGAMI: a software suite for activated ion mobility mass spectrometry (aIM-MS) applied to multimeric protein assemblies. Int. J. Mass Spectrom. 427, 20–28 (2018).
Acknowledgements
This research was supported by the National Science Centre of Poland (NCN) under the Opus grant no. 2019/33/B/ST4/02428 (to A.K.), Preludium no. 2018/31/N/ST4/01909 and Etiuda no. 2020/36/T/ST4/00404 (to. M.D.P.D). A.K and M.D.P.D would like to thank Michał Tracz for performing the bottom-up MS experiments. We acknowledge the support of EPSRC through the strategic equipment award EP/T019328/1, and Waters Corporation for their continued support of mass spectrometry research within the Michael Barber Centre for Collaborative Mass Spectrometry.
Author information
Authors and Affiliations
Contributions
M.D.P.D., A.K., and P.B. conceived the study. M.D.P.D. performed the experimental sections, data analysis and interpretation. M.D.P.D., A.K., and P.B. drafted the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peris-Díaz, M.D., Krężel, A. & Barran, P. Deciphering the safeguarding role of cysteine residues in p53 against H2O2-induced oxidation using high-resolution native mass spectrometry. Commun Chem 8, 13 (2025). https://doi.org/10.1038/s42004-024-01395-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-024-01395-w