Abstract
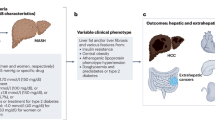
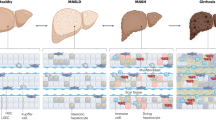
Metabolic-dysfunction-associated steatotic liver disease (MASLD, previously known as non-alcoholic fatty liver disease or NAFLD) is a prevalent and heterogeneous condition affecting nearly 30% of the global population. MASLD is defined as excessive hepatic lipid accumulation with at least one feature of insulin resistance, with potential progression to metabolic dysfunction-associated steatohepatitis, cirrhosis and hepatocellular carcinoma. The disease often coexists with insulin resistance and cardiovascular and chronic kidney diseases. Human genetics has shed light on MASLD predisposition and its causal association with type 2 diabetes and insulin resistance, enabling the field to progress towards precision-medicine therapeutics. Convergent selection of somatic mutations in genes involved in glucose and lipid metabolism in cirrhotic livers suggests adaptive responses to gluco-lipotoxicity that influence end-stage liver disease. Recently, two distinct types of MASLD, with specific clinical trajectories, were identified on the basis of partitioned polygenic risk scores. Future studies are needed to integrate this knowledge, enabling earlier detection, risk stratification and targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Szczepaniak, L. S. et al. Magnetic resonance spectroscopy to measure hepatic triglyceride content: prevalence of hepatic steatosis in the general population. Am. J. Physiol. Endocrinol. Metab. 288, E462–E468 (2005).
Targher, G., Valenti, L. & Byrne, C. D. Metabolic dysfunction-associated steatotic liver disease. N. Engl. J. Med. 393, 683–698 (2025).
Younossi, Z. et al. Global burden of NAFLD and NASH: trends, predictions, risk factors and prevention. Nat. Rev. Gastroenterol. Hepatol. 15, 11–20 (2018).
Ferguson, D. & Finck, B. N. Emerging therapeutic approaches for the treatment of NAFLD and type 2 diabetes mellitus. Nat. Rev. Endocrinol. 17, 484–495 (2021).
Sanyal, A. J. et al. Prospective study of outcomes in adults with nonalcoholic fatty liver disease. N. Engl. J. Med. 385, 1559–1569 (2021).
Younossi, Z. M. et al. The global epidemiology of nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH): a systematic review. Hepatology 77, 1335–1347 (2023).
Israelsen, M. et al. Validation of the new nomenclature of steatotic liver disease in patients with a history of excessive alcohol intake: an analysis of data from a prospective cohort study. Lancet Gastroenterol. Hepatol. 9, 218–228 (2024).
Younossi, Z. M. et al. The economic and clinical burden of nonalcoholic fatty liver disease in the United States and Europe. Hepatology 64, 1577–1586 (2016).
Rinella, M. E. et al. A multisociety Delphi consensus statement on new fatty liver disease nomenclature. J. Hepatol. 79, 1542–1556 (2023).
Mancina, R. M. et al. The MBOAT7-TMC4 variant rs641738 Increases risk of nonalcoholic fatty liver disease in individuals of European descent. Gastroenterology 150, 1219–1230 (2016).
Buch, S. et al. A genome-wide association study confirms PNPLA3 and identifies TM6SF2 and MBOAT7 as risk loci for alcohol-related cirrhosis. Nat. Genet. 47, 1443–1448 (2015).
Kozlitina, J. et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 46, 352–356 (2014).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Ciardullo, S. et al. Impact of MASLD and MetALD on clinical outcomes: A meta-analysis of preliminary evidence. Liver Int. 44, 1762–1767 (2024).
Younossi, Z. M., Kalligeros, M. & Henry, L. Epidemiology of metabolic dysfunction-associated steatotic liver disease. Clin. Mol. Hepatol. 31, S32–S50 (2024).
Tinajero, M. G. & Malik, V. S. An update on the epidemiology of type 2 diabetes: a global perspective. Endocrinol. Metab. Clin. North Am. 50, 337–355 (2021).
Díaz, L. A. et al. Disparities in steatosis prevalence in the United States by Race or Ethnicity according to the 2023 criteria. Commun. Med. 4, 219 (2024).
Kubiliun, M. J., Cohen, J. C., Hobbs, H. H. & Kozlitina, J. Contribution of a genetic risk score to ethnic differences in fatty liver disease. Liver Int. 42, 2227–2236 (2022).
Kawaguchi, T. et al. Genetic polymorphisms of the human PNPLA3 gene are strongly associated with severity of non-alcoholic fatty liver disease in Japanese. PLoS ONE 7, e38322 (2012).
Kozlitina, J. & Sookoian, S. Global epidemiological impact of PNPLA3 I148M on liver disease. Liver Int. 45, e16123 (2024).
Romeo, S., Sanyal, A. & Valenti, L. Leveraging human genetics to identify potential new treatments for fatty liver disease. Cell Metab. 31, 35–45 (2020).
Cherubini, A., Della Torre, S., Pelusi, S. & Valenti, L. Sexual dimorphism of metabolic dysfunction-associated steatotic liver disease. Trends Mol. Med. 30, 1126–1136 (2024).
Balakrishnan, M. et al. Women have a lower risk of nonalcoholic fatty liver disease but a higher risk of progression vs men: a systematic review and meta-analysis. Clin. Gastroenterol. Hepatol. 19, 61–71 (2021).
Toniutto, P. et al. Role of sex in liver tumor occurrence and clinical outcomes: a comprehensive review. Hepatology 79, 1141–1157 (2024).
Cherubini, A. et al. Interaction between estrogen receptor-α and PNPLA3 p.I148M variant drives fatty liver disease susceptibility in women. Nat. Med. 29, 2643–2655 (2023).
Zhou, J., Bai, L., Zhang, X. J., Li, H. & Cai, J. Nonalcoholic fatty liver disease and cardiac remodeling risk: pathophysiological mechanisms and clinical implications. Hepatology 74, 2839–2847 (2021).
Hagström, H. et al. Cardiovascular risk factors in non-alcoholic fatty liver disease. Liver Int. 39, 197–204 (2019).
Shang, Y., Nasr, P., Widman, L. & Hagström, H. Risk of cardiovascular disease and loss in life expectancy in NAFLD. Hepatology 76, 1495–1505 (2022).
Mantovani, A. et al. Non-alcoholic fatty liver disease and risk of incident chronic kidney disease: an updated meta-analysis. Gut 71, 156–162 (2022).
Peng, H. et al. Nonalcoholic fatty liver disease and cardiovascular diseases: a Mendelian randomization study. Metabolism 133, 155220 (2022).
Au Yeung, S. L., Borges, M. C., Wong, T. H. T., Lawlor, D. A. & Schooling, C. M. Evaluating the role of non-alcoholic fatty liver disease in cardiovascular diseases and type 2 diabetes: a Mendelian randomization study in Europeans and East Asians. Int J. Epidemiol. 52, 921–931 (2023).
Jamialahmadi, O., et al. Partitioned polygenic risk scores identify distinct types of metabolic dysfunction-associated steatotic liver disease. Nat. Med. 30, 3614–3623 (2024).
Raverdy, V., et al. Data-driven cluster analysis identifies distinct types of metabolic dysfunction-associated steatotic liver disease. Nat. Med. 30, 3624–3633 (2024).
Romeo, S., et al. Clinical staging to guide management of metabolic disorders and their sequelae: a European Atherosclerosis Society consensus statement. Eur. Heart J. https://doi.org/10.1093/eurheartj/ehaf314 (2025).
Ahmed, A., Cule, M., Bell, J. D., Sattar, N. & Yaghootkar, H. Differing genetic variants associated with liver fat and their contrasting relationships with cardiovascular diseases and cancer. J. Hepatol. 81, 921–929 (2024).
Chen, Y. et al. Genome-wide association meta-analysis identifies 17 loci associated with nonalcoholic fatty liver disease. Nat. Genet. 55, 1640–1650 (2023).
Burza, M. A. et al. PNPLA3 I148M (rs738409) genetic variant and age at onset of at-risk alcohol consumption are independent risk factors for alcoholic cirrhosis. Liver Int. 34, 514–520 (2013).
Trépo, E., Romeo, S., Zucman-Rossi, J. & Nahon, P. PNPLA3 gene in liver diseases. J. Hepatol. 65, 399–412 (2016).
Azariadis, K. et al. PNPLA3 I148 M genetic variant in autoimmune hepatitis characterises advanced disease at diagnosis and reduced survival free of cirrhotic events and liver-related mortality. J. Transl. Autoimmun. 9, 100243 (2024).
Luukkonen, P. K., et al. The PNPLA3 I148M variant increases ketogenesis and decreases hepatic de novo lipogenesis and mitochondrial function in humans. Cell Metab. 35, 1887–1896 (2023).
Mancina, R. M. et al. Paradoxical dissociation between hepatic fat content and de novo lipogenesis due to PNPLA3 sequence variant. J. Clin. Endocrinol. Metab. 100, E821–E825 (2015).
Wang, Y., et al. PNPLA3(148 M) is a gain-of-function mutation that promotes hepatic steatosis by inhibiting ATGL-mediated triglyceride hydrolysis. J. Hepatol. 82, 871–881 (2024).
Pingitore, P. et al. Recombinant PNPLA3 protein shows triglyceride hydrolase activity and its I148M mutation results in loss of function. Biochim. Biophys. Acta 1841, 574–580 (2014).
He, S. et al. A sequence variation (I148M) in PNPLA3 associated with nonalcoholic fatty liver disease disrupts triglyceride hydrolysis. J. Biol. Chem. 285, 6706–6715 (2010).
Huang, Y., Cohen, J. C. & Hobbs, H. H. Expression and characterization of a PNPLA3 isoform (I148M) associated with nonalcoholic fatty liver disease. J. Biol. Chem. 286, 37085–37093 (2011).
Luukkonen, P. K. et al. Human PNPLA3-I148M variant increases hepatic retention of polyunsaturated fatty acids. JCI Insight 4, e127902 (2019).
Johnson, S. M. et al. PNPLA3 is a triglyceride lipase that mobilizes polyunsaturated fatty acids to facilitate hepatic secretion of large-sized very low-density lipoprotein. Nat. Commun. 15, 4847 (2024).
Lindén, D. et al. Pnpla3 silencing with antisense oligonucleotides ameliorates nonalcoholic steatohepatitis and fibrosis in Pnpla3 I148M knock-in mice. Mol. Metab. 22, 49–61 (2019).
Armisen, J., et al. AZD2693, a PNPLA3 antisense oligonucleotide, for the treatment of MASH in 148 M homozygous participants: two randomized phase I trials. J. Hepatol. 83, 31–42 (2025).
Fabbrini, E. et al. Phase 1 tTrials of PNPLA3 siRNA in I148M homozygous patients with MAFLD. N. Engl. J. Med. 391, 475–476 (2024).
Prill, S. et al. The TM6SF2 E167K genetic variant induces lipid biosynthesis and reduces apolipoprotein B secretion in human hepatic 3D spheroids. Sci. Rep. 9, 11585 (2019).
Mahdessian, H. et al. TM6SF2 is a regulator of liver fat metabolism influencing triglyceride secretion and hepatic lipid droplet content. Proc. Natl Acad. Sci. USA 111, 8913–8918 (2014).
Smagris, E., Gilyard, S., BasuRay, S., Cohen, J. C. & Hobbs, H. H. Inactivation of Tm6sf2, a gene defective in fatty liver disease, impairs lipidation but not secretion of very low density lipoproteins. J. Biol. Chem. 291, 10659–10676 (2016).
Dongiovanni, P. et al. Transmembrane 6 superfamily member 2 gene variant disentangles nonalcoholic steatohepatitis from cardiovascular disease. Hepatology 61, 506–514 (2015).
Tanaka, Y. et al. LPIAT1/MBOAT7 depletion increases triglyceride synthesis fueled by high phosphatidylinositol turnover. Gut 70, 180–193 (2020).
Xia, M., Chandrasekaran, P., Rong, S., Fu, X. & Mitsche, M. A. Hepatic deletion of Mboat7 (LPIAT1) causes activation of SREBP-1c and fatty liver. J. Lipid Res. 62, 100031 (2021).
Speliotes, E. K. et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7, e1001324 (2011).
Jamialahmadi, O. et al. Exome-wide association study on alanine aminotransferase identifies sequence variants in the GPAM and APOE associated with fatty liver disease. Gastroenterology 160, 1634–1646 (2021).
Ghouse, J. et al. Integrative common and rare variant analyses provide insights into the genetic architecture of liver cirrhosis. Nat. Genet. 56, 827–837 (2024).
Emdin, C. A. et al. A missense variant in Mitochondrial Amidoxime Reducing Component 1 gene and protection against liver disease. PLoS Genet. 16, e1008629 (2020).
Dutta, T. et al. Mitochondrial amidoxime-reducing component 1 p. Ala165Thr increases protein degradation mediated by the proteasome. Liver Int. 44, 1219–1232 (2024).
Hou, W. et al. Biochemical and functional characterization of the p. A165T missense variant of mitochondrial amidoxime-reducing component 1. J. Biol. Chem. 300, 107353 (2024).
Wu, M. et al. Fatty liver disease protective MTARC1 p. A165T variant reduces the protein stability of MTARC1. Biochem. Biophys. Res. Commun. 702, 149655 (2024).
Ciociola, E., et al. MARC1 downregulation reduces hepatocyte lipid content by increasing beta-oxidation. Clin. Mol. Hepatol. 31, 445–459 (2024).
Lindén, D. & Romeo, S. Therapeutic opportunities for the treatment of NASH with genetically validated targets. J. Hepatol. 79, 1056–1064 (2023).
Dongiovanni, P. et al. Causal relationship of hepatic fat with liver damage and insulin resistance in nonalcoholic fatty liver. J. Intern. Med. 283, 356–370 (2018).
Chen, J., Zhou, H., Jin, H. & Liu, K. Role of inflammatory factors in mediating the effect of lipids on nonalcoholic fatty liver disease: a two-step, multivariable Mendelian randomization study. Nutrients 14, 4434 (2022).
Bianco, C. et al. Non-invasive stratification of hepatocellular carcinoma risk in non-alcoholic fatty liver using polygenic risk scores. J. Hepatol. 74, 775–782 (2021).
Harrison, S. A. et al. Resmetirom for nonalcoholic fatty liver disease: a randomized, double-blind, placebo-controlled phase 3 trial. Nat. Med. 29, 2919–2928 (2023).
Alfawaz, S., Burzangi, A. & Esmat, A. Mechanisms of non-alcoholic fatty liver disease and beneficial effects of semaglutide: a review. Cureus 16, e67080 (2024).
Luukkonen, P. K. et al. Hydroxysteroid 17-β dehydrogenase 13 variant increases phospholipids and protects against fibrosis in nonalcoholic fatty liver disease. JCI Insight 5, e132158 (2020).
Romeo, S. et al. Morbid obesity exposes the association between PNPLA3 I148M (rs738409) and indices of hepatic injury in individuals of European descent. Int J. Obes. 34, 190–194 (2010).
Stender, S. et al. Adiposity amplifies the genetic risk of fatty liver disease conferred by multiple loci. Nat. Genet. 49, 842–847 (2017).
Jamialahmadi, O. et al. Genome-wide interaction study with BMI identifies CYP7A1 and GIPR as genetic modulators of MASLD. Clin. Mol. Hepatol. https://doi.org/10.3350/cmh.2025.0159 (2025).
Pelusi, S. et al. Prevalence and determinants of liver disease in relatives of italian patients with advanced MASLD. Clin. Gastroenterol. Hepatol. 22, 2231–2239 (2024).
Pelusi, S. et al. Rare pathogenic variants predispose to hepatocellular carcinoma in nonalcoholic fatty liver disease. Sci. Rep. 9, 3682 (2019).
Baselli, G. A. et al. Rare ATG7 genetic variants predispose patients to severe fatty liver disease. J. Hepatol. 77, 596–606 (2022).
Brunt, E. M. et al. Complexity of ballooned hepatocyte feature recognition: defining a training atlas for artificial intelligence-based imaging in NAFLD. J. Hepatol. 76, 1030–1041 (2022).
Baselli, G., Romeo, S. & Valenti, L. Reply to: “Lack of hepatic autophagy promotes severity of liver injury but not steatosis”: ATG7 genetic variants behave as fatty liver disease progression modifiers. J. Hepatol. 77, 1459–1461 (2022).
Mashek, D. G. Hepatic lipid droplets: a balancing act between energy storage and metabolic dysfunction in NAFLD. Mol. Metab. 50, 101115 (2021).
Mantovani, A. et al. Association between primary hypothyroidism and metabolic dysfunction-associated steatotic liver disease: an updated meta-analysis. Gut 73, 1554–1561 (2024).
Sinha, R. A. Thyroid hormone-mediated selective autophagy and its implications in countering metabolic dysfunction-associated steatotic liver disease. Endocrinol. Metab. 39, 686–692 (2024).
Zeng, X. F. et al. The role of dietary modification in the prevention and management of metabolic dysfunction-associated fatty liver disease: an international multidisciplinary expert consensus. Metabolism 161, 156028 (2024).
Verweij, N. et al. Germline mutations in CIDEB and protection against liver disease. N. Engl. J. Med. 387, 332–344 (2022).
Valenti, L., Bugianesi, E., Pajvani, U. & Targher, G. Nonalcoholic fatty liver disease: cause or consequence of type 2 diabetes?. Liver Int. 36, 1563–1579 (2016).
Younossi, Z. M. et al. The global epidemiology of nonalcoholic fatty liver disease and nonalcoholic steatohepatitis among patients with type 2 diabetes. Clin. Gastroenterol. Hepatol. 22, 1999–2010 (2024).
Huang, D. Q. et al. Type 2 diabetes, hepatic decompensation, and hepatocellular carcinoma in patients with non-alcoholic fatty liver disease: an individual participant-level data meta-analysis. Lancet Gastroenterol. Hepatol. 8, 829–836 (2023).
Huang, D. Q. et al. Fibrosis progression rate in biopsy-proven nonalcoholic fatty liver disease among people with diabetes versus people without. Diabetes 165, 463–472 (2023).
Tamaki, N. et al. Glycemic control target for liver and cardiovascular events risk in metabolic dysfunction-associated steatotic liver disease. Hepatol. Res 54, 753–762 (2024).
Huang, Y. et al. A feed-forward loop amplifies nutritional regulation of PNPLA3. Proc. Natl Acad. Sci. USA 107, 7892–7897 (2010).
Najjar, S. M., Caprio, S. & Gastaldelli, A. Insulin clearance in health and disease. Annu Rev. Physiol. 85, 363–381 (2023).
Kotronen, A., Juurinen, L., Tiikkainen, M., Vehkavaara, S. & Yki-Järvinen, H. Increased liver fat, impaired insulin clearance, and hepatic and adipose tissue insulin resistance in type 2 diabetes. Gastroenterology 135, 122–130 (2008).
Valenti, L. et al. Increased expression and activity of the transcription factor FOXO1 in nonalcoholic steatohepatitis. Diabetes 57, 1355–1362 (2008).
Rametta, R. et al. Increased insulin receptor substrate 2 expression is associated with steatohepatitis and altered lipid metabolism in obese subjects. Int J. Obes. 37, 986–992 (2013).
Valenti, L. et al. Hepatic notch signaling correlates with insulin resistance and nonalcoholic fatty liver disease. Diabetes 62, 4052–4062 (2013).
Rametta, R. et al. Hepatic steatosis and PNPLA3 I148M variant are associated with serum Fetuin-A independently of insulin resistance. Eur. J. Clin. Invest. 44, 627–633 (2014).
Milani, I. et al. Hepatokines and MASLD: the GLP1–Ras–FGF21–Fetuin-A crosstalk as a therapeutic target. Int. J. Mol. Sci. 25, 10795 (2024).
Ng, S. W. K. et al. Convergent somatic mutations in metabolism genes in chronic liver disease. Nature 598, 473–478 (2021).
Goswami, A., Bhargava, N., Dadhich, S. & Kulamarva, G. Insulin resistance in euglycemic cirrhosis. Ann. Gastroenterol. 27, 237–243 (2014).
Sveinbjornsson, G. et al. Multiomics study of nonalcoholic fatty liver disease. Nat. Genet. 54, 1652–1663 (2022).
Wang, Z. et al. Positive selection of somatically mutated clones identifies adaptive pathways in metabolic liver disease. Cell 186, 1968–1984 (2023).
Kotiyal, S. & Evason, K. J. Exploring the interplay of telomerase reverse transcriptase and β-catenin in hepatocellular carcinoma. Cancers 13, 4202 (2021).
Marchetti, A., et al. Impact of clonal hematopoiesis of indeterminate potential on hepatocellular carcinoma in individuals with steatotic liver disease. Hepatology 80, 816–827 (2024).
Wong, W. J. et al. Clonal haematopoiesis and risk of chronic liver disease. Nature 616, 747–754 (2023).
Caddeo, A. & Romeo, S. Precision medicine and nucleotide-based therapeutics to treat MASH. Clin. Mol. Hepatol. 31, S76–S93 (2024).
Mancina, R. M. et al. PSD3 downregulation confers protection against fatty liver disease. Nat. Metab. 4, 60–75 (2022).
Sakuma, I. et al. Lysophosphatidic acid triggers inflammation in the liver and white adipose tissue in rat models of 1-acyl-sn-glycerol-3-phosphate acyltransferase 2 deficiency and overnutrition. Proc. Natl Acad. Sci. USA 120, e2312666120 (2023).
Lindén, D. et al. Liver-directed overexpression of mitochondrial glycerol-3-phosphate acyltransferase results in hepatic steatosis, increased triacylglycerol secretion and reduced fatty acid oxidation. FASEB J. 20, 434–443 (2006).
Keating, S. E. et al. High-intensity interval training is safe, feasible and efficacious in nonalcoholic steatohepatitis: a randomized controlled trial. Dig. Dis. Sci. 68, 2123–2139 (2023).
Stine, J. G. et al. NASHFit: a randomized controlled trial of an exercise training program to reduce clotting risk in patients with NASH. Hepatology 76, 172–185 (2022).
Mardinoglu, A. et al. An integrated understanding of the rapid metabolic benefits of a carbohydrate-restricted diet on hepatic steatosis in humans. Cell Metab. 27, 559–571 (2018).
Browning, J. D. et al. Short-term weight loss and hepatic triglyceride reduction: evidence of a metabolic advantage with dietary carbohydrate restriction. Am. J. Clin. Nutr. 93, 1048–1052 (2011).
Taskinen, M. R. et al. Adverse effects of fructose on cardiometabolic risk factors and hepatic lipid metabolism in subjects with abdominal obesity. J. Intern. Med. 282, 187–201 (2017).
Speliotes, E. K. & Schneider, C. V. PNPLA3 I148M interacts with environmental triggers to cause human disease. Liver Int. 45, e16106 (2024).
Chen, V. L. et al. Genetic risk accentuates dietary effects on hepatic steatosis, inflammation and fibrosis in a population-based cohort. J. Hepatol. 81, 379–388 (2024).
Santoro, N. et al. Hepatic fat accumulation is modulated by the interaction between the rs738409 variant in the PNPLA3 gene and the dietary Omega6/Omega3 PUFA intake. PLoS ONE 7, e37827 (2012).
Nobili, V. et al. Influence of dietary pattern, physical activity, and I148M PNPLA3 on steatosis severity in at-risk adolescents. Genes Nutr. 9, 392 (2014).
Sevastianova, K. et al. Genetic variation in PNPLA3 (adiponutrin) confers sensitivity to weight loss-induced decrease in liver fat in humans. Am. J. Clin. Nutr. 94, 104–111 (2011).
Dongiovanni, P. et al. Statin use and non-alcoholic steatohepatitis in at risk individuals. J. Hepatol. 63, 705–712 (2015).
Ahmed, O. et al. Statin-associated regulation of hepatic PNPLA3 in patients without known liver disease. J. Intern. Med. 297, 47–59 (2025).
Nobili, V., Bedogni, G., Donati, B., Alisi, A. & Valenti, L. The I148M variant of PNPLA3 reduces the response to docosahexaenoic acid in children with non-alcoholic fatty liver disease. J. Med. Food 16, 957–960 (2013).
Vell, M. S. et al. Omega-3 intake is associated with liver disease protection. Front. Public Health 11, 1192099 (2023).
Pillai, S. et al. The PNPLA3 I148M variant is associated with transaminase elevations in type 2 diabetes patients treated with basal insulin peglispro. Pharmacogenomics J. 18, 487–493 (2018).
Urias, E. et al. PNPLA3 risk allele association with ALT response to semaglutide treatment. Gastroenterology 166, 515–517 (2024).
Moretti, V., Romeo, S. & Valenti, L. The contribution of genetics and epigenetics to MAFLD susceptibility. Hepatol. Int. 18, 848–860 (2024).
Wand, H. et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 591, 211–219 (2021).
Nobili, V. et al. A 4-polymorphism risk score predicts steatohepatitis in children with nonalcoholic fatty liver disease. J. Pediatr. Gastroenterol. Nutr. 58, 632–636 (2014).
Donati, B. et al. MBOAT7 rs641738 variant and hepatocellular carcinoma in non-cirrhotic individuals. Sci. Rep. 7, 4492 (2017).
De Vincentis, A. et al. A polygenic risk score to refine risk stratification and prediction for severe liver disease by clinical fibrosis scores. Clin. Gastroenterol. Hepatol. 20, 658–673 (2022).
Innes, H. et al. Performance of routine risk scores for predicting cirrhosis-related morbidity in the community. J. Hepatol. 77, 365–376 (2022).
Bianco, C., Tavaglione, F., Romeo, S. & Valenti, L. Genetic risk scores and personalization of care in fatty liver disease. Curr. Opin. Pharmacol. 61, 6–11 (2021).
Vujkovic, M. et al. A multiancestry genome-wide association study of unexplained chronic ALT elevation as a proxy for nonalcoholic fatty liver disease with histological and radiological validation. Nat. Genet. 54, 761–771 (2022).
Acknowledgements
S.R. was supported by the Swedish Cancerfonden (22 2270 Pj), the Swedish Research Council (Vetenskapsradet (VR), 2023-02079), the Swedish state under the Agreement between the Swedish government and the county councils (the ALF agreement, ALFGBG-965360), the Swedish Heart Lung Foundation (20220334), the Novo Nordisk Distinguished Investigator Grant - Endocrinology and Metabolism (NNF23OC0082114), the Novo Nordisk Project grants in Endocrinology and Metabolism (NNF24OC0091535) and AstraZeneca Agreement for Research. L.V. was supported by the Italian Ministry of Health (Ministero della Salute), Ricerca Finalizzata 2021 RF-2021-12373889, Italian Ministry of Health, Ricerca Finalizzata PNRR 2022 ‘RATIONAL: Risk stratification of nonalcoholic fatty liver’ PNRR-MAD-2022-12375656, the European Union, H2020-ICT-2018-20/H2020-ICT-2020-2 programme ‘Photonics’ under grant agreement No. 101016726 – REVEAL, the European Union, HORIZON-MISS-2021-CANCER-02-03 programme ‘Genial’ under grant agreement ‘101096312’, the Italian Ministry of Research (MUR) PNRR – M4 - C2 ‘National Center for Gene Therapy and Drugs based on RNA Technology’ CN3, Spoke 4 ‘ASSET: A sex-specific approach to NAFLD targeting’, and the PRIN 2022 MUR: ‘Disentangling genetic, epigenetic and hormonal regulation of Fe/heme metabolism in the gender-specific nature of NAFLD (DEFENDER)’. The funders had no role in the present work.
Author information
Authors and Affiliations
Contributions
All authors contributed to the content and organization of this Review; to writing, editing and/or revising the manuscript; and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
S.R. received the research grant from AstraZeneca for basic-science research on steatotic liver disease and has been consulting for AstraZeneca, GSK, Celgene Corporation, Ribo-cure AB¸ Ultragenyx, Amgen, Sanofi, Wave Life Sciences, Novartis, Chiesi and Pfizer in the past 5 years; declares equity in Heptabio; and is an inventor on a patent titled “Method for treating fatty liver disease”, on PSD3, US application number 17,480266 filed on 21 September 2021. L.V. has been consulting for Novo Nordisk, Pfizer, Boehringer Ingelheim, Resalis, MSD and speaker for Viatris, Novo Nordisk, GSK. R.M.M. has nothing to declare.
Peer review
Peer review information
Nature Metabolism thanks Stefan Stender, Michele Vacca and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mancina, R.M., Valenti, L. & Romeo, S. Human genetics of steatotic liver disease: insights into insulin resistance and lipid metabolism. Nat Metab 7, 2199–2211 (2025). https://doi.org/10.1038/s42255-025-01394-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42255-025-01394-8
This article is cited by
-
Umbilical Cord Mesenchymal Stem Cells and Wnt Pathway Modulation: A Novel Therapy for Liver Cirrhosis and Steatosis
Tissue Engineering and Regenerative Medicine (2026)