Abstract
Complex systems usually feature interactions not only between pairwise entities but also among three or more entities. Hypergraph can effectively characterize these higher-order interactions. Meanwhile, all higher-order interactions can also be projected onto a number of lower-order interactions. Determining whether all higher-order interactions must be considered or if they can be approximated by lower-order interactions with minimal loss remains an open question. We propose a method to decompose higher-order structures in a stepwise way, thereby allowing to explore the impacts of hyperedges of any order. Experiments suggest that in some networks, incorporating higher-order interactions significantly enhances the accuracy of link prediction, while in others, the effect is insignificant. Therefore, the role of higher-order interactions varies in different types of networks. Overall, since the improvement in predictive accuracy provided by higher-order interactions is significant in some networks, we believe that the study of higher-order interactions is valuable.

Similar content being viewed by others
Introduction
From individual interactions in social networks to species symbiosis in ecosystems, and from stock market fluctuations to information flow through the Internet, real-world complex systems exhibit diverse behavioral patterns and time-varying dynamics through multi-level, multi-scale interactions1,2,3. The complexity of such systems does not solely arise from the intricacy of their operational rules but primarily stems from the interactions among constituent entities4,5. Networks, as a powerful mathematical tool, have been widely adopted to model these interactions by representing entities as nodes and pairwise relationships as links6,7. However, traditional network models often fall short in capturing the full spectrum of real-world interactions, which frequently involve collective, higher-order relationships among multiple entities rather than simple dyadic connections8,9,10,11,12. For instance, in scientific collaboration networks, projects involving multiple researchers cannot be fully characterized by pairwise interactions alone; higher-order models better encapsulate the collaborative dynamics of such multilateral teams13. Similarly, in commercial transactions, the involvement of intermediaries (e.g., third-party agents) alongside buyers and sellers necessitates a framework that accounts for multi-stakeholder interactions, offering a more accurate representation of business relationships13. In ecosystems, interactions between two microbial species are often regulated by other species14,15,16,17. For example, species A produces an antibiotic to inhibit species B, while a third species, C, secretes an enzyme that degrades this antibiotic, thereby reducing the inhibitory effect of species A on species B. This pattern of interactions among three microbial species cannot be adequately captured by pairwise interactions, as species C introduces an additional regulatory layer that influences the interaction between species A and B17. Beyond these examples, higher-order architectures are pervasive across disciplines: biochemical reactions frequently involve multiple substrates or enzymatic intermediaries18, and proteins assemble into functional complexes through multi-molecular interactions19.
The significance of higher-order interactions has already been noted by many scientists. For example, they enable more accurate predictions of drug combinations that may cause adverse side effects—effects that do not occur when the individual drugs are administered separately20,21. They are also essential for understanding the interdependencies among groups of neurons in avalanche dynamics22, and have been shown to enhance the accuracy of visual response predictions in the neurons of anesthetized cats23. Despite these empirical successes, Zhang et al.24 and Benson et al.25 have pointed out that the variability in the number of nodes contained in higher-order hyperedges can pose both complexity and sophistication challenges to the network analysis. While graph neural networks (GNNs) are highly effective in processing pairwise interactions, traditional GNNs often struggle to capture directly higher-order interactions26. Recent advances in hypergraph neural networks (HNNs) (see the survey27 and related references therein) have shown promise in addressing these limitations. Considering the additional complexity involved in analyzing higher-order interactions, it remains to be determined whether the introduction of these interactions is a significant innovation or merely a costly topological game28,29.
In this study, we employ link prediction, a fundamental task in network science, as a starting point for quantitatively analyzing the impact of higher-order interactions. Link prediction aims to predict the links that exist but have not yet been observed, or that will appear in the future, based on the observed network structure30,31,32. Hyperedge prediction is a natural extension of link prediction to hypergraphs, with the prediction object being extended from pairwise links to higher-order hyperedges33. Hyperedge prediction has become an active branch in the hypergraph study, with various prediction algorithms proposed24,34,35,36,37,38,39. Recently, Yoon et al.40 proposed the concept of n-projected graphs, where each node represents a set of n − 1 nodes, and an edge is formed between two such sets if their union contains exactly n nodes and there exists at least one hyperedge that contains all these n nodes. They argued that aggregating information from 2-projected up to n-projected graphs naturally extends the pairwise projected graph of a hypergraph (named as “collaboration networks” in early literature41). Their experiments show that incorporating a 3-projected graph into the pairwise projection significantly improves prediction accuracy, with diminishing returns at higher orders. Moreover, if one considers a very high-order projection, the prediction accuracy may decline. While Yoon et al.’s work offers a valuable framework for understanding the role of higher-order interactions in link prediction, their projection-based approach generates an aggregated graph containing a large number of new composite nodes, each representing a collection of several original nodes, which significantly increases computational and analytical complexity. Furthermore, their method does not cleanly strip away the effects of higher-order interactions in a stepwise way. For example, if one aims to retain information up to 5-order interactions and thus analyze the effects of 6-order and 6+-order interactions, this cannot be achieved by aggregating 2-projected to 5-projected graphs. This is because hyperedges of orders 3 through 5 are also projected into pairwise relationships. Therefore, regardless of how many orders are taken into account, the method of Yoon et al. will project all higher-order interactions to pairwise interactions, except that the set of nodes is different. Building on this insight, we propose a more applicable method: the n-reduced graph. This method preserves the structural information of lower-order hyperedges while stepwise decomposing higher-order interactions. Using this method, we can quantitatively assess the contribution of higher-order interactions and perform efficient and effective hyperedge prediction.
Results
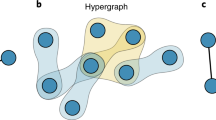
A hypergraph is denoted as G(V, E), where V = {v1, v2, …, vN} denotes the set of nodes, and E = {e1, e2, …, eM} denotes the set of hyperedges42,43. Unlike a traditional link that represents a connection between only two nodes, a hyperedge can capture the relationship among multiple nodes. Specifically, a hyperedge eα can involve two or more nodes, and its order, denoted by kα, is defined as the number of nodes eα contains. The number of hyperedges containing node vi is defined as the hyperdegree of vi, denoted as di. Hyperedge prediction aims to predict unobserved hyperedges based on the observed structure. Specifically, this algorithm tries to identify whether any unobserved hyperedge ec ⊆ 2V⧹E is indeed a true hyperedge33. Further details, including the n-reduced method, hyperedge prediction algorithms, data splitting and sampling strategies, and evaluation metrics, are presented in the Methods section.
Higher-order interactions enhance prediction
This study utilizes 12 real-world hypergraphs spanning diverse domains. As hyperedges containing ten or more nodes are relatively rare and computationally expensive to process, this study–like that of Yoon et al.40–focuses only on hyperedges with order k ≤ 10. Table 1 shows basic statistical properties of these twelve real-world hypergraphs, including their domain, number of nodes and hyperedges, and average hyperedges order. Detailed information for each hypergraph is included in Supplementary Note 1. Figure S1 (see Supplementary Note 1) illustrates the distribution of hyperedge orders. It is evident that lower-order hyperedges constitute the majority, while the number of hyperedges decreases significantly as the order increases. Figure S2 (see Supplementary Note 1) presents the cumulative distribution of hyperdegrees. These distributions exhibit a broad range and display similar characteristics to power-law distributions, though they cannot be precisely characterized by power laws44,45.
In a given hypergraph, all hyperedges of order k > n are decomposed into multiple n-order hyperedges by the n-reduced operator (see Methods). Clearly, the larger the value of n, the more higher-order structural information is preserved. As described in the “Data Splitting and Sampling” section of the Methods, to ensure consistency in evaluating the training and testing of hyperedge structures, hyperedges of order k > n are excluded from the test set. This ensures that predictions focus exclusively on hyperedges with k ≤ n. Figures 1 and S3 (see Supplementary Note 2) depict the trend of AUC as n increases across six hypergraphs. In all hypergraphs, AUC generally increases with increasing n, indicating that higher-order interactions contribute positively to the prediction accuracy. However, as n further increases, the change in AUC tends to level off, suggesting that the marginal benefit of incorporating additional higher-order information becomes smaller at larger values of n.
a–f show the results corresponding to datasets email-Eu, threads-math-sx, NDC-classes, iAF1260b, Nematode, and Pubmed, respectively. The solid line represents the average AUC over ten independent experiments, and the shaded area indicates the range of the average AUC plus and minus the standard deviation.
As shown in Figs. 1 and S3 (see Supplementary Note 2), the average AUC (see Methods) shows an upward trend for every hypergraph, suggesting that higher-order interaction information significantly enhances the accuracy of hyperedge prediction. However, directly using these results to quantify the contribution of higher-order interactions to the predictive performance is not rigorous. This is because hyperedges in the test set vary with different n, making the reasons underlying different AUC values very complicated. To eliminate confusion caused by varying test sets, we further evaluate the prediction performance for hyperedges of fixed order k within the test set. For example, we compare the predictive accuracies for hyperedges of order k = 3 across different n-reduced graphs (n ≥ 3). This ensures the consistency of the test set and more clearly reveals the impact of higher-order structural information.
Figures 2 and S4 (see Supplementary Note 2) illustrate the trend of AUC values as n increases for hyperedges of different orders k. For k ≥ 3, most networks (excluding email-Enron and DAWN) show a significant increase in AUC with the presence of more and more higher-order information. In NDC-classes and NDC-substances, the enhancement in prediction accuracy with higher-order information is evident. For example, when k = 3, as n increases from 3 to 7, AUC rises from 0.46 to 0.72 and from 0.68 to 0.87, respectively. In these cases, decomposing higher-order hyperedges into lower-order ones, even if the resulting hyperedges are still of order no less than 3, will reduce the predictive accuracy for 3-order hyperedges, indicating that higher-order structural information plays a crucial role in hyperedge prediction. For k = 2, for the networks NDC-classes, iAF1260b, Nematode, DBLP, NDC-substances and Pubmed, the higher-order interactions significantly improve the prediction accuracy of 2-order hyperedges, and for the other six networks, the increase of AUC is relatively flat or fluctuates with the addition of higher-order information. As shown in Fig. 2b, the increase in AUC tends to be flat with the increasing n, suggesting that reaching a certain level, the benefits of higher-order information diminish. Conversely, in other plots of Fig. 2, AUC continues to rise with increasing n, indicating that retaining more higher-order information can significantly enhance prediction performance in these networks. Additionally, we applied this method to predict hyperedges of order k > n. Different from the case k ≤ n, AUC values do not exhibit any clear trend as n increases (see Supplementary Note 3 for details). In summary, higher-order interactions generally contribute positively to hyperedge prediction; however, their impact varies across different networks. In some networks, they play a vital role, while in others, their contributions are negligible.
a–f show the results corresponding to datasets email-Eu, threads-math-sx, NDC-classes, iAF1260b, Nematode, and Pubmed, repectively. The shade of color indicates the value of AUC, with darker colors corresponding to larger AUC values.
Comparing n-reduced method with n-projected method
Recently, ref. 40 proposed an intriguing but distinctly different method known as the n-projected graph. This method is also useful for analyzing the impact of higher-order interactions on hyperedge prediction. Given a hypergraph G(V, E), its n-projected graph is denoted as \({G}_{n}^{p}=({V}_{n}^{p},{E}_{n}^{p})\), where the node set \({V}_{n}^{p}\) consists of all (n − 1)-node subsets of V, and each subset is represented as a single node in \({V}_{n}^{p}\), that is
In \({V}_{n}^{p}\), node vn and node un are connected if they satisfy the following two conditions: (1) vn and un differ by exactly one element among the n − 1 elements each contains, i.e., ∣un ∪ vn∣ = n; (2) the union of vn and un is fully contained in at least one hyperedge in E. Accordingly,
For a given n, the graph Gp(n) used for hyperedge prediction is constructed as a direct aggregation from the 2-projected graph up to the n-projected graph, denoted as \({G}^{p}(n):= \left({G}_{2}^{p},{G}_{3}^{p},\cdots \,,{G}_{n}^{p}\right)\). It is worth noting that for any n ≥ 2, Gp(n) contains only pairwise interactions and its complexity primarily comes from the quantity and heterogeneity of nodes. As each \({G}_{n}^{p}\) represents a union of sets rather than multisets, any duplicate instances of a hyperedge e will be excluded. Although the n-projected and n-reduced graphs are conceptually similar in that they both decompose higher-order interactions, they differ fundamentally in how this decomposition is performed. Therefore, we are particularly interested in determining which method preserves more useful structural information. A natural assumption is that the one retaining more useful information will yield better performance in hyperedge prediction. For comparison, we use the same six features and logistic regression model to predict hyperedges in the n-projected graph, following the method proposed by ref. 40. Based on an edge’s attribute in Gp(n), we can identify its order in the original hypergraph. For instance, an edge between two sets of four nodes corresponds to a five-order hyperedge in the original hypergraph. Figures 3 and S6 (see Supplementary Note 4) compare the predictive performance of these two methods across hyperedge orders ranging from 3 to 6, with n = 7. The results show that the n-reduced method significantly outperforms the n-projected method across all twelve real networks and different orders. Specifically, the best AUC achieved by the n-projected graph is 0.74, with an average AUC of 0.62, whereas the n-reduced graph achieves a best AUC of 0.94 and an average AUC of 0.83. These findings suggest that if higher-order interactions need to be reduced for convenient and efficient analysis or some other reasons, the method of the n-reduced graph is superior in retaining informative higher-order structure.
a–f show the results corresponding to datasets email-Eu, threads-math-sx, NDC-classes, iAF1260b, Nematode, and Pubmed, respectively. Here, n is set to 7, and all results are averaged over ten independent experiments. The y-axis represents the order k of the predicted hyperedges. The dark blue bars denote the average AUC for the n-reduced method, while the magenta bars represent the average AUC for the n-projected method. To provide a clear comparison of prediction accuracy between the two methods, the length of each bar is scaled according to the AUC value of each method as a proportion of the sum of AUC values of both methods. For instance, if the AUC for the n-reduced method is 0.8 and the AUC for the n-projected method is 0.7, the lengths of the dark blue and magenta bars would be 8/15 and 7/15, respectively.
We have also leveraged other state-of-the-art algorithms, including hyperedge prediction using resource allocation (HPRA)37, neural hypergraph link prediction (NHP)46, and nonuniform hyper-network embedding with dual mechanism (NHNE)47. Detailed results are provided in Supplementary Note 5, which consistently demonstrate that our findings remain robust across these various methods.
Discussion
Before hypergraphs became popular, bipartite graphs48,49 and collaboration networks41,50 were widely used to represent higher-order interactions. For instance, to describe scientific collaborations, bipartite graphs utilize two types of nodes (one for authors and one for articles) and connect each article to its authors. Collaboration networks simplify this by using pairwise links between scientists who have co-authored at least one article. Although bipartite graphs and hypergraphs are mathematically equivalent, bipartite graphs introduce additional complexity due to their heterogeneous node types. Collaboration networks reduce complexity by projecting higher-order interactions into pairwise links, which inevitably leads to information loss. While edge weights can help retain some details, they cannot fully capture the richer structure of hypergraphs51,52,53. Despite this, collaboration networks remain useful because they are easier to analyze than hypergraphs. A critical question arises: In which scenarios is it necessary to retain all information from higher-order interactions, and in which cases can they be represented as pairwise interaction networks, such as collaboration networks? If higher-order interactions cannot be accurately represented as pairwise interaction networks, can they be represented as lower-order interactions to reduce the complexity of the analysis?
The key to addressing these questions lies in quantifying the impact of higher-order interactions in a given scenario. For example, in a network where tasks such as link prediction, community detection, identification of critical nodes, and spreading prevalence estimation are considered, how can we ascertain whether it is necessary to consider hyperedges of 5 and 5+ orders? A straightforward but overly simplistic approach is to remove all hyperedges with k > 4 and compare the performance on specific tasks before and after the removal. However, this approach is too crude, because even if we apply the unweighted pairwise projection, some information about 5-order and 5+-order interactions will be retained. That is to say, if we cannot analyze these higher-order hyperedges because of the limited computational resources, we can at least project them into multiple pairwise interactions rather than remove them entirely. A less aggressive method is to directly compare hypergraphs with their pairwise projections54. However, this results in a significant loss of information, and we cannot focus solely on the impacts of hyperedges of orders k ≥ 5, since 3-order and 4-order interactions are also projected onto pairwise interactions. The optimal approach is to employ stepwise decomposition to retain as much information as possible. The effect of any information that has to be lost should then be attributed to the influence of higher-order interactions. For example, to analyze the effect of hyperedges of 5 and 5+ orders, we should strive to retain their information through interactions of orders 2 to 4 before conducting comparative analyses.
The aforementioned idea is applied in both n-projected and n-reduced graphs to stepwise decompose higher-order interactions. However, they are different. The n-projected operator ultimately projects all higher-order interactions into pairwise interactions, introducing a large number of heterogeneous nodes to retain information about the higher-order hyperedges as much as possible. In contrast, the n-reduced operator represents higher-order interactions through lower-order interactions while keeping the set of nodes unchanged. Although these two methods share the same starting point, the n-reduced graph may be more suitable as an analyzing tool, because researchers seem to be less inclined to deal with heterogeneous nodes. Otherwise, bipartite graphs would be more popular than hypergraphs for representing higher-order interactions.
In this study, we compare the n-projected and n-reduced methods by employing link prediction as an entry point to quantify the influence of higher-order interactions. Link prediction is chosen because it serves as a more fundamental task compared to the analysis of specific networked dynamics. Although our method has yielded promising results in hyperedge prediction, its applicability to other tasks–such as community detection and the identification of critical nodes–remains to be further exploration. From the experiment in this study, there are two main conclusions: First, higher-order interactions generally have a significant and positive effect on predictive accuracy, although their effectiveness may vary across different networks and is not universally guaranteed. Second, compared to the n-projected method, the n-reduced method proposed in the study retains more higher-order information. This is evidenced by the fact that the n-reduced method achieves a higher accuracy in hyperedge prediction under the same conditions.
In conclusion, we offer three specific suggestions based on our findings. First, our results show that higher-order interactions have a significant and positive impact on hyperedge prediction. Therefore, when computationally feasible, researchers are encouraged to prioritize the use of hypergraphs to represent higher-order interactions, rather than reducing them to pairwise interactions. Second, we observed that the influence of higher-order interactions varies significantly across different real-world networks. Therefore, in order to draw more comprehensive and reliable conclusions, further analysis should be conducted on domain-specific and topology-specific hypergraphs. This may even include the distinction of which hyperedges bring richer information in a given hypergraph55. In addition, we propose a toy model to investigate whether a small fraction of all possible nested lower-order hyperedges can effectively retain the structural information of higher-order interactions. As shown in Supplementary Note 6, retaining only 20% of reduced hyperedges–specifically those involving large-degree nodes–can achieve very close prediction accuracy to the original n-reduction method. Finally, as the n-reduced operator can effectively retain higher-order interaction information, we recommend researchers use the n-reduced graphs as an analytical tool to quantify the role of higher-order interactions on specific dynamics (e.g., propagation and synchronization)56,57,58,59 and other graph mining tasks (e.g., critical nodes identification and community detection)60,61,62.
As with any research, this study has its limitations. The n-reduced method is heuristic in nature and lacks a rigorous mathematical or physical theory to quantify how much information it remains. Therefore, an important yet challenging direction for future study is how to design a rigorous theory. Maybe one can first consider a statistical theory on hypergraph configuration model63. Benson et al.25 attempted to use lower-order interaction data to predict the closure of simplicial complexes, and thus, another issue for future study is to extend the current stepwise decomposition method for simplicial complexes. Additionally, future investigations could benefit from exploring diffusion models or statistics-driven approaches. These methods may provide deeper theoretical insights into the effectiveness of hypergraph-based analyses, thereby addressing some of the current limitations.
Methods
The n-reduced graph
In order to analyze the role of all hyperedges with orders larger than a threshold order, we propose the following method, referred to as the n-reduced graph. Given a hypergraph G(V, E), its corresponding n-reduced graph, denoted by Gn(V, En), is defined as
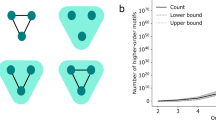
As defined above, the hyperedge set En comprises two parts: (1) all hyperedges in G with kα≤n are retained in En; (2) the hyperedges with kα > n are decomposed into \((\begin{array}{c}{k}_{\alpha }\\ n\end{array})\) hyperedges, each of which has an order of n. Note that when n = 2, Gn is equivalent to the unweighted pairwise projection of G; and when \(n={\max }_{\alpha }({k}_{\alpha })\), Gn is equivalent to the original hypergraph G. Unlike the n-projected graph, the n-reduced graph retains the node set and all hyperedges with order no greater than n, while decomposing hyperedges of order greater than n using sets of hyperedges with order n. In summary, the n-reduced graph is an approximated representation of the original hypergraph that consists of hyperedges of orders no greater than n, with the goal of minimizing information loss, which is particularly suitable for evaluating the role of hyperedges with orders larger than a threshold. Figure 4 illustrates the construction process of a three-reduced graph. This hypergraph consists of eight nodes and seven hyperedges, with the orders of six hyperedges (e1, e2, e3, e4, e5, and e6) not exceeding 3. These hyperedges are directly retained in the 3-reduced graph. For the hyperedge e7 with kα > 3, we generate all three-order hyperedges by traversing all triples from the set of nodes in the hyperedge e7, which are subsequently added to the three-reduced graph.
a shows the original structure, b illustrates the 3-reduction process, and c displays the reorganized structure. This hypergraph contains eight nodes and seven hyperedges. Among them, the orders of e2, e3, and e5 are 2, the orders of e1, e4 and e6 are 3, and the order of e7 is 4. When n = 3, hyperedge e7 is decomposed into all combinations of its nodes, say {v1, v2, v3}, {v1, v2, v5}, {v1, v3, v5}, and {v2, v3, v5}.
Features and classifier
This study applies three local similarity features and three weighted features to evaluate the likelihood of a set of nodes to form a hyperedge. The former measures whether the nodes in the set are all adjacent to some other nodes (two nodes are adjacent if they appear together in at least one hyperedge), while the latter measures the closeness between pairs of nodes. In brief, the former treats the candidate node set as a whole, whereas the latter views it as the sum of multiple pairwise relationships.
In this study, local similarity features are direct extensions of classic similarity indices used in link prediction64,65,66,67,68,69. (1) Common neighbor index: The common neighbors of a hyperedge e are the common neighbors of all nodes in e, that is
where N(v) represents the neighbor set of the node v. (2) Jaccard coefficient: The Jaccard coefficient is a normalization index that divides the number of common neighbors among all nodes in hyperedge e by the total number of neighbors of any nodes in e:
(3) Adamic–Adar index: The Adamic–Adar index penalizes high-degree nodes by dividing the degrees of common neighbors, as
Weighted features measure the closeness among the candidates node set by using different averages of the weights of direct connections between node pairs. Let Wuv denote the weight of the connection between nodes u and v, which is defined as the number of hyperedges containing both u and v. Clearly, if u and v are adjacent, then Wuv > 0, otherwise, Wuv = 0. For any candidate node set e, let Ee represent the set of pairs of adjacent nodes in e, that is
Note that u ∈ e and v ∈ e do not necessarily imply Wuv > 0, as e may not be a hyperedge in the original hypergraph. Accordingly, we can calculate the average weight of all adjacent node pairs in e using three different methods. (4) Geometric mean:
(5) Harmonic mean:
(6) Arithmetic mean:
This study integrates the six features into a hyperedge feature vector and utilizes the Logistic regression to train the model70. That is, the feature vector of a candidate node set e can be represented as a linear combination of \([{{{\rm{CN}}}}(e),{{{\rm{JC}}}}(e),{{{\rm{AA}}}}(e),{{{\rm{GM}}}}(e),{{{\rm{HM}}}}(e),{{{\rm{AM}}}}(e)]\), and then this feature vector is used to learn the scoring function f(e), such that:
where ϵ is the binary classification threshold of f(e), used to determine whether e is a potential hyperedge.
Data splitting and sampling
This study randomly splits each dataset into a training set and a test set in an 8:2 ratio. The training set is utilized to train the model, while the test set is used to evaluate the performance. For a given n, hyperedges in the training set with orders greater than n are decomposed into multiple n-order hyperedges using the n-reduced operator. It is worth noting that, we do not reduce the hyperedges in the test set in order to preserve the integrity of the original true hyperedges, which can avoid introducing potentially misleading hyperedges. To enhance the model’s generalization capability, 5-fold cross-validation is performed on the training set. Specifically, the training set is randomly divided into five equal-sized subsets. In each time, one subset is designated as the validation set, while the remaining four subsets are used to calculate feature vectors, maintaining an 8:2 ratio between training and validation data. This process is repeated five times, ensuring that each subset is treated as the validation set once.
To address the significant disparity between the number of candidate hyperedges and true hyperedges in hyperedge prediction (note that, this disparity is much larger than that in link prediction for pairwise interaction networks, because the number of possible hyperedges in a hypergraph is about 2N, much larger than N2 in a pairwise interaction network), this study adopts a negative sampling strategy, which generates non-existent hyperedges (negative samples) to balance with the missing hyperedges (positive samples) in both size and distribution38,71. For each positive hyperedge e in the validation set or the test set, we randomly remove one node v0 from e and then randomly select a node from the neighbors of e to replace v0. If the resulting set of nodes, eneg, does not belong to the hyperedge set En, it is considered to be a valid negative sample. This negative sampling method ensures that the generated negative samples are structurally similar to positive hyperedges, thereby increasing the difficulty of prediction. Consequently, this method effectively enhances the model’s generalization capability, enabling it to more accurately distinguish between existent and non-existent hyperedges.
Evaluation metrics
Hyperedge prediction is a specialized binary classification task aimed at accurately distinguishing between positive and negative samples. To evaluate the predictive performance of the model, we utilize the Area Under the ROC Curve (AUC) as the evaluation metric72. AUC is widely used for evaluating classification models, and our previous works73,74,75,76 demonstrate that AUC provides superior discriminability and greater information content compared to other commonly used metrics. The range of AUC is [0, 1]. If the prediction is assigned completely at random, the AUC value will be 0.5. A higher AUC indicates better predictive performance. Although some studies suggest that AUC may exhibit evaluation bias due to imbalanced positive and negative samples77,78,79, the negative sampling method employed in this study ensures a balanced number of positive and negative samples, thereby mitigating these issues. To enhance the robustness of the results, we conducted 10 independent experiments and used the average AUC as the final outcome.
Data availability
All data associated with this study are accessible at https://github.com/jackyjh/n-reduction-graph.
Code availability
The code used to analyze the data is available at https://github.com/jackyjh/n-reduction-graph.
References
Sterman, J. D. Learning in and about complex systems. Syst. Dyn. Rev. 10, 291–330 (1994).
Arthur, W. B. Complexity and the economy. Science 284, 107–109 (1999).
Ladyman, J., Lambert, J. & Wiesner, K. What is a complex system? Eur. J. Philos. Sci. 3, 33–67 (2013).
Kauffman, S. A. At Home in the Universe: The Search for Laws of Self-Organization and Complexity (Oxford Univ. Press, 1996).
Hidalgo, C. A. et al. The principle of relatedness. In Proc. Ninth International Conference on Complex Systems 451–457 (Springer Press, 2018)
Barabási, A. L. Network Science (Cambridge Univ. Press, 2016).
Newman, M. E. J. Networks. (Oxford Univ. Press, 2018).
Battiston, F. et al. Networks beyond pairwise interactions: structure and dynamics. Phys. Rep. 874, 1–92 (2020).
Battiston, F. et al. The physics of higher-order interactions in complex systems. Nat. Phys. 17, 1093–1098 (2021).
Battiston, F. & Petri, G. Higher-order Systems (Springer Press, 2022).
Bianconi, G. Higher-order Networks (Cambridge Univ. Press, 2021).
Bick, C., Gross, E., Harrington, H. A. & Schaub, M. T. What are higher-order networks? SIAM Rev. 65, 686–731 (2023).
Bonacich, P., Holdren, A. C. & Johnston, M. Hyper-edges and multidimensional centrality. Soc. Netw. 26, 189–203 (2004).
Wootton, J. T. Indirect effects in complex ecosystems: recent progress and future challenges. J. Sea Res. 48, 157–172 (2002).
Werner, E. E. & Peacor, S. D. A review of trait-mediated indirect interactions in ecological communities. Ecology 84, 1083–1100 (2003).
Poisot, T., Stouffer, D. B. & Gravel, D. Beyond species: why ecological interaction networks vary through space and time. Oikos 124, 243–251 (2015).
Bairey, E., Kelsic, E. D. & Kishony, R. High-order species interactions shape ecosystem diversity. Nat. Commun. 7, 12285 (2016).
Klamt, S., Haus, U. U. & Theis, F. Hypergraphs and cellular networks. PLoS Comput. Biol. 5, e1000385 (2009).
Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Nguyen, D. A., Nguyen, C. H. & Mamitsuka, H. Central-smoothing hypergraph neural networks for predicting drug-drug interactions. IEEE Trans. Neural Netw. Learn. Syst. 35, 11620–11625 (2023).
Vaida, M. & Purcell, K. Hypergraph link prediction: learning drug interaction networks embeddings. In 2019 18th IEEE International Conference on Machine Learning and Applications 1860–1865 (IEEE Press, 2019).
Yu, S. et al. Higher-order interactions characterized in cortical activity. J. Neurosci. 31, 17514–17526 (2011).
Ganmor, E., Segev, R. & Schneidman, E. Sparse low-order interaction network underlies a highly correlated and learnable neural population code. Proc. Natl Acad. Sci. USA 108, 9679–9684 (2011).
Zhang, M., Cui, Z., Jiang, S. & Chen, Y. Beyond link prediction: predicting hyperlinks in adjacency space. In Proc. Thirty-Second AAAI Conference on Artificial Intelligence 4430-4437 (AAAI Press, 2018)
Benson, A. R., Abebe, R., Schaub, M. T., adbabaie, A. & Kleinberg, J. Simplicial closure and higher-order link prediction. Proc. Natl Acad. Sci. USA 115, E11221–E11230 (2018).
Zhang, M. & Chen, Y. Link prediction based on graph neural networks. In Proc. 32nd International Conference on Neural Information Processing Systems 5171–5181 (Curran Associates Inc. Press, 2018).
Kim, S. et al. A survey on hypergraph neural networks: an in-depth and step-by-step guide. In Proc. 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining 6534–6544 (ACM Press, 2024).
Wolf, M. M., Klinvex, A. M. & Dunlavy, D. M. Advantages to modeling relational data using hypergraphs versus graphs. In 2016 IEEE High Performance Extreme Computing Conference 1–7 (IEEE Press, 2016).
Torres, L., Blevins, A. S., Bassett, D. & Eliassi-Rad, T. The why, how, and when of representations for complex systems. SIAM Rev. 63, 435–485 (2021).
Lü, L. & Zhou, T. Link prediction in complex networks: a survey. Phys. A 390, 1150–1170 (2011).
Martínez, V., Berzal, F. & Cubero, J. C. A survey of link prediction in complex networks. ACM Comput. Surv. 49, 1–33 (2016).
Zhou, T. Progresses and challenges in link prediction. iScience 24, 103217 (2021).
Chen, C. & Liu, Y. Y. A survey on hyperlink prediction. In IEEE Transactions on Neural Networks and Learning Systems 1–17 (IEEE Press, 2023).
Xu, Y, Rockmore, D, Kleinbaum, A. M. Hyperlink prediction in hypernetworks using latent social features. In Proc. 16th International Conference on Discovery Science 324–339 (Springer Press, 2013).
Tu, K., Cui, P., Wang, X., Wang, F. & Zhu, W. Structural deep embedding for hyper-networks. In Proc. Thirty-Second AAAI Conference on Artificial Intelligence 426–433 (AAAI Press, 2018).
Zhang, R., Zou, Y. & Ma, J. Hyper-SAGNN: a self-attention based graph neural network for hypergraphs. In Proc. International Conference on Learning Representations (2020).
Kumar, T., Darwin. K., Parthasarathy. S. & Ravindran, B. HPRA: hyperedge prediction using resource allocation. In Proc. 12th ACM Conference on Web Science 135–143 (ACM Press, 2020).
Hwang, H., Lee, S., Park, C. & Shin, K. AHP: learning to negative sample for hyperedge prediction. In Proc. 45th International ACM SIGIR Conference on Research and Development in Information Retrieval 2237–2242 (ACM Press, 2022).
Contisciani, M., Battiston, F. & Bacco, C. D. Inference of hyperedges and overlapping communities in hypergraphs. Nat. Commun. 13, 7229 (2022).
Yoon, S., Song, H., Shin, K. & Yi, Y. How much and when do we need higher-order information in hypergraphs? A case study on hyperedge prediction. In Proc. Web Conference 2020 2627–2633 (ACM Press, 2020).
Newman, M. E. J. The structure of scientific collaboration networks. Proc. Natl Acad. Sci. USA 98, 404–409 (2001).
Berge, C. Hypergraphs: Combinatorics of Finite Sets (Elsevier Press, 1989).
Bretto, A. Hypergraph Theory: An Introduction (Springer Press, 2013).
Holme, P. Rare and everywhere: perspectives on scale-free networks. Nat. Commun. 10, 1016 (2019).
Broido, A. D. & Clauset, A. Scale-free networks are rare. Nat. Commun. 10, 1017 (2019).
Yadati, N. et al. NHP: neural hypergraph link prediction. In Proc. 29th ACM International Conference on Information & Knowledge Management 1705–1714 (ACM Press, 2020).
Huang, J., Chen, C., Ye, F., Hu, W. & Zheng, Z. Nonuniform hyper-network embedding with dual mechanism. ACM Trans. Inf. Syst. 38, 28 (2020).
Lambiotte, R. & Ausloos, M. Uncovering collective listening habits and music genres in bipartite networks. Phys. Rev. E 72, 066107 (2005).
Shang, M. S., Lü, L., Zhang, Y. C. & Zhou, T. Empirical analysis of web-based user-object bipartite networks. EPL 90, 48006 (2010).
Zhang, P. P. et al. Model and empirical study on some collaboration networks. Phys. A 360, 599–616 (2006).
Newman, M. E. J. Scientific collaboration networks. II. Shortest paths, weighted networks, and centrality. Phys. Rev. E 64, 016132 (2001).
Zhou, T., Ren, J., Medo, M. & Zhang, Y. C. Bipartite network projection and personal recommendation. Phys. Rev. E 76, 046115 (2007).
Wang, Y. & Kleinberg, J. From graphs to hypergraphs: hypergraph projection and its reconstruction. In Proc. International Conference on Learning Representations (2024).
Iacopini, I., Petri, G., Barrat, A. & Latora, V. Simplicial models of social contagion. Nat. Commun. 10, 2485 (2019).
Musciotto, F., Battiston, F. & Mantegna, R. N. Detecting informative higher-order interactions in statistically validated hypergraphs. Commun. Phys. 4, 218 (2021).
Majhi, S., Perc, M. & Ghosh, D. Dynamics on higher-order networks: a review. J. R. Soc. Interface 19, 20220043 (2022).
Boccaletti, S. et al. The structure and dynamics of networks with higher order interactions. Phys. Rep. 1018, 1–64 (2023).
Zhang, Y., Lucas, M. & Battiston, F. Higher-order interactions shape collective dynamics differently in hypergraphs and simplicial complexes. Nat. Commun. 14, 1605 (2023).
Wang, W. et al. Epidemic spreading on higher-order networks. Phys. Rep. 1056, 1–70 (2024).
Zeng, Y., Huang, Y., Ren, X. L. & Lü, L. Identifying vital nodes through augmented random walks on higher-order networks. Inf. Sci. 679, 121067 (2024).
Xiao, J. & Xu, X. K. Community detection from fuzzy and higher-order perspectives. EPL 144, 11003 (2023).
Liu, Y., Fan, Y. & Zeng, A. Higher-order interactions disturb community detection in complex networks. Phys. Lett. A 494, 129288 (2024).
Philip, S. C. Configuration models of random hypergraphs. J. Complex Netw. 8, cnaa018 (2020).
Barabási, A. L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Liben-Nowell, D. & Kleinberg, J. The link-prediction problem for social networks. J. Am. Soc. Inf. Sci. Technol. 58, 1019–1031 (2007).
Zhou, T., Lü, L. & Zhang, Y. C. Predicting missing links via local information. Eur. Phys. J. B 71, 623–630 (2009).
Newman, M. E. J. Clustering and preferential attachment in growing networks. Phys. Rev. E 64, 025102(R) (2001).
Jaccard, P. Distribution de la flore alpine dans le Bassin des Dranses et dans quelques régions voisines. Bull. Soc. Vaud. Sci. Nat. 37, 241–272 (1901).
Adamic, L. A. & Adar, E. Friends and neighbors on the web. Soc. Netw. 25, 211–230 (2003).
Hosmer, D. W., Lemeshow, S. & Sturdivant, R. X. Applied Logistic Regression (John Wiley & Sons Press, 2013).
Patil, P., Sharma, G. & Murty, M. N. Negative sampling for hyperlink prediction in networks. In 24th Pacific-Asia Conference on Knowledge Discovery and Data Mining 607–619 (Springer Press, 2020).
Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 27, 861–874 (2006).
Zhou, T. Discriminating abilities of threshold-free evaluation metrics in link prediction. Phys. A 615, 128529 (2023).
Jiao, X. et al. Comparing discriminating abilities of evaluation metrics in link prediction. J. Phys. Complex. 5, 025014 (2024).
Wan, S., Bi, Y., Jiao, X. & Zhou, T. Quantifying discriminability of evaluation metrics in link prediction for real networks. Preprint at arXiv. 2409.20078 (2024).
Bi, Y., Jiao, X., Lee, Y. L. & Zhou, T. Inconsistency of evaluation metrics in link prediction. PNAS Nexus 3, 498 (2024).
Lobo, J. M., Jiménez-Valverde, A. & Real, R. AUC: a misleading measure of the performance of predictive distribution models. Glob. Ecol. Biogeogr. 17, 145–151 (2007).
Yang, Y., Lichtenwalter, R. N. & Chawla, N. V. Evaluating link prediction methods. Knowl. Inf. Syst. 45, 751–782 (2015).
Chen, J., Muscoloni, A., Abdelhamid, I., Wu, Y. & Cannistraci, C. V. Generalizing the AUC–ROC for unbalanced data, early retrieval and link prediction evaluation. Preprints. 202209.0277 (2024).
Leskovec, J., Kleinberg, J. & Faloutsos, C. Graph evolution: densification and shrinking diameters. ACM Trans. Knowl. Discov. Data 1, 2–es (2007).
Yin, H., Benson, A. R., Leskovec, J. & Gleich, D. F. Local higher-order graph clustering. In Proc. 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 555–564 (ACM Press, 2017)
Zachary, A. K. et al. BiGG models: a platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Res. 44, D515–D522 (2016).
Dallas, T. A. et al. Gauging support for macroecological patterns in helminth parasites. Glob. Ecol. Biogeogr. 27, 1437–1447 (2018).
Carlson, C. J., Zipfel, C. M., Garnier, R. & Bansal, S. Global estimates of mammalian viral diversity accounting for host sharing. Nat. Ecol. Evol. 3, 1070–1075 (2019).
Sen, P. et al. Collective classification in network data. AI Mag. 29, 93 (2008).
Ley, M. The DBLP computer science bibliography: evolution, research issues, perspectives. In International Symposium on String Processing and Information Retrieval 1–10 (Springer Press, 2002).
Acknowledgements
The authors are supported by the National Natural Science Foundation of China under Grant Nos. 42361144718 and T2293771, and STI 2030-Major Project under Grant No. 2024ZD0523903.
Author information
Authors and Affiliations
Contributions
T.Z. proposed the study. J.B., Y.B. and T.Z. designed the study. J.B. performed the experiments. Y.B. prepared the figures. J.B., Y.B. and T.Z. analyzed the data. J.B., Y.B. and T.Z. wrote and edited the manuscript. J.B., Y.B. and T.Z. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Physics thanks Tim LaRock and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bian, J., Zhou, T. & Bi, Y. Unveiling the role of higher-order interactions via stepwise reduction. Commun Phys 8, 228 (2025). https://doi.org/10.1038/s42005-025-02157-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42005-025-02157-3