Abstract
Background
Hepatocellular carcinoma (HCC), a leading cause of cancer-related deaths globally, poses significant challenges in early detection. Improved diagnostic accuracy can drastically influence patient outcomes, emphasizing the need for innovative, non-invasive biomarkers.
Methods:
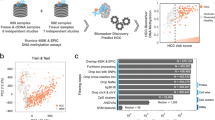
This study utilized a cohort of 402 participants, including healthy controls, chronic hepatitis patients, and HCC patients from Bangladesh, to evaluate DNA methylation signatures in peripheral blood mononuclear cells (PBMC). We performed targeted next-generation sequencing on selected genes previously identified to assess their methylation dynamics. The development of M8 and M4 scores was based on these dynamics, using Receiver Operating Characteristic (ROC) analysis to determine their effectiveness in detecting early-stage HCC alongside existing markers such as epiLiver and alpha-fetoprotein (AFP).
Results:
Integration of M8 and M4 scores with epiLiver and AFP significantly enhances diagnostic sensitivity for early-stage HCC. The M4+epiLiver score achieves a sensitivity of 79.4% in Stage A HCC, while combining M4 with AFP increases sensitivity to 88.2–95.7% across all stages, indicating a superior diagnostic performance compared to each marker used alone.
Conclusions:
Our study confirms that combining gene methylation profiles with established diagnostic markers substantially improves the sensitivity of detecting early-stage HCC. This integrated diagnostic approach holds promise for advancing non-invasive cancer diagnostics, potentially leading to earlier treatment interventions and improved survival rates for high-risk patients.
Plain Language Summary
Liver cancer is one of the top causes of cancer death worldwide, and finding it early is crucial for successful treatment. This research focuses on using a simple blood test to look for specific DNA changes that signal the early stages of liver cancer. We tested this method on a diverse group of people from Bangladesh, including those already at high risk for liver cancer due to chronic liver infections. By combining this new blood test with other existing tests, we were able to detect liver cancer more accurately and earlier than by using traditional methods alone. This approach could make it easier and less invasive to find liver cancer early, offering a better chance for effective treatment and a hopeful prognosis for those at risk.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the most common primary liver cancer and a leading cause of cancer-related deaths worldwide, presenting significant challenges in early detection1,2,3. Traditional diagnostic methods, such as ultrasonography, CT, and MRI, often lack sensitivity for early-stage HCC, while biopsies carry risks and discomfort4,5,6,7. Alpha-fetoprotein (AFP), a commonly used serological marker, lacks satisfactory sensitivity in early-stage HCC4.
Examining cancer-specific DNA methylation markers in Circulating tumor DNA (ctDNA) is currently proposed as a preferred method for early detection, but this method faces limitations such as low abundance and technical complexity, which hinders its effectiveness in early cancer detection8,9,10. Highly sensitive techniques required to handle ctDNA add technical complexity. ctDNA is usually fragmented, and the process of bisulfite conversion for DNA methylation markers further complicates analyses due to its harsh conditions, which can lead to DNA degradation11. Recent advances suggest that enzymatic conversion might mitigate these limitations by preserving the quality and quantity of cfDNA, offering a promising alternative for methylation analysis12,13. This underscores the need for other robust and accessible diagnostic options. Peripheral blood mononuclear cells (PBMCs) provide a stable and abundant source of DNA, potentially carrying methylation signatures indicative of early-stage HCC.
Our recent research9 identified five DNA methylation biomarkers in plasma cell-free DNA (plasma-cfDNA) that are highly sensitive and specific for detecting HCC. While effective, these biomarkers demonstrate reduced sensitivity at the early stages of the disease, particularly at a 100% specificity threshold. The technical complexities associated with bisulfite conversion of plasma-cfDNA further complicate their use, prompting us to explore more feasible diagnostic alternatives. This includes combining plasma cfDNA with PBMCs to enhance early detection.
HCC’s relationship with the immune system is well-documented, with the disease known to induce identifiable changes in the host’s immune system, supporting the use of PBMCs for diagnosis14,15,16,17,18,19. Unlike ctDNA, PBMCs’ stable and ample DNA content simplifies the technical process of developing and validating molecular markers for HCC detection and staging.
In a previous study20, we analyzed the methylome of a diverse group of participants, including healthy individuals and patients with chronic hepatitis B, C, and various stages of HCC. This study identified distinct changes in the DNA methylation patterns of PBMCs and T cells, effectively differentiating early-stage HCC from controls and chronic hepatitis cases. These methylation disparities became more pronounced with the progression of cancer. We successfully identified and validated several methylation markers in PBMCs and T cells that are effective in detecting early stages of HCC.
Among the genes scrutinized, STAP1 and AHNAK emerged as promising candidates for further validation owing to their potential biomarker properties. Their significance was later affirmed in two additional patient cohorts from Beijing, thereby underlining the potential utility of these methylation markers in early HCC detection20.
STAP1 (Signal Transducing Adaptor Protein 1) and AHNAK (Neuroblast Differentiation-associated Protein) have been previously implicated in various biological processes relevant to cancer21. STAP1 was reported to recruit signaling pathway-related proteins involved in the B cell receptor signal transduction pathways responsible for immune regulation, potentially playing a role in tumorigenesis and immune responses21. AHNAK functions as a tumor suppressor in breast cancer by inhibiting cell growth via potentiation of the TGFβ (Transforming Growth Factor Beta) signaling pathway, which regulates cell growth, differentiation, and immune response22 and also lung tumor development23.
Building on these preliminary findings, we expanded our investigation to encompass a more extensive panel of genes. This selection was informed by their significance and stage-dependency, as evidenced in our previous study20, which showed promise in further preliminary analyses. This panel includes NBPF3 (Neuroblastoma Breakpoint Family, member 3). Although direct connections between NBPF3 and cancer have not been conclusively established, other members of the NBPF gene family have been implicated in oncogenesis, suggesting a potential, yet unconfirmed, role for NBPF3 in cancer development24.
GLRX2 (Glutaredoxin 2) plays a central role in mitochondrial redox regulation25. It is involved in response to oxidative stress and redox signaling in mitochondria26. GLRX2 functions as a glutathione-dependent oxidoreductase that is activated by oxidative stress27. GLRX2 facilitates the maintenance of mitochondrial redox homeostasis, protects cells from oxidative damage, and inhibits mitochondrially mediated apoptosis27. While GLRX2’s primary role is in redox homeostasis, it might indirectly influence immune responses due to its antioxidative function. Its potential involvement in cancer stems from the redox balance’s significance in DNA protection and tumor cell survival. Among the genes included in our panel for further investigation are RNMTL1P1 (RNA Methyltransferase Like 1 Pseudogene 1), VMP1 (Vacuole Membrane Protein 1) and MAP1LC3B2 (Microtubule Associated Protein 1 Light Chain 3 Beta 2). We also study cg0943517, located in an open sea region, far from any known genes.
This validation study embarks on a meticulous evaluation of an expanded 8-gene methylation marker panel in a well-characterized, independent cohort of 402 subjects spanning the spectrum of CHB and HCC (stage 0/A to D). By employing bisulfite next-generation sequencing for in-depth analyses, we aim to examine the clinical utility of these markers in HCC early detection and staging. This pivotal step seeks to facilitate timely therapeutic interventions and enhance patient prognoses through a more comprehensive understanding of methylation markers.
Although early detection of cancer could save lives, it is not generally employed as a public health measure since currently approved tests have a high rate of false positives and false negatives. Motivated by the aspiration to ease the diagnostic journey for HCC patients, often burdened by invasive procedures, we deliberately opt for a non-invasive blood-based assay. By leveraging the methylation signature in PBMCs, this study endeavors to introduce a novel, minimally invasive avenue for HCC detection and monitoring. The ensuing analysis aims to inch closer to a clinically viable, blood-based methylation assay for HCC, positioned to significantly augment current diagnostic and monitoring paradigms. Of particular interest is discovering a test that has a very high NPV and high sensitivity for early stages, which is currently unavailable.
In summary, the current study demonstrates that combining the M4 score (cumulative score based on the methylation levels of four genes: MAP1LC3B2, NBPF3, RNMTL1P1, and STAP1) with existing diagnostic tools such as AFP significantly enhances diagnostic sensitivity for early-stage HCC. Specifically, M4 + AFP achieves a sensitivity of 88.2% in Stage A and up to 95.7% across all stages. These results underscore the potential of integrating PBMC methylation markers with current serological and plasma-based markers to provide a more comprehensive and effective diagnostic approach for early-stage HCC.
Methods
Clinical study design
This observational study registered under ClinicalTrials.gov ID: NCT03483922, builds upon our prior work, as detailed in reference (11), employing the same cohort of 402 participants from the Dhaka area. These included 49 healthy controls, 51 patients with Chronic Hepatitis B, and 302 patients diagnosed with HCC across stages 0 to D (HCC 0 n = 2, HCC A n = 32, HCC B n = 86, HCC C n = 106, HCC D n = 76). Participants were compensated, and the study protocol was approved by the Institutional Review Board (IRB) of ICDDR,B under protocol number PR-18025. This approval covered both the current study, and the previous study described in our previously published paper9. All relevant ethical regulations were followed, and informed consent was obtained from all participants before their inclusion in the study.
The exclusion criteria for the study were as follows: unwillingness or inability to provide informed consent, unwillingness or inability to comply with the protocol requirements, participation in a different clinical trial investigating a vaccine, drug, medical device, or medicinal procedure less than four weeks preceding the current study, planned participation in another clinical trial during the present study period, known cases of cirrhosis, any other known inflammatory diseases (bacterial or viral infection with the exception of hepatitis B or C), known cases of diabetes, asthma, autoimmune disease, any other diagnosed cancer, and for healthy controls, any known inflammatory or infectious diseases including Hepatitis B and Hepatitis C, any diagnosis of chronic disease, cancer medication use or drugs of abuse. Furthermore, blood samples from non-HCC patients were similarly derived at ICDDR,B with exclusion criteria being the diagnosis of any liver disease, diagnosis of HCC or liver metastasis of other cancers.
For comprehensive demographic details of the participants and other information regarding this clinical study, refer to Table 19.
Participants provided informed consent for DNA methylation biomarker research. Blood sample collection and PBMC isolation were conducted at ICDDR,B in Dhaka, Bangladesh, before being shipped to HKG Epitherapeutics for further analysis. The lab team at HKG Epitherapeutics was blinded to the identity of the samples throughout the analytic procedures. Subsequently, data analysis was carried out in Montreal and shared with ICDDR,B, who relayed the results to the respective clinical personnel.
Methodological approach for sample allocation for training and validation cohorts
We randomly allocated 402 samples into training and validation cohorts, as detailed in Supplementary Data 1 and 2, using a “stratified random sampling” method28,29 (https://doi.org/10.5281/zenodo.10359896) using a 70:30 split ratio. The training cohort included 34 controls, 74 Stage C, and 53 Stage D patients (Supplementary Data 1), while the validation cohort comprised 15 controls, 51 CHB patients, and HCC patients across stages A (34), B (86), C (32), and D (23) (Supplementary Data 2).
To ensure that our methylation markers specifically identify HCC and are not confounded by CHB, we validated the specificity of the markers on “naïve” samples of CHB. This approach that validated our markers on a completely naive CHB cohort gives us higher confidence that the selected markers are HCC-specific markers and that they differentiate HCC from CHB. In addition, we assume that a certain fraction of the CHB patients might have converted to early HCC and were not yet clinically diagnosed. By including them in the training session, we might have confounded the “control” sample.
Additionally, we focused only on the later stages of cancer in the training set and included early stages (A and B) only in the validation set. Again, early stages samples were “naïve” to the markers trained on late-stage cancers, increasing our confidence in the robustness of the markers for early stages of cancer. The inclusion of early-stage cases (34) only in the validation cohort tests whether a model developed on later-stage data applies to early-stage disease. This approach ensures that the markers are effective across all stages of HCC, which is vital for early detection, where timely interventions can dramatically impact patient outcomes.
Data imputation
The empty values within the dataset were addressed through imputation, where the method of filling missing values was selected based on the nature of the data for each variable. For variables with a normal distribution or believed to have data missing at random, mean imputation was utilized. In cases of skewed data, median imputation was employed as a more robust measure of central tendency. For categorical variables, mode imputation was adopted to represent the most frequent category. Each imputation method was chosen with consideration to preserve the integrity of the data to the extent possible while preparing a comprehensive dataset for analysis.
Blood Collection, PBMC Separation, and DNA Extraction
Blood was collected in 9-ml tubes containing K3-EDTA and processed within one hour. Plasma and peripheral blood monocyte separation was performed according to GE Healthcare Cat No 71-7167-00 protocol. The PBMCs were frozen at −80 °C for a maximum of 5 months before being shipped on dry ice to the lab. Upon arrival at the lab, the cells were thawed, and DNA was extracted by the previously described guanidine isothiocyanate method30, followed by binding to silica magnetic beads, washing with 80% ethanol and elution with water.
Targeted bisulfite sequencing and methylation analysis
Bisulfite conversion was conducted utilizing the EZ-96 DNA Methylation MagPrep kit (D5041, Zymo Research, Irvine, CA, USA), which was followed by two sequential rounds of polymerase chain reaction. For the inaugural round, we employed primers incorporating an anchoring sequence along with sequences targeting a specified region encompassing the following genes and their corresponding CGs: GLRX2 (cg09182724), STAP1 (cg04398282), AHNAK1 (cg14171514), cg09435170, VMP1 (cg02782634), NBPF3 (cg14018420), RNMTL1P1 (cg08365438), MAP1LC3B2 (cg22164177). The amplification was executed using the Bio-Rad C1000 Touch Thermal Cycler (Bio-Rad Laboratories, CA, USA). The primers utilized for this procedure are available upon request. A 5 µL aliquot of the initial PCR amplification product was subjected to a secondary round of PCR amplification employing primers imbued with indexes for sample barcoding (primer sequences available upon request). Following amplification, PCR products were combined into a pooled library, which was subsequently purified on two occasions utilizing AMPure XP Beads (Beckman Coulter Life Sciences, CA, USA). Quantitative assessment of the purified library was performed via Real-Time PCR employing the NEBNext® Library Quant Kit for Illumina (New England Biolabs, MA, USA). Barcoded libraries, encompassing all samples, were sequenced on the Illumina platform utilizing the MiSeq Reagent Nano Kit V2 and a 250×2 paired-end sequencing protocol (Illumina, CA, USA).
For our targeted bisulfite sequencing experiments, we utilized Trim-galore (parameters: trim_galore –illumina –paired --fastqc) (https://zenodo.org/record/5127899#.Y7RxfOzMJqs) to trim sequencing adapters and low-quality data from the raw, paired-end reads and obtain clean data for subsequent analysis. We then aligned the clean data to eight reference genomes GLRX2 (cg09182724), STAP1 (cg04398282), AHNAK1 (cg14171514), cg09435170, VMP1 (cg02782634), NBPF3 (cg14018420), RNMTL1P1 (cg08365438), MAP1LC3B2 (cg22164177) using Bismark31 To mitigate the impact of PCR amplification biases and sequencing errors and to ensure a more accurate representation of the methylation status across the targeted genomic regions, deduplication was performed. This process was carried out using a custom script, which iterates through each pair of reads, identifies, and removes duplicates based on their UMIs (Unique Molecular Identifiers) and alignment positions. The full script is accessible on Zenodo https://doi.org/10.5281/zenodo.1000213132.
Using Bismark methylation extractor (bismark_methylation_extractor --p --bedGraph --counts --scaffolds --no_overlap), we calculated the methylation level at each CpG site by extracting methylation information from the aligned reads.
epiLiver Test Description
The epiLiver test utilizes plasma samples to analyze DNA methylation markers for the early detection of HCC, as described earlier9. This test examines specific CpG sites associated with key genes, including CHFR, VASH2, CCNJ, GRID2IP, and F12, using targeted bisulfite next-generation sequencing. It generates two primary scores: HCC-detect which includes DNA genes CHFR, VASH2, CCNJ, GRID2IP genes, designed for maximum sensitivity in identifying HCC, and HCC-spec, which focuses on the F12 gene and is optimized for high specificity by differentiating HCC from other liver conditions. This high-throughput assay ensures high specificity and sensitivity in detecting HCC among the study cohort.
Methodological Approach for Integrating Plasma-Based AFP and epiLiver Scores with PBMC-Derived M4 and M8 Markers in HCC Detection
In our investigation, detailed in ref. 9, we assessed the utility of Alpha-Fetoprotein (AFP) levels in plasma as a predictor for HCC. The AFP measurements, as detailed in our earlier work, were conducted using Chemiluminescence Immunoassay. Among our cohort of 302 HCC patients, AFP data were available for 282 individuals, while 20 cases did not have this information (details provided in Table 1 of Cheishvili et al. 9). Additionally, we included AFP levels from 22 Chronic Hepatitis B (CHB) patients and one healthy individual for comparative analysis.
We adhered to the AFP threshold of 400 ng/mL, a standard in HCC diagnostics. This threshold is informed by a substantial body of literature, including a comprehensive review that sifted through 29,828 articles, ultimately including 59 studies and one review covering 11,731 confirmed HCC patients and 21,972 controls33. In our methodology, a sample was deemed indicative of HCC when AFP levels exceeded this 400 ng/mL threshold.
Additionally, we integrated the analysis with the M4, M8, and epiLiver scores. The M4 and M8 scores are composite metrics derived from the methylation levels of specific genes to predict the presence of HCC. M8 Score includes eight genes: AHNAK1, GLRX2, MAP1LC3B2, NBPF3, RNMTL1P1, STAP1, VMP1, and cg09435170. Each gene’s methylation level is multiplied by a regression coefficient, and the products are summed to generate the M8 score. This score is designed to effectively discriminate between control and HCC cases. M4 score is a simplified version, incorporating only four genes from the M8 set: MAP1LC3B2, NBPF3, RNMTL1P1, and STAP1. Similar to the M8 score, the methylation levels of these four genes are weighted by their respective regression coefficients and summed to create the M4 score. Both scores were developed and validated using Receiver Operating Characteristic (ROC) analysis to determine their effectiveness in detecting early-stage HCC.
epiLiver score, based on the combined probability of “HCC-detect” and “HCC-spec9,” were assessed against a threshold of 1.955, associated with 100% specificity. This integration enabled a comprehensive comparison of the predictive efficacies of M4, M8, epiLiver, and AFP within the same cohort and control group while accounting for instances of missing AFP data.
Furthermore, we investigated a combined diagnostic approach. In this methodology, a sample was identified as HCC-positive if either the AFP level surpassed 400 ng/mL or if the prediction based on the epiLiver, M4, or M8 scores indicated HCC. This approach, seeking to enhance the accuracy of HCC detection, effectively leverages the strengths of both biochemical (AFP) and molecular (epiLiver, M4, M8) markers.
Statistics and reproducibility
Power analysis and sample size determination
In our prior work20, we identified specific CGs that accurately distinguished early HCC stages from chronic hepatitis B and C, as well as from healthy controls. Power calculations were carried out to determine the minimal sample size necessary for a follow-up clinical trial, targeting a power of 0.8. The analyses showed the requisite sample sizes for detecting the given delta methylation between non-cancerous (Controls, CHB, CHC) and all four HCC stages: 4 for GLRX2 (cg09182724) (delta −0.12598485), 9 for STAP1 (cg04398282) (delta 0.100254597), 9 for AHNAK (cg14171514) (delta −0.104702096), 7 for cg09435170 (delta −0.111705118), 1 for VMP1 (cg02782634) (delta −0.128756528), 3 for NBPF3 (cg14018420) (delta 0.09283704), 4 for RNMTL1P1 (cg08365438) (delta −0.054604218), and 4 for MAP1LC3B2 (cg22164177) (delta −0.130729187). The power analyses delineated the requisite sample sizes to detect the given delta methylation between non-cancerous (Controls, CHB, CHC) and all four HCC stages for each CG. The cohort used in our validation study, which consisted of 49 controls, 51 CHB, 34 HCC-0/HCC_A, 86 HCC_B, 106 HCC_C, and 76 HCC_D subjects, significantly exceeds the calculated sample sizes, thus providing robust validation for our study findings.
Cross-validation approach for model evaluation in cancer prediction
Cross-validation was executed using the “tidyverse” and “caret” R packages, following to the same analytical framework as outlined in our previous publication9,34 (https://doi.org/10.5281/zenodo.7823332).
The process involved a validation set approach, where the dataset was randomly divided into training and validation subsets. For this study, the data comprised 49 healthy controls, 51 CHB (chronic hepatitis B) patients, and 302 cancer individuals categorized across stages A to D. The cross-validation procedure was performed three times, each with a unique seed (123, 234, and 586), ensuring the training data encompassed 50% of the entire dataset in each instance. This approach allowed for the computation of model performance metrics, including R-squared (R2), Root Mean Square Error (RMSE), and Mean Absolute Error (MAE), as detailed in the results section. Leveraging the models derived from these three distinct training datasets, we evaluated the precision with which the model could predict cancer in the remaining validation dataset, a comparative analysis of which is presented in Supplementary Fig. 1.
All statistical analyses were conducted using R software (version 4.3.2), Prism (GraphPad Software, version 10.3.1) and Python (version 3.11.5).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Comparative analysis of gene-specific methylation profiles across HCC stages
After dividing the dataset into training and validation cohorts as described in the Methods, we conducted a Receiver Operating Characteristic (ROC) analysis on the median methylation levels of each gene within the training cohort. This analysis compared controls with combined Stages C and D to determine each gene’s discriminative capability. We established stringent thresholds for each gene, at specificities ranging from 97.06% to 100%, as detailed in Table 2. These thresholds were then applied in the validation cohort to assess the markers’ performance across various HCC stages and control groups.
In this validation study, conducted with a cohort from Bangladesh, we employed targeted next-generation sequencing (NGS) to analyze the median methylation levels of specific genes, thereby extending our comparison to a very different ethnic population from our prior study on a Chinese cohort20. Our analysis focused on genes AHNAK (cg14171514), STAP1 (cg04398282), NBPF3 (cg14018420), RNMTL1P1 (cg08365438), VMP1 (cg02782634), GLRX2 (cg09182724), MAP1LC3B2 (cg22164177), as well as the non-gene-specific cg09435170. These genes were selected based on their demonstrated performance as classifiers of HCC in our previous study20. This strategy enabled not only the validation of the clinical significance of these markers but also determining their performance across different ethnic groups.
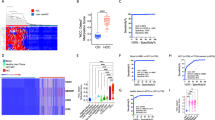
The amplification strategy expanded our analysis beyond the specific CG sites previously identified using the Illumina 450 K array20. We assessed the median methylation levels for each gene’s respective amplicons (Fig. 1A). As described in our previous findings20, STAP1 was highly significant. Additionally, a subset of genes including RNMTL1P1, GLRX2, NBPF3, and MAP1LC3B2, showed significant differences across all stages compared to the control group with p-values less than 0.0001, as depicted in Fig. 1A. VMP1 showed significant differences in all stages except for Stage A. Conversely, cg09435170 displayed no significant differences between the stages and the control group, except for Stage A where it was significant. AHNAK was significantly different when comparing Stage A to the control group (p = 0.0041) and showed borderline significance in Stage B (p = 0.1069).
For genes represented by a single CG site (STAP1 and cg0943517), methylation levels of that specific site were analyzed. Each data point represents an individual sample’s methylation value. Statistical comparisons were made using a nonparametric ANOVA (Kruskal-Wallis test) followed by Dunn’s multiple comparison test. Comparisons were made between the control groups (n = 49) and each disease stage: CHB (Chronic Hepatitis B) (n = 51), Stage A (n = 34), Stage B (n = 86), Stage C (n = 106), and Stage D (n = 76) for each gene. P-values indicating significant differences are noted accordingly. Source data are provided as a Source Data file.
In the validation cohort, each gene’s performance varied across different stages of HCC and control groups. Notably, NBPF3 showed the highest sensitivity across all HCC stages, with sensitivity values of 55.88% in HCC-A, 56.98% in HCC-B, 68.75% in HCC-C, and 60.87% in HCC-D. This was coupled with high specificity in CHB (96.08%) and control groups (93.33%).
RNMTL1P1 also displayed notable sensitivity, especially in the early stages, with values of 61.76% in HCC-A and 68.60% in HCC-B. However, its specificity was slightly lower than NBPF3.
STAP1, maintaining 100% specificity in the validation cohort, showed varying sensitivity across HCC stages, with 17.65% in HCC-A, 29.07% in HCC-B, 15.63% in HCC-C, and 26.09% in HCC-D.
Conversely, AHNAK and GLRX exhibited lower sensitivity across HCC stages. AHNAK’s sensitivity in the early stages A and B was 8.82% and 8.14%, respectively, while GLRX showed minimal sensitivity across all HCC stages, peaking at 2.94% in HCC-A. Nonetheless, both genes maintained high specificity.
The remaining genes, VMP, MAP1LC3B2, and cg09435170, showed mixed results. Their specificity and sensitivity varied across different HCC stages. For a detailed breakdown of each gene’s performance metrics across different stages and control groups, please refer to Table 2. This gene-specific analysis implied that the sensitivity of a single gene is insufficient, and a combinatory model ought to be developed.
Sequential refinement of gene scores for enhanced discrimination of HCC stages
Our approach to developing a robust metric for early detection and staging of HCC began with including all eight genes. We performed multilinear regression analysis on the training cohort (refer to Supplementary Data 1), employing a binary classification system: controls were labeled as zero, and all cancer stages as one. This analysis led to the development of a combined score, the M8-score, as detailed in Supplementary Data 3 (Summary output of 8 genes). The M8-score, calculated by aggregating the products of each gene’s median methylation level and its corresponding regression coefficient, effectively discriminated between control and cancer stages in the validation cohort. In this model, each coefficient reflects the importance of individual genes in distinguishing between different disease states.
Initial analysis using the Kruskal-Wallis test assessed the overall significant differences between methylation scores of the control and various disease stages (CHB, Stage A to D) in the validation cohort. Subsequently, Dunn’s multiple comparisons test was employed as a post-hoc analysis to pinpoint significant specific pairwise differences among the groups, as illustrated in Fig. 2A, B and detailed in Supplementary Data 2. This analysis revealed no significant difference between the control group and CHB (p > 0.9999). However, the differences between the control group and the cancer stages (Stages A to D) were significant, with increasing levels of significance correlating with the progression of the cancer stage (p-values from 0.0012 to <0.0001). These results emphasize the potential of our identified methylation markers in differentiating early and advanced stages of HCC from the control group and CHB.
A Scatter Plots of Methylation Scores (M8 and M4): Top Panel (Training Cohort): Displays the distribution of M8 and M4 scores for Stage C (n = 74) and Stage D (n = 53) of HCC (Hepatocellular Carcinoma) compared to the control group (n = 34). Each dot represents an individual sample, with horizontal lines indicating the median methylation score per group. Bottom Panel (Validation Cohort): Illustrates the distribution of M8 and M4 scores across all stages of HCC (Stage A (n = 34), Stage B (n = 86), Stage C (n = 32), Stage D (n = 23)), CHB (Chronic Hepatitis B) (n = 51), and control (n = 15) groups. Statistical analysis was performed using the Kruskal–Wallis test for multiple comparisons. Source data are provided as a Source Data file. B ROC Analysis Training Cohort: ROC curves for M8 (top left) and M4 (top right) combine stages C and D of HCC and are compared against the combined Control and CHB groups. Bottom Panel (Validation Cohort): Showcases ROC curves for M8 (bottom left) and M4 (bottom right), aggregating stages A, B, C, and D of HCC, juxtaposed with the combined Control and CHB groups for analysis.
After establishing the M8-score, we tested whether we could reduce the number of genes in our model. We used the median methylation of the genes (AHNAK, NBPF3, RNMTL1P1, VMP1, GLRX2, MAP1LC3B2) (Supplementary Data 1) alongside the single CG methylation levels of STAP1 and cg09435170, as presented in Supplementary Data 4. We used a multivariable linear regression analysis with the eight genes as independent variables to determine which genes are significant classifiers of HCC. We identified four significant genes (NBPF3, RNMTL1P1, MAP1LC3B2, and STAP1). We used a multilinear regression with these four genes to create the M4-score, a collective metric composed of the weighted coefficient of each of the four genes (refer to Fig. 2 and Supplementary Data 3).
To compare the performance of the M8 and M4 scores, both training and validation cohorts were examined. Initially, the Kruskal-Wallis test was conducted to assess overall differences between groups. Following this, Dunn’s multiple comparisons test was utilized to further appraise the discriminative efficacy of these scores across these cohorts, thoroughly examining their predictive capacities in differentiating HCC stages (Fig. 2A).
ROC analysis was then conducted within the training cohort to evaluate the discriminative ability of M8 and M4 scores to distinguish controls from HCC stages and established thresholds at 100% specificity and highest sensitivity for each M-score (Fig. 2B, Table 3).
In the validation cohort, both M8 and M4 scores demonstrated high specificity (100%) in the control group. Notably, the sensitivity for early-stage HCC (Stage A) was higher for the M4-score (73.53%) as compared to the M8 score (67.65%). Similar trends were observed in later stages, with the M4 score generally outperforming the M8 score.
A critical observation was that our M-scores results classified a subset of the CHB group (9.8% for the M8-score and 17.63% for the M4-score) as HCC. Given that 15%-40% of CHB individuals may develop HCC in their lifetime, particularly in Asian populations35, like our Bangladeshi cohort, this finding is significant. Alternatively, these results might just simply reflect the reduced specificity of the markers for CHB. To differentiate these two possibilities, a follow-up study is required.
Cross-validation analysis of model robustness in classifying HCC
Cross-validation was performed as outlined in the methods section, utilizing models derived from three distinct training datasets, each generated using different random seeds (123, 234, and 586). For the first training set (seed 123), the model exhibited an R-squared (R2) value of 1, with a Root Mean Square Error (RMSE) of 1.456573 and a Mean Absolute Error (MAE) of 1.358391. Similarly, the second training set (seed 234) yielded an R2 of 1, RMSE of 1.403377, and MAE of 1.34692. The third training set (seed 586) further corroborated the consistency of the model, demonstrating an R2 of 1, RMSE of 1.446507, and MAE of 1.406228. These results, indicating significant statistical differences, were consistent across all training instances, underscoring the robustness of the M8 and M4 model predictions in differentiating HCC cases from healthy control and chronic hepatitis B groups (Supplementary Fig. 1).
Enhanced HCC detection: integrating M8 and M4 PBMC methylation scores with epiLiver ctDNA analysis
To improve the diagnostic accuracy for HCC, our study implemented an integrated approach combining M8 and M4 scores, derived PBMC methylation markers (current study), with the epiLiver score based on ctDNA analysis9. This strategy aimed to harness the strengths of each marker, enhancing overall sensitivity while maintaining high specificity.
For each participant in the validation cohort, both the M-score and the epiLiver score were applied using the most stringent threshold for each score. Specifically, the M-scores were set at a threshold corresponding to 100% specificity based on ROC analysis of the training cohort. Similarly, the epiLiver score threshold was set at 100% specificity, using a sum of probabilities threshold of >1.995. At this stringent level, the sensitivity of the epiLiver score was lower than what was previously reported using a 95% specificity threshold9.
A positive cancer diagnosis was confirmed if either the M-score or the epiLiver score exceeded their respective thresholds. This combined methodology, referred to as M8+epiLiver and M4+epiLiver in our analysis (refer to Table 3), was specifically designed to improve the sensitivity of HCC detection without compromising specificity.
This integrated approach enhanced sensitivity for early-stage HCC. For instance, in Stage A HCC, the sensitivity of the M4+epiLiver score reached 79.4%, an improvement over the 73.5% for M4 alone and 26.5% for epiLiver. In Stage B HCC, the sensitivity of M4+epiLiver increased to 86.0%, exceeding the individual performances of epiLiver (48.8%), M8 (70.9%), and M4 (75.6%). These findings highlight the benefit of merging plasma-cfDNA markers with PBMC-derived markers.
An important aspect of our study was that the combined M8+epiLiver and M4+epiLiver scores were 100% specific. This is crucial for avoiding false-positive diagnoses, thereby enhancing the reliability of these tests.
In summary, the combination of epiLiver and PBMC M-scores represents a significant advancement. By integrating these scores, we increase sensitivity, particularly in the crucial early stages of HCC, without compromising specificity.
AFP and its integration with epiLiver, M8, and M4 scores
We next evaluated whether combining AFP, a classic HCC marker, with either epiLiver or the PBMC M4 and M8 scores and their combinations enhance the sensitivity of detection of HCC. Table 3 presents the performance metrics of AFP alone, as well as combined scores of M8, M4 with either epiLiver and AFP or just with AFP.
AFP alone demonstrated a moderate sensitivity across HCC stages, particularly in the early stages (Stage A), with a sensitivity of 62.5%, and the sensitivity improved when AFP was combined with either PBMC or ctDNA markers. For example, the combination of either M8 + AFP or M4 + AFP showed enhanced sensitivities of 85.3% and 88.2%, respectively, in Stage A, which is notably higher than AFP alone or the M4-score (73.53%) and the M8 score (67.65%) on their own. These combinations also performed exceptionally well in later stages (B–D), consistently achieving sensitivities between 90.7-96.9%, which is substantially higher than the 59.5-80.0% range observed with AFP alone or M4 and M8 alone (Supplementary Data 5 for summary).
Overall, the integration of PBMC M-scores with epiLiver and AFP significantly enhanced the sensitivity across all HCC stages while maintaining a high specificity, particularly among CHB patients. Among the various combinations, M4-epiLiver+AFP and M8-epiLiver+AFP demonstrated the highest sensitivity across later HCC stages, while M4 + AFP was highly sensitive at the early stage with no further enhancement in combination with epiLiver. The strong performance in early-stage HCC detection is crucial for timely clinical intervention. All combinations maintained 100% specificity.
We compared the reported sensitivities for traditional imaging techniques like ultrasound, either alone or in combination with alpha-fetoprotein (AFP), with ‘M4 + AFP’ and ‘M4-epiLiver+AFP’ markers highlighting the high performance of the combined methylation and AFP blood-based biomarkers described here (Supplementary Data 5).
We evaluated the diagnostic efficacy of the M4 + AFP combination for early-stage HCC, Stage A, by calculating the Positive Predictive Value (PPV) and Negative Predictive Value (NPV) as discussed earlier36. These calculations were informed by the prevalence of Hepatitis B Virus (HBV) and Hepatitis C Virus (HCV) in Bangladesh, as these conditions are known risk factors for HCC. The prevalence of HBV in the Bangladeshi population is estimated at approximately 5.4%, while HCV prevalence is around 0.84%37,38. Utilizing the sensitivity and specificity values of the M4 + AFP combination for Stage A HCC, as delineated in Table 4, along with the HBV and HCV prevalence rates, we calculated the PPV and NPV. The PPV reflects the likelihood that individuals with a positive M4 + AFP test truly have early-stage HCC, while the NPV indicates the likelihood that those with a negative test result genuinely do not have the condition.
Additionally, we calculated PPV and NPV independent of prevalence39. The results, presented in Table 5 when combining CHB patients and controls, demonstrate varying PPV and NPV across different markers and HCC stages, providing further insights into their diagnostic performance. For example, the M4 + AFP combination exhibited a PPV of 74% and an NPV of 86% for early-stage HCC (HCC-A). When the PPV is calculated when taking into consideration only healthy people as the control group, then the PPV across all tests is 100% (Table 5). This results from the detection of HCC by our test in HCB patients. It is unclear whether these are false negatives or true converters from HCB to HCC. Our calculations reveal a PPV of 100% and an NPV of 99.3% when considering the HBV prevalence. In the context of HCV prevalence, the PPV remains at 100%, with the NPV slightly higher at 99.9%.
In conclusion, the combination of M4 or M8 scores with either epiLiver and AFP markers, or both, significantly enhances HCC diagnostics, improving detection capabilities early in the disease and across all stages of HCC. Our findings indicate that M4 markers, which require fewer regions for analysis, perform comparably to M8 markers, suggesting an optimized approach that reduces complexity without compromising diagnostic efficacy. This streamlined methodology not only maximizes sensitivity but also maintains high specificity.
Specificity of M4 and M8 scores for HCC comparison to breast and prostate cancer
To validate the specificity of the genes included in the M4 and M8 scores for detecting HCC, we applied the M4 and M8 scores to DNA methylation data from PBMC derived from prostate and breast cancer patients. We used data from our earlier studies that examined breast cancer markers in T cells40 and prostate cancer methylation41. Although these data were derived from T cells, our previous study demonstrated equal performance whether T cells or PBMC were used20. We inspected the same CpGs for which methylation data were available in these two datasets. We observed that while AHNAK, STAP1, and cg09435170 sites displayed significant methylation differences between breast cancer (BrCa) and control groups, the significance was modest and much less pronounced than what we observed in HCC (Fig. 1A). For AHNAK, the p-value was 0.0218, for STAP1, the p-value was 0.02, and for cg09435170, the p-value was 0.02. STAP1 showed hypermethylation, while AHNAK and cg09435170 showed hypomethylation in BrCa. This trend paralleled our observations in liver cancer in PBMCs (current study) (Supplementary Fig. 2). Other examined sites, including VMP1 (P = 0.068), did not reach statistical significance.
We further investigated HCC genes in prostate cancer data. We examined AHNAK, STAP1, RNMTL1P1, VMP1, and MAP1LC3B2 (Fig. S3) and found no statistical significance when comparing healthy control individuals to prostate adenocarcinoma. Though this observation is quite encouraging and suggests that our HCC markers may be specific to HCC, it still requires more validation on larger datasets and across more cancer types.
Discussion
Our study focused on evaluating the diagnostic potential of PBMC-derived methylation markers in HCC, with an emphasis on high sensitivity across different stages of the disease. The obvious advantage of using white blood cells rather than cfDNA is the technical challenge in quantifying DNA methylation in extremely scarce plasma-cfDNA and the relative abundance of white blood cell DNA. Initially, we investigated a classifier incorporating a weighted coefficient of eight genes, known as the M8 score. Subsequently, we used a more streamlined model comprising four key genes, termed the M4 score. This simplification pursued two primary objectives: firstly, to enhance cost-effectiveness and ease of analysis by reducing the number of genes involved; secondly, to explore whether a smaller, more targeted set of significant genes could retain or even enhance the model’s predictive accuracy for HCC.
The M4 score demonstrated improved sensitivity across various stages of HCC compared to the M8 score, with sensitivities of 73.5% for Stage A, 75.6% for Stage B, 78.1% for Stage C, and 95.7% for Stage D, indicating its effectiveness in accurately detecting HCC from early to advanced stages. Given the reduced number of genes in the M4 score, developing a multiplex PCR or digital PCR assay could offer a faster and more cost-effective alternative to methylation-specific sequencing. This approach would streamline the diagnostic process, making it more accessible for routine clinical use while maintaining the accuracy of the M4 score.
The integration of M-scores with the previously described epiLiver9 significantly enhanced sensitivity without compromising specificity. Specifically, in Stage A HCC, the sensitivity of the M4+epiLiver score at a 100% specificity threshold reached 79.4%, compared to 73.5% for M4 alone and 26.5% for epiLiver. In Stage B HCC, this sensitivity further improved to 86.0% for M4+epiLiver, outperforming the individual performances of epiLiver (48.8%), M8 (70.9%), and M4 (75.6%). Additionally, in later stages C and D, the M4+epiLiver scores showed sensitivities of 93.8% and 95.7%, respectively, indicating a consistent enhancement across various HCC stages. These results highlight the diagnostic advantage of combining cfDNA methylation and PBMC DNA methylation. By maintaining a high specificity threshold of 100% for each biomarker set, we achieved enhanced sensitivity while ensuring the robustness and reliability of the diagnostic approach.
Combining M8 and M4 scores with epiLiver and AFP markers improved sensitivity without sacrificing specificity, and it particularly excelled in early-stage HCC detection. The M4 + AFP combination, for instance, achieved a sensitivity of 88.2% in early-stage HCC (Stage A/0), significantly higher than M4 alone (73.5%), AFP alone (62.5%) or epiLiver (26.5%). This increased sensitivity was consistent across all HCC stages, with the M4 + AFP reaching 89.5% in Stage B and the M4-epiLiver-AFP combination showing sensitivities of 96.9% and 95.7% in Stages C and D. These findings suggest that the combination of PBMC-based markers with AFP could offer a diagnostic approach for early HCC identification, especially in populations at risk due to HBV and HCV.
In addition to the sensitivity and specificity metrics, the Positive Predictive Value (PPV) and Negative Predictive Value (NPV) are critical for assessing the real-world utility of our M4 + AFP combination for early-stage HCC detection. These values are especially significant in clinical settings where disease prevalence influences diagnostic accuracy.
To contextualize these findings within the Bangladeshi population, where Hepatitis B Virus (HBV) and Hepatitis C Virus (HCV) are prevalent risk factors for HCC, we calculated the PPV and NPV. For HBV prevalence, the M4 + AFP combination shows a PPV of 100% and an NPV of 99.3%. In the context of HCV prevalence, these values remain impressively high, with a PPV of 100% and an NPV of 99.9%. These results underscore the M4 + AFP combination’s effectiveness as a non-invasive diagnostic tool for early HCC detection, particularly in populations at risk due to HBV and HCV.
Therefore, while traditional methods such as AFP alone may fall short in sensitivity and specificity, our integrated approach employing the M4 + AFP combination overcomes these limitations.
Our anecdotal observations from a limited sample of two patients with very early-stage HCC (Stage 0) suggested that the PBMC-based M scores might have potential for early detection. These results, while preliminary, indicate a possible advantage of PBMC markers in early-stage HCC detection, warranting further investigation.
In our study, we observed that several chronic hepatitis B (CHB) patients had M scores within the range typically indicative of HCC. This observation may be linked to the well-documented increased risk of HCC among individuals with CHB. Specifically, the M8 and M4 scores identified potential HCC in 9.8% and 17.6% of CHB patients, respectively. These percentages are notable, considering epidemiological data suggest that 15%-40% of CHB patients may develop HCC in their lifetime35, particularly in regions like Bangladesh, where our cohort was based. Such findings might underscore the potential utility of M scores in the early detection of HCC among high-risk groups, which is crucial for timely intervention that can significantly improve patient outcomes. However, it is important to recognize that these findings on their own do not confirm the presence of HCC. The presence of elevated M scores in CHB patients might alternatively reflect the reduced specificity of our markers within this subgroup. Without follow-up data, we cannot conclusively determine whether the elevated M scores represent true HCC cases or false positives. Therefore, these results emphasize the need for further studies to assess the specificity of these markers and to confirm their predictive value in this patient population. Prospective studies will be essential to clarify these findings.
Like any study, ours has its set of limitations. The number of participants, though adequate as per statistical power calculations, and the varied characteristics within the chronic hepatitis B group may affect the accuracy and reliability of our results. Also, the age differences among the healthy controls, hepatitis B, and HCC patients, coupled with the study being conducted solely in Bangladesh, may limit how applicable our findings are to other populations. However, the fact that markers that were originally detected in a Chinese population performed so well in genetically distant Indian populations is encouraging.
One notable limitation is the availability of AFP data. AFP testing was conducted based on clinical indications at the physician’s discretion, resulting in AFP data from only one healthy individual. Additionally, while AFP data were available for a subset of the CHB group, this does not provide a comprehensive control group for comparison. This limitation impacts the calculation of specificity for the combined biomarker and necessitates a cautious interpretation of the results. The absence of AFP data for the majority of healthy individuals and the selective availability in the CHB group prevents meaningful comparison and accurate specificity estimation.
Our study’s cross-sectional design underscores the need for longitudinal studies to fully grasp the changes in methylation across different stages of HCC and chronic hepatitis B evolution.
Additionally, our study did not examine how inflammation or other liver diseases might influence the sensitivity and specificity of the M4 and M8 biomarkers. This question would need to be resolved by studying samples from patients with other liver diseases or chronic inflammatory conditions. However, we do show that common chronic liver inflammatory conditions such as chronic hepatitis B could be differentiated from Stage A HCC using these markers.
While our study demonstrated promising results for the early detection of HCC using specific DNA methylation markers, it’s crucial to consider the specificity of these markers for HCC versus other cancers. Nevertheless, analysis of DNA methylation in T cells from prostate and breast cancer suggests specificity of our markers for HCC. Testing the marker panel across a diverse set of cancers will clarify its specificity and enhance its clinical utility.
The identified methylation markers offer a potential augmentation to existing HCC diagnostic algorithms, particularly in early detection, and perhaps replacing cfDNA markers altogether with PBMC markers in combination with AFP. Integration might be envisioned through a diagnostic panel that melds these markers with existing imaging and serological assays, enhancing accuracy in diagnosis and risk stratification. However, a structured pathway toward clinical translation, akin to the integration of High-Sensitivity Troponin T (hs-TnT) in Acute Coronary Syndrome diagnostic algorithms42,43, is imperative. This entails clinical validation of the markers, algorithm development, guideline revisions, clinician training, and continuous evaluation, addressing challenges at each juncture for successful implementation.
Looking ahead, a more extensive validation of the identified methylation markers in larger and heterogeneous cohorts is imperative. Longitudinal analysis could further illuminate the methylation dynamics and the pivotal role these markers play in HCC progression. Delving deeper into the interaction between methylation markers and immune modulation could unveil more layers, offering a clearer diagnostic picture of HCC.
The bar graph presented illustrates the predictive performance of different score combinations in identifying HCC (Hepatocellular Carcinoma). The combinations encompass standalone M-scores (M8, M5, M3, M2), M-scores combined with epiLiver (M8+epiLiver, M5+epiLiver, M3+epiLiver, M2+epiLiver), and a further integration with AFP (Alpha-Fetoprotein) (M8+epiLiver+AFP, M5+epiLiver+AFP, M3+epiLiver+AFP, M2+epiLiver+AFP). Note: in the combinations with AFP, control samples did not have AFP data. Therefore, predictions for control samples in these combinations were derived from the M-scores combined with epiLiver only, excluding AFP. Source data are provided as a Source Data file.
In summary, our study illuminates the potential of PBMC methylation markers in broadening the diagnostic spectrum for HCC. The high sensitivity rates, particularly at the best-performing M4-epiLiver and M4 + AFP score, underscore the prospect of devising a more precise and early detection paradigm for HCC. This methodology could significantly propel timely clinical interventions, potentially ameliorating the prognosis for patients with HCC.
Data availability
The data supporting the findings of this study have been deposited in the Sequence Read Archive (SRA) PRJNA1067580 and are accessible under controlled conditions. Acquiring the data necessitates a material transfer agreement (MTA) and is restricted solely to academic utilization. For further details and to request data access, please contact David Cheishvili at david.cheishvili@epimedtech.com. The source data for Figs. 1, 2, and 3 are available in Supplementary Data 6.
Code availability
All codes are available on Zenodo:
The R code for cross-validation can be downloaded: https://doi.org/10.5281/zenodo.782333234
The code for UMI BAM deduplication is also available:
https://doi.org/10.5281/zenodo.1000213132
The code for stratified random sampling is accessible at: https://doi.org/10.5281/zenodo.1035989629.
References
Dhanasekaran, R., Limaye, A. & Cabrera, R. Hepatocellular carcinoma: current trends in worldwide epidemiology, risk factors, diagnosis, and therapeutics. Hepat. Med. 4, 19–37 (2012).
Yang, J. D. et al. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 16, 589–604 (2019).
Yin, Z. et al. Multipotent mesenchymal stromal cells play critical roles in hepatocellular carcinoma initiation, progression and therapy. Mol. Cancer 17, 178 (2018).
Bruix, J., Sherman, M. & American Association for the Study of Liver. Management of hepatocellular carcinoma: an update. Hepatology 53, 1020–1022 (2011).
Chen, T. Circulating non-coding RNAs as potential diagnostic biomarkers in hepatocellular carcinoma. J. Hepatocell. Carcinoma 9, 1029–1040 (2022).
Kirnap, M. et al. Evaluation of safety and efficacy of liver biopsy following liver transplant. Exp. Clin. Transpl. 13, 312–314 (2015).
Rockey, D. C. et al. Liver biopsy. Hepatology 49, 1017–1044 (2009).
Tang, J. C. et al. Circulating tumor DNA in hepatocellular carcinoma: trends and challenges. Cell Biosci. 6, 32 (2016).
Cheishvili, D. et al. A high-throughput test enables specific detection of hepatocellular carcinoma. Nat. Commun. 14, 3306 (2023).
Andersson, D. et al. Ultrasensitive circulating tumor DNA analysis enables precision medicine: experimental workflow considerations. Expert Rev. Mol. Diagn. 21, 299–310 (2021).
Liu, Y. et al. Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution. Nat. Biotechnol. 37, 424–429 (2019).
Vaisvila, R. et al. Enzymatic methyl sequencing detects DNA methylation at single-base resolution from picograms of DNA. Genome Res. 31, 1280–1289 (2021).
Ahn, J. et al. Differentially hypomethylated cell-free DNA and coronary collateral circulation. Clin. Epigenet. 4, 140 (2022).
Blair, G. E. & Cook, G. P. Cancer and the immune system: an overview. Oncogene 27, 5868 (2008).
Ehrlich, P. Über den jetzigen Stand der Karzinomforschung. 1908, Amsterdam.
Varayathu, H. et al. Combination strategies to augment immune check point inhibitors efficacy—implications for translational research. Front. Oncol. 11, 559161 (2021).
Vaddepally, R. K. et al. Review of indications of FDA-approved immune checkpoint inhibitors per NCCN guidelines with the level of evidence. Cancers (Basel) 12, 738 (2020).
Yu, Y. & Cui, J. Present and future of cancer immunotherapy: a tumor microenvironmental perspective. Oncol. Lett. 16, 4105–4113 (2018).
Luke, J. J. et al. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat. Rev. Clin. Oncol. 14, 463–482 (2017).
Zhang, Y. et al. The signature of liver cancer in immune cells DNA methylation. Clin. Epigenet. 10, 8 (2018).
Yang, X. et al. Signal-transducing adaptor protein 1 (STAP1) in microglia promotes the malignant progression of glioma. J. Neurooncol. 164, 127–139 (2023).
Lee, I. H. et al. Ahnak functions as a tumor suppressor via modulation of TGFbeta/Smad signaling pathway. Oncogene 33, 4675–4684 (2014).
Park, J. W. et al. AHNAK loss in mice promotes type II pneumocyte hyperplasia and lung tumor development. Mol. Cancer Res 16, 1287–1298 (2018).
Vandepoele, K. et al. A novel gene family NBPF: intricate structure generated by gene duplications during primate evolution. Mol. Biol. Evol. 22, 2265–2274 (2005).
Johansson, C. et al. Reversible sequestration of active site cysteines in a 2Fe-2S-bridged dimer provides a mechanism for glutaredoxin 2 regulation in human mitochondria. J. Biol. Chem. 282, 3077–3082 (2007).
Seco-Cervera, M. et al. Thioredoxin and glutaredoxin systems as potential targets for the development of new treatments in Friedreich’s Ataxia. Antioxidants (Basel) 9, 1257 (2020).
Coppe, A. et al. Genome evolution in the cold: Antarctic icefish muscle transcriptome reveals selective duplications increasing mitochondrial function. Genome Biol. Evol. 5, 45–60 (2013).
Kernan, W. N. et al. Stratified randomization for clinical trials. J. Clin. Epidemiol. 52, 19–26 (1999).
Cheishvili, D. Clinical validation of peripheral blood mononuclear cell DNA methylation markers for accurate early detection of hepatocellular carcinoma. https://doi.org/10.5281/zenodo.10359896 (2023).
Coombs, L. M. et al. Simultaneous isolation of DNA, RNA, and antigenic protein exhibiting kinase activity from small tumor samples using guanidine isothiocyanate. Anal. Biochem 188, 338–343 (1990).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Cheishvili, D. UMIBamDeduplicator. https://doi.org/10.5281/zenodo.10002131 (2023).
Zhang, J. et al. The threshold of alpha-fetoprotein (AFP) for the diagnosis of hepatocellular carcinoma: a systematic review and meta-analysis. PLoS ONE 15, e0228857 (2020).
Cheishvili, D. R script for linear regression analysis with cross-validation using the tidyverse and caret libraries (2023).
Lu, S. et al. Secular Trends and Geographic Variations of Hepatitis B Virus and Hepatitis C Virus‐associated Hepatocellular Carcinoma in Taiwan. Int. J. Cancer 119, 1946–1952 (2006).
Tenny, S. & Hoffman, M. R. Prevalence (StatPearls, 2023).
Mahtab, M. A. et al. Epidemiology of hepatitis B virus in Bangladeshi general population. Hepatobiliary Pancreat. Dis. Int. 7, 595–600 (2008).
Mamun Al, M. et al. Prevalence and risk factors of asymptomatic hepatitis C virus infection in Bangladesh. J. Clin. Exp. Hepatol. 1, 13–16 (2011).
Trevethan, R. Sensitivity, specificity, and predictive values: foundations, pliabilities, and pitfalls in research and practice. Front. Public Health 5, 307 (2017).
Parashar, S. et al. DNA methylation signatures of breast cancer in peripheral T-cells. BMC Cancer 18, 574 (2018).
Mehdi, A. et al. DNA methylation signatures of Prostate Cancer in peripheral T-cells. BMC Cancer 20, 588 (2020).
Tan, W. C. J. et al. The Asia-Pacific Society of Cardiology (APSC) Expert Committee Consensus Recommendations for Assessment of Suspected Acute Coronary Syndrome Using High-Sensitivity Cardiac Troponin T in the Emergency Department. Circ. J. 84, 136–143 (2020).
Zhelev, Z. et al. Diagnostic accuracy of single baseline measurement of Elecsys Troponin T high-sensitive assay for diagnosis of acute myocardial infarction in emergency department: systematic review and meta-analysis. BMJ 350, h15 (2015).
Author information
Authors and Affiliations
Contributions
D.C. contributed to conceptualization, investigation, formal analysis, writing (original draft)., and visualization. M.S. was involved in conceptualization, investigation, writing (review and editing), and supervision. C.W. contributed to the methodology. W.A.K. and M.A.M. provided resources and project administration. M.M.K., M.G.K., N.J., P.C.D., A.K.Y., A.I., D.C.D., S.M.N.A., S.A., MR contributed resources to the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: D.C. and C.W. are inventors on a patent application (PCT/IB2019/055855) titled “Methods for predicting or detecting hepatocellular carcinoma”, which includes the tests disclosed in the paper. M.S. is a shareholder in HKG Epitherapeutics Ltd. All other authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheishvili, D., Wong, C., Karim, M.M. et al. Clinical validation of peripheral blood mononuclear cell DNA methylation markers for accurate early detection of hepatocellular carcinoma in Asian patients. Commun Med 4, 220 (2024). https://doi.org/10.1038/s43856-024-00652-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-024-00652-2