Abstract
Conotoxins are small, disulfide-rich peptides that display exceptional affinity and selectivity for ion channels and receptors, making them valuable templates for therapeutic development. However, their optimization remains challenging due to the limited diversity of naturally occurring variants and the labor-intensive nature of conventional engineering strategies. Here, we present CreoPep, a deep learning-based generative framework specifically developed to design and optimize conotoxins targeting defined receptors. CreoPep integrates masked language modeling with a progressive masking scheme and employs an augmentation pipeline that combines physics-based energy screening with temperature-controlled multinomial sampling. This enables the generation of structurally and functionally diverse peptide variants while retaining essential pharmacological features. Structural analysis shows that CreoPep-generated variants adopt both conserved and previously unobserved binding modes, including disulfide-deficient forms. Together, these findings establish CreoPep as a powerful computational-experimental framework for the rational design of conotoxin-based peptides and provide a foundation for extending similar approaches to other peptide families.
Similar content being viewed by others
Introduction
Target-specific peptides are biomolecules that bind with high affinity and specificity to biological targets, such as membrane receptors or ion channels, thereby modulating their function (Fig. 1a–c)1,2. Among these, conotoxins, bioactive peptides derived from marine cone snail venom, exhibit exceptional potency and selectivity across various ion channel families, including voltage-gated calcium (Cav) channels, nicotinic acetylcholine receptors (nAChRs), and voltage-gated sodium (Nav) channels3,4,5,6. Their precision in regulating neural activity makes conotoxins promising scaffolds for therapeutic development3,7.
a ω-conotoxin features three disulfide bonds. An example is ω-conotoxin MVIIA, which targets Cav2.2 channels (PDBid: 7MIX). b α-conotoxin contains two disulfide bonds. Examples include α-conotoxin ImI and Vc1.1, which primarily target nAChRs (PDBid: 7KOO). c µ-conotoxin has three disulfide bonds. An example is µ-conotoxin KIIIA, which blocks voltage-gated sodium channels (PDBid: 6J8E). d Saturation mutagenesis, where each amino acid residue in the loop1 and loop2 regions of the ImI sequence is replaced by one of the other 19 natural amino acids. Yellow lines indicate disulfide bonds. e Point mutations are introduced at specific positions identified through alanine scanning, targeting either key or non-key or key residues. f Target-specific peptide design based on deep learning, directly generating high-potency ImI mutants. g Schematic diagram of sequence space representation of ImI mutants. The purple dot represents the wild-type peptide, the orange dots represent peptides generated by point mutations, the large purple circle indicates the exploration range of point mutations, the gray dots represent peptides produced by saturation mutagenesis, and the pink dots represent peptides generated by deep learning (DL) methods.
Conotoxins are stabilized by conserved disulfide bonds and classified into more than 30 distinct structural frameworks, each associated with characteristic pharmacological profiles8. For example, ω-conotoxins target Cav channels9 (Fig. 1a), α-conotoxins modulate nAChRs10 (Fig. 1b), and μ-conotoxins block Nav channels11 (Fig. 1c). Despite their conserved structural scaffolds, conotoxins display extraordinary sequence diversity, with an estimated 1 million bioactive variants existing in nature. However, fewer than 1% (~10,000) have been sequenced, and only a small subset characterized functionally. This striking diversity highlights their therapeutic potential2,3,12, exemplified by ω-conotoxin MVIIA13,14, a 25-residue peptide from Conus magus venom that selectively inhibits N-type Cav channels and is approved for chronic pain treatment. However, naturally derived peptides often exhibit suboptimal activity, unintended off-target effects, and limited metabolic stability, posing barriers to clinical translation1. Addressing these challenges typically involves extensive mutagenesis and chemical modification, a labor-intensive process that underscores the need for more efficient peptide engineering strategies15,16,17.
Current peptide engineering approaches generally fall into two categories: saturation mutagenesis (Fig. 1d) and targeted point mutation (Fig. 1e). Saturation mutagenesis substitutes selected positions with all 20 natural amino acids, followed by structural prediction (e.g., RosettaFold18, AlphaFold319) and binding affinity estimation (e.g., FoldX20, FlexPepDock21, HPEPDOCK22). Although capable of identifying high-affinity mutants and exploring broad sequence space, this method is computationally expensive and yields relatively few viable candidates. In contrast, targeted point mutation approaches, such as alanine scanning are more efficient but rely heavily on prior structural knowledge and provide limited exploration of sequence diversity, making them less suited for de novo design. Notably, both strategies depend on fixed backbone conformations, restricting access to novel structural motifs—a key limitation in advancing peptide therapeutics.
Recent advances in deep learning are beginning to transform drug discovery, providing new opportunities to accelerate peptide and protein design23,24,25,26,27,28,29,30,31,32,33. Models trained to predict protein–peptide interactions (PpIs), can rapidly screen large peptide libraries. For example, Lei et al. developed a convolutional neural networks (CNNs) integrated with self-attention to predict PpIs directly from sequence data while identifying peptide binding residues34. However, the applicability of such models is constrained by the limited availability of high quality training data. Beyond screening, deep generative models can directly design peptide analogs with improved affinity. Deng et al. introduced a reinforcement learning-based model, RLpMIEC, which integrates peptide-MHC interaction energy spectra and sequence features to generate peptides with high binding affinity for MHC-I molecules35. Similarly, Chen et al. applied a gated recurrent unit (GRU)-based variational autoencoder (VAE) with Metropolis-hastings (MH) sampling to generate peptide inhibitors targeting β-catenin and NF-κB essential modulator36. Despite promising results, these methods often rely on target-specific datasets and explore limited conformational diversity, restricting their generalizability.
In this study, we develop a computational strategy tailored specifically for the design and optimization of conotoxin peptides (Fig. 1f, g). We introduce CreoPep, a conditional generative framework based on a masked language model inspired by diffusion principles37. CreoPep uses a progressive masking (PM) strategy that incrementally increases masked tokens during training (adding noise) and gradually predicts them during generation (denoising). This enables the model to better learn the relationship between peptide sequence, structure, and function. To further enhance data quality, we integrate CreoPep with a peptide augmentation pipeline, incorporating multiple rounds of physics-based binding energy screening to construct a large pseudo-labeled dataset for improved training. CreoPep also incorporates temperature-controlled multinomial sampling to ensure that generated peptides maintain both structural diversity and specificity. We validate the efficacy of CreoPep by designing conotoxin peptides targeting the α7 and α4β2 nAChRs and demonstrate encouraging potency in electrophysiological assays. Collectively, our results establish CreoPep as a robust computational-experimental framework that streamlines the design, optimization, and experimental validation of conotoxins. By leveraging deep learning and advanced generative modeling, CreoPep accelerates the discovery of target-specific conotoxins and lays the foundation for extending similar approaches to other peptide families, with broad implications for drug development, synthetic biology, and personalized medicine.
Results
Improving conotoxin binding through mutant design
Current conotoxin optimization strategies focus primarily on enhancing the potency of known sequences through iterative mutagenesis. For example, α-conotoxin ImI, a compact 12-residue peptide, has been subjected to extensive point mutation studies, albeit with limited success, likely due to its evolutionarily constrained structure. To explore alternative optimization strategies, we first implemented saturation mutagenesis on the ImI/α7 nAChR system. By systematically replacing residues in both loop 1 and loop 2 with all 20 natural amino acids, we generated 1000 variants and assessed their binding free energy changes (ΔΔG) using physics-based screening (Fig. 1d). This screen identified 65 mutants (6.5% hit rate) with improved binding affinity relative to wild-type ImI (Supplementary Data 1), demonstrating the potential but limited efficiency of this approach. Furthermore, we used ProteinMPNN24 to design 1000 peptides based on the wild-type ImI backbone and calculated their ΔΔG values using FoldX (Supplementary Data 1). It can be found that only 88 of these designed peptides had a ∆∆G ≤ 0.5 kcal mol−1 (Supplementary Table 1).
To improve the success rate, we next applied a constrained mutation strategy by fixing three experimentally validated critical residues (D5, P6, R7) while randomizing the remaining positions38. This approach increased the number of favorable mutants to 253 (Supplementary Data 1), confirming that incorporating structural knowledge can guide more effective optimization. However, this improvement in efficiency came at the cost of reduced exploration across the broader sequence-structure landscape. These findings highlight a fundamental trade-off in conotoxin engineering: while structural constraints enhance hit rates, they concurrently restrict access to novel structural motifs. To overcome this limitation, we aim to develop an advanced peptide design framework that balances hit rate and conformational diversity, enabling the efficient generation of functional variants while expanding the accessible chemical and structural space.
Development of the generative model CreoPep for conotoxin mutation design
To overcome limitations in current conotoxin design, we developed CreoPep, a deep learning-based conditional generative model that leverages evolutionary information to efficiently explore conotoxin chemical space (Fig. 2a). CreoPep is built on ProtBert’s masked language modeling (MLM) framework39, pretrained on large-scale protein sequence datasets via self-supervised learning. It generates structurally diverse and pharmacologically relevant conotoxin variants through three key innovations. First, we introduced a progressive masking (PM) scheme that replaces the conventional fixed 15% masking rate with a dynamic schedule that gradually increases the number of masked tokens during training. This enables more nuanced and context-aware learning of sequence discrepancies. During sequence generation, CreoPep uses this framework to iteratively predict one masked residue at a time until a complete conotoxin sequence is reconstructed. Second, the training data for each instance included: (1) a subtype label (one of the 53 subtypes), (2) a potency label (high or low), (3) a wild-type conotoxin, and (4) auxiliary conotoxins (randomly selected peptides share the same subtype and potency as the wild-type). Subtype and potency labels provide explicit functional constraints, while auxiliary sequences offer implicit functional patterns to guide model learning. Third, all inputs except the auxiliary conotoxins were encoded using the PM scheme; auxiliary conotoxins were instead encoded via a multilayer perceptron (MLP). The combined input features were fused using a convolutional neural network (CNN) to capture higher-order representations.
a Schematic representation of the CreoPep framework. Left panel (top to bottom): training dataset format and feature extraction methods for each component, training phase, and generation phase. Right panel: mutant screening workflow using physics-based screening for peptides generated by CreoPep. b Illustration of three core functional capabilities of CreoPep. c Data augmentation pipeline used to enhance training diversity and model performance.
CreoPep integrates three key tasks into a single framework: label prediction, conditional generation, and optimization generation (Fig. 2b). (1) Label prediction, where subtype and potency are masked and inferred from the conotoxin sequence and optional auxiliary peptides. (2) Conditional generation, where masked residues are predicted iteratively under the guidance of target subtype/potency labels and auxiliary peptides to construct functionally tailored sequences. (3) Optimization generation, where multiple candidates are generated conditionally, evaluated via the label prediction module, and filtered to retain only those with improved potency scores and enhanced receptor-specific confidence relative to the inputs. This unified architecture enables efficient, functionally guided navigation of conotoxin sequence space, facilitating the design of novel peptide variants with improved binding affinity and functional specificity.
Data augmentation guided by energy-based filtering
To train CreoPep, we curated 2088 conotoxins with experimentally determined targets and potency values (IC50) from the ConoServer database8, encompassing seven receptor families divided into 53 subtypes. Potency was classified as high (IC50 ≤ 1000 nM) or low (IC50 > 1000 nM). However, the dataset exhibited sparsity and redundancy (Supplementary Fig. 1); for example, only 67 high-potency conotoxins targeted the α7 nAChR subtype, and most of these were mutants derived from a small number of wild-type sequences via residue scanning, leading to low sequence diversity. To address these limitations, we developed an iterative augmentation pipeline that integrates CreoPep with physics-based method (Fig. 2a, c), enabling progressive expansion of the training dataset through in silico generation and energy-based filtering while preserving pharmacological relevance.
FoldX was selected for its well-established accuracy in predicting binding free energy changes (ΔΔG)40, and its generalizability was validated across seven conotoxin-receptor systems using experimental IC50 data from ConoServer (Supplementary Tables 2–15). As summarized in Supplementary Table 16, FoldX achieved >75% accuracy in five systems, with lower performance in α-conotoxin Vc1.1/α9α10 and ImI/α7 (<70% accuracy). Notably, the ImI/α7 system exhibited 100% sensitivity, correctly identifying all potency-enhancing mutants. Comparable predictive performance for μ-conotoxin KIIIA/Nav1.2 and α-conotoxin LvIA/AChBP, while exceptional specificity (100%) was achieved in the ω-conotoxin MVIIA/Cav2.2 and α-conotoxin AuIB/α3β4 systems. Overall predictive reliability was further supported by the Matthews correlation coefficient (MCC) analysis, with the highest performance recorded in the LvIA/AChBP system (MCC = 0.83). These results confirm FoldX as a selective and robust screening filter for high-potency variants in our pipeline.
CreoPep was initially trained on the 2088 conotoxin sequence dataset, then used to generate 1000 mutant variants for each of eight distinct conotoxin/receptor subtype systems. To control the diversity of generated sequences, a polynomial sampling strategy with a temperature factor (τ) was applied. In the first generation round, τ was set to 1.5 to maximize peptide sequence diversity. FoldX then computed ΔΔG values between generated mutants and their corresponding wild-type peptides. Mutants with ΔΔG ≤ 0.5 kcal mol−1 were retained as high potency candidates, while sequences with ΔG(mutant) ≥ 0 kcal mol−1 were labeled low potency (see “Methods”). Both sets were merged into the training pool to retrain CreoPep. This procedure was repeated over five rounds, with τ gradually reduced from 1.5 to 1.1 to refine the search toward higher-fidelity variants. The final augmented training set consisted of 18,355 conotoxins (Supplementary Data 2), significantly increasing the size of the CreoPep training set.
Design of high-potency conotoxin mutants with controllable diversity via CreoPep
The high sequence conservation and limited number of potent conotoxin mutants in the training set constrain the diversity of generated sequences. However, excessive diversification may compromise potency. To address this trade-off, we implemented and evaluated a data augmentation pipeline designed to balance potency and diversity in the generated conotoxin mutants.
We calculated ΔΔG values for 1000 mutants across eight systems at various stages of the augmentation process (Fig. 3, violin plot). From stages C1 to C5, the ΔΔG distributions in most systems progressively shifted toward lower values, indicating an increased frequency of high-potency conotoxins (ΔΔG ≤ 0.5 kcal mol−1). After data augmentation, the number of favorable mutations in C5 is significantly higher than that in C1 (Supplementary Fig. 2). In contrast, the KIIIA/Nav1.2 system exhibited a shift toward higher ΔΔG values, suggesting reduced efficacy in potency optimization. Nonetheless, the system still yielded approximately 20 favorable mutants, outperforming random generation (Supplementary Table 1).
The x-axis represents different stages of random mutation and data augmentation, while the y-axis shows the ΔΔG values calculated using physics-based method. Violin plots display the distribution of ΔΔG values for 1000 peptide mutants, where the width of the plot at any given ΔΔG value reflects the density of data points. The median value represented by the black horizontal line, and a dashed horizontal line representing the mean within the box, the lower and upper quartiles delineating the borders of the box, and the vertical black lines indicating the 1.5 interquartile range. A line graph overlays the plots to indicate the average hamming distance between each mutant and the wild-type peptide. For each plot, the left y-axis represents ΔΔG values, the right y-axis corresponds to Hamming distance, and the right panel shows the representative complex structures for each system.
To quantify sequence diversity, we calculated the Hamming distance, a measure of sequence divergence (range: 0–1) between each mutant and their corresponding wild-type peptide (Fig.3, line plot). From stages C1 to C3, the average Hamming distance increased across all systems, reflecting a model bias toward exploring sequence diversity. Beyond C3, this trend reversed, except for minor fluctuations in MVIIA/Cav2.2 and AuIB/α3β4 systems, indicating a shift toward potency optimization. Meanwhile, with each round of data augmentation, the sequence diversity of the mutants shows a significant improvement compared to C1 (Supplementary Table 17 and Supplementary Fig. 3).
Importantly, the temperature factor (τ) was initially set to 1.5 at stage C1 and gradually reduced to 1.1 by C5. Despite this reduction, the Hamming distance did not decline immediately (C1–C3), likely due to the incorporation of highly diverse mutants generated in C1 were incorporated into subsequent training cycles, thereby amplifying overall dataset diversity. During C1–C3, this cumulative diversity outweighed the decreasing influence of τ. From C3–C5, however, the model exhibited a smooth transition from diversity exploration to potency refinement. Overall, these results demonstrate that CreoPep’s data augmentation pipeline effectively increases the number of high-potency mutants while maintaining enhanced sequence diversity.
Classification and generative performance assessment of CreoPep
To evaluate the classification performance of CreoPep throughout the iterative data augmentation process, we assessed the model’s prediction accuracy for subtype, potency, and target at each stage on its corresponding validation dataset. Given their unique pharmacological profiles, conotoxins can exhibit multitarget potency. Consequently, the model must be capable of recognizing and outputting multiple potential labels rather than being constrained to a single optimal prediction. To enable a comprehensive performance evaluation, we used the TopK accuracy metric, where a prediction is considered correct if the true label is among the top K predicted labels with the highest probabilities. The TopK accuracy of CreoPep for subtype prediction at each stage is shown in Fig. 4a. In the final stage, the model achieved a Top1 to Top5 accuracy of 76.63%, 90.85%, 95.92%, 96.73%, and 97.22%, respectively, on its validation set. For potency prediction, the final model achieved an accuracy of 93.36% (Supplementary Table 18). As shown in Fig. 4b, all models across different stages demonstrated Top1 accuracy above 95% for target prediction on their respective validation sets (Supplementary Table 19).
a Accuracy of CreoPep in subtype and potency prediction tasks. b Accuracy of CreoPep in target prediction. c–e Average pLDDT score, TM-score, and Hamming distance of 1000 mutants generated by CreoPep. f–h Comparison of isoelectric point, net charge, GRAVY (hydrophobicity) distributions among the wild-type conotoxin, high potency conotoxins, and 1000 CreoPep-generated mutants. i–p Feature distributions in the latent space for the wild-type conotoxin, high potency conotoxins, and 1000 CreoPep-generated mutants across eight systems.
To assess the model’s generative performance, we independently generated 1000 conotoxin mutants for each of the eight subtype systems using the final model, and evaluated their foldability, novelty, physicochemical properties, and latent-space features. Structural foldability, predicted using OmegaFold41, yielded average predicted local distance difference test (pLDDT) scores42 ranging from 58.93 (Nav1.2) to 70.43 (α7 nAChR) (Fig. 4c), suggesting reasonable structural viability, though slightly below the training set average of 71.46. Novelty analysis showed that the generated mutants exhibited low structural similarity to wild-type peptides (template modeling (TM) score43 < 0.5, Fig. 4d) and high sequence divergence (Hamming distance > 0.3, Fig. 4e). Mutants derived from μ-conotoxin MVIIA/Cav2.2 demonstrated especially high novelty with TM-scores < 0.2 and Hamming distance > 0.7. Analysis of physicochemical properties revealed that the generated mutants retained essential features of their wild-type counterparts, including isoelectric point (Fig. 4f), net charge (Fig. 4g), and the grand average of hydropathy (GRAVY) index44 (Fig. 4h), with distributions closely matching those of high-potency training set peptides. Consistent trends were observed across other data augmentation cycles (Supplementary Fig. 4). Finally, embeddings generated using evolutionary scale modeling-2 (ESM-245) and visualized through uniform manifold approximation and projection (UMAP46) visualization (Fig. 4i–p) confirmed strong latent-space overlap between the generated and wild-type peptides, particularly for the α3β4, α3β2, and α9α10 nAChR subtypes. This suggests that the model preserves evolutionary and structural features in its generative outputs. We also compared the latent-space distribution of mutants generated via random saturation mutagenesis to those produced during various augmentation stages (Supplementary Fig. 5). Randomly saturated mutants showed minimal overlap with the training set peptides, while the augmented mutants initially clustered with the training data (C1) and then gradually dispersed into novel regions, forming distinct clusters. Notably, after C3, a subset of mutants reconverged with the training set cluster, consistent with the trends observed in Fig. 3. Taken together, these findings demonstrate that our model is capable of generating structurally viable and novel conotoxin analogs that preserve key physicochemical and functional characteristics of the wild-type peptides.
Experimental validation of conotoxin ImI mutants generated via CreoPep
We trained the final version of CreoPep on 18,355 conotoxins generated through our data augmentation pipeline. Focusing on the ImI/α7 nAChR system selected for its experimental tractability, we generated 1000 candidate mutants under high-potency conditions (τ = 1). Candidates were initially ranked by their predicted potency probabilities, and the top 60 candidates were selected for structural analysis. For each peptide-α7 complex, five confirmations were predicted using AlphaFold3, followed by binding energy calculations with FoldX. Fifteen mutants consistently exhibited stronger binding than the wild-type ImI. Visual inspection of their binding modes in PyMOL further refined this list to 13 high-confidence candidates (CP_α7_1 to CP_α7_13), which were selected for experimental validation.
Using solid-phase peptide synthesis, we successfully produced all 13 candidate conotoxins for functional characterization. Two-electrode voltage clamp electrophysiology on human (h) α7 nAChR revealed that seven peptides showed substantial inhibitory activity (>50% blockade) at 10 µM (Fig. 5a). Notably, two designed peptides, CP_α7_1 and CP_α7_6, retained strong inhibition even at 1 µM. Quantitative concentration–response analysis confirmed submicromolar IC50 values for these top candidates, 405.1 ± 28.7 nM for CP_α7_1 and 504.6 ± 29.8 nM for CP_α7_6 (mean ± SD, n = 6) (Fig. 5b). The potency of CP_α7_1 is slightly superior to that of the wild-type, while the potency of CP_α7_6 is comparable to that of ImI47 (Supplementary Table 20), validating CreoPep’s effectiveness in generating highly potent nAChR antagonists (Supplementary Data 3).
a Bar graph showing the inhibitory effects of 13 candidate conotoxins (at 1 and 10 µM) on ACh-evoked peak current amplitudes mediated by human (h) α7 nAChRs. Whole-cell currents were evoked by 100 µM ACh (mean ± SD, n = 4–8). The dashed line indicates 50% inhibition of the peak current amplitude. b Concentration–response relationships for CP-α7-1 and CP-α7-6 inhibition of ACh-evoked currents at (h) α7 nAChRs. Current amplitudes (mean ± SD; n = 6) were normalized to the response elicited by 100 µM ACh alone. c Latent space distribution of 1000 mutants (gray dots), ImI, high potency α7-targeting conotoxins (red dots), low potency α7-targeting conotoxins (green dots), and the 13 candidate conotoxins (large yellow dots for high potency; small yellow dots for low potency). The contour plot represents the density distribution of the 1000 mutants, with darker colors indicating higher density in that region. d Multiple sequence alignment of ImI with the 13 candidate conotoxins. Among them, colors reflect the physicochemical properties of amino acids, using the Zappo protein color scheme: pink for aliphatic/hydrophobic, orange for aromatic, blue for positive, red for negative, green for hydrophilic, magenta for conformationally special, yellow for cysteine, and gray for positions without mutations. e Graphical representation of multiple sequence alignment of 1000 CreoPep-generated mutants, showing the frequency of amino acid variations at each position relative to the ImI sequence.
To evaluate CreoPep’s capacity for exploring chemical space, we mapped the 13 candidate conotoxin mutants within the latent space of the ImI/α7 system. The mutants formed three distinct and well-separated clusters (Fig. 5c). Interestingly, most candidate conotoxins co-localized with ImI, with four high-potency peptides forming a tightly packed subcluster. Notably, CP_α7_6 was located in a region distant from the main cluster, in a densely populated area of the latent space. Excitingly, both CP_α7_1 and CP_α7_6 broke the well-known S-X-P motif4, which is highly conserved in conotoxins (Fig. 5d). These findings highlight CreoPep’s ability to explore chemical space and discover novel, high potency peptide variants. An integrated computational-experimental analysis demonstrated CreoPep’s robust capacity to identify functional residues in target-specific peptides. Multiple sequence alignment (Fig. 5d) and variation frequency analysis (Fig. 5e) identified four conserved non-cysteine residues (P6, R7, W10, and R11) as essential for activity. These residues were preserved across all high-potency mutants except CP_α7_6; their mutation consistently abolished potency. Conversely, mutation-tolerant regions clustered at residues 1, 4, 5, and 9 (Fig. 5e). All active mutants included substitutions at these locations, with position D5 displaying the highest mutational flexibility, six of seven potent variants carried modifications at this site. Furthermore, comparative analysis revealed significantly greater disulfide bond variability in the ImI/α7 system compared to seven other systems (Supplementary Fig. 6). This observation challenges the traditional view that disulfide bonds are rigid and conserved structural elements essential for α-conotoxin bioactivity. However, it aligns with Tabassum et al.47 who reported that, except for ImI, disulfide-deficient variants of Vc1.1 and AuIB showed diminished or abolished nAChR inhibitory potency. The strong concordance between CreoPep’s predictions and experimental results underscores its exceptional ability to identify critical functional residues, distinguish nonessential positions, accurately model mutation tolerance, and guide rational peptide design through targeted modification of variable regions while preserving core functional motifs.
Structural basis of high-potency mutant binding
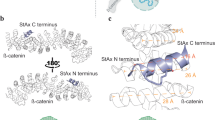
Electrophysiological characterization identified CP_α7_1 and CP_α7_6 as the most potent hα7 nAChR inhibitors among the CreoPep-designed variants. AlphaFold3-predicted structures, further refined by MD simulations, revealed that both mutants employ distinct strategies to stabilize the α7(+)α7(−) interface. CP_α7_1 preserves ImI’s disulfide bonds (Cys2–Cys8, Cys3–Cys14) and core binding motifs, whereas CP_α7_6 achieves comparable potency via a completely reengineered interaction network, despite lacking disulfide connectivity (Fig. 5d). In the CP_α7_1/hα7 nAChR complex, key stabilizing interactions include hydrogen bonds (I1-Y188, Y4-E162, D5-Y188, A9-Q117), hydrophobic packing (P6 with Y93/W149/W55/L119), and salt bridges (R7-D197, R11-E193) (Fig. 6a–e). In contrast, CP_α7_6-ha7 complex features nine hydrogen bonds (spanning T2-S113 to A12-K145), hydrophobic contacts involving Y5-L109, P6-Y188/Y195, L8-W149, and W11-L169, as well as a critical salt-bridge (R7-D164) supported by additional hydrogen bonds (R7-S36/Q57) (Fig. 6f–j). Despite substantial sequence divergence, both variants achieve interfacial stabilization through distinct interaction networks. This demonstrates CreoPep’s capability to uncover both conservative and non-conservative binding modes without sacrificing potency, highlighting its potential for rational peptide engineering.
a Visualization of the overall binding mode between CP_α7_1 and the hα7 nAChR. b–e Details interactions individual amino acid residues of CP_α7_1 with the hα7 nAChR. f Visualization of the binding mode between CP_α7_6 and the hα7 nAChR. g–j Detailed interaction of individual amino acid residues of CP_α7_6 with the hα7 nAChR binding pocket.
Experimental validation of conotoxin GID mutants generated via CreoPep
To further validate the generalizability of CreoPep across different systems, we conducted biological activity validation on the GID/α4β2 nAChR system. It is a greater challenge because the majority of mutants of GID that have been synthesized and tested are inactive, indicating the highly specific interaction between GID and the α4β2 receptor48,49,50. Based on experience gained from the ImI/α7 nAChR system, we retained key conserved residues predicted by CreoPep (I1, C5, C6, P9, C11, R12, N14, C19) during the screening of GID mutants, while keeping the rest of the process unchanged (Fig. 7a).
a Graphical representation of multiple sequence alignment of 1000 CreoPep-generated mutants, showing the frequency of amino acid variations at each position relative to the GID sequence. b Bar graph showing the inhibitory effects of 10 candidate conotoxins (at 1 and 10 µM) on ACh-evoked peak current amplitudes mediated by human (h) α4β2 nAChRs. Whole-cell currents were evoked by 3 µM ACh (mean ± SD, n = 4–8). The dashed line indicates 50% inhibition of the peak current amplitude. c Multiple sequence alignment of GID with the 10 candidate conotoxins. Among them, colors reflect the physicochemical properties of amino acids, using the Zappo protein color scheme: pink for aliphatic/hydrophobic, orange for aromatic, blue for positive, red for negative, green for hydrophilic, magenta for conformationally special, yellow for cysteine, and gray for positions without mutations. d Concentration–response relationships for CP-α4β2-2 inhibition of ACh-evoked currents at (h) α4β2 nAChRs. Current amplitudes (mean ± SD; n = 6) were normalized to the response elicited by 100 µM ACh alone. e Bar graph showing the inhibitory effects of CP-α4β2-2 (at 1 and 10 µM) on ACh-evoked peak current amplitudes mediated by human (h) α7 and α3β2 nAChRs. f Constructed complex models of CP-α4β2-2 bound to (h) α4β2 nAChR and their binding modes.
Functional characterization of the top 10 candidate peptides from 1000 CreoPep-generated GID mutants revealed a 30% hit rate for inhibiting human α4β2 nAChR at a concentration of 10 µM. Among these, CP_α4β2_2 exhibited the strongest inhibitory activity (>50% blockade) (Fig. 7b, c). Further quantitative concentration–response analysis confirmed that this mutant achieved a IC50 value of 3.5 µM (95% CI; 3.3–3.8) (Fig. 7d), which is comparable to data reported by Abba E. Leffler et al.48. The potency of CP_α4β2_2 is slightly lower than that of the wild-type GID, which contains unnatural amino acid modifications48 (Supplementary Table 20). These results show that CreoPep performs well even in this highly specific system.
Additionally, CreoPep has broken through the limitations of traditional peptide optimization, enabling the exploration of “riskier” mutation spaces and the design of highly diverse functional variants. For instance, CP_α4β2_1 maintained activity with an A10R mutation, whereas a similar mutation (A10Q) reported previously abolished activity. Such functional preservation may arise from compensatory mutations at multiple sites (up to six) (Fig. 7c). However, an excessive number of mutations and significant changes in physicochemical properties can also lead to a higher probability of disruption of the active conformation (loss of activity in CP_α4β2_3, CP_α4β2_4, CP_α4β2_9, and CP_α4β2_10). Notably, all three active mutants showed amino acid substitutions in the N-terminal tail, a region previously shown to be critical for α4β2 activity48. However, CP_α4β2_2 retained inhibitory activity comparable to wild-type GID despite mutations at two positions in this region. Subtype selectivity analysis indicated CP_α4β2_2 exhibited strong inhibition of both α7 (95.5% blockade) and α3β2 (92.96% blockade) nAChR subtypes at 1 µM (n = 4–8; Fig. 7e), showing functional selectivity similar to that of GID. Structural analysis revealed that in the D → R mutation, the R3 formed a new electrostatic interaction network with D170, successfully compensating for the effects of the polarity reversal. Meanwhile, the introduction of P at position 4 may disrupt the original secondary structure, thereby making the conformation of positions 1–3 more flexible and increasing the likelihood of electrostatic interactions with surrounding amino acids (Fig. 7f). This represents a “riskier” attempt that would be challenging to achieve through manual design, demonstrating potential of CreoPep in exploring “riskier” mutation spaces.
Discussion
Target-specific conotoxin peptides offers substantial promise for therapeutic development due to their high affinity and specificity for molecular targets. However, the clinical translation of natural conotoxins is often hindered by suboptimal potency, limited selectivity, and poor metabolic stability. Traditional optimization strategies, such as saturation mutagenesis and targeted point mutation, can partially improve the peptide activity, but these methods are inefficient, labor-intensive, and heavily reliant on domain expertise. Accordingly, there is a critical need for more efficient strategies capable of generating potent conotoxin variants with high sequence diversity.
In this study, we introduce CreoPep, a deep learning-based conditional generation framework that leverages a masked language model with a progressive masking (PM) strategy for efficient design and optimization of conotoxin peptides. The key innovation of CreoPep lies in its ability to generate diverse peptides guided by target and potency labels, while maintaining sequence diversity and biological functionality through a temperature-controlled multinomial sampling approach. In addition, CreoPep incorporates an iterative data augmentation pipeline using FoldX-based binding energy screening, which expands the training dataset, enhances model performance, and provides mechanistic insights—particularly under few-shot learning settings. Notably, CreoPep functions as an efficient computational alternative to traditional alanine scanning, enabling the identification of critical functional residues. Electrophysiological validation further confirmed the model’s effectiveness: 7 out of 13 designed conotoxin candidates showed relatively high potency against the hα7 nAChR. Among these, variants CP_α7_1 and CP_α7_6 exhibited sub-micromolar inhibitory activity, with IC50 values of 405.1 and 504.6 nM, respectively. Additionally, 3 out of 10 designed variants demonstrated inhibitory activity against the hα4β2 nAChR. These findings highlight CreoPep’s capacity to generate potent and functionally relevant conotoxin mutants with therapeutic potential.
Despite its advantages, CreoPep also has limitations. Among the ImI-derived mutants, only a single variant showed marginal improvement in potency compared to the wild-type peptide, underscoring the inherent difficulty of surpassing the potency of short, naturally optimized conotoxins. Incorporation of nonnatural amino acids may be necessary to enhance such scaffolds. Because CreoPep is based on the BERT architecture, its vocabulary can easily be expanded to include user-defined tokens representing non-canonical residues. Beyond potency and receptor subtype, additional constraints such as toxicity, stability, or pharmacokinetic properties could also be incorporated through appropriate predictive scoring models.
The potency class framework used in this study may make also limit CreoPep’s ability to identify the mutants with highly favorable ΔΔG values. Employing more stringent thresholds during data augmentation (e.g., ΔΔG ≤ −5 kcal mol−1) could improve discrimination but would require generating substantially larger peptide libraries, increasing computational costs. Future work could therefore focus on reformulating CreoPep as a multi-class or regression-based model to better capture graded potency and facilitate the identification of top-performing mutants without prohibitive computational overhead. Finally, the data augmentation process in CreoPep relies on complex structure prediction and binding affinity estimation steps. The selection of tools in this pipeline, such as replacing FoldX binding energy scoring with alternative metrics like the ipTM score from AlphaFold3 or energy scoring from Rosetta, could further improve the quality of generated peptides and the success rate of downstream validation.
In conclusion, CreoPep represents a robust, deep learning-driven platform for the rational design of target-specific conotoxin therapeutics. Future developments could focus on three main directions: (1) incorporating nonnatural amino acids to expand chemical space, (2) generating multi-target peptides to address complex disease networks, and (3) integrating patient-specific biological data to enable personalized peptide therapeutics. These developments would further strengthen CreoPep’s potential to drive the next-generation peptide drug discovery.
Methods
Data collection and processing
We collected 2088 conotoxins from the ConoServer database8, with each entry containing target information and potency data (Supplementary Fig. 1). Each training sample required a subtype label, a potency label, a wild-type conotoxin, and up to three auxiliary conotoxins. The auxiliary conotoxins are randomly selected from a set which has the same subtype and potency as the wild-type conotoxin. The initial training set was obtained from Conoserver and exhibited data imbalance (Supplementary Fig. 1a, b). We performed data augmentation across eight complex systems, assigning pseudo-labels based on binding energy calculated by FoldX (Supplementary Fig. 1c, d). The expanded datasets from each round contained 4471, 7330, 10,499, 14,072, and 18,355 peptide sequences, respectively.
Model implementations
The CreoPep model architecture integrates four core components: the encoder block (for labels and wild-type conotoxin sequence representation learning), the MLP block (for processing auxiliary conotoxin features), the CNN block (for dimensionality reduction after feature fusion), and the decoder block (for final prediction). CreoPep first extracts wild-type conotoxin sequence, as well as subtype and potency labels, using features through the encoder block. This block is based on the pre-trained protein language model ProtBert. The BERT encoder of ProtBert is frozen, and an additional trainable BERT encoder without an embedding layer is added, enabling the model to adapt to specific tasks while leveraging pre-trained knowledge. The MLP block integrates an embedding layer and a Xavier-initialized multilayer perceptron (linear layer + leakyReLU + linear layer) to extract auxiliary conotoxin features. Subsequently, these features are concatenated and then reduced in dimension by a 1 × 1 convolutional CNN block. Finally, the fused features are input into the decoder block to generate the final prediction. The training protocols are listed in Supplementary Table 21 and the pseudocode of the core training process is shown in Supplementary Table 22.
Loss function
We proposed a masked cross-entropy loss function to handle multi-task token prediction. For a sequence with n positions, the raw cross-entropy loss \({{{\mathcal{L}}}}_{{{\rm{CE}}}}({y}_{i},{\hat{y}}_{i})\) is computed at each position i. We then apply three binary mask matrices - the \([{\mbox{MASK}}]\) token position mask \({\mathbb{M}}_{{\rm{mask}}} \in \{{\mathrm{0,1}}\}^{n}\), label mask \({{\mathbb{M}}}_{{{\rm{label}}}}\in \{{\mathrm{0,1}}\}^{m}\), and sequence mask \({{\mathbb{M}}}_{{{\rm{seq}}}}\in \{{\mathrm{0,1}}\}^{p}\)—to compute task-specific losses:
Where ‘\(\circ\)’ denotes element-wise multiplication (Hadamard product). The final composite loss is given by the weighted sum:
Here, \(\alpha ,\beta ,\gamma \in {{\mathbb{R}}}^{+}\) are tunable hyperparameters that control each task’s contribution to the overall optimization objective, with α = 1, β = 1/2, and γ = 1/3. This loss function ensures that the model prioritizes position-level correctness while also incorporating label and sequence-level constraints with reduced weighting.
Mask ratio setting
To evaluate the effect of different masking ratios on the generative model, we tested values ranging from 10% to 100% in 10% increments. All sequences were first padded to a fixed length of L = 54 tokens. For each masking ratio \(r\in \{{\mathrm{0.1,0.2}},\ldots ,1.0\}\), we computed the number of masked positions as:
where t is computed using rounding to the nearest integer and t = 54 represents complete masking (100%) and t = 27 corresponds to the 50% masking condition. Under a fixed 100-epoch training regime, all model variants achieved convergence as shown in Fig. S7, with asterisks marking their respective minimum loss values. The optimal performance was observed at r = 0.5 (50% masking, t = 27 positions), yielding the lowest loss of 0.729. This 50% masking configuration was consequently adopted for final model training.
Sampling strategy
In the data augmentation pipeline, diversity in the generated sequences is ensured by introducing multinomial sampling during the peptide generation phase. Meanwhile, a temperature factor τ is incorporated into the sampling process to control the degree of diversity in the generated peptides. Specifically, a higher τ value results in a smoother probability distribution, thereby increasing the diversity of the sampled peptides. The calculation formula is as follows:
Here, \({z}_{i}\) denotes the logit (raw score) for the i-th class, while \({z}_{j}\) represents the logit for all \(V\) classes, with \(j\) ranging from 1 to \(V\). The output \({\mbox{P}}({y}_{i})\) is the resulting probability distribution. Subsequently, a class is sampled from this distribution using multinomial sampling:
Evaluation metrics for FoldX
We evaluated FoldX’s performance using four standard metrics: accuracy, sensitivity, specificity, and MCC (Matthews correlation coefficient), with their calculation formulas shown as follows.
where TP, TN, FP and FN denote the number of true positives, true negatives, false positives and false negatives, respectively. The terms of positive (P), negative (N), true (T) and false (F) were defined as follows: (1) positive: IC50 decreased or no change in experimental data; (2) negative: IC50 increased in experimental data; (3) true: predicted ∆∆G class is consistent with experimental data; (4) false: predicted ∆∆G class is not consistent with experimental data. The complex structure that achieved optimal performance across all four evaluation metrics was selected as the final system structure. Although α-conotoxin GID contains nonnatural amino acids that are incompatible with FoldX’s input format, we included it in subsequent data augmentation tasks due to the importance of the GID/α4β2 system. For data augmentation, we used GID[(GLA)4E, O16P] as the representative complex structure of the GID/α4β2 system, but we did not assess FoldX’s performance on this particular complex.
Binding energy calculations
We utilized official Python bindings of FoldX (pyfoldx) to calculate peptide-receptor binding free energies. Using the wild-type complex structure as a template, we first optimize the structure using FoldX’s repair function for energy minimization. We then calculate the binding free energy of wild-type peptide (ΔGwild-type) using the getInterfaceEnergy function. For each mutant, the following steps were executed: (i) mutation site identification through sequence alignment; (ii) stepwise introduction of amino acid substitutions; (iii) local energy optimization following each mutation; (iv) global relaxation of the fully mutated complex. The binding free energy of each mutant (ΔGmutant) was then computed with getInterfaceEnergy, and the binding energy difference (ΔΔG = ΔGmutant- ΔGwild-type) was determined. Here, ∆Gmutant refers to the binding energy of the mutant complex itself, while ∆Gwild-type refers to that of the wild-type complex. During dataset expansion, we established robust classification criteria based on prior studies. Following Wu et al.40 definition, in which ΔΔG = ± 1 kcal mol−1 indicates comparable binding affinity, we adopted a more stringent threshold (ΔΔG ≤ 0.5 kcal mol−1) to identify favorable mutations. These were labeled as high-potency mutants (positive samples) in our training dataset. For low-potency mutants (negative samples), we applied a conservative cutoff (ΔGmutant > 0 kcal mol−1) to reduce the likelihood of false negatives and enhance dataset reliability.
Molecular dynamics simulations of mutants CP_α7_1 and CP_α7_6 with hα7 nAChR
The initial complex structures of the CP_α7_1 and CP_α7_6 mutants bound to the hα7 nAChR were predicted using AlphaFold3. Based on this structure, the C-terminal amidation modifications of the mutants were performed using PyMOL. MD simulations were performed with the AMBER 22 software package using the ff19SB force field51. Each complex was solvated in a truncated octahedral periodic box. The system was initially heated from 50 to 300 K over 100 ps in the NVT ensemble, with the solute restrained by a harmonic force of 5 kcal mol−1 Å−2. Subsequently, the simulation transitioned to the NPT ensemble, during which the harmonic restraints were gradually reduced from 5 to 0 kcal mol−1 Å−2 over another 100 ps. For the production phase, a 50 ns MD simulation was carried out with the temperature and pressure maintained at 300 K and 1 bar, respectively. After the MD simulation, the MD trajectories were analyzed using VMD (http://www.ks.uiuc.edu/), and root mean square deviation (RMSD) values were calculated to assess structural stability.
Peptide synthesis and purification
The linear peptide was synthesized on a 0.1 mmol scale using solid-phase peptide synthesis (SPPS) with Rink-Amide resin. After swelling and deprotecting the resin using 20% piperidine for 2 h, Fmoc-protected amino acids were coupled using HCTU (4.0 equiv) and DIPEA (8.0 equiv) for 1 h, followed by deprotection with 20% piperidine for 30 min. Between each step, the resin was washed three times with DMF and twice with DCM. For peptides containing two disulfide bonds, regioselective oxidation was achieved by incorporating Fmoc-Cys(Acm)-OH at the CysI and CysIII positions and Fmoc-Cys(Trt)-OH at CysII and CysIV positions. Following peptide chain assembly and terminal Fmoc removal, the peptide was cleaved from the resin using TFA/TIPS/H2O (90:5:5) for 3 h, concentrated, precipitated in cold ether, and analyzed by LC-MS. The first disulfide bond was formed by slowly adding 0.8 equiv of 2,2’-dithiodipyridine in 10 mL of methanol to the crude peptide solution over 30 min with constant stirring. The product was purified by preparative RP-HPLC. Subsequently, iodine (5 equiv) was used to simultaneously remove the Acm protecting groups and facilitate disulfide bond formation between CysI and CysIII. The reaction was quenched with ascorbic acid until the solution became colorless. All crude peptides were purified by semi-preparative RP-HPLC on a C18 column using a gradient of 0.05% TFA in H2O/MeCN (flow rate: 6 mL min-1) with the following gradient: 0 min, 100% solvent A; 5 min, 85% solvent A; 20 min, 70% solvent A; 40 min, 60% solvent A solvent A: H2O/MeCN (90:10, v/v); solvent B: H2O/MeCN (10:90, v/v). The purity and molecular weight of the final products were confirmed by analytical RP-HPLC and LC/MS, respectively. All peptides had a purity of greater than 95% (Supplementary Figs. 8–30).
Circular dichroism (CD)
At room temperature in a nitrogen atmosphere, the CD spectrum was measured using a Jasco J-810 spectropolarimeter with wavelengths between 300 and 185 nm, with a 1.0 mm path length cell, 1.0 nm bandwidth and a response time of 2 s, averaging three scans. All peptides were dissolved in water at a concentration of 0.15 mg/mL (Supplementary Figs. 31 and 32).
Xenopus laevis oocyte preparation and microinjection
All procedures were approved by the Animal Ethics Committees of the Victor Chang Cardiac Research Institute, Sydney, and the University of Wollongong (project no. AE 20/17). Human α7 nAChR clone, provided by Prof. Jon Lindstrom (University of Pennsylvania), and plasmid constructs of human α4 and β2 nAChR subunits were linearized, and cRNA was transcribed in vitro using the SP6/T7 mMessage mMachine kit (Ambion). Stage V–VI oocytes (1200–1300 μm) were harvested from 5-year-old female Xenopus laevis (Nasco), anesthetized with 1.7 mg mL−1 ethyl 3-aminobenzoate methanesulfonate (pH 7.4), defolliculated with 1.5 mg mL−1 collagenase Type II (Worthington) in OR-2 solution (82.5 NaCl, 2 KCl, 1 MgCl₂, 5 HEPES, pH 7.4), and microinjected with 10 ng of human α7 or α4β2 cRNAs (verified by spectrophotometry and gel electrophoresis) using Drummond glass pipettes. Injected oocytes were incubated at 18 °C in ND96 solution (96 NaCl, 2 KCl, 1 CaCl₂, 1 MgCl₂, 5 HEPES, pH 7.4) supplemented with 5% FBS, 50 mg L−1 gentamicin, and 10,000 U mL−1 penicillin-streptomycin (Gibco).
Oocyte two-electrode voltage clamp recording and data analysis
Electrophysiological recordings were performed 2–5 days post-injection using a GeneClamp 500B amplifier and pClamp9 software (Molecular Devices), with a holding potential of −80 mV. Recording electrodes (0.3–1 MΩ resistance) were pulled from GC150T-7.5 borosilicate glass (Harvard Apparatus) and filled with 3 M KCl. Perfusion was carried out at 2 mL/min (Legato 270 syringe pump; KD Scientific) in an OPC-1 chamber (<20 μL; Automate Scientific) using ND96 solution. After rinsing, oocytes were exposed to three applications of 3 or 100 μM ACh (the EC₅₀ for human α4β2 and α7 nAChRs, respectively), each followed by a 3 min washout with ND96. Peptide testing involved 5 min static incubation (perfusion off), followed by co-application with ACh during perfusion. All peptides were dissolved in ND96 supplemented with 0.1% BSA. Current amplitudes (ACh alone vs. ACh + peptide) were analyzed using Clampfit 10.7 (Molecular Devices), with peptide activity expressed as the ratio of evoked currents. Data from 4 to 8 oocytes are presented as mean ± SD. Concentration–response curves were generated by nonlinear regression to determine IC₅₀ values (95% CI), and statistical significance was assessed using ANOVA (p < 0.05; GraphPad Prism 9).
Data availability
The model weights of CreoPep are available on Zenodo at this link (https://zenodo.org/records/15192592). The training sets, along with the generated peptides and their corresponding ΔΔG values from various data augmentation stages, can be accessed at this GitHub repository (https://github.com/gc-js/CreoPep/tree/main/data). Additionally, the source files and codes used for plotting are available at https://github.com/gc-js/CreoPep/tree/main/plot.
Code availability
The code is available in our GitHub project at https://github.com/gc-js/CreoPep. (https://zenodo.org/records/18034331).
References
Muttenthaler, M., King, G. F., Adams, D. J. & Alewood, P. F. Trends in peptide drug discovery. Nat. Rev. Drug Discov. 20, 309–325 (2021).
Wang, L. et al. Therapeutic peptides: current applications and future directions. Signal Transduct Target Ther. 7, 48 (2022).
Jin, A.-H. et al. Conotoxins: chemistry and biology. Chem. Rev. 119, 11510–11549 (2019).
Lewis, R. J., Dutertre, S., Vetter, I. & Christie, M. J. Conus venom peptide pharmacology. Pharmacol. Rev. 64, 259–298 (2012).
Terlau, H. & Olivera, B. M. Conus venoms: a rich source of novel ion channel-targeted peptides. Physiol. Rev. 84, 41–68 (2004).
Pei, S. et al. Conotoxins targeting voltage-gated sodium ion channels. Pharmacol. Rev. 76, 828–845 (2024).
Akondi, K. B. et al. Discovery, synthesis, and structure–activity relationships of conotoxins. Chem. Rev. 114, 5815–5847 (2014).
Kaas, Q., Yu, R., Jin, A.-H., Dutertre, S. & Craik, D. J. ConoServer: updated content, knowledge, and discovery tools in the conopeptide database. Nucleic Acids Res. 40, D325–D330 (2012).
Gao, S., Yao, X. & Yan, N. Structure of human Cav2.2 channel blocked by the painkiller ziconotide. Nature 596, 143–147 (2021).
Noviello, C. M. et al. Structure and gating mechanism of the a7 nicotinic acetylcholine receptor. Cell 184, 2121–2134 (2021).
Pan, X. et al. Molecular basis for pore blockade of human Na+ channel Nav1.2 by the μ-conotoxin KIIIA. Science 363, 1309–1313 (2019).
Fosgerau, K. & Hoffmann, T. Peptide therapeutics: current status and future directions. Drug Discov. Today 20, 122–128 (2015).
Olivera, B. M. et al. Neuronal calcium channel antagonists. Discrimination between calcium channel subtypes using omega-conotoxin from Conus magus venom. Biochemistry 26, 2086–2090 (1987).
Miljanich, G. P. Ziconotide: neuronal calcium channel blocker for treating severe chronic pain. Curr. Med. Chem. 11, 3029–3040 (2004).
Kurtzhals, P., Østergaard, S., Nishimura, E. & Kjeldsen, T. Derivatization with fatty acids in peptide and protein drug discovery. Nat. Rev. Drug Discov. 22, 59–80 (2023).
Xiao, W. et al. Advance in peptide-based drug development: delivery platforms, therapeutics and vaccines. Signal Transduct Target Ther. 10, 74 (2025).
Ji, X., Nielsen, A. L. & Heinis, C. Cyclic peptides for drug development. Angew. Chem. Int. Ed. 63, e202308251 (2024).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Delgado, J., Radusky, L. G., Cianferoni, D. & Serrano, L. FoldX 5.0: working with RNA, small molecules and a new graphical interface. Bioinformatics 35, 4168–4169 (2019).
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta FlexPepDock web server—high resolution modeling of peptide–protein interactions. Nucleic Acids Res. 39, W249–W253 (2011).
Zhou, P., Jin, B., Li, H. & Huang, S.-Y. HPEPDOCK: a web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res. 46, W443–W450 (2018).
Watson, J. L. et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023).
Dauparas, J. et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Vázquez Torres, S. et al. De novo designed proteins neutralize lethal snake venom toxins. Nature 639, 225–231 (2025).
Liu, H. et al. De novo design of self-assembling peptides with antimicrobial activity guided by deep learning. Nat. Mater. 24, 1295–1306 (2025).
Cao, Q. et al. Designing antimicrobial peptides using deep learning and molecular dynamic simulations. Brief. Bioinform. 24, bbad058 (2023).
Zhao, Y. et al. A unified deep framework for peptide–major histocompatibility complex–T cell receptor binding prediction. Nat. Mach. Intell. 7, 650–660 (2025).
Dadonaite, B. et al. Spike deep mutational scanning helps predict success of SARS-CoV-2 clades. Nature 631, 617–626 (2024).
Ferruz, N. & Höcker, B. Controllable protein design with language models. Nat. Mach. Intell. 4, 521–532 (2022).
Gao, Y. et al. Pan-peptide meta Learning for T-cell receptor–antigen binding recognition. Nat. Mach. Intell. 5, 236–249 (2023).
Huang, J. et al. Identification of potent antimicrobial peptides via a machine-learning pipeline that mines the entire space of peptide sequences. Nat. Biomed. Eng. 7, 797–810 (2023).
Vázquez Torres, S. et al. De novo design of high-affinity binders of bioactive helical peptides. Nature 626, 435–442 (2024).
Lei, Y. et al. A deep-learning framework for multi-level peptide–protein interaction prediction. Nat. Commun. 12, 5465 (2021).
Deng, Q. et al. RLpMIEC: high-affinity peptide generation targeting major histocompatibility complex-I guided and interpreted by interaction spectrum-navigated reinforcement learning. J. Chem. Inf. Model. 64, 6432–6449 (2024).
Chen, S. et al. Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations. Nat. Commun. 15, 1611 (2024).
Ho, J., Jain, A. & Abbeel, P. Denoising diffusion probabilistic models. Advances in neural information processing systems 33, 6840–6851 (2020).
Yu, R., Craik, D. J. & Kaas, Q. Blockade of neuronal α7-nAChR by α-conotoxin ImI explained by computational scanning and energy calculations. PLoS Comput. Biol. 7, e1002011 (2011).
Elnaggar, A. et al. ProtTrans: Towards cracking the language of life’s code through self-supervised learning. IEEE Trans. Pattern Anal. Mach. Intell. 44, 7112–7127 (2021).
Wu, X. et al. Computational design of α-conotoxins to target specific nicotinic acetylcholine receptor subtypes. Chem. A Eur. J. 30, e202302909 (2024).
Wu, R. 2022. High-resolution de novo structure prediction from primary sequence. Preprint at bioRxiv https://doi.org/10.1101/2022.07.21.500999 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Xu, J. & Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5? Bioinformatics 26, 889–895 (2010).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://doi.org/10.48550/arXiv.1802.03426 (2020).
Tabassum, N. et al. Role of CysI –CysIII disulfide bond on the structure and activity of α-conotoxins at human neuronal nicotinic acetylcholine receptors. ACS Omega 2, 4621–4631 (2017).
Millard, E. L. et al. Inhibition of neuronal nicotinic acetylcholine receptor subtypes by α-conotoxin GID and analogues. J. Biol. Chem. 284, 4944–4951 (2009).
Leffler, A. E. et al. Discovery of peptide ligands through docking and virtual screening at nicotinic acetylcholine receptor homology models. Proc. Natl. Acad. Sci. USA 114, E8100–E8109 (2017).
Nicke, A. et al. Isolation, structure, and activity of GID, a novel α4/7-conotoxin with an extended N-terminal sequence. J. Biol. Chem. 278, 3137–3144 (2003).
Case, D. A. et al. AmberTools. J. Chem. Inf. Model. 63, 6183–6191 (2023).
Acknowledgements
We gratefully acknowledge our laboratory members for their insightful discussions. This work was supported by the National Natural Science Foundation of China (Grant No. U25A20178, 82122064, 82473832, 62373172), Fundamental Research Funds for the Central Universities (202441004), Qingdao Science and Technology Demonstration Project (25-1-1-sfgc-2-nsh), Jinan Innovation Team Project (202333022), Qingdao National Laboratory for Marine Science and Technology (2022QNLM030003-4), Shandong Supporting Funds for Talents (2022GJJLJRC02-046), and an Australian Research Council Discovery Project Grant (DP150103990 to DJA).
Author information
Authors and Affiliations
Contributions
C.G. conducted data collection, deep learning design, peptide synthesis, data analysis, and wrote the original draft. H.-S.T. conducted electrophysiology experiments, data curation. L.L. conducted peptide synthesis, data analysis, and wrote the original draft. Z.Z. conducted deep learning design, peptide synthesis. Z.H. conducted data collection and peptide synthesis. B.A. conducted electrophysiology experiments. Y.W. conducted data collection. T.J., W.C., and S.C. obtained the resources for the study. D.J.A. revised and edited the manuscript. R.Y. conceptualized the study, acquired the resources, and wrote the original draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Yu Kang and the other anonymous reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ge, C., Tae, HS., Lu, L. et al. A deep learning framework (CreoPep) for target-specific design and optimization of conotoxin peptides. Commun Chem 9, 82 (2026). https://doi.org/10.1038/s42004-025-01885-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01885-5