Abstract
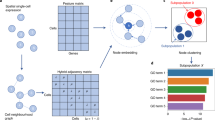
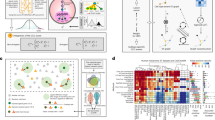
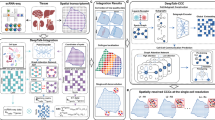
Cell–cell interactions (CCI), driven by distance-dependent signalling, are important for tissue development and organ function. While imaging-based spatial transcriptomics offers unprecedented opportunities to unravel CCI at single-cell resolution, current analyses face challenges such as limited ligand–receptor pairs measured, insufficient spatial encoding and low interpretability. We present GITIII (graph inductive bias transformer for intercellular interaction investigation), a lightweight, interpretable, self-supervised graph transformer-based model that conceptualizes cells as words and their surrounding cellular neighbourhood as context that shapes the meaning or state of the central cell. GITIII infers CCI by examining the correlation between a cell’s state and its niche, enabling us to understand how sender cells influence the gene expression of receiver cells, visualize spatial CCI patterns, perform CCI-informed cell clustering and construct CCI networks. Applied to four spatial transcriptomics datasets across multiple species, organs and platforms, GITIII effectively identified and statistically interpreted CCI patterns in the brain and tumour microenvironments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The mouse PMC MERFISH dataset is available at ftp://download.brainimagelibrary.org:8811/02/26/02265ddb0dae51de/. The SEA-AD MERFISH dataset is available at https://registry.opendata.aws/allen-sea-ad-atlas/. The NSCLC CosMx dataset is available at https://nanostring.com/products/cosmx-spatial-molecular-imager/ffpe-dataset/nsclc-ffpe-dataset/. The breast cancer Xenium dataset is available via 10X Genomics at https://www.10xgenomics.com/products/xenium-in-situ/preview-dataset-human-breast. The ligand–receptor databases of CellChat and NeuronChat are publicly available, and an integrated version is available via GitHub at https://github.com/lugia-xiao/GITIII.
Code availability
GITIII is implemented in Python using PyTorch. The code and tutorial are available via GitHub at https://github.com/lugia-xiao/GITIII and via Zenodo at https://doi.org/10.5281/zenodo.17476318 (ref. 85). The code to reproduce the results in this paper is available via GitHub at https://github.com/lugia-xiao/GITIII_reproducible and via Zenodo at https://doi.org/10.5281/zenodo.17476330 (ref. 86).
References
Armingol, E., Baghdassarian, H. M. & Lewis, N. E. The diversification of methods for studying cell-cell interactions and communication. Nat. Rev. Genet. 25, 381–400 (2024).
Armingol, E. et al. Context-aware deconvolution of cell-cell communication with Tensor-cell2cell. Nat. Commun. 13, 3665 (2022).
Graeber, T. G. & Eisenberg, D. Bioinformatic identification of potential autocrine signaling loops in cancers from gene expression profiles. Nat. Genet. 29, 295–300 (2001).
Liu, Z., Sun, D. & Wang, C. Evaluation of cell-cell interaction methods by integrating single-cell RNA sequencing data with spatial information. Genome Biol. 23, 218 (2022).
Wang, T., Li, B., Nelson, C. E. & Nabavi, S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinforma. 20, 40 (2019).
Zhao, W., Johnston, K. G., Ren, H., Xu, X. & Nie, Q. Inferring neuron-neuron communications from single-cell transcriptomics through NeuronChat. Nat. Commun. 14, 1128 (2023).
Arnol, D., Schapiro, D., Bodenmiller, B., Saez-Rodriguez, J. & Stegle, O. Modeling cell-cell interactions from spatial molecular data with spatial variance component analysis. Cell Rep. 29, 202–211 (2019).
Cang, Z. et al. Screening cell-cell communication in spatial transcriptomics via collective optimal transport. Nat. Methods 20, 218–228 (2023).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Li, H. et al. Decoding functional cell-cell communication events by multi-view graph learning on spatial transcriptomics. Brief Bioinform. 24, bbad359 (2023).
Li, Z., Wang, T., Liu, P. & Huang, Y. SpatialDM for rapid identification of spatially co-expressed ligand-receptor and revealing cell–cell communication patterns. Nat. Commun. 14, 3995 (2023).
Pham, D. et al. Robust mapping of spatiotemporal trajectories and cell-cell interactions in healthy and diseased tissues. Nat. Commun. 14, 7739 (2023).
Schrod, S. et al. Spatial cellular networks from omics data with SpaCeNet. Genome Res 34, 1371–1383 (2024).
Shao, X. et al. Knowledge-graph-based cell–cell communication inference for spatially resolved transcriptomic data with SpaTalk. Nat. Commun. 13, 4429 (2022).
So, E., Hayat, S., Nair, S. K., Wang, B. & Haibe-Kains, B. GraphComm predicts cell cell communication using a graph based deep learning method in single cell RNA sequencing data. Sci. Rep. 15, 36914 (2025).
Wang, H. et al. SOAPy: a Python package to dissect spatial architecture, dynamics, and communication. Genome Biol. 26, 80 (2025).
Wu, D., Gaskins, J. T., Sekula, M. & Datta, S. Inferring cell-cell communications from spatially resolved transcriptomics data using a Bayesian Tweedie model. Genes 14, 1368 (2023).
Yang, W. et al. Deciphering cell-cell communication at single-cell resolution for spatial transcriptomics with subgraph-based graph attention network. Nat. Commun. 15, 7101 (2024).
Browaeys, R. et al. MultiNicheNet: a flexible framework for differential cell-cell communication analysis from multi-sample multi-condition single-cell transcriptomics data. Preprint at bioRxiv https://doi.org/10.1101/2023.06.13.544751 (2023).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. 20, 180–219 (2025).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Janesick, A. et al. High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis. Nat. Commun. 14, 8353 (2023).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genet 53, 1698–1711 (2021).
Liu, J. et al. CytoSignal detects locations and dynamics of ligand-receptor signaling at cellular resolution from spatial transcriptomic data. Preprint at bioRxiv https://doi.org/10.1101/2024.03.08.584153 (2024).
Yan, L., Cheng, J., Nie, Q. & Sun, X. Dissecting multilayer cell-cell communications with signaling feedback loops from spatial transcriptomics data. Genome Res 35, 1400–1414 (2025).
Fischer, D. S., Schaar, A. C. & Theis, F. J. Modeling intercellular communication in tissues using spatial graphs of cells. Nat. Biotechnol. 41, 332–336 (2023).
Wang, H. et al. CGCom: a framework for inferring cell-cell communication based on graph neural network. Preprint at bioRxiv https://doi.org/10.1101/2023.11.10.566642 (2023).
Yuan, Y. & Bar-Joseph, Z. GCNG: graph convolutional networks for inferring gene interaction from spatial transcriptomics data. Genome Biol. 21, 300 (2020).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Lanjakornsiripan, D. et al. Layer-specific morphological and molecular differences in neocortical astrocytes and their dependence on neuronal layers. Nat. Commun. 9, 1623 (2018).
Ben Haim, L. & Rowitch, D. H. Functional diversity of astrocytes in neural circuit regulation. Nat. Rev. Neurosci. 18, 31–41 (2017).
Batiuk, M. Y. et al. Identification of region-specific astrocyte subtypes at single cell resolution. Nat. Commun. 11, 1220 (2020).
Bayraktar, O. A. et al. Astrocyte layers in the mammalian cerebral cortex revealed by a single-cell in situ transcriptomic map. Nat. Neurosci. 23, 500–509 (2020).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
O’Dea, M. R. & Hasel, P. Are we there yet? Exploring astrocyte heterogeneity one cell at a time. Glia 73, 619–631 (2025).
Mathys, H. et al. Single-cell multiregion dissection of Alzheimer’s disease. Nature 632, 858–868 (2024).
Singhal, V. et al. BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis. Nat. Genet. 56, 431–441 (2024).
Molina-Gonzalez, I. et al. Astrocyte-oligodendrocyte interaction regulates central nervous system regeneration. Nat. Commun. 14, 3372 (2023).
Cubelos, B. et al. Cux-2 controls the proliferation of neuronal intermediate precursors of the cortical subventricular zone. Cereb. Cortex 18, 1758–1770 (2008).
Nagata, I., Keilhauer, G. & Schachner, M. Neuronal influence on antigenic marker profile, cell shape and proliferation of cultured astrocytes obtained by microdissection of distinct layers from the early postnatal mouse cerebellum. Brain Res. 389, 217–232 (1986).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
Gao, C., Jiang, J., Tan, Y. & Chen, S. Microglia in neurodegenerative diseases: mechanism and potential therapeutic targets. Signal Transduct. Target Ther. 8, 359 (2023).
Leng, F. & Edison, P. Neuroinflammation and microglial activation in Alzheimer disease: where do we go from here?. Nat. Rev. Neurol. 17, 157–172 (2021).
Leng, K. et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat. Neurosci. 24, 276–287 (2021).
Stogsdill, J. A. et al. Pyramidal neuron subtype diversity governs microglia states in the neocortex. Nature 608, 750–756 (2022).
Depp, C., Doman, J. L., Hingerl, M., Xia, J. & Stevens, B. Microglia transcriptional states and their functional significance: context drives diversity. Immunity 58, 1052–1067 (2025).
Fang, R. et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56–62 (2022).
Suzuki, T. et al. CUX2 deficiency causes facilitation of excitatory synaptic transmission onto hippocampus and increased seizure susceptibility to kainate. Sci. Rep. 12, 6505 (2022).
Fujita, Y., Nakanishi, T., Ueno, M., Itohara, S. & Yamashita, T. Netrin-G1 regulates microglial accumulation along axons and supports the survival of layer V neurons in the postnatal mouse brain. Cell Rep. 31, 107580 (2020).
Yaguchi, K., Nishimura-Akiyoshi, S., Kuroki, S., Onodera, T. & Itohara, S. Identification of transcriptional regulatory elements for Ntng1 and Ntng2 genes in mice. Mol. Brain 7, 19 (2014).
McNamara, N. B. et al. Microglia regulate central nervous system myelin growth and integrity. Nature 613, 120–129 (2023).
Whalley, K. Neuronal neighbours tune microglial identity. Nat. Rev. Neurosci. 23, 582–583 (2022).
Bryan, K. J. et al. Expression of CD74 is increased in neurofibrillary tangles in Alzheimer’s disease. Mol. Neurodegener. 3, 13 (2008).
Su, H., Na, N., Zhang, X. & Zhao, Y. The biological function and significance of CD74 in immune diseases. Inflamm. Res 66, 209–216 (2017).
Olah, M. et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat. Commun. 11, 6129 (2020).
Bill, R. et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science 381, 515–524 (2023).
Yang, Q. et al. Single-cell RNA sequencing reveals the heterogeneity of tumor-associated macrophage in non-small cell lung cancer and differences between sexes. Front. Immunol. 12, 756722 (2021).
Lichtenberger, B. M. et al. Autocrine VEGF signaling synergizes with EGFR in tumor cells to promote epithelial cancer development. Cell 140, 268–279 (2010).
Sharma, S. V., Bell, D. W., Settleman, J. & Haber, D. A. Epidermal growth factor receptor mutations in lung cancer. Nat. Rev. Cancer 7, 169–181 (2007).
Lazaratos, A. M., Annis, M. G. & Siegel, P. M. GPNMB: a potent inducer of immunosuppression in cancer. Oncogene 41, 4573–4590 (2022).
Saha, S. K. et al. KRT19 directly interacts with beta-catenin/RAC1 complex to regulate NUMB-dependent NOTCH signaling pathway and breast cancer properties. Oncogene 36, 332–349 (2017).
Astsaturov, I. et al. Synthetic lethal screen of an EGFR-centered network to improve targeted therapies. Sci. Signal 3, ra67 (2010).
Xu, S. et al. Using clusterProfiler to characterize multiomics data. Nat. Protoc. 19, 3292–3320 (2024).
DeLeon-Pennell, K. Y., Barker, T. H. & Lindsey, M. L. Fibroblasts: the arbiters of extracellular matrix remodeling. Matrix Biol. 9, 1–7 (2020).
Lai, Y. et al. Interleukin-8 induces the endothelial cell migration through the activation of phosphoinositide 3-kinase-Rac1/RhoA pathway. Int J. Biol. Sci. 7, 782–791 (2011).
Denisenko, T. V., Pivnyuk, A. D. & Zhivotovsky, B. p53-autophagy-metastasis link. Cancers 10, 148 (2018).
Jung, H., Seong, H. A. & Ha, H. Critical role of cysteine residue 81 of macrophage migration inhibitory factor (MIF) in MIF-induced inhibition of p53 activity. J. Biol. Chem. 283, 20383–20396 (2008).
Pavlyukov, M. S. et al. Apoptotic cell-derived extracellular vesicles promote malignancy of glioblastoma via intercellular transfer of splicing factors. Cancer Cell 34, 119–135 (2018).
Chen, F. et al. Extracellular vesicle-packaged HIF-1alpha-stabilizing lncRNA from tumour-associated macrophages regulates aerobic glycolysis of breast cancer cells. Nat. Cell Biol. 21, 498–510 (2019).
Horrevorts, S. K. et al. Glycan-modified apoptotic melanoma-derived extracellular vesicles as antigen source for anti-tumor vaccination. Cancers 11, 1266 (2019).
He, Y. et al. FOXA1 overexpression suppresses interferon signaling and immune response in cancer. J. Clin. Invest. 131, e147025 (2021).
Schultz, S. et al. Progression-specific genes identified in microdissected formalin-fixed and paraffin-embedded tissue containing matched ductal carcinoma in situ and invasive ductal breast cancers. BMC Med. Genomics 11, 80 (2018).
Takahashi, Y., Kitadai, Y., Bucana, C. D., Cleary, K. R. & Ellis, L. M. Expression of vascular endothelial growth factor and its receptor, KDR, correlates with vascularity, metastasis, and proliferation of human colon cancer. Cancer Res. 55, 3964–3968 (1995).
Hanley, C. J., Henriet, E., Sirka, O. K., Thomas, G. J. & Ewald, A. J. Tumor-resident stromal cells promote breast cancer invasion through regulation of the basal phenotype. Mol. Cancer Res. 18, 1615–1622 (2020).
Ma, L. H. et al. Graph inductive biases in transformers without message passing. Proc. Mach. Learn. Res. 202, 23321–23337 (2023).
Littman, R. et al. Joint cell segmentation and cell type annotation for spatial transcriptomics. Mol. Syst. Biol. 17, e10108 (2021).
Fu, X. et al. BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data. Nat. Commun. 15, 509 (2024).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Lin, Y. et al. scClassify: sample size estimation and multiscale classification of cells using single and multiple reference. Mol. Syst. Biol. 16, e9389 (2020).
Xiao, X. GITIII code. Zenodo https://doi.org/10.5281/zenodo.17476318 (2025).
Xiao, X. GITIII paper results code. Zenodo https://doi.org/10.5281/zenodo.17476330 (2025).
Acknowledgements
We thank J. Y. H. Yang and Y. Lin for generously sharing their data generated using different cell segmentation methods. X.X. and Z.W. were supported by the National Institutes of Health (NIH; grant no. R01LM014087). L.Z. was supported by the NIH (grant nos. R56AG074015 and P30AG066508) and Women’s Health Research Center at Yale. H.Z. was supported by the NIH (grant nos. R56AG074015 and U01HG013840).
Author information
Authors and Affiliations
Contributions
X.X. and Z.W. conceptualized the study. X.X. designed and implemented the model algorithm, analysed the data and created the figures. L.Z., H.Z. and Z.W. refined the analyses and aided result interpretation. X.X. and Z.W. wrote the paper. Z.W. supervised the project. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Ellis Patrick, Qiangfeng Cliff Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Spatial gene expression in subgroups identified by GITIII and spatial distribution of CCI influencing Rorb, Gfap, and Cux2 in the mouse PMC dataset.
a, Super type annotation of astrocytes by Zhang et al. b, Spatial distribution of astrocyte subgroups and excitatory neurons. Specifically, astrocyte subgroup 1 is located in layer 1, subgroup 2 is located in layers 2 and 3, subgroup 3 is located in layers 4 and 5, subgroup 4 is located in layer 6, and subgroup 5 is located in the white matter. c, Spatial distribution of astrocyte subgroups and oligodendrocytes. d, Spatial expression patterns of Cux2, Lama3, Igfbp5, and Gfap in astrocytes. Astrocyte subgroup 2 located in layers 2 and 3 are physically close to L2/3 IT neurons and had higher expression of Cux2 and Lama3. In contrast, astrocyte subgroup 5 located in the white matter are physically close to oligodendrocytes and had higher expression of Gfap and Igfbp5 compared to other astrocytes. e, Super type annotation of L2/3 IT neurons by Zhang et al. f, Spatial distribution of L2/3 IT subgroups. Specifically, subgroup 1 is located in the upper region of layers 2 and 3, closer to layer 1, while subgroups 2 and 3 are located in the deeper region of layers 2 and 3, closer to layer 4. g, Spatial distribution of L2/3 IT subgroups and other excitatory neurons. h, Spatial expression patterns of Camk2d, Trp53i11, Rorb, and Sulf2 in L2/3 IT neurons. L2/3 IT subgroups 2 and 3 located in the deeper region of layers 2 and 3 are physically close to L4/5 IT neurons and had higher expression of Rorb and Sulf2, respectively, compared to other L2/3 IT cells. i, Spatial distribution of interactions from L4/5 IT to L2/3 IT neurons influencing the expression of Rorb. j, Spatial distribution of interactions from oligodendrocytes to astrocytes influencing the expression of Gfap. k, Spatial distribution of interactions from L2/3 IT neurons to astrocytes influencing the expression of Cux2.
Extended Data Fig. 2 Spatial distribution of microglia and L2/3 IT neurons in the SEA-AD dataset.
a, The strength of interactions influencing the expression of RORB. b, Spatial distribution of microglia subgroups and excitatory neurons. Specifically, microglia subgroup 1 is located in layers 2 and 3, subgroup 2 in layers 4 and 5, and subgroup 3 in layer 6. c, Super type annotation of L2/3 IT neurons in Gabitto et al. d, UMAP visualization of slides using the inferred network strength as features, with colour indicating slide from patients with or without dementia.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–8 and Figs. 1–8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, X., Zhang, L., Zhao, H. et al. Inferring spatial single-cell-level interactions through interpreting cell state and niche correlations learned by self-supervised graph transformer. Nat Mach Intell 8, 42–58 (2026). https://doi.org/10.1038/s42256-025-01161-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s42256-025-01161-0
This article is cited by
-
Identifying spatial single-cell-level interactions with graph transformer
Nature Machine Intelligence (2026)