Abstract
Microfluidics experiments offer high-resolution insights into transport and chemical processes in porous media, yet direct measurement of evolving concentration profiles remains challenging. Numerical simulations can serve as virtual probes but are labor-intensive and computationally expensive. Here, we develop a physics-based machine learning toolbox that transforms such simulations into efficient and scalable virtual probes. Central to our toolbox is the non-intrusive reduced basis method, supported by the U-Net and the Convolutional Autoencoder, which learns mappings from experimental images and physical parameters to concentration profiles. By incorporating physics into its construction, the toolbox delivers accurate predictions with a limited number of training samples. Applied to two microfluidics experiments with different base patterns, the toolbox predicts spatio-temporal concentration profiles, effective diffusivities, and locations with a high probability of precipitation. This paves the way for digital twins that enable real-time analysis and tuning of experiments on the fly.
Similar content being viewed by others
Introduction
Microfluidics has been widely adopted in geoscience applications, including nuclear waste disposal and carbon sequestration, to investigate transport and chemical processes at the pore scale1,2,3,4,5,6,7,8,9. These investigations provide critical insights into phenomena, such as radionuclide transport in subsurface nuclear waste repositories and carbon dioxide mineralization during sequestration, that inform decision-making and risk assessment3,4,5,6,7,8,9,10,11,12,13,14,15,16. Microfluidics experiments typically utilize micromodels that replicate natural porous-media processes under controlled conditions, including temperature and pressure2,17,18,19,20,21,22. When coupled with in situ techniques such as Raman spectroscopy and optical microscopy, these experiments offer high-resolution and non-destructive qualitative insights into the transport and chemical processes, for instance by capturing the evolution of pore structures and mineral consumption3,4,6,7,9,23,24,25. However, quantitative measurements, particularly of aqueous concentrations, remain challenging due to the lack of optimum probing devices1,26,27,28,29,30,31,32.
To overcome this limitation, quantitative measurements are often carried out through process-based numerical simulations, such as the Lattice–Boltzmann method30,33,34 and the finite volume method10,27,28,29,31,32. These simulations serve as virtual probes for estimating aqueous concentrations within microfluidics devices, enabling direct analysis of transport and chemical processes. Probing concentration profiles during experiments is essential when guiding imaging tools is necessary to locate fast events over large samples, such as mineral nucleation in porous media or reactor chambers30,35,36, while real-time parameter adjustments and fast analysis are required to investigate specific processes and optimize subsequent experiments37,38.
Up to now, real-time quantitative measurements and analyses are still not feasible, meaning that experiments and numerical simulations (that serve as the probes) cannot be conducted side by side27,28. The key limitations are: (1) labor-intensive image segmentation required to process numerous experimental images for defining numerical simulation domains, and (2) high computational costs associated with performing numerical simulations on each image due to the fine discretizations needed to resolve complex microstructures. Machine learning is a promising approach to address these challenges. Many studies utilize Convolutional Neural Networks (CNNs), trained in a data-driven manner, to directly map experimental images to effective properties39,40,41,42,43,44,45,46,47,48,49. The training data is generated through manual segmentation of experimental images, followed by performing numerical simulations, such as Lattice–Boltzmann or finite volume methods, on each image39,46,47,49. However, such approaches are not designed to estimate transport or chemical states, limiting their applicability as experimental probes. Additionally, this method requires a large number of training samples, which are costly to generate.
To address these limitations, we develop a physics-based machine-learning toolbox that emphasizes (1) process explainability, capturing concentration profiles, subsequently deriving effective properties and saturation index, and (2) efficient construction by incorporating physics into the machine learning model, ensuring accurate predictions with a small number of training samples and minimizing computational costs. Through these concepts, this toolbox enables experimentalists to perform fast image segmentation, efficient predictive modeling of aqueous concentrations, evaluation of effective properties, and identification of locations with a high probability of precipitation.
The core method in this toolbox is the non-intrusive reduced basis (NI-RB) method that provides accurate and efficient mapping from physical parameters to physical states (in our case, concentration profiles)50,51,52,53,54,55. It is achieved by reducing degrees of freedom of a physical problem and maintaining its structure in basis functions50,51,52,53,54,55. This approach is different from a widely used approach, the Physics-Informed Neural Network (PINN), where the physical problem is encoded in the loss function56,57,58. The PINN is, however, reported to suffer from the gradient pathology problem59, an expensive training process due to complex loss landscape60,61,62, and requires more training samples than the NI-RB method50,52,63. Since the NI-RB method is inefficient when dealing with high-dimensional parameters such as images, we combine this method with the U-Net and the Convolutional Autoencoder (CAE).
The U-Net is employed to binarize the experimental images, while the CAE subsequently reduces the dimensionality of the binarized outputs. This low-dimensional representation facilitates efficient mapping to concentration profiles via the NI-RB. In addition, the CAE is also employed to generate additional binary images, thereby augmenting the training samples for the NI-RB model training. After training the CAE model and verifying that its decoder can accurately reconstruct the input binarized images, the resulting latent space captures the structural features and variability present in the training samples. By sampling from this latent space, additional binary images (statistically similar to those in the training samples) can be generated. The use of CAE for dimensionality reduction and data augmentation has been demonstrated in numerous studies64,65,66,67, including those focused on porous media applications65,67.
We apply our toolbox on microfluidics experiments aiming for predicting locations with a high probability of precipitation, and understanding effective diffusivity changes on evolving structures driven by chemical reactions. These experiments necessitate probing on multiple species concentrations to predict the saturation index, and tracer concentration to estimate the effective diffusivity. Numerical simulations based on the Lattice–Boltzmann method are used as probes for multiple species and tracer concentrations before and after chemical reactions. In these experiments, we induced celestine (SrSO4) precipitation in homogeneous circular grain patterns and natural rock patterns. Time-lapse optical microscopy captured pore structure alterations before and after reaction. Our toolbox performs fast segmentation utilizing the U-Net to transform optical microscopy images into simulation domains. The CAE uses the U-Net outputs to generate more training samples for the NI-RB model and reduce the dimensionality of the U-Net outputs before they enter the NI-RB model. The NI-RB model then provides quantitative multiple species and tracer concentrations, subsequently estimates saturation index (SI), locations with a high probability of precipitation, and effective diffusivity. Our approach demonstrates robustness, requiring a small number of training samples and short training time. This framework lays the foundation for creating digital twins of microfluidics experiments, enabling real-time physics and chemistry analyses in which experimental conditions can be adjusted dynamically. Such capabilities would allow on-the-fly modification of boundary conditions during experiments and further reduce the number of physical experiments required to explore specific phenomena.
Results
Physics-based machine learning framework
Framework capabilities
The physics-based machine learning framework in our toolbox replicates the step-by-step workflow of process-based numerical modeling, thereby preserving the original modeling workflow while significantly reducing computational time. The overall implementation uses our open-source Python library Chip Analyzer and Calculator (cac). The core idea is to map RGB optical microscopy images and time to the distribution functions (physical solutions) of Lattice–Boltzmann simulations. Concentration profiles are obtained from these distribution functions and subsequently used to compute the saturation index (SI) and effective diffusivity. Importantly, this mapping is achieved using a limited number of training samples. The use of images as inputs to the toolbox and the challenge of limited training samples are especially important in the context of microfluidics experiments for two main reasons: (1) accurate analysis and interpretation of experimental results require direct numerical simulations on the experimental images68, and (2) generating training data for machine learning models is both computationally expensive and labor-intensive, as it involves segmenting optical microscopy images and running Lattice–Boltzmann simulations on the segmented outputs69.
The toolbox provides two modules: (1) prediction of multiple species concentration profiles at the current time step, followed by prediction of the saturation index (SI) at the next time step to identify locations with a high probability of precipitation (workflow marked with green arrows in Fig. 1), and (2) prediction of tracer concentration at the current time, followed by estimation of effective diffusivity. This framework relies on a common assumption in coupled transport-chemical modeling: geometry changes induced by chemical reactions evolve more slowly than the transport processes30,68,69,70.
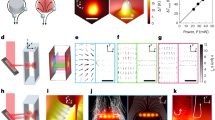
Purple arrows indicate the measurement process performed via optical microscopy from t1 to t2. Green arrows represent calculations using multiple injected species within the microfluidics device to obtain concentration profiles at t1, as well as computation of the saturation index to predict precipitation locations and growth directions at t2. a The workflow begins with image acquisition using optical microscopy. Representative RGB images acquired at t1 (b) and t2 (e) are shown for the chip with a uniform pattern (noted as “uniform") and the chip with a rock pattern (noted as “rock"). c At t1, the acquired RGB images are processed by the toolbox to predict multiple species concentrations; exemplary Sr2+ and \({{{{{\rm{SO}}}}}_{4}}^{2-}\) concentrations for the chip with a uniform pattern. d These predicted concentrations are then used to compute the saturation index, enabling prediction of growth directions (chip with uniform pattern) and identification of regions with a high probability of precipitation (chip with rock pattern). e The predictions are validated against the RGB images acquired at t2.
The saturation index (SI) is defined as69
where Q is the ion activity product of the species involved in dissolution or precipitation, Ksp is the solubility product constant of the corresponding solid. SI > 0 indicates supersaturation and a tendency for precipitation, SI = 0 denotes equilibrium between solid and solution, and SI < 0 denotes undersaturation. Given that species concentrations cannot be directly measured, a prediction produced by the toolbox is considered correct if the predicted region with SI > 0 at t2 corresponds to newly precipitated crystals or to the growth of crystals already present at t1.
Overview of the framework
We rely on the non-intrusive reduced basis (NI-RB) method as the key approach, as this method guarantees accuracy and efficiency (Fig. 2)50,53,54,55,63,71. There are two challenges in the implementation: (1) the high-dimensional nature of RGB experimental images requires a large number of training samples for the NI-RB model to achieve accurate predictions, and (2) the reliance on manual segmentation limits the availability of training data, thereby reducing model performance. We address these challenges by pre-processing the RGB experimental images and applying dimensionality reduction prior to input into the NI-RB model, using two different machine learning methods: the U-Net and the CAE (see Fig. 2).
The black dashed line separates two measurement times, denoted as t1 and t2. Purple arrows indicate the measurement process, and green arrows represent calculations within the toolbox. a At time t1, the workflow begins with an RGB experimental image acquired via optical microscopy. b This image is passed to a U-Net to generate a binary segmentation. c The segmented image is then compressed by the encoder of a Convolutional Autoencoder (CAE) into a vector of latent parameters, β, which is concatenated with the time. The CAE decoder is also used to augment training samples for the non-intrusive reduced basis (NI-RB) method. The concatenated vector is fed into the NI-RB model to predict the distribution functions of the Lattice Boltzmann simulations. d These predicted distribution functions are post-processed to obtain multiple species concentrations at t1. e The saturation index is computed to identify locations with a high probability of precipitation at t2. f Locations with saturation index (SI) above zero, indicating a high probability of precipitation, are validated against the newly acquired RGB image at t2. This workflow is then repeated for subsequent time steps.
First, the U-Net (see Fig. 2 for illustration and “Methods” section for mathematical formulation) performs fast segmentation of RGB experimental images, converting them into binary images. It is trained on a limited dataset comprising RGB experimental images from optical microscopy and their manually segmented counterparts, as the generation of these training samples is labor-intensive. The U-Net is specifically selected for this task as its encoding-decoding architecture with skip connections, which allows it to achieve accurate segmentation even with a limited number of training samples, prior to augmentations72.
Next, the segmented binary images (from the same dataset used for training the U-Net model) are used to train a CAE model (see Fig. 2 for illustration and “Methods” section for mathematical formulation). This model serves two key purposes: the encoder reduces the dimensionality of binary image inputs before going into the NI-RB model, while the decoder generates additional binary images to augment the training samples for the NI-RB model training. This method helps overcome the challenge of limited training samples for building the NI-RB model.
The multi-component Lattice–Boltzmann simulations are then performed on the binary images generated through the trained CAE’s decoder. The encoded binary images, time, and corresponding solutions of Lattice–Boltzmann simulations form the training dataset for constructing the NI-RB model (see Fig. 2 for illustration and “Methods” section for mathematical formulation). The NI-RB model enables fast predictions of distribution functions, which in turn facilitate fast predictions of concentration, effective diffusivity, and saturation index (SI) (see Fig. 2), owing to its architecture of only two dense layers with a small number of neurons in each layer50,53,54,55,63,71. The number of NI-RB models constructed corresponds to the number of injected species and the tracer. In our experiments, injecting SrCl2 and Na2SO4 results in four ionic species and one tracer, and therefore five NI-RB models. While only a single U-Net and a single CAE model are required. Finally, the predicted multiple species and tracer concentrations can be used to probe the microfluidics experiments, providing a quantitative understanding of the experimental results and the prediction of locations with a high probability of precipitation.
Note that this framework is general and can be applied to map images and physical parameters to a variety of chemical and transport states, as well as effective properties. By transforming different physical and chemical processes into Boltzmann transport form, diverse state variables can be derived. However, the output of the NI-RB model remains the distribution functions. This ensures that the overall structure of the toolbox remains unchanged, regardless of the specific chemical or transport processes being modeled through the Lattice–Boltzmann simulations.
Chip with rock pattern case
In this case, the collected experimental results were randomly split into training, validation, and prediction samples, with five images used for training, three for validation, and one for prediction. Note that this original dataset is used to train only the U-Net and the CAE models. It is subsequently expanded via the trained CAE model to generate a larger dataset for training the NI-RB model. This larger dataset has a size of 450, randomly split into 300 training and 100 validation samples.
We applied our toolbox to a prediction sample from the chip with rock pattern case, as shown in Fig. 3. The optimum hyperparameters for the U-Net model are listed in Table 1. Using this optimum model, the prediction achieves a Jaccard index of 0.9933, indicating a good agreement between the predicted and reference segmentations. This accuracy is further supported by the difference image, which reveals only a few scattered red pixels (see the first row of Fig. 3). The U-Net model generated this prediction in 0.84 s on a Dell Latitude laptop equipped with 32 GB RAM and a 13th Gen Intel i7-1365U processor (12 CPUs). Training was also efficient, requiring only 24 min for a single model on an NVIDIA A100 GPU with 512 GB RAM using the JURECA high-performance computing (HPC) infrastructure.
CAE denotes the Convolutional Autoencoder, and NI-RB denotes the non-intrusive reduced basis method. Red pixels in the images shown in the U-Net and CAE sections indicate the differences between the reference and the predicted images. The blue frame marks the image boundaries. a RGB image obtained via optical microscopy. b U-Net section showing the reference image, the U-Net prediction, and their difference (left to right). c CAE section showing the reference image, the CAE prediction, and their difference (left to right). d NI-RB tracer section showing the reference solution, the predicted solution, and the absolute error (left to right).
For the CAE model, we found that the same encoder depth as in the U-Net model was sufficient. For the latent space, we used a dense layer with 16 neurons. The optimal hyperparameters, including learning rate, batch size, and number of epochs, are reported in Table 1. The CAE results, shown in the second row of Fig. 3, demonstrate acceptable accuracy (a Jaccard index of 0.9867), especially considering the complexity of optimizing both architecture and hyperparameters with a limited number of training samples. When the trained decoder can reconstruct the input images, this indicates that the latent space can accurately parameterize any images, bounded by the variability present in the training set73,74,75.
With the selected architecture and hyperparameters, the trained CAE delivered a prediction in 0.17 s using a CPU on a Dell Latitude laptop. The training process required 11.8 min (for a single model) on a GPU within the JURECA HPC infrastructure.
Estimating effective diffusivity
Once fast segmentation using the U-Net model and dimensionality reduction using the CAE model are established, the resulting reduced (latent) parameters can be used directly as inputs to the NI-RB models. Importantly, the same reduced parameters are used for both tracer and multiple species concentrations prediction without any need to retrain or modify the U-Net or the CAE. Only the NI-RB model changes, depending on which concentration is being predicted.
The training samples for constructing the NI-RB model for tracer concentration and effective diffusivity estimation are obtained using Lattice–Boltzmann simulations, each requiring 24 h on a single CPU core on the JURECA HPC infrastructure. In total, this amounts to 10,800 CPU core-hours. The Lattice–Boltzmann simulations employ a D3Q7 setup, where the degrees of freedom per simulation are 7 (number of Qs) × 20 (number of timesteps) × 512 (image pixel width) × 512 (image pixel height) = 36,700,160.
We applied a cutoff of 1 × 10−5 during the Proper Orthogonal Decomposition step, resulting in 250 basis functions. This cutoff was selected based on a sensitivity analysis of the cutoff with respect to the NI-RB model estimation error (see Supplementary Fig. S1), which showed no further significant accuracy improvement upon increasing the number of basis functions. Moreover, at larger cutoffs, the NI-RB models were unable to capture geometry changes due to chemical reaction. With the chosen cutoff 1 × 10−5, the resulting L2 estimation error is 1 × 10−3, which is sufficient for this case. The optimal hyperparameters are listed in Table 1. The trained NI-RB model can predict a new sample in 0.14 s using a CPU on a Dell Latitude laptop, corresponding to a speedup of more than 6 × 105 compared to the full Lattice–Boltzmann simulation. Training is also efficient, requiring only 2.56 min per model on a CPU within the JURECA HPC infrastructure.
Furthermore, we estimated the effective diffusivity of the prediction case using the distribution functions predicted by the NI-RB model. As shown in Fig. 3, the predicted effective diffusivity is close to the reference values. The complete workflow, from an RGB experimental image to the effective diffusivity and tracer concentration profile, takes in total 3.81 s on a CPU in a Dell Latitude laptop. This prediction time is shorter than the experimental image acquisition interval of 10 min, enabling evaluation of tracer diffusion direction and effective diffusivity alongside the experiments.
Predicting locations with a high probability of precipitation
We trained four additional NI-RB models to predict the concentrations of Sr2+, Cl−, Na+, and \({{{{{\rm{SO}}}}}_{4}}^{2-}\) based on multi-component Lattice–Boltzmann simulations that mimic the experimental setup (see Multi-component Lattice–Boltzmann simulation subsection). Compared to the tracer simulation, more time steps were required to achieve similar accuracy and convergence, resulting in an output dimensionality of 7 (number of Qs) × 300 (number of timesteps) × 512 (image pixel width) × 512 (image pixel height) = 550,502,400 degrees of freedom for each model.
We applied the same Proper Orthogonal Decomposition cutoff of 1 × 10−5 to all NI-RB models, resulting in 321, 325, 330, and 354 basis functions for the Sr2+, Cl−, Na+, and \({{{{{\rm{SO}}}}}_{4}}^{2-}\) models, respectively. The cutoff was selected following the same rationale as for the NI-RB Tracer model (see Supplementary Fig. S4 for the spectra decay profile for each model). The resulting L2 estimation errors and the optimal hyperparameters are provided in Table 1. Each trained NI-RB model predicts a new sample in 0.003 s on a CPU (Dell Latitude laptop), corresponding to a speedup of more than 9 × 104 relative to the full Lattice–Boltzmann simulation. Training is also computationally efficient, requiring only 14–15 min per model on a CPU within the JURECA HPC infrastructure.
The prediction case is treated as a future event that we aim to forecast using the existing training samples. To do so, we selected an experimental image recorded 10 min prior to the prediction target (Fig. 4). This image serves as the simulation domain. Fast segmentation using the U-Net model and dimensionality reduction using the CAE model were then applied, as shown in the first row of Fig. 4. After performing chemical equilibrium calculations, the red regions (SI > 0) in Fig. 4 indicate locations where precipitation is thermodynamically favored and thus likely to occur in the next experimental frame, 10 min later. These predicted regions correspond closely to the actual locations where precipitation or crystal growth is subsequently observed.
Green arrows denote calculations performed within the toolbox. The dashed red lines mark the locations where crystals are growing. a RGB image obtained via optical microscopy 10 min prior to the prediction image and used as in the training samples. b Segmentation result from the U-Net. c Predicted multiple species concentrations obtained using the non-intrusive reduced basis method. d Post-processed species concentrations identifying locations with a high probability of precipitation. e RGB image acquired via optical microscopy at the prediction time, confirming precipitation at the predicted locations.
Chip with uniform pattern case
This case differs from the previous one because the RGB experimental images contain noticeable noise (see Fig. 5). We explicitly included this noise in the training dataset so that the machine learning models can learn to recognize and handle it. In this case, four images were used for training, two for validation, and one for prediction. This dataset is then expanded via the trained CAE model to generate a larger dataset for training the NI-RB model. This larger dataset has a size of 150, randomly split into 100 training and 50 validation samples.
CAE denotes the Convolutional Autoencoder, and NI-RB denotes the non-intrusive reduced basis method. Red pixels in the images shown in the U-Net and CAE sections indicate the differences between the reference and the predicted images. The blue frame marks the image boundaries. a RGB image obtained via optical microscopy. b U-Net section showing the reference image, the U-Net prediction, and their difference (left to right). c CAE section showing the reference image, the CAE prediction, and their difference (left to right). d NI-RB tracer section showing the reference solution, the predicted solution, and the absolute error (left to right).
We applied our toolbox to a prediction sample, as shown in Fig. 5. The optimal hyperparameters for the U-Net model are listed in Table 2. Using this optimized model, the prediction achieved a Jaccard index of 0.9803, indicating strong agreement between the predicted and reference segmentations. This is further supported by the difference image, which shows only a few scattered red pixels (see the first row of Fig. 5). The U-Net produced this prediction in 0.51 s on a Dell Latitude laptop with 32 GB RAM and a 13th Gen Intel i7-1365U processor (12 CPUs). Training was also efficient, requiring only 21 min for a single model on an NVIDIA A100 GPU with 512 GB RAM on the JURECA HPC system.
The CAE model uses the same encoder depth as the U-Net model. The optimal hyperparameters are reported in Table 2. The CAE results, shown in the second row of Fig. 5, demonstrate strong performance with a Jaccard index of 0.9763. The trained CAE generated a prediction in 0.12 s on a Dell Latitude laptop CPU. Training required 24 min (for a single model) on a GPU within the JURECA HPC infrastructure.
Estimating effective diffusivity
For constructing the training samples, we ran Lattice–Boltzmann simulations, each requiring 24 h on a single CPU core on the JURECA HPC infrastructure. In total, this amounts to 10,800 CPU core-hours. The Lattice–Boltzmann simulations employ a D3Q7 setup, where the degrees of freedom per simulation are 7 (number of Qs) × 16 (number of timesteps) × 256 (image pixel width) × 256 (image pixel height) = 9,175,040.
We applied a cutoff of 1 × 10−5 during the Proper Orthogonal Decomposition step, resulting in 88 basis functions. The cutoff was selected following the same rationale as in the chip with rock pattern case. Compared with the chip with a rock pattern case, the NI-RB tracer model here uses fewer basis functions with the same cutoff, reflecting the simpler geometry changes. Such simple geometry changes do not require a larger number of basis functions. With the cutoff of 1 × 10−5, the L2 estimation error is 6.0 × 10−4, which is sufficient for this case. The optimal hyperparameters are listed in Table 2. The trained NI-RB model can predict a new sample in 0.04 s using a CPU on a Dell Latitude laptop, corresponding to a speedup of more than 1 × 106 compared to the full Lattice–Boltzmann simulation. Training is also efficient, requiring only 4.79 min per model on a CPU within the JURECA HPC infrastructure.
Figure 5 shows that the predicted effective diffusivity is the same as the reference values. The complete workflow, from an RGB experimental image to the effective diffusivity and tracer concentration profile, takes a total of 1.27 s on a CPU in a Dell Latitude laptop. This prediction time is shorter than the experimental image acquisition interval of 10 min, enabling evaluation of tracer diffusion direction and effective diffusivity alongside the experiments.
Predicting locations with a high probability of precipitation
We trained four additional NI-RB models to predict the concentrations of Sr2+, Cl−, Na+, and \({{{{{\rm{SO}}}}}_{4}}^{2-}\) based on multi-component Lattice–Boltzmann simulations that mimic the experimental setup (see Multi-component Lattice–Boltzmann simulation subsection). The same Proper Orthogonal Decomposition cutoff of 1 × 10−5 (following the same rationale as in the chip with rock pattern case) is applied to all NI-RB models, resulting in 94, 95, 131, and 144 basis functions for the Sr2+, Cl−, Na+, and \({{{{{\rm{SO}}}}}_{4}}^{2-}\) models, respectively. The resulting L2 estimation errors and optimal hyperparameters are summarized in Table 2. Each trained NI-RB model predicts a new sample in 0.002 s on a CPU (Dell Latitude laptop), corresponding to a speedup of more than 1 × 105 relative to the full Lattice–Boltzmann simulation. Training required only 5–7 min per model on a CPU within the JURECA HPC infrastructure.
The prediction case is treated as a future event that we aim to forecast using the existing training samples. To do so, we selected an experimental image recorded 10 min prior to the prediction target (Fig. 6). This image serves as the simulation domain. After performing chemical equilibrium calculations, the red regions (SI > 0) in Fig. 6 indicate locations where precipitation is thermodynamically favored and thus likely to occur in the next experimental frame, 10 min later. These predicted regions correspond closely to the actual directions where crystal growth is subsequently observed.
Green arrows denote calculations performed within the toolbox. The dashed red lines mark the locations where crystals are growing. a RGB image obtained via optical microscopy 10 min prior to the prediction image and used as in the training samples. b Segmentation result from the U-Net. c Predicted multiple species concentrations obtained using the non-intrusive reduced basis method. d Post-processed species concentrations identifying locations with a high probability of precipitation. e RGB image acquired via optical microscopy at the prediction time, confirming crystal growth at the predicted locations.
Discussion
Our toolbox ensures process explainability at each step, enabling error identification and user supervision. This design prevents any inaccuracies originating from the data-driven components from being misinterpreted as physical anomalies. Instead, the toolbox enables users to trace the source of errors, facilitating targeted improvements and promoting reliable, physically consistent interpretation of experimental results. With this explainability, the predicted concentration can be confidently used as a probe for experiments. Furthermore, this toolbox will enable experimentalists to tune the experimental conditions and analyze processes on-the-fly.
The toolbox delivers accurate predictions of concentration and locations with a high probability of precipitation using a limited number of experimental results. It is due to the integrated use of the U-Net, the CAE, and the NI-RB methods. The U-Net architecture, with its skip connections, enables fast and accurate segmentation from just a few RGB experimental images, minimizing the labor effort required for manual segmentation72. The CAE model addresses the challenge of learning accurate mappings from images and scalar parameters to chemical and transport states by reducing the dimensionality and enabling the generation of a large number of synthetic training samples. Although the CAE can generate many such samples, the NI-RB model maintains computational efficiency by requiring only a relatively small number for training. Moreover, the NI-RB method maintains the structure of the physical problem within its basis functions, allowing accurate predictions with a small number of training samples52,53,55,63,76.
There are other methods, such as Karhunen–Loeve and the Active Subspaces method, that are commonly used for dimensionality reduction71,77,78,79, serving the same purpose as the CAE model. However, these methods are not efficient when applied to complex images78,80. In contrast, models like the generative adversarial network (GAN), the Variational Autoencoder (VAE), and the Variational Autoencoder-Generative Adversarial Network (VAE-GAN) methods have been used for reconstruction of porous media images (can also facilitate dimensionality reduction), which aligns with our objective67,81,82. These methods typically require a large number of training samples and substantial computational resources. Thus, we adopt the CAE model by carefully selecting the architecture to operate effectively with a limited number of training samples and reasonable computational resources.
The predictive capability of the toolbox is inherently limited to cases similar to those represented in the training dataset. This limitation stems from the data-driven nature of both the U-Net and CAE models, which require substantial variability in the training samples. Achieving wide coverage of possible prediction cases demands significant labor effort to manually segment images, which is necessary to increase the variability of the training samples.
The number of basis functions in the NI-RB model is relatively high compared to their typical use in previous studies50,51,53,54,63. This is because the model must capture geometric variations in every realization. In addition, the presence of small precipitates requires higher accuracy, further increasing the number of basis functions needed.
The Lattice–Boltzmann-based toolbox developed in this work offers a highly adaptable framework for simulating micro-scale transport phenomena. A key advantage lies in its structural consistency. As long as the governing equations of processes of interest can be formulated within the Boltzmann transport framework, the core implementation remains largely unchanged. This stands in contrast to classical numerical methods (such as the finite volume or finite element methods), which often require substantial code modification when extended to new physical and chemical processes. As a result, this toolbox is a flexible and transferable platform that can accommodate a wide range of phenomena with minimal restructuring.
For the current saturation index post-processing, we do not employ a surrogate model because the chemical system is relatively simple and the equilibrium calculations are fast (approximately 6.9 × 10−4 s per calculation on a single CPU core of a Dell laptop) and inexpensive. However, for more complex chemical systems (or systems involving chemical kinetics) and for larger image sizes, a surrogate model becomes necessary to maintain computational efficiency30. In future work, we plan to integrate a chemical surrogate model into our physics-based machine learning framework to further accelerate the chemical equilibrium calculations and, consequently, the saturation index predictions.
Because multiple machine learning models are applied sequentially, errors can propagate from the U-Net model through the CAE and ultimately to the NI-RB model, potentially affecting the final outputs. For example, in the chip with uniform pattern case, the CAE model mispredicts the geometry of the precipitate in the right corner (see Fig. 5). This misprediction artificially blocks species diffusion from the bottom boundary that would otherwise propagate toward the right side of the domain. As a result, the predicted precipitation location slightly underestimates growth in the right corner. Nevertheless, the framework still correctly captures the location with a high probability of precipitation in the left corner. Therefore, the propagated error remains acceptable and does not significantly alter the overall interpretation of the results.
Future developments will focus on extending the capability of our framework to handle advective and fast reaction regimes, where strong coupling between the Lattice–Boltzmann solver and the chemical module becomes essential. This advancement is particularly important because fully coupled chemical calculations and Lattice–Boltzmann simulations are computationally expensive and therefore not feasible to run side-by-side with experiments. Looking further ahead, with optimized training procedures, the toolbox could evolve into a versatile platform capable of rapidly predicting a broad spectrum of physicochemical processes amenable to Lattice–Boltzmann modeling (e.g., multiphase flow, heat transfer, electroosmosis, electrophoresis). Although the use of the NI-RB method within our framework is promising, the presence of multiple physical processes, strongly coupled with chemical processes, may give rise to shock fronts and discontinuities, which would require additional treatment to maintain the accuracy and efficiency of the NI-RB method83,84,85,86. Such developments would represent a major step toward a unified, fast, physics-based simulation framework for lab-on-a-chip systems, with the potential to accelerate experimental design, optimization, and discovery.
Our framework will also be extended to assist in-situ synchrotron-based experiments on 4-D porous structures. This capability is particularly important because synchrotron beam time is limited and the beam field of view is often smaller than the size of the porous structure being imaged87,88. When imaging nucleation or crystal growth events in porous structures, it is highly beneficial to identify potential locations of these events in advance so that the beam can be directed to the most relevant regions. This targeted imaging strategy can help avoid inefficient use of beam time and reduce the need to repeat experiments multiple times. The underlying idea is to enable fast spatio-temporal concentration predictions using our framework. For this purpose, the U-Net and CAE models would need to be extended to accommodate three-dimensional images through the use of 3D convolutions, while the NI-RB model would need to handle significantly larger input and output dimensionalities. As a consequence, a larger number of training samples would be required. This extension would become even more challenging in the presence of multiphysics coupling, fast chemical reactions, and hysteresis.
Methods
Microfluidics experimental and numerical modeling setup
Experimental setup
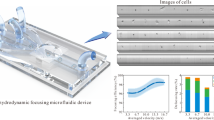
Figure 7 presents experimental and numerical setups for both chips with rock and a uniform pattern. In both experiments, a solution of 50 mM SrCl2 and 50 mM Na2SO4 was injected into the chip inlet using a syringe pump (Fig. 7a), at flow rates of 50 μL/min for the chip with rock pattern (Fig. 7b) and 1000 nL/min for the chip with uniform pattern (Fig. 7c), creating a parallel flow of SrCl2 and Na2SO4, as illustrated in Fig. 73. The temperature is maintained at 25 ∘C and pH 5.63. This parallel flow drove diffusion of SrCl2 and Na2SO4 toward the center of the chip, where mixing and reaction occurred, resulting in SrSO4 precipitation9,89. In the snapshot area, the system was dominated by diffusion, with flux arising perpendicular to the parallel flow9,89.
Red arrows denote the flow of SrCl2, blue arrows denote the flow of Na2SO4, and purple arrows indicate the observation process. a Experimental setup consisting of an optical microscope, a syringe pump injecting 50 mM SrCl2 and 50 mM Na2SO4 at 50 μL/min for the chip with rock pattern and 1000 nL/min for the chip with uniform pattern, the microfluidic devices, and the outflow3. b Transport of SrCl2 and Na2SO4 within the borosilicate glass chip, referred to as the chip with rock pattern. The dashed box marks the region where SrCl2 diffuses from the top and Na2SO4 from the bottom. c Transport of SrCl2 and Na2SO4 within the polydimethylsiloxane (PDMS) chip, referred to as the chip with uniform pattern. The dashed box marks the region where SrCl2 diffuses from the top and Na2SO4 from the bottom. d Preparation for Lattice–Boltzmann simulations, in which images of the snapshot region are acquired at multiple time steps and subsequently segmented. e For each segmented image, a multi-component Lattice–Boltzmann simulation is performed using boundary conditions consistent with the experimental setup, and f A Lattice–Boltzmann simulation with a conservative tracer is also performed using fixed boundary values of 1.0 mM at the top and 0.0 mM at the bottom, from which effective diffusivity is derived. The same numerical setup is applied to the PDMS experiments.
Monitoring of the evolving geometry was performed using high-resolution optical microscopy and confocal Raman spectroscopy3,69. The optical microscope resolution was approximately 0.6 × 0.6 × 1 μm33,69. In both experiments, images capturing the geometric evolution were taken every 15 min and 10 min, respectively3. Further experimental details can be found in Lönartz et al.3.
Numerical modeling setup
We performed two types of simulations under a diffusive regime: a multi-component Lattice–Boltzmann simulation for describing the experiments and a Lattice–Boltzmann simulation with a conservative tracer for estimating effective diffusivity. We assumed that chemical reactions occur more slowly than the transport processes30,68,69,70. This means that the physical time follows the time when an image is captured by the optical microscope, and the transport simulation is always performed until stationary for each image30,69,70. A comprehensive discussion about this assumption is presented in Molins et al.68 and Lichtner70.
For the multi-component simulations as shown in Fig. 7e, we used four components that affect the formation of SrSO4 precipitates: Sr2+, Cl−, Na+, and \({{{{{\rm{SO}}}}}_{4}}^{2-}\). Note that, throughout the experiment, the pH is constant at 5.6; hence, the H+ ion is not included in the transport equation. The same boundary conditions as in the experimental setup are used at the top and at the bottom part of the snapshot area. In a chip with a rock pattern, a constant injection of SrCl2 at the top creates a constant supply of 50 mM Sr2+ and 100 mM Cl− ions, while a constant injection of Na2SO4 at the bottom creates a constant supply of 100 mM Na+ and 50 mM \({{{{{\rm{SO}}}}}_{4}}^{2-}\) ions. These translate into Dirichlet boundary conditions 50 mM Sr2+ and 100 mM Cl− at the top, and 100 mM Na+ and 50 mM \({{{{{\rm{SO}}}}}_{4}}^{2-}\) at the bottom. The same condition and numerical setting are also applied to the chip with a uniform pattern case.
For the tracer simulation shown in Fig. 7f, we used a conservative tracer which did not trigger any chemical reactions or change the geometry. The boundary conditions for this simulations are of the Dirichlet type for both top and bottom boundary.
There are two bottleneck tasks here: image segmentation and Lattice–Boltzmann simulation. Image segmentation is a labor-intensive task because microfluidics experimental images are often complex, making them challenging to be automatically labeled using existing algorithms90,91,92. The Lattice–Boltzmann simulation is computationally expensive and requires a significant amount of time when performed on complex structures33,93,94.
Multi-component Lattice–Boltzmann simulation
The Lattice–Boltzmann simulations solve for spatio-temporal multi-component and tracer concentration profiles within the microfluidics devices3,27,30. Considering a system with pure diffusion, the governing equation of mass transport is3,30:
where J [mol/m2/s] is the mass flux and D [m2/s] is the mass diffusivity. The subscription i denotes the component, such as Sr2+, Cl−, Na+, \({{{{{\rm{SO}}}}}_{4}}^{2-}\), and tracer. We used 1.56 × 10−9 [m2/s] for the diffusivity of Sr2+, 1.31 × 10−9 [m2/s] for the diffusivity of Cl−, 1.93 × 10−9 [m2/s] for the diffusivity of Na+, 8.46 × 10−10 [m2/s] for the diffusivity of \({{{{{\rm{SO}}}}}_{4}}^{2-}\), and 2.2 × 10−9 [m2/s] for the diffusivity of tracer. The Lattice–Boltzmann simulation solves for k distribution functions fk, depending on the scheme employed95. For both the rock and uniform patterns in our case, we used the D3Q7 scheme3. The governing equations for the Lattice–Boltzmann simulation for each component can be expressed as follows95:
where r [m] denotes the position vector, e is the unit discrete velocities, δx [m] is the grid size, δt,D [s] is the time step, cD = δx/δt,D [m/s] is the lattice speed, and S is the source term (in our case S = 0). τD [-] is the dimensionless relaxation time, given by τD = 4D/cDδx + 0.5. The subscript k = 0, 1, 2, …, 6 corresponds to the directions in the Q = 7 scheme (D3Q7). The superscript eq denotes equilibrium distribution functions. The equilibrium distribution functions are defined as follows33,95:
with
The local concentration of component i, Ci(r, t) is calculated by Ci = ∑kfk. The local flux along the x direction is calculated through fk:
In our case, the fluid-solid interfaces, such as fluid-base solid and fluid-precipitate interfaces, are set as the zero normal flux boundary condition. The conventional bounce-back rule in the Lattice–Boltzmann method is therefore employed.
For the tracer simulation, when the simulation reaches a stationary, the effective diffusivity is determined through the following equation:
where ∑Jx/S is the flux through the unit cross-section of the chip, and L is the length of the chip. Our setup for the Lattice–Boltzmann simulation is illustrated in Fig. 7.
U-Net
The U-Net model is a machine learning model characterized by a symmetrical architecture of interconnected encoders and decoders72. The input to the model for the chip with rock pattern case is an image with dimensions of 512 × 512 and three channels (RGB), while the output is a binary image with dimensions of 512 × 512 pixels and a single channel (see Fig. 8). For the chip with uniform pattern case, input and output dimensions of 256 × 256 pixels are used. Each encoder comprises a 2 × 2 two-dimensional max pooling layer, followed by two 3 × 3 two-dimensional convolutional layers, each succeeded by a batch normalization layer and a Rectified Linear Unit (ReLU) activation function. Note that no padding is applied in the convolutional layers, which requires the input image dimensions (width and height) to be a factor of two. Each encoder downsamples the input image by a factor of two and doubles the number of output channels. Consequently, the number of filters in the convolutional layers doubles at each subsequent encoder stage.
Black arrows indicate the information flow from an input RGB image (acquired using optical microscopy) to the corresponding binary image used later for the Lattice–Boltzmann simulations. The architecture shown corresponds to the chip with a rock pattern chip with an input size of 512 × 512 pixels. A similar architecture is used for a chip with a uniform pattern, with an input size of 256 × 256 pixels.
Each decoder consists of two 3 × 3 two-dimensional transposed convolutional layers, each succeeded by a batch normalization layer and a Rectified Linear Unit (ReLU) activation function, and a 2 × 2 two-dimensional up-sampling layer. Each decoder is connected to its corresponding encoder through a concatenate operation (skip connection), which transfers information from the second convolutional layer of the encoder to the first transposed convolutional layer of the decoder. Following the operation in its encoder counterpart, each decoder performs upsampling by a factor of two and halves the number of channels. As a result, the number of filters in the convolutional layers is halved at each subsequent decoder stage. The final block consists of a 3 × 3 two-dimensional convolutional layer and a sigmoid layer, producing a binary image based on probability values. Probability values greater than 0.5 are classified as 1.0 (solid), while values less than or equal to 0.5 are classified as 0.0 (pore space). For a detailed mathematical formulation of each operation, readers are encouraged to visit Bishop and Nasrabadi96.
An optimum U-Net model is constructed in a two-loop approach:
-
The inner loop. This inner loop focuses on obtaining optimum weights and biases through minimizing the binary cross-entropy loss between predictions and ground truth on training samples:
$${\min}_{{{{\bf{W}}}},{{{\bf{b}}}}}{L}_{\log },{{{\rm{tr}}}}=-\frac{1}{{N}_{{{{\rm{tr}}}}}}\displaystyle {\sum }_{i=1}^{{N}_{{{{\rm{tr}}}}}}\displaystyle {\sum }_{k=1}^{K}{y}_{i,k}\log Pr\,{({y}_{i,k}=1)}_{i,k},$$(8)where W is the weights of a U-Net model, b is the biases of a U-Net model, Ntr denotes the number of training samples, K denotes the image dimension, y is the value of the true label (in this case, either 0 or 1), and Pr denotes the predicted probability of having yi,k = 1. The subscription log denotes logarithmic form, tr denotes training samples, i denotes the enumeration for the training samples, and k denotes the enumeration for image dimension.
-
The outer loop. This outer loop focuses on selecting optimum architecture and hyperparameters through minimizing the binary cross-entropy loss between predictions and ground truth on validation samples:
$${\min}_{{{{\boldsymbol{\theta }}}}}{L}_{\log ,{{{\rm{va}}}}}=-\frac{1}{{N}_{{{{\rm{ts}}}}}}\displaystyle {\sum }_{i=1}^{{N}_{{{{\rm{va}}}}}}\displaystyle {\sum }_{k=1}^{K}{y}_{i,k}\log Pr\,{({y}_{i,k}=1)}_{i,k}$$(9)gg where θ is the hyperparameters and parameterized architecture of a U-Net model. The subscription va denotes validation samples. To solve Eq. (9), we use a random search algorithm96,97 and select the model with the lowest binary cross-entropy loss as the optimal model.
The hyperparameters to be optimized include the learning rate, batch size, and number of epochs, while the architectural parameters are the encoder depth and the number of filters in the first encoder layer. The U-Net architecture is parameterized using the encoder depth and the number of filters in the first encoder layer98, based on the established principles of the U-Net architecture: each successive encoder reduces the spatial resolution of the input by half while doubling the number of filters, and each successive decoder performs the inverse operation, upsampling by half and halving the number of filters. Thus, the architecture can be fully defined by specifying the number of encoders (which directly determines the number of decoders due to symmetry) and the number of filters in the first encoder. A hundred parallel optimization steps using a random search algorithm96,97 is employed to select the optimum hyperparameters.
We used the TensorFlow Python library99 to construct the U-Net model. Automatic augmentation, including rotation, translation, and flipping, embedded in the generator function of the TensorFlow Python library99, is also applied to enrich the training samples.
Scaling the input and output images is critical for successful training. In our case, scaling is performed by dividing each pixel value in the input and output image arrays by 255, the maximum possible pixel intensity.
Another key consideration is the variability of the training samples besides data augmentation. A trained U-Net cannot generalize to cases beyond those represented in these training samples.
Convolutional autoencoder
The CAE is an encoder-decoder model that uses the same image as both its input and output. The model seeks to replicate the input image while parameterizing it in a latent space, serving as a non-linear dimensionality reduction method78. For a chip with a rock pattern case, the CAE model takes a 512 × 512-pixel binary image generated by the trained U-Net model, attempting to replicate it at the output. While for the chip with a uniform pattern case, the input and output dimensions of the CAE model are 256 × 256 pixels. It is important to note that CAE models are prone to overfitting and can suffer from inefficient training, often resulting in excessive computational resource demands or necessitating a large number of training samples78,100. Careful attention must be given to the input dimensionality and the complexity of the model architecture100. This limitation explains why we do not directly use the RGB images captured by the optical microscope as input and output for the CAE model.
Guided by the U-Net model architecture, we adopt a similar successive downsampling-upsampling approach (see Fig. 9). However, to mitigate overfitting, we simplify the encoder and decoder architecture. Each encoder consists of a 3 × 3 two-dimensional convolutional layer with a stride of 2 and a ReLU activation function, while each decoder comprises a 3 × 3 two-dimensional transpose convolutional layer with a stride of 2 and a ReLU activation function. For a detailed mathematical formulation of each operation, readers are encouraged to visit Padula et al.78 and Bishop and Nasrabadi96.
Black arrows indicate the information flow from an input binary image to an output binary image. The architecture shown corresponds to the chip with a rock pattern chip with an input size of 512 × 512 pixels. A similar architecture is used for a chip with a uniform pattern, with an input size of 256 × 256 pixels.
The key difference from the U-Net lies in the stride used in the convolutional layers. In the U-Net, starting from an RGB image with higher dimensionality, multiple operations within an encoder (two convolutions followed by max pooling) are necessary to extract features effectively, resulting in a large number of parameters. In contrast, the CAE starts with a binary image, allowing us to replace the two convolutions and max pooling with a single convolution with a stride of 2. This approach maintains the same output size while resulting in a smaller number of trainable parameters. The same principle is applied to the decoder design. The design for the number of filters also follows that of the U-Net model.
We construct an optimum CAE model using a two-loop approach:
-
The inner loop. This inner loop focuses on obtaining optimum weights and biases by minimizing the mean square error between predictions and ground truth on training samples:
$${\min }_{{{{\bf{W}}}},{{{\bf{b}}}}}{L}_{{{{\rm{tr}}}}}=\frac{1}{{N}_{{{{\rm{tr}}}}}\times K}{\sum }_{i=1}^{{N}_{{{{\rm{tr}}}}}}{\sum }_{k=1}^{K}{({y}_{i,k}-{\widehat{y}}_{i,k})}^{2},$$(10)where the hat denotes prediction.
-
The outer loop. This outer loop focuses on selecting optimum architecture and hyperparameters through minimizing the binary cross-entropy loss between predictions and ground truth on validation samples:
$${\min }_{{{{\boldsymbol{\theta }}}}}{L}_{{{{\rm{va}}}}}=\frac{1}{{N}_{{{{\rm{va}}}}}\times K}{\sum }_{i=1}^{{N}_{{{{\rm{va}}}}}}{\sum }_{k=1}^{K}{({y}_{i,k}-{\widehat{y}}_{i,k})}^{2},$$(11)To solve Eq. (11), we use a random search algorithm96,97 and select the model with the lowest mean square error as the optimal model.
For hyperparameter optimization, we focus on learning rate, batch size, and number of epochs, as the architecture is optimized separately. Scaling and data augmentation are performed similarly to the ones used in U-Net optimization. A hundred parallel optimization steps using a random grid search algorithm are employed to find the optimum hyperparameters.
The CAE model can be trained either sequentially after the U-Net model or in parallel, as both models share the same training and validation sets. The main challenge lies in selecting optimal hyperparameters while simultaneously determining the optimum encoder-decoder architecture. Our strategy begins with fixing the architecture to facilitate hyperparameter optimization. Once the hyperparameters are tuned, we iteratively refine the architecture to improve the CAE’s accuracy. A recommended starting point is to adopt an architecture similar to that of the trained U-Net model, but with skip connections removed.
Non-intrusive reduced-basis method
The non-intrusive reduced basis (NI-RB) method is a model order reduction (MOR) technique aiming for reducing spatial and temporal degrees of freedom53,54,55. It approximates the solution of a physics-based model as a linear combination of coefficients and basis functions53,54,55. Following the formulation in refs. 101,102,103, the reduced-order model approximates solutions of Lattice–Boltzmann simulation, providing a map from parameters β and time t to aggregated distribution functions \({{{\bf{f}}}}({{{\bf{r}}}},t)\in {{\mathbb{R}}}^{7\times {N}_{t}\times {N}_{r}}\) (stacked from f0 to f6), where Nt is the number of timesteps. In our case, we adopt a non-intrusive formulation, replacing the Galerkin projection with a NN model50,51,52,53,54,55. The mathematical formulation of this NI-RB formulation is as follows:
where r is the reduced dimension, α is the reduced basis coefficients, β is the parameters, and V is the basis functions. The subscription k denotes the enumeration from 0 to the reduced dimension. We further demonstrate in Supplementary Fig. S3 that constructing an NI-RB model that maps to f(r, t) yields more accurate concentration predictions than directly mapping to the spatio-temporal concentration.
This NI-RB method becomes inefficient when dealing with high-dimensional parameters71. Since our input here is a binary image (which is high-dimensional), we connect the NI-RB model with the trained encoder from the CAE model. Therefore, β is the latent space of the CAE model.
We perform Proper Orthogonal Decomposition (via randomized Singular Value Decomposition) on the snapshots of f(r, t) to construct the basis function53,54,55,71. The basis functions are selected from the eigenvectors (after the decomposition) through an energy criterion54:
where λ is the eigenvalue and ϵ is the tolerance. The subscription k denotes the enumeration for eigenvalues. Investigating the Proper Orthogonal Decomposition cutoff with respect to the NI-RB model estimation error is essential to determine the optimal cutoff. In our cases, a cutoff of 1 × 10−5 was applied to all NI-RB models.
Once the basis functions are selected, we construct a NN model to map parameters β and time t to reduced basis coefficients α, see Fig. 10. This mapping can be expressed mathematically as follows96:
where h denotes the tanh activation function. The size of W and b are determined by the number of dense layers and the number of neurons in each dense layer.
Black arrows indicate the information flow from an input binary image to an output binary image. The figure shown corresponds to the chip with a rock pattern chip with an input size of 512 × 512 pixels. A similar construction approach is used for a chip with a uniform pattern, with an input size of 256 × 256 pixels. a Trained decoder from the Convolutional Autoencoder (CAE) model, used to augment existing segmented images for the construction of the non-intrusive reduced basis (NI-RB) model. b Trained encoder from the CAE model, used to reduce the dimensionality of input images. c Combined latent space of the CAE model and time. d Reduced basis coefficients obtained via Proper Orthogonal Decomposition (POD) on the Lattice–Boltzmann solutions. e Basis functions obtained via POD on the Lattice–Boltzmann solutions.
The training for the Neural Network model uses a two-loop approach:
-
The inner loop. This inner loop focuses on obtaining optimum weights and biases through minimizing the mean square error between predicted reduced basis coefficients \(\widehat{{{{\boldsymbol{\alpha }}}}}\) and ground truth α on training samples:
$${\min }_{{{{\bf{W}}}},{{{\bf{b}}}}}{L}_{{{{\rm{tr}}}}}=\frac{1}{{N}_{{{{\rm{tr}}}}}\times r}{\sum }_{i=1}^{{N}_{{{{\rm{tr}}}}}}{\sum }_{k=0}^{r}{({\alpha }_{i,k}-{\widehat{\alpha }}_{i,k})}^{2},$$(15) -
The outer loop. This outer loop focuses on selecting optimum architecture and hyperparameters through minimizing the mean square error between predicted distribution functions \(\widehat{{{{\bf{f}}}}}({{{\bf{r}}}},t)\) and ground truth f(r, t):
$${\min }_{{{{\boldsymbol{\theta }}}}}{L}_{{{{\rm{va}}}}}=\frac{1}{{N}_{{{{\rm{va}}}}}\times 7\times {N}_{t}\times {N}_{r}}{\sum }_{i=1}^{{N}_{{{{\rm{tr}}}}}}{\sum }_{k=1}^{7\times {N}_{t}\times {N}_{r}}{({f}_{i,k}-{\widehat{f}}_{i,k})}^{2},$$(16)
This outer loop error becomes the approximation error of the non-intrusive reduced basis model. The hyperparameters include the number of neurons in each layer of the NN model, learning rate, batch size, and the number of epochs. For hyperparameter optimization, we use the Bayesian Optimization with Hyperband (BOHB) algorithm104. This approach improves the efficiency of the Bayesian optimization algorithm by combining it with the Hyperband algorithm, thereby reducing the number of optimization steps required104.
Once the non-intrusive reduced basis model is constructed, we can use it to calculate spatio-temporal multiple components and tracer concentration in the post-processing step (see Fig. 2) as follows3:
For estimating effective diffusivity, we post-process the predicted tracer concentration using the following formulation3:
where Ctracer,in = 1.0 [mol/m3] is the inlet concentration at the top boundary, Ctracer,out = 0.0 [mol/m3] is the outlet concentration at the bottom boundary, and Nx is the number of lattices in the direction of diffusion (For chip with rock pattern case, it is 512, while for chip with uniform pattern case, it is 256).
An important aspect of the training process is scaling the inputs and outputs, as proper scaling ensures robust model performance. We apply standard scaling to the inputs and minmax scaling to the outputs105.
Another critical aspect is the sampling process. The training samples must be representative of the entire range of parameters. We employ the Latin Hypercube Sampling method106 to sample the latent space and subsequently generate geometries for the Lattice–Boltzmann simulations.
The number of NI-RB models constructed corresponds to the number of transported components, plus an additional model for the tracer. Although each model receives the same set of input parameters, it predicts a different output response. Importantly, the models can be trained in parallel, as they are independent of one another.
Predicting locations with a high probability of precipitation and growth direction
To predict locations with a high probability of precipitation or growth directions, we post-process the stationary multiple species concentrations predicted by the NI-RB models using chemical equilibrium calculations. Chemical equilibrium is computed by minimizing the Gibbs free energy to obtain the equilibrium concentrations of all species, following the formulation described in Leal et al.107. These calculations are performed using the open-source software Reaktoro108.
For our experimental cases, we use the PHREEQC database within Reaktoro together with the WATEQ Debye-Hückel model to compute aqueous species activities109. Reaktoro takes the multicomponent concentrations predicted by the NI-RB models as input and returns the saturation index of SrSO4, which is then used to assess possible precipitation locations or growth directions. Each calculation took 6.9 × 10−4 s.
Regions in which the predicted saturation index of SrSO4 exceeds zero indicate a high probability of newly precipitated SrSO4 crystals or the growth of existing crystals. We use these regions as a criterion to evaluate the predictive accuracy of the toolbox. When the predicted regions cover most of these events, as qualitatively assessed from the subsequent image, the toolbox prediction is considered accurate.
Data availability
The Python scripts used to generate training samples and construct the machine learning models, along with the results for both rock and uniform patterns, are published at https://doi.org/10.5281/zenodo.18768617. Raw simulation data is available upon request.
Code availability
The codes for in this toolbox are parts of open-source Chip Analyzer and Calculator (cac) Python library, published in our Github https://github.com/FZJ-RT/cac.git. These codes are based on several open-source Python libraries, such as Keras, Tensorflow, Horovod, Scikit-learn, OpenCV, bohb-hpo, and Dask. The codes for performing the Lattice–Boltzmann simulation are available in the Github repository https://github.com/FZJ-RT/lbm.git. The detailed implementations for plotting, training, and calculating concentration and effective diffusivity are published in https://doi.org/10.5281/zenodo.18768617.
References
Roman, S., Rembert, F., Kovscek, A. R. & Poonoosamy, J. Microfluidics for geosciences: metrological developments and future challenges. Lab a Chip 25, 4273–4289 (2025).
Scheidweiler, D. et al. Spatial structure, chemotaxis and quorum sensing shape bacterial biomass accumulation in complex porous media. Nat. Commun. 15, 191 (2024).
Lönartz, M. I., Yang, Y., Deissmann, G., Bosbach, D. & Poonoosamy, J. Capturing the dynamic processes of porosity clogging. Water Resour. Res. 59, e2023WR034722 (2023).
Poonoosamy, J. et al. Microfluidic investigation of pore-size dependency of barite nucleation. Commun. Chem. 6, 250 (2023).
Ling, B. et al. Probing multiscale dissolution dynamics in natural rocks through microfluidics and compositional analysis. Proc. Natl. Acad. Sci. USA 119, e2122520119 (2022).
Poonoosamy, J. et al. A lab on a chip experiment for upscaling diffusivity of evolving porous media. Energies 15, 2160 (2022).
Poonoosamy, J. et al. A lab-on-a-chip approach integrating in-situ characterization and reactive transport modelling diagnostics to unravel (Ba, Sr) SO4 oscillatory zoning. Sci. Rep. 11, 23678 (2021).
Steefel, C. I. Reactive transport at the crossroads. Rev. Mineral. Geochem. 85, 1–26 (2019).
Tartakovsky, A. M., Redden, G., Lichtner, P. C., Scheibe, T. D. & Meakin, P. Mixing-induced precipitation: experimental study and multiscale numerical analysis. Water Resour. Res. 44, 1–19 (2008).
Molins, S. & Knabner, P. Multiscale approaches in reactive transport modeling. Rev. Mineral. Geochem. 85, 27–48 (2019).
Maher, K. & Mayer, K. U. Tracking diverse minerals, hungry organisms, and dangerous contaminants using reactive transport models. Elem. Int. Mag. Mineral. Geochem. Petrol. 15, 81–86 (2019).
Prommer, H., Sun, J. & Kocar, B. D. Using reactive transport models to quantify and predict groundwater quality. Elem. Int. Mag. Mineral. Geochem. Petrol. 15, 87–92 (2019).
De Windt, L. & Spycher, N. F. Reactive transport modeling: a key performance assessment tool for the geologic disposal of nuclear waste. Elem. Int. Mag. Mineral. Geochem. Petrol. 15, 99–102 (2019).
Meile, C. & Scheibe, T. D. Reactive transport modeling of microbial dynamics. Elem. Int. Mag. Mineral. Geochem. Petrol. 15, 111–116 (2019).
Lichtner, P. C., Steefel, C. I. & Oelkers, E. H. Reactive Transport in Porous Media. (eds Lichtner, P. C., Steefel, C. I. & Oelkers, E. H.) (De Gruyter, Berlin, Boston, 1996).
Steefel, C. I., DePaolo, D. J. & Lichtner, P. C. Reactive transport modeling: An essential tool and a new research approach for the earth sciences. Earth Planet. Sci. Lett. 240, 539–558 (2005).
Lei, W. et al. Advancing sustainable energy solutions with microfluidic porous media. Lab on a Chip 25, 3374–3410 (2025).
Morais, S. et al. Studying key processes related to CO2 underground storage at the pore scale using high pressure micromodels. React. Chem. Eng. 5, 1156–1185 (2020).
Zhao, B. et al. Comprehensive comparison of pore-scale models for multiphase flow in porous media. Proc. Natl. Acad. Sci. USA 116, 13799–13806 (2019).
Zhao, B., MacMinn, C. W. & Juanes, R. Wettability control on multiphase flow in patterned microfluidics. Proc. Natl. Acad. Sci. USA 113, 10251–10256 (2016).
Porter, M. L. et al. Geo-material microfluidics at reservoir conditions for subsurface energy resource applications. Lab a Chip 15, 4044–4053 (2015).
Lenormand, R., Touboul, E. & Zarcone, C. Numerical models and experiments on immiscible displacements in porous media. J. Fluid Mech. 189, 165–187 (1988).
Musabbir Rahman, R. et al. A novel microfluidic approach to quantify pore-scale mineral dissolution in porous media. Sci. Rep. 15, 6342 (2025).
Nitta, N. et al. Raman image-activated cell sorting. Nat. Commun. 11, 3452 (2020).
Poonoosamy, J. et al. Microfluidic flow-through reactor and 3D Raman imaging for in situ assessment of mineral reactivity in porous and fractured porous media. Lab a Chip 20, 2562–2571 (2020).
Fay, M. E. et al. iclots: open-source, artificial intelligence-enabled software for analyses of blood cells in microfluidic and microscopy-based assays. Nat. Commun. 14, 5022 (2023).
Noiriel, C. & Soulaine, C. Pore-scale imaging and modelling of reactive flow in evolving porous media: Tracking the dynamics of the fluid–rock interface. Transp. Porous Media 140, 181–213 (2021).
Soulaine, C., Maes, J. & Roman, S. Computational microfluidics for geosciences. Front. Water 3, 643714 (2021).
Deng, H., Tournassat, C., Molins, S., Claret, F. & Steefel, C. A pore-scale investigation of mineral precipitation driven diffusivity change at the column-scale. Water Resour. Res. 57, e2020WR028483 (2021).
Prasianakis, N. I. et al. Neural network based process coupling and parameter upscaling in reactive transport simulations. Geochim. et. Cosmochim. Acta 291, 126–143 (2020).
Noiriel, C., Steefel, C. I., Yang, L. & Ajo-Franklin, J. Upscaling calcium carbonate precipitation rates from pore to continuum scale. Chem. Geol. 318, 60–74 (2012).
Molins, S., Silin, D., Trebotich, D. & Steefel, C. Direct pore-scale numerical simulation of precipitation and dissolution. Mineral. Mag. 75, 1487 (2011).
Yang, Y. et al. Pore-scale modeling of water and ion diffusion in partially saturated clays. Water Resour. Res. 60, e2023WR035595 (2024).
Prasianakis, N., Curti, E., Kosakowski, G., Poonoosamy, J. & Churakov, S. Deciphering pore-level precipitation mechanisms. Sci. Rep. 7, 13765 (2017).
Cavanaugh, J., Whittaker, M. L. & Joester, D. Crystallization kinetics of amorphous calcium carbonate in confinement. Chem. Sci. 10, 5039–5043 (2019).
Jiang, S. & ter Horst, J. H. Crystal nucleation rates from probability distributions of induction times. Cryst. Growth Des. 11, 256–261 (2011).
Lashkaripour, A. et al. Design automation of microfluidic single and double emulsion droplets with machine learning. Nat. Commun. 15, 83 (2024).
Volk, A. A. et al. Alphaflow: autonomous discovery and optimization of multi-step chemistry using a self-driven fluidic lab guided by reinforcement learning. Nat. Commun. 14, 1403 (2023).
Yong, W. P. et al. Multiscale upscaling study for CO2 storage in carbonate rocks using machine learning and multiscale imaging. In Offshore Technology Conference Asia, D021S007R002 (OTC, 2024).
Jiang, F. et al. Upscaling permeability using multiscale X-ray-CT images with digital rock modeling and deep learning techniques. Water Resour. Res. 59, e2022WR033267 (2023).
Elmorsy, M., El-Dakhakhni, W. & Zhao, B. Generalizable permeability prediction of digital porous media via a novel multi-scale 3D convolutional neural network. Water Resour. Res. 58, e2021WR031454 (2022).
Liu, M., Kwon, B. & Kang, P. K. Machine learning to predict effective reaction rates in 3D porous media from pore structural features. Sci. Rep. 12, 5486 (2022).
Siavashi, J., Najafi, A., Ebadi, M. & Sharifi, M. A CNN-based approach for upscaling multiphase flow in digital sandstones. Fuel 308, 122047 (2022).
Wang, Z. et al. Pore-scale study of mineral dissolution in heterogeneous structures and deep learning prediction of permeability. Phys. Fluids 34, 116609 (2022).
Alqahtani, N. J. et al. Flow-based characterization of digital rock images using deep learning. SPE J. 26, 1800–1811 (2021).
Menke, H. P., Maes, J. & Geiger, S. Upscaling the porosity–permeability relationship of a microporous carbonate for Darcy-scale flow with machine learning. Sci. Rep. 11, 2625 (2021).
Santos, J. E. et al. Computationally efficient multiscale neural networks applied to fluid flow in complex 3D porous media. Transp. Porous Media 140, 241–272 (2021).
Niu, Y., Mostaghimi, P., Shabaninejad, M., Swietojanski, P. & Armstrong, R. T. Digital rock segmentation for petrophysical analysis with reduced user bias using convolutional neural networks. Water Resour. Res. 56, e2019WR026597 (2020).
Wu, H., Fang, W.-Z., Kang, Q., Tao, W.-Q. & Qiao, R. Predicting effective diffusivity of porous media from images by deep learning. Sci. Rep. 9, 20387 (2019).
Santoso, R., Degen, D., Knapp, D., Pechnig, R. & Wellmann, F. Entropy production as a comprehensive indicator to address epistemic uncertainties–a case study of the Hague, Netherlands. In Proc. of the 49th Workshop on Geothermal Reservoir Engineering, 1–10 (Stanford University, 2024).
Degen, D., Cacace, M. & Wellmann, F. 3D multi-physics uncertainty quantification using physics-based machine learning. Sci. Rep. 12, 17491 (2022).
Degen, D. et al. Perspectives of physics-based machine learning for geoscientific applications governed by partial differential equations. Geosci. Model Dev. Discuss. 2023, 1–50 (2023).
Swischuk, R., Mainini, L., Peherstorfer, B. & Willcox, K. Projection-based model reduction: formulations for physics-based machine learning. Comput. Fluids 179, 704–717 (2019).
Wang, Q., Hesthaven, J. S. & Ray, D. Non-intrusive reduced order modeling of unsteady flows using artificial neural networks with application to a combustion problem. J. Comput. Phys. 384, 289–307 (2019).
Hesthaven, J. S. & Ubbiali, S. Non-intrusive reduced order modeling of nonlinear problems using neural networks. J. Comput. Phys. 363, 55–78 (2018).
Karniadakis, G. E. et al. Physics-informed machine learning. Nat. Rev. Phys. 3, 422–440 (2021).
Raissi, M., Yazdani, A. & Karniadakis, G. E. Hidden fluid mechanics: learning velocity and pressure fields from flow visualizations. Science 367, 1026–1030 (2020).
Raissi, M., Perdikaris, P. & Karniadakis, G. E. Physics-informed neural networks: a deep learning framework for solving forward and inverse problems involving nonlinear partial differential equations. J. Comput. Phys. 378, 686–707 (2019).
Wang, S., Teng, Y. & Perdikaris, P. Understanding and mitigating gradient flow pathologies in physics-informed neural networks. SIAM J. Sci. Comput. 43, A3055–A3081 (2021).
Chuang, P.-Y. & Barba, L. A. Predictive limitations of physics-informed neural networks in vortex shedding. Preprint at https://doi.org/10.48550/arXiv.2306.00230 (2023).
Chuang, P.-Y. & Barba, L. A. Experience report of physics-informed neural networks in fluid simulations: pitfalls and frustration https://doi.org/10.48550/arXiv.2205.14249 (2022).
Krishnapriyan, A., Gholami, A., Zhe, S., Kirby, R. & Mahoney, M. W. Characterizing possible failure modes in physics-informed neural networks. Adv. Neural Inf. Process. Syst. 34, 26548–26560 (2021).
Santoso, R., Degen, D., Cacace, M. & Wellmann, F. State-of-the-art physics-based machine learning for hydro-mechanical simulation in geothermal applications. In European Geothermal Congress, 1–10 (European Geothermal Congress, 2022).
Nageswara Rao, B. et al. Convolutional autoencoder-based deep learning for intracerebral hemorrhage classification using brain CT images. Cogn. Neurodyn. 19, 1–13 (2025).
Jun, H. & Cho, Y. Repeatability enhancement of time-lapse seismic data via a convolutional autoencoder. Geophys. J. Int. 228, 1150–1170 (2022).
Delgado, J. M. D. & Oyedele, L. Deep learning with small datasets: using autoencoders to address limited datasets in construction management. Appl. Soft Comput. 112, 107836 (2021).
Shams, R., Masihi, M., Boozarjomehry, R. B. & Blunt, M. J. Coupled generative adversarial and auto-encoder neural networks to reconstruct three-dimensional multi-scale porous media. J. Pet. Sci. Eng. 186, 106794 (2020).
Molins, S. et al. Simulation of mineral dissolution at the pore scale with evolving fluid-solid interfaces: review of approaches and benchmark problem set. Comput. Geosci. 25, 1285–1318 (2021).
Poonoosamy, J. et al. A microfluidic experiment and pore scale modelling diagnostics for assessing mineral precipitation and dissolution in confined spaces. Chem. Geol. 528, 119264 (2019).
Lichtner, P. C. The quasi-stationary state approximation to coupled mass transport and fluid-rock interaction in a porous medium. Geochim. et. Cosmochim. Acta 52, 143–165 (1988).
O’Leary-Roseberry, T., Villa, U., Chen, P. & Ghattas, O. Derivative-informed projected neural networks for high-dimensional parametric maps governed by PDEs. Comput. Methods Appl. Mech. Eng. 388, 114199 (2022).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical image computing and computer-assisted intervention–MICCAI 2015: 18th international conference, Munich, Germany, October 5-9, 2015, proceedings, part III 18, 234–241 (Springer, 2015).
Coscia, D., Demo, N. & Rozza, G. Generative adversarial reduced order modelling. Sci. Rep. 14, 3826 (2024).
Romor, F., Stabile, G. & Rozza, G. Non-linear manifold reduced-order models with convolutional autoencoders and reduced over-collocation method. J. Sci. Comput. 94, 74 (2023).
Jo, H., Pan, W., Santos, J. E., Jung, H. & Pyrcz, M. J. Machine learning assisted history matching for a deepwater lobe system. J. Pet. Sci. Eng. 207, 109086 (2021).
Willcox, K. & Segundo, B. The role of computational science in digital twins. Nat. Comput. Sci. 4, 147–149 (2024).
Li, Y. et al. Subspace enhanced active learning method for high-dimensional reliability analysis. Struct. Multidiscip. Optim. 68, 1–26 (2025).
Padula, G., Romor, F., Stabile, G. & Rozza, G. Generative models for the deformation of industrial shapes with linear geometric constraints: model order and parameter space reductions. Comput. Methods Appl. Mech. Eng. 423, 116823 (2024).
Tezzele, M., Fabris, L., Sidari, M., Sicchiero, M. & Rozza, G. A multifidelity approach coupling parameter space reduction and nonintrusive POD with application to structural optimization of passenger ship hulls. Int. J. Numer. Methods Eng. 124, 1193–1210 (2023).
Almotiri, J., Elleithy, K. & Elleithy, A. Comparison of autoencoder and principal component analysis followed by neural network for e-learning using handwritten recognition. In 2017 IEEE Long Island Systems, Applications and Technology Conference (LISAT), 1–5 (IEEE, 2017).
Zhang, T., Tu, H., Xia, P. & Du, Y. Reconstruction of porous media using an information variational auto-encoder. Transp. Porous Media 143, 271–295 (2022).
Mosser, L., Dubrule, O. & Blunt, M. J. Reconstruction of three-dimensional porous media using generative adversarial neural networks. Phys. Rev. E 96, 043309 (2017).
Pichi, F., Ballarin, F., Rozza, G. & Hesthaven, J. S. An artificial neural network approach to bifurcating phenomena in computational fluid dynamics. Comput. Fluids 254, 105813 (2023).
Discacciati, N., Hesthaven, J. S. & Ray, D. Controlling oscillations in high-order discontinuous galerkin schemes using artificial viscosity tuned by neural networks. J. Comput. Phys. 409, 109304 (2020).
Ray, D. & Hesthaven, J. S. Detecting troubled-cells on two-dimensional unstructured grids using a neural network. J. Comput. Phys. 397, 108845 (2019).
Forti, D. & Rozza, G. Efficient geometrical parametrisation techniques of interfaces for reduced-order modelling: application to fluid–structure interaction coupling problems. Int. J. Comput. Fluid Dyn. 28, 158–169 (2014).
Kim, F. H. et al. Synchrotron 4-dimensional imaging of two-phase flow through porous media. MRS Adv. 1, 2757–2761 (2016).
Berg, S. et al. Real-time 3d imaging of Haines jumps in porous media flow. Proc. Natl. Acad. Sci. USA 110, 3755–3759 (2013).
Zhao, C. et al. Microfluidic experiments of biological CaCO3 precipitation in transverse mixing reactive environments. Acta Geotech. 18, 5299–5318 (2023).
Poonoosamy, J. et al. A radiochemical lab-on-a-chip paired with computer vision to unlock the crystallization kinetics of (Ba, Ra) SO4. Sci. Rep. 14, 9502 (2024).
Guo, K. et al. An artificial intelligence-assisted digital microfluidic system for multistate droplet control. Microsyst. Nanoeng. 10, 138 (2024).
Meldrum, F. C. & O’Shaughnessy, C. Crystallization in confinement. Adv. Mater. 32, 2001068 (2020).
Ansumali, S., Karlin, I., Arcidiacono, S., Abbas, A. & Prasianakis, N. Hydrodynamics beyond Navier-Stokes: exact solution to the Lattice Boltzmann hierarchy. Phys. Rev. Lett. 98, 124502 (2007).
Chen, S. & Doolen, G. D. Lattice Boltzmann method for fluid flows. Annu. Rev. Fluid Mech. 30, 329–364 (1998).
Yang, Y. & Wang, M. Pore-scale modeling of chloride ion diffusion in cement microstructures. Cem. Concr. Compos. 85, 92–104 (2018).
Bishop, C.M. & Nasrabadi, N.M.: Pattern Recognition and Machine Learning vol. 4. (Springer, 2006)
Yang, L. & Shami, A. On hyperparameter optimization of machine learning algorithms: theory and practice. Neurocomputing 415, 295–316 (2020).
Inc., T. M. U-net https://www.mathworks.com/help/vision/ref/unet.html?#responsive_offcanvas. (2024).
Abadi, M. et al. TensorFlow: Large-scale machine learning on heterogeneous systems https://www.tensorflow.org/. Software available from tensorflow.org. (2015).
Ahmed, S. E., San, O., Rasheed, A. & Iliescu, T. Nonlinear proper orthogonal decomposition for convection-dominated flows. Phys. Fluids 33, 121702 (2021).
Koellermeier, J., Krah, P., Reiss, J. & Schellin, Z. Model order reduction for the 1D Boltzmann-BGK equation: identifying intrinsic variables using neural networks. Microfluid. Nanofluidics 28, 16 (2024).
Mohammadi Arani, R., Dehghan, M. & Abbaszadeh, M. Proper orthogonal decomposition–Lattice Boltzmann method: simulating the air pollutant problem in street canyon areas. SIAM J. Sci. Comput. 44, B885–B909 (2022).
Brown, L., Li, J., Calo, V. M., Ghommem, M. & Efendiev, Y. Lattice Boltzmann flow simulations with applications of reduced order modeling techniques. In IPTC 2014: International Petroleum Technology Conference, cp–395 (European Association of Geoscientists & Engineers, 2014).
Falkner, S., Klein, A. & Hutter, F. Combining hyperband and Bayesian optimization. In NIPS 2017 Bayesian Optimization Workshop (NIPS, 2017).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
McKay, M. D., Beckman, R. J. & Conover, W. J. A comparison of three methods for selecting values of input variables in the analysis of output from a computer code. Technometrics 42, 55–61 (2000).
Leal, A. M., Kulik, D. A., Smith, W. R. & Saar, M. O. An overview of computational methods for chemical equilibrium and kinetic calculations for geochemical and reactive transport modeling. Pure Appl. Chem. 89, 597–643 (2017).
Leal, A. M. Reaktoro: An open-source unified framework for modeling chemically reactive systems. https://reaktoro.org (2015).
Truesdell, A. H. & Jones, B. F. WATEQ, a computer program for calculating chemical equilibria of natural waters. J. Res. U.S. Geol. Surv. 2, 233–248 (1974).
Acknowledgements
The main research leading to these results has received funding from the European Research Council through the project GENIES (ERC, grant agreement 101040341). The authors also acknowledge funding from the Helmholtz AI project through the T6 project (ZT-1-PF-5-084). The authors gratefully acknowledge computing time on the supercomputer JURECA [https://doi.org/10.17815/jlsrf-7-182] at Forschungszentrum Jülich under grant no. paj2406, paj2413, tripm, jiek63, and magie.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
R.S. contributes in conceptualization, methodology, software, formal analysis, validation, visualization, and writing. Y.Y. contributes in methodology, software, formal analysis, validation, and writing. M.L. contributes in experimental data. J.P. contributes in conceptualization, data curation, experiments, funding, supervision, and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Physics thanks thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Santoso, R., Yang, Y., Lönartz, M. et al. Physics-based machine learning toolbox for probing concentration under diffusive regime in microfluidics devices. Commun Phys 9, 106 (2026). https://doi.org/10.1038/s42005-026-02590-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42005-026-02590-y