Abstract
Background
Neural recordings capture crucial pathophysiological processes along the dementia continuum. However, cross-center variability in recording techniques and paradigms limit their generalizability and diagnostic power, preventing clinical use. We here propose a computational approach enabling cross-center classification even in the presence of completely different clinical pipelines.
Methods
We leveraged a digital twin model to derive digital biomarkers linking neurodegeneration mechanisms to alterations in neural activity across multiple recording modalities. We tested the generalizability of digital biomarkers through cross-center classification of Mild Cognitive Impairment (MCI) and healthy subjects in two independent clinics. The two datasets presented different recording techniques (EEG and MEG), preprocessing modalities, recruitment criteria and diagnostic guidelines. Digital biomarkers derived from one clinic were tested for classifying patients in the other clinic and vice versa employing a transfer learning approach.
Results
Digital biomarkers outperform standard biomarkers in the MCI vs healthy classification in both separate datasets (83% vs 58% for EEG dataset and 75% vs 68% for MEG dataset). Moreover, they achieve accurate and consistent cross-center classification (77–78% accuracy), while standard biomarkers perform poorly in the generalization attempt (56–65%). Additionally, digital biomarkers reliably predict global cognitive status across clinics across both datasets ( p < 0.01), while standard biomarkers present no correlation.
Conclusions
Digital biomarkers generalize across recording techniques and datasets, enabling a cross-modal and cross-center classification of a patient’s condition. These biomarkers offer a robust measure of patient-specific neurodegeneration, mapping neural recordings anomalies into a common framework of underlying structural alterations. The vast differences between the two datasets support the applicability of this approach also in the presence of high inter-center variability.
Plain language summary
People with pathological cognitive decline lose memory, reasoning and communication ability, developing diseases such as dementia. Non-invasive methods that record activity in the brain have been investigated to see whether they can be used to identify people with pathological cognitive decline. Ideally these methods would require minimal equipment and reduce patient discomfort, but their results often vary greatly between clinics. We used patient-specific computational brain models, called digital brain twins, to find possible digital markers of pathological cognitive decline. We evaluated whether these markers could be used to diagnose pathological clinical cognitive decline across two groups of people in different clinics. Our results demonstrate that digital markers generalize well across different clinical settings and recording methods. They could provide better indicators of pathological cognitive decline than currently used alternatives and so improve identification of people with pathological cognitive decline.
Similar content being viewed by others
Introduction
The functional characterization of prodromal dementia conditions constitutes a fundamental step to support an early diagnosis of cognitive decline and for the understanding of its pathophysiology1. Non-invasive recordings2,3,4, combined with computational techniques5,6,7,8,9, are promising candidates for this task, due to their high temporal resolution and reduced discomfort for the patient. While many works have explored the combination of non-invasive recordings and machine learning methods for dementia classification10,11,12,13 this approach ultimately failed to reach clinical stage. Main obstacles include the lack of interpretability and reproducibility of these methods14,15, and poor generalizability, as models are often trained on monocentric datasets16.
Predicting clinical evolution for each patient requires the assessment of the status of individual pathophysiological processes. Hence, one cannot rely on data features alone, which lead to correlational associations by construction. Digital twins, i.e., personalized models of brain activities, have been proposed to overcome this limitation, as they encapsulate previous pathophysiological knowledge about disease evolution and exploit it to predict the individual case17. However, the use of digital twins to derive biomarkers of pathological conditions has been rarely attempted18,19.
Here, we use digital twins as a physiologically-informed way to merge datasets across different modalities and clinical centers. We employed the previously validated Digital Alzheimer’s Disease Diagnosis (DADD) model18,20, reproducing cortical activity at several levels of dementia severity18 to derive patient-specific digital biomarkers of connectivity and synaptic degeneration representing Mild Cognitive Impairment (MCI) condition. Here, ‘digital biomarkers’ refer to model-derived neurodegeneration parameters estimated from non-invasive recordings.
Digital biomarkers were used in a transfer learning approach to classify MCI patients and healthy subjects across non-overlapping cohorts from two independent clinics. Data acquisition in the two cohorts presented different recording techniques, preprocessing and postprocessing paradigms, recruitment criteria and diagnostic guidelines. This enabled us to test the ability of our approach to perform multicentric analysis also in the presence of high inter-center variability. The classifying performance of these digital biomarkers was compared with that of standard electrophysiological biomarkers like power spectral density (PSD) and functional connectivity (FC). Digital biomarkers enable an accurate cross-modal and cross-center classification, outperforming standard electrophysiological biomarkers and paving the way for clinical applications.
Methods
Participants recruitment (Dataset 1)
A total of 44 patients diagnosed with Mild Cognitive Impairment (16 males and 28 females; mean age 73.88 ± 8.32 years; mean education 10.14 ± 3.97 years) were recruited from Careggi University Hospital, Florence, Italy. Participants were enrolled in the PREVIEW project21 (ClinicalTrials.gov Identifier: NCT05569083), a longitudinal study focused on prodromal forms of dementia. Classification into cognitive categories was conducted based on extensive clinical and neuropsychological assessments, adhering to the NIA-AA criteria for diagnosing mild cognitive impairment22. Inclusion criteria are reported in Supplementary Materials, “Inclusion criteria (Dataset 1)” paragraph. A control group of 17 participants (10 males and 7 females) was also enrolled. MCI and control participants were not matched for age (65.19 ± 4.42 years in the control group, U = 123.0, p = 0.00001, Mann–Whitney test) or education (13.95 ± 4.03 years, U = 576.0, p = 0.001, Mann–Whitney test). We ensured that these discrepancies constituted no confounding factor for the results, by assessing the correlations between age and education levels and the standard EEG and MEG biomarkers used for classification tasks and for the derivation of digital biomarkers. These correlation analyses found no statistical significance (results are reported in Supplementary Table 1).
Dataset 1 recordings were obtained using electroencephalography (EEG), with no source reconstruction. Recruitment and EEG recordings followed the Declaration of Helsinki and the guidelines of the Committee on Human Experimentation at Careggi University Hospital, Florence, Italy. The study was approved by the local Institutional Review Board (Careggi University Hospital, reference number: 15691oss), and all patients provided informed consent.
Participants recruitment (Dataset 2)
A total of 32 patients diagnosed with amnestic Mild Cognitive Impairment (aMCI) (18 males and 14 females; 21 with single-domain aMCI and 11 with multiple-domain aMCI; mean age 71.31 ± 6.83 years; mean education 10.54 ± 4.33 years) were recruited from the Center of Cognitive and Memory Disorders at the Hermitage Capodimonte Clinic in Naples, Italy. All participants were right-handed native Italian speakers, with MCI diagnoses made following the criteria established by the National Institute on Aging-Alzheimer’s Association22. Inclusion criteria are reported in Supplementary Materials, “Inclusion criteria (Dataset 2)” paragraph. A control group of 32 participants (19 males and 13 females), matched for age (69.9 ± 5.61 years) and education (12.96 ± 4.56 years), was also enrolled.
Dataset 2 recordings were obtained using magnetoencephalography (MEG) with source reconstruction. For all MCI patients and 27/32 controls both source-reconstructed MEG data and neuropsychological assessments data were available and were then included in the present study. The study protocol received approval from the ethics committee “Comitato Etico Campania Centro” (Protocol number: 93 C.E./Reg. n.14-17OSS), with all participants providing informed consent in written form as per the Declaration of Helsinki.
Signals recordings and preprocessing (Dataset 1)
EEG data were collected from subjects who met the inclusion criteria outlined in the study. The EEG recordings were conducted while subjects were seated in a comfortable position with eyes closed. Signals were collected using the 64-channels Galileo-NT system (E.B. Neuro S.p.a.), with sensor topography following the extended 10/20 system. Unipolar signals were recorded at a sampling rate of 512 Hz, and electrode impedances, monitored throughout EEG acquisition, were maintained in the 7–10 kOhm range. Portions of the signal where impedance deviated outside of the 7–10 kOhm range were discarded.
The EEG pre-processing pipeline was implemented on MATLAB with the commercially available EEGLAB Toolbox23, following the PREP pipeline24, followed by Independent Component Analysis (ICA) artifact rejection. This includes: (i) band-pass filtering in the 0.5–45 Hz range using a Butterworth filter, (ii) identification and subsequent removal of noisy channels, (iii) average re-referencing, (iv) visual rejection of high-amplitude artifacts and (v) ICA-based detection and elimination of artefactual components. The details of each step are reported in Supplementary Materials, “Signal preprocessing (Dataset 1)” paragraph.
Signals recordings and preprocessing (Dataset 2)
Data were collected using a magnetoencephalography (MEG) system with 154 magnetometers and nine reference sensors in a magnetically shielded room (ATB, Biomag, ULM, Germany) to minimize external noise. Head position under the helmet was defined with Fastrack (Polhemus), digitizing four anatomical landmarks (nasion, left and right preauricular points, and the vertex) and the positions of four reference coils attached to the subject’s head. Each subject underwent two recordings (3.5 min each) with a 1-min rest period, in a resting state with eyes closed. Cardiac activity and eye movements were also recorded. Preprocessing was similar to that described by Liparoti et al.25. Source reconstruction was based on the volume conduction models described by Nolte26, using the automatic anatomical labeling (AAL) atlas27. The procedure is reported in Supplementary Materials, “Signal preprocessing (Dataset 2)” paragraph.
Electrophysiological metrics
For both datasets, we extracted relative power spectral density (PSD) values in standard frequency bands (delta: 0–4 Hz, theta: 4–8 Hz, alpha: 8–13 Hz, beta: 13–30 Hz, gamma: 30–45 Hz). Functional connectivity (FC) was assessed using both standard Pearson correlation or amplitude envelope correlation (which is extremely robust to volume conduction effects28), choosing the most informative one after dataset merging (standard Pearson correlation). For both metrics, we considered only significant correlation values (p < 0.05). FC was computed within standard frequency bands and across the whole-band range (0.5–45 Hz as signals were band-pass filtered within this frequency interval). We computed electrophysiological metrics in both datasets with the same technique, but separately for EEG (Dataset 1) in sensor space and MEG with source reconstruction (Dataset 2) in source space. To quantify the role of preprocessing techniques, in the Supplementary Materials “Comparison between preprocessed and raw signals (both datasets)” paragraph, we also report the results of statistical analysis and digital biomarkers derivation based on electrophysiological metrics derived from raw signals (i.e., without preprocessing) in both datasets.
DADD model and simulated signal
The DADD digital twin model used in this study has been previously described in detail in previous publications from our group18,20. Briefly, it represents the brain cortex as a network of 76 interacting regions using The Virtual Brain (TVB) platform29. Neural activity within each region of interest (ROI) is simulated using a mean-field approximation based on a modified Jansen-Rit neural mass model30, These ROIs are connected through a high-fidelity structural connectivity matrix derived from human tractography data31.
There is a notable discrepancy between the number of ROIs in the model parcellation (n = 76) and both the EEG sensors (n = 65) and the ROIs of the atlas used for source reconstruction of MEG signals (n = 90). This discrepancy arises from the choice of structural connectivity data, which is independent of the functional atlas used for source reconstruction, and of course from the EEG cap used in experimental recordings. This allowed us to test another aspect of the flexibility of the model, i.e., its ability to bridge different parcellation schemes and recording modalities (e.g., EEG in sensor space and MEG in source space).
The model output thus consists of simulated time series representing neural dynamics in each ROI at different levels of structural neurodegeneration. Within the model, neurodegeneration is modeled by three parameters affecting both synaptic transmission and connectivity.
Synaptic degeneration is governed by the lp (from local parameter, since it affects local activity) parameter with the equation:
Connectivity degeneration is described by the cp and np parameters, according to the equation:
The synaptic degeneration parameter (lp) affects only the local dynamics of the neural mass, while connectivity degeneration (cp) and neuroplasticity (np) influence the structural connectivity matrix. Biophysically, lp reflects cortical hypo-inhibition and hyperexcitation due to early Abeta deposition32 in regions corresponding to early Braak stages, mainly frontal and temporal regions33. The cp parameter portrays white matter atrophy34. The np parameter describes the neuroplastic effects on connective rewiring35. The values in the equations are derived from biophysical constraints and can be consulted in a previous work from our group18.
Additionally, we considered the squared sum between the connectivity and synaptic degeneration parameter, referred as total degeneration parameter, according to the equation:
Statistics and reproducibility
Comparisons across groups were conducted using the Mann–Whitney U test, suitable for comparing two independent groups when assumptions of normality and equal variances are not met. Classification performances were statistically compared using McNemar’s test applied to confusion matrices (See Supplementary Materials, “Machine learning pipeline, classification and cross-classification” paragraph).
Outliers were defined as datapoints that ranged more than 3\(\sigma\) from the nearest datapoint, where \(\sigma\) is the standard deviation of the sample.
Correlation between standard and digital biomarkers with MMSE scores was evaluated with Scipy’s linregress command. MMSE scores were age-corrected prior to the analysis. The correction was made by computing the slope of the regression line \({S}_{{Age}/{MMSE}}\) between age and MMSE (across both datasets), and then adding to the MMSE score the weighted age value, according to the formula:
Where \({abs}\) represents the absolute value. Education presented no significant relationship with MMSE across both datasets (r = −0.04, p = 0.81) and was thus excluded from the correction of MMSE scores. All computational and data analysis pipelines were implemented in Python, with simulations conducted using the Virtual Brain software, and analyzed with standard Python packages such as SciPy, NumPy, and Pandas. Machine learning pipeline was implemented using Scikit-Learn and is reported in Supplementary Materials, “Machine learning pipeline (classification and cross-classification)”.
Results
We analyzed two distinct datasets (Fig. 1A). The first comprised 17 healthy controls (CTR) subjects and 44 MCI patients, that were recruited at the Neurological Department of Careggi Hospital in Florence, Italy and underwent resting-state electroencephalography (EEG).
A Participants from two different datasets and multi-modal recording techniques were considered. Both datasets comprised healthy controls (CTR) and mild cognitive impairment (MCI) patients, enrolled with different criteria from two independent clinics. The two datasets employed different recording techniques, with Dataset 1 using EEG and Dataset 2 employing MEG with source reconstruction. B Biomarkers were derived for each patient, using either standard analysis of electrophysiological recordings or personalized brain models (digital biomarkers). C Biomarkers performance was assessed by using them as classifying features between healthy and MCI condition. Leveraging a transfer-learning approach, the generalizability of such classifications was also assessed, by employing the classifier trained on Dataset 1 to classify patients of Dataset 2 (and vice versa). Performances of standard and digital biomarkers were then compared.
The second dataset comprised 27 CTR subjects and 32 amnestic MCI patients, that underwent magnetoencephalography (MEG) at the Hermitage Hospital in Naples, Italy. Patients in the two datasets presented no statistical differences in biological sex composition (chi-square test = 2.22, p = 0.14), age (Mann–Whitney U-test=479.0, p = 0.068) and education (U = 628.5, p = 0.89). We evaluated the cross-modal and cross-center classification performance of digital biomarkers compared to standard biomarkers (Fig. 1B). Using a transfer learning approach, we trained machine learning classifiers on patients from one dataset and tested their ability to classify patients from the other dataset (Fig. 1C).
Digital Biomarkers discriminate between diagnostic categories across both datasets
EEG and MEG signals were pre-processed prior to statistical analysis (see Methods, “Signals recordings and preprocessing” paragraphs). We extracted the same standard biomarkers from the recordings of the two datasets (see “Electrophysiological metrics” paragraph). We then used standard biomarkers to derive digital biomarkers following a procedure already validated on EEG data18,20 (see Supplementary Materials, “Determination of personalized digital biomarkers” paragraph). We computed the mutual information between the categorical variable of patients’ clinical condition (CTR vs MCI) and both standard and digital biomarkers after merging the two datasets (see Methods, “Mutual Information” paragraph). We identified the beta/theta PSD ratio as the most informative standard biomarker (MI = 0.21 [0.04, 0.32]), while the most informative digital biomarker was found to be the squared sum of synaptic degeneration and connectivity degeneration parameter (total degeneration parameter, MI = 0.21 [0.09, 0.24]).
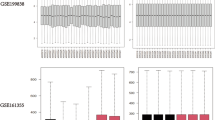
Subsequently, we ran statistical analyses to compare the values of the most informative standard and digital biomarkers across conditions in the two datasets. For Dataset 1, the most informative standard biomarker (beta/theta ratio) did not display a significant difference between the two conditions (U = 502, p = 0.054, d = 0.2, Fig. 2A). However, the total parameter digital biomarker presented significant differences across groups (U = 196, p = 0.010, d = −0.9). For Dataset 2, the beta/theta ratio did significantly differ across groups but with a moderate effect size (U = 622, p = 0.004, d = 0.5). The total degeneration parameter was also significant, presenting a larger effect size (U = 257, p = 0.006, d = −0.8, Fig. 2B).
A Boxplot comparison of most informative standard biomarker and most informative digital biomarker across groups (Dataset 1, 17 HC and 44 MCI). Only total degeneration was found to significantly differ across groups. B Boxplot comparison of most informative standard biomarker and most informative digital biomarker across groups (Dataset 2, 27 HC and 32 MCI). The beta/theta PSD ratio (see Main Text) was found to significantly differ across groups, although with moderate size effect. Total degeneration displayed similar significance and larger size effect. C No significant correlation with corrected MMSE was found for standard biomarkers (the highest correlation was found with beta/theta ratio, r = 0.13, p = 0.15). D Synaptic degeneration presented significant anti-correlation with corrected MMSE (r = −0.27, p = 0.009). Reported values in C and D were normalized before plotting. Notation: In the boxplots, whiskers represent 1.5 times the interquartile range and median is highlighted by horizontal black line. In linear regression plots, shaded areas represent error bands with confidence intervals (2.5%–97.5%). Significance notation: * stands for p < 0.05; ** stands for p < 0.01, two-sided Mann–Whitney test.
Digital Biomarkers predict global cognitive status of patients (MMSE)
We then investigated the relationship between identified biomarkers (both standard and digital) and global cognitive status measured with the (age-)corrected Mini-Mental State Examination (MMSE) in MCI patients, merging the two datasets. This was done with a linear regression analysis between the candidate biomarkers (either standard or digital) and MMSE scores corrected for patient’s age (see Methods, “Statistics and Reproducibility” paragraph). Notably, while for standard biomarkers no significant correlation was found (the highest correlation was found for the beta/theta ratio, r = 0.13, p = 0.15, see Fig. 2C), the synaptic degeneration digital biomarker showed a significant negative correlation with corrected MMSE (r = −0.27, p = 0.009, Fig. 2D). In other words, greater synaptic damage predicted by our model correlated with poorer global cognitive performance during clinical assessments. We tested the robustness of this prediction by bootstrapping 100 times the merged dataset to analyze the resulting distribution of r-values (Supplementary Fig. 1). The r-values for the digital biomarkers’ linear regression presented bimodality (Hartigan’s dip test, p < 0.001), reflecting the diverse nature of patients in the two datasets. Bimodality was not observed in linear regression r-values for standard biomarkers. This bimodality was further confirmed by running the same linear-regression analysis between corrected MMSE and both standard and digital biomarkers separately for the two datasets (Supplementary Fig. 2). No significant correlation with corrected MMSE was found for standard biomarkers in Dataset 1 (the highest correlation was found for alpha band power, r = 0.10, p = 0.28), while the connectivity degeneration parameter showed significant correlation (r = −0.29, p = 0.036). In Dataset 2, no standard biomarker predicted global cognitive state with significance (highest correlation was found with beta/theta ratio, r = 0.14, p = 0.23). Among digital biomarkers, the synaptic degeneration parameter strongly predicted corrected MMSE in MCI patients (r = −0.47, p = 0.004).
Digital Biomarkers classify patients in intra-clinical and cross-clinical settings, always outperforming standard biomarkers
Next, we assessed the performance of both standard and digital biomarkers in classifying healthy controls versus MCI patients. We evaluated the classification performance of both standard and digital biomarkers using a nested cross-validation approach (see Supplementary Materials, “Machine Learning Pipeline (Classification and Cross-Classification)” paragraph). We incorporated a feature selection routine in the training set to identify the most informative features for both groups of biomarkers. We then compared classification performances. The classifying features for standard biomarkers are reported in the Supplementary Materials “Mutual Information” paragraph, while all digital biomarkers were considered as potential features in the digital biomarker set. Multiple machine learning algorithms were tested (see Supplementary Materials, “Machine learning pipeline (classification and cross-classification)” paragraph), selecting the one that obtained the best performance for all classifications (reported in Supplementary Table 2). In Dataset 1, the digital biomarkers outperformed the standard biomarkers in terms of mean accuracy (0.83 ± 0.07 vs 0.58 ± 0.04), and area under the receiver operator characteristic (ROC) curve (AUC, 0.79 vs 0.62, Fig. 3A). Statistical test on the performance of both classifiers confirmed the superiority of digital biomarkers (p = 0.006, McNemar’s test). Similar results were obtained for Dataset 2, with the digital biomarkers showing higher accuracy 0.75 ± 0.03 vs 0.68 ± 0.04) and AUC (0.77 vs 0.69, Fig. 3B), even if the differences between classifiers were not statistically significant (p = 0.22, McNemar’s test). These findings suggest that digital biomarkers are more effective and generalizable in distinguishing between healthy and MCI patients across different datasets, and that the digitalization leads to similar classification performances starting from either MEG or EEG (as opposed to standard classification that was clearly superior in MEG).
A Biomarker performance in healthy-MCI classification for Dataset 1. Digital model-based biomarkers presented higher values in mean accuracy (left, p = 0.006) and area under the roc curve (right). B Biomarker performance in healthy-MCI classification for Dataset 2. Also for Dataset 2 digital model-based biomarkers outperformed standard biomarkers in mean accuracy (left) and area under the roc curve (right), although with no statistical significance (p = 0.22). C Cross-centric classification performance based on standard and digital biomarkers (training on Dataset 1, testing on Dataset 2). Digital biomarkers presented higher accuracy (left, p = 0.0001) and higher area under the ROC curve (right), while standard biomarkers presented slightly above chance performances. D Cross-centric classification performance based on standard and digital biomarkers (training on Dataset 2, testing on Dataset 1). Also in this case, digital biomarkers presented higher accuracy (left, p = 0.031) and a slightly higher area under the ROC curve (right). Notation: error bars represent standard deviation values, while bar height represents mean value (100 bootstrap iterations). Both mean and standard deviation were computed after 100 bootstrap iterations for all panels. Significance notation: * stands for p < 0.05; ** stands for p < 0.01, two-sided McNemar’s test.
We finally tested the performance of both standard and digital biomarkers in supporting the cross-classification of patients (see Supplementary Materials, “Machine learning pipeline (classification and cross-classification” section).
For cross-classifications from Dataset 1 to Dataset 2, digital biomarkers achieved higher accuracy when compared with standard biomarkers (0.78 ± 0.03 vs 0.56 ± 0.04) and AUC (0.74 vs 0.53, Fig. 3C). Superior performances were confirmed by McNemar’s test (p = 0.0001). Similarly, for cross-classifications from Dataset 2 to Dataset 1, digital biomarkers demonstrated superior performance (p = 0.031, McNemar’s test), with higher accuracy (0.77 ± 0.03 vs 0.65 ± 0.06) and AUC (0.76 vs 0.70, Fig. 3D).
Discussion
Overall, digital biomarkers demonstrated strong cross-classification capabilities, in contrast to the poor performance observed in standard biomarkers. These findings mark a significant advance toward integrating non-invasive recordings and computational methods into clinical practice. Additionally, by reconstructing the structural mechanisms underlying various functional and dynamic alterations, digital biomarkers exhibited great generalizability across recording types.
The DADD model replicates neural activity at various levels of neurodegeneration by incorporating parameters that describe multiscale pathological mechanisms related to dementia progression. Namely, the synaptic degeneration parameters capture hyperexcitation effects due to early amyloid-beta deposition36. The connectivity degeneration parameter portrays white matter atrophy due to axonal damage34,37. The neuroplasticity parameter captures the connective rewiring operated by the brain to cope with connectivity insults38. These parameters were able to distinguish between healthy and MCI patients in both datasets with consistent performance, outperforming both MEG and EEG and mapping alterations observed in both recording techniques to a common framework.
Reproducibility is a pivotal requirement in clinical studies15, and digital biomarkers allowed to obtain consistent results using the same classification techniques on different datasets adopting different inclusion criteria, recording modalities, pre-processing and post-processing techniques. Such reproducibility resulted to be impossible for standard biomarkers, which achieved variable results on the two different datasets with overall low performance, coherently with multi-centric studies conducted with similar methodology39.
As expected, MEG data provided better classification performance compared to EEG. However, when the classification was performed using digital biomarkers, this difference between datasets faded. This suggests that the extraction of digital biomarkers might also be used to “clean” noisier data (as in EEG), bringing the level of classification to a level comparable to the gold-standard (MEG). This might have major practical implications given the relatively low-cost and wide-spread availability of EEG, as opposed to MEG which is typically only available in large research facilities.
Moreover, while no standard biomarker correlated with patients’ cognitive status, digital biomarkers predicted corrected MMSE scores with high significance, linking synaptic degeneration to cognitive decline severity. Notably, in Dataset 1, the connectivity degeneration parameter (cp) was found to anti-correlate with corrected MMSE. Patients in this cohort are MCI of no specific type. The cp parameter captures the disconnection effects, which are widely documented to correlate with cognitive decline due to age or disease34,40. On the contrary, in Dataset 2 the synaptic degeneration was found to strongly anti-correlate with corrected MMSE. Patients in Dataset 2 are amnestic MCI, known to present higher degeneration of temporal lobes related to amyloid deposition when compared to non-amnestic MCI41. Crucially, the synaptic degeneration parameter selectively alters synaptic transmission in prefrontal and temporal lobes (see Methods, “DADD Model and Simulated Signals” paragraph). Hence, digital biomarkers not only correlate with cognitive status, but also offer interpretable prediction on the macro- and micro-structural alterations leading to cognitive decline in MCI patients.
We have already attempted a CTR-MCI classification on patients of Dataset 1 in a previous work18 by using standard biomarkers, obtaining better results compared with the ones discussed here. The EEG biomarkers used for classification in that study were selected specifically for Dataset 1 through a tailored feature selection process. However, the need for common electrophysiological metrics to characterize both datasets led to a significant drop in classification performance within this dataset, once again highlighting a lack of generalizability of standard biomarkers.
This limitation was evident in both intra- and inter-dataset classifications. In the intra-dataset classification, Dataset 2 showed higher mean accuracy (0.68) compared to Dataset 1 (0.58), possibly due to the difference in classification power between EEG (Dataset 1) and MEG (Dataset 2), as mentioned above. For inter-dataset classification, transferring the classification from Dataset 2 to Dataset 1 achieved greater mean accuracy (0.65) than the reverse (0.56 for Dataset 1 → 2). In contrast, digital biomarkers delivered more consistent results, with similar performances in intra-dataset classification mean accuracy (0.83 for Dataset 1, 0.75 for Dataset 2) and inter-dataset classification mean accuracy (0.78 for Dataset 1 → 2 and 0.77 for Dataset 2 → 1).
The use of computational models in personalized medicine is an emerging field, and their application for diagnostic and prognostic purposes remains relatively unexplored—particularly in fields like dementia. One notable exception is the study by Triebkorn et al.6, where a computational brain model was employed to improve classification between healthy controls, individuals with mild cognitive impairment (MCI), and Alzheimer’s disease patients leading to a 0.10 increase in F1-score.
Another promising avenue for patient classification across the cognitive decline continuum involves the use of deep neural networks. Several studies have leveraged this approach, notably Fouladi et al.11 reported a 92% accuracy in classifying healthy individuals versus those with MCI or AD using EEG data and a convolutional neural network. Sibilano et al.10 tackled earlier stages, reporting a 0.76 F1-score in distinguishing SCD from MCI, and 0.54 in a multiclass classification task involving healthy, SCD, and MCI individuals.
While deep learning approaches can achieve high accuracy, they often lack interpretability and require large datasets—an obstacle in clinical contexts where extensive data collection is not always feasible. Moreover, cross-center classification has rarely been attempted using these techniques, with the notable exemption of Watanabe et al.39, that however focused on classifying different dementia forms with varying results (AUC ranging from 0.508 to 0.920).
No study has ever attempted the cross-generalization of biomarkers derived from non-invasive recordings or computational models in classifying cognitive decline patients. The recent work by Watanabe et al.39 identified biomarkers that could discriminate between different dementia types from electroencephalographic recordings, enrolling patients from three different centers. However, the model was trained on overt cases of dementia and was not generalized with a cross-classification.
It is worth noting that this study presents several limitations. While our model effectively captures key neurophysiological mechanisms associated with cognitive decline progression, it is important to acknowledge that cognitive decline is a multifaceted condition influenced by numerous biological, psychological, and environmental factors—many of which cannot be fully represented within a computational framework. Our model currently focuses on capturing early neurodegeneration processes underlying pre-clinical or early clinical condition like subjective cognitive decline and MCI. Further iterations of the model should aim at adapting to a greater array of conditions, including overt dementia states like Alzheimer’s disease. Additionally, our study was designed to assess the robustness of model-derived parameters across two independent cohorts acquired from different medical facilities. These cohorts differed not only in recording modality (EEG vs. MEG), but also in preprocessing pipelines, acquisition protocols, and recruitment criteria. This allowed us to test the capability of the model to generalize across datasets presenting a vast array of differences. While the model parameters consistently outperformed standard biomarkers, which showed marked variability between cohorts, the numerous differences between centers introduce potential confounding factors, complicating the interpretation of why certain standard metrics failed to generalize. Importantly, although our work leverages two distinct datasets, the term “cross-centric” in this context refers to the application of the model across different modalities and sites, rather than to multiple datasets using the same modality across different centers. In the latter frame, a multicentric design would require using the same approach to, for example, two EEG and two MEG datasets independently recorded at different sites.
Ensuring that candidate biomarkers can support an accurate cross-classification is crucial for the clinical application of computational and machine learning models. Digital biomarkers provided by our model demonstrated high cross-classification accuracy, providing robust and interpretable classification. The fact that the two datasets employed different recruitment criteria and multimodal recordings techniques, underlies once more the potential role of digital biomarkers in realistic clinical settings. Our work paves the way for the translation of non-invasive recordings and computational models into clinical practice.
Data availability
Data reported in Fig. 2 can be found in.csv format in Supplementary Data 1–4. Anonymized demographic data (including MMSE and Age) can be provided by alberto.mazzoni@santannapisa.it (Dataset 1, EEG) and pierpaolo.sorrentino@univ-amu.fr (Dataset 2, MEG) upon reasonable request, from a qualified academic investigator.
Code availability
Code for the digital twin simulation and digital biomarkers derivation can be found in https://github.com/LoreAma/Code_AD_simulations/. Statistical Analysis was implemented in Python using standard commercially available packages such as Numpy, Scipy and Scikit-Learn.
References
Rasmussen, J. & Langerman, H. Alzheimer’s disease—why we need early diagnosis. Degener. Neurol. Neuromuscul. Dis. 9, 123–130 (2019).
Horvath, A. et al. EEG and ERP biomarkers of Alzheimer’s disease: a critical review. Front. Biosci. Landmark Ed. 23, 183–220 (2018).
Gouw, A. A. Clinical applications of EEG/MEG in AD: diagnosis, prognosis and treatment monitoring. Alzheimers Dement 19, e073238 (2023).
Meghdadi, A. H. et al. EEG and ERP biosignatures of mild cognitive impairment for longitudinal monitoring of early cognitive decline in Alzheimer’s disease. PLoS One 19, e0308137 (2024).
Stefanovski, L. et al. Linking molecular pathways and large-scale computational modeling to assess candidate disease mechanisms and pharmacodynamics in Alzheimer’s disease. Front. Comput. Neurosci. 13, 54 (2019).
Triebkorn, P. et al. Brain simulation augments machine-learning–based classification of dementia. Alzheimers Dement. Transl. Res. Clin. Interv. 8, e12303 (2022).
Van Nifterick, A. M. et al. A multiscale brain network model links Alzheimer’s disease-mediated neuronal hyperactivity to large-scale oscillatory slowing. Alzheimers Res. Ther. 14, 101 (2022).
Martínez-Cañada, P. et al. Combining aperiodic 1/f slopes and brain simulation: An EEG/MEG proxy marker of excitation/inhibition imbalance in Alzheimer’s disease. Alzheimers Dement. Diagn. Assess. Dis. Monit. 15, e12477 (2023).
Cabrera-Álvarez, J. et al. A Multiscale Closed-Loop Neurotoxicity Model of Alzheimer's Disease Progression Explains Functional Connectivity Alterations. eNeuro 11, ENEURO.0345-23.2023 (2024).
Sibilano, E. et al. An attention-based deep learning approach for the classification of subjective cognitive decline and mild cognitive impairment using resting-state EEG. J. Neural Eng. 20, 016048 (2023).
Fouladi, S., Safaei, A. A., Mammone, N., Ghaderi, F. & Ebadi, M. J. Efficient deep neural networks for classification of Alzheimer’s disease and mild cognitive impairment from Scalp EEG recordings. Cogn. Comput. 14, 1247–1268 (2022).
Kim, N. H., Yang, D. W., Choi, S. H. & Kang, S. W. Machine learning to predict brain amyloid pathology in pre-dementia Alzheimer’s disease using QEEG features and genetic algorithm heuristic. Front. Comput. Neurosci. 15, 755499 (2021).
Martin, S. A., Townend, F. J., Barkhof, F. & Cole, J. H. Interpretable machine learning for dementia: a systematic review. Alzheimers Dement 19, 2135–2149 (2023).
Koenig, T., Smailovic, U. & Jelic, V. Past, present and future EEG in the clinical workup of dementias. Psychiatry Res. Neuroimaging 306, 111182 (2020).
Munafò, M. R. et al. A manifesto for reproducible science. Nat. Hum. Behav. 1, 1–9 (2017).
Cassani, R., Estarellas, M., San-Martin, R., Fraga, F. J. & Falk, T. H. Systematic review on resting-state EEG for Alzheimer’s disease diagnosis and progression assessment. Dis. Markers 2018, 5174815 (2018).
D’Angelo, E. & Jirsa, V. The quest for multiscale brain modeling. Trends Neurosci 45, 777–790 (2022).
Amato, L. G. et al. Personalized modeling of Alzheimer’s disease progression estimates neurodegeneration severity from EEG recordings. Alzheimers Dement. Diagn. Assess. Dis. Monit. 16, e12526 (2024).
Wang, H. E. et al. Delineating epileptogenic networks using brain imaging data and personalized modeling in drug-resistant epilepsy. Sci. Transl. Med. 15, eabp8982 (2023).
Amato, L. G. et al. Digital twins and non-invasive recordings enable early diagnosis of Alzheimer’s disease. Alzheimers Res. Ther. 17, 125 (2025).
Mazzeo, S. et al. PRedicting the EVolution of SubjectIvE Cognitive Decline to Alzheimer’s disease with machine learning: the PREVIEW study protocol. BMC Neurol 23, 300 (2023).
Albert, M. S. et al. The diagnosis of mild cognitive impairment due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7, 270–279 (2011).
Delorme, A. & Makeig, S. EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J. Neurosci. Methods 134, 9–21 (2004).
Bigdely-Shamlo, N., Mullen, T., Kothe, C., Su, K.-M. & Robbins, K. A. The PREP pipeline: standardized preprocessing for large-scale EEG analysis. Front. Neuroinformatics 9, 16 (2015).
Liparoti, M. et al. Functional brain network topology across the menstrual cycle is estradiol dependent and correlates with individual well-being. J. Neurosci. Res. 99, 2271–2286 (2021).
Nolte, G. The magnetic lead field theorem in the quasi-static approximation and its use for magnetoencephalography forward calculation in realistic volume conductors. Phys. Med. Biol. 48, 3637 (2003).
Gong, G. et al. Mapping anatomical connectivity patterns of human cerebral cortex using in vivo diffusion tensor imaging tractography. Cereb. Cortex 19, 524–536 (2009).
Bastos, A. M. & Schoffelen, J.-M. A tutorial review of functional connectivity analysis methods and their interpretational pitfalls. Front. Syst. Neurosci. 9, 175 (2016).
Sanz Leon, P. et al. The virtual brain: a simulator of primate brain network dynamics. Front. Neuroinformatics 7, 10 (2013).
Jansen, B. H. & Rit, V. G. Electroencephalogram and visual evoked potential generation in a mathematical model of coupled cortical columns. Biol. Cybern. 73, 357–366 (1995).
Sanz-Leon, P., Knock, S. A., Spiegler, A. & Jirsa, V. K. Mathematical framework for large-scale brain network modeling in The Virtual Brain. Neuroimage 111, 385–430 (2015).
Ren, S.-Q. et al. Amyloid β causes excitation/inhibition imbalance through dopamine receptor 1-dependent disruption of fast-spiking GABAergic input in anterior cingulate cortex. Sci. Rep. 8, 302 (2018).
Braak, H. & Braak, E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. (Berl.) 82, 239–259 (1991).
Ohlhauser, L., Parker, A. F., Smart, C. M. & Gawryluk, J. R. White matter and its relationship with cognition in subjective cognitive decline. Alzheimers Dement. Diagn. Assess. Dis. Monit. 11, 28–35 (2019).
Vance, D. E., Roberson, A. J., McGuinness, T. M. & Fazeli, P. L. How neuroplasticity and cognitive reserve: protect cognitive functioning. J. Psychosoc. Nurs. Ment. Health Serv. 48, 23–30 (2010).
Maestú, F., de Haan, W., Busche, M. A. & DeFelipe, J. Neuronal excitation/inhibition imbalance: core element of a translational perspective on Alzheimer pathophysiology. Ageing Res. Rev. 69, 101372 (2021).
Pietroboni, A. M. et al. The role of Amyloid-β in white matter damage: possible common pathogenetic mechanisms in neurodegenerative and demyelinating diseases. J. Alzheimers Dis. 78, 13–22 (2020).
Stam, C. J. Hub overload and failure as a final common pathway in neurological brain network disorders. Netw. Neurosci. 8, 1–23 (2024).
Watanabe, Y. et al. A deep learning model for the detection of various dementia and MCI pathologies based on resting-state electroencephalography data: a retrospective multicentre study. Neural Netw 171, 242–250 (2024).
O’Sullivan, M. et al. Evidence for cortical “disconnection” as a mechanism of age-related cognitive decline. Neurology 57, 632–638 (2001).
Csukly, G. et al. The differentiation of amnestic type MCI from the non-amnestic types by structural MRI. Front. Aging Neurosci. 8, 52 (2016).
Acknowledgements
L.G.A., M.L., and A.M. were supported and partially funded by #NEXTGENERATIONEU (NGEU) and funded by the Ministry of University and Research (MUR), National Recovery and Resilience Plan (NRRP), project EBRAINS-Italy (IR0000011) - European Brain ReseArch INfrastructureS-Italy (DN. 101 16.06.2022), project MNESYS (PE0000006) – A Multiscale integrated approach to the study of the nervous system in health and disease (DN. 1553 11.10.2022). R.M., G.S., E.T.L., P.S. were supported and funded by Ministero Sviluppo Economico; Contratto di sviluppo industriale “Farmaceutica e Diagnostica” (CDS 000606); European Union “NextGenerationEU,” (Investimento 3.1.M4. C2) of PNRR. L.G.A., M.L., V.M., G.G., A.G., V.B. and A.M. were partially funded by Tuscany Region - PRedicting the EVolution of SubjectIvE Cognitive Decline to Alzheimer’s Disease With machine learning – PREVIEW CUP.D18D20001300002.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Stavros Dimitriadis and Seyed-Mahdi Khaligh-Razavi for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Amato, L.G., Minino, R., Lassi, M. et al. Digital twins support cross-modal and cross-centric classification of mild cognitive impairment. Commun Med 6, 30 (2026). https://doi.org/10.1038/s43856-025-01281-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01281-z