Abstract
Antibiotic resistance is rapidly emerging as one of the most critical health threats, with resistant microorganisms progressively diminishing the effectiveness of established antibiotics. As a result, the development of therapeutic approaches that effectively target resistant pathogens is of utmost importance. In this study, we developed inhibitors for APH(2”)-IVa, a bacterial kinase conveying resistance to aminoglycoside antibiotics. Starting from a hit of a fragment-based screening, we explored the inhibitory motif by structure-based design, ultimately leading to a series of triazole analogues. Advanced analogues displayed promising ADME properties, emerging selectivity vs a panel of human kinases, permeability in both Gram-positive and Gram-negative bacteria, and a moderate antibiotic efficacy for clinical strains of P. aeruginosa. Taken together, our results suggest inhibition of bacterial kinases could be a promising option to reinstall the efficacy of aminoglycoside antibiotics.
Similar content being viewed by others
Introduction
Antibiotic resistance is one of the most pressing public health challenges nowadays. Aminoglycosides are extensively used in the clinic, due to their effectiveness against both Gram-positive and Gram-negative species. However, emerging resistance toward aminoglycosides increasingly jeopardizes the efficacy of this class of antibiotics and demands for additional therapeutic modalities1. Given the essential role of aminoglycosides in treating infections caused by pathogens listed as critical and high priority by the World Health Organization (WHO)2, strategies to safeguard the clinical efficacy of aminoglycosides are urgently needed to address the growing threat of multidrug-resistant (MDR) bacteria. Resistance to aminoglycosides can occur through multiple mechanisms, among which deactivation by bacterial enzymes is one of the most prominent3. Aminoglycoside-modifying enzymes encompass three main classes: i) acetyltransferases (AACs), ii) nucleotidyltransferases (ANTs), and iii) phosphotransferases (APHs). APHs inactivate aminoglycosides by catalyzing the transfer of a phosphoryl group from a phosphoryl donor, ATP or GTP, to a hydroxyl group present on aminoglycosides4,5. Targeting APHs holds potential as a target to tackle resistance since (i) APHs are often implicated in MDR phenotypes, especially in high-risk pathogens; (ii) APH phosphorylation often impacts a broad range of aminoglycosides, therefore APHs are considered clinically more relevant6. In light of these considerations, developing small molecule inhibitors of APHs could pave the way to counteracting aminoglycoside resistance7.
In the past, significant efforts have been dedicated to discovering compounds targeting APHs. Key strategies have included repurposing existing eukaryotic protein kinase inhibitors, targeting novel allosteric pockets, and designing novel antibiotics or de novo APH inhibitors.
The structural similarity between APHs and eukaryotic protein kinases (ePKs) has inspired the repurposing of ePK inhibitors to address APH-mediated resistance. Shakya et al. characterized the inhibition of different APHs by diverse protein kinase inhibitors (PKI), revealing differences in binding modes compared to ePKs and identifying compounds that restore aminoglycoside efficacy against resistant bacteria8. Several natural products show activity against APHs. Flavonoids like quercetin, a broad-spectrum kinase inhibitor, were found to bind to several APH isoforms7.
In a study by Daigle et al., the interactions of kinase inhibitors from three distinct structural classes—indole carbazoles, flavonoids, and isoquinoline sulfonamides—were evaluated against several APH enzymes. Notably, members of the isoquinoline sulfonamide class demonstrated significant inhibitory activity against APH(3’)-IIIa and the bifunctional AAC(6’)-APH(2’) enzyme, both found in S. aureus and Enterococci. However, despite their in vitro potency, these compounds failed to restore aminoglycoside efficacy in bacterial culture assays. Nevertheless, the findings provided an important proof-of-concept for targeting APHs with kinase-directed scaffolds9.
In a separate investigation, the casein kinase 1 inhibitor CKI-7 exhibited an inhibition constant (Kᵢ) of approximately 65 µM against APH(3’)-IIIa10, further supporting the viability of this strategy.
Building upon this concept, subsequent studies explored additional ePK inhibitors with distinct scaffolds and mechanisms of kinase selectivity. For instance, the anthrapyrazolone SP600125, a canonical JNK inhibitor, was found to inhibit APH(3′)-Ia from E. coli in vitro and exhibited modest synergy when combined with kanamycin. However, this effect was insufficient to fully restore antibacterial activity. Still, its performance as a chemical probe reinforced the druggability of the APH ATP-binding pocket and its potential as a target for rational inhibitor design.
Extending this rationale further, the pyrazolopyrimidine (PP) inhibitors 1-NA-PP1 and 1-NM-PP1, originally developed as selective kinase inhibitors for prostate cancer research and found to be inactive against wild-type ePKs due to steric clashes with a “gatekeeper” residue in the ATP-binding site11, were shown to bind APH(3’)-Ia (Ki = 21.5 µM and Ki = 34.4 µM, respectively). When tested in a hyperpermeable E. coli ΔtolC ΔbamB strain expressing aphA1, both inhibitors were able to almost completely restore kanamycin effectiveness. However, they failed to resensitize wild-type E. coli strains expressing aphA1, likely due to reduced permeability and/or active efflux12.
Exploiting distinct allosteric sites often offers a way to bypass cross-reactivity between bacterial phosphotransferases and mammalian kinases, which commonly share conserved ATP-binding domains.
The docking study performed by Leban et al. identified compound NL8 as an ATP-competitive APH(3’)-IIIa inhibitor, and NL6, which showed activity against both APH(3’))-IIIa from S. aureus and Enterococci and APH(2”)-IVa from E. casseliflavus. NL6, identified as a non-competitive APH inhibitor was further optimized, unfortunately resulting into analogs which displayed lower efficiency13.
In a recent study by Kaplan et al., a combination of an in silico screening and molecular dynamics simulations led to the discovery of compound EK3, currently the most potent allosteric inhibitor of APH(2”)-IVa identified to date. Although EK3 stood out as a promising allosteric inhibitor, the authors emphasize that further structure-based optimization is required to improve its in vitro potency8.
Alternative strategies to counteract enzyme-mediated antibiotic resistance have focused on developing next-generation antibiotics, such as plazomicin14, whose structure has been chemically modified to specifically evade inactivation by some AMEs15,16. Despite its significant antibacterial potential against MDR Enterobacteriaceae, plazomicin faces several limitations that restrict its clinical use. These include susceptibility to certain AMEs, including APH(2”)-IVa16, and challenges in achieving sustainable market access due to financial and commercial hurdles17. Another example is represented by propylamycin and analogs thereof, developed with the aim of reducing ototoxicity and overcoming inactivation by APH(3’) enzymes18.
Further alternative approaches have utilized a combinatorial library of designed ankyrin repeat (AR) proteins as allosteric inhibitors of APH(3’)-IIIa, representing promising starting points for rational drug design19,20. Although progress has been made, their clinical application remains distant and requires further studies.
The de novo design of APH inhibitors is still in its early stages, with structure-based design and virtual screening emerging as promising but still underexplored strategies. For example, a library of 1,3-diamines was rationally designed to mimic the 2-deoxystreptamine ring of the aminoglycoside pharmacophore without the sugar backbone, showing competitive inhibition of key AMEs such as ANT(2”) and APH(3’)21.
More recently, a virtual screening performed by Parulekar et al. identified ZINC71575479 as a competitive inhibitor of APH(5) that could completely inhibit APH(5) from the multidrug-resistant organism Bacillus subtilis subsp. subtilis strain RK. In silico toxicity studies showed that ZINC71575479 has an acceptable toxicity profile, indicating its potential as a lead compound for developing inhibitors targeting APH enzymes22.
In the accompanying paper23, we report a fragment-based screening in which we identified the 7-azaindole 1 and the pyridine-2-amines like 2 as ATP-competitive inhibitors of APH(2”)-IVa, APH(3’)-IIa from K. pneumoniae and APH(3’)-IIb from P. aeruginosa (Fig. 1). Here, we present a structure-activity relationship (SAR) study to explore the scope of these scaffolds as APH inhibitors and as potential anti-resistance agents.
Crystal structures of APH(2”)-IVa in complex with 1 (A, PDB 9QOD) and 2 (B, PDB 9QNQ) and their corresponding omit maps contoured at a sigma level of ± 1. Inhibitors are represented in grey sticks, water molecule as red sphere and residues involved in interactions are shown as yellow lines. Interactions are shown as dashed lines: Van der Waals interactions in grey, hydrogen bonds in blue and ionic bonds in orange.
Results
Based on the cocrystal structure of 1 with APH(2”)-IVa (Fig. 1A), we initially focused on the functionalization of position 3’ of the pyrazole ring of 1, by assessing the tolerability toward various substitutions. The addition of a residue at this position would extend the molecule toward the catalytic triads consisting of Lys46, Glu60 and Asp217, which play a role in the coordination of the phosphates of the endogenous substrates ATP or GDP24. Therefore, engaging into new interactions could improve binding affinity of the resulting APH inhibitors.
To investigate the tolerability toward substitution in position 3’ of the pyrazole ring, we first synthesized the methyl-substituted analog 10. Toward this, the commercially available 3 was treated with m-CPBA to provide N-oxide 4, which was in turn regioselectively brominated to yield 5. Treatment of the latter with benzenesulfonyl chloride gave 6 which was further subjected to a Sonogashira reaction with but-3-yn-2-ol 7 to give intermediate 8 in 80% yield. Subsequent oxidation with Dess-Martin Periodinane (DMP) successfully converted 8 into the ynone 9, obtained in 50% yield. Finally, treatment with hydrazine and H2SO4 gave the desired pyrazole-containing analogue 10 in 61% yield (Fig. 2A). We also pursued the triazole derivative 14a, as the additional nitrogen atom aligned with the desired exit vector in the 3’-position. Toward this end, 6 was subjected to a Sonogashira reaction with trimethylsilylacetylene 11 to provide 12 in 79% yield. Subsequently, treatment with K2CO3 successfully removed both the silyl and the phenylsulfonyl groups yielding 13 in 73% yield. Finally, Cu-catalyzed reaction of alkyne 13 with NaN3 provided the desired triazole 14a (Fig. 2A).
A Synthesis of 7-azaindole derivatives 10 and 14a:a) a) m-CPBA, DME:n-Hexane (2:1, v/v), rt; b) tetramethylammonium bromide, methanesulfonic anhydride, DMF, -10 °C to rt; c) c) benzenesulfonyl chloride; NaH, DMF, 0 °C to rt; d) Pd(dppf)Cl2, TMEDA/DMF (1:1) 90 °C; e) DMP, DCM, 0 °C to rt; f) NH2NH2 • H2O, H2SO4, NaOH, EtOH, reflux to rt; g) Pd(PPh3)4, TMEDA/DMF (1:1) 80 °C; h) K2CO3, MeOH, rt; i) NaN3, sodium ascorbate, CuSO4(H2O)5, H2O/t-BuOH (1:1), rt. B Synthesis of the pyridine-2-amine derivatives 19 and 22. a) NCS, DMF, -20 °C to rt; b) Pd(dppf)Cl2, XPhos, K2CO3, dioxane/H2O (6:1), 80 °C, 4 h; c) 3 M HCl, MeOH; d) CuI, PPh3, Pd(OAc)2, ACN/Et3N, 85 °C, 20 h; e) Jones reagent, acetone, 0 °C to rt; f) NH2NH2•H2O, EtOH.
In parallel, we tested the versatility of the pyridine-2-amine scaffold (Fig. 2B) by synthesizing 19 and 22 as direct analogs of 1, 10, and 2. Toward this, the commercially available 4-bromopyridin-2-amine 15 was treated with N-chlorosuccinimide (NCS) and the desired regioisomer 16 was isolated in 31% yield (Fig. 2B). Suzuki coupling between 16 and the commercially available 1-(tetrahydro-2H-pyran-2-yl)-5-(4,4,5,5-tetramethyl-1,3,2-dioxaborolan-2-yl)-1H-pyrazole 17 gave 18 in moderate yield (57%). Finally, treatment with 3 M HCl successfully cleaved the THP protecting group and afforded the desired aminopyridine derivative 19 in 67% yield. 22 was synthesized following the same pathway used for 10 (Fig. 2B).
The effects of 10, 14a, 19 and 22 on the thermostability and activity of APH(2”)-IVa were measured by thermal shift assay and using the Pyruvate Kinase-Lactate Dehydrogenase-coupled assay (Table 1). Overall, these orthogonal assays correlated, provided a reliable assessment of the inhibitory activity of the compounds, and confirmed the ATP-competitive mode of inhibition.
10 showed an inhibition constant comparable to that of 1 (Ki = 388 nM, Table 1), while the inhibition efficacy of the unsubstituted triazole analogue 14a slightly dropped. The enzyme inhibition assay revealed a 52- and 25- fold decrease in binding affinity for pyridine-2-amine analogs 19 and 22 when compared to 1. As a result, we discontinued further exploration of the pyridine-2-amine scaffold (Table 1 and Supplementary Fig. 6).
Intrigued by the performance of 14a, we solved the co-crystal structure in complex with APH(2”)-IVa (Fig. 3). This confirmed the desired binding mode and revealed the nitrogen in position 3’ as a promising exit vector to engage residues such as Asp217, Glu60 and Lys46. Therefore, a first series of triazole analogs (14b-l) was synthesized (Fig. 4A), following the same route used for 14a and using the commercially available azides (Fig. 4C) The acid derivative 14 d was further amidated or esterified to provide 23, 24, 25 and 26 (Fig. 4).
Crystal structure of APH(2”)-IVa in complex with 14a (PDB 9QOT) and its corresponding omit map contoured at a sigma level of ± 1. Inhibitor is represented in grey sticks, residues involved in interactions are shown as yellow lines and water molecule is represented as red sphere. Interactions are shown as dashed lines: van der Waals interactions in grey, hydrogen bonds in blue and ionic bonds in orange.
A a) Azides, sodium ascorbate, CuSO4(H2O)5, H2O/t-BuOH (1:1), rt. B a) di(1H-imidazol-1-yl)methanone, NH3, DMF, rt; b) Benzylamine or p-methoxybenzylamine, DIPEA, HATU, DMF, 25 °C. C Commercially available azides used for derivatization. D Custom-made azides used for derivatization.
The effects of triazole analogs were measured as before by thermal shift assay, and in the enzymatic-coupled assay (Table 2). The latter revealed that introducing the charged carboxylic acid (14 d, Ki = 14,500 nM) compromised binding, whereas unsubstituted amide derivative 24 (Ki = 799 nM) and the ester 23 (Ki = 425 nM) improved binding compared to 14a. The presence of the hydroxyl group in compound 14 d probably works against this stacking. In general, introducing aromatic residues was not tolerated, with the notable exception of 14j (Ki = 376 nM). We were pleased to observe that the alcohol 14c (Ki = 458 nM) had improved affinity compared to 14a (Table 2).
In light of the promising biochemical activities of the 7-azaindole triazoles, we determined their first-line ADME properties to assess their principal suitability as potential drugs (Table 3). With the exception of 14e, 14 f, 14 g, 14i, all tested 7-azaindole triazoles were highly stable in murine and human microsomes. Most compounds were stable in human plasma, but several of them were only partially stable in mouse plasma, which could be attributed to slightly different enzymes available, highlighting the importance of the multiparameter optimization. Several compounds were highly bound to human plasma proteins, which was slightly less pronounced for mouse plasma proteins. A certain extent of plasma protein binding is highly accepted, as it is thought to lower renal clearance, because only the free fraction will be excreted. By contrast, high plasma protein binding (>95-99%) is associated with the concern of low efficacy as a result of only a small free fraction. However, also drugs on the market, like dalbavancin, harbor high plasma protein binding (>99%) and are still effective25. An even higher plasma protein binding capacity has been observed for a drug class in preclinical development, cystobactamids, harboring a plasma protein binding of 100% in in vitro assays. Despite that, efficacy was observed leading to the hypothesis that plasma protein binding capacities might be altered during infection26. Although plasma protein binding was high here, overall, the inhibitors had favorable ADME properties, with good metabolic and plasma stability and an acceptable plasma protein binding in light of the indication for treatment of bacterial infections.
The crystal structures of APH(2”)-IVa in complex with compounds 14b, 23, 24, 14 f, and 14k confirmed a binding mode similar to 14a. These structures helped us explore structurally the binding pocket (Fig. 5). Indeed, even if these compounds were not better at inhibiting the enzyme, they showed interactions with the loop of the β-sheet located just above the ATP binding site. For example, the presence of the methoxy group in compound 23 favors stacking on the strands 1 and 2 of the neighbouring beta sheet. The presence of the hydroxyl group in compound 14d probably works against this stacking. Moreover, the short and/or rigid extensions do not allow flexibility to interact with the loop that is highly flexible itself. This suggests that by adding a longer extension to these compounds, we could potentially stabilize them with this flexible loop and make new interactions with the protein that may increase their inhibitory effect.
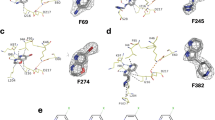
Crystal structures of APH(2”)-IVa in complex with 14a (A, PDB 9QOT),14b (B, PDB 9QOX), 23 (C, PBD 9RL1), 24 (D, PBD 9QOW), 14 f (E, PDB 9QOZ) and 14k (F, PDB 9QOU) and their corresponding omit maps contoured at a sigma level of ± 1. Inhibitors are represented in grey sticks, residues involved in interactions are shown as yellow lines. Interactions are shown as dashed lines: van der Waals interactions in grey, hydrogen bonds in blue and ionic bonds in orange.
To further improve affinity, we systematically introduced methyl groups at the hydroxy ethylene substituent of 14b by employing a set of substituted azidoethanol building blocks (Fig. 4D), yielding 14b1-b4 (Fig. 4A)27. This indeed improved affinity in all four cases when compared to 14b (Table 4).
The promising affinity and excellent ADME profile of 14c motivated us to further synthesize analogs 14c1-c6, containing substituted azidopropanol units including the diols 14 m and 14n. Gratifyingly, all analogs retained high binding to APH(2”)-IVa ( < 500 nM, Table 4). Slight improvements over 14c were observed for several analogs such as 14c1 (Ki = 325 nM), 14c3 (Ki = 375 nM), 14c4 (Ki = 367 nM), and 14c6 (Ki = 348 nM).
Inspired by the results, we solved the crystal structures of APH(2”)-IVa in complex with some of the synthesised analog compounds (14b1, 14c1, 14c3-c6, 14 m and 14n), showing interactions with the loop of the β-sheet located just above the binding pocket (Fig. 6). The extension arm of these compounds is stabilised via Van der Waals interactions with either the backbone or the side chains of the residues forming this loop. The fact that the efficacy of most of these compounds is not diminished compared to 14c is promising as this region of the protein can be highly flexible. Moreover, considering that we can stabilise this loop without losing effect, we can now focus on targeting specifically the residues located at the junction of the ATP and the aminoglycoside binding sites to increase selectivity.
Crystal structures of APH(2”)-IVa in complex with 14b1 (A, PDB 9QP6), 14c1 (B, PDB 9QP0), 14c3 (C, PDB 9QP1), 14c4 (D, PDB 9QP2), 14c5 (E, PDB 9QP3), 14c6 (F, PDB 9QP5) 14m (G, PDB 9QP7), 14n (H, PDB 9QPA) and their corresponding omit maps contoured at a sigma level of ± 1. Inhibitors are represented in grey sticks, residues involved in interactions are shown as yellow lines and water molecules are represented as red spheres. Interactions are shown as dashed lines: van der Waals interactions in grey, hydrogen bonds in blue and ionic bonds in orange.
Exemplarily, compounds 14c2, 14c3 and 14c4 were characterized for their ADME properties (Table 3). This confirmed that the modifications in the triazole substituent part retained the excellent drug-like profile of 14c.
Additionally, the most advanced APH inhibitors were tested in a broad kinase screening to assess the selectivity versus human kinases.28,29 Toward this goal, a panel of 101 purified human kinases and common kinase off-targets were tested in a thermal shift assay. The major human off-targets identified were BIKE, AAK1, AurA, MSK1_b, MAP3K5, DRAK1 and AURKB (Table 5 and Supplementary Table 2). We assume that the affinity for the off-targets lies in the high micromolar to millimolar range. As an example, a waterfall plot illustrating the activity of 1 is provided in the Supplementary Fig. 2. Notably, Erk-2 was only minimally inhibited by 1 (Supplementary Fig. 3), consistent with the cocrystal structure of 1 with Erk-2 (PDB 9QQJ, Supplementary Fig. 5), which revealed an opposite/reversed binding mode to that observed when 1 interacts with APHs. Inhibition of these human targets has to be considered in assays with human or mammalian cells. Further development of these APH inhibitors should aim at improving the selectivity ratio vs these human kinases.
Based on the promising biochemical activity and the positive ADME assessment, diazole 10 and the triazoles 14a, 14b and 14c were further characterized in several bacterial uptake/permeability assays. First, we validated our uptake assay using ciprofloxacin as an established positive control30 with the wild-type E. coli and the ΔtolC strain, deficient of an active efflux pump. As previously described, higher concentrations of ciprofloxacin were observed in the cytoplasm when the ΔtolC strain was used (Supplementary Fig. 1). Gratifyingly, the alcohols 14b and 14c, but not 10 and only partially 14a showed cellular uptake for wild-type E. coli (Fig. 7A). Primarily, compounds were detected in the periplasm, but only in very low concentration in the cytoplasm. When using the E. coli ΔtolC strain, higher concentration of all four compounds were found in the cyto- and periplasm (Fig. 7B). Next, we investigated the cellular uptake for other Gram-negative bacteria, such as K. pneumoniae (wild type) and P. aeruginosa (wild type). It was observed that slightly higher concentrations compared to E. coli were detected in the cytoplasm, but also in the periplasm. Overall, compound levels remained low (Fig. 7C, D). Whereas the alcohols 14b and 14c had slightly higher cellular uptake in K. pneumoniae, 10 and 14a had a more favorable uptake in P. aeruginosa (Fig. 7D). We hypothesize that this might be attributed to slight differences in transporters and the cell membrane in these different bacterial pathogens. Moreover, ten-fold higher concentrations in cytoplasm, membrane and periplasm were observed for all compounds tested with P. aeruginosa. We can only hypothesize that this might be attributed to a better attachment to the membrane resulting in subsequent higher concentrations being taken. As Gram-positive strains typically harbor structurally different efflux pumps, i.e. RND family efflux pumps, such as AcrAB / tolC, which are only present in Gram-negative bacteria and mainly responsible for efflux of aminoglycosides, as well as a different membrane structure31,32,33 we investigated cellular uptake in E. faecalis and methicillin-resistant S. aureus (MRSA). Moreover, we tested three additional compounds, i.e. 14c2, 14c3 and 14c4. As expected, high levels in periplasm and cytoplasm were observed for all tested compounds. Compounds 14c2, 14c3 and 14c4 had higher levels in cytoplasm compared to periplasm, for both E. faecalis and MRSA (Fig. 7E, F). Thus, it can be concluded that the structural variations introduced in 14c2, 14c3 and 14c4 led to an increased uptake into cytoplasm in the Gram-positive strains tested herein compared to the first-generation derivatives 10, 14a, 14b and 14c.
A Uptake assay for 14a, 10, 14b and 14c in E. coli. B Uptake assay for 14a, 10, 14b and 14c in E. coli ΔtolC strain. C) Uptake assay for 14a, 10, 14b and 14c in K. pneumoniae. D) Uptake assay for 14a, 10, 14b and 14c in P. aeruginosa. E) Uptake assay for 14a, 10, 14b, 14c, 14c2, 14c3 and 14c4 in E. faecalis. F) Uptake assay for 14a, 10, 14b, 14c, 14c2, 14c3 and 14c4 in methicillin-resistant S. aureus. Assays were conducted in biological triplicate. The results are displayed as mean and standard deviation. Concentrations were assessed in membrane, periplasm, cytoplasm and cell lysate (from left to right). Blue bars 14a, red bars 10, green bars 14b, purple bars 14c, orange bars 14c2, black bars 14c3, brown bars 14c4.
Since 10 had a more favorable uptake in P. aeruginosa (Fig. 7D), we monitored its effect on bacterial growth of the clinical P. aeruginosa strain C0214 (Fig. 8A). Increasing kanamycin concentrations progressively inhibited growth, with a MIC of 64 µg mL-1. The addition of 10 did not change the MIC value but retarded bacterial growth in a concentration-dependent manner, which was most evident for a kanamycin concentration of 32 µg mL-1 (Fig. 8A and Supplementary Fig. 4A). As expected, the addition of compound 10 alone did not affect bacterial growth (first column of the checkerboard assay and black curves in Supplementary Fig. 4A). Similar results were observed for compound 14n and 14c (Fig. 8B, C and Supplementary Fig. 4B, C). Overall, these results indicate a slight effect the APH inhibitor on the bacteria growth at the concentration below the MIC of kanamycin A. However, the impact of these compounds on bacterial growth remains moderate. This weak effect observed on bacteria may be due to the high intracellular concentration of ATP which competes with the APH inhibitors that have sub-micromolar affinities. Further optimization is therefore still needed to improve their efficacy.
MIC determinations with P. aeruginosa C0214 in the presence of increasing concentrations of kanamycin A (from left to right) and of 10 A, 14n B or 14c C (from top to bottom). Cells of the table are stained with a linear color gradient according to OD600nm, from white at 0 to dark blue at the maximum value in the plate, indicated next to the color scale. The first column shows the effect of inhibitors in the absence of kanamycin A. The last column represents the negative control (medium with inhibitors but without bacteria). Growth curves are shown in Supplementary Fig. 4.
Conclusion
Developing targeted strategies to counter antibacterial resistance mechanisms is of utmost importance to restore efficacy of established antibiotics. In this context, inhibiting APHs represents a compelling approach to directly tackle a key antibiotic resistance mechanism. By concomitantly administering the aminoglycoside and APH inhibitor, antibiotic efficacy may be achieved by neutralizing one of the most relevant mechanism of bacterial resistance to aminoglycosides. Moreover, the co-administration of APH inhibitors with aminoglycosides might help to prevent the emergence of resistance development in strains that have not yet acquired APH activity.
Starting from the fragment screening hit 1, we explored the pyrazole ring of 1. This resulted in 10 or the triazole analogs 14a or 14c, which potently inhibited APH and displayed good ADME profile, incl. a more favorable uptake in multiple, particularly Gram-positive, bacterial strains. Moreover, several of the advanced compounds showed beneficial properties and enhanced the efficacy of the aminoglycoside kanamycin on a clinical P. aeruginosa strain expressing APH(3’)-IIb, although further optimization will be necessary before potential clinical application.
A broad profiling of human kinases revealed that sparing human kinases is in general possible. However, some human kinases such as Bike, AurA, or MSK_b displayed cross-reactivity, which must be addressed in future optimization studies.
In summary, we show that the bacterial kinases such as APHs are valid targets to tackle antibiotic resistance that can be reversed with drug-like molecules and are highly suitable for structure-based optimization.
Methods
Synthetic procedures, NMR spectra and analytical data for the synthesized compounds, details of the X-ray crystallographic analyses, ADME and uptake studies, bacterial assays and kinase screening are given as Supplementary Information.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and Supplementary Data. Raw data used to generate figures and tables are provided as supplementary material in a spreadsheet named Supplementary Data 1.
References
Krause, K. M., Serio, A. W., Kane, T. R. & Connolly, L. E. Aminoglycosides: An overview. Cold Spring Harb. Perspect. Med. 6, 1–18 (2016).
Antimicrobial Resistance Division (AMR), I. I. and R. C. (IRC). WHO bacterial priority pathogens list, 2024: Bacterial pathogens of public health importance to guide research, development and strategies to prevent and control antimicrobial resistance. 72 (2024).
Wright, G. D. Aminoglycoside-modifying enzymes. Curr. Opin. Microbiol. 2, 499–503 (1999).
Daigle, D. M., McKay, G. A., Thompson, P. R. & Wright, G. D. Aminoglycoside antibiotic phosphotransferases are also serine protein kinases. Chem. Biol. 6, 11–18 (1999).
Ramirez, M. S. & Tolmasky, M. E. Aminoglycoside modifying enzymes. Drug Resist. Updat. 13, 151–171 (2010).
Shi, K., Caldwell, S. J., Fong, D. H. & Berghuis, A. M. Prospects for circumventing aminoglycoside kinase mediated antibiotic resistance. Front. Cell. Infect. Microbiol. 4, 1–17 (2013).
Magaña, A. J. et al. Restoring susceptibility to aminoglycosides: identifying small molecule inhibitors of enzymatic inactivation. RSC Med. Chem. 14, 1591–1602 (2023).
Kaplan, E. et al. APH Inhibitors that Reverse Aminoglycoside Resistance in Enterococcus casseliflavus. ChemMedChem 20, 1–11 (2025).
Daigle, D. M., McKay, G. A. & Wright, G. D. Inhibition of aminoglycoside antibiotic resistance enzymes by protein kinase inhibitors. J. Biol. Chem. 272, 24755–24758 (1997).
Fong, D. H. & Berghuis, A. M. Crystallization and preliminary crystallographic analysis of 3′-aminoglycoside kinase type IIIa complexed with a eukaryotic protein kinase inhibitor, CKI-7. Acta Crystallogr. Sect. D. Biol. Crystallogr. 60, 1897–1899 (2004).
Tandon, M. et al. New Pyrazolopyrimidine Inhibitors of Protein Kinase D as Potent Anticancer Agents for Prostate Cancer Cells. PLoS ONE 8, (2013).
Stogios, P. J. et al. Structure-guided optimization of protein kinase inhibitors reverses aminoglycoside antibiotic resistance. Biochem. J. 454, 191–200 (2013).
Leban, N., Kaplan, E., Chaloin, L., Godreuil, S. & Lionne, C. Kinetic characterization and molecular docking of novel allosteric inhibitors of aminoglycoside phosphotransferases. Biochim. Biophys. Acta - Gen. Subj. 1861, 3464–3473 (2017).
Shaeer, K. M., Zmarlicka, M. T., Chahine, E. B., Piccicacco, N. & Cho, J. C. Plazomicin: A Next-Generation Aminoglycoside. Pharmacotherapy 39, 77–93 (2019).
Golkar, T. et al. Structural basis for plazomicin antibiotic action and resistance. Commun. Biol. 4, 1–8 (2021).
Cox, G. et al. Plazomicin Retains Antibiotic Activity against Most Aminoglycoside Modifying Enzymes. ACS Infect. Dis. 4, 980–987 (2018).
Wells, N., Nguyen, V. K. & Harbarth, S. Novel insights from financial analysis of the failure to commercialise plazomicin: Implications for the antibiotic investment ecosystem. Humanit. Soc. Sci. Commun. 11, 1–13 (2024).
Lubriks, D. et al. Synthesis and Antibacterial Activity of Propylamycin Derivatives Functionalized at the 5′′- And Other Positions with a View to Overcoming Resistance Due to Aminoglycoside Modifying Enzymes. ACS Infect. Dis. 7, 2413–2424 (2021).
Amstutz, P. et al. Intracellular kinase inhibitors selected from combinatorial libraries of designed ankyrin repeat proteins. J. Biol. Chem. 280, 24715–24722 (2005).
Kohl, A. et al. Allosteric inhibition of aminoglycoside phosphotransferase by a designed ankyrin repeat protein. Structure 13, 1131–1141 (2005).
Welch, K. T. et al. Discovery of non-carbohydrate inhibitors of aminoglycoside-modifying enzymes. Bioorg. Med. Chem. 13, 6252–6263 (2005).
Parulekar, R. S., Barale, S. S. & Sonawane, K. D. Antibiotic resistance and inhibition mechanism of novel aminoglycoside phosphotransferase APH(5) from B. subtilis subsp. subtilis strain RK. Braz. J. Microbiol. 50, 887–898 (2019).
Kowalewski, J. et al. Fragment-based drug design of a bacterial kinase inhibitor capable of increasing antibiotic sensitivity of clinical isolates. Commun. Chem. Under revision. https://doi.org/10.1038/s42004-025-01795-6.
Shakya, T. et al. A small molecule discrimination map of the antibiotic resistance kinome. Chem. Biol. 18, 1591–1601 (2011).
Turner, N. A., Xu, A., Zaharoff, S., Holland, T. L. & Lodise, T. P. Determination of plasma protein binding of dalbavancin. J. Antimicrob. Chemother. 77, 1899–1902 (2022).
Kohnhäuser, D. et al. Optimization of the Central α-Amino Acid in Cystobactamids to the Broad-Spectrum, Resistance-Breaking Antibiotic CN-CC-861. J. Med. Chem. 67, 17162–17190 (2024).
Buffa, V. et al. Conformational Plasticity and Binding Affinity Enhancement Controlled by Linker Derivatization in Macrocycles. Angew. Chem. - Int. Ed. 64, e202418512 (2025).
Fedorov, O., Niesen, F. H., Knapp, S. Kinase inhibitor selectivity profiling using differential scanning fluorimetry. Methods Mol. Biol. 795, (2012).
Amrhein, J. A., Berger, L. M., Balourdas, D. I., Joerger, A. C., Menge, A., Kramer, A., Frischkorn, J. M., Berger, B. T., Elson, L., Kaiser, A., Schubert-Zsilavecz, M., Muller, S., Knapp, S. & Hanke, T. Synthesis of Pyrazole-Based Macrocycles Leads to a Highly Selective Inhibitor for MST3. J. Med Chem. 67, 674–690 (2024).
Prochnow, H. et al. Subcellular Quantification of Uptake in Gram-Negative Bacteria. Anal. Chem. 91, 1863–1872 (2019).
Kumawat, M. et al. Role of bacterial efflux pump proteins in antibiotic resistance across microbial species. Microb. Pathog. 181, 106182 (2023).
Thakur, V., Uniyal, A. & Tiwari, V. A comprehensive review on pharmacology of efflux pumps and their inhibitors in antibiotic resistance. Eur. J. Pharmacol. 903, 174151 (2021).
Blair, J. M. A., Richmond, G. E. & Piddock, L. J. V. Multidrug efflux pumps in gram-negative bacteria and their role in antibiotic resistance. Future Microbiol. 9, 1165–1177 (2014).
Acknowledgements
K.R. thanks Prof. Dr. Mark Brönstrup for providing the E. coli ΔtolC strain. Moreover, K.R. thanks Kimberley Vivien Sander, Janine Schreiber and Jennifer Wolf for excellent technical assistance. Furthermore, K.R. and F.H. thank the German Federal Ministry of Education and Research (BMBF, SIAM-APH, No. 16GW0235, grant number: 01KI2126B) for funding. Work in the lab of K.R. is funded by the German Center for Infection Research (DZIF, TTU 09.719). J.K., M.T., J.-F.G. and C.L. thank the ANR (ANR-19-AMRB-0001), the Association Vaincre la Mucoviscidose (RF20220503015), and the Association Grégory Lemarchal for financial support. J.K. is thankful to Fonds France-Canada pour la Recherche and Mitacs for travelling grants. The CBS is part of the ChemBioFrance research infrastructure and is a member of the FranceBioImaging (FBI) and the French Infrastructure for Integrated Structural Biology (FRISBI), two national infrastructures supported by the Agence Nationale de la Recherche (ANR-10-INBS-0004 and ANR-10-INBS-0005, respectively). The authors thank Dr. Muriel Gelin for determining the structure of Erk-2 in complex with compound 1. The authors thank Prof. Gerard D. Wright for the donation of the clinical strains from Hamilton General Hospital (Hamilton, ON, Canada).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conceptualization: F.H., V.B., J.K., C.L., J.-F.G., K.R.; Draft Writing: V.B. Formal Analysis: F.H., V.B., R.D, J.K., C.L., C.D.-M., J-F.G., K.R.; Funding acquisition: F.H., C.L., J-F.G., K.R, S.K.; Investigation: V.B., J.K., R.D., A.K., M.-R., M.T., G.Q., M.C.; Supervision: F.H., C.L., J-F.G., K.R.; Reviewing and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Yinhu Wang, Mark McLaughlin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Buffa, V., Kowalewski, J., Qi, G. et al. Targeting bacterial kinases as a strategy to counteract antibiotic resistance. Commun Chem 8, 390 (2025). https://doi.org/10.1038/s42004-025-01794-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01794-7
This article is cited by
-
Fragment-based drug design of a bacterial kinase inhibitor capable of increasing the antibiotic sensitivity of clinical isolates
Communications Chemistry (2025)