Abstract
Drug repositioning holds promise for discovering new therapeutic applications for existing drugs, accelerating drug development and reducing associated costs. However, current methodologies encounter difficulties in managing diverse network representations, tackling cold start issues, and handling intrinsic attribute representations. Here we introduce a Unified Knowledge-Enhanced deep learning framework for Drug Repositioning (UKEDR), which integrates knowledge graph embedding, pre-training strategies, and recommendation systems to address these challenges. To overcome the cold start issue, UKEDR utilizes a semantic similarity-driven embedding approach. Our evaluations show that UKEDR performs better than various baselines, including classical machine learning, network-based, and deep learning approaches. In cold start scenarios, it demonstrates an improved capability in handling unseen nodes and generalizing to new compounds. The model also demonstrates strong robustness on imbalanced datasets and shows excellent generalization capabilities in specific drug-centric and disease-centric cold-start scenarios, validating its potential for real-world applications.
Similar content being viewed by others
Introduction
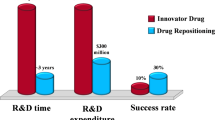
The traditional drug development process is notorious for its prolonged duration and significant financial costs, encompassing pivotal phases from initial target discovery to conclusive clinical trials. This complex journey typically takes 10–15 years and accrues expenses from $500 million–$2 billion1. Despite extensive efforts, the success rate remains notably low, with over 90% of projects ending in failure2. In contrast, drug repositioning offers a more efficient alternative. This approach identifies novel therapeutic uses for existing drugs, leveraging established, safe, and effective medications, reducing risks and costs. Additionally, drug repositioning benefits from an accelerated development timeline by utilizing existing clinical data. However, despite interest from academia and industry, successful instances of drug repositioning are still relatively rare, often occurring serendipitously.
To address this challenge, a range of methodologies has been developed, which can be classified into three categories: machine learning (ML), network-based, and deep learning (DL) methods. Classical ML models approach drug repositioning as a binary classification problem, employing standard classification models or matrix-based models for prediction, such as SVM3, logistic regression4, random forest5, WKNKN6, KMDR7, DRRS8, BNNR9, and DisDrugPred10. In contrast, network-based methods have been introduced to enhance drug repositioning predictions, overcoming the limitations of classical ML models. Hu et al.11 constructed a disease-drug network by meticulously analyzing the genomic expression profiles of diseases and drugs, utilizing this network to predict drug-disease interactions. Cheng et al.12 developed three supervised inference methods, DBSI, TBSI, and NBI, based on complex network theory to predict drug-target interactions for drug repositioning. Luo et al.13 proposed the MBiRW method, which creates a heterogeneous network using drug and disease similarity networks, as well as known drug-disease interactions, and then uses the BiRW algorithm to make predictions. DL methods leverage intricate interaction networks among drugs, targets, and diseases, representing these associations as nodes and edges within a network. By leveraging techniques such as network analysis and graph neural networks, these methods model and analyze these interaction networks, enabling the prediction of drug-disease interactions. A noteworthy example is DeepDR14, a network-based DL method tailored for drug repositioning, which integrates multiple drug-related networks to improve prediction accuracy. However, extracting valuable information from diverse medication repositioning data sources remains challenging. To address this, methods like Relational Graph Convolutional Networks (RGCN)15, Heterogeneous Graph Transformer16, and Heterogeneous Attention Networks (HAN)17 have been introduced. These approaches aim to model the complex relationships between drugs, targets, and diseases within heterogeneous networks, thereby integrating diverse information to enhance the accuracy of drug repositioning predictions. Recently, researchers have explored the potential of Knowledge Graphs (KGs) in addressing the drug repositioning problem. One approach is GrEDeL, which utilizes a biomedical KG embedding model based on PubMed abstracts and employs the translation-based mode TransE18 to embed entities. DREAMwalk utilizes a “semantic multi-layer guilt-by-association” approach and introduces a teleport operation to address the imbalance in node quantities in biomedical knowledge graphs, enhancing drug repositioning19. KGCNH is a knowledge graph convolutional network for drug repositioning that combines semantic and topological information. It uses a relation-aware attention mechanism and a Gumbel-SoftMax heuristic search to improve drug-disease association predictions20. EKGDR utilizes graph neural networks for end-to-end embedding of the drug knowledge graph, enabling it to learn the underlying intents of drug-disease interactions and aggregate relational messages across multihop paths21. In parallel, the synergy between KGs and Recommender Systems (RS) has become a vibrant research area, as KGs provide rich side information to alleviate data sparsity and improve recommendation interpretability. These hybrid approaches range from using KG embeddings as input features to more advanced GNN-based models like KGAT22, which jointly learn embeddings and user preferences.
Machine learning methods struggle to capture the biological mechanisms underlying drug action. Network-based approaches have introduced relationships such as drug-protein, drug-side effect, and drug-GO, which partially address this issue. However, the integration of multiple networks makes it challenging to effectively combine these representations. Knowledge graph (KG) approaches, particularly those leveraging powerful Graph Neural Networks, have emerged as a state-of-the-art paradigm. For instance, models like LAGCN23 and DRHGCN24 effectively fuse heterogeneous graph information using sophisticated GCN architectures with inter- and intra-domain modules. Other advanced methods, such as FuHLDR25 fuse higher-order meta-path information with lower-order relations, while models like HeTDR26 integrate network topology with features from text mining.
However, a common principle unites these powerful graph-centric models: their predictive capabilities are fundamentally rooted in the entities and structures present within the graph. This leads to a critical limitation, often acknowledged within the literature itself. This dependency on pre-existing graph structures is an acknowledged limitation of many GNN-based approaches. For instance, it has been noted that models like DRHGCN cannot be applied to novel diseases for which association data is unavailable, as their feature generation is contingent on such information24. This defines a critical challenge: the prediction for novel entities that are entirely absent from the knowledge graph, a scenario often termed the ‘out-of-graph’ cold start. Furthermore, these models primarily derive relational representations from the graph’s topology, often overlooking the rich intrinsic attribute representations of the entities themselves. These combined limitations—a dependency on pre-existing graph structures and an absence of attribute-level understanding—significantly hinder the real-world applicability of purely graph-based approaches.
To address these two issues, we propose the UKEDR model, which introduces pre-training to tackle the lack of attribute representations. For drugs, we utilize molecular SMILES and carbon spectral data for contrastive learning, while for diseases, we fine-tune a large language model using textual descriptions. These pre-trained models allow us to obtain attribute representations for any new molecule or disease. To address the cold start problem, we search for nodes similar to those unseen in the current pre-trained space and map them into the knowledge graph embedding space, using these nodes to derive the relational representations of unseen nodes. Traditional dot products are insufficient for predicting drug repositioning effectively; therefore, we incorporate an attention-based recommendation algorithm to better integrate relational and attribute representations for drug repositioning.
In this work, we demonstrate that our proposed framework, UKEDR, consistently outperforms various baseline methods across three standard benchmark datasets. The model exhibits exceptional generalization capabilities, particularly in challenging cold-start scenarios. In a realistic simulation of predicting clinical trial outcomes from approved drug data, UKEDR improves the area under the receiver operating characteristic curve (AUC) by 39.3% over the next-best model. Furthermore, our framework shows strong robustness on highly imbalanced data and, critically, can make accurate predictions for drugs and diseases that are entirely new to the knowledge graph. Our analyses reveal that this superior performance is driven by the synergistic combination of graph-based relational embeddings and rich, pre-trained intrinsic features.
Results
Systematic module configuration exploration of UKEDR
Traditional drug repositioning methods often rely on simple dot products or single-layer neural networks for feature interactions, which may not fully capture the complex relationships between drugs and diseases. While recent studies have attempted to integrate knowledge graph embedding with recommendation systems, they typically employ random or experience-based combinations, lacking a systematic evaluation of different module combinations.
To determine the optimal module configuration for UKEDR, we conducted a systematic exploration of various knowledge graph embedding methods and recommendation system (RS) approaches, as illustrated in Fig. 1 and Supplementary Fig. 2. The introduction of recommendation systems aims to better model the interaction between relationship features and attribute features, moving beyond the limitations of traditional approaches. Interestingly, our analysis revealed that despite the integration of knowledge graph and pre-training representations, the choice of knowledge graph embedding methods had minimal impact on the model’s ultimate performance (Fig. 1c, d, Supplementary Fig. 3). Instead, the selection of downstream recommendation algorithms emerged as the critical factor influencing overall effectiveness (Fig. 1e, f, Supplementary Fig. 4). This pattern is consistent across both AUC and AUPR metrics.
a, b Distribution of AUC and AUPR values across different KGE-RS models sorted by decreasing mean performance, with global averages indicated by dashed lines. c, d Interaction effects between KGE methods and RS algorithms show the dominant influence of RS choice on both AUC and AUPR performance. e, f Alternative perspective demonstrating the consistent performance patterns of RS methods across various KGE techniques. g, h Model stability comparison using coefficients of variation for AUC and AUPR, with lower values indicating more stable performance across experimental runs.
Our analysis demonstrates that AFM recommendation systems consistently outperformed other algorithms across all knowledge graph embedding methods, with AUC values above 0.95 and AUPR values above 0.96 (Fig. 1a, b). These top-performing configurations also exhibited superior stability, as evidenced by their low coefficients of variation (Fig. 1g, h). Using PairRE as a representative case, the PairRE_AFM configuration demonstrated superior performance (AUC = 0.95, AUPR = 0.96), showing statistically significant improvements (p < 0.05) over alternatives such as PairRE_BPR and PairRE_NFM. Based on these findings, we selected PairRE as our knowledge graph embedding model due to its exceptional scalability and implemented attentional factorization machines (AFM) as the recommender system for drug repositioning prediction. The superior performance of AFM can be attributed to its ability to capture complex feature interactions through attention mechanisms, providing more sophisticated modeling of drug-disease associations compared to traditional methods.
To ensure robust and comprehensive feature representation, we developed a systematic feature extraction pipeline that integrates complementary deep neural architectures. For disease representation, we introduced DisBERT, a domain-specific language model obtained by fine-tuning BioBERT on an extensive corpus of over 400,000 disease-related text descriptions. This specialized training process enables DisBERT to capture subtle nuances and complex semantic patterns specific to disease manifestations, significantly enhancing the quality of disease feature representation compared to generic biomedical language models. Complementing the disease representation framework, we leveraged the CReSS model27 for drug feature extraction, establishing a balanced dual-stream architecture for feature learning. This carefully engineered feature extraction framework provides high-quality drug and disease representations for the subsequent PairRE and AFM modules, where each component contributes its specialized capabilities to the overall system. The systematic integration of these components enables our model to effectively capture both the broad semantic context of diseases and the fine-grained molecular characteristics of drugs, providing a solid foundation for accurate drug repositioning predictions.
UKEDR achieves superior performance across diverse benchmarks and challenging scenarios
In this study, we evaluated the performance of various models across a comprehensive set of scenarios designed to test both standard performance and robustness under challenging conditions. For the standard evaluation, we benchmarked the models using three datasets: RepoAPP, RepoClin, and RepoData. To maintain a balanced dataset, we randomly sampled negative samples to achieve a 1:1 positive-to-negative sample ratio. The evaluation metrics, including AUC and Area Under the Precision-Recall Curve (AUPR) values, were derived from five-fold cross-validation for each model. This validation process was replicated five times to ensure the reliability of the results (Fig. 2 and Supplementary Table 5).
a, b represent the AUC and AUPR on the RepoData, respectively; c, d represent the AUC and AUPR on the RepoAPP, respectively; e, f represent the AUC and AUPR on the RepoClin, respectively; g, h represent the AUC and AUPR in the scenario of the drug cold start, respectively; i, j represent the AUC and AUPR in the scenario of the disease cold start, respectively; k, l represent the AUC and AUPR in the scenario of the general cold start, respectively.
In the standard benchmark scenario, UKEDR achieved the best performance across all three datasets, with EKGDR and DeepDR ranking closely behind. Specifically, in RepoAPP, UKEDR delivered the best results (AUC = 0.958, AUPR = 0.968), followed by EKGDR (AUC = 0.937, AUPR = 0.945). Notably, the impact of each model on RepoAPP was less prominent compared to RepoClin and RepoData. The violin plots revealed that the UKEDR model exhibited a more stable and uniform distribution of results, whereas the HAN, KEGDR, DREAMwalk, and KNN models showed a more scattered pattern, indicating higher variance in their predictions.
To simulate a realistic drug discovery process, we evaluated the models in a general cold start scenario, where we trained our model on RepoAPP (approved drugs) and subsequently evaluated its performance on RepoClin (drugs in clinical trials). This approach uses data from approved drugs to train the model and predict the results of ongoing clinical trials, offering several advantages such as realistic prediction scenarios, reduced bias, and improved generalizability. In this scenario, UKEDR demonstrated superior performance with the highest scores (AUC = 0.950 ± 0.007, AUPR = 0.961 ± 0.005), significantly outperforming all other models. The next-best performer was EKGDR (AUC = 0.682 ± 0.011, AUPR = 0.746 ± 0.007), followed by DeepDR (AUC = 0.621 ± 0.025, AUPR = 0.687 ± 0.017). Notably, UKEDR’s performance exhibited an improvement of 39.3% in AUC and 28.8% in AUPR compared to the second-best model, EKGDR. This substantial performance gap underscores UKEDR’s exceptional capability in handling unseen nodes and generalizing to new compounds.
To further probe the model’s robustness, we conducted evaluations under even more stringent conditions. First, we assessed performance on the RepoAPP110 dataset, which features a 1:10 positive-to-negative sample ratio to mimic real-world class imbalance. In this setting, UKEDR achieved an AUC of 0.963 and an AUPR of 0.839. This AUPR score is particularly significant, as it substantially outperforms other leading methods, highlighting UKEDR’s ability to effectively identify true positive associations in a highly imbalanced environment where the AUPR metric is more informative. Second, we dissected the cold-start problem into two specific scenarios. In the drug cold-start scenario, where the model must predict interactions for entirely new drugs, UKEDR achieved an outstanding AUC of 0.795 ± 0.009 and AUPR of 0.812 ± 0.010, dramatically surpassing all baseline models. This underscores the critical role of our pre-trained drug feature module. Similarly, in the disease cold-start scenario, UKEDR also led the performance with an AUC of 0.736 ± 0.013 and AUPR of 0.739 ± 0.013, validating the effectiveness of our pre-trained disease features in capturing the semantic essence of unseen diseases.
Robustness evaluation of UKEDR with the additional dataset
To systematically evaluate the robustness of UKEDR, we conducted experiments on Cdataset, a widely adopted benchmark dataset in drug repositioning research initially presented by Luo et al.13. This comprehensive dataset encompasses 2532 validated drug-disease associations, comprising interactions between 663 drugs and 409 diseases. We implemented a rigorous five-fold cross-validation protocol while maintaining a balanced 1:1 positive-to-negative sample ratio. This experimental design ensures a thorough performance assessment under realistic conditions. The experimental results demonstrated the model’s consistent and superior performance across multiple evaluation metrics (Fig. 3, Supplementary Table 6).
a, b Represent the AUC and AUPR on the additional dataset, respectively.
The cross-validation results revealed that UKEDR achieved robust performance metrics, with AUC values ranging from 0.869 to 0.907, and AUPR scores varying between 0.830 and 0.930. These results were benchmarked against several state-of-the-art models to assess comparative performance. UKEDR exhibited AUC scores comparable to established models such as EKGDR (0.883–0.912) and DeepDR (0.865–0.896), while demonstrating particularly notable performance in AUPR metrics.
Significantly, UKEDR’s AUPR performance (0.830–0.930) showed marginal superiority over EKGDR (0.866–0.901), a distinction particularly relevant given AUPR’s enhanced sensitivity in evaluating imbalanced dataset performance. While other contemporary models, including HGT and KGCNH, demonstrated promising results, their overall performance metrics did not exceed those achieved by UKEDR.
The robust performance of UKEDR is evidenced by its consistent achievement across all cross-validation folds, maintaining competitive AUC values while exhibiting superior AUPR metrics. This consistency, coupled with the model’s successful performance on the benchmark dataset, substantiates UKEDR’s capability to effectively process diverse data distributions while maintaining high predictive accuracy across various evaluation criteria. These findings collectively validate the robustness and adaptability of the UKEDR framework in handling complex drug-response prediction tasks.
The combination of knowledge graph embedding and pre-training boost the performance of UKEDR
The effective integration of different feature types is crucial for enhancing model performance in drug repositioning tasks. However, before combining knowledge graph embedding and pre-trained features, it is essential to understand their intrinsic characteristics and potential complementarity. This understanding would provide theoretical support for their integration and guide the design of more effective models.
To this end, we conducted a detailed analysis of similarity matrices for drugs and diseases across both feature modalities (Supplementary Fig. 5 and Fig. 4a, b). Our analysis revealed distinct and potentially complementary patterns between these feature types. Knowledge graph features generated similarity matrices with notably lower values, showing mean similarities of 0.162 (std: 0.051) for diseases and 0.183 (std: 0.050) for drugs. The majority of KGE similarities clustered within narrow ranges, indicating their specialization in capturing precise relational information between entities. This concentrated distribution pattern aligns with the fundamental nature of knowledge graphs, which excel at representing explicit relationships and structural information.
a Distribution of normalized disease similarity matrices derived from KGE and Pretrain methods. b Distribution of normalized drug similarity matrices from KGE and Pretrain methods. c ROC curves from ablation studies comparing the performance of different model variants. d Precision-recall curves showing the predictive power of different model configurations in ablation experiments.
In contrast, pre-trained features exhibited significantly higher similarity values with broader distributions. Disease pre-trained features showed a mean similarity of 0.606 (std: 0.135), while drug pre-trained features demonstrated a mean of 0.480 (std: 0.159). This higher-value, broader distribution suggests their capability to capture intrinsic molecular and semantic properties that might not be explicitly encoded in knowledge graphs. The substantial difference in distribution patterns between KGE and pre-trained features strongly supports our hypothesis that the two feature types capture fundamentally different aspects of drug-disease relationships. This is further confirmed by the near-zero correlation coefficients between the two modalities (Pearson correlation of −0.0064 for diseases and −0.0092 for drugs, both with p values ≈ 0), demonstrating they provide statistically independent information. This statistical independence makes their integration particularly valuable for drug repositioning tasks, as each modality contributes unique and complementary information to the model. To rigorously test this hypothesis and quantify the potential benefits of feature integration, we designed a comprehensive ablation study comparing various model configurations. We systematically evaluated five variants: KGE (utilizing only Knowledge Graph Embedding), PRE (incorporating only Drug and Disease Pre-training), KGE_Cress (combining KGE with Drug Pre-training), KGE_Disbert (integrating KGE with Disease Pre-training), and the complete UKEDR model integrating all components. All variants employed AFM as the downstream classifier to ensure fair comparison (Fig. 4c, d). The results provided strong empirical support for our hypothesis, with the complete UKEDR model achieving superior performance (AUC = 0.953, AUPR = 0.964) compared to all partial implementations. Particularly revealing was the performance progression observed through the variants: the intermediate performance levels of KGE_Cress and KGE_Disbert, while superior to single-feature variants, still fell short of the complete model’s capabilities. This stepwise improvement pattern demonstrates that each feature type contributes unique and valuable information to the model’s understanding of drug-disease relationships.
These findings not only validate our initial hypothesis about feature complementarity but also provide important insights into the mechanism of successful drug repositioning models. The observed synergy between knowledge graph embedding and pre-trained features suggests that effective drug repositioning requires both the relational context captured by knowledge graphs and the intrinsic properties encoded in pre-trained representations. This understanding could guide future developments in drug repositioning methodologies and similar biomedical applications where multiple types of features are available.
UKEDR suggests drug/disease biological mechanisms and potential novel drugs for various diseases
To elucidate the performance characteristics of knowledge graph embeddings and pretrained features in drug recommendation systems, we conducted a systematic multidimensional analysis of the prediction results from KGE, pretrained, and their combined models. Probability density distributions (Fig. 5a) and distributional statistical features (Fig. 5b) revealed significant differences among the three models: the KGE model exhibited a near-symmetric distribution centered at 0.316 (mean 0.340); the pretrained model demonstrated a highly right-skewed distribution with the majority of predictions clustered in lower score segments (mean 0.241, median merely 0.008); while the combined model presented intermediate distributional characteristics (mean 0.110, median 0.032). Percentile analysis (Fig. 5c) further quantified this phenomenon, showing that pretrained-related models maintained low prediction scores up to the 80th percentile, followed by a steep exponential increase. Inter-model correlation analysis (Fig. 5d–f) indicated that KGE and the combined model exhibited significant correlation (r = 0.625), substantially exceeding the correlation degrees of KGE-pretrained (r = 0.362) and pretrained-combined models (r = 0.372)—a finding suggesting that KGE features may play a more dominant role in the fusion architecture. Overlap analysis of high-confidence predictions (Fig. 5g) further confirmed that KGE and the combined model maintained higher consistency (51.4%) in top 10% predictions, significantly surpassing the consistency between KGE-pretrained (35.8%) and pretrained-combined models (34.7%). High-frequency drug recommendation analysis (Fig. 5h, i) provided a practical perspective, showing that the combined model could identify the most diverse high-frequency drug candidates (229 types), followed by the KGE model (218 types), with the pretrained model identifying the fewest (155 types); high-frequency drugs commonly recommended by all three models constituted only 77 types, with each model maintaining a significant proportion of unique recommendations. These findings hold important implications, suggesting that feature fusion strategies not only integrate the predictive advantages of both foundational models but also effectively mitigate the “head effect” commonly observed in recommendation systems, significantly enhancing recommendation diversity—a characteristic of particular value for discovering potential novel therapeutic approaches in the field of drug repositioning.
a Probability density distribution of prediction scores showing distinct distributional characteristics. b Violin plots quantifying prediction score statistics with median and mean values. c Percentile analysis of score distribution reveals model-specific ranking patterns. d–f Correlation scatter plots demonstrating pairwise relationships between KGE-Pretrained, KGE Combined, and Pretrained-Combined models with Pearson correlation coefficients. g Overlap percentage analysis of top 10% predictions between model pairs. h Comparison of high-frequency drug recommendations (frequency ≥10) across models. i Venn diagram showing overlap of high-frequency drug recommendations between the three prediction models.
The average degree of drugs in the prediction results derived from knowledge graph embedded features is observed to decrease compared to that of pre-trained features (Supplementary Table 7). This indicates that knowledge graph embedded features assist the model in identifying items with a lower degree, while pre-trained features aid in recognizing items with a higher degree. This difference arises from the fact that the knowledge graph enriches the model with semantic information, enhancing its understanding of node relationships. On the other hand, the pre-trained model assigns higher weights to frequently occurring words during training, making it more likely to recognize items with higher degrees that are more prevalent in text data. Furthermore, when combining the two features, the standard deviation in the Knowledge Graph+Pre-train results is reduced compared to using pre-training alone. This suggests that the integration of the two features moderates or smooths the data’s variability, leading to more stable results. Additionally, the range of degrees in the combined results falls between the ranges observed in the Knowledge Graph and Pre-training results alone, suggesting that the combined use of Knowledge Graph and pre-trained features can cover a wider coverage of degree distributions, likely due to their complementary strengths in capturing item relevance. In conclusion, these analyses suggest that knowledge graphs and pre-trained features provide complementary information, enhancing the model’s comprehensive understanding of associative relationships between items.
We respectively identified the top 20 relationship pairs in the prediction rankings based on KGE features and KGE combined with pretraining features. Subsequently, we extracted a relevant subset of the knowledge graph corresponding to the drugs and diseases involved in these relationship pairs. This subset is designated as KGE Ripple1 (KGE features) and KGE_Pretrain Ripple1 (KGE with pretraining features) (Supplementary Figs. 6. and 7). While both KGE Ripple1 and KGE_Pretrain Ripple1 demonstrate dispersed structures with relatively sparse connectivity, KGE_Pretrain Ripple1 exhibits a 17.5% higher connectivity compared to KGE Ripple1 (with average degrees of 2.97 and 2.52, respectively). Additionally, its betweenness centrality decreases by 9.0% (0.000422 vs. 0.000384). Pretraining enhances the model’s ability to identify diverse pathways between drugs and diseases while decreasing node centrality, resulting in more distributed information flow across the network rather than concentration through key nodes. These changes suggest that pretrained features enhance knowledge graph structure and drug-disease relationship representations, thereby facilitating the dispersed propagation of information within the graph. These structural alterations may contribute to improving the performance of prediction models as they provide richer contextual information and a more balanced flow of information.
Moreover, we constructed UKEDR based on the benchmark to predict new drug-disease therapeutic associations. We excluded the positive samples that appeared in both the training and test sets. The top 15 predicted association scores are presented in Table 1. The prediction results indicate that DB00678 (Losartan) has the potential to treat C0020443 (Hypercholesterolemia). This finding is supported by the study reported by Tershakovec et al.28. To further clarify the predicted results of our model, we selected four representative cases and visualized some of their associated nodes in the knowledge graph (Fig. 6). Glutamine (DB00130) modulates several signaling pathways, including SMP0000365, SMP0000536, and SMP0000072, thereby influencing critical molecular targets such as 3’,5’-cyclic-AMP phosphodiesterase 4D (PDE4D, Q08499), N-acetyl-D-glucosamine kinase (NAGK, Q9UJ70), and MYC proto-oncogene (MYC, P01106). This intricate network of interactions indicates a potential role for L-Glutamine in the therapeutic management of prostate cancer (C0376358) (refer to Fig. 6a). Similarly, Fluticasone propionate (DB00588) affects multiple pathways and proteins, including Cytochrome P450 3A4 (CYP3A4, P08684), ATP-dependent translocase ABCB1 (ABCB1, P08183), and Cytochrome P450 2C8 (CYP2C8, P10632). These targets are shared with other compounds such as DB01182, DB00358, DB00207, DB00468, and others. Emerging evidence suggests that these drugs have a significant association with falciparum malaria (C0024535), proposing a potential therapeutic application of DB00588 in its treatment (see Fig. 6b). Furthermore, targets P40124, P14632, P13797, Q3SYV4, and P13020 are correlated with disease C0029456, as well as with diseases C0860207, C4279912, C3658290, C1262760, and C0019193. The known efficacy of DB00709 in treating these conditions indicates it may exert its therapeutic effects through these targets, indicating its potential use for C0029456 (Fig. 6c). Additionally, DB00678 interacts with several proteins, including P22309, P06684, P30556, P08183, P02768, and O95342. These proteins are associated with the disease C0020443 and the pathway map04976. The data indicate potential drug synergy an enhanced metabolism of Abemaciclib when co-administered with Cerivastatin (DB00439) (see Fig. 6d).
a–d Network topology highlighting four representative drug-disease predictions. Nodes represent different entities: drugs (blue diamonds), protein targets (purple circles), biological pathways (red hexagons), and diseases (green squares). Edge colors indicate binding affinity quartiles: green (Q1, <25%), yellow (Q2, 25–50%), red (Q3, 50–75%), and orange (Q4, >75%). Node size is proportional to degree centrality. The network topology reveals the complex interplay between molecular targets and biological pathways in drug-disease associations, supporting the predicted therapeutic relationships.
The knowledge graph visualization reveals the intricate molecular mechanisms underlying drug-disease associations through a comprehensive network of protein interactions and signaling pathways. This interpretability bridges computational predictions with biological mechanisms, providing valuable insights for experimental validation and enhancing the precision of drug repositioning predictions.
Computational efficiency analysis
To assess the practical feasibility of UKEDR, we evaluated its computational efficiency on our largest benchmark dataset, RepoData. All evaluations were conducted on a workstation equipped with a single NVIDIA GeForce RTX 4090 GPU with 24GB of VRAM. The training process of UKEDR mainly involves knowledge graph embedding (KGE) and training the final recommendation model. The most computationally intensive stage, KGE training on our full knowledge graph, required approximately 1.23 hours. In contrast, the training for the final recommendation system (AFM) was significantly faster, completing in under 10 minutes. For inference, generating association scores for the entire test set took approximately 6 minutes, which is highly efficient for large-scale prediction tasks.
While classical models like Random Forest (RF) are computationally faster, they lack the capacity to model complex relationships and handle cold-start scenarios effectively, as demonstrated by the significant performance gap in our results. The complexity of UKEDR is admittedly higher than some deep learning models like DeepDR, primarily due to the integration of large pre-trained models and a massive knowledge graph. However, this architectural complexity is a necessary and justified trade-off for achieving state-of-the-art performance, especially in the critical cold-start scenarios where other models are limited.
Discussion
The UKEDR model represents a significant advancement in drug repositioning, not merely by improving metrics on standard benchmarks, but by fundamentally addressing the true cold-start problem for out-of-graph entities—a critical limitation for many state-of-the-art GNN-based methods. This unique capability is rooted in its innovative dual-feature architecture, which fuses graph-based relational embeddings with powerful, pre-trained intrinsic attributes derived directly from molecular and textual data. The power of this approach was most rigorously validated in our most stringent cold-start scenarios. The model’s outstanding performance when predicting entirely new drugs (AUC = 0.795, AUPR = 0.812) and new diseases (AUC = 0.7364, AUPR = 0.7393)—scenarios where many graph-centric models would falter—provides direct evidence of its superior generalization ability for entities independent of their presence in the knowledge graph.
Furthermore, UKEDR’s robustness is substantiated across a wider range of challenging conditions. In the realistic simulation of training on approved drugs and testing on clinical trial drugs, the model demonstrated remarkable performance improvements of 39.3% in AUC and 28.8% in AUPR compared to the next-best performer, EKGDR. Moreover, its exceptional performance on the highly imbalanced RepoAPP110 dataset (AUPR = 0.839) showcases its ability to maintain high precision when negative samples are abundant. Collectively, these comprehensive evaluations confirm that UKEDR is not only a solution for the critical cold-start challenge but also a robust and highly practical framework for real-world drug discovery.
The dual-feature architecture of UKEDR addresses the fundamental limitation of incomplete attribute representation in traditional methods. By combining knowledge graph embeddings with pre-trained features, our model captures both explicit relational information and implicit semantic properties. The knowledge graph component encodes structured relationships between entities, while the pre-training component extracts rich molecular and disease characteristics from textual descriptions. Our analysis reveals that these complementary information sources significantly enhance the model’s predictive accuracy and robustness. The observed synergy between these features suggests that comprehensive drug repositioning requires both structural understanding of biological networks and detailed characterization of molecular properties. Furthermore, UKEDR provides interpretable predictions through its knowledge graph structure, enabling researchers to trace the reasoning behind each prediction. This interpretability is crucial for clinical applications, as it allows validation of predicted drug-disease associations through established biological mechanisms.
Despite these advances, several important challenges remain in the field of computational drug repositioning. The construction and maintenance of comprehensive knowledge graphs continue to be labor-intensive processes, potentially introducing inconsistencies and incomplete representations of biological relationships. The optimal integration of heterogeneous data sources remains challenging, particularly in capturing complex, non-linear interactions between different feature types. Additionally, the computational resources required for large-scale knowledge graphs and real-time updates present practical challenges for widespread implementation. These limitations highlight the need for continued methodological development in automated knowledge graph construction, adaptive feature integration, and computational efficiency.
Looking forward, several promising research directions emerge from our findings. The development of automated methods for knowledge graph construction and validation could significantly enhance the scalability and accuracy of drug repositioning models. Integration of additional data modalities, including clinical records, genomic data, and pathway information, could provide richer contextual information for predictions. Advanced feature learning approaches, particularly in self-supervised learning and causal reasoning, offer potential pathways for improving prediction accuracy and biological relevance. Furthermore, the application of uncertainty quantification methods and privacy-preserving techniques could facilitate the clinical implementation of these models.
Methods
Benchmark and baseline
To establish the authentic relationships between drugs and diseases in the real world, we adopted the methodology proposed by Brown and Patel, as outlined in RepoDB29. This methodological framework was selected for its robust approach to relationship mapping and its demonstrated effectiveness in pharmacological research.
In the data acquisition phase, we systematically curated information from two authoritative sources. The primary source was DrugCentral, a comprehensive database containing regulatory-approved and commercially available pharmaceutical compounds. From this repository, we meticulously extracted drug-disease pairs, which carry high credibility due to their validation through rigorous clinical trials and market authorization processes. The second major data source was the AACT (Aggregate Analysis of ClinicalTrials.gov) database, a structured repository derived from ClinicalTrials.gov. The AACT database provides extensive clinical trial records, encompassing detailed information about trial designs, participant demographics, intervention protocols, and outcomes, enabling us to capture the full spectrum of drug-disease relationships under clinical investigation.
Our data processing methodology followed a stringent protocol: From DrugCentral, we specifically focused on marketed drugs with well-defined therapeutic indications, which were systematically organized into the RepoAPP dataset. Concurrently, we extracted drug-disease relationship pairs from the AACT database, carefully documenting relationships at various clinical trial phases, which formed the RepoClin dataset. These complementary datasets represent both established therapeutic relationships and potential therapeutic applications under investigation.
To develop a more comprehensive knowledge base, we implemented a systematic update of the original RepoDB. This update process involved the integration of both RepoAPP and RepoClin datasets, employing rigorous data cleaning and standardization protocols. We applied sophisticated normalization techniques to ensure consistency across different data sources and implemented quality control measures to maintain data integrity. The resulting RepoData (as detailed in Table 2) represents a significant enhancement over previous versions, incorporating not only an expanded set of drug-disease relationship pairs but also including confidence metrics for each relationship, which is particularly valuable for drug repositioning studies.
During the assessment of our model, we divided the datasets into three distinct subsets: training, validation, and testing. Specifically, 80% of the known drug-disease pairs were allocated to the training set, while 20% were evenly distributed between the validation and test sets. The dataset maintains a balanced positive-to-negative sample ratio of 1:1. To ensure a robust evaluation of our model’s performance, we utilized a five-fold cross-validation and repeated the process five times. This approach aimed to minimize the influence of random partitioning on the model’s performance, resulting in a more accurate and comprehensive assessment. We compared our method with several existing techniques, including RF30, LR4, KNN31, DeepDR14, HAN17, HGT16, RGCN15, DREAMwalk19, KGCNH20, and EKGDR21. (Supplementary Table 1).
Robustness and specific cold-start scenario analysis
To further assess the robustness and real-world applicability of our proposed model, we designed additional experiments to address two critical aspects raised during the review process: performance on imbalanced datasets and a more granular analysis of cold-start scenarios. These evaluations aim to simulate more realistic conditions and provide deeper insights into the model’s generalization capabilities.
First, to address the concern that a balanced 1:1 positive-to-negative sample ratio may not reflect real-world data distributions, we constructed an imbalanced dataset, designated as RepoAPP110. This dataset was derived from the original RepoAPP by sampling negative instances to create a 1:10 positive-to-negative ratio. Evaluating our model on this skewed dataset allows for a more stringent test of its ability to identify true drug-disease associations amidst a large number of unknown or negative pairs, a common challenge in drug repositioning. For this experiment, we performed five-fold cross-validation.
Second, to conduct a more fine-grained investigation of the model’s performance in cold-start situations, we established two distinct scenarios beyond the general setup of training on RepoAPP and testing on RepoClin. The first, termed the drug cold-start scenario, was designed to assess the model’s capacity to make predictions for novel compounds. In this setup, the test set exclusively contains associations involving drugs that were not present in the training set, thereby measuring generalization based on intrinsic features learned via pre-training. The second, the disease cold-start scenario, conversely, evaluates the model’s performance on associations involving diseases that were absent from the training set. This setup specifically tests the model’s ability to generalize to new therapeutic areas by leveraging the semantic features of diseases. For both specific cold-start scenarios, a five-fold cross-validation process was implemented and repeated five times to ensure the stability and reliability of the results.
Knowledge graph construction
To build a comprehensive knowledge graph, we collected data from numerous well-established public datasets, including DrugBank32, DisGeNET33, DrugCentral34, CTD35, UMLS36, UniProt37, PDBbind38, PubChem39, etc. Each of these databases offers a unique perspective, thereby enhancing the graph’s comprehensiveness and breadth of relationships (Supplementary Table 2). We implemented a systematic data processing pipeline to ensure data quality and consistency. The pipeline begins with raw data extraction, followed by a multi-step cleaning process: (1) standardizing entity identifiers using established systems (InChIKey for compounds and drugs, UMLS for diseases, and UniProt for proteins); (2) removing duplicate entries through exact matching of standardized identifiers; and (3) filtering out entities and relationships lacking proper identifier mappings or sufficient supporting evidence. For relationship curation, we validated interaction types across different data sources and merged equivalent relationships while preserving source information to maintain data provenance.
The resulting knowledge graph encompasses 2,326,366 entities, includes nine entity types: targets, chemicals, genes, diseases, proteins, GO terms, pathways, variants, and drugs. Notably, we identified 157,338,703 relationships among these entities (Supplementary Table 3, Supplementary Table 4 and Supplementary Fig. 1). To our knowledge, this represents the largest knowledge graph currently utilized in drug repositioning.
Module configuration selection
To ascertain the most effective knowledge graph embedding and recommendation methods for drug repositioning, we conducted an exhaustive comparison of well-established methods in both categories. This involved eight knowledge graph embedding models, namely AutoSF40, DistMult41, TransE18, ComplEx42, InterHT43, TripleRE44, PairRE45, and RotatE46 (Supplementary1. KGE method), along with six recommendation algorithms: BPR47, ConvNCF48 DMF49, FM50, NFM51, and AFM52 (Supplementary2. Recommendation method). To comprehensively evaluate their performance, we combined each embedding method with each recommendation algorithm, resulting in 48 unique models. For evaluation, we employed BioKG53, an existing knowledge graph encompassing a diverse array of entity types (i.e., diseases, proteins, drugs, side effects, and protein functions) and relevant relations (i.e., drug-drug interactions, protein-protein interactions, drug-protein interactions, drug-side effects, and function-function relations). BioKG was chosen due to its manageable size, consisting of 1.7 million entities and 2.3 million relations, which allows for efficient training times when compared to our larger constructed knowledge graph. This facilitated a thorough evaluation of each model combination while minimizing computational overhead. To determine significant differences among the models, we conducted a 5-fold cross-validation using the RepoAPP dataset, repeated five times. The obtained results were analyzed using ANOVA to determine the presence of statistically significant differences among the models.
Pipeline of the UKEDR
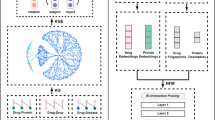
UKEDR incorporates an innovative multi-component architectural design that integrates advanced technologies, including knowledge graph embeddings, contrastive learning, and pre-trained feature extraction (Fig. 7). The technical pipeline encompasses four critical components: a cold-start processing and prediction system, knowledge graph feature extraction, drug pre-trained feature learning, and disease pre-trained feature extraction.
a Cold start process and prediction via a recommendation system; b drug and disease feature extraction based KGE model; c drug pre-train feature-based contrastive learning; and d disease pre-train featur- based DisBERT.
In the cold-start processing and prediction component (Fig. 7a), the model employs recommendation system strategies to address prediction challenges for novel drugs or diseases. This component initially determines whether nodes exist within the knowledge graph; for absent nodes, it implements cold-start processing to acquire both knowledge graph and pre-trained representations. Through the integration of attention mechanisms and graph neural networks, the system effectively handles prediction tasks for new entities lacking historical interaction data.
The knowledge graph feature extraction module (Fig. 7b) utilizes efficient graph embedding techniques to map drug and disease entities into a low-dimensional vector space. This module effectively captures structured information from the knowledge graph by learning vector representations of entities and their relationships. These structured feature representations provide an essential knowledge foundation that effectively supports subsequent drug repositioning prediction tasks.
The drug pre-trained feature learning module (Fig. 7c) introduces an innovative contrastive learning framework. This module leverages molecular structure information and carbon spectrum data to obtain latent drug representations through contrastive learning approaches. The contrastive learning strategy ensures proximity relationships between similar drugs in the feature space while enhancing the model’s comprehension of molecular characteristics.
The disease pre-trained feature extraction module (Fig. 7d) is derived from fine-tuning the BioBERT model using extensive textual data, specifically designed to process disease-related textual descriptions and clinical information. Through pre-training tasks, this module learns disease semantic features, effectively capturing complex information including disease phenotypes and pathological mechanisms.
These components collaborate in a staged manner to construct a comprehensive drug repositioning prediction framework. During the feature extraction phase, the model obtains multimodal representations from knowledge graphs, molecular structures, and disease information, subsequently integrating these heterogeneous features through a designed fusion strategy for use as input in the prediction phase of drug-disease association prediction. This multi-source feature-based architecture not only effectively addresses the cold-start problems faced by traditional methods but also provides more comprehensive and accurate predictions through the integrated utilization of diverse information sources.
Disease pre-train feature extraction using BioBERT
BioBERT54, a pre-trained deep learning language model designed specifically for biomedical text-mining tasks, builds upon the foundation of BERT and was trained on biomedical texts sourced from PubMed and PMC. This model aims to overcome the limitations inherent in generic language models by demonstrating, through fine-tuning for various tasks, such as named entity recognition, relation extraction, and text classification. In our investigation, we initialized the model with pre-training parameters from BioBERT and fine-tuned it using disease text descriptions sourced from MseqDR55 and UMLS36 to elicit effective representations of disease features.
Using WordPiece tokenization and relation extraction techniques, we obtained subword representations for all diseases. Given the constraints on training costs, we fine-tuned the representations of diseases and genes as anonymous entities, employing a batch size of 32 and a learning rate of 2 × e−5. Following the refinement process leading to the creation of DisBERT, we stabilized its parameters and leveraged disease descriptions as input to generate disease-specific features. As a result, each disease is uniquely represented by a 768-dimensional feature vector.
Drug pre-train feature extraction using contrastive learning
To extract comprehensive drug features for our repositioning task, we leveraged the CReSS model27, a powerful contrastive learning framework that effectively integrates molecular structure information with spectroscopic data. This model was selected for its demonstrated ability to overcome the limitations of traditional methods based on predefined drug similarity and molecular descriptors. The framework’s sophisticated approach to molecular representation learning, which combines 13 C nuclear magnetic resonance (NMR) spectral data with SMILES string information, aligns well with our need for robust drug feature extraction.
The CReSS framework implements a symmetric contrastive learning process through several key steps. Initially, both SMILES and NMR data are encoded and normalized:
where \({f}_{s}\) and \({f}_{n}\) are the SMILES and NMR encoders, respectively. The similarity scores are then computed as:
where τ is the temperature parameter controlling the separation margin between positive and negative pairs.
The symmetric contrastive loss is formulated as:
In our implementation, we utilized a hybrid architecture consisting of a CNN-MLP network for NMR encoding:
where \({h}_{l}\) represents the features at layer l, and \({b}_{l}\) is the bias term. The SMILES processing employs a Transformer-based network with initial embedding:
where \({W}_{e}\) is the token embedding matrix, PE adds positional encoding, and X represents the input SMILES sequence. The final SMILES embedding is obtained through:
where \({e}_{s}\in {{\mathbb{R}}}^{768}\) is the final feature vector.
This framework provides several key advantages for drug repositioning. The normalization step ensures stable similarity computations, while the temperature-scaled logits help control the learning dynamics. The symmetric loss combines both SMILES-to-NMR and NMR-to-SMILES matching, ensuring bidirectional consistency in the learned representations. By processing SMILES sequences through the Transformer architecture and incorporating NMR spectral data, we obtained rich molecular representations that effectively capture both local chemical environments and global structural properties.
For our drug repositioning study, the dual-modal approach effectively captures both structural and experimental properties of drug molecules. The SMILES sequences are transformed into 300 × 768-dimensional embedding matrices, which are then processed to generate the final 768-dimensional feature vectors. These comprehensive representations, learned through the symmetric contrastive framework, served as robust inputs for our subsequent drug repositioning analysis, enabling more accurate predictions of drug-disease associations.
KGE feature extract
PairRE45 uses paired vectors to improve the representation of relations in knowledge graph embedding. PairRE allows for adaptive adjustment of the margin in the loss function, enabling it to effectively handle intricate relationships like many-to-many or one-to-many. Additionally, it encodes diverse relation patterns, including symmetry/antisymmetry, inverse relationships, and composition. PairRE is effective in capturing the complex relationships in drug repositioning.
In our approach, we utilize paired vectors to represent each relation and use an embedding learn to derive a low-dimensional representation of these vectors. Specifically, for a given triple \((h,r,t)\), we represent \(h\) and \(t\) as vectors \({{{\bf{h}}}}\) and \({{{\bf{t}}}}\), and represent the relation \(r\) as a pair of vectors \({{{{\bf{r}}}}}^{H}\) and \({{{{\bf{r}}}}}^{T}\). We then use the distance function \({f}_{r}({{{\bf{h}}}},{{{\bf{t}}}})\) to compute the score of the triple \((h,r,t)\), which is used to define the loss function. The distance function \({f}_{r}({{{\bf{h}}}},{{{\bf{t}}}})\) is defined as follows (Eq. 10):
where \(\circ\) denotes the Hadamard product (i.e., element-wise product) between two vectors.
The PairRE loss function consists of two parts. The first part calculates the score of the known positive sample \((h,r,t)\) and maps it to the range\([0,1]\) using the sigmoid function\(\sigma\). Specifically, the loss function \({{{{\rm{L}}}}}_{1}\) for positive samples is defined as (see Eq. 11):
Here, \(\gamma\) is a margin value that controls the distance between positive and negative samples.
The second part calculates the score of negative samples \(({h ^{\prime}},r,{t ^{\prime}})\) and maps it to the range \([0,1]\) using the sigmoid function. The loss function \({L}_{2}\) for negative samples is defined as (see Eq. 12):
Here, \({h}_{i}^{\prime}\) and \({t}_{i}^{\prime}\) are one of the head and tail entities from the positive sample \({{{{\rm{p}}}}}({{{{\rm{h}}}}}_{i}^{\prime}{,{{{\rm{r}}}},{{{\rm{t}}}}}_{i}^{\prime})\) is the sampling probability for negative samples, and \({{{\rm{n}}}}\) is the number of negative samples.
The total loss function \(L\) is the weighted sum of the loss functions for positive and negative samples (see Eq. 13):
Here,\(\alpha\) is a hyperparameter that controls the weight of the negative sample loss function. Finally, we use optimization algorithms such as stochastic gradient descent to minimize \(L\), update model parameters, and learn the embeddings of the knowledge graph.
Prediction via a recommendation system
While traditional drug repositioning models often rely on simple interaction functions like the dot product, this approach has a significant limitation: it treats all feature interactions as equally important. This assumption is often too simplistic to capture the complex and nuanced relationships between drugs and diseases. The need to move beyond the dot product towards more expressive interaction models is a well-established concept in modern recommendation systems56. To address this, our study implemented the AFM model to enhance prediction accuracy52. AFM’s core advantage lies in its attention mechanism, which learns to assign different weights to different feature interactions, thereby allowing the model to focus on the most predictive signals and creating a more powerful and interpretable interaction framework. The AFM employs low-order feature vector decomposition to reduce computational complexity compared to MLP methods. To generate the drug feature vector, we merged the drug features derived from the knowledge graph with pre-trained drug features. Similarly, we combined the disease features derived from the knowledge graph with representations obtained from DisBERT to produce the disease representation vector. The mathematical expressions for the concatenated drug and disease feature vectors are outlined in Eqs. 14 and 15:
where \({{{\bf{d}}}}{{{{\bf{r}}}}}_{k}\) represents the drug feature derived from knowledge graph embedding, \({{{\bf{d}}}}{{{{\bf{r}}}}}_{p}\) as the pre-trained drug feature, \({{{\bf{d}}}}{{{{\bf{s}}}}}_{k}\) represents the disease feature from knowledge graph embedding, and \({{{\bf{d}}}}{{{{\bf{s}}}}}_{p}\) represents the disease feature obtained from DisBERT, and \({{{\bf{d}}}}{{{\bf{s}}}}\) and \({{{\bf{d}}}}{{{\bf{r}}}}\) represent the representations of disease and drug, respectively. The symbol \({||}\) represents the vector concatenation operation.
The sample feature vector \({{{\bf{x}}}}\) can be represented as (see Eq. 16):
where \(m\) and \(n\) are the numbers of feature dimensions in \({{{\bf{d}}}}{{{\bf{s}}}}\) and \({{{\bf{d}}}}{{{\bf{r}}}}\), respectively. \({x}_{dsj}\) and \({x}_{dri}\) represent the \(j-th\) and \(\,i-th\) feature dimensions of the disease feature vector and drug feature vector, respectively. As the feature dimensionality of \({{{\bf{d}}}}{{{\bf{s}}}}\) and \({{{\bf{d}}}}{{{\bf{r}}}}\) may differ, there is no need to specify which dimension to concatenate along during the vector concatenation operation
We utilized an attention-based pooling layer to compress feature interactions in the embedding space, taking into account their relative importance. This approach generates a k-dimensional output vector, which is used for prediction through a linear combination of learned weights. The AFM model can be formulated as follows (see Eq. 17):
where \(n\) is the number of features, \({w}_{0}\) and \({w}_{i}\) are bias terms and weights respectively for linear regression, \({{{\bf{x}}}}\) is the input feature vector, \({{{\bf{p}}}}\) is the weight vector for the attention-based pooling layer, \({v}_{i}\) is the latent factor vector for feature \(i\), \(\odot\) denotes element-wise multiplication, and\({a}_{ij}\) is the attention weight, used to calculate the interaction weight between feature \(i\) and feature\(j\). It is obtained by normalizing the intermediate weight \({a}_{ij}^{{\prime} }\) using the SoftMax function over all feature pairs in the input sample. The calculation formula is (see Eq. 18):
where \({ {\mathcal R} }_{x}\) is the set of all feature pairs in the input sample, \({{{\bf{h}}}}\) is the weight vector, \({{{\bf{W}}}}\) is the weight matrix, \({{{\bf{b}}}}\) is the bias vector used in the attention-based pooling layer. The rectified linear unit (ReLU) activation function is utilized in neural networks to introduce nonlinearity.
L2 regularization is applied to the weight matrix W to prevent overfitting in the attention network component. The optimized objective function combines mean squared error and L2 regularization. Stochastic gradient descent is employed to optimize this objective function. The objective function is as follows (see Eq. 19):
where λ controls the regularization strength.
Handling cold start issue
The cold start problem is a significant challenge in drug repositioning, where the model needs to make predictions for new entities (drugs or diseases) that do not appear in the training knowledge graph. Traditional knowledge graph embedding methods struggle with this scenario as they can only learn representations for entities present in the graph during training. This limitation severely restricts their practical application in drug repositioning, where the ability to handle new entities is crucial.
To address this challenge, we propose a novel semantic similarity-driven embedding approach that leverages both pre-trained embeddings and knowledge graph structure. Our method consists of two key components: (1) a similarity-guided representation strategy for initializing unseen nodes, and (2) a feature concatenation mechanism that combines both structural and semantic information.
For the initialization of unseen nodes, we develop a representation learning strategy based on semantic similarity distribution. When encountering a new entity not present in the knowledge graph, we first project it into the pre-trained semantic space. We then construct its representation by leveraging the collective knowledge from semantically similar entities that exist in both spaces. Specifically, the initial representation of the unseen node is computed as the centroid of the most similar existing entities, as shown in Eq. 20:
where\({h}_{unseen}\) is the initial representation of the unseen node, \({{{{\mathcal{N}}}}}_{k}\) represents the set of k most semantically similar entities, and \({h}_{i}\) denotes the knowledge graph embedding of entity i.
This similarity-guided initialization strategy effectively bridges the semantic space and graph space, ensuring robust representation learning for unseen entities. By leveraging semantic similarities from the pre-trained space and maintaining consistency with existing knowledge graph embeddings, our approach captures essential structural patterns while enabling seamless integration of new entities into the established knowledge graph framework.
The final representation of an entity is obtained by concatenating its initialized representation with its pre-trained features, as defined in Eq. 21:
where \({h}_{final}\) is the final representation, \({h}_{pretrained}\) is the pre-trained feature vector, and \(\oplus\) denotes the concatenation operation. This concatenation mechanism preserves both the structural information from the knowledge graph and the semantic information from pre-trained features.
Through this semantic similarity-driven approach, we effectively address the cold start problem by enabling our model to generate meaningful representations for previously unseen entities. When a new drug or disease is encountered, our method can immediately construct its representation and integrate it into the existing knowledge graph structure, allowing for immediate prediction capabilities without the need for retraining the entire model.
Data availability
The datasets supporting this study are publicly available from the GitHub repository at https://github.com/LiKun8732/UKEDR and the Zenodo archive at https://zenodo.org/records/16735421.
Code availability
The source code supporting this study are publicly available from the GitHub repository at https://github.com/LiKun8732/UKEDR and the Zenodo archive at https://zenodo.org/records/16735421.
References
Gao, Z., Ding, P. & Xu, R. KG-Predict: a knowledge graph computational framework for drug repurposing. J. Biomed. Inform. 132, 104133 (2022).
Zeng, X., Tu, X., Liu, Y., Fu, X. & Su, Y. Toward better drug discovery with knowledge graph. Curr. Opin. Struct. Biol. 72, 114–126 (2022).
Wang, Y., Chen, S., Deng, N. & Wang, Y. Drug repositioning by kernel-based integration of molecular structure, molecular activity, and phenotype data. PLoS One 8, e78518 (2013).
Gottlieb, A., Stein, G. Y., Ruppin, E. & Sharan, R. PREDICT: a method for inferring novel drug indications with application to personalized medicine. Mol. Syst. Biol. 7, 496 (2011).
Cao, D. S. et al. Computational prediction of drug target interactions using chemical, biological, and network features. Mol. Inform. 33, 669–681 (2014).
Ezzat, A., Zhao, P., Wu, M., Li, X. L. & Kwoh, C. K. Drug-target interaction prediction with graph regularized matrix factorization. IEEE/ACM Trans. Comput. Biol. Bioinform. 14, 646–656 (2017).
Kuang, Q. et al. A kernel matrix dimension reduction method for predicting drug-target interaction. Chemom. Intell. Lab. Syst. 162, 104–110 (2017).
Luo, H. et al. Computational drug repositioning using low-rank matrix approximation and randomized algorithms. Bioinformatics 34, 1904–1912 (2018).
Yang, M., Luo, H., Li, Y. & Wang, J. Drug repositioning based on bounded nuclear norm regularization. Bioinformatics 35, i455–i463 (2019).
Xuan, P. et al. Drug repositioning through integration of prior knowledge and projections of drugs and diseases. Bioinformatics 35, 4108–4119 (2019).
Hu, G. & Agarwal, P. Human disease-drug network based on genomic expression profiles. PLoS One 4, e6536 (2009).
Cheng, F. et al. Prediction of drug-target interactions and drug repositioning via network-based inference. PLoS Comput. Biol. 8, e1002503 (2012).
Luo, H. et al. Drug repositioning based on comprehensive similarity measures and bi-random walk algorithm. Bioinformatics 32, 2664–2671 (2016).
Zeng, X. et al. deepDR: a network-based deep learning approach to in silico drug repositioning. Bioinformatics 35, 5191–5198 (2019).
Schlichtkrull, M. et al. Modeling relational data with graph convolutional networks. Semantic Web (eswc 2018) (eds Gangemi, A. et al.) vol. 10843, 593–607 (2018).
Hu, Z., Dong, Y., Wang, K. & Sun, Y. Heterogeneous graph transformer. Web Conference 2020: Proceedings of the World Wide Web Conference (www 2020) 2704–2710 (2020).
Wang, X. et al. Heterogeneous graph attention network. Web Conference 2019: Proceedings of the World Wide Web Conference (www 2019) 2022–2032 (2019).
Bordes, A., Usunier, N., Garcia-Duran, A., Weston, J. & Yakhnenko, O. Translating embeddings for modeling multi-relational data. In: Advances in Neural Information Processing Systems. 26. Curran Associates, Inc.; 2013. Accessed March, https://papers.nips.cc/paper/2013/hash/1cecc7a77928ca8133fa24680a88d2f9-Abstract.html (2023).
Bang, D., Lim, S., Lee, S. & Kim, S. Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers. Nat. Commun. 14, 3570 (2023).
Du, X., Sun, X. & Li, M. Knowledge Graph Convolutional Network with Heuristic Search for Drug Repositioning. J. Chem. Inf. Model. 64, 4928–4937 (2024).
Tayebi, J. & BabaAli, B. EKGDR: An End-to-End Knowledge Graph-Based Method for Computational Drug Repurposing. J. Chem. Inf. Model 64, 1868–1881 (2024).
Wang, X., He, X., Cao, Y., Liu, M. & Chua, T. S KGAT: Knowledge Graph Attention Network for Recommendation. In: Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining. 950–958. https://doi.org/10.1145/3292500.3330989 (2019).
Yu, Z., Huang, F., Zhao, X., Xiao, W. & Zhang, W. Predicting drug–disease associations through layer attention graph convolutional network. Brief. Bioinform 22, bbaa243 (2021).
Cai, L. et al. Drug repositioning based on the heterogeneous information fusion graph convolutional network. Brief. Bioinform 22, bbab319 (2021).
Zhao, B. W. et al. Fusing Higher and Lower-Order Biological Information for Drug Repositioning via Graph Representation Learning. IEEE Trans. Emerg. Top. Comput. 12, 163–176 (2024).
Jin, S. et al. HeTDR: Drug repositioning based on heterogeneous networks and text mining. Patterns 2, 100307 (2021).
Yang, Z. et al. Cross-Modal Retrieval between 13C NMR Spectra and Structures for Compound Identification Using Deep Contrastive Learning. Anal. Chem. 93, 16947–16955 (2021).
Tershakovec, A. M. et al. Effect of LDL cholesterol and treatment with losartan on end-stage renal disease in the RENAAL study. Diab. Care 31, 445–447 (2008).
Brown, A. S. & Patel, C. J. A standard database for drug repositioning. Sci. Data 4, 170029 (2017).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Altman, N. S. An introduction to Kernel and nearest-neighbor nonparametric regression. Am. Stat. 46, 175–185 (1992).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Piñero, J. et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 48, D845–D855 (2020).
Avram, S. et al. DrugCentral 2023 extends human clinical data and integrates veterinary drugs. Nucleic Acids Res. 51, D1276–D1287 (2023).
Davis, A. P. et al. Comparative Toxicogenomics Database (CTD): update 2021. Nucleic Acids Res. 49, D1138–D1143 (2021).
Bodenreider, O. The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32, D267–D270 (2004).
The UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531 (2023).
Wang, R., Fang, X., Lu, Y., Yang, C. Y. & Wang, S. The PDBbind database: methodologies and updates. J. Med. Chem. 48, 4111–4119 (2005).
Kim, S. et al. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res. 49, D1388–D1395 (2021).
Zhang, Y., Yao, Q., Dai, W. & Chen, L. AutoSF: searching scoring functions for knowledge graph embedding. 2020 IEEE 36th International Conference on Data Engineering (icde 2020) 433–444 (2020).
Yang, B., Yih, W. T., He, X., Gao, J. & Deng, L. Embedding entities and relations for learning and inference in knowledge bases. arXiv https://doi.org/10.48550/arXiv.1412.6575 (2015).
Trouillon, T., Welbl, J., Riedel, S., Gaussier, E. & Bouchard, G. Complex Embeddings for Simple Link Prediction. International Conference on Machine Learning, vol. 48 (2016).
Wang, B. et al. InterHT: knowledge graph embeddings by interaction between head and tail entities. arXiv https://doi.org/10.48550/arXiv.2202.04897 (2022).
Yu, L. et al. Triplere: knowledge graph embeddings via triple relation vectors. Available at: https://vixra.org/abs/2112.0095 (2023).
Chao, L., He, J., Wang, T. & Chu, W. PairRE: knowledge graph embeddings via paired relation vectors. 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing (acl-Ijcnlp 2021), Vol. 1, 4360–4369 (2021).
Sun, Z., Deng, Z. H., Nie, J. Y. & Tang, J. RotatE: knowledge graph embedding by relational rotation in complex space. arXiv https://doi.org/10.48550/arXiv.1902.10197 (2019).
Rendle, S., Freudenthaler, C., Gantner, Z. & Schmidt-Thieme, L. BPR: Bayesian personalized ranking from implicit feedback. In:Proceedings of the Twenty-Fifth Conference on Uncertainty in Artificial Intelligence. UAI ’09. AUAI Press; 452–461 (2009).
He, X. et al. Outer product-based neural collaborative filtering. Published online 2227–2233 (2018).
Xue, H. J., Dai, X., Zhang, J., Huang, S. & Chen, J. Deep matrix factorization models for recommender systems. Published online 3203–3209 (2017).
Rendle, S. et al. Factorization machines. 2010 IEEE International Conference on Data Mining, 995–1000 (2010).
He, X. & Chua, T. S. Neural factorization machines for sparse predictive analytics. In: Proceedings of the 40th International ACM SIGIR Conference on Research and Development in Information Retrieval. SIGIR ’17. Association for Computing Machinery; 355–364 (2017).
Xiao, J. et al. Attentional factorization machines: learning the weight of feature interactions via attention networks. Proceedings of the Twenty-Sixth International Joint Conference on Artificial Intelligence, 3119–3125 (2017).
Walsh, B., Mohamed, S. K. & Nováček, V. BioKG: a knowledge graph for relational learning on biological data. In: Proceedings of the 29th ACM International Conference on Information & Knowledge Management. CIKM ’20. Association for Computing Machinery; 3173–3180 (2020).
Lee, J. et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2020).
Falk, M. J. et al. Mitochondrial Disease Sequence Data Resource (MSeqDR): a global grass-roots consortium to facilitate deposition, curation, annotation, and integrated analysis of genomic data for the mitochondrial disease clinical and research communities. Mol. Genet. Metab. 114, 388–396 (2015).
He, X. et al. Neural collaborative filtering. Proceedings of the 26th International Conference on World Wide Web, 173–182 (2017).
Wang, J. H., Devalia, J. L., Sapsford, R. J. & Davies, R. J. Effect of corticosteroids on release of RANTES and sICAM-1 from cultured human bronchial epithelial cells, induced by TNF-alpha. Eur. Respir. J. 10, 834–840 (1997).
Lister, A. et al. Nrf2 is overexpressed in pancreatic cancer: implications for cell proliferation and therapy. Mol. Cancer 10, 37 (2011).
MacManus, C. F. et al. Interleukin-8 signaling promotes translational regulation of cyclin D in androgen-independent prostate cancer cells. Mol. Cancer Res. 5, 737–748 (2007).
Rutschmann, S. et al. PanR1, a dominant negative missense allele of the gene encoding TNF-α (Tnf), does not impair lymphoid development1. J. Immunol. 176, 7525–7532 (2006).
Schuldt, K. et al. A -436c>a polymorphism in the human FAS gene promoter associated with severe childhood malaria. PLoS Genet. 7, e1002066 (2011).
Jiang, X. et al. Protective effect of FGF21 on type 1 diabetes-induced testicular apoptotic cell death probably via both mitochondrial- and endoplasmic reticulum stress-dependent pathways in the mouse model. Toxicol. Lett. 219, 65–76 (2013).
Toshino, A. et al. Expression of keratinocyte transglutaminase in cornea of vitamin A-deficient rats. Curr. Eye Res. 30, 731–739 (2005).
Preview : Journal Type Zhang, G. et al. Oxymatrine hydrazone (OMTH) synthesis and its protective effect for rheumatoid arthritis through downregulation of MEK/NF-κB pathway. Environ. Toxicol. 36, 2448–2453 (2021).
Suarez-Martinez, E., Husain, K. & Ferder, L. Adiponectin expression and the cardioprotective role of the vitamin D receptor activator paricalcitol and the angiotensin converting enzyme inhibitor enalapril in ApoE-deficient mice. Ther. Adv. Cardiovasc. Dis. 8, 224–236 (2014).
Zhu, X., Li, J., Liu, F. & Uetrecht, J. P. Involvement of T helper 17 cells in D-penicillamine–induced autoimmune disease in brown Norway rats. Toxicol. Sci. 120, 331–338 (2011).
Coe, B. P. et al. Neurodevelopmental disease genes implicated by de novo mutation and copy number variation morbidity. Nat. Genet. 51, 106–116 (2019).
Armenia, J. et al. The long tail of oncogenic drivers in prostate cancer. Nat. Genet. 50, 645–651 (2018).
Wang, M. et al. Large-scale association analysis in Asians identifies new susceptibility loci for prostate cancer. Nat. Commun. 6, 8469 (2015).
Spanagel, R. The clock gene Per2 influences the glutamatergic system and modulates alcohol consumption. Nat. Med. 11, 35–42 (2005).
Acknowledgements
This work was supported by the National Science and Technology Major Project of the Ministry of Science and Technology of China [2023ZD0507104]; National Natural Science Foundation of China [22173118, 22220102001, 22307112 and 82304316]; Hunan Provincial Science Fund for Distinguished Young Scholars [2021JJ10068]; the Science and Technology Innovation Program of Hunan Province [2021RC4011]; The Natural Science Foundation of Hunan Province [2022JJ80104]; and The 2020 Guangdong Provincial Science and Technology Innovation Strategy Special Fund [2020B1212030006, Guangdong-Hong Kong-Macau Joint Lab]. Funding for open access charge: HKBU Strategic Development Fund project [SDF19-0402-P02]. We acknowledge Haikun Xu and the High-Performance Computing Center of Central South University for support. The study was approved by the university’s review board. This work was supported in part by the Center for Computational Biology and Bioinformatics, Furong Laboratory, and Bioinformatics Center, Xiangya Hospital, Central South University.
Author information
Authors and Affiliations
Contributions
T.H. and D.C. contributed to the paper revision and experimental design. K.L. developed the method. K.L., X.Y. and L.Y. wrote the code. K.L. performed the analysis. K.L., L.F., J.Y., Q.Y. and C.W. wrote the paper. Y.D., D.J. and C.W. co-supervised the project and helped with troubleshooting. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, K., Yi, J., Ye, Q. et al. A fused deep learning approach to transform drug repositioning. Commun Chem 8, 334 (2025). https://doi.org/10.1038/s42004-025-01712-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01712-x