Abstract
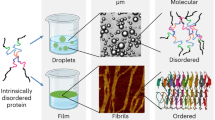
Intrinsically disordered protein regions (IDRs) play a key role in the formation of biomolecular condensates, a ubiquitous mode of cellular compartmentalization, but the underlying microscopic details remain unclear. Here, microsecond-level molecular dynamics simulations and fractal formalism are employed to study at atomistic resolution a model dense phase composed of 24 copies of a C-terminal 73-residue arginine- and glycine-rich IDR (RGG3) of fused in sarcoma (FUS) protein in the absence of RNA. RGG3 displays a highly dynamic behavior in the dense phase with only a small configurational entropy loss and a minor slowdown in diffusion as compared to the dilute phase. Despite rapid mixing, short contact residence times and structurally heterogenous binding interfaces in the dense phase, RGG3 exhibits a distinct dynamic binding mode, with statistically defined interaction motifs and a robust multi-scale topology of self-associated protein clusters. An analysis of bound water suggests that solvent entropy may significantly contribute to the thermodynamics of condensate formation. Our results demonstrate how a well-defined organization of the disordered protein dense phase across scales emerges from highly heterogenous, transient interactions at the molecular level.
Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. A combined Supplementary Information PDF includes Supplementary Figs. 1–3 and Supplementary Table 1. Supplementary Movie 1, initial and final conformations from MD simulations (Supplementary Data 1) are available as Supplementary Data.
References
Wright, P. E. & Dyson, H. J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 16, 18–29 (2015).
Babu, M. M. The contribution of intrinsically disordered regions to protein function, cellular complexity, and human disease. Biochem. Soc. Trans. 44, 1185–1200 (2016).
Tsang, B., Pritišanac, I., Scherer, S. W., Moses, A. M. & Forman-Kay, J. D. Phase separation as a missing mechanism for interpretation of disease mutations. Cell 183, 1742–1756 (2020).
McAffee, D. B. et al. Discrete LAT condensates encode antigen information from single pMHC:TCR binding events. Nat. Commun. 13, 7446 (2022).
Bondos, S. E., Dunker, A. K. & Uversky, V. N. Intrinsically disordered proteins play diverse roles in cell signaling. Cell Commun. Signal. 20, 20 (2022).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Boeynaems, S. et al. Phase separation in biology and disease; current perspectives and open questions. J. Mol. Biol. 435, 167971 (2023).
Pappu, R. V., Cohen, S. R., Dar, F., Farag, M. & Kar, M. Phase transitions of associative biomacromolecules. Chem. Rev. 123, 8945–8987 (2023).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Lafontaine, D. L. J., Riback, J. A., Bascetin, R. & Brangwynne, C. P. The nucleolus as a multiphase liquid condensate. Nat. Rev. Mol. Cell Biol. 22, 165–182 (2021).
Zhu, L. & Brangwynne, C. P. Nuclear bodies: the emerging biophysics of nucleoplasmic phases. Curr. Opin. Cell Biol. 34, 23–30 (2015).
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal. 14, 1–1 (2016).
Lyon, A. S., Peeples, W. B. & Rosen, M. K. A framework for understanding the functions of biomolecular condensates across scales. Nat. Rev. Mol. Cell Biol. 22, 215–235 (2021).
Uversky, V. N. Recent developments in the field of intrinsically disordered proteins: intrinsic disorder-based emergence in cellular biology in light of the physiological and pathological liquid-liquid phase transitions. Annu. Rev. Biophys. 50, 135–156 (2021).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Holehouse, A. S. & Kragelund, B. B. The molecular basis for cellular function of intrinsically disordered protein regions. Nat. Rev. Mol. Cell Biol. 25, 187–211 (2024).
Nomura, T. et al. Intranuclear aggregation of mutant FUS/TLS as a molecular pathomechanism of amyotrophic lateral sclerosis. J. Biol. Chem. 289, 1192–1202 (2014).
Mészáros, B., Erdős, G. & Dosztányi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 46, W329–W337 (2018).
Emenecker, R. J., Griffith, D. & Holehouse, A. S. Metapredict: a fast, accurate, and easy-to-use predictor of consensus disorder and structure. Biophys. J. 120, 4312–4319 (2021).
Bugge, K. et al. Interactions by disorder–a matter of context. Front. Mol. Biosci. 7, https://doi.org/10.3389/fmolb.2020.00110 (2020).
Fuxreiter, M. Classifying the binding modes of disordered proteins. Int. J. Mol. Sci. 21, 8615–8615 (2020).
Jahn, L. R., Marquet, C., Heinzinger, M. & Rost, B. Protein embeddings predict binding residues in disordered regions. Sci. Rep. 14, 13566 (2024).
Darling, A. L. & Uversky, V. N. Intrinsic disorder and posttranslational modifications: the darker side of the biological dark matter. Front. Genet. 9, https://doi.org/10.3389/fgene.2018.00158 (2018).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Vernon, R. M. et al. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. eLife 7, https://doi.org/10.7554/eLife.31486 (2018).
Saar, K. L. et al. Learning the molecular grammar of protein condensates from sequence determinants and embeddings. Proc. Natl. Acad. Sci. USA 118, e2019053118 (2021).
Flock, T., Weatheritt, R. J., Latysheva, N. S. & Babu, M. M. Controlling entropy to tune the functions of intrinsically disordered regions. Curr. Opin. Struct. Biol. 26, 62–72 (2014).
Soranno, A. et al. Integrated view of internal friction in unfolded proteins from single-molecule FRET, contact quenching, theory, and simulations. Proc. Natl. Acad. Sci. USA 114, E1833–E1839 (2017).
Fleck, M., Polyansky, A. A. & Zagrovic, B. PARENT: a parallel software suite for the calculation of configurational entropy in biomolecular systems. J. Chem. Theory Comput. 12, 2055–2065 (2016).
Fleck, M., Polyansky, A. A. & Zagrovic, B. Self-consistent framework connecting experimental proxies of protein dynamics with configurational entropy. J. Chem. Theory Comput. 14, 3796–3810 (2018).
Polyansky, A. A., Gallego, L. D., Efremov, R. G., Köhler, A. & Zagrovic, B. Protein compactness and interaction valency define the architecture of a biomolecular condensate across scales. eLife 12, e80038 (2023).
Moses, D., Ginell, G. M., Holehouse, A. S. & Sukenik, S. Intrinsically disordered regions are poised to act as sensors of cellular chemistry. Trends Biochem. Sci. 48, 1019–1034 (2023).
Ghosh, C., Nagpal, S. & Muñoz, V. Molecular simulations integrated with experiments for probing the interaction dynamics and binding mechanisms of intrinsically disordered proteins. Curr. Opin. Struct. Biol. 84, 102756 (2024).
Piana, S., Donchev, A. G., Robustelli, P. & Shaw, D. E. Water dispersion interactions strongly influence simulated structural properties of disordered protein states. J. Phys. Chem. B 119, 5113–5123 (2015).
Piana, S., Robustelli, P., Tan, D., Chen, S. & Shaw, D. E. Development of a force field for the simulation of single-chain proteins and protein–protein complexes. J. Chem. Theory Comput. 16, 2494–2507 (2020).
Gopal, S. M. et al. Conformational preferences of an intrinsically disordered protein domain: a case study for modern force fields. J. Phys. Chem. B 125, 24–35 (2021).
Wang, W. Recent advances in atomic molecular dynamics simulation of intrinsically disordered proteins. Phys. Chem. Chem. Phys. 23, 777–784 (2021).
Kasahara, K., Terazawa, H., Takahashi, T. & Higo, J. Studies on molecular dynamics of intrinsically disordered proteins and their fuzzy complexes: a mini-review. Comput. Struct. Biotechnol. J. 17, 712–720 (2019).
Bastida, A., Zúñiga, J., Fogolari, F. & Soler, M. A. Statistical accuracy of molecular dynamics-based methods for sampling conformational ensembles of disordered proteins. Phys. Chem. Chem. Phys. https://doi.org/10.1039/D4CP02564D (2024).
Zhu, J., Salvatella, X. & Robustelli, P. Small molecules targeting the disordered transactivation domain of the androgen receptor induce the formation of collapsed helical states. Nat. Commun. 13, 6390 (2022).
Galvanetto, N. et al. Extreme dynamics in a biomolecular condensate. Nature 619, 876–883 (2023).
Rauscher, S. & Pomès, R. The liquid structure of elastin. eLife 6, e26526 (2017).
Paloni, M., Bailly, R., Ciandrini, L. & Barducci, A. Unraveling molecular interactions in liquid-liquid phase separation of disordered proteins by atomistic simulations. J. Phys. Chem. B 124, 9009–9016 (2020).
Flores-Solis, D. et al. Driving forces behind phase separation of the carboxy-terminal domain of RNA polymerase II. Nat. Commun. 14, 5979 (2023).
Dignon, G. L., Zheng, W., Best, R. B., Kim, Y. C. & Mittal, J. Relation between single-molecule properties and phase behavior of intrinsically disordered proteins. Proc. Natl. Acad. Sci. USA 115, 9929–9934 (2018).
Chou, H. Y. & Aksimentiev, A. Single-protein collapse determines phase equilibria of a biological condensate. J. Phys. Chem. Lett. 11, 4923–4929 (2020).
Bremer, A. et al. Deciphering how naturally occurring sequence features impact the phase behaviours of disordered prion-like domains. Nat. Chem. 14, 196–207 (2022).
Lotthammer, J. M., Ginell, G. M., Griffith, D., Emenecker, R. J. & Holehouse, A. S. Direct prediction of intrinsically disordered protein conformational properties from sequence. Nat. Methods 21, 465–476 (2024).
Milles, S. & Lemke, E. A. Mapping multivalency and differential affinities within large intrinsically disordered protein complexes with segmental motion analysis. Angew. Chem. Int. Ed. 53, 7364–7367 (2014).
Fung, H. Y. J., Birol, M. & Rhoades, E. IDPs in macromolecular complexes: the roles of multivalent interactions in diverse assemblies. Curr. Opin. Struct. Biol. 49, 36–43 (2018).
Martin, E. W. et al. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science 367, 694–699 (2020).
Sipko, E. L., Chappell, G. F. & Berlow, R. B. Multivalency emerges as a common feature of intrinsically disordered protein interactions. Curr. Opin. Struct. Biol. 84, 102742 (2024).
De La Cruz, N. et al. Disorder-mediated interactions target proteins to specific condensates. Mol. Cell https://doi.org/10.1016/j.molcel.2024.08.017 (2024).
Saar, K. L. et al. Protein condensate atlas from predictive models of heteromolecular condensate composition. Nat. Commun. 15, 5418 (2024).
Alshareedah, I. et al. Sequence-specific interactions determine viscoelasticity and ageing dynamics of protein condensates. Nat. Phys. https://doi.org/10.1038/s41567-024-02558-1 (2024).
Sundaravadivelu Devarajan, D. et al. Sequence-dependent material properties of biomolecular condensates and their relation to dilute phase conformations. Nat. Commun. 15, 1912 (2024).
Kozak, F. et al. An atomistic view on the mechanism of diatom peptide-guided biomimetic silica formation. Adv. Sci. 2401239, https://doi.org/10.1002/advs.202401239 (2024).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Kar, M. et al. Solutes unmask differences in clustering versus phase separation of FET proteins. Nat. Commun. 15, 4408 (2024).
Gil-Garcia, M. et al. Local environment in biomolecular condensates modulates enzymatic activity across length scales. Nat. Commun. 15, 3322 (2024).
Crozat, A., Åman, P., Mandahl, N. & Ron, D. Fusion of CHOP to a novel RNA-binding protein in human myxoid liposarcoma. Nature 363, 640–644 (1993).
Tan, A. Y., Riley, T. R., Coady, T., Bussemaker, H. J. & Manley, J. L. TLS/FUS (translocated in liposarcoma/fused in sarcoma) regulates target gene transcription via single-stranded DNA response elements. Proc. Natl. Acad. Sci. USA 109, 6030–6035 (2012).
Ederle, H. & Dormann, D. TDP-43 and FUS en route from the nucleus to the cytoplasm. FEBS Lett. 591, 1489–1507 (2017).
Schoen, M. et al. Super-resolution microscopy reveals presynaptic localization of the ALS/FTD related protein FUS in hippocampal neurons. Front. Cell. Neurosci. 9, 496–496 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Deng, Q. et al. FUS is phosphorylated by DNA-PK and accumulates in the cytoplasm after DNA damage. J. Neurosci. 34, 7802–7813 (2014).
Banani, S. F. et al. Compositional control of phase-separated cellular bodies. Cell 166, 651–663 (2016).
Yoshizawa, T. et al. Nuclear import receptor inhibits phase separation of FUS through binding to multiple sites. Cell 173, 693–705 (2018).
Hofweber, M. et al. Phase separation of FUS is suppressed by its nuclear import receptor and arginine methylation. Cell 173, 706–719 (2018).
Qamar, S. et al. FUS phase separation is modulated by a molecular chaperone and methylation of arginine cation-π interactions. Cell 173, 720–734 (2018).
Schuster, B. S. et al. Identifying sequence perturbations to an intrinsically disordered protein that determine its phase-separation behavior. Proc. Natl. Acad. Sci. USA 117, 11421–11431 (2020).
Zheng, W. et al. Molecular details of protein condensates probed by microsecond long atomistic simulations. J. Phys. Chem. B 124, 11671–11679 (2020).
Hong, Y. et al. Hydrophobicity of arginine leads to reentrant liquid-liquid phase separation behaviors of arginine-rich proteins. Nat. Commun. 13, 7326 (2022).
Baade, I. et al. The RNA-binding protein FUS is chaperoned and imported into the nucleus by a network of import receptors. J. Biol. Chem. 296, 100659–100659 (2021).
Wake, N. et al. Expanding the molecular grammar of polar residues and arginine in FUS phase separation. Nat. Chem. Biol. 21, 1076–1088 (2025).
Burke, K. A., Janke, A. M., Rhine, C. L. & Fawzi, N. L. Residue-by-residue view of in vitro FUS granules that bind the C-terminal domain of RNA polymerase II. Mol. Cell 60, 231–241 (2015).
Murthy, A. C. et al. Molecular interactions underlying liquid−liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Brady, J. P. et al. Structural and hydrodynamic properties of an intrinsically disordered region of a germ cell-specific protein on phase separation. Proc. Natl. Acad. Sci. USA 114, E8194–E8203 (2017).
Kang, J., Lim, L. & Song, J. ATP enhances at low concentrations but dissolves at high concentrations liquid-liquid phase separation (LLPS) of ALS/FTD-causing FUS. Biochem. Biophys. Res. Commun. 504, 545–551 (2018).
Ruff, K. M., Roberts, S., Chilkoti, A. & Pappu, R. V. Advances in understanding stimulus-responsive phase behavior of intrinsically disordered protein polymers. J. Mol. Biol. 430, 4619–4635 (2018).
Monti, M. et al. catGRANULE 2.0: accurate predictions of liquid-liquid phase separating proteins at single amino acid resolution. Genome Biol. 26, 33 (2025).
von Bülow, S., Siggel, M., Linke, M. & Hummer, G. Dynamic cluster formation determines viscosity and diffusion in dense protein solutions. Proc. Natl. Acad. Sci. USA 116, 9843–9852 (2019).
Fennell, C. J., Ghousifam, N., Haseleu, J. M. & Gappa-Fahlenkamp, H. Computational signaling protein dynamics and geometric mass relations in biomolecular diffusion. J. Phys. Chem. B 122, 5599–5609 (2018).
Kamagata, K. et al. Structure-dependent recruitment and diffusion of guest proteins in liquid droplets of FUS. Sci. Rep. 12, 7101 (2022).
Ranganathan, S. & Shakhnovich, E. The physics of liquid-to-solid transitions in multi-domain protein condensates. Biophys. J. 121, 2751–2766 (2022).
Ebbinghaus, S. et al. An extended dynamical hydration shell around proteins. Proc. Natl. Acad. Sci. USA 104, 20749–20752 (2007).
Rodier, F., Bahadur, R. P., Chakrabarti, P. & Janin, J. Hydration of protein–protein interfaces. Proteins Struct. Funct. Bioinform. 60, 36–45 (2005).
Huggins, David, J. Quantifying the entropy of binding for water molecules in protein cavities by computing correlations. Biophys. J. 108, 928–936 (2015).
Kumar, K. et al. Cation–π interactions in protein–ligand binding: theory and data-mining reveal different roles for lysine and arginine. Chem. Sci. 9, 2655–2665 (2018).
Fuxreiter, M. & Tompa, P. Fuzziness: Structural Disorder in Protein Complexes (eds Fuxreiter, M. & Tompa, P.) 1–14 (Springer, 2012).
Bernadó, P. & Blackledge, M. A self-consistent description of the conformational behavior of chemically denatured proteins from NMR and small angle scattering. Biophys. J. 97, 2839–2845 (2009).
Rekhi, S. et al. Expanding the molecular language of protein liquid–liquid phase separation. Nat. Chem. 16, 1113–1124 (2024).
Kar, M. et al. Phase-separating RNA-binding proteins form heterogeneous distributions of clusters in subsaturated solutions. Proc. Natl. Acad. Sci. USA 119, e2202222119 (2022).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct. Funct. Bioinform. 78, 1950–1958 (2010).
Henriques, J. & Skepö, M. Molecular dynamics simulations of intrinsically disordered proteins: on the accuracy of the TIP4P-D water model and the representativeness of protein disorder models. J. Chem. Theory Comput. 12, 3407–3415 (2016).
Koder Hamid, M., Månsson, L. K., Meklesh, V., Persson, P. & Skepö, M. Molecular dynamics simulations of the adsorption of an intrinsically disordered protein: force field and water model evaluation in comparison with experiments. Front. Mol. Biosci. 9, https://doi.org/10.3389/fmolb.2022.958175 (2022).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Abraham, M. J, van der Spoel, D., Lindahl, E., Hess, B. & the GROMACS development team, GROMACS User Manual version 5.1.4, www.gromacs.org (2016). https://manual.gromacs.org/5.1.4/manual-5.1.4.pdf. Accessed 13 Mar 2026.
Posit team (2025). RStudio: Integrated Development Environment for R. Posit Software, PBC, Boston, MA. http://www.posit.co.
Wickham, H. ggplot2. (Springer International Publishing, 2016). https://doi.org/10.1007/978-3-319-24277-4.
Jaki, T. & Wolfsegger, M. J. Estimation of pharmacokinetic parameters with the R package PK. Pharm. Stat. 10, 284–288 (2011).
Källberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 7, 1511–1522 (2012).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Hess, B. Determining the shear viscosity of model liquids from molecular dynamics simulations. J. Chem. Phys. 116, 209–217 (2002).
Yeh, I.-C. & Hummer, G. System-size dependence of diffusion coefficients and viscosities from molecular dynamics simulations with periodic boundary conditions. J. Phys. Chem. B 108, 15873–15879 (2004).
Gajarska, Z. & Lohninger, H. Savitzky-Golay Filter - Coefficients, (2021). http://www.hyperspectral-imaging.org/cc_savgol_coeff.html. Accessed 13 Mar 2026.
Taylor, J. R. An Introduction to Error Analysis : the Study of Uncertainties in Physical Measurements, 2nd edn (University Science Books, 1997).
King, B. M., Silver, N. W. & Tidor, B. Efficient calculation of molecular configurational entropies using an information theoretic approximation. J. Phys. Chem. B 116, 2891–2904 (2012).
Morán, J., Fuentes, A., Liu, F. & Yon, J. FracVAL: an improved tunable algorithm of cluster–cluster aggregation for generation of fractal structures formed by polydisperse primary particles. Comput. Phys. Commun. 239, 225–237 (2019).
Acknowledgements
Production simulations as well as parts of the analyses were carried out on the Vienna Scientific Cluster. This research was funded in part by the Austrian Science Fund (FWF) [grant DOI: 10.55776/P30680, 10.55776/P30550]. For open access purposes, the author has applied a CC BY public copyright license to any author accepted manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
A.A.P., B.Z. designed the study. B.F., A.A.P. performed the simulations, data analysis and visualization. All authors wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Polyansky, A.A., Frühbauer, B. & Žagrović, B. Dynamic yet well-defined organization of the FUS RGG3 dense phase. Commun Chem (2026). https://doi.org/10.1038/s42004-026-01974-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-026-01974-z