Abstract
Background
The post-acute sequelae of SARS-CoV-2 (PASC), also known as long COVID, remain a significant health issue that is incompletely understood. Predicting which acutely infected individuals will develop long COVID is challenging due to the absence of established biomarkers, clear disease mechanisms, or well-defined sub-phenotypes. Machine learning (ML) models may address this gap by leveraging clinical data to enhance diagnostic precision.
Methods
Clinical data, including antibody titers and viral load measurements collected at the time of hospital admission, are used to predict the likelihood of acute COVID-19 progressing to long COVID. Machine learning models are trained and evaluated for predictive performance. Feature importance analysis is performed to identify the most influential predictors.
Results
The machine learning models achieve median AUROC values ranging from 0.64 to 0.66 and AUPRC values between 0.51 and 0.54, demonstrating predictive capabilities. Low antibody titers and high viral loads at hospital admission emerge as the strongest predictors of long COVID outcomes. Comorbidities—such as chronic respiratory, cardiac, and neurologic diseases—and female sex are also identified as significant risk factors.
Conclusions
Machine learning models identify patients at risk for developing long COVID based on baseline clinical characteristics. These models guide early interventions, improve patient outcomes, and mitigate the long-term public health impacts of SARS-CoV-2.
Plain language summary
Long COVID, or post-acute sequelae of SARS-CoV-2, is a prolonged health condition that can occur after acute COVID-19 infection. However, the ability to predict who will develop long COVID remains limited due to the absence of clear tests or biomarkers. We looked at patients’ medical information, including the amount of virus in their body at hospital admission, and how strong their immune response was. Using computer programs that can find hidden patterns in large sets of data, we discovered that people with a weaker immune response, higher amounts of virus, certain long term health problems and women are more likely to develop long COVID. This study highlights that computer-based tools could help doctors identify high-risk patients early and provide care that may prevent long-term complications.
Similar content being viewed by others
Introduction
The definition of post-acute sequelae of SARS-CoV-2 infection (PASC), or post-COVID-19 syndrome (Long COVID) has been evolving since its initial characterization. The United States Centers for Disease Control and Prevention (CDC) define long COVID as a range of overlapping physical and cognitive symptoms that present for 4 weeks or more after the onset of infection, and may last for weeks, months or even years1. Approximately, one in seven US adults have experienced at least one symptom associated with long COVID2, while estimates in the United Kingdom suggest that around 2 million individuals have been affected by this condition3.
Identifying factors that are predictive of long COVID remains a significant challenge4. Machine learning (ML) models have shown great success in identifying key features from routine laboratory measurements, patient demographics, clinical parameters based on electronic health records (EHR) for the diagnosis, prognosis and prediction of mortality in acute COVID-19 disease5,6,7,8,9,10,11. Huyut et al. explored various combinations of laboratory biomarkers from routine blood measurements to predict COVID-19 risk and mortality across different patient cohorts. Their analysis identified elevated levels of procalcitonin in combination with D-dimer, erythrocyte sedimentation rate (ESR), direct bilirubin (D-Bil) and ferritin as biomarkers predictive of mortality in COVID-195,7. In a separate study incorporating blood biomarkers alongside patient demographics and comorbidities, they found that older age combined with elevated international normalized ratio (INR), D-dimer and serum ferritin levels were significant risk factors for COVID-19 related death6,12.
While few studies have attempted to use ML approaches to develop predictive models for long COVID outcomes using EHRs, these studies have varied in their population characteristics and definitions of long COVID13,14. Potential predictive factors include age, sex, specific symptoms, comorbidities and health care utilization patterns. However, most published models have relied on non-curated EHR data and have not incorporated SARS-CoV-2 specific laboratory markers, such as antibody titers or viral load measurements.
In this study, we present a predictive model for long COVID using a large, well-characterized cohort of patients with COVID-19 who were hospitalized across the United States early in the pandemic. This cohort is part of the IMmuno Phenotyping Assessment in a COVID-19 Cohort (IMPACC) study. Our model integrates demographic, clinical, and laboratory data collected at the time of hospitalization for acute SARS-CoV-2 infection, aiming to improve the identification of patients at risk for developing long COVID. Although comorbidities and laboratory tests have been associated with acute COVID-19 outcomes, there remains a critical need to define which early clinical features predict the development of post-acute sequelae (PASC). This study addresses that gap by leveraging machine learning models to identify baseline predictors of long COVID in hospitalized adults.
Methods
Study design and setting
The study followed the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines for reporting observational studies15. The design of the IMPACC study has been previously published16. This study was registered at clinicaltrials.gov (NCT0438777). Biologic samples collected consisted of blood and mid-turbinate nasal swabs (self, or staff collected; with step-by-step instructions provided to both staff and study participants). The timepoints were as follows: enrollment (Day 1), and Days 4, 7, 14, 21, and 28 post hospital admission (and if feasible, for discharged participants, Days 14, and 28) and 3, 6, 9 and 12 months after discharge. Only baseline data was used for this modeling. Selected information was available after discharge for self-reported vaccination status, recurrent SARS-CoV-2 infection, and persistence of symptoms, as well as mortality.
Study participants
Patients 18 years and older admitted to 20 US hospitals (affiliated with 15 academic institutions) between May 2020 and March 2021 were enrolled within 72 h of hospital admission for COVID-19 infection. Only confirmed positive SARS-CoV-2 PCR and symptomatic cases attributable to COVID-19 infection were followed longitudinally. Participants did not receive compensation for study participation while inpatient, and subsequently were offered compensation during outpatient follow-up visits. The study was designed to enroll participants of both sexes, and sex at birth was collected based on self-report or care giver report.
Data collection and study variables
Specific data elements were acquired via review of electronic medical records during the inpatient period: participant demographic characteristics (age, sex at birth, race, ethnicity) comorbidities and body mass index (BMI), presenting signs/symptoms and onset, baseline diagnostic investigations (predefined laboratory values, and radiographic findings), baseline oxygen- and ventilatory-support requirement [either a) not requiring supplemental oxygen, b) requiring oxygen, c) non-invasive ventilation, or high-flow oxygen, or d) invasive mechanical ventilation, and/or extracorporeal membrane oxygenation (ECMO)], relevant clinical outcomes such as length of stay and complications as well as medications used to treat COVID-19 while inpatient.
In addition, self-reported symptoms and standardized patient reported outcome surveys were assessed quarterly for the duration of the study up to 12 months after initial hospital discharge. Patient-reported data was collected using a comprehensive digital remote monitoring tool, in the form of a mobile application developed by My Own Med, Inc. Along with the mobile application, an administrative portal was developed to collect information by study personnel during site visits or via telephone interviews by a study coordinator to ensure real-time electronic data capture. The surveys administered at these remote visits included reporting of the following symptoms: upper respiratory symptoms (sore throat, conjunctivitis/red eyes), cardiopulmonary symptoms (shortness of breath (dyspnea), cough), systemic symptoms (fever, chills, fatigue/malaise, muscle aches (myalgia)), neurologic symptoms (loss of smell/taste (anosmia/ageusia), headache), and gastrointestinal symptoms (nausea/vomiting).
In addition, the functional assessments of general health and the evaluation of deficits in specific health domains were conducted using validated Patient-Reported Outcome (PRO) measures, including: EQ-5D-5L, a standardized, self-administered instrument that describes and quantifies health-related quality of life17,18,19,20. Patient-Reported Outcomes Measurement Information System (PROMIS). The PROMIS measures administered included: PROMIS® Item Bank v2.0—Physical Function and PROMIS Item Bank v2.0—Cognitive Function, two computer adaptive surveys with tailored questionnaires based on item response theory21. PROMIS Scale v1.2 - Global Health Mental 2a and PROMIS Item Bank v1.0 - Psychosocial Illness Impact – Positive - Short Form 8a, two surveys with fixed questions18,19,20,22. PROMIS Pool v1.0—Dyspnea Time Extension computer adaptive instrument for participants who reported shortness of breath23,24,25. This 7-item questionnaire assesses whether there has been a meaningful increase or decrease in the duration of time needed by an adult to perform a given task in the past 7 days compared to 3 months ago due to shortness of breath.
For all PROMIS measures, scoring was based on PROMIS standardized instructions and conversion to a t-statistic26. Health Recovery Score: Overall health was also assessed by a health recovery score utilizing a Visual Analog Scale of 1-100 to indicate overall physical and mental function compared to pre-COVID function.
All data were reviewed centrally to ensure accuracy and consistency. Any data concerns were resolved by querying the site. The full study data collection forms for the quarterly outpatient surveys are provided in the online supplement (Surveys Administered).
The full study data collection forms and deidentified data are available upon request.
Assays
SARS-CoV-2 viral load was assessed by a central laboratory from nasal swab samples at each time point by RT-PCR of the viral N1, and N2 genes (see online supplement)27. Anti-SARS-CoV-2 spike (S), and receptor binding domain (RBD) antibodies were quantified by enzyme-linked immunosorbent assay (ELISA) in serum specimens28.
Statistics and reproducibility
We report median (interquartile range, IQR) for continuous variables and frequency (percent) for categorical variables. We examined bivariate associations between demographic and clinical factors and the two PRO clusters with minimal group representing non long COVID and deficit group representing long COVID using Wilcoxon rank sum test for continuous variables and chi-square test for categorical variables. P < 0.05 was considered statistically significant.
Machine learning modeling feature selection
We used all the features available at the time of hospital admission, including patient demographics, comorbidities, symptoms, baseline clinical characteristics, baseline lab measurements. In addition, we used viral levels (N1 Ct) and antibody measurements (anti-Spike IgG) both collected at the time of (Visit 1) of hospital admission. In total we had access to 93 features. Categorical variables were converted to one-hot encoded. We excluded participants with missing data for any of the variables to ensure the integrity and completeness of the dataset. After this exclusion process, a total of 385 participants remained available for inclusion in the modeling analysis. Feature selection is a commonly used technique in predictive machine learning models to reduce the feature space by removing irrelevant and redundant features. We performed feature selection on only a training data set using LASSO (Least Absolute Shrinkage and Selection Operator) logistic regression. LASSO includes the l1 norm of feature coefficients as a penalty term to the loss function which forces the coefficients for those features with weak associations to zero. We chose features with non-zero coefficients using 10-fold cross validation. To further reduce the features, we repeated this process 100 times and retained only those features that appeared >80 times out of 100 repetitions. We used only those features as input to train all the machine learning models. The features anti-Spike IgG, viral load N1 CT are modeled as continuous variables and all other features as binary variables.
Outcome variable: long COVID phenotypes
Long COVID outcomes were defined using previously established patient-reported outcome (PRO) clusters derived from longitudinal assessments conducted up to 12 months post-hospital discharge. These clusters were generated using latent class and clustering analysis and included four distinct phenotypes: MIN: Individuals with no or minimal functional deficits, COG: Individuals with predominant cognitive or mental health deficits, PHY: Individuals with predominant physical deficits, MLT: Individuals with multi-domain or widespread deficits. For the purposes of our analysis, we consolidated the three deficit clusters (COG, PHY, and MLT) into a single group representing individuals with long COVID symptoms (referred to as the Deficit group). The MIN cluster was used to define individuals without significant post-acute sequelae (referred to as the Minimal group). This binary classification—Minimal vs. Deficit—was used as the primary outcome variable in all predictive modeling analyses.
Machine learning model development
We trained machine learning models to predict the risk of developing a long COVID phenotype, “Deficit” versus non-long COVID phenotype, “Minimal”. The R package, Caret, was used to train, fine-tune and build final models. Hyperparameters for each model were tuned using tuneGrid functionality from caret package with three-fold cross-validation, set to optimize the area under the receiver operating characteristic curve (AUROC). Each model’s performance was evaluated using AUROC and AUPRC (area under precision-recall curve) on set-aside testing data, repeated fifteen times. To interpret the models, we computed the relative importance of all the features included in the model training using varImp functionality from caret.
Ethics approval
NIAID staff conferred with the Department of Health and Human Services Office for Human Research Protections (OHRP) regarding potential applicability of the public health surveillance exception [45CFR46.102(l) (2)] to the IMPACC study protocol. OHRP concurred that the study satisfied criteria for the public health surveillance exception, and the IMPACC study team sent the study protocol, and participant information sheet for review, and assessment to institutional review boards (IRBs) at participating institutions. Twelve institutions elected to conduct the study as public health surveillance, while 3 sites with prior IRB-approved biobanking protocols elected to integrate and conduct IMPACC under their institutional protocols (The University of Texas at Austin, IRB 2020-04-0117; University of California San Francisco, IRB 20-30497; Case Western Reserve University, IRB STUDY20200573) with informed consent requirements. Participants enrolled under the public health surveillance exclusion were provided information sheets describing the study, samples to be collected, and plans for data de-identification, and use. Those that requested not to participate after reviewing the information sheet were not enrolled.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Demographics and descriptive statistics
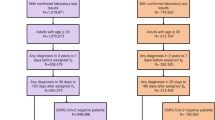
1164 participants were enrolled between May 5th, 2020 and March 19th, 2021 and followed up to 28 days post hospitalization, then at quarterly intervals for up to 12 months post hospital discharged. As previously reported, 590/702 (84%) who were alive at least 3 months post hospital discharge completed at least one quarterly set of symptom and patient-reported outcome (PRO) surveys. This group differed from non-responders to the surveys by having shorter hospitalization or fewer limitations at discharge (42% versus 57%). PASC was defined in this cohort as previously described with patient reported outcomes of clinical deficits (Ozonoff et al.)29. Data from 385 participants was available for modeling after removing missing data. Of these 385, 238 (61.8%) had minimal or no deficit by PRO, while 147 (38.2%) did have a significant deficit and were therefore labeled as long COVID. Supplementary Table 1 details the demographics, clinical characteristics, baseline radiographic and laboratory findings of this subset of the cohort. Median age was 57 years (IQR 20.0) and did not differ between those with and without evidence of long COVID. The analysis cohort was predominantly male (n = 241; 63%), though more females than males reported long COVID (47% vs. 33%, p = 0.005). Slightly more participants reporting long COVID were white 59% vs 52%, p = 0.049), but ethnicity did not differ significantly. 93% of participants had at least one comorbidity, and the presence of a co-morbidity was higher in those with long COVID (p < 0.001). Hypertension, diabetes mellitus, chronic lung disease (not asthma) and chronic cardiac disease were the most common comorbid conditions reported, and all were more prevalent in participants classified as having long COVID. The majority of the analysis cohort (86%; n = 332) had a body mass index (BMI) above 25 kg/m2, and class 3 obesity (BMI > 40) was more commonly seen in the participants with long COVID (20% vs 11%, p = 0.041). Upon admission, 157 (41%) had an elevated baseline C-reactive protein (CRP) ( ≥ 10 mg/L) and 280 (73%) had infiltrate(s) on chest imaging, but there were no differences between those with long COVID and those without. Similarly, the majority required some ventilatory or oxygenation support on admission (79% or n = 305) but baseline gas exchange and respiratory ordinal score were not associated with evidence of long COVID. Use of steroids and, remdesivir also were not associated with long COVID prevalence, nor was the prevalence of complications during the hospital stays. 186 (48%) out of 385 reported at least one symptom during the quarterly surveys, most commonly cardiopulmonary (cough or dyspnea) (33%), followed by gastrointestinal, neurologic or upper respiratory (32%) or systemic (fever, fatigue, myalgia, chills) (26%), and all reported symptoms were more common in those with long COVID as defined by PROs. As the study was conducted between May 2020 and March 2021, none of the participants had received a COVID-19 vaccine prior to admission.
Machine learning models using variables collected at the time of COVID-19 hospitalization predict risk of developing long COVID
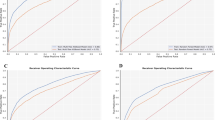
We developed a series of machine learning models for predicting the risk of developing a long COVID phenotype versus not developing a long COVID phenotype29. Models were built based on the clinical variables, symptoms, antibody (anti-Spike IgG) and viral load (N1 Ct) measurements collected at the time of hospital admission (baseline) for acute SARS-CoV-2 infection. A schematic of the machine learning model development pipeline is shown in Fig. 1. First, data were split into stratified training and testing datasets using a random 70:30 split, respectively, where each dataset had equal representation of those with and without subsequent development of a long COVID phenotypes. Next, we selected relevant features by performing feature selection on the training data set using LASSO regression. Models were built on the training data set and model performances were then assessed on the training data and subsequently on the test data set repeated 15 times which was entirely independent from the training data on which the models were constructed. Model performances showed accurate prediction on the training data, as expected, with mean AUROC values ranging from 0.766 to 0.846 (bGLM = 0.780, NNET = 0.766, RF = 0.846, SVM = 0.796, XGB = 0.836) and mean AUPRC (bGLM = 0.688, NNET = 0.669, RF = 0.802, SVM = 0.729, XGB = 0.773) values ranging from 0.669 to 0.802 (supplementary Fig. 1). When the models were then tested on the independent test dataset, all five models still achieved a significant predictive performance in terms of mean AUROC, with values ranging from 0.64 to 0.66 (bGLM = 0.657, NNET = 0.658, RF = 0.657, SVM = 0.651, XGB = 0.637). Similarly, AUPRC values ranged from 0.51 to 0.54 (bGLM = 0.526, NNET = 0.531, RF = 0.538, SVM = 0.533, XGB = 0.51) (Fig. 2 and supplementary Fig. 2). These results demonstrate that a parsimonious set of baseline clinical features can robustly predict subsequent development of a long COVID phenotype.
Schematic of the machine learning models development for predicting long COVID.
Distribution of (a) AUROC and (b) AUPRC values for all the models.
Feature importance
The relative importance score of all features for all models were investigated to understand which variables were important to predict long COVID (Fig. 3). All models demonstrated that lower antibody titers (anti-Spike IgG) and higher viral loads (N1 Ct) measured at hospital admission were the most statistically important baseline features in identifying patients that go on to develop long COVID symptoms. Among other features, diabetes mellitus, chronic respiratory disease, chronic cardiac disease, chronic neurologic disease and being female were significant risk factors for long COVID. Conversely, symptoms of cough, chest pain, sore throat were indicative of lower risk of developing long COVID.
a Dot plot showing scaled importance of all features included in the models predictive of long COVID. The size of the circle shows the relative importance of features. b Forest plot showing the univariate model odds ratios for the same features. c Boxplots showing SARS-CoV-2 viral levels (N1 CT) and antibody titers (Spike IgG) measured at the hospital admission by long COVID outcomes, Minimal vs Deficit clusters. Shown are median values (horizontal lines), interquartile ranges (boxes), and 1.5 IQR (whiskers), as well as individual points. The lower CT values indicate higher viral loads, the y-axis reversed.
Discussion
Developing a reliable predictive model of long COVID/PASC has proved challenging, largely due to the absence of a specific symptom-based definition for this condition29,30,31,32, unclear understanding of pathophysiology, and absence of validated biomarkers. Four years after the emergence of SARS-CoV-2, few studies have attempted to create a predictive model of long COVID. Antony et al. suggested that factors such as advanced age, female sex, certain comorbidities and medications administered, have a predictive value in determining the likelihood of developing long COVID13. This is consistent with the findings of Pfaff et al., who demonstrated that the most important features predictive of long COVID include post-COVID-19 respiratory symptoms, healthcare utilization rates, age as well as selected pre-existing conditions14. Finally, Sudre et al. relied on symptoms collected from patients during the early phase of their COVID-19 to determine their likelihood of developing long COVID. They reported that a symptom burden of five or more symptoms during the first week of infection was a strong predictor of PASC. Fatigue, headache, dyspnea, hoarse voice, and myalgia carried the most weight in predicting progression to PASC4,14. These studies highlight important components of predicting PASC but are limited in unique ways. The study by Antony et al. lacked diversity and relied on billing codes (likely underestimating PASC to a 0.3% prevalence). The study by Pfaff et al. included patients seeking care in long COVID clinics mostly within the pulmonary department and therefore is not generalizable to different PASC phenotypes. The third study by Sudre et al. was based on self-reported diagnosis and data entered by patients who were app users and limited to spring of 2020. None of these studies integrated clinical laboratory data specific to SARS-CoV-2 such as antibody titer and viral load.
Our study is based on a well-characterized prospective multicenter cohort that includes laboratory values, curated clinical data, as well as patient-reported symptoms and outcomes. Long COVID in the IMPACC cohort previously identified 4 clusters of patients based on reported clinical deficits. The predictive model proposed in this study focuses on baseline SARS-CoV-2 viral load and antibody measurements as features with the highest importance in terms of predictive value of progression to PASC. Many studies have postulated that an elevated viral load at time of COVID-19 positivity is correlated with worse clinical outcomes33,34,35 and emerging data suggest an association between viral load and Long COVID29,36,37. Moreover, the relationship between viral load and development of complications has already been established for other viruses, especially HIV. An elevated initial viral load on diagnosis of HIV was an accurate predictor of an earlier progression to AIDS38. Additionally, a high CSF HIV viral load was one of the most important predictors of progression to dementia in the pre-ART era38,39. Nonetheless, according to our data, antiviral therapy (limited to remdesivir in our study) was not associated with a decrease in PASC risk. Passive and active immunization was rarely received in our population (8.3% received convalescent plasma and vaccines were not commonly available), not allowing us to draw any conclusion on the benefit of early antibody in preventing PASC. In addition, steroid use in the acute phase was not associated with a reduction in PASC, and abnormal CRP levels on admission did not differ among those who developed PASC and those who did not.
While certain comorbidities are associated with PASC (e.g., chronic pulmonary, cardiac or neurologic diseases), it is unclear based on our data if some of the PASC disease burden could be misattributed to COVID-19 or if COVID-19 accentuates these pre-existing conditions. We also found that female sex was associated with PASC consistent with other studies40. Severity of disease on admission was not associated with PASC in our model, and as noted by others PASC also occurs after mild to moderate COVID41,42.
Our study has many strengths including the use of advanced ML models to integrate baseline clinical characteristics, antibody titers, and viral loads to predict long COVID outcomes. It enables early identification of individuals at risk, which could allow for more timely interventions and personalized care strategies. Our study employed five distinct ML algorithms, and each of these models captures different relationships between features and long COVID outcomes, which provides robustness and helps validate consistent signals across different frameworks. The comprehensive feature importance analysis reveals the key predictors, and our data demonstrate robustness of models across multi-center data. We also note the following limitations. First, our cohort was enrolled early in the pandemic and does not fully reflect how COVID-19 currently presents (in patients with hybrid immunity, exposed to variants of concern and having access to more antivirals, introduction of vaccines); in addition, because our study was designed at the onset of the pandemic, we did not comprehensively capture all symptoms that have been assessed to define PASC, though the PRO measures we chose are a reflection of current health status across functional domains and also include a comparison to pre-illness baseline. Second, we did not include additional cohorts (e.g: non-hospitalized COVID-19 cohort nor a hospitalized cohort without COVID-19) and the exclusion of mild to moderate COVID-19 affects the generalizability of our findings. Third, we did not include in our modeling changes of clinical and laboratory data in the acute phase of illness, as our goal was to create a framework for stratification upon presentation to care. A further limitation is that viral load measurements were taken at the time of hospital admission, which may have occurred at different points in the course of infection for different participants. As a result, the predictive value of viral load may be confounded by time since infection, potentially underestimating its relationship with long COVID outcomes. A limitation of ML models in predicting long COVID outcomes is their dependance on the quality and comprehensiveness of the input data, which can lead to biased or incomplete predictions if key variables or new symptoms are not captured. Moreover, ML models may struggle with generalizability, as models trained on data from early in the pandemic may not perform as well on data reflecting current patient characteristics, including those with hybrid immunity or exposed to newer variants.
Long COVID has a substantial public health impact. Our study integrates multiple data available at presentation to predict risk of PASC. Our findings suggest that a functional antiviral Ab immune response contributes to viral clearance and may decrease the occurrence of PASC. Our results highlight the benefit of measuring both antibody responses and viral load during the acute phase for the early identification of patients at high risk for PASC, which may facilitate early interventions.
Data availability
The IMPACC Data Sharing Plan is designed to enable the widest dissemination of data, while also protecting the privacy of the participants and the utility of the data by de-identifying and masking potentially sensitive data elements. All IMPACC data including those generated in this study have been deposited in the Immunology Database and Analysis Portal (ImmPort), a NIAID Division of Allergy, Immunology and Transplantation funded data repository under accession code SDY1760. All raw and processed data are available under restricted access to comply with the NIH public data sharing policy for IRB-exempted public health surveillance studies, Access can be obtained via AccessClinicalData@NIAID (https://accessclinicaldata.niaid.nih.gov/study-viewer/clinical_trials). Additional guidelines for access are outlined on ImmPort (https://docs.immport.org/home/impaccslides). The source data for Figs. 2 and 3 are accessible from Supplementary Data 1 and 2.
Code availability
All codes for the analyses and tables generated by this study have been deposited in the Bitbucket repository https://bitbucket.org/kleinstein/impacc-public-code/src/master/predict_model_PASC_manuscript/ and are publicly available as of the date of publication. The code is also stored in Zenodo and can be accessed via the https://doi.org/10.5281/zenodo.1712785243.
Change history
23 February 2026
A Correction to this paper has been published: https://doi.org/10.1038/s43856-026-01425-9
References
CDC. Long COVID Basics. https://www.cdc.gov/coronavirus/2019-ncov/long-term-effects/index.html (2025).
National Center for Health Statistics. U.S. Census Bureau HPS, 2022–. Long COVID. https://www.cdc.gov/nchs/covid19/pulse/long-covid.htm (2025).
Office for National Statistics. Prevalence of Ongoing Symptoms Following Coronavirus (COVID-19) Infection in the UK. https://www.ons.gov.uk (2025).
Sudre, C. H. et al. Attributes and predictors of long COVID. Nat. Med. 27, 626–631 (2021).
Huyut, M. T., Velichko, A. & Belyaev, M. Detection of risk predictors of COVID-19 mortality with classifier machine learning models operated with routine laboratory biomarkers. Appl. Sci. 12, 12180 (2022).
Huyut, M. T. & Huyut, Z. Effect of ferritin, INR, and D-dimer immunological parameters levels as predictors of COVID-19 mortality: a strong prediction with the decision trees. Heliyon 9, e14015 (2023).
Huyut, M. T. & İlkbahar, F. Corrigendum to ‘The effectiveness of blood routine parameters and some biomarkers as a potential diagnostic tool in the diagnosis and prognosis of Covid-19 disease’ [Int. Immunopharmacol. 98 (2021) 107838]. Int. Immunopharmacol. 123, 110823 (2023).
Huyut, M. T. & Velichko, A. Diagnosis and prognosis of COVID-19 disease using routine blood values and LogNNet neural network. Sensors 22, 4820 (2022).
Huyut, M. T. & Üstündağ, H. Prediction of diagnosis and prognosis of COVID-19 disease by blood gas parameters using decision trees machine learning model: a retrospective observational study. Med. Gas Res. 12, 60–66 (2022).
Gao, Y. et al. Machine learning based early warning system enables accurate mortality risk prediction for COVID-19. Nat. Commun. 11, 5033 (2020).
He, F., Page, J. H., Weinberg, K. R. & Mishra, A. The development and validation of simplified machine learning algorithms to predict prognosis of hospitalized patients with COVID-19: Multicenter, retrospective study. J. Med. Internet Res. 24, e31549 (2022).
Mertoglu, C. et al. COVID-19 is more dangerous for older people and its severity is increasing: a case-control study. Med. Gas Res. 12, 51–54 (2022).
Antony, B. et al. Predictive models of long COVID. EBioMedicine 96, 104777 (2023).
Pfaff, E. R. et al. Identifying who has long COVID in the USA: a machine learning approach using N3C data. Lancet Digit Health 4, e532–e541 (2022).
von Elm, E. et al. The Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement: guidelines for reporting observational studies. J. Clin. Epidemiol. 61, 344–349 (2008).
IMPACC Manuscript Writing Team & IMPACC Network Steering Committee. Immunophenotyping assessment in a COVID-19 cohort (IMPACC): a prospective longitudinal study. Sci. Immunol. 6, eabf3733 (2021).
Devlin, N. J. & Brooks, R. EQ-5D and the EuroQol group: past, present and future. Appl. Health Econ. Health Policy 15, 127–137 (2017).
Salsman, J. M. et al. Assessing psychological well-being: self-report instruments for the NIH Toolbox. Qual. Life Res. 23, 205–215 (2014).
Lai, J.-S., Garcia, S. F., Salsman, J. M., Rosenbloom, S. & Cella, D. The psychosocial impact of cancer: evidence in support of independent general positive and negative components. Qual. Life Res. 21, 195–207 (2012).
Hays, R. D., Bjorner, J. B., Revicki, D. A., Spritzer, K. L. & Cella, D. Development of physical and mental health summary scores from the patient-reported outcomes measurement information system (PROMIS) global items. Qual. Life Res. 18, 873–880 (2009).
Lai, J.-S., Wagner, L. I., Jacobsen, P. B. & Cella, D. Self-reported cognitive concerns and abilities: two sides of one coin? Psychooncology 23, 1133–1141 (2014).
National Institutes of Health. Patient-Reported Outcomes Measurement Information System (PROMIS®). https://commonfund.nih.gov/promis (2025).
Choi, S. W., Victorson, D. E., Yount, S., Anton, S. & Cella, D. Development of a conceptual framework and calibrated item banks to measure patient-reported dyspnea severity and related functional limitations. Value Health 14, 291–306 (2011).
Yount, S. E. et al. Brief, valid measures of dyspnea and related functional limitations in chronic obstructive pulmonary disease (COPD). Value Health 14, 307–315 (2011).
Irwin, D. E. et al. Correlation of PROMIS scales and clinical measures among chronic obstructive pulmonary disease patients with and without exacerbations. Qual. Life Res. 24, 999–1009 (2015).
PROMIS. Dyspnea Scoring Manual. https://www.healthmeasures.net/images/PROMIS/manuals/Scoring_Manuals_/PROMIS_Dyspnea_Scoring_Manual.pdf (2021).
Centers for Disease Control and Prevention. Research Use Only 2019-novel Coronavirus (2019-ncov) Real-Time RT-PCR Primers And Probes. https://stacks.cdc.gov/view/cdc/88834 (2020).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020).
Ozonoff, A. et al. Features of acute COVID-19 associated with post-acute sequelae of SARS-CoV-2 phenotypes: results from the IMPACC study. Nat. Commun. 15, 216 (2024).
Nalbandian, A. et al. Post-acute COVID-19 syndrome. Nat. Med. 27, 601–615 (2021).
Pan, D. & Pareek, M. Toward a universal definition of post-covid-19 condition-how do we proceed? JAMA Netw Open 6, e235779 (2023).
Deer, R. R. et al. Characterizing long COVID: deep phenotype of a complex condition. EBioMedicine 74, 103722 (2021).
Fajnzylber, J. et al. SARS-CoV-2 viral load is associated with increased disease severity and mortality. Nat. Commun. 11, 5493 (2020).
Shenoy, S. SARS-CoV-2 (COVID-19), viral load and clinical outcomes; lessons learned one year into the pandemic: a systematic review. Pediatr. Crit. Care Med. 10, 132–150 (2021).
Pujadas, E. et al. SARS-CoV-2 viral load predicts COVID-19 mortality. Lancet Respir Med. 8, e70 (2020).
Su, Y. et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell 185, 881–895.e20 (2022).
Girón Pérez, D. A. et al. Post-COVID-19 syndrome in outpatients and its association with viral load. Int. J. Environ. Res. Public Health 19, 15145 (2022).
Saag, M. S. et al. HIV viral load markers in clinical practice. Nat. Med. 2, 625–629 (1996).
Brew, B. J., Pemberton, L., Cunningham, P. & Law, M. G. Levels of human immunodeficiency virus type 1 RNA in cerebrospinal fluid correlate with AIDS dementia stage. J. Infect. Dis. 175, 963–966 (1997).
Tsampasian, V. et al. Risk factors associated with post-covid-19 condition: a systematic review and meta-analysis. JAMA Intern. Med. 183, 566–580 (2023).
Townsend, L. et al. Persistent fatigue following SARS-CoV-2 infection is common and independent of severity of initial infection. PLoS ONE 15, e0240784 (2020).
Chan Sui Ko, A. et al. Number of initial symptoms is more related to long COVID-19 than acute severity of infection: a prospective cohort of hospitalized patients. Int. J. Infect. Dis. 118, 220–223 (2022).
Acknowledgements
This study is being supported by grants R01AI104870, R01AI132774, R01AI135803, R01AI145835, U19AI057229, U19AI062629, U19AI077439, U19AI089992,U19AI090023, U19AI118608, U19AI118610, U19AI125357, U19AI128910, U19AI128913, U54AI142766, U19AI089992, U24AI52179 from the National Institute of Allergy and Infectious Diseases (NIAID), a part of the U.S. National Institutes of Health(NIH), and P51 OD011132, S10 OD026799 from NIH.
Author information
Authors and Affiliations
Consortia
Contributions
N.D.J., H.S., S.T.W., A.O., P.B., M.C.A., N.R., A.A., L.B., C.S.C., G.M., C.B.C., V.S., C.H., K.C.N., J.S., A.S., D.C., N.I.A.H., C.B., S.C.B., E.M., and V.S.M. were involved in the study concept and design. H.S., S.T.W., A.O., C.M., N.D.J., H.V.B., M.C.A., D.E., S.L., F.K.R., J.D.-A., P.B., and N.R. contributed to the study methodology. C.M., N.D.J., H.S., S.T.W., H.V.B., S.L., F.K.R., L.B.R., S.K.S., C.R.L., W.E., S.E.B., H.T.M. were involved in data curation. L.B., C.S.C., G.M., C.B.C., N.R., V.S., C.H., K.C.N., J.M., A.S., D.C., N.I.A.H., C.B., S.C.B., and E.M. contributed to the investigation. C.M., N.D.J., H.VB., S.L., A.O., J.-D.A., J.M., and V.S.M. provided software. A.O., C.M., N.D.J., H.V.B., M.C.A., D.E., S.L., F.K.R., J.Q., C.S., R.D., and C.P.S. were involved in formal analysis. A.O., C.M., N.D.J., H.V.B., M.C.A., D.E., S.L., and F.K.R. were involved in data validation, visualization, or replication of analyses. N.D.J, H.S., S.T.W., M.C.A., J.S., N.R., and P.B. drafted the manuscript. A.O., J.S., N.D.J., C.M., C.S., C.B.C., M.K., L.B., A.S., F.K.R., H.V.B., D.E., S.L., A.F.S., V.S., D.H., R.M., S.K., O.L., C.B., E.H., D.J.E., B.P.U., K.C.N., M.D., C.H., W.M., N.I.A.H., J.M., M.A.A., S.C.B., D.C., F.KH., L.I.E., E.M., G.M., R.S., J.D.-A., B.P.E., A.A., E.R., M.C.A., P.B., H.S., L.N.G., N.R., and C.P.S. critically reviewed and revised the manuscript. P.B., A.A., A.O., S.K., B.P.E., N.R., E.R., J.-D.-A., and J.M. were involved in project administration, and A.O., J.S., N.D.J., C.M., C.S.C., C.B.C., M.K., L.B., A.S., F.K.R., H.V.B., D.E., S.L., A.F.S., V.S., D.H., R.M., S.K., O.L., C.B., E.H., D.E., B.P.U., K.C.N., M.D., C.H., W.M., N.I.A.H., J.M., M.A.A., S.C.B., D.C., F.K.H., L.I.E., E.M., G.M., R.S., J.D.-A., B.P.E., A.A., E.R., M.C.A., P.B., and N.R. provided study resources. O.L., R.S., E.H., A.F.S., W.M., M.D., B.P.U., E.F., D.J.E., R.M., D.H., F.K.H., J.M., M.K., M.A.A., and L.I.E. obtained funding. P.B., M.C.A., and N.R. provided supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: The Icahn School of Medicine at Mount Sinai has filed patent applications relating to SARS-CoV-2 serological assays and NDV-based SARS-CoV-2 vaccines which list F.K. as co-inventor. Mount Sinai has spun out a company, Kantaro, to market serological tests for SARS-CoV-2. F.K. has consulted for Merck and Pfizer (before 2020), and is currently consulting for Pfizer, Seqirus, 3rd Rock Ventures, Merck and Avimex. The Krammer laboratory is also collaborating with Pfizer on animal models of SARS-CoV-2. V.S. is a co-inventor on a patent filed relating to SARS-CoV-2 serological assays (the “Serology Assays”). O.L. is a named inventor on patents held by Boston Children’s Hospital relating to vaccine adjuvants and human in vitro platforms that model vaccine action. His laboratory has received research support from GlaxoSmithKline (GSK). C.B.C. serves as a consultant to bioMerieux and is funded for a grant from Bill & Melinda Gates Foundation. J.A.O. is a consultant at Knocean Inc. Jessica Lasky-Su serves as a scientific advisor of Precion Inc. S.R.H., G.M. and K.W. are employees of Metabolon Inc. V.S.M. is a current employee of MyOwnMed. N.R. reports contracts with Lilly, Immorna, Vaccine Company and Sanofi for COVID-19 clinical trials and serves as a consultant for ICON, EMMES, Imunon, CyanVac for consulting on safety for COVID19 clinical trials. A.R. is a current employee of Immunai Inc. Steven Kleinstein is a consultant related to ImmPort data repository for Peraton. Nathan Grabaugh is a consultant for Tempus Labs and the National Basketball Association. Akiko Iwasaki is a consultant for 4BIO, Blue Willow Biologics, Revelar Biotherapeutics, RIGImmune, Xanadu Bio, Paratus Sciences. M.K. receives research funds paid to her institution from NIH, ALA; Sanofi, Astra-Zeneca for work in asthma, serves as a consultant for Astra-Zeneca, Sanofi, Chiesi, GSK for severe asthma; is a co-founder and CMO for RaeSedo, Inc, a company created to develop peptidomimetics for treatment of inflammatory lung disease. E.M. received research funding from Babson Diagnostics, honorarium from Multiple Sclerosis Association of America and has served on advisory boards of Genentech, Horizon, Teva and Viela Bio. C.C. receives research funding from NIH, FDA, DOD, Roche-Genentech and Quantum Leap Healthcare Collaborative as well as consulting services for Janssen, Vasomune, Gen1e Life Sciences, NGMBio, and Cellenkos. Wade Schulz was an investigator for a research agreement, through Yale University, from the Shenzhen Center for Health Information for work to advance intelligent disease prevention and health promotion; collaborates with the National Center for Cardiovascular Diseases in Beijing; is a technical consultant to Hugo Health, a personal health information platform; cofounder of Refactor Health, an AI-augmented data management platform for health care; and has received grants from Merck and Regeneron Pharmaceutical for research related to COVID-19. G.A.M. received research grants from Rehdhill, Cognivue, Pfizer, and Genentech, and served as a research consultant for Gilead, Merck, Viiv/GSK, and Jenssen. L.N.G. received research funding paid to her institution from Pfizer, Inc. E.M. is an Editorial Board Member for Communications Medicine and Guest Editor for the Post COVID-19 condition/Long COVID Collection, but was not involved in the editorial review or peer review, nor in the decision to publish this article. L.N.G. is a Guest Editor for the Post COVID-19 condition/Long COVID Collection, but was not involved in the editorial review or peer review, nor the decision to publish this article.
Peer review
Peer review information
Communications Medicine thanks Sabra Klein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Doni Jayavelu, N., Samaha, H., Wimalasena, S.T. et al. Machine learning models predict long COVID outcomes based on baseline clinical and immunologic factors. Commun Med 6, 1 (2026). https://doi.org/10.1038/s43856-025-01230-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01230-w
This article is cited by
-
Associated factors and predictive nomogram of long COVID: a cross-sectional study in China
BMC Pulmonary Medicine (2026)